Introduction
Clinical studies have revealed that the human
epidermal growth factor receptor 2 (HER2) gene is amplified
in 20–30% of all breast cancers (1),
and in ~90–95% of these cases, overexpression is a direct result of
gene amplification (2,3). The HER2 protein is a 185-kDa
transmembrane growth factor receptor with tyrosine kinase activity
involved in cellular signaling, which regulates cell growth and
development (4). HER2 gene
amplification or overexpression in breast cancer is a prognostic
factor and predictive of a more aggressive clinical course for the
patient (5). It is associated with
high tumor-grade, hormone receptor-negative tumors, lymph node
metastasis (6), increased risk of
recurrence after surgery, poor response to conventional
chemotherapy and shortened survival (7,8). In
addition, diagnostic assays for HER2 expression in breast cancer
have also a high predictive value (1)
and are important in therapeutic decision-making. Notably, the
HER2 gene product, p185HER2/neu, represents a target for
specific therapy with the humanized recombinant monoclonal antibody
trastuzumab (Herceptin®; Genentech, Inc., South San
Francisco, CA, USA) (9,10).
The efficacy of therapeutic regimens that include
trastuzumab, administered in combination with conventional
chemotherapy in both the metastatic and the adjuvant setting,
requires the accurate determination of HER2 status, since the
presence of this alteration is the criteria to determine the
patient eligibility for trastuzumab treatment (11,12).
Trastuzumab therapy improves survival rate among women with
metastatic or localized HER2-positive breast cancer (7,9,11,12). One
year of treatment provides a significant disease-free and overall
survival benefit, and is the standard of care (13). Analysis requires the application of
methods performed on archival formalin-fixed, paraffin-embedded
tissue (3). There are two
complementary pathological diagnostic tests in current clinical use
to determine HER2 status in breast cancer: Fluorescence in
situ hybridization (FISH) to evaluate HER2 gene
amplification and immunohistochemistry (IHC) to detect protein
overexpression; they examine different aspects of the biology of
HER2-driven cancer (14).
Approximately 80% of newly diagnosed invasive breast
cancers (IBCs) are tested for HER2 using IHC and 20% are tested
using FISH (15,16). The American Society of Clinical
Oncology (ASCO) and the College of American Pathologists (CAP)
recommend treating patients whose tumor test is IHC 3+ or
FISH-positive with trastuzumab, whereas patients whose tumors are
IHC 0 or 1+ or FISH-negative are treated with standard
chemotherapy. The same guidelines recommend also researching gene
amplification by FISH in tumors scoring 2+ (17). Recently, the 2013 ASCO/CAP guidelines
recommend either using IHC assays for initial evaluation of HER2
status followed by reflex testing by FISH of certain IHC
categories, or the primary use of FISH in initial testing (18). The agreement between IHC 3+ and FISH
amplification should be ≥95% (17).
The presence of ‘incomplete membrane staining that is faint/barely
perceptible and within >10% of tumor cells’ in those cases
scoring 1+, or the absence of HER2 protein immunoexpression, are
due to the absence of HER2 gene amplification in the
majority of cases (19); however, a
minor but significant number of cases that have faint/perceptible
IHC expression (score 1+) (20) or no
HER2 protein expression but exhibit HER2 gene amplification,
have been observed worldwide in different cohorts of patients
(7). These data were confirmed by
extensive internal and external international quality assurance of
HER2 testing (21). Discordance
between test results may be present in patients with unfavorable
tumor characteristics, including high histological grade, high
proliferative index and negative or low hormone receptor expression
(20), thus leading to false-negative
results for HER2 status. In consequence, patients with a
false-negative tumor result would be denied the clinical benefits
of trastuzumab or other HER2-targeted therapies (22).
In fact, the aim of the ASCO/CAP guidelines update
in 2013 was to focus on accurate HER2 testing to ensure access to
high-quality cancer biomarker tests that would aid specialists to
match the right treatments with the right patients (18). Therefore, even if the absolute number
of HER2-positive cases detected could be very low, it would be
clinically useful to test HER2 gene amplification in
selected tumors with adverse features scoring 1+ by IHC.
The aim of the present study is to assess the
incidence of HER2 gene amplification in selected tumors with
adverse features that scored 1+ by IHC.
Materials and methods
Patients
A total of 331 consecutive IBCs observed between
January and December 2013 were tested by IHC for HER2, of which, 42
tumors scored 3+ (13%); 43 tumors scored 2+ (13%), of which 12
cases exhibited HER2 amplification; 102 tumors scored 1+ (31%); and
144 tumors scored 0 (43.5%). In total, 75 out of 102 (73.5%) IBC
cases scoring 1+ by IHC, which occurred in women who underwent
surgery at the National Cancer Research Institute ‘Giovanni Paolo
II’ (Bari, Italy), were selected for the study. The other 27 out of
102 cases were detected on metastatic sites or core biopsies and
were discarded. A total of 48 out of 75 (64%) IBC cases (patients'
median age, 60.75 years) were selected according to ≥1 unfavorable
tumor characteristics, and subsequently tested by FISH.
Ethics statement
The present study was approved by the Institutional
Review Board of the National Cancer Research Institute ‘Giovanni
Paolo II’. Before undergoing routine surgery, all patients signed
an informed consent form authorizing the use of the removed
biological tissue for research purposes according to ethical
standards.
