Introduction
The human epidermal growth factor receptor 2 (HER2)
gene (also known as erbB-2 or neu) is a member of the epidermal
growth factor receptor (EGFR) family (1,2). Located
on chromosome 17q21, it codes for a transmembrane glycoprotein with
tyrosine kinase activity (3). HER2 is
markedly homologous with EGFR, which serves an important role in
the regulation of cell growth, differentiation and invasion
(2,3).
It is well documented that HER2 amplification and/or overexpression
is associated with a worse clinical outcome in breast cancer
(4–9).
HER2-targeted therapy, i.e., using trastuzumab, markedly improves
the survival outcome of patients with HER2-positive breast cancer
(10–13).
We recently reported that the HER2 Ile655Val
polymorphism is significantly associated with the survival outcome
of patients with HER2-positive breast cancer, indicating that
patients with the Val variant exhibit breast cancer with an
aggressive phenotype, but are more sensitive to trastuzumab
treatment (14). In the present
study, another common polymorphism (HER2 Pro1170Ala) of HER2 was
investigated. This polymorphism is located in the tail region of
the HER2 protein, and codes for either Pro (CCC) or Ala (GCC) at
position 1170 (15). Previous studies
have suggested that this polymorphism affects HER2 activity by
altering tail construction and function (2,16–18). A previous study indicated that the
HER2 Pro1170Ala variant is associated with an increased risk of
developing lung cancer in Korean women (19). However, no association between the
HER2 Pro1170Ala polymorphism and the risk of developing breast or
endometrial cancer was identified in further studies (15,20–23). A
recent study identified that the Pro1170Ala polymorphism is
associated with a risk of trastuzumab cardiotoxicity (24). Currently, to the best of our
knowledge, no investigation has been conducted into whether the
HER2 Pro1170Alapolymorphism affects breast cancer survival.
Therefore, in the present study, the incidence of
the HER2 Pro1170Ala polymorphism was determined in a cohort of
3,305 women with operable primary breast cancer, and the
association between the HER2 Pro1170Ala polymorphism and survival
was investigated; it was further determined whether the HER2
Pro1170Ala polymorphism was associated with survival in
HER2-positive and HER2-negative patients, respectively.
Materials and methods
Patients
A total of 3,430 female patients with operable
primary breast cancer (stages I–III) were treated at the Breast
Center, Peking University Cancer Hospital (Beijing, China) between
January 2005 and October 2011. Of these 3,430 patients, 125 were
excluded from the present study, as the HER2 Pro1170Ala genotype
was not identified due to the poor quality of the DNA samples of 77
patients and as survival data were not available for 48 patients.
Consequently, 3,305 patients were included in the present study.
Ages at diagnosis of the patients ranged between 21 and 90 years,
with a median age of 50 years. The stage of the tumors was
classified according to the tumor-node-metastasis classification of
the Union Internationale Contre Le Cancer (25,26). Tumor
size was defined as the maximum tumor diameter measured using
ultrasound at the time of diagnosis. Tumor grade, tumor size,
estrogen receptor (ER) status, progesterone receptor (PR) status
and adjuvant therapy were obtained from the review of medical
records, and are presented in Table
I. This study was conducted in accordance with the ethical
principles of the Declaration of Helsinki and approved by the
Research and Ethics Committee of Peking University Cancer Hospital.
The project number was 2011KT12. All patients provided written
informed consent.
 | Table I.Association between the HER2
Prol170A1a genotype and clinicopathological characteristics. |
Table I.
Association between the HER2
Prol170A1a genotype and clinicopathological characteristics.
