Introduction
Cluster of differentiation (CD)44 is a transmembrane
glycoprotein that functions as an adhesion molecule for
extracellular matrices, primarily hyaluronan (1,2). The
expression of CD44 in cancer cells has also been implicated in cell
migration, invasion and metastasis (1,2).
Furthermore, CD44 has recently been recognized as a major marker
for cancer stem cells (CSCs) in several types of cancer (2,3).
The CD44 gene consists of 20 exons, 10 of which
(exons 1–5 and 16–20) encode the shortest and ubiquitously
expressed standard isoform of CD44 (CD44s) (1,2). The other
10 exons (exons 6–15, also referred to as exons v1-v10) are variant
exons inserted by alternative splicing in various combinations,
thereby generating various variant isoforms of CD44 (CD44v)
(1,2).
The insertion of variant exons lengthens the extracellular domain
and alters the binding and signaling properties of CD44. The
expression of CD44v is restricted to a few epithelial tissues,
including several types of cancer (2). Although extensive research has been
conducted, the association between the expression of CD44v and
tumor aggressiveness remains controversial (1,2,4–6).
CD44v8-10, also known as CD44R1 and CD44E, is one of
the isoforms of CD44v, and contains the variant exons 13–15
(v8-v10) (1,2). Clinical studies have demonstrated that
CD44v8-10 is expressed in various human epithelial cancers,
including breast, lung, colon, bladder and gastric cancer, and its
expression is correlated with metastasis (7–10).
Furthermore, recent studies suggest that the expression of
CD44v8-10 is associated with CSC-like phenotypes (11,12).
The bone is one of the most frequent organs to be
affected by metastatic cancer (13).
Bone metastases cause devastating bone pain and skeletal-related
events, including pathological fractures, spinal compression and
hypercalcemia, thereby severely deteriorating the quality of life
of patients (13). However, the
mechanisms underlying bone metastases have not yet been fully
elucidated.
Clinical studies have suggested that a positive
correlation exists between the expression of CD44 and bone
metastasis (14,15). Furthermore, our recent preclinical
study demonstrated that CD44, particularly CD44s, confers CSC-like
properties to cancer cells and enhances their metastatic potential
to bone (16). However, the roles of
CD44v in the development of bone metastases have yet to be
explored. Thus, the present study examined the differential roles
of CD44s and CD44v, particularly CD44v8-10, in bone metastases
using a well-characterized animal model.
Materials and methods
Cell cultures
The human breast cancer cell lines MDA-MB-231 and
MCF-7 were obtained from the American Type Culture Collection
(Manassas, VA, USA). The human lung cancer cell line A549 was
obtained from the Health Science Research Resources Bank (Osaka,
Japan). Cells were cultured in Dulbecco's modified Eagle's medium
(DMEM; Sigma-Aldrich; Merck Millipore, Darmstadt, Germany)
supplemented with 10% fetal bovine serum (FBS; Thermo Fisher
Scientific, Inc., Waltham, MA, USA) and 100 µg/ml kanamycin sulfate
(Meiji Seika Kaisha, Ltd., Tokyo, Japan), and were maintained at
37°C in a humidified atmosphere of 5% CO2 in air.
Overexpression of ESRP1 and
CD44v8-10
The complementary (c)DNA of human epithelial
splicing regulatory protein 1 (ESRP1) in the pOTB7 vector was
purchased from Dharmacon (GE Healthcare Bio-Sciences, Pittsburgh,
PA, USA). Human CD44v8-10 cDNA was reverse transcribed and
amplified from total RNA isolated from MDA-MB-231 cells
overexpressing the ESRP1 gene. Total RNA was isolated using a High
Pure RNA Isolation kit (Roche Diagnostics, Basel, Switzerland).
cDNA was synthesized from 5 µg of total RNA using PrimeScript
Reverse Transcriptase (Takara Bio, Inc., Shiga, Japan). The primer
sequences for CD44v8-10 were 5′-CGGACACCATGGACAAGTTT-3′ (forward)
and 5′-TCCAACGGTTGTTTCTTTCC-3′ (reverse). PCR was performed using
PrimeSTAR GXL DNA polymerase (Takara Bio, Inc.) with a TaKaRa PCR
Thermal Cycler Dice (Takara Bio, Inc.) for 30 cycles (98°C for 10
sec, 60°C for 15 sec and 68°C for 90 sec). Both cDNAs were
subcloned into the pLVSIN-CMV Pur vector (Takara Bio, Inc.) and
transduced into cells using a Lenti-X Lentiviral Expression System
(Takara Bio, Inc.). As a control, empty vector (EV) was transduced
into the cells. Colonies resistant to puromycin (0.25–1.00 µg/ml;
Sigma-Aldrich; Merck Millipore) were isolated and cloned.
Reverse transcription-polymerase chain
reaction (RT-PCR)
Total cellular RNA was isolated using a High Pure
RNA Isolation kit. cDNA was synthesized from 5 µg of total RNA
using PrimeScript Reverse Transcriptase. Two sets of PCR primers
were used to detect CD44 (8). One set
(primer set 1, forward 5′-TCCCAGACGAAGACAGTCCCTGGAT-3′ and reverse
5′-CACTGGGGTGGAATGTGTCTTGGTC-3′) detected all the isoforms of CD44,
while the other set (primer set 2, forward
5′-GACAGAATCCCTGCTACCAATA-3′ and reverse
5′-ATGTGTCTTGGTCTCCTGATAA-3′) specifically detected CD44v8-10 and
CD44v10. The primer sequences for glyceraldehyde-3-phosphate
dehydrogenase (GAPDH) were 5′-CATGGAGAAGGCTGGGGCTC-3′ (forward) and
5′-CACTGACACGTTGGCAGTGG-3′ (reverse). PCR was conducted using PCR
Master Mix (Promega Corporation, Madison, WI, USA) with a TaKaRa
PCR Thermal Cycler Dice for 25 cycles (at 95, 55 and 72°C for 30
sec), or for 20 cycles in the case of GAPDH. The PCR products were
separated on 2% agarose gels containing ethidium bromide and
visualized under UV light. The sizes of the fragments were
confirmed by comparison with a 100-bp DNA ladder (Takara Bio,
Inc.).
Flow cytometry
Cells were harvested by trypsinization and incubated
with a phycoerythrin (PE)-conjugated anti-panCD44 antibody (1:100
dilution; 12-0441-81; eBioscience, Inc., San Diego, CA, USA) for 40
min at 4°C. Flow cytometric analysis was performed using a flow
cytometer (Cytomics FC 500; Beckman Coulter, Inc., Brea, CA, USA).
Data were analyzed using FlowJo software (version 7.6.5; FlowJo,
LLC, Ashland, OR, USA).