IHC analysis
Samples were tested by IHC to observe the expression
of estrogen receptor (ER) and progesterone receptor (PgR), and to
evaluate the Ki-67 cellular proliferation index. Hormone receptors
for estrogen and progesterone were tested using monoclonal rabbit
anti-human estrogen receptor α (clone SP1; 1:60 dilution; Dako,
Glostrup, Denmark) and monoclonal mouse anti-human progesterone
receptor (clone PgR 636; 1:100 dilution; Dako) respectively,
whereas Ki-67 was detected using monoclonal mouse anti-human Ki-67
antigen [clone mindbomb E3 ubiquitin protein ligase 1 (MIB-1); 1:80
dilution; Dako]. ER, PgR and Ki-67 immunostaining were confined to
the nucleus. ER, PgR and Ki-67 index were scored according to the
St. Gallen International Breast Cancer Conference guidelines
(23,24): ER and PgR receptors were scored as
negative/positive when no/any staining was present in the tumor,
while Ki-67-labelling index was considered high when staining was
present in >30% of tumor cells, intermediate when it was 16–30%,
and low when it was ≤15%. In the present study, two subgroups were
considered: when Ki-67-labelling index was >30%, it was
considered high, whereas when Ki-67 was ≤30%, it was considered
low. All samples were also analyzed by IHC using the HerceptTest™
kit (Dako) according to the manufacturers' protocol. Cytoplasmic
immunoreactivity was ignored. HER2 was scored as 0, 1+, 2+ or 3+ in
accordance with the ASCO/CAP guidelines, also adopted by the
Italian Society of Pathological Anatomy and Diagnostic Cytology -
Italian Division of the International Academy of Pathology
(SIAPEC-IAP) (18): 0, no staining
observed or membrane staining that is incomplete or faint/barely
perceptible in ≤10% of tumor cells; 1+, incomplete membrane
staining that is faint/barely perceptible within >10% of tumor
cells; 2+, circumferential membrane staining that is incomplete
and/or weak/moderate within >10% of tumor cells, or complete and
circumferential membrane staining that is intense within ≤10% of
tumor cells; and 3+, circumferential membrane staining that is
complete and intense within >10% of tumor cells.
FISH for gene amplification
FISH was conducted using a dual HER2/Cep17 probe
(Path Vysion HER2 DNA Probe kit; Abbott Molecular, Inc., Des
Plaines, IL, USA), combining a HER2 gene probe (190 kb
Spectrum Orange-directly labelled DNA probe) with a centromeric
enumeration probe for chromosome 17 (CEP17; 5.4 kb Spectrum
Green-directly labelled fluorescent DNA probe specific for the
chromosome 17 α satellite DNA sequence). Unstained sections of
target tissue (4-µm-thick) were cut from paraffin-embedded blocks.
The sections were baked overnight at 56°C. Subsequently, the
paraffin was removed from the sections with a 15-min wash in warm
xylene at 60°C and 3 × 15-min washes in xylene. The samples were
dehydrated twice in 100% ethanol for 5 min and dried in the HYBrite
instrument (Abbott Molecular, Inc.) at 45°C. The sections were
fixed in methanol:acetic acid (3:1) for 12 min, dried in the
HYBrite instrument at 45°C, and then immersed in 0.2 M HCl for 10
min, in purified water for 3 min and in a 2X saline sodium citrate
(SSC) wash buffer for 3 min. The slides were then placed in a
pretreatment sodium thiocyanate solution for 25 min at 84°C and
rinsed in purified water for 3 min, followed by rinsing in 2X SSC
wash buffer for 3 min. After incubation in a protease solution at
37°C for 15 min, the enzymatic reaction was stopped by placing the
slides in deionized water for 3 min and air dried. Next, the slides
were dehydrated through graded alcohols, and 10 µl
PathVysion® HER2/CEP17 probe was applied to the
sections. The slides were coverslipped, sealed with rubber cement,
and the probe/target tissue was then co-denatured for 5 min at 75°C
using the HYBrite instrument and allowed to hybridize overnight at
37°C. The coverslip was carefully removed in a 1X SSC/0.1% NP-40
solution, and to remove non-specifically bound probe, the slides
were washed in 1X SSC/0.1% NP-40 for 5 min. Stringency wash was
performed with 2X SSC/0.3% NP-40 for 5 min at room temperature, and
then at 72°C for 3 min. The sections were washed twice in 1X
SSC/0.1% NP-40. Slides were air dried in the dark, counterstained
with 10 µl 4′,6-diamidino-2-phenylindole (DAPI) and
coverslipped.
FISH analysis and interpretation
FISH analysis was performed using an epifluorescence
microscope (BX-UCB; Olympus Corporation, Tokyo, Japan) with
appropriate filters for Spectrum Orange and Green, a triple
bandpass filter set, and an ultraviolet filter for DAPI nuclear
counterstain. Normal (×10) and oil fluorescence objectives (×60 and
×100) were used for the analysis. Analysis was performed in a dark
room, and the DAPI filter set and a low-power objective were used
to confirm areas of invasive carcinoma. Using the triple bandpass
filter set and a 60X oil objective, the presence of CEP17 signals
in ≥75% of cancer cell nuclei was confirmed. Only tumor cells with
non-overlapping nuclei were scored. The signals were recorded with
a charge-coupled device camera (Olympus Corporation), and analysis
of the signal pattern was performed with CytoVision®
software (version 4.5.4; Leica Microsystems, Inc., Buffalo Grove,
IL, USA).