| Characteristic | Overall | Pro/Pro, n (%) | Pro/Ala, n (%) | Ala/Ala, n (%) | P-value |
|---|
| Total | 3,305 | 955 (29) | 1,679 (51) | 671 (20) | 0.175a |
| Age at diagnosis,
years |
|
|
|
| 0.771b |
| ≤40 | 593 | 174 (18) | 305 (18) | 114 (17) |
|
|
>40 | 2,712 | 781 (82) | 1,374 (82) | 557 (83) |
|
| Tumor size, cm |
|
|
|
| 0.547b |
| ≥2 | 1,733 | 499 (55) | 872 (54) | 362 (57) |
|
|
<2 | 1,412 | 403 (45) | 734 (46) | 275 (43) |
|
|
Unknown | 160 |
|
|
|
|
| Tumor grade |
|
|
|
| 0.312b |
| I | 414 | 130 (16) | 199 (14) | 85 (15) |
|
| II | 2,045 | 583 (72) | 1,033 (72) | 429 (73) |
|
| III | 381 | 99 (12) | 210 (14) | 72 (12) |
|
|
Unknown | 465 |
|
|
|
|
| Lymph nodes |
|
|
|
| 0.239b |
|
Positive | 1,154 | 323 (35) | 580 (36) | 251 (39) |
|
|
Negative | 2,030 | 605 (65) | 1,031 (64) | 394 (61) |
|
|
Unknown | 121 |
|
|
|
|
| ER status |
|
|
|
| 0.699b |
|
Positive | 2,290 | 662 (72) | 1,154 (70) | 474 (72) |
|
|
Negative | 934 | 261 (28) | 486 (30) | 187 (28) |
|
|
Unknown | 81 |
|
|
|
|
| PR status |
|
|
|
| 0.636b |
|
Positive | 2,040 | 594 (64) | 1,027 (62) | 419 (63) |
|
|
Negative | 1185 | 328 (36) | 615 (38) | 242 (37) |
|
|
Unknown | 80 |
|
|
|
|
| HER2 status |
|
|
|
| 0.941b |
|
Positive | 728 | 205 (23) | 370 (23) | 153 (23) |
|
|
Negative | 2,442 | 696 (77) | 1247 (77) | 499 (77) |
|
|
Unknown | 135 |
|
|
|
|
| Trastuzumab
use |
|
|
|
| 0.374b |
|
Yes | 146 | 44 (5) | 79 (5) | 23 (3) |
|
| No | 3,159 | 911 (95) | 1,600 (95) | 648 (97) |
|
| Adjuvant
therapy |
|
|
|
| 0.495b |
| C | 1,133 | 313 (33) | 593 (35) | 227 (34) |
|
| E | 623 | 189 (20) | 316 (19) | 118 (18) |
|
|
C+E | 1,248 | 357 (37) | 620 (37) | 271 (40) |
|
|
None | 301 | 96 (10) | 150 (9) | 55 (8) |
|
HER2 Pro1170Ala genotyping
Genomic DNA was isolated from peripheral blood
leukocytes using phenol-chloroform. In brief, peripheral blood
leukocytes were mixed with equal volumes of a phenol-chloroform
mixture to remove protein contaminants, then precipitated with 100%
ethanol. Amplification of DNA fragments was performed using
polymerase chain reaction (PCR) using a Gene Cycler™ thermocycler
(Bio-Rad Laboratories, Inc., Hercules, CA, USA) in a 20-µl solution
containing 30 ng genomic DNA, 2.5 mM MgCl2, 0.8 mM
dNTPs, 1.0X PCR buffer, 0.5 µM forward and reverse primers, and
1.25 units AmpliTaq DNA polymerase (Promega Corporation, Madison,
WI, USA). The reaction conditions were an initial 94°C for 5 min to
activate Taq DNA polymerase, followed by 35 cycles of denaturation
at 94°C for 30 sec, annealing at 60°C for 30 sec and extension at
72°C for 45 sec, and a final extension at 72°C for 10 min. The
forward primer sequence was 5′-CCTGCCCTCTGAGACTGATG-3′ and the
reverse primer sequence was 5′-GTTCCTCTTCCAACGAGGCT-3′. The HER2
Pro1170Ala genotype was detected by direct sequencing in our
laboratory. All fragments were sequenced using a BigDye Terminator
Cycle Sequencing kit and ABI 3730 automated sequencer (both Applied
Biosystems; Thermo Fisher Scientific, Inc., Waltham, MA, USA). HER2
gene (ID 2064) was used as the reference gene with primer sequences
5′-ATGGAGCTGGCGGCCTTGT-3′. In order to avoid potential
contamination, each set of PCR contained a negative and 3 positive
controls, a negative control without DNA template and 3 positive
controls (known Pro/Pro, Pro/Ala or Ala/Ala genotype, respectively)
performed simultaneously. A total of 30% of the cases were
genotyped in duplicate and results were fully concordant.