Western blotting
Total protein lysates were prepared in lysis buffer
[20 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (pH 7.4),
150 mM NaCl, 1 mM ethylene glycol-bis(β-aminoethyl
ether)-N,N,N',N'-tetraacetic acid, 1.5 mM MgCl2, 10%
glycerol, 1% Triton X-100, 10 µg/ml aprotinin, 10 µg/ml leupeptin,
1 mM phenylmethylsulfonyl fluoride and 0.2 mM sodium
orthovanadate]. Equal amounts of proteins were separated on 7.5%
sodium dodecyl sulfate-polyacrylamide gel electrophoresis and
transferred onto nitrocellulose membranes (Bio-Rad Laboratories,
Inc., Hercules, CA, USA). The membranes were then immunoblotted
with the following antibodies: Anti-panCD44 [1:500 dilution
(17)], anti-CD44v9 (1:500 dilution;
LKG-M001; Cosmo Bio, Tokyo, Japan), anti-ESRP1 (1:500 dilution;
HPA023720; Atlas Antibodies AB, Stockholm, Sweden) and anti-β-actin
(1:1,000 dilution; A00702; GenScript, Tokyo, Japan), and visualized
with a horseradish peroxidase-conjugated anti-mouse immunoglobulin
(Ig)G (1:10,000 dilution; 715-035-150; Jackson ImmunoResearch
Laboratories, Inc., West Grove, PA, USA) or anti-rabbit IgG
antibody (1:10,000 dilution; 711-035-152; Jackson ImmunoResearch
Laboratories, Inc.), followed by enhanced chemiluminescence (ECL).
ECL solution was made by dissolving 0.2 mM p-coumaric acid
(Sigma-Aldrich; Merck Millipore), 2.5 mM luminol (Sigma-Aldrich;
Merck Millipore) and 0.02% hydrogen peroxide (Wako Pure Chemical
Industries, Ltd., Osaka, Japan) in 100 mM Tris-HCl (pH 8.5). The
expression of β-actin was used as a loading control.
Cell proliferation in monolayer
cultures
Cells (1,000 cells/well) were plated in a 96-well
plate in growth medium and cultured for 72 h. Subsequently, cell
proliferation was determined using Cell Proliferation Reagent WST-1
(Roche Diagnostics). Absorbance was measured using iMark Microplate
Absorbance Reader (Bio-Rad Laboratories, Inc.) at a wavelength of
450 nm.
Wound healing assay
Cells (1×105 cells/well) were plated in a
24-well plate in growth medium and incubated for 24 h. Upon
confirming the formation of a complete monolayer, cells were
wounded by scratching lines with a standard 200-µl plastic pipette
tip. Cell migration into the wound area was observed with a
phase-contrast microscope after 24 or 48 h. The percentage of the
filled wound area was calculated as follows: Filled wound area (%)
= (original wound area-remaining wound area)/original wound area ×
100.
Cell invasion assay
Cell invasion assays were conducted using 24-well
Corning BioCoat Matrigel Invasion Chambers (BD Biosciences,
Franklin Lakes, NJ, USA). In brief, cells were seeded into the
upper inserts (2.5×104 cells/insert) in DMEM
supplemented with 1% FBS. The outer wells were filled with DMEM
containing 10% FBS as a chemoattractant. After 24-h incubation, the
membranes were stained with hematoxylin and mounted on slides.
Invading cells were counted in five randomly selected microscopic
fields. Data are expressed as the number of invaded
cells/field.
Tumor sphere formation
Cells were plated in ultra-low attachment 24-well
plates (Corning, Inc., Corning, NY, USA) at a density of 1,000
cells/well and cultured in DMEM/F12 (Sigma-Aldrich; Merck
Millipore) supplemented with 2% MACS Supplement B27 PLUS (Miltenyi
Biotec GmbH, Bergisch Gladbach, Germany), 40 ng/ml recombinant
human fibroblast growth factor-2 (ProSpec, East Brunswick, NJ, USA)
and 20 ng/ml recombinant human epidermal growth factor
(Sigma-Aldrich; Merck Millipore). The cells were replenished with
fresh medium every 3 days. After culturing for 6–12 days, the
number of tumor spheres with a diameter of >100 µm was counted
by light microscopy. Data are expressed as the number of tumor
spheres/well.
Animal experiments
Under anesthesia with pentobarbital (0.05 mg/g body
weight; Kyoritsu Seiyaku Corporation, Tokyo, Japan), MDA-MB-231
cell clones [1×105 cells/0.1 ml phosphate-buffered
saline (PBS)] were injected into the left cardiac ventricle of
athymic nude mice (KSN/Slc, female, 4-week-old, n=5/group; Japan
SLC, Inc., Shizuoka, Japan). All mice were sacrificed 4 weeks after
cell inoculation. All animal experiments were approved by the
Animal Management Committee of Matsumoto Dental University (Nagano,
Japan). Mice were housed at 28°C in a 12-h light:dark cycle and
were fed and watered ad libitum.
Histomorphometric and
immunohistochemical analyses
Histomorphometry
Paraffin sections were prepared by conventional
methods and were stained with hematoxylin and eosin.
Histomorphometric analysis of the tumor burden in bone was
conducted as described previously (16). Data are expressed as the tumor area
(mm2).
Immunohistochemistry
Paraffin sections were deparaffinized in xylene and
rehydrated in a graded series of ethanol. Endogenous peroxidase
activity was blocked by immersing the sections in 0.3%
H2O2 in methanol for 15 min. After blocking
with 3% bovine serum albumin in PBS for 30 min, sections were
incubated with primary antibodies overnight at 4°C. Primary
antibodies against panCD44 (17) and
CD44v9 (LKG-M001; Cosmo Bio) were applied at a dilution of 1:500.
Indirect immunohistochemical staining was performed using a
Histofine Simple Stain MAX PO kit (Nichirei Biosciences, Inc.,
Tokyo, Japan). Chromogen was developed using Liquid DAB+ (Dako,
Glostrup, Denmark). The slides were counterstained with
hematoxylin.
Statistical analysis
Data are expressed as the mean ± standard error of
the mean. Data were analyzed by analysis of variance followed by a
Tukey's test for determining differences between the groups, using
Mini StatMate software (version 1.1; ATMS, Tokyo, Japan). P<0.05
was considered to indicate a statistically significant
difference.
Results
Effects of ESRP1 overexpression on
bone metastases
ESRP1 is an epithelial-specific splicing factor that
regulates the alternative splicing of several genes, including CD44
(18–20). The overexpression of ESRP1 has been
reported to cause a splicing switch from CD44s to CD44v (18–20). In
order to examine the differential roles of CD44s and CD44v8-10 in
bone metastases, the current study employed an approach in which
the ESRP1 gene was stably transduced into MDA-MB-231 human breast
cancer cells and A549 human lung cancer cells (MDA/ESRP1 and
A549/ESRP1, respectively). While mock-transduced cells (MDA/EV and
A549/EV) predominantly expressed CD44s, the overexpression of ESRP1
reduced the expression of CD44s, but increased that of CD44v
(Fig. 1A). RT-PCR analysis confirmed
that the dominant isoform of CD44v expressed in MDA/ESRP1 and
A549/ESRP1 cells was CD44v8-10 (Fig.