A total of 60 nuclei from two distinct areas of the
invasive carcinoma were scored for green and red signals for each
section; red signals represent HER2 gene copies, green
signals represent CEP17 gene copies. The mean number of
CEP17 and HER2 signals was recorded, and the results were expressed
as a ratio of red to green signals.
In agreement with the ASCO/CAP guidelines (18), which have also been adopted by
SIAPEC-IAP, HER2 ratio-based amplification was considered. Gene
amplification was evaluated as present when the HER2/CEP17 ratio
was ≥2 or when the mean HER2 copy number was ≥6.
Before and during the study there was a
between-laboratory quality assessment exercise involving the
circulation of control sections.
Statistical methods
The baseline characteristics of the study population
were calculated, and the results were expressed as frequencies and
percentages for the categorical variables.
Comparisons of clinical parameters between the
groups of interest were performed with the Pearson χ2
test or the Fisher's exact test, when appropriate, for categorical
variables. P<0.05 was considered to indicate a statistically
significant difference. All the analyses were performed using the
Statistical Analysis System software (SAS Institute, Cary, NC,
USA).
Results
A total of 48 IBC samples with unfavorable tumor
characteristics, including high histological grade (G3) according
to the Elston-Ellis classification (25), high proliferative index, lymph node
positivity, presence of peritumoral vascular invasion and negative
hormone receptor expression, were selected.
Clinicopathological data, including histological
grade, peritumoral vascular invasion, lymph node status, Ki-67
index, ER and PgR status are shown in Table I.
 | Table I.Tumor characteristics in 48 breast
cancer cases scoring 1+ by immunohistochemistry for HER2
expression. |
Table I.
Tumor characteristics in 48 breast
cancer cases scoring 1+ by immunohistochemistry for HER2
expression.
| Tumor
characteristics | IDC histotype | ILC histotype | Total (n) | % |
|---|
| Histological grade
(G3) | 19 | 3 | 22 | 46 |
| Peritumoral
vascular invasion | 23 | 4 | 27 | 56 |
| Lymph
node-positive | 27 | 5 | 32 | 78 |
| Ki-67 >30% | 22 | 1 | 23 | 48 |
| ER-negative | 3 | 0 | 3 | 6 |
| PgR-negative | 10 | 0 | 10 | 21 |
In total, 42 out of 48 cases (87.5%) exhibited a
histological diagnosis of infiltrating ductal carcinoma (IDC) and 6
cases (12.5%) were diagnosed as infiltrating lobular carcinoma
(ILC). A total of 22 out of 48 tumors (46%) displayed high
histological grade (G3) (Fig. 1A),
and 27 cases (56%) exhibited peritumoral vascular invasion.
Regarding lymph node status, 41 patients out of 48 had axillary
lymph node dissection, of which, 32 (78%) were node-positive, 9
were node-negative and 7 had no axillary lymph node dissection. A
total of 23 cases (48%) had a high proliferative index (Ki-67,
>30%) (Fig. 1B), while 3 (6%) and
10 (21%) cases were negative for ER and PgR expression,
respectively. FISH was performed on 48 HER2 samples scoring 1+ by
IHC (Fig. 1C) with unfavorable tumor
characteristics, and 7 IDCs out of 48 (14.6%) exhibited HER2
amplification (Fig. 1D); all the 7
samples displayed a high proliferative index (Ki-67, >30%) and
5/7 had high histological grade. No amplification was detected in
any of the 6 ILCs (Table II). In
total, 42 tumors scored 3+ by IHC and 43 cases scored 2+, 12 of
which resulted amplified by FISH, indicating that 54 cases (16%)
out of 331 were overexpressed and/or amplified. A total of 7 cases
(2%) cases scoring 1+ by IHC out of 331 exhibited HER2
amplification, indicating that, in total, 61 (18%) tumors out of
331 were overexpressed and/or amplified.