Assessment of HER2 status
HER2 status was obtained from a review of pathology
reports. The HER2 status was determined using immunostaining
according to a standard method (27):
A score of 0 and 1+ was considered negative and score of 3+ was
considered positive; a score of 2+ was further evaluated using
fluorescence in situ hybridization using a Vysis CLL FISH
Probe kit (Abbott Laboratories, Abbott Park, IL, USA), according to
the manufacturer's protocol.
Statistical analysis
Statistical analysis was performed using SPSS
software for Windows (version 20.0; IBM SPSS, Armonk, NY, USA). The
associations between the HER2 Pro1170Ala genotype variants and
clinicopathological characteristics in the entire cohort were
evaluated using Pearson's χ2 test. Survival curves were
derived from Kaplan-Meier estimator analysis and the differences
between the curves were compared using log-rank tests.
Recurrence-free survival (RFS) was defined as the time between the
date of pathological diagnosis and the date of locoregional
recurrence or metastasis, distant metastasis or mortality from
breast cancer. Distant recurrence-free survival (DRFS) was defined
as the time between diagnosis and the occurrence of distant
metastasis or mortality, for which breast cancer was the primary or
underlying cause. Multivariate survival analysis was performed to
identify independent prognostic variables in the patients with
HER2-negative breast cancer using tumor grade, tumor size, lymph
node status, ER status, PR status, whether adjuvant therapy was
used or not and the HER2 Pro1170Ala genotype as covariates. All