1B). Flow cytometric analysis using an antibody that recognizes
panCD44 revealed that the total amount of CD44 expressed was rarely
affected by ESRP1 (Fig. 1C).
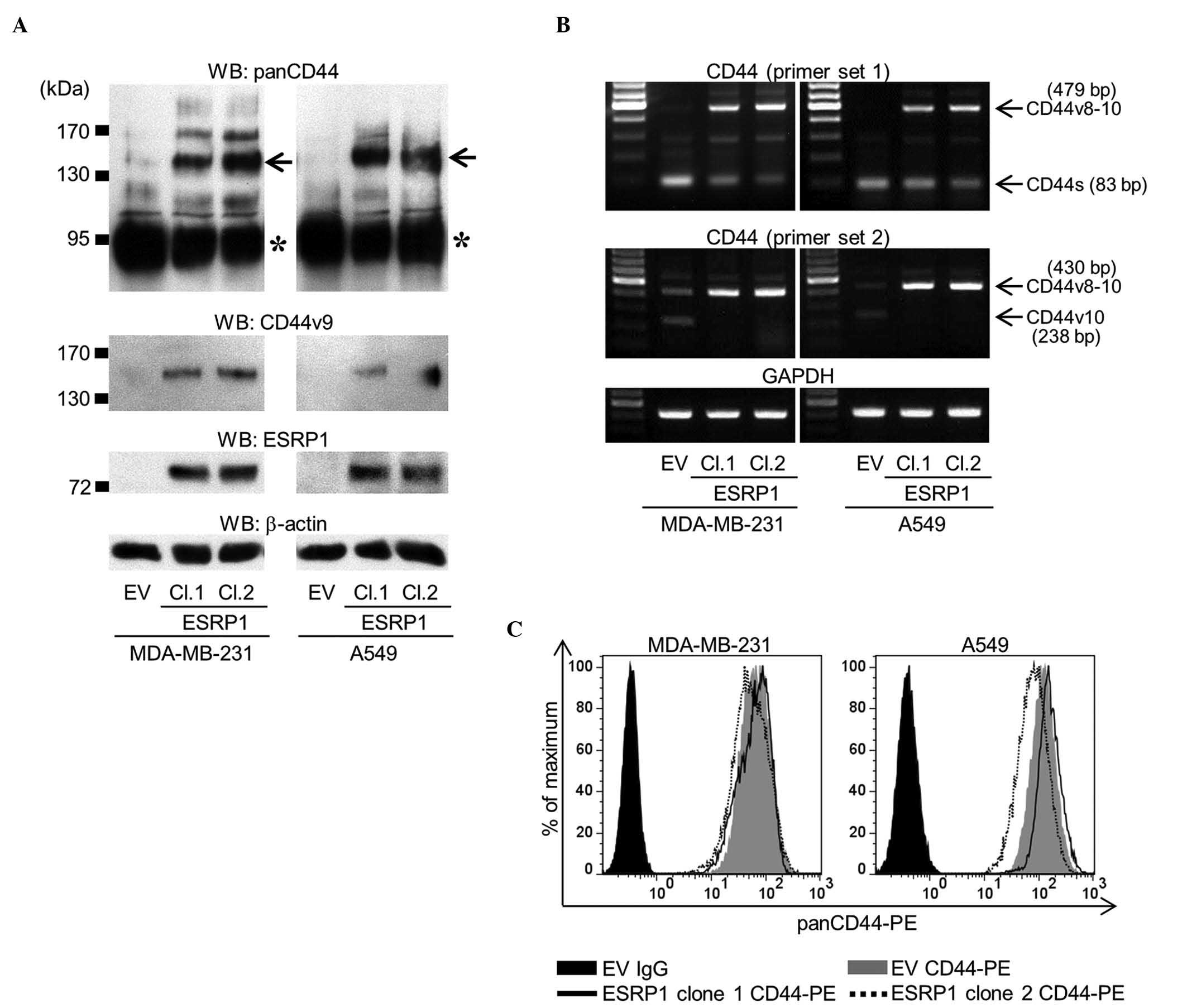 | Figure 1.Establishment of MDA-MB-231 and A549
cell clones overexpressing ESRP1. (A) The expression of panCD44,
CD44v9 and ESRP1 was determined by western blot analysis (asterisk,
CD44s; arrow, CD44v8-10). Two ESRP1-overexpressing clones (clones 1
and 2) of each cell line were examined in subsequent experiments,
and compared with empty vector mock-transduced cells. (B) Messenger
RNA expression of CD44 was determined by reverse
transcription-polymerase chain reaction using two sets of primers.
Primer set 1 detected all isoforms of CD44, while primer set 2
specifically detected CD44v8-10 and CD44v10. (C) Flow cytometric
analysis of the total amount of CD44 protein was determined using
an anti-panCD44 antibody. WB, western blot; CD, cluster of
differentiation; CD44v, variant isoform of CD44; ESRP1, epithelial
splicing regulatory protein 1; EV, empty vector; Cl, clone; CD44s,
standard isoform of CD44; GAPDH, glyceraldehyde-3-phosphate
dehydrogenase; PE, phycoerythrin; Ig, immunoglobulin. |
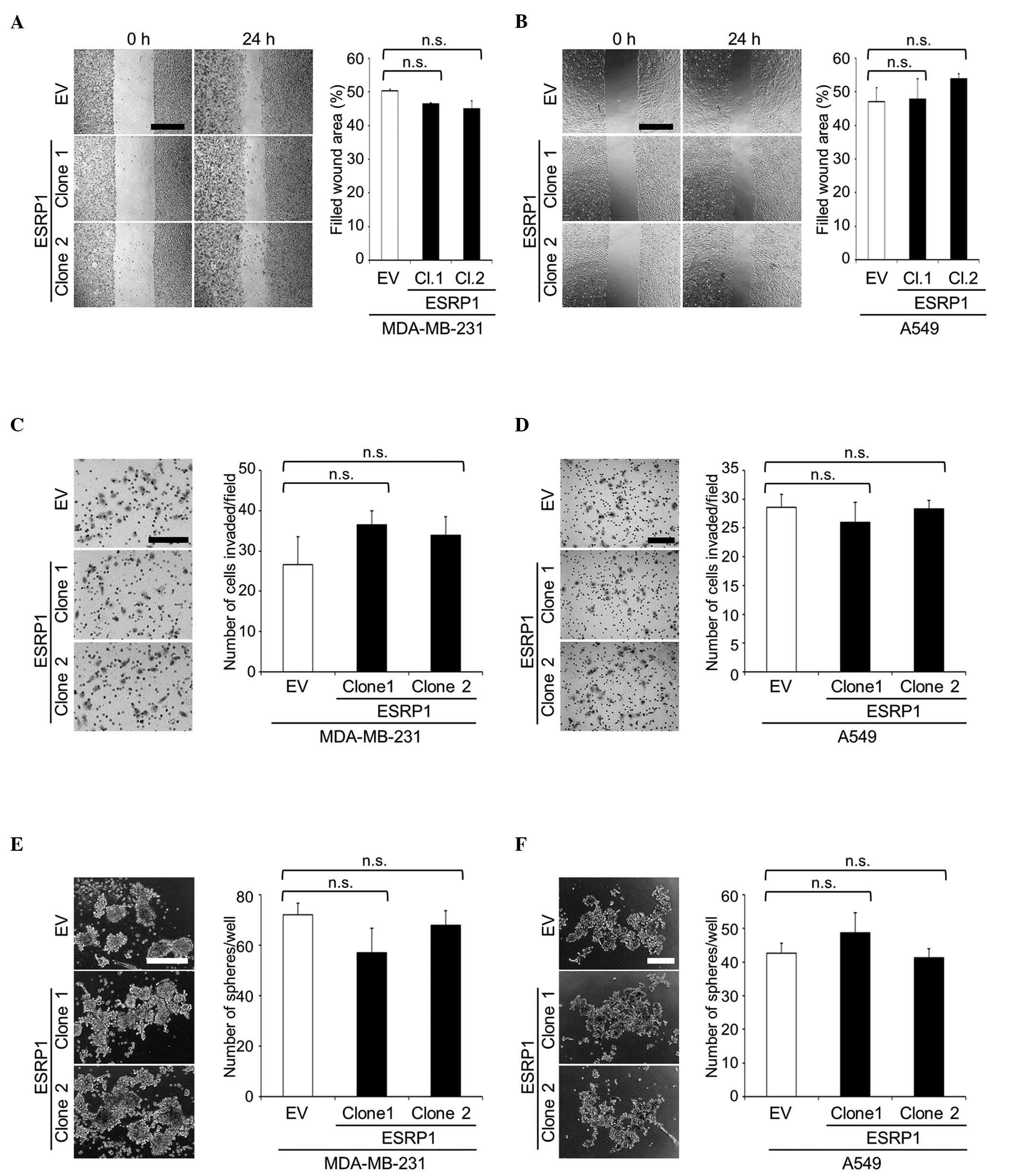
Using these clones, cellular functions were
evaluated by in vitro assays. WST-1 assay revealed no
significant differences in cell proliferation between mock- and
ESRP1-transduced cells (data not shown). In addition, ESRP1 did not
significantly affect cell migration or invasion, as determined in
wound healing and matrigel invasion assays, respectively (Fig. 2A-D). Since CD44 is a widely recognized
marker for CSCs (2,3), CSC-like properties were assessed by
