Introduction
Breast cancer is the second most common cancer
worldwide and is the leading cause of cancer-associated mortality
in women, accounting for ~15% of all cancer-associated mortalities
in women (1). Breast cancer is
heterogeneous and can be classified into several subtypes,
including luminal, human epidermal growth factor 2 (HER2) and
triple-negative breast cancer (TNBC) subtypes, based on the
expression of estrogen receptor (ER) and progesterone receptor (PR)
and the receptor tyrosine kinase erbB-2 (HER2) (2,3). Thus, it
is important to understand the molecular mechanisms involved in the
development and the acquisition of malignancy in breast tumor, and
develop more effective treatments for breast cancer patients.
The development, local invasion or metastasis of
breast cancer is involved in the dysregulation, mutation and
epigenetic mechanism of various genes (4). The dysregulated genes include coding RNA
and non-coding RNA, such as microRNAs (miRNAs) (5). miRNAs are able to silence gene
expression by targeting complementary regions of mRNAs and
inhibiting protein translation, which is critical in normal and
abnormal biological processes, including cancer (6). Dysregulation of miRNAs has been observed
in breast cancer and is associated with tumor growth, drug
resistance and metastasis (7).
Therefore, therapeutic strategies based on modulating the
expression levels of miRNAs are promising approaches for breast
cancer treatment.
miR-125b is dysregulated in a variety of tumors;
however, as miR-125b is either upregulated or downregulated in
different tumors, this suggests that the oncogenic and tumor
suppressive potential of miR-125b is dependent on the type of
cancers (8). In addition, previous
studies have shown a different role of miR-125b in breast cancer.
Ferracin et al found a downregulation of miR-125b in
metastatic breast cancers (9), which
may account for hypermethylation of the miR-125b promoter (10). Feliciano et al found that
miR-125b acted as a tumor suppressor in breast tumorigenesis via
its direct targets glutamyl amino peptidase, casein kinase II-α,
cyclin-J and multiple epidermal growth factor-like domains 9
(11). In addition,
miR-125b-overexpressing breast cancer cells were impaired in their
anchorage-dependent growth and exhibited reduced migration and
invasion capacities (12). However,
miR-125b can also induce metastasis by targeting StAR related lipid
transfer domain containing 13 (STARD13) in MCF-7 and MDA-MB-231
breast cancer cells (13). Our
previous study demonstrated that upregulation of miR-125b conferred
a chemoresistant phenotype by targeting B-cell lymphoma 2
antagonist killer 1 (Bak1) (14), and
other previous data showed that miR-125b could maintain cancer
stem-like side population fraction (15). Circulating miR-125b expression was
associated with chemotherapeutic resistance of breast cancer
(16). Due to these different
arguments, the role of miR-125b in breast cancer requires
additional studying.
In the present study, the expression of miR-125b and
clinicopathological correlation in breast cancer tissues was
investigated by in situ hybridization (ISH). The association
between miR-125b expression and the molecular subtype of breast
cancer was analyzed. It was found that miR-125b expression is
elevated in breast cancer tissues compared to that in non-cancerous
tissues, and associated with clinical tumor node-metastasis (TNM)
stages, predicting a poor prognosis. In addition, miR-125b
expression is positively correlated with HER2 expression and
significantly associated with the tumor size, lymph node metastasis
status and TNM stage in HER2-positive breast cancer patients. The
current study provides clinical data to demonstrate the oncogenic
potential of miR-125b, particularly in HER2-positive human breast
cancer. miR-125b may be a good prognostic marker combined with HER2
in human breast cancer.
Materials and methods
Tissue samples and clinical data
In total, 221 paraffin-embedded breast cancer and 49
paraffin-embedded non-cancerous breast tissue samples obtained
between November 2001 and September 2012 at The Second Xiangya
Hospital of Central South University (Changsha, China) were used.
All the tissue samples were formed into 9 slices in a tissue
microarray, as previously described (17), with each sample in duplicate or
triplicate. Clinicopathological characteristics of breast cancer
patients were recorded including name, gender, age, occupation,
ethnicity, clinical TNM stage, recurrence, pathology diagnosis,
molecular subtype and chemoradiotherapy strategies. All the patient
information was anonymized prior to analysis. The profile of
clinicopathological characteristics of the breast cancer patients
is shown in Table I. The age of
patients ranged between 23 and 71 years. All 221 patients with
breast cancer had valid follow-up data, of which 30 cases had used
therapeutic strategies containing paclitaxel. The overall survival
(OS) was defined as the time between diagnosis and the date of
death or the date last known alive. The present study was approved
by the Committee on the Ethics of Central South University. All
individuals participating or their families provided written
informed consent.
 | Table I.Clinicopathological characteristics
of breast cancer patients. |
Table I.
Clinicopathological characteristics
of breast cancer patients.
| Characteristic | Value, % (n) |
|---|
| Age, mean ± SD | 46±0.66 |
| Gender |
|
|
Male | 0.4 (1/221) |
|
Female | 99.6 (220/221) |
| Tumor size |
|
|
T1-2 | 57.0 (126/221) |
|
T3-4 | 43.0 (95/221) |
| Nodal
metastasis |
|
|
Present | 77.8 (172/221) |
|
Absent | 22.2 (49/221) |
| Distant
metastasis |
|
|
Present | 11.8 (26/221) |
|
Absent | 88.2 (195/221) |
| TNM stage |
|
| I | 5.9 (13/221) |
| II | 58.8 (130/221) |
|
III | 24.4 (54/221) |
| IV | 10.9 (24/221) |
ISH
The ISH probe used for detecting miR-125b labeled
digoxin was designed and synthesized by Sangon Biotech Co., Ltd.