statistical tests were two-sided, and P<0.05 was considered to
indicate a statistically significant difference.
Results
Patient characteristics
The clinicopathological characteristics of the 3,305
patients examined are presented in Table
I. The incidence of the HER2 Pro1170Ala genotype was determined
in these 3,305 patients: 29% (955/3,305) were homozygous for the
Pro/Pro genotype, 51% (1,679/3,305) were heterozygous for the
Pro/Ala genotype and 20% (671/3,305) were homozygous for the
Ala/Ala genotype. The frequency of the variants conformed to the
Hardy-Weinberg equilibrium (P=0.175).
No significant association between the Pro1170Ala
polymorphism and age at diagnosis, tumor size, tumor grade, lymph
node status, ER status, PR status, HER2 status, adjuvant therapy or
trastuzumab treatment was identified in this cohort of 3,305
patients (Table I).
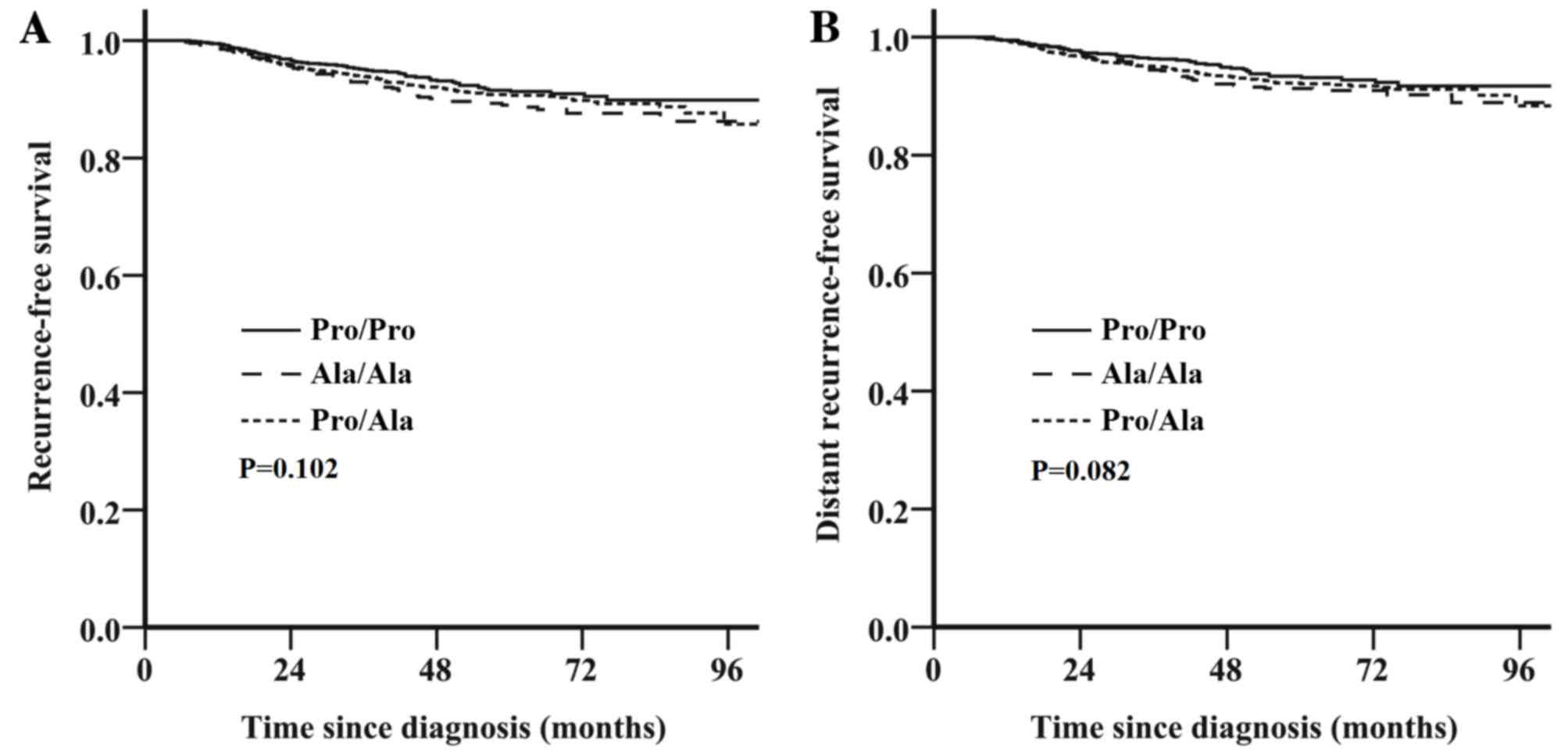
Survival outcome is not associated
with the HER2 Pro1170Ala genotype for the entire cohort
Follow-up data were available for all 3,305
patients; the median follow-up period was 53 months (range, 2–110
months). The estimated 5-year RFS and DRFS in the 3,305 patients
were 90.9% [95% confidence interval (CI), 89.7–92.1) and 92.4% (95%
CI, 91.4–93.4), respectively. No significant difference in survival
was identified between the HER2 Pro1170Ala genotypes in the entire
cohort of 3,305 patients; patients with the Pro/Ala or Ala/Ala
genotype exhibited a similar survival outcome to those with the
Pro/Pro genotype [RFS: unadjusted hazard ratio (HR), 1.26; 95% CI,
0.96–1.65; P=0.102; Fig. 1A; DRFS:
unadjusted HR, 1.31; 95% CI, 0.97–1.79; P=0.082; Fig. 1B].