sphere formation assay. The ability of ESRP1-overexpressing cells
to form tumor spheres in suspension cultures was similar to that of
the control cells (Fig. 2E and
F).
The bone-metastatic potential of MDA/ESRP1 cells
in vivo was examined using the intracardiac injection model.
In accordance with the data obtained in vitro, ESRP1 did not
cause significant differences in the development of bone metastases
(Fig. 3A and B), although the
expression of CD44v8-10 in MDA/ESRP1 cells was maintained at a high
level in bone (Fig. 3C).
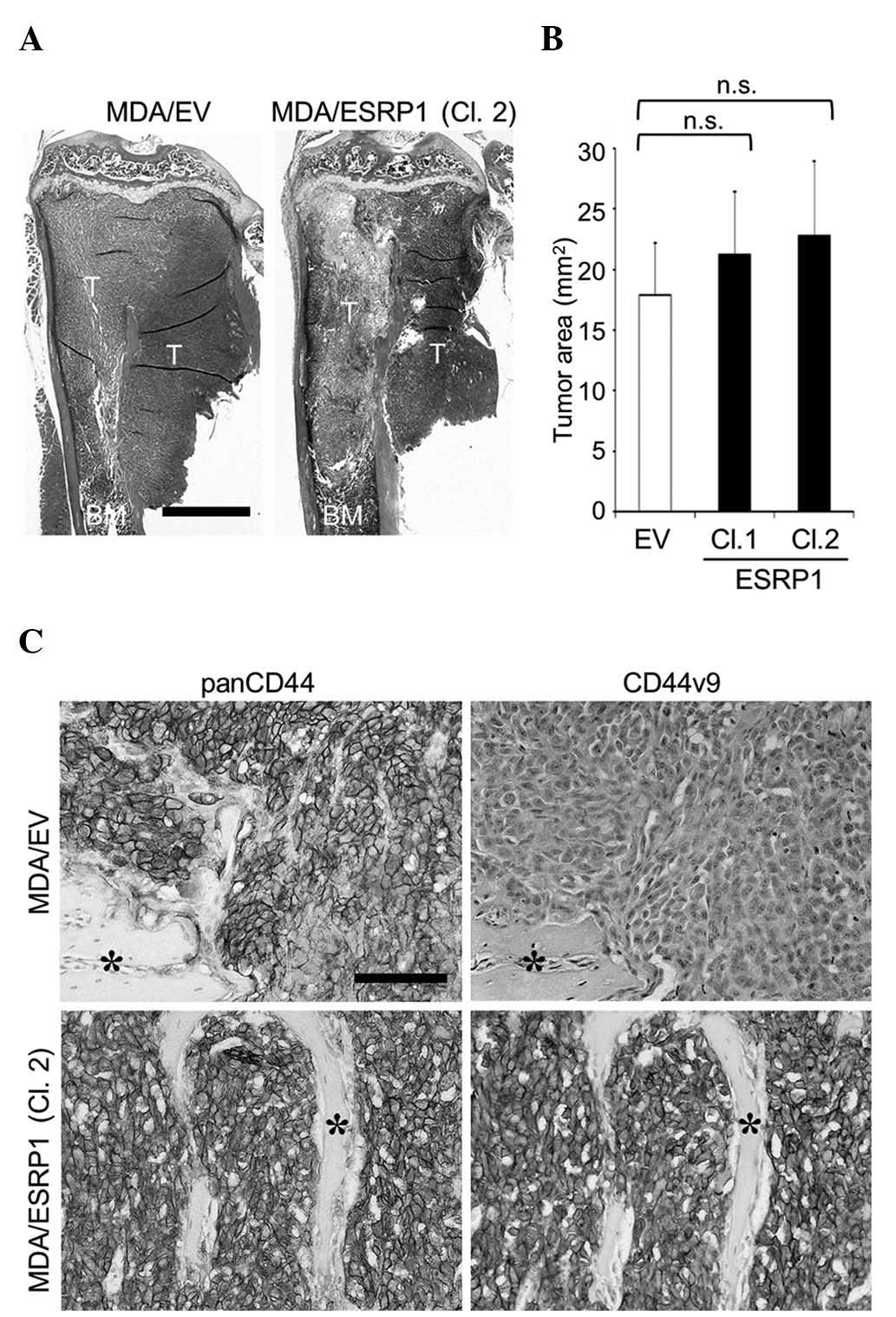 | Figure 3.Effects of ESRP1 overexpression on the
development of bone metastases of MDA-MB-231 cells. (A)
Representative histological pictures of bone metastases
(hematoxylin and eosin staining; scale bar, 1 mm). (B)
Histomorphometric analysis of the tumor burden in bone. Data are
expressed as tumor area (mm2; n=5/group). (C)
Immunohistochemical detection of the expression of panCD44 and
CD44v9 in cancer cells colonized in bone (asterisk, bone; scale
bar, 100 µm). CD, cluster of differentiation; CD44v, variant
isoform of CD44; ESRP1, epithelial splicing regulatory protein 1;
EV, empty vector; Cl, clone; T, tumor; BM, bone marrow; n.s., not
significant. |
Effects of CD44v8-10 overexpression on
bone metastases
To further examine the roles of CD44v8-10, an
alternative approach was undertaken in which the CD44v8-10 gene was
stably transduced into MDA-MB-231 cells to generate MDA/CD44v cells
(Fig. 4). MDA/CD44v cells exhibited
no significant changes in cell proliferation, migration, invasion
or tumor sphere formation in vitro (Fig. 5A, C and E), or in the development of
bone metastases in mice (Fig. 6).