(Shanghai, China). Slices were processed using Enhanced Sensitive
ISH Detection kit I (catalogue no., MK1030; Wuhan Boster Biological
Technology, Ltd., Wuhan, China) according to the manufacturer's
protocol. The kit contains prehybridization solution, normal goat
serum, biotin-antidigoxin IgG, streptavidin-biotin-complex and
biotin-peroxidase. Slides were deparaffinized and hydrated with
xylene twice (each for 10 min), put through an ethanol gradient
(100, 95, 90, 80 and 70%, each for 5 min), and rinsed with
distilled water (dH2O). The slides then were treated
with 3% H2O2 for 15 min and washed twice with
dH2O. The slides were then treated with pepsin solution
for 10 min at 37°C, and washed three times with 0.5 M
phosphate-buffered saline (PBS) for 5 min and once with
dH2O for 10 min at room temperature. Following 3 h
incubation with prehybridization solution at 37°C, slides were
incubated with miR-125b probe (2 µg/ml; Sangon Biotech Co., Ltd.)
overnight at 55°C. On the next day, slides were incubated in 2X
saline sodium citrate (SSC) for 30 min at 37°C, washed once with
0.5X SSC for 15 min, and then washed 3 times with 0.2X SSC for 10
min. Following 30 min blockade with normal goat serum at 37°C,
slices were incubated with biotin-antidigoxin IgG for 90 min at
37°C and washed 4 times with 0.5 M PBS for 5 min, followed by
Streptavidin-Biotin-Complex and biotin-peroxidase incubation for 30
min at 37°C. Subsequent to washing in 0.5 M PBS for 20 min, the
slides were visualized with 3,3′-diaminobenzidine (Fuzhou Maixin
Biotech Co., Ltd., Fuzhou, China) for 5 min and counterstained with
hematoxylin for 90 sec. The slides were mounted and dried. Images
of slides were captured with an Olympus BX51 microscope
(magnification, ×200; Olympus Corporation, Tokyo, Japan).
Evaluation of staining
The slides were evaluated by two independent
pathologists under a light microscope (magnification, ×200; BX51;
Olympus Corporation). miR-125b staining intensity was scored as 0
(no staining was determined as negative, −), 1 (light yellow
staining was determined as weak, +), 2 (yellow staining was
determined as moderate, ++) and 3 (tan staining was determined as
strong, +++). The extent of staining was scored as 0–1.0 (0–100%).
The final staining score (0–3) was calculated as the multiplication
of the intensity score and extent score. The expression of miR-125b
was scored as high expression (≥1) or low expression (<1). To
compare the expression of miR-125b between normal and tumor
tissues, the score of miR-125b expression was normalized to the
average score of miR-125b in normal tissues.
Statistical analysis
GraphPad Prism 5 software (Graphpad Software, Inc.,
La Jolla, CA, USA) was used to perform statistical analysis. The
data are presented as the mean ± standard deviation. The difference
of miR-125b expression between breast cancer and non-cancerous
breast tissue was analyzed using Student's t-test. The
correlation between miR-125b expression and HER2 expression was
analyzed using Spearman's rank correlation analysis. The
contingency data was analyzed by using χ2 or Fisher's
exact test. The OS estimates over time were calculated using the
Kaplan-Meier method with log-rank test. A value of P<0.05 was
considered to indicate a statistically significant difference.
Results
Association between the expression of
miR-125b and clinicopathological features of breast cancer
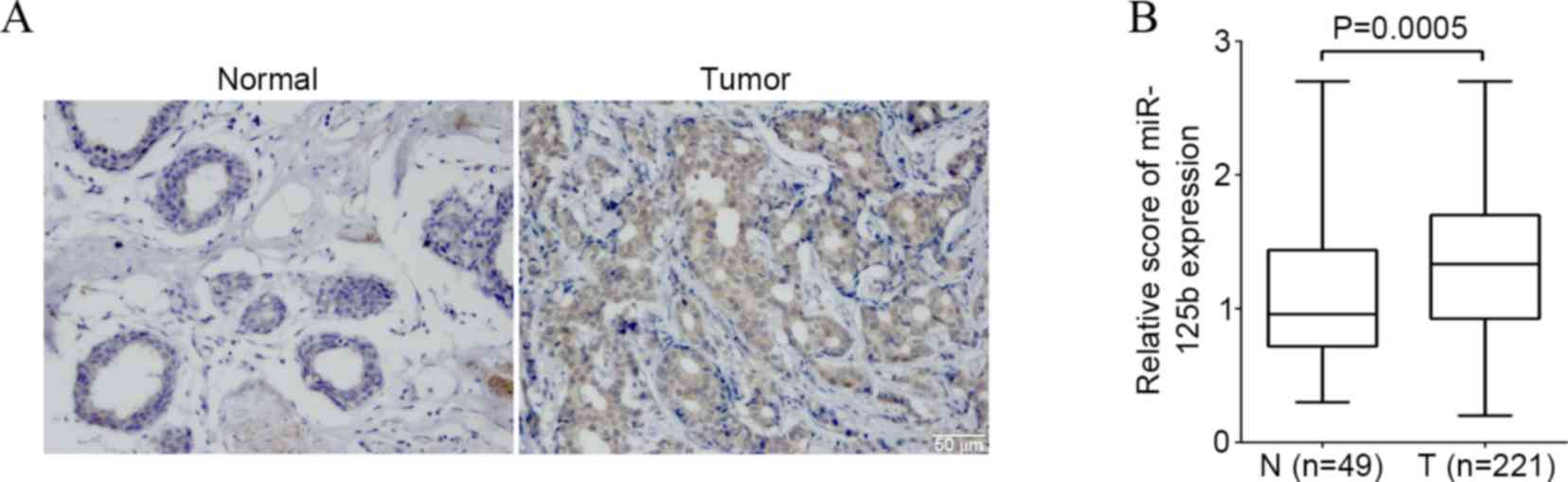
The present study detected the expression of
miR-125b in breast cancer and non-cancerous breast control tissue
by ISH. Expression of the miR-125b was found in the cytoplasm and
nucleus of breast cancer cells (Fig.