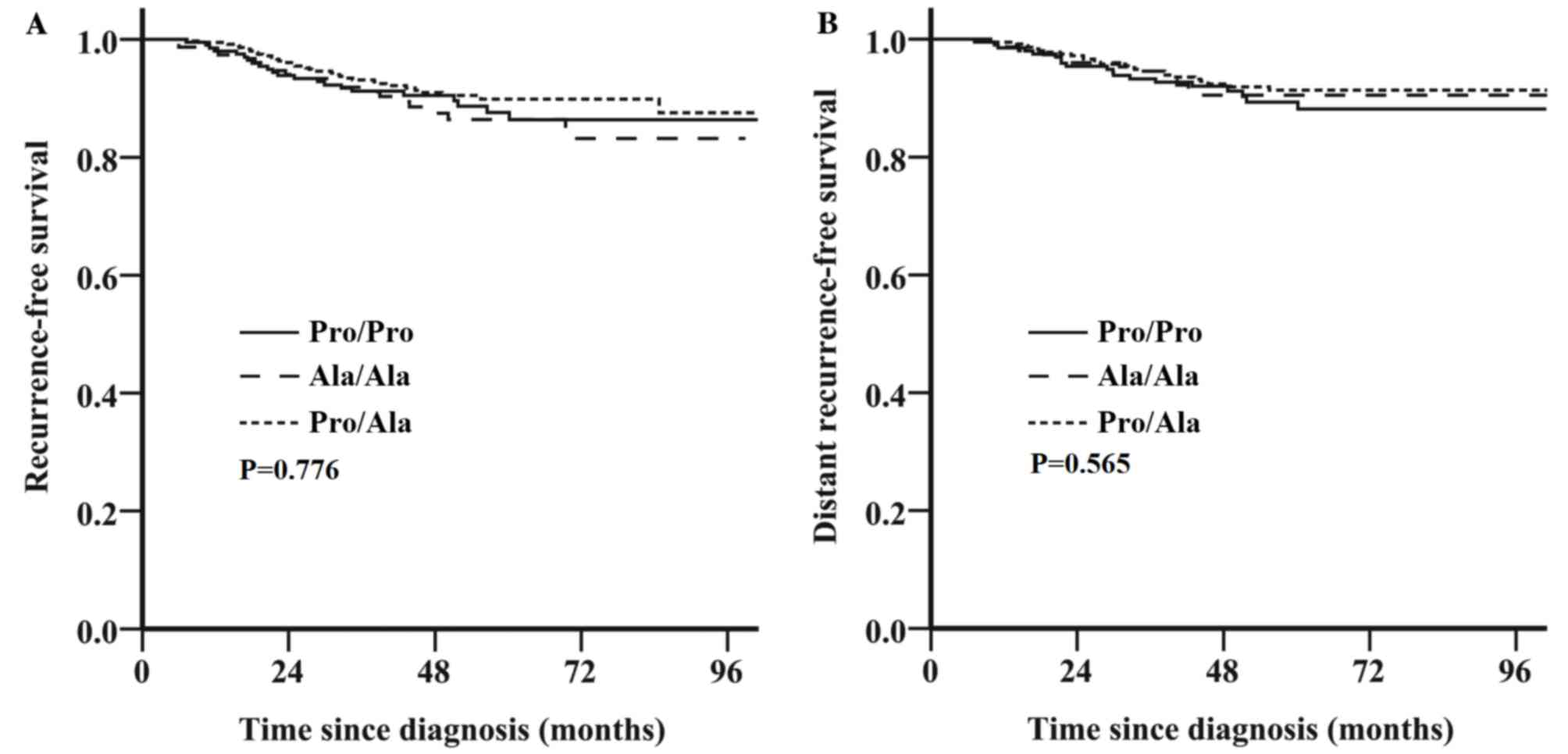
Survival outcome is associated with
the HER2 Pro1170Ala genotype for HER2-negative breast cancer, but
not HER2-positive breast cancer
HER2 status was available for 3,170/3,305 patients.
Of these, 728 (23%) were diagnosed withHER2-positive breast cancer
and 2,442 (77%) were diagnosed with HER2-negative breast cancer.
The association between the HER2 Pro1170Ala polymorphism and
survival was analyzed in patients with HER2-positive and
HER2-negative tumors, respectively. Among the patients with
HER2-positive breast cancer, no significant association was
identified between the HER2 Pro1170Ala genotype and survival (RFS:
Pro/Ala or Ala/Ala genotype vs. Pro/Pro genotype; unadjusted HR,
0.93; 95% CI, 0.56–1.54; P=0.776; Fig.