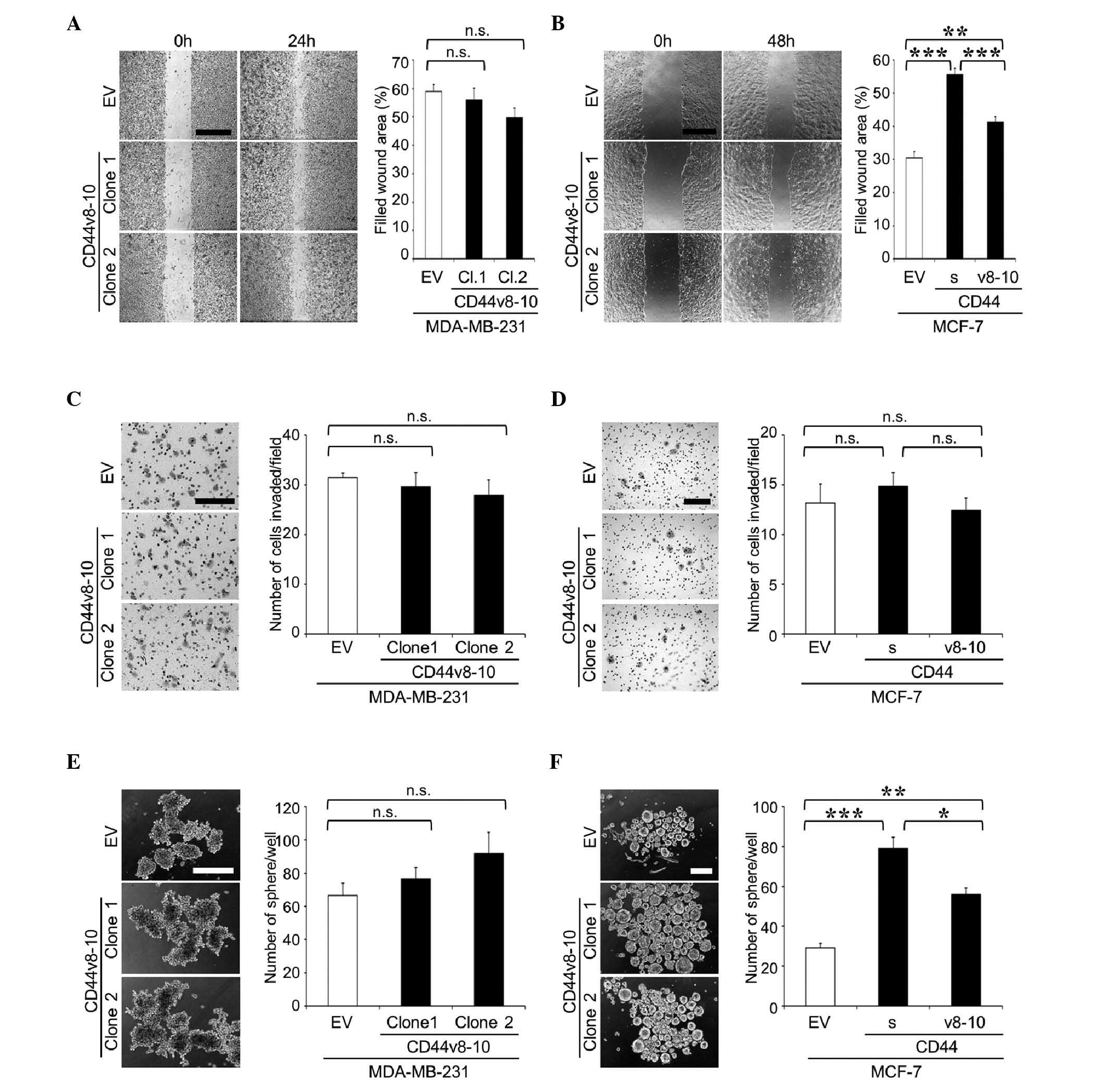 | Figure 5.Cell migration, cell invasion and
tumor sphere formation of MDA-MB-231 and MCF-7 cell clones
overexpressing CD44v8-10. (A and B) Cell migration of (A)
MDA-MB-231 and (B) MCF-7 clones was determined by wound healing
assay. Representative microscopic images at 0 and 24 h (MDA-MB-231)
or 48 h (MCF-7) after wounding are shown on the left side of the
panel (scale bar, 500 µm). Data are expressed as the percentage of
filled wound area. **P<0.01, ***P<0.001 vs. control. (C and
D) Cell invasion of (C) MDA-MB-231 and (D) MCF-7 clones was
determined by matrigel invasion assay. Representative microscopic
images of membranes are shown on the left side of the panel (scale
bar, 200 µm). Data are expressed as the number of cells
invaded/field. (E and F) Tumor sphere formation of (E) MDA-MB-231
and (F) MCF-7 clones in suspension cultures. Representative
microscopic images are shown on the left side of the panel (scale
bar, 200 µm). Data are expressed as the number of tumor
spheres/well. *P<0.05, **P<0.01, ***P<0.001 vs. control.