1A). The percentage of high miR-125b expression in the breast
cancer and the noncancerous breast control tissue was 68.3%
(151/221) and 57.1% (28/49) respectively. There was a significantly
higher score of the miR-125b expression in breast cancer compared
to that of non-cancerous breast control tissue (P=0.0005) (Fig. 1B).
The current study additionally investigated the
association between the expression of the miR-125b and
clinicopathological features of breast cancer including age,
gender, tumor size, lymph node metastasis status, distant
metastasis, and clinical TNM stage in a univariate χ2
test. As shown in Table II, no
significant differences were observed between miR-125b expression
and age, gender, tumor size, lymph node metastasis status or
distant metastasis of breast cancer (P>0.05). However, miR-125b
expression was significantly associated with the clinical TNM stage
(P=0.02).
 | Table II.Association between miR-125b
expression and clinicopathological characteristics in breast
cancer. |
Table II.
Association between miR-125b
expression and clinicopathological characteristics in breast
cancer.
|
| miR-125b
expression, n (%) |
|
|---|
|
|
|
|
|---|
| Variables | High (score
≥1) | Low (score
<1) | P-value |
|---|
| Total | 151 | 70 |
|
| Age, mean ± SD | 47.4±1.2 | 45.9±0.8 | 0.29 |
| Gender |
|
| 1.00 |
|
Male | 1 (100.0) | 0 (0.0) |
|
|
Female | 150 (68.2) | 70 (31.8) |
|
| Tumor size |
|
| 0.46 |
|