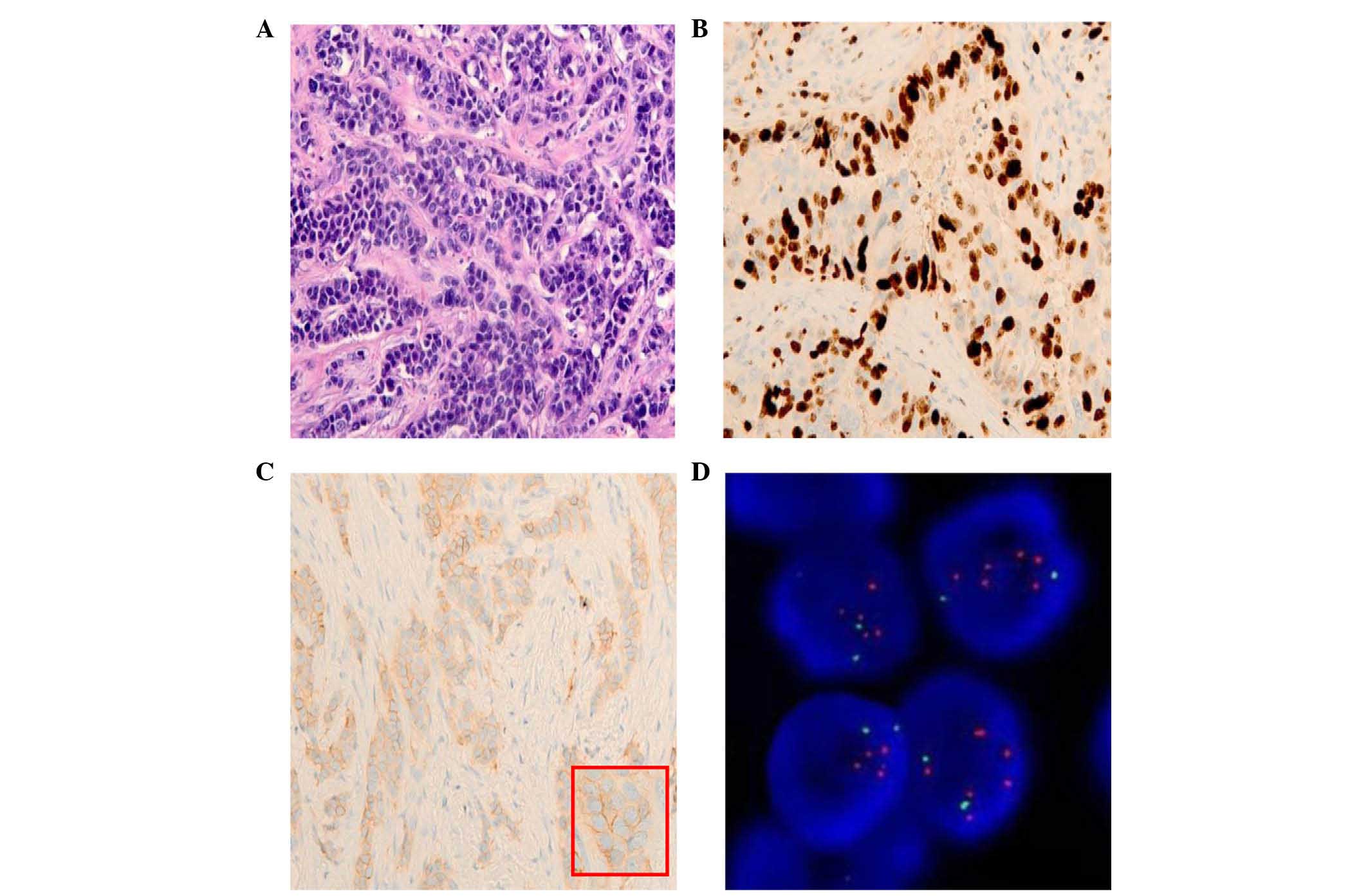 | Figure 1.Representative images of H&E and
IHC staining and FISH test in an IBC. (A) H&E staining of an
IBC with high histological grade (G3) (magnification, ×20). (B)
High Ki-67 expression in an invasive breast tumor (magnification,
×20). (C) IHC HER2 protein expression in an IBC (HER2 score, 1+;
magnification, ×20). An enlarged image of HER2 protein expression
is presented on the bottom right part of the image (red frame;
magnification, ×40). (D) HER2 gene amplification tested by
FISH. Red signals represent HER2 gene copies, while green
signals represent chromosome enumeration probe 17 copies (oil
fluorescence objective; magnification, ×60). H&E, hematoxylin
and eosin; IHC, immunohistochemistry; FISH, fluorescence in
situ hybridization; IBC, invasive breast carcinoma; HER2, human
epidermal growth factor receptor 2. |
 | Table II.FISH results in 48 breast cancer
cases scoring 1+ by immunohistochemistry for HER2 expression. |
Table II.
FISH results in 48 breast cancer
cases scoring 1+ by immunohistochemistry for HER2 expression.
|
| IDC histotype
(n=42) | ILC histotype
(n=6) |
|---|
|
|
|
|
|---|
| Tumor
characteristics | FISH+, n
(%) | FISH−, n
(%) | FISH+, n
(%) | FISH−, n
(%) |
|---|
| Histological grade
(G3) | 5
(12) | 14 (33) | 0 (0) | 3 (50) |
| Peritumoral
vascular invasion | 3 (7) | 20 (48) | 0 (0) | 4 (67) |
| Lymph
node-positive | 3 (7) | 24 (57) | 0 (0) | 5 (83) |
| Ki-67 >30% | 7
(17) | 15 (36) | 0 (0) | 1 (17) |
| ER-negative | 1 (2) | 2 (5) | 0 (0) | 0 (0) |
| PgR-negative | 3 (7) | 7 (17) | 0 (0) | 0 (0) |
In 42 IDC samples, statistical analysis evidenced a
significant association between histological grade and high
proliferative index as detected by MIB-1 (P=0.0200), whereas no
association was noted regarding peritumoral vascular invasion or
presence of metastases (Table III).
Additionally, in 48 HER2 breast cancer samples scoring 1+, a
significant association between the presence of gene amplification
and high proliferative index was also observed (P=0.0033) (Table IV).
 | Table III.Association between histological
grade and unfavorable tumor characteristics in 42 infiltrating
ductal carcinoma cases scoring 1+ by immunohistochemistry for HER2
expression. |
Table III.