n.s., not significant; CD, cluster of differentiation; CD44v,
variant isoform of CD44; EV, empty vector; Cl, clone; CD44s,
standard isoform of CD44. |
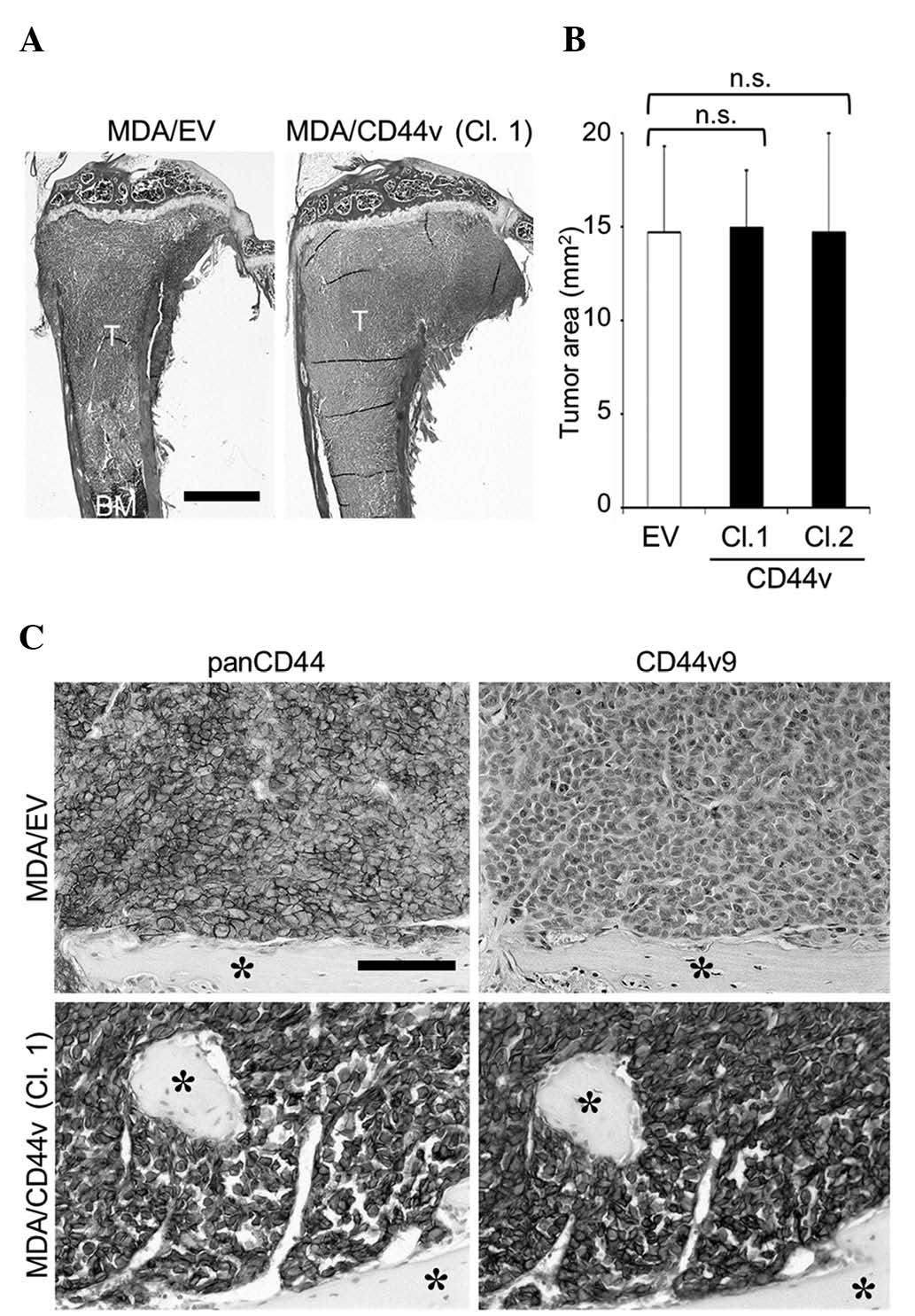 | Figure 6.Effects of CD44v8-10 overexpression on
the development of bone metastases of MDA-MB-231 cells. (A)
Representative histological pictures of bone metastases
(hematoxylin and eosin staining; scale bar, 1 mm). (B)
Histomorphometric analysis of the tumor burden in bone. Data are
expressed as tumor area (mm2; n=5/group). (C)
Immunohistochemical detection of panCD44 and CD44v9 expression in
cancer cells colonized in bone (asterisk, bone; scale bar, 100 µm).
CD, cluster of differentiation; CD44v, variant isoform of CD44; EV,
empty vector; Cl, clone; T, tumor; BM, bone marrow; n.s., not
significant. |
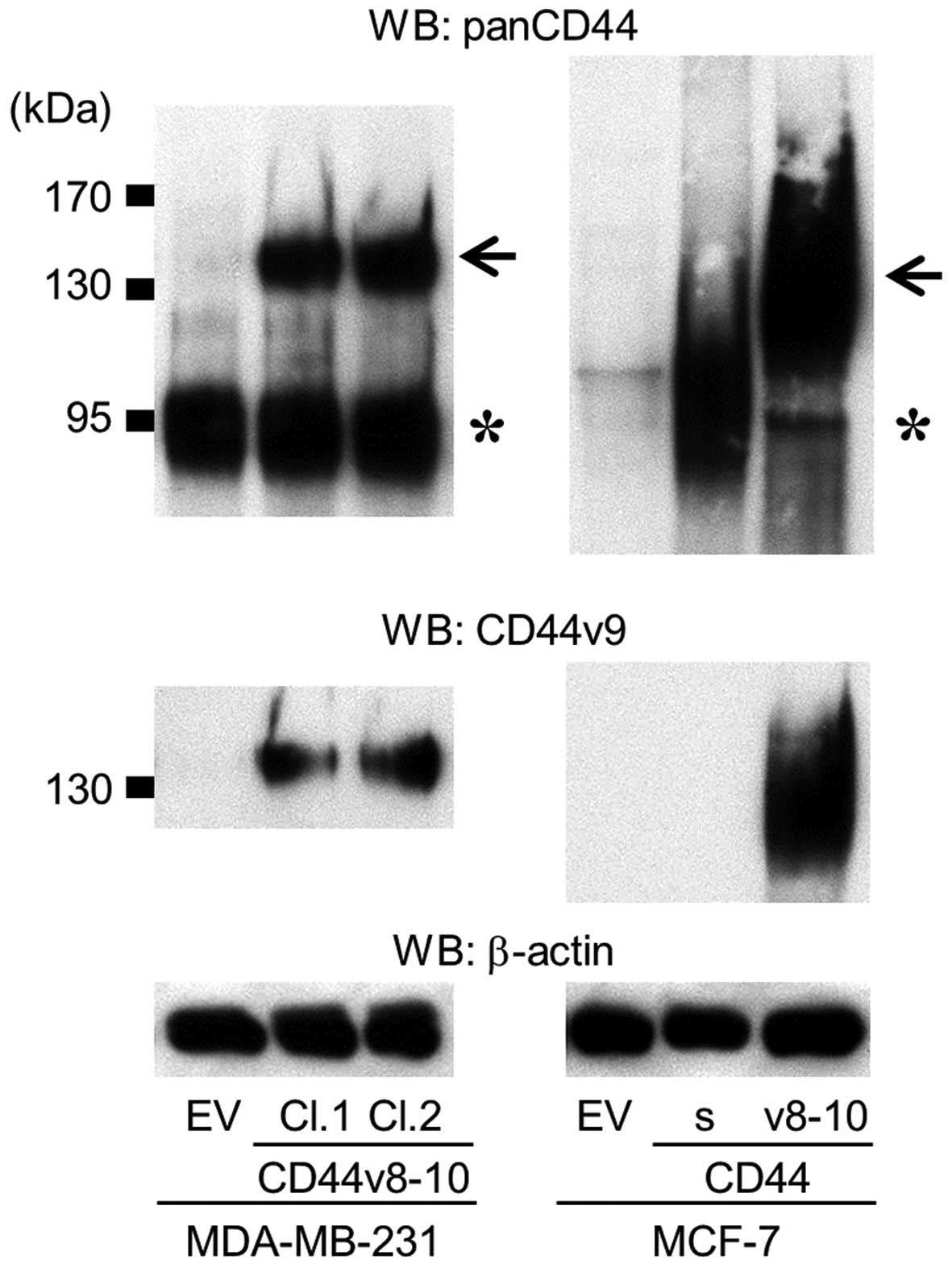
The effects of the overexpression of CD44v8-10 were
similarly tested in MCF-7 human breast cancer cells (MCF-7/CD44v;
Fig. 4). In contrast to MDA-MB-231
and A549 cells, MCF-7 cells expressed very low levels of any
isoform of CD44 (Fig. 4). As reported
previously (16), the overexpression
of CD44s promoted cell migration and sphere formation in MCF-7
cells in vitro (Fig. 5B and
F). MCF-7/CD44v also exhibited enhanced cell migration and
sphere formation compared with the control cells (Fig. 5B and F), while neither CD44s nor
CD44v8-10 stimulated cell invasion (Fig.