T1-T2 | 89 (70.6) | 37 (29.4) |
|
|
T3-T4 | 62 (65.3) | 33 (34.7) |
|
| Nodal
metastasis |
|
| 0.23 |
|
Present | 121 (70.3) | 51 (29.7) |
|
|
Absent | 30 (61.2) | 19 (38.8) |
|
| Distant
metastasis |
|
| 0.26 |
|
Present | 15 (57.7) | 11 (42.3) |
|
|
Absent | 136 (69.7) | 59 (30.3) |
|
| TNM stage |
|
| 0.02a |
|
I–II | 90 (62.9) | 53 (37.1) |
|
|
III–IV | 61 (78.2) | 17 (21.8) |
|
| ER expression, n
(%) |
|
| 0.46 |
|
Positive | 84 (66.1) | 43 (33.9) |
|
|
Negative | 67 (71.3) | 27 (28.7) |
|
| PR expression, n
(%) |
|
| 0.38 |
|
Positive | 82 (65.1) | 44 (34.9) |
|
|
Negative | 69 (72.6 | 26 (27.4) |
|
| HER2 expression, n
(%) |
|
| 0.034a |
|
Positive | 105 (73.4) | 38 (26.6) |
|
|
Negative | 46 (59.0) | 32 (41.0) |
|
High miR-125b expression has a poor
prognosis in HER2-positive breast cancer patients
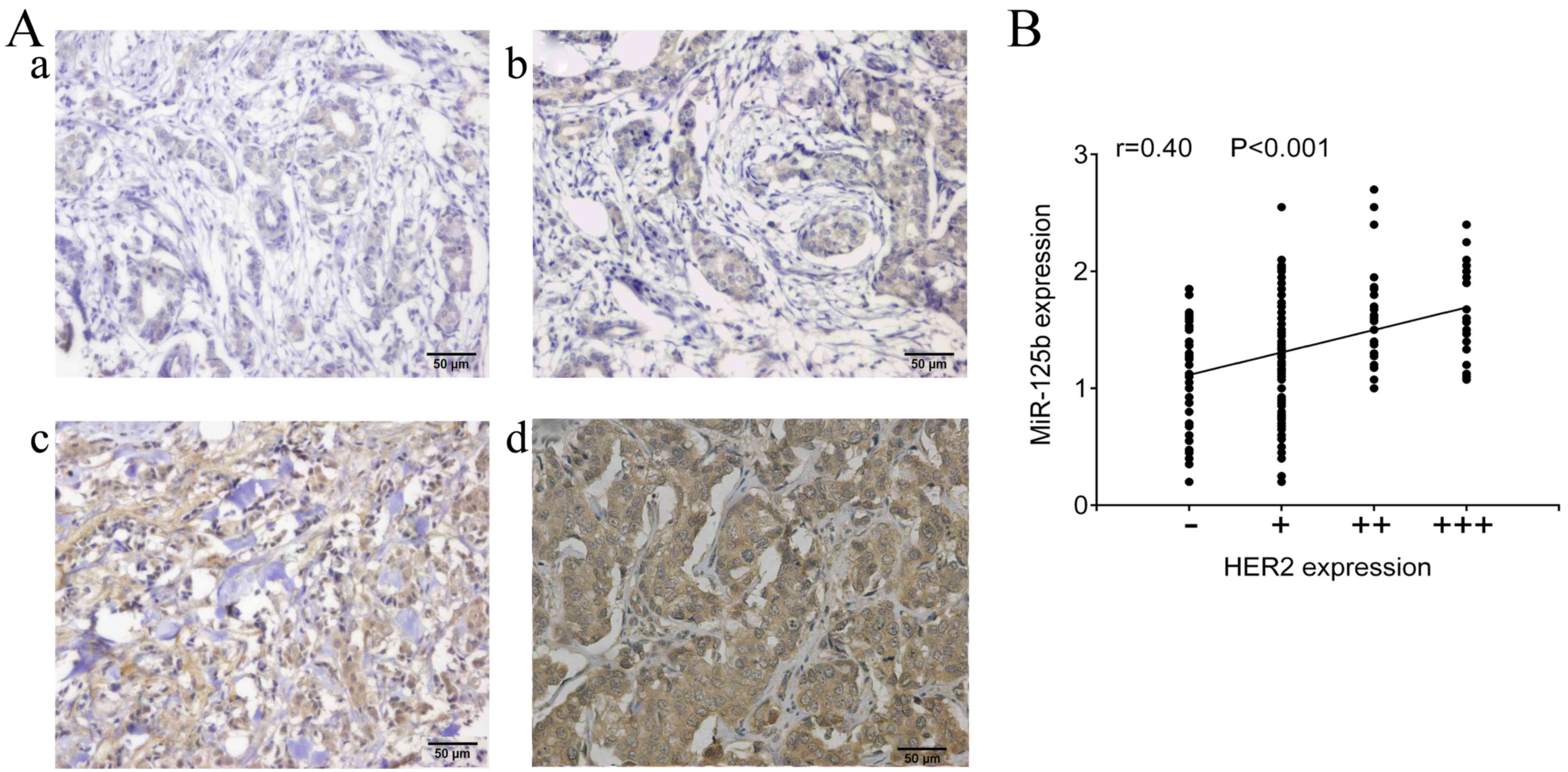
As shown in Table II,
although no significant correlation was identified between the
expression of miR-125b and expression of ER and PR (P>0.05),
there was a significant association between expression of miR-125b
and HER2 in breast cancer patients (P=0.034). In addition, the
expression of miR-125b was positively correlated with the HER2
expression (r=0.4, P<0.001; Fig. 2A
and B).
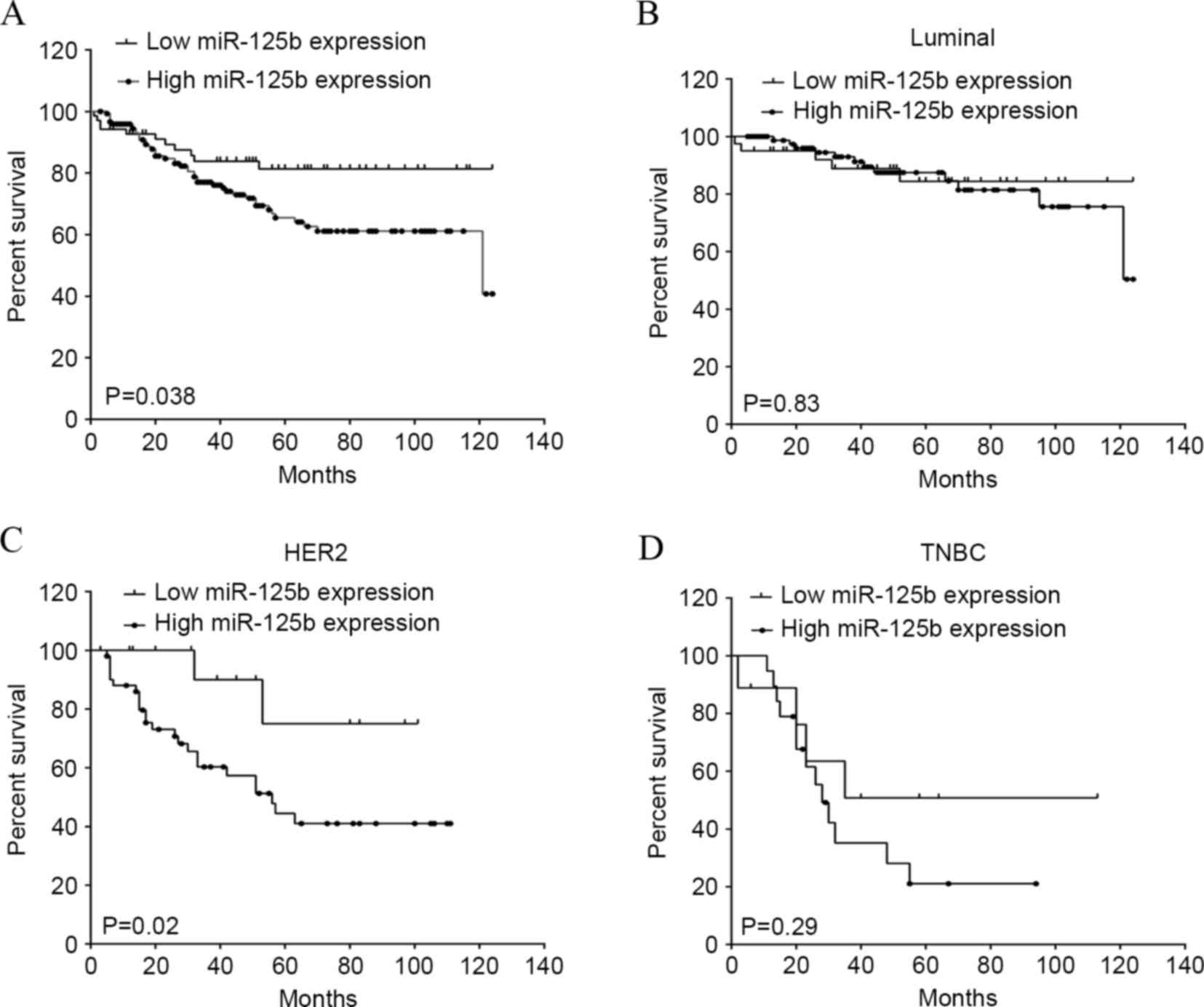
The present study additionally analyzed the
association between miR-125b expression and clinical outcomes. All
221 breast cancer patients were included in the survival curves.