2A; DRFS: Unadjusted HR, 0.85; 95% CI, 0.49–1.48; P=0.565;
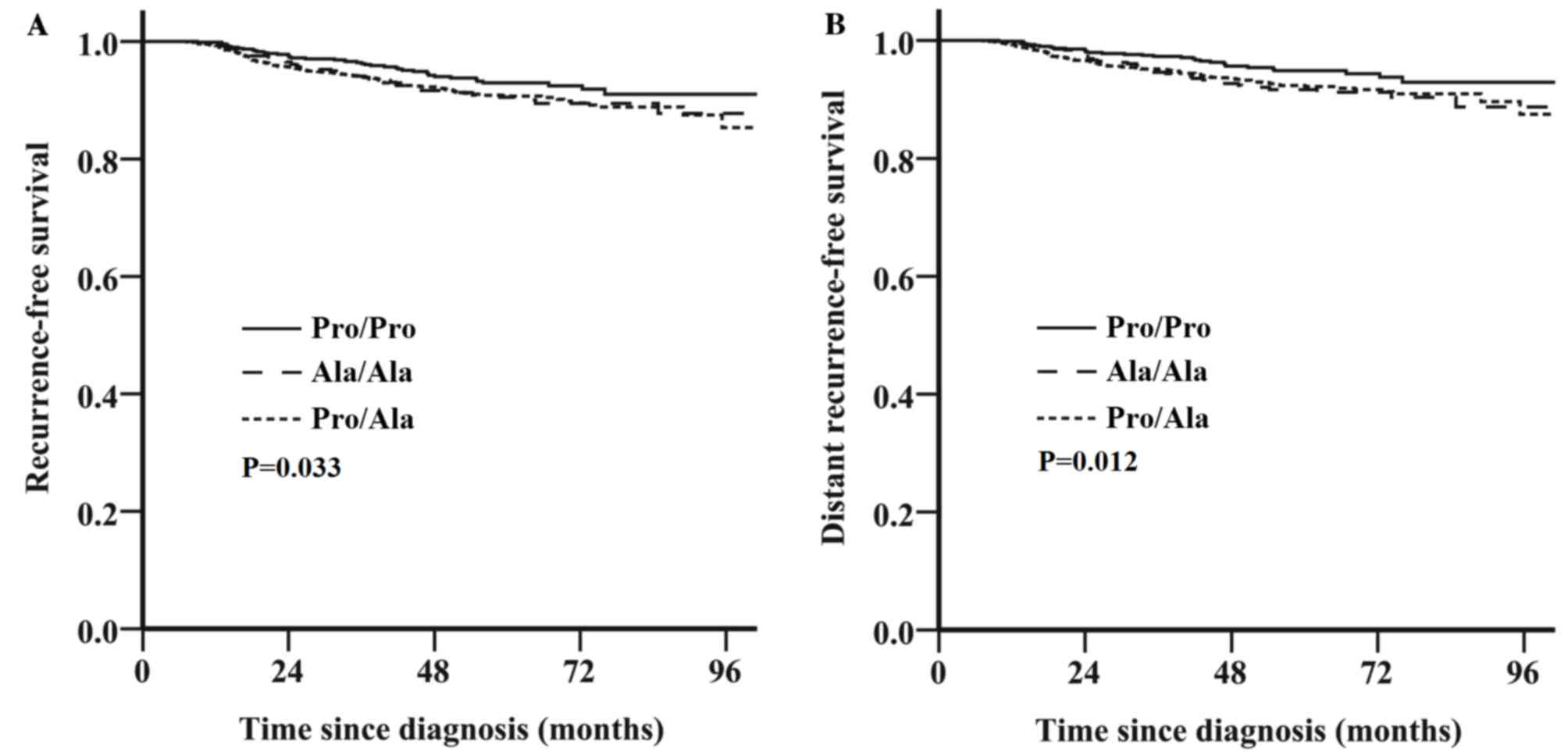
Fig. 2B). By contrast, among patients
with HER2-negative tumors, those with the Pro/Ala or Ala/Ala
genotype exhibited decreased RFS (unadjusted HR, 1.45; 95% CI,
1.03–2.04; P=0.033; Fig. 3A) and DRFS
(unadjusted HR, 1.65; 95% CI, 1.11–2.44; P=0.012; Fig. 3B) compared with those with the Pro/Pro
genotype. Furthermore, multivariate analysis revealed that the
Pro/Ala or Ala/Ala genotype was a near significant unfavorable
factor for RFS (adjusted HR, 1.46; 95% CI, 1.00–2.15; P=0.053;
Table II) and a significantly
unfavorable factor for DRFS (adjusted HR, 1.63; 95% CI, 1.05–2.53;
P=0.029; Table II) after adjustment
for age, tumor grade, tumor size, lymph node status, ER status, PR
status and adjuvant therapy.
 | Table II.Multivariate analysis of RFS and DRFS
in the 2,442 patients with HER2-negative breast cancer. |
Table II.