Association between histological
grade and unfavorable tumor characteristics in 42 infiltrating
ductal carcinoma cases scoring 1+ by immunohistochemistry for HER2
expression.
|
|
| High histological
grade (n=19) | Low histological
grade (n=23) |
|
|---|
|
|
|
|
|
|
|---|
| Variable | Category | n | % | n | % | P-value |
|---|
| Peritumoral
vascular invasionc | 0 | 10 | 52.63 | 9 | 39.13 | 0.3816a |
|
| 1 | 9 | 47.37 | 14 | 60.87 |
|
| Ki-67
>30%d | 0 | 5 | 26.32 | 14 | 60.87 | 0.0251a |
|
| 1 | 14 | 73.68 | 9 | 39.13 |
|
| Lymph node
statuse | 0 | 6 | 42.86 | 3 | 13.64 | 0.1111b |
|
| 1 | 8 | 57.14 | 19 | 86.36 |
|
 | Table IV.Association between unfavorable tumor
characteristics and HER2 amplification in 48 breast cancer cases
scoring 1+ by immunohistochemistry for HER2 expression. |
Table IV.
Association between unfavorable tumor
characteristics and HER2 amplification in 48 breast cancer cases
scoring 1+ by immunohistochemistry for HER2 expression.
|
|
| FISH=0
(n=41)a | FISH=1
(n=7)a |
|
|---|
|
|
|
|
|
|
|---|
| Variable | Category | n | % | n | % |
P-valueb |
|---|
| Histological
gradec | L | 24 | 58.54 | 2 |
28.57 | 0.2226 |
|
| H | 17 | 41.46 | 5 |
71.43 |
|
|
Histotyped | 0 | 35 | 85.37 | 7 | 100.00 | 0.5725 |
|
| 1 | 6 | 14.63 | 0 |
0.00 |
|
| Peritumoral
vascular invasione | 0 | 17 | 41.46 | 4 |
57.14 | 0.6830 |
|
| 1 | 24 | 58.54 | 3 |
42.86 |
|
| Ki-67
>30%f | 0 | 25 | 60.98 | 0 |
0.00 | 0.0033 |
|
| 1 | 16 | 39.02 | 7 | 100.00 |
|
| Lymph node
statusg | 0 | 6 | 17.14 | 3 |
50.00 | 0.1075 |
|
| 1 | 29 | 82.86 | 3 |
50.00 |
|
Discussion
With the aim of reaching high-quality personalized
medicine, one of the major aims of the 2013 ASCO/CAP guidelines
update (18) was to aid breast cancer
specialists to accurately classify patients for HER2-targeted
treatment, thus avoiding false-negative and false-positive HER2
results, as false-negative patients may be denied biological
treatment, while false-positive cases may receive potentially
toxic, costly and ineffective treatment (26).
Despite the axiom ‘the right treatment with the
right patient’ reported in the ASCO/CAP 2013 updated guideline
recommendations (18), Iorfida et
al (20) demonstrated that a
considerable percentage (13%) of cases that scored 1+ for HER2
protein expression by IHC and tested for gene-copy ratio by FISH
exhibited gene amplification. This issue evidenced the possibility
to deny an effective therapy in a subset of breast cancer
patients.
This observation led to the following open
questions: i) Could the percentage of IHC 1+/FISH+ cases
be higher than expected?; ii) is the reason for this disagreement
technical (quality of the determination) or biological (subset of
not-overexpressed/amplified tumors)?; iii) could the selection of
certain unfavorable tumor characteristics be helpful in identifying
these cases?; and iv) should the selection criteria for the FISH
test be reconsidered?
In an attempt to address the above questions, the
present study selected 48 IBCs scoring 1+ by IHC for HER2 protein
expression according to biopathological parameters of clinical
aggressiveness, as previously reported by Iorfida et al
(20). These cases exhibited a
disagreement between absent or very low HER2 protein expression
detected by IHC and the presence of gene amplification detected by
FISH in 7 IDCs (14.6%). Consistent with previous reports, the
present data revealed that there is ~15% of
IHC−/FISH+ cases in the selected subset of 48
tumors (20,27). Regarding the causes of IHC/FISH
disagreement, Perez et al (28) described from a technical point of view
the discordance in HER2 results between local and central
laboratories participating in clinical trials. Furthermore, the
ASCO/CAP guidelines emphasized the requirement for very stringent
controls, particularly for new laboratories or when a new assay is
adopted (29). Regarding the present
study, it should be highlighted that the Molecular Pathology
Laboratory (Department of Pathology, National Cancer Research
Institute ‘Giovanni Paolo II’) is well inserted, with good results,
in the National Italian program for HER2 testing both for FISH
[control quality (CQ) FISH HER2 SIAPEC-IAP] and for IHC (Nordic
CQ), promoted by SIAPEC-IAP (18).
Other than technical issues, biological features
could also be important, such as the importance of heterogeneity in
determining the discrepancy in HER2 status within the tumor or
between the tumor and its metastasis (30). Several studies have also demonstrated
cell-to-cell heterogeneity of HER2 gene amplification and HER2
protein expression at highly variable rates (1–50%), depending on
the methodologies and the sample set used (31–34).