The median OS time was 40 months, with a range of 2–124 months. The
OS rate of patients with high miR-125b expression was significantly
decreased compared to the survival of patients with low miR-125b
expression (P=0.038; hazard ratio, 0.55; 95% CI, 0.31–0.96;
Fig. 3A). To investigate the
association between miR-125b and molecular subtypes of breast
cancer, all 221 breast cancer cases were classified into the
following groups: Luminal (high miR-125b expression, n=81; low
miR-125b expression, n=38), identified as ER+ and/or
PR+; HER2 (high miR-125b expression, n=51; low miR-125b
expression, n=20), identified as ER/PR− and
HER2+; and TNBC (high miR-125b expression, n=19; low
miR-125b expression, n=12), identified as ER/PR− and
HER2− (18). There was no
significant difference between the OS rate of patients with high
miR-125b expression and that of patients with low miR-125b
expression in the molecular luminal (Fig.
3B) and TNBC (Fig. 2D) subtypes.
Notably, the OS rate of patients with high miR-125b expression was
significantly reduced compared to the survival of patients with low
miR-125b expression in molecular subtypes of HER2 (P=0.02; hazard
ratio, 0.38; 95% CI 0.16–0.92; Fig.
3C), indicating an association between miR-125b and HER2
receptor expression. Thus, the current study additionally analyzed
the association between the expression of miR-125b and
clinicopathological features of HER2-positive breast cancer. A
total of 143 patients withHER2-positive breast cancer were
included. As shown in Table III, no
significant differences were observed between expression of
miR-125b and age, gender and distant metastasis of HER2-positive
breast cancer patients (P>0.05). However, miR-125b expression
was significantly associated with the tumor size (P=0.03), lymph
node metastasis status (P=0.03) and TNM stage (P=0.02).
 | Table III.Association between miR-125b
expression and clinicopathological characteristics in HER2-positive
breast cancer. |
Table III.
Association between miR-125b
expression and clinicopathological characteristics in HER2-positive
breast cancer.
|
| miR-125b
expression, n (%) |
|
|---|
|
|
|
|
|---|
| Variables | High (score
≥1) | Low (score
<1) | P-value |
|---|
| Total, n | 105 | 38 |
|
| Age, mean ± SD | 46.7±1.06 | 48.2±1.86 | 0.49 |
| Gender |
|
|
|
|
Male | 0 (0.0) | 0 (0.0) | 1.00 |
|
Female | 105 (73.4) | 38 (26.6) |
|
| Tumor size |
|
| 0.03a |
|
T1-2 | 59 (67.0) | 29 (33.0) |
|
|
T3-4 | 46 (83.6) | 9 (16.4) |
|
| Nodal
metastasis |
|
| 0.03a |
|
Present | 88 (77.9) | 25 (22.1) |
|
|
Absent | 17 (56.7) | 13 (43.3) |
|
| Distant
metastasis |
|
| 0.72 |
|
Present | 9 (69.2) | 4 (30.8) |
|
|
Absent | 96 (73.8) | 34 (26.2) |
|
| TNM stage |
|
| 0.02a |
|
I–II | 57 (66.3) | 29 (33.7) |
|
|
III–IV | 48 (84.2) | 9 (15.8) |
|
High miR-125b expression predicts a
poor prognosis in breast cancer patients treated with
paclitaxel
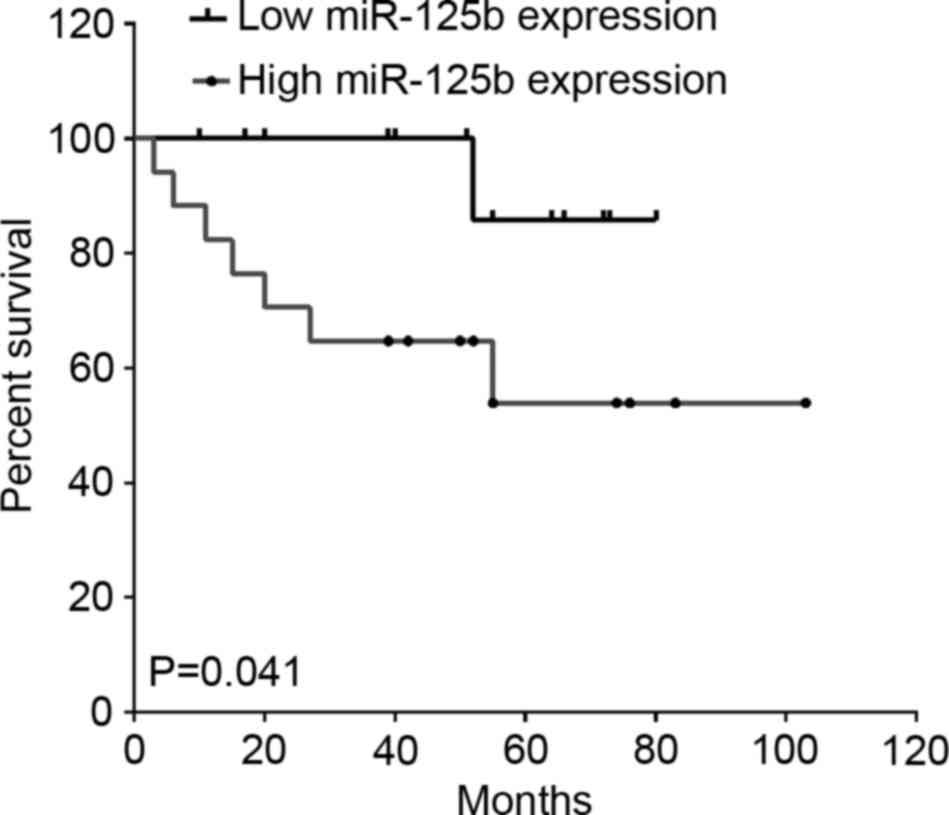
To investigate the association between miR-125b and
paclitaxel treatment in breast cancer, among the 221 breast cancer
patients, 30 cases that used therapeutic strategies containing
paclitaxel were included. As shown in Fig. 4, high miR-125b expression indicated a
lower OS rate in patients treated with paclitaxel compared with
those of patients with low miR-125b expression (P=0.041; hazard
ratio, 0.23; 95% CI, 0.06–0.95; Fig.