Multivariate analysis of RFS and DRFS
in the 2,442 patients with HER2-negative breast cancer.
|
| RFS | DRFS |
|---|
|
|
|
|
|---|
| Variable | HR (95% CI) | P-value | HR (95% CI) | P-value |
|---|
| Pro1170Ala
genotype |
| Pro/Ala
or Ala/Ala vs. Pro/Pro | 1.46
(1.00–2.00) | 0.053 | 1.63
(1.05–2.05) | 0.029 |
| Age, years |
| ≤40 vs.
>40 | 1.40
(0.96–2.96) | 0.081 | 1.42
(0.94–2.94) | 0.101 |
| Tumor size, cm |
| ≥2 vs.
<2 | 2.71
(1.84–4.84) | <0.001 | 2.58
(1.69–3.69) | <0.001 |
| Tumor grade |
| III vs.
I/II | 1.17
(0.71–1.71) | 0.548 | 1.13
(0.66–1.66) | 0.653 |
| Lymph nodes |
|
Positive vs. negative | 3.57
(2.52–5.52) | <0.001 | 4.29
(2.89–6.89) | <0.001 |
| ER status |
|
Negative vs. positive | 1.64
(1.04–2.04) | 0.033 | 1.66
(1.00–2.00) | 0.047 |
| PR status |
|
Negative vs. positive | 1.89
(1.22–2.22) | 0.004 | 1.82
(1.13–2.13) | 0.014 |
| Adjuvant
therapy |
| Therapy
vs. none | 1.63
(0.78–3.78) | 0.191 | 2.10
(1.00–4.00) | 0.048 |
Discussion
To the best of our knowledge, the present study is
the first to investigate the association between the HER2
Pro1170Ala polymorphism and survival outcome in a large cohort of
patients with breast cancer. Although the HER2 Pro1170Ala genotype
was not identified to be associated with survival outcome in the
entire cohort of 3,305 patients with breast cancer or in the 728
patients with HER2-positive breast cancer, this polymorphism was
identified to be significantly associated with survival outcome in
the 2,442 patients with HER2-negative breast cancer. The Pro/Ala or
Ala/Ala genotype was associated with a decreased RFS and DRFS
compared with the Pro/Pro genotype in the HER2-negativesubgroup,
and the Pro/Ala or Ala/Ala genotype was identified as an
independent unfavorable factor for DRFS, indicating that the Ala
variant led to a more aggressive phenotype compared with the Pro
variant among HER2-negative patients.
HER2 protein consists of four domains: The
extracellular region, the transmembrane domain, the tyrosine kinase
domain and the C-terminal tail (28).
The Pro1170Ala polymorphism is located in the tail coding region of
HER2. The C-terminal tail serves a critical role in the regulation
of the enzyme activity of the kinase (2,16,17,29,30). As a
non-synonymous coding variant, the Ala variant of the HER2
Pro1170Ala genotype may alter the spatial conformation of the tail
region and may affect tyrosine kinase activity (18,31).
A previous study suggested that the Ala variant
increased the risk of lung cancer (19), indicating that the Ala variant may
promote HER2 activity. In the present study, an association between
the HER2 Pro1170Ala polymorphism and survival was identified in
patients with HER2-negative breast cancer, but not in patients with
HER2-positive breast cancer. The underlying molecular mechanism for
this difference remains unclear; however, one possibility is that,
although the HER2 Pro1170Ala polymorphism may alter the HER2
activity, it is not sufficient to influence HER2 activity when the
HER2 gene is amplified or overexpressed. Although patients with
HER2-negative breast cancer have a more favorable survival outcome
compared with that of patients with HER2-positive breast cancer, a
minority of patients with HER2-negative breast cancer may exhibit
metastases after treatment (5–7).