However determined, molecular heterogeneity of HER2 gene
amplification is recognized in ≥4–5% of breast cancers (32,35) and
appears to be more frequent in advanced disease.
In the present study, the detection of areas of
heterogeneity in the same tumor was present both for IHC and FISH
in the 7 IHC−/FISH+ tumors analyzed. Notably,
a number of IHC−/FISH+ areas were present
where the sample was less well fixed, as observed by hematoxylin
and eosin staining, or a delay in the time of fixation was
recorded. These observations led to further emphasize the
importance of a correct pre-analytical phase (36) in order to obtain HER2 gene
amplification and protein overexpression level in concert. However,
it must be considered that variability in HER2 testing may arise
from pre-analytic, analytic and post-analytic factors according to
the testing method (28).
The main purpose of the current study was to
identify various unfavorable tumor characteristics that could
distinguish discordant cases, particularly those that were not
overexpressed/amplified. Discordance in HER2 results is often
present among tumor cases that are selected according to
unfavorable and histological factors. As reported in previous
studies, the presence of certain biopathological factors such as
peritumoral vascular invasion, high histological grade and high
proliferative index, appears to play a fundamental role in the
identification of these cases (20,27). In
the subset of 48 patients selected in the present study, the
incidence of amplified HER2 cases rose to 14.6%, and proved to be
significantly associated with an elevated cellular kinetic index.
In the present study, when the Ki-67 index was >30%, it was
considered high. It must be highlighted that numerous cut-off
values have been proposed in the literature (37,38), and
that the reproducibility of the test for Ki-67 is still far from
being elevated, due to an important inter-observer and
inter-laboratory variability (38).
Recently, Goldhirsch et al (39) classified the Ki-67 index into three
classes, considering as cut-off value the presence of ≥20% of
neoplastic, invasive Ki-67-positive cells. Furthermore, Iorfida
et al (20), utilized a
cut-off value of 14%, including also cases with moderate
proliferative activity, according to the aforementioned
Goldhirsch's classification. The present study adopted a cut-off
value of 30% (23), which is higher
and, in the authors' opinion, more restrictive, in order to
identify tumors with a more aggressive phenotype.
The present data suggest that it could be advisable
to perform the FISH test in IHC 1+ breast cancers, in order to
identify HER2-positive cases that could be misclassified as
HER2-negative, thus denying those patients the opportunity of
benefitting from HER2-targeted therapy. According to the current
ASCO/CAP guidelines (18), FISH is
not performed in HER2 cases scoring 1+. It is clear that if the
current guidelines were applied, these patients would be denied
biological therapy from which they could benefit. However, the
magnitude of the problem in terms of cost/benefit ratio must be
defined. In the present cases, the global incidence of
HER2-positive cases (overexpressed and/or amplified) was 16%, and
there was a IHC−/FISH+ disagreement of 2%
(7/331), which led to the same increment of HER2-positive cases
(18%). Thus, this is the risk, according to our experience.
In terms of cost/benefit ratio, this 2% increment in
the study population does not appear to be sufficient to justify
extending the FISH test to HER2 cases scoring 1+ by IHC. However,
~7 patients from our Institute, who could have benefited from
HER2-targeted therapy, had treatment denied to them or initiated
too late.
In conclusion, based on the biopathological
parameters discussed in the present study, and particularly a high
proliferative index, the results from the current study suggest
that there is a higher probability of identifying tumors scoring 1+
by IHC that exhibit HER2 amplification by FISH, thus aiding
the selection of patients who are suitable for HER2-targeted
therapy according to an acceptable cost/benefit ratio.
Acknowledgements
The authors would like to thank Mrs. Milena Zambetti
(Department of Pathology, National Cancer Research Centre ‘Giovanni
Paolo II’, Bari, Italy) for her technical assistance, Dr Caroline
Oakley (Scientific Directorate, National Cancer Research Centre
‘Giovanni Paolo II’) for the revision of the present manuscript and
Dr Giusi Graziano (Scientific Directorate, National Cancer Research
Centre ‘Giovanni Paolo II’) for the statistical analyses.
References
|
1
|
Powell WC, Hicks DG, Prescott N, Tarr SM,
Laniauskas S, Williams T, Short S, Pettay J, Nagle RB, Dabbs DJ, et
al: A new rabbit monoclonal antibody (4B5) for the
immunohistochemical (IHC) determination of the HER2 status in
breast cancer: Comparison with CB11, fluorescence in situ
hybridization (FISH), and interlaboratory reproducibility. Appl
Immunohistochem Mol Morphol. 15:94–102. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Slamon DJ, Godolphin W, Jones LA, Holt JA,
Wong SG, Keith DE, Levin WJ, Stuart SG, Udove J, Ullrich A, et al:
Studies of the HER-2/neu proto-oncogene in human breast and ovarian
cancer. Science. 244:707–712. 1989. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Pauletti G, Godolphin W, Press MF and
Slamon DJ: Detection and quantitation of HER-2/neu gene
amplification in human breast cancer archival material using
fluorescence in situ hybridization. Oncogene. 13:63–72.
1996.PubMed/NCBI
|
|
4
|
Yarden Y: Biology of HER2 and its
importance in breast cancer. Oncology. 61(Suppl 2): S1–S13. 2001.