5D). MCF-7 cells have been demonstrated to possess low
metastatic potential and rarely metastasize to bone (16). Our previous study revealed that
overexpression of CD44s did not enhance the development of bone
metastases (16). Similarly, in the
present study, overexpression of CD44v8-10 failed to potentiate
metastatic potential and developed few lesions in bone (data not
shown).
Discussion
Accumulating evidence indicates that CD44 is
involved in various cancer phenotypes, including enhanced cell
proliferation, migration, invasion and metastasis (1,2).
Consistent with these findings, the present authors recently
reported that the expression of CD44 in cancer cells increases
tumorigenicity, cell migration and invasion, thereby promoting
cancer metastasis to bone (16).
However, it currently remains unknown whether CD44s and CD44v make
differential contributions to the development of bone
metastases.
Similar to the overexpression of CD44s, that of
CD44v8-10 in MCF-7 cells (which rarely express any isoforms of
CD44) significantly increased cell migration and tumor sphere
formation. These results suggest that CD44v8-10 has the potential
to promote tumor progression in a similar manner to CD44s. The
overexpression of ESRP1, which caused a switch in alternative
splicing from CD44s to CD44v (mainly CD44v8-10) did not change the
phenotypes or metastatic potential of MDA-MB-231 or A549 cells.
Considering that ESRP1 did not change the total amount of CD44, the
contribution of CD44v8-10 to the development of bone metastases is
likely comparable to that of CD44s. However, the overexpression of
CD44v8-10 did not further enhance the aggressiveness of MDA-MB-231
cells. Since MDA-MB-231 cells endogenously express high level of
CD44s, the efficacy of endogenous CD44s to enhance bone metastases
may have already reached its maximum level.
As discussed above, although cell migration and
tumor sphere formation were stimulated in MCF-7/CD44v cells, these
effects were less prominent than those observed in MCF-7/CD44s
cells (Fig. 5B and F). One
explanation for these results is that the expression levels of
CD44s and CD44v8-10 may not be completely the same. Another
possibility is that there are functional differences between CD44s
and CD44v8-10. CD44s has been demonstrated to interact with its
major ligand, hyaluronan, more effectively than CD44v8-10 (21). In addition, the CD44v segment is known
to include binding sites for several growth factors (2). These differences may become visible
under certain conditions.
Our previous study revealed that the overexpression
of CD44s in MCF-7 cells failed to increase bone metastases,
although cell migration and tumorigenicity were significantly
enhanced (16). Similar results were
obtained with MCF-7/CD44v cells, suggesting that solely CD44v8-10
is not sufficient to cause bone metastases.
Recent studies have proposed a critical contribution
of cancer stem-like cells to the development of metastases
(22). CD44 is one of the most
well-recognized markers for CSCs (2,3).
Consistent with this notion, our previous study demonstrated that
CD44s confers stem cell-like phenotypes to cancer cells, thus
leading to the increased formation of bone metastases (16). The present study, using MCF-7 cells,
revealed that CD44v8-10 also enhanced sphere formation, suggesting
that CD44v8-10, in addition to CD44s, has the ability to confer
stem cell-like properties to cancer cells and may be a marker for
CSCs.
It has been suggested that CD44v is predominantly
expressed in epithelial cells, whereas CD44s is mainly expressed in
mesenchymal cells (19). Furthermore,
the induction of epithelial-mesenchymal transition (EMT) is
accompanied by a shift from CD44v to CD44s (19). In contrast to these findings, Yae
et al reported that the expression of the CD44 isoforms is
not associated with that of EMT markers (20). In the present study, RT-PCR analysis
also revealed that the overexpression of ESRP1 and CD44v8-10 did
not cause consistent changes in the expression of the epithelial
marker E-cadherin or the mesenchymal marker vimentin in cancer
cells (data not shown). Thus, further studies are required to
clarify the molecular mechanism of the association between the
switch in CD44 isoform expression and EMT.
ESRP1 has been demonstrated to regulate the
alternative splicing of genes other than CD44, including fibroblast
growth factor receptor 2, catenin delta-1 (CTNND1) and enabled
homolog (ENAH) (18). The current
study confirmed by RT-PCR analysis that the overexpression of ESRP1
also changed the splicing patterns of CTNND1 and ENAH (data not
shown). The involvement of the ESRP1-induced splicing switch in
genes other than CD44 requires further investigation.
In conclusion, the present results collectively
suggest that CD44v8-10 promotes tumor aggressiveness and bone
metastases to a similar extent to CD44s, while their functional
differences have yet to be elucidated in detail. Thus, CD44v8-10,
in addition to CD44s, could be a potential therapeutic target for
the treatment of bone metastases.
Acknowledgements
The present study was partly supported by grants
from the Japan Society for the Promotion of Science (grant nos.,
JP21390505, JP23659883 and JP15K11093; Tokyo, Japan; Grants-in-Aid
for Scientific Research) and the Naito Foundation (Tokyo,
Japan).