Therefore, genotyping of the HER2 Pro1170Ala polymorphism may be
useful for identifying the relatively high-risk patients among all
patients with HER2-negative breast cancer. In conclusion, the
results of the present study demonstrated that, among the patients
with HER2-negative breast cancer, the HER2 Pro1170Ala polymorphism
is significantly associated with survival, with the Ala variant
exhibiting an aggressive phenotype and decreased survival outcome.
Further independent studies are required to confirm these
findings.
Acknowledgements
The present study was supported by the 973 project
(grant no. 2013CB911004), the National Science and Technology
Support Program (grant no. 2014BAI09B08) and the National Natural
Science Foundation of China (grant nos. 30973436 and 81071629).
Glossary
Abbreviations
Abbreviations:
|
HER2
|
human epidermal growth factor receptor
2
|
|
EGFR
|
epidermal growth factor receptor
|
|
ER
|
estrogen receptor
|
|
PR
|
progesterone receptor
|
|
RFS
|
recurrence-free survival
|
|
DRFS
|
distant recurrence-free survival
|
|
HR
|
hazard ratio
|
|
CI
|
confidence interval
|
References
|
1
|
Schreiber AB, Libermann TA, Lax I, Yarden
Y and Schlessinger J: Biological role of epidermal growth
factor-receptor clustering. Investigation with monoclonal
anti-receptor antibodies. J Biol Chem. 258:846–853. 1983.PubMed/NCBI
|
|
2
|
Lemmon MA, Schlessinger J and Ferguson KM:
The EGFR family: Not so prototypical receptor tyrosine kinases.
Cold Spring Harb Perspect Biol. 6:a0207682014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Akiyama T, Sudo C, Ogawara H, Toyoshima K
and Yamamoto T: The product of the human c-erbB-2 gene: A
185-kilodalton glycoprotein with tyrosine kinase activity. Science.
232:1644–1646. 1986. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Slamon D, Clark G, Wong S, Levin W,
Ullrich A and McGuire W: Human breast cancer: Correlation of
relapse and survival with amplification of the HER-2/neu oncogene.
Science. 235:177–182. 1987. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Toikkanen S, Helin H, Isola J and Joensuu
H: Prognostic significance of HER-2 oncoprotein expression in
breast cancer: A 30-year follow-up. J Clin Oncol. 10:1044–1048.
1992. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Seshadri R, Firgaira FA, Horsfall DJ,
McCaul K, Setlur V and Kitchen P: Clinical significance of
HER-2/neu oncogene amplification in primary breast cancer. The
South Australian Breast Cancer Study Group. J Clin Oncol.
11:1936–1942. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Sjogren S, Inganäs M, Lindgren A, Holmberg
L and Bergh J: Prognostic and predictive value of c-erbB-2
overexpression in primary breast cancer, alone and in combination
with other prognostic markers. J Clin Oncol. 16:462–469. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Scorilas A, Yotis J, Pateras C, Trangas T
and Talieri M: Predictive value of c-erbB-2 and cathepsin-D for
Greek breast cancer patients using univariate and multivariate
analysis. Clin Cancer Res. 5:815–821. 1999.PubMed/NCBI
|
|
9
|
Ludovini V, Gori S, Colozza M, Pistola L,
Rulli E, Floriani I, Pacifico E, Tofanetti FR, Sidoni A, Basurto C,
et al: Evaluation of serum HER2 extracellular domain in early
breast cancer patients: Correlation with clinicopathological
parameters and survival. Ann Oncol. 19:883–890. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Piccart-Gebhart MJ, Procter M,
Leyland-Jones B, Goldhirsch A, Untch M, Smith I, Gianni L, Baselga
J, Bell R, Jackisch C, et al: Trastuzumab after adjuvant
chemotherapy in HER2-positive breast cancer. N Engl J Med.
353:1659–1672. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Gianni L, Dafni U, Gelber RD, Azambuja E,
Muehlbauer S, Goldhirsch A, Untch M, Smith I, Baselga J, Jackisch
C, et al: Treatment with trastuzumab for 1 year after adjuvant
chemotherapy in patients with HER2-positive early breast cancer: A
4-year follow-up of a randomised controlled trial. Lancet Oncol.
12:236–244. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Romond EH, Perez EA, Bryant J, Suman VJ,
Geyer CE Jr, Davidson NE, Tan-