View Article : Google Scholar
|
|
5
|
Slamon DJ, Clark GM, Wong SG, Levin WJ,
Ullrich A and McGuire WL: Human breast cancer: Correlation of
relapse and survival with amplification of the HER-2/neu oncogene.
Science. 235:177–182. 1987. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Eccles SA: The role of c-erbB-2/HER2/neu
in breast cancer progression and metastasis. J Mammary Gland Biol
Neoplasia. 6:393–406. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Dendukuri N, Khetani K, McIsaac M and
Brophy J: Testing for HER2-positive breast cancer: A systematic
review and cost-effectiveness analysis. CMAJ. 176:1429–1434. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ross JS and Fletcher JA: The HER-2/neu
oncogene in breast cancer: Prognostic factor, predictive factor,
and target for therapy. Stem Cells. 16:413–428. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cobleigh MA, Vogel CL, Tripathy D, Robert
NJ, Scholl S, Fehrenbacher L, Wolter JM, Paton V, Shak S, Lieberman
G and Slamon DJ: Multinational study of the efficacy and safety of
humanized anti-HER2 monoclonal antibody in women who have
HER2-overexpressing metastatic breast cancer that has progressed
after chemotherapy for metastatic disease. J Clin Oncol.
17:2639–2648. 1999.PubMed/NCBI
|
|
10
|
Pauletti G, Dandekar S, Rong H, Ramos L,
Peng H, Seshadri R and Slamon DJ: Assessment of methods for
tissue-based detection of the HER-2/neu alteration in human breast
cancer: A direct comparison of fluorescence in situ hybridization
and immunohistochemistry. J Clin Oncol. 18:3651–3664.
2000.PubMed/NCBI
|
|
11
|
Romond EH, Perez EA, Bryant J, Suman VJ,
Geyer CE Jr, Davidson NE, Tan-Chiu E, Martino S, Paik S, Kaufman
PA, et al: Trastuzumab plus adjuvant chemotherapy for operable
HER2-positive breast cancer. N Engl J Med. 353:1673–1684. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Piccart-Gebhart MJ, Procter M,
Leyland-Jones B, Goldhirsch A, Untch M, Smith I, Gianni L, Baselga
J, Bell R, Jackisch C, et al: Trastuzumab after adjuvant
chemotherapy in HER2-positive breast cancer. N Engl J Med.
353:1659–1672. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Goldhirsch A, Gelber RD, Piccart-Gebhart
MJ, de Azambuja E, Procter M, Suter TM, Jackisch C, Cameron D,
Weber HA, Heinzmann D, et al: 2 years versus 1 year of adjuvant
trastuzumab for HER2-positive breast cancer (HERA): An open-label,
randomised controlled trial. Lancet. 382:1021–1028. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bloom KJ and Cote RJ: Counterpoint: Both
immunohistochemistry and fluorescence in situ hybridization play
important roles for HER2 evaluation. Clin Chem. 57:983–985. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ross JS, Slodkowska EA, Symmans WF,
Pusztai L, Ravdin PM and Hortobagyi GN: The HER-2 receptor and
breast cancer: Ten years of targeted anti-HER-2 therapy and
personalized medicine. Oncologist. 14:320–368. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Garrison LP Jr, Lalla D, Brammer M,
Babigumira JB, Wang B and Perez EA: Assessing the potential
cost-effectiveness of retesting IHC0, IHC1+, or FISH-negative early
stage breast cancer patients for HER2 status. Cancer.
119:3113–3122. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wolff AC, Hammond ME, Schwartz JN, Hagerty
KL, Allred DC, Cote RJ, Dowsett M, Fitzgibbons PL, Hanna WM, Langer
A, et al: American Society of Clinical Oncology/College of American
Pathologists guideline recommendations for human epidermal growth
factor receptor 2 testing in breast cancer. J Clin Oncol.
25:118–145. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wolff AC, Hammond ME, Hicks DG, Dowsett M,
McShane LM, Allison KH, Allred DC, Bartlett JM, Bilous M,
Fitzgibbons P, et al: Recommendations for human epidermal growth
factor receptor 2 testing in breast cancer: American Society of
Clinical Oncology/College of American Pathologists clinical
practice guideline update. J Clin Oncol. 31:3997–4013. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Brunello E, Bogina G, Bria E, Vergine M,
Zamboni G, Pedron S, Daniele I, Furlanetto J, Carbognin L, Marconi
M, et al: The identification of a small but significant subset of
patients still targetable with anti-HER2 inhibitors when affected
by triple negative breast carcinoma. J Cancer Res Clin Oncol.
139:1563–1568. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Iorfida M, Dellapasqua S, Bagnardi V,
Cardillo A, Rotmensz N, Mastropasqua MG, Bottiglieri L, Goldhirsch
A, Viale G and Colleoni M: HER2-negative (1+) breast cancer with
unfavorable prognostic features: To FISH or not to FISH? Ann Oncol.