Glossary
Abbreviations
Abbreviations:
|
CD
|
cluster of differentiation
|
|
CD44s
|
standard isoform of CD44
|
|
CD44v
|
variant isoform of CD44
|
|
CSCs
|
cancer stem cells
|
|
CTNND1
|
catenin delta-1
|
|
DMEM
|
Dulbecco's modified Eagle's medium
|
|
ECL
|
enhanced chemiluminescence
|
|
EMT
|
epithelial-mesenchymal transition
|
|
ENAH
|
enabled homolog
|
|
ESRP1
|
epithelial splicing regulatory protein
1
|
|
EV
|
empty vector
|
|
FBS
|
fetal bovine serum
|
|
GAPDH
|
glyceraldehyde-3-phosphate
dehydrogenase
|
|
PBS
|
phosphate-buffered saline
|
|
PE
|
phycoerythrin
|
|
RT-PCR
|
reverse transcription-polymerase chain
reaction
|
References
|
1
|
Ponta H, Sherman L and Herrlich PA: CD44:
From adhesion molecules to signalling regulators. Nat Rev Mol Cell
Biol. 4:33–45. 2003. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Zöller M: CD44: Can a cancer-initiating
cell profit from an abundantly expressed molecule? Nat Rev Cancer.
11:254–267. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Visvader JE and Lindeman GJ: Cancer stem
cells: Current status and evolving complexities. Cell Stem Cell.
10:717–728. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Günthert U, Hofmann M, Rudy W, Reber S,
Zöller M, Haussmann I, Matzku S, Wenzel A, Ponta H and Herrlich P:
A new variant of glycoprotein CD44 confers metastatic potential to
rat carcinoma cells. Cell. 65:13–24. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Friedrichs K, Franke F, Lisboa BW, Kügler
G, Gille I, Terpe HJ, Hölzel F, Maass H and Günthert U: CD44
isoforms correlate with cellular differentiation but not with
prognosis in human breast cancer. Cancer Res. 55:5424–5433.
1995.PubMed/NCBI
|
|
6
|
Seelentag WK, Günthert U, Saremaslani P,
Futo E, Pfaltz M, Heitz PU and Roth J: CD44standard and variant
isoform expression in human epidermal skin tumors is not correlated
with tumor aggressiveness but down-regulated during proliferation
and tumor de-differentiation. Int J Cancer. 69:218–224. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Olsson E, Honeth G, Bendahl PO, Saal LH,
Gruvberger-Saal S, Ringnér M, Vallon-Christersson J, Jönsson G,
Holm K, Lövgren K, et al: CD44 isoforms are heterogeneously
expressed in breast cancer and correlate with tumor subtypes and
cancer stem cell markers. BMC Cancer. 11:4182011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Okamoto I, Morisaki T, Sasaki J, Miyake H,
Matsumoto M, Suga M, Ando M and Saya H: Molecular detection of
cancer cells by competitive reverse transcription-polymerase chain
reaction analysis of specific CD44 variant RNAs. J Natl Cancer
Inst. 90:307–315. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yamaguchi A, Urano T, Goi T, Saito M,
Takeuchi K, Hirose K, Nakagawara G, Shiku H and Furukawa K:
Expression of a CD44 variant containing exons 8 to 10 is a useful
independent factor for the prediction of prognosis in colorectal
cancer patients. J Clin Oncol. 14:1122–7112. 1996.PubMed/NCBI
|
|
10
|
Yamaguchi A, Saito M, Goi T, Iida A,
Takeuchi K, Hirose K, Nakagawara G, Urano T, Furukawa K and Shiku
H: Expression of CD44 variant exons 8-10 in gastric cancer. Jpn J
Cancer Res. 86:1166–1171. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lau WM, Teng E, Chong HS, Lopez KA, Tay
AY, Salto-Tellez M, Shabbir A, So JB and Chan SL: CD44v8-10 is a
cancer-specific marker for gastric cancer stem cells. Cancer Res.
74:2630–2641. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zeng Y, Wodzenski D, Gao D, Shiraishi T,
Terada N, Li Y, Vander Griend DJ, Luo J, Kong C, Getzenberg RH and
Kulkarni P: Stress-response protein RBM3 attenuates the stem-like
properties of prostate cancer cells by interfering with CD44
variant splicing. Cancer Res. 73:4123–4133. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Weilbaecher KN, Guise TA and McCauley LK:
Cancer to bone: A fatal attraction. Nat Rev Cancer. 11:411–425.
2011. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Abraham BK, Fritz P, McClellan M,
Hauptvogel P, Athelogou M and Brauch H: Prevalence of
CD44+/CD24-/low cells in breast cancer may not be associated with
clinical outcome but may favor distant metastasis. Clin Cancer Res.
11:1154–1159. 2005.PubMed/NCBI
|
|
15
|
Balic M, Lin H, Young L, Hawes D, Giuliano
A, McNamara G, Datar RH and Cote RJ: Most early disseminated cancer
cells detected in bone marrow of breast cancer patients have a
putative breast cancer stem cell phenotype. Clin Cancer Res.
12:5615–5621. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Hiraga T, Ito S and Nakamura H: Cancer
stem-like cell marker CD44 promotes bone metastases by enhancing
tumorigenicity, cell motility, and hyaluronan production. Cancer
Res. 73:4112–4122. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Nakamura H, Kato R, Hirata A, Inoue M and
Yamamoto T: Localization of CD44 (hyaluronan receptor) and
hyaluronan in rat mandibular condyle. J Histochem Cytochem.
53:113–120. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Warzecha CC, Sato TK, Nabet B, Hogenesch
JB and Carstens RP: ESRP1 and ESRP2 are epithelial
cell-type-specific regulators of FGFR2 splicing. Mol Cell.
33:591–601. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Brown RL, Reinke LM, Damerow MS, Perez D,
Chodosh LA, Yang J and Cheng C: CD44 splice isoform switching in
human and mouse epithelium is essential for epithelial-mesenchymal
transition and breast cancer progression. J Clin Invest.
121:1064–1074. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yae T, Tsuchihashi K, Ishimoto T, Motohara
T, Yoshikawa M, Yoshida GJ, Wada T, Masuko T, Mogushi K, Tanaka H,
et al: Alternative splicing of CD44 mRNA by ESRP1 enhances lung
colonization of metastatic cancer cell. Nat Commun. 3:8832012.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bartolazzi A, Jackson D, Bennett K, Aruffo
A, Dickinson R, Shields J, Whittle N and Stamenkovic I: Regulation
of growth and dissemination of a human lymphoma by CD44 splice
variants. J Cell Sci. 108:1723–1733. 1995.PubMed/NCBI
|
|
22
|
Oskarsson T, Batlle E and Massagué J:
Metastatic stem cells: Sources, niches, and vital pathways. Cell
Stem Cell. 14:306–321. 2014. View Article : Google Scholar : PubMed/NCBI
|