4).
Discussion
miR-125b is located at chromosome 11q24 and
chromosome 21q21, the so-called fragile sites, which are commonly
deleted in lung, ovary and cervical cancer, indicating a functional
loss of miR-125b in these tumor types (19). It has been demonstrated that miR-125b
is dysregulated in a broad variety of tumors. It is downregulated
in head and neck tumors, oral squamous cell carcinomas,
osteosarcomas and gliomas (20–23).
Hypermethylation in the promoter regions of miR-125b appears to
block miR-125b expression levels in ovarian cancer (24) and breast cancer (10). However, enhanced miR-125b expression
was also observed in colorectal cancer, leukemia, gastric,
follicular and pancreatic cancers and certain brain tumor-derived
glioma cell lines, which are associated with poor prognoses
(25–28). In the present study, it was found that
miR-125b expression was significantly increased in breast cancer
tissues compared to those of noncancerous tissues, and high
miR-125b expression indicated a poor prognosis in breast cancer
patients. Tang et al demonstrated that upregulation of
miR-125b was able to activate the metastatic activities of breast
cancer cells in vivo and in vitro by inducing breast
cancer cells to obtain epithelial and mesenchymal characteristics
while regulating the reorganization of actin cytoskeleton through
the STARD13-Ras homologue gene family member A-Rho-associated
protein kinase signaling pathway (13). Consistently, the present results
showed that miR-125b expression was correlated with clinical TNM
stages. miR-125b-mediated breast cancer metastasis appears to
account for the elevated stem cell-like side population and
enhanced cancer stem cells properties (14). However, the upregulation of miR-125b
is regulated by mechanisms that are not well understood in breast
cancer. In B-cell progenitor acute lymphoblastic leukemia,
translocation t(11;14)(q24;q32) leads to a significant upregulation
of miR-125b (29,30). It is possible that chromosomal
translocations may lead to aberrantly high miR-125b expression in
breast and other types of tumors. Additional studies are required
to determine the precise causes of aberrant miR-125b expression in
cancer.
The present study additionally analyzed the
association between miR-125b expression and ER, PR and HER2.
miR-125b expression was positively correlated with HER2, but not ER
and PR. Notably, high miR-125b expression was significantly
correlated with tumor size and TNM stages in HER2-positive breast
cancer patients, along with a poor prognosis. Although studies have
shown that HER2 is inversely correlated with miR-125b in gastric
adenocarcinomas and ovarian cancer (31,32), and
acts as a target of miR-125b in small cell lung cancer (33) and endometrial cancer (34), there is no miR-125b-mediated
downregulation of HER2 in miR-125b-transfected LNCaP and cds1
prostate cancer cells (35).
Additionally our previous study did not identify downregulation of
HER2 by miR-125b in breast cancer BT-474, BT-474M1 and SKBr3 cells
(15). A reasonable explanation is
that miRNAs exhibit their functions dependent on their diverse
target genes in a cell type-specific and phenotype-specific manner
(36). Due to the high heterogeneity
of breast cancer, miR-125b may target different genes in various
breast cancer cell lines. In addition, miR-125b can directly
interact with the tumor suppressor gene p53, and target it in
humans and zebrafish. Other molecules belonging to the p53 network
such as Bak1, puma and cell cycle regulators are also targeted by
miR-125b (37). Thus, miR-125b can
interfere with oncogenic signaling by inhibiting key components of
the p53 network, acting as an oncogenic miRNA.
In addition, miR-125b also plays a role in
chemoresistance in breast cancer. It was previously found that
overexpression of miR-125b caused a marked inhibition of
paclitaxel-induced apoptosis and increased resistance to paclitaxel
by targeting Bak1 in breast cancer cells (15). The present study shows that high
miR-125b expression is a marker of poor prognosis in breast cancer
patients that are treated with paclitaxel. Wang et al
(16) showed that high miR-125b
expression had an increased percentage of proliferating cells and
decreased percentage of apoptotic cells subsequent to neoadjuvant
chemotherapy in breast cancer patients, and reducing the miR-125b
level sensitized breast cancer cells to chemotherapy. Previously,
Vilquin et al (38)
demonstrated that ectopic overexpression of miR-125b is sufficient
to confer MCF-7 cell resistance to letrozole and anastrozole by
targeting and activating the AKT/mammalian target of rapamycin
pathway, which appears to be estrogen-independent. In addition,
elevated miR-125b expression levels are a novel marker for poor
prognosis in letrozole resistant breast cancer. Furthermore,
elevated miR-125b levels in circulation may be a marker for early
breast cancer detection (7).
Clinically, elevated expression of miR-125b is associated with
non-pathological complete response in breast cancer patients that
received taxane-anthracycline-based neoadjuvant chemotherapy
(39).