23:1371–1372. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bartlett JM, Ibrahim M, Jasani B, Morgan
JM, Ellis I, Kay E, Connolly Y, Campbell F, O'Grady A, Barnett S
and Miller K: External quality assurance of HER2 FISH and ISH
testing: Three years of the UK national external quality assurance
scheme. Am J Clin Pathol. 131:106–111. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Tinoco G, Warsch S, Glück S, Avancha K and
Montero AJ: Treating breast cancer in the 21st century: Emerging
biological therapies. J Cancer. 4:117–132. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Goldhirsch A, Ingle JN, Gelber RD, Coates
AS, Thürlimann B and Senn HJ: Panel members: Thresholds for
therapies: Highlights of the St Gallen International Expert
Consensus on the primary therapy of early breast cancer 2009. Ann
Oncol. 20:1319–1329. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Jalava P, Kuopio T, Juntti-Patinen L,
Kotkansalo T, Kronqvist P and Collan Y: Ki67 immunohistochemistry:
A valuable marker in prognostication but with a risk of
misclassification: Proliferation subgroups formed based on Ki67
immunoreactivity and standardized mitotic index. Histopathology.
48:674–682. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Elston CW and Ellis IO: Pathological
prognostic factors in breast cancer. I. The value of histological
grade in breast cancer: Experience from a large study with
long-term follow-up. Histopathology. 19:403–410. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Rakha EA, Starczynski J, Lee AH and Ellis
IO: The updated ASCO/CAP guideline recommendations for HER2 testing
in the management of invasive breast cancer: A critical review of
their implications for routine practice. Histopathology.
64:609–615. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lambein K, Van Bockstal M, Denys H and
Libbrecht L: 2013 update of the American Society of Clinical
Oncology/College of American Pathologists guideline for human
epidermal growth factor receptor 2 testing: Impact on
immunohistochemistry-negative breast cancers. J Clin Oncol.
32:1856–1857. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Perez EA, Cortés J, Gonzalez-Angulo AM and
Bartlett JM: HER2 testing: Current status and future directions.
Cancer Treat Rev. 40:276–284. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wolff AC, Hammond ME, Hicks DG, Dowsett M,
McShane LM, Allison KH, Allred DC, Bartlett JM, Bilous M,
Fitzgibbons P, et al: Recommendations for human epidermal growth
factor receptor 2 testing in breast cancer: American Society of
Clinical Oncology/College of American Pathologists clinical
practice guideline update. Arch Pathol Lab Med. 138:241–256. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Wu JM, Halushka MK and Argani P:
Intratumoral heterogeneity of HER-2 gene amplification and protein
overexpression in breast cancer. Hum Pathol. 41:914–917. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Vance GH, Barry TS, Bloom KJ, Fitzgibbons
PL, Hicks DG, Jenkins RB, Persons DL, Tubbs RR and Hammond ME:
College of American Pathologists: Genetic heterogeneity in HER2
testing in breast cancer: Panel summary and guidelines. Arch Pathol
Lab Med. 133:611–612. 2009.PubMed/NCBI
|
|
32
|
Bartlett AI, Starcyznski J, Robson T,
Maclellan A, Campbell FM, van de Velde CJ, Hasenburg A, Markopoulos
C, Seynaeve C, Rea D and Bartlett JM: Heterogeneous HER2 gene
amplification: Impact on patient outcome and a clinically relevant
definition. Am J Clin Pathol. 136:266–274. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Lee S, Jung W, Hong SW and Koo JS:
Evaluation of intratumoral HER-2 heterogeneity by fluorescence in
situ hybridization in invasive breast cancer: A single institution
study. J Korean Med Sci. 26:1001–1006. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ohlschlegel C, Zahel K, Kradolfer D, Hell
M and Jochum W: HER2 genetic heterogeneity in breast carcinoma. J
Clin Pathol. 64:1112–1126. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Starczynski J, Atkey N, Connelly Y,
O'Grady T, Campbell FM, di Palma S, Wencyk P, Jasani B, Gandy M and
Bartlett JM: UKNEQAS: HER2 gene amplification in breast cancer: A
rogues' gallery of challenging diagnostic cases: UKNEQAS
interpretation guidelines and research recommendations. Am J Clin
Pathol. 137:595–605. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Bussolati G, Annaratone L and Maletta F:
The pre-analytical phase in surgical pathology. Recent Results
Cancer Res. 199:1–13. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Dowsett M, Nielsen TO, A'Hern R, Bartlett
J, Coombes RC, Cuzick J, Ellis M, Henry NL, Hugh JC, Lively T, et
al: Assessment of Ki67 in breast cancer: Recommendations from the
International Ki67 in breast cancer working group. J Natl Cancer
Inst. 103:1656–1664. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Tashima R, Nishimura R, Osako T, Nishiyama
Y, Okumura Y, Nakano M, Fujisue M, Toyozumi Y and Arima N:
Evaluation of an optimal cut-off point for the Ki-67 index as a
prognostic factor in primary breast cancer: A retrospective study.
PLoS One. 10:e01195652015. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Goldhirsch A, Winer EP, Coates AS, Gelber
RD, Piccart-Gebhart M, Thürlimann B and Senn HJ: Panel members:
Personalizing the treatment of women with early breast cancer:
Highlights of the St Gallen International Expert Consensus on the
Primary Therapy of Early Breast Cancer 2013. Ann Oncol.
24:2206–2223. 2013. View Article : Google Scholar : PubMed/NCBI
|