Overall, the present study provides evidence that
elevated miR-125b expression predicts a poor prognosis, as well as
a poor clinical responsiveness of paclitaxel-based neoadjuvant
chemotherapy, and is associated with tumor size and TNM stage in
HER2-positive breast cancer. Thus, identification of miR-125b may
be a potential molecular biomarker for prediction of the clinical
outcome in breast cancer patients, particularlyHER2-positive cases
that receive paclitaxel-based neoadjuvant chemotherapy.
Acknowledgements
This study was supported by grants obtained from the
National Natural Science Foundation of China (grant nos. 81328019
and 81572748) and the Natural Science Foundation of Hunan Province
(grant no. 2015JJ2148).
References
|
1
|
Torre LA, Bray F, Siegel RL, Ferlay J,
LortetTieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Eroles P, Bosch A, Pérez-Fidalgo JA and
Lluch A: Molecular biology in breast cancer: Intrinsic subtypes and
signaling pathways. Cancer Treat Rev. 38:698–707. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Goldhirsch A, Wood WC, Coates AS, Gelber
RD, Thürlimann B and Senn HJ: Panel members: Strategies for
subtypes-dealing with the diversity of breast cancer: Highlights of
the St. Gallen international expert consensus on the primary
therapy of early breast cancer 2011. Ann Oncol. 22:1736–1747. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Byler S, Goldgar S, Heerboth S, Leary M,
Housman G, Moulton K and Sarkar S: Genetic and epigenetic aspects
of breast cancer progression and therapy. Anticancer Res.
34:1071–1077. 2014.PubMed/NCBI
|
|
5
|
Kontorovich T, Levy A, Korostishevsky M,
Nir U and Friedman E: Single nucleotide polymorphisms in miRNA
binding sites and miRNA genes as breast/ovarian cancer risk
modifiers in Jewish high-risk women. Int J Cancer. 127:589–597.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Baffa R, Fassan M, Volinia S, O'Hara B,
Liu CG, Palazzo JP, Gardiman M, Rugge M, Gomella LG, Croce CM and
Rosenberg A: MicroRNA expression profiling of human metastatic
cancers identifies cancer gene targets. J Pathol. 219:214–221.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Matamala N, Vargas MT, GonzalezCampora R,
Minambres R, Arias JI, Menendez P, AndresLeon E, GomezLopez G,
Yanowsky K, CalveteCandenas J, et al: Tumor microRNA expression
profiling identifies circulating microRNAs for early breast cancer
detection. Clin Chem. 61:1098–1106. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Banzhaf-Strathmann J and Edbauer D: Good
guy or bad guy: The opposing roles of microRNA 125b in cancer. Cell
Commun Signal. 12:302014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ferracin M, Bassi C, Pedriali M, Pagotto
S, D'Abundo L, Zagatti B, Corrà F, Musa G, Callegari E, Lupini L,
et al: miR-125b targets erythropoietin and its receptor and their
expression correlates with metastatic potential and ERBB2/HER2
expression. Mol Cancer. 12:1302013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhang Y, Yan LX, Wu QN, Du ZM, Chen J,
Liao DZ, Huang MY, Hou JH, Wu QL, Zeng MS, et al: miR-125b is
methylated and functions as a tumor suppressor by regulating the
ETS1 proto-oncogene in human invasive breast cancer. Cancer Res.
71:3552–3562. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Feliciano A, Castellvi J, ArteroCastro A,
Leal JA, Romagosa C, HernandezLosa J, Peg V, Fabra A, Vidal F,
Kondoh H, et al: miR-125b acts as a tumor suppressor in breast
tumorigenesis via its novel direct targets ENPEP, CK2-α, CCNJ, and
MEGF9. PLoS One. 8:e762472013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Scott GK, Goga A, Bhaumik D, Berger CE,
Sullivan CS and Benz CC: Coordinate suppression of ERBB2 and ERBB3
by enforced expression of micro-RNA miR-125a or miR-125b. J Biol
Chem. 282:1479–1486. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tang F, Zhang R, He Y, Zou M, Guo L and Xi
T: MicroRNA-125b induces metastasis by targeting STARD13 in MCF-7
and MDA-MB-231 breast cancer cells. PLoS One. 7:e354352012.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang HJ, Guo YQ, Tan G, Dong L, Cheng L,
Li KJ, Wang ZY and Luo HF: miR-125b regulates side population in
breast cancer and confers a chemoresistant phenotype. J Cell
Biochem. 114:2248–2257. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhou M, Liu Z, Zhao Y, Ding Y, Liu H, Xi
Y, Xiong W, Li G, Lu J, Fodstad O, et al: MicroRNA-125b confers the
resistance of breast cancer cells to paclitaxel through suppression
of pro-apoptotic Bcl-2 antagonist killer 1 (Bak1) expression. J
Biol Chem. 285:21496–21507. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang H, Tan G, Dong L, Cheng L, Li K, Wang
Z and Luo H: Circulating MiR-125b as a marker predicting
chemoresistance in breast cancer. PLoS One. 7:e342102012.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Fan SQ, Ma J, Zhou J, Xiong W, Xiao BY,
Zhang WL, Tan C, Li XL, Shen SR, Zhou M, et al: Differential
expression of Epstein-Barr virus-encoded RNA and several
tumor-related genes in various types of nasopharyngeal epithelial
lesions and nasopharyngeal carcinoma using tissue microarray
analysis. Hum Pathol. 37:593–605. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Sorlie T, Tibshirani R, Parker J, Hastie
T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, et
al: Repeated observation of breast tumor subtypes in independent
gene expression data sets. Proc Natl Acad Sci USA. 100:8418–8423.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Calin GA, Sevignani C, Dumitru CD, Hyslop
T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M,
et al: Human microRNA genes are frequently located at fragile sites
and genomic regions involved in cancers. Proc Natl Acad Sci USA.
101:2999–3004. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Nakanishi H, Taccioli C, Palatini J,
FernandezCymering C, Cui R, Kim T, Volinia S and Croce CM: Loss of
miR-125b-1 contributes to head and neck cancer development by
dysregulating TACSTD2 and MAPK pathway. Oncogene. 33:702–712. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Henson BJ, Bhattacharjee S, O'Dee DM,
Feingold E and Gollin SM: Decreased expression of miR-125b and
miR-100 in oral cancer cells contributes to malignancy. Genes
Chromosomes Cancer. 48:569–582. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Liu LH, Li H, Li JP, Zhong H, Zhang HC,
Chen J and Xiao T: miR-125b suppresses the proliferation and
migration of osteosarcoma cells through down-regulation of STAT3.
Biochem Biophys Res Commun. 416:31–38. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Smits M, Wurdinger T, van het Hof B,
Drexhage JA, Geerts D, Wesseling P, Noske DP, Vandertop WP, de
Vries HE and Reijerkerk A: Myc-associated zinc finger protein (MAZ)
is regulated by miR-125b and mediates VEGF-induced angiogenesis in
glioblastoma. FASEB J. 26:2639–2647. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
He J, Xu Q, Jing Y, Agani F, Qian X,
Carpenter R, Li Q, Wang XR, Peiper SS, Lu Z, et al: Reactive oxygen
species regulate ERBB2 and ERBB3 expression via miR-199a/125b and
DNA methylation. EMBO Rep. 13:1116–1122. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Bousquet M, Harris MH, Zhou B and Lodish
HF: MicroRNA miR-125b causes leukemia. Proc Natl Acad Sci USA.
107:21558–21563. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Willimott S and Wagner SD: miR-125b and
miR-155 contribute to BCL2 repression and proliferation in response
to CD40 ligand (CD154) in human leukemic B-cells. J Biol Chem.
287:2608–2617. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Xia HF, He TZ, Liu CM, Cui Y, Song PP, Jin
XH and Ma X: MiR-125b expression affects the proliferation and
apoptosis of human glioma cells by targeting Bmf. Cell Physiol
Biochem. 23:347–358. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Le MT, Teh C, ShyhChang N, Xie H, Zhou B,
Korzh V, Lodish HF and Lim B: MicroRNA-125b is a novel negative
regulator of p53. Genes Dev. 23:862–876. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Bousquet M, Quelen C, Rosati R, MansatDe
MV, La Starza R, Bastard C, Lippert E, Talmant P,
Lafage-Pochitaloff M, Leroux D, et al: Myeloid cell differentiation
arrest by miR-125b-1 in myelodysplastic syndrome and acute myeloid
leukemia with the t(2;11)(p21;q23) translocation. J Exp Med.
205:2499–2506. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chapiro E, Russell LJ, Struski S, Cavé H,
Radford-Weiss I, Valle VD, Lachenaud J, Brousset P, Bernard OA,
Harrison CJ and Nguyen-Khac F: A new recurrent translocation
t(11;14)(q24;q32) involving IGH@ and miR-125b-1 in B-cell
progenitor acute lymphoblastic leukemia. Leukemia. 24:1362–1364.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Fassan M, Pizzi M, Realdon S, Balistreri
M, Guzzardo V, Zagonel V, Castoro C, Mastracci L, Farinati F, Nitti
D, et al: The HER2-miR125a5p/miR125b loop in gastric and esophageal
carcinogenesis. Hum Pathol. 44:1804–1810. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
He J, Jing Y, Li W, Qian X, Xu Q, Li FS,
Liu LZ, Jiang BH and Jiang Y: Roles and mechanism of miR-199a and
miR-125b in tumor angiogenesis. PLoS One. 8:e566472013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Yagishita S, Fujita Y, Kitazono S, Ko R,
Nakadate Y, Sawada T, Kitamura Y, Shimoyama T, Maeda Y, Takahashi
F, et al: Chemotherapy-regulated microRNA-125-HER2 pathway as a
novel therapeutic target for trastuzumab-mediated cellular
cytotoxicity in small cell lung cancer. Mol Cancer Ther.
14:1414–1423. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Shang C, Lu YM and Meng LR: MicroRNA-125b
down-regulation mediates endometrial cancer invasion by targeting
ERBB2. Med Sci Monit. 18:BR149–BR155. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Shi XB, Xue L, Yang J, Ma AH, Zhao J, Xu
M, Tepper CG, Evans CP, Kung HJ and White RW deVere: An
androgen-regulated miRNA suppresses Bak1 expression and induces
androgen-independent growth of prostate cancer cells. Proc Natl
Acad Sci USA. 104:19983–19988. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Schwarzenbacher D, Balic M and Pichler M:
The role of microRNAs in breast cancer stem cells. Int J Mol Sci.
14:14712–14723. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Le MT, ShyhChang N, Khaw SL, Chin L, Teh
C, Tay J, O'Day E, Korzh V, Yang H, Lal A, et al: Conserved
regulation of p53 network dosage by microRNA-125b occurs through
evolving miRNA-target gene pairs. PLoS Genet. 7:e10022422011.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Vilquin P, Donini CF, Villedieu M, Grisard
E, Corbo L, Bachelot T, Vendrell JA and Cohen PA: MicroRNA-125b
upregulation confers aromatase inhibitor resistance and is a novel
marker of poor prognosis in breast cancer. Breast Cancer Res.
17:132015. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zheng Y, Li S, Boohaker RJ, Liu X, Zhu Y,
Zhai L, Li H, Gu F, Fan Y, Lang R, et al: A microRNA expression
signature in taxane-anthracycline-based neoadjuvant chemotherapy
response. J Cancer. 6:671–677. 2015. View Article : Google Scholar : PubMed/NCBI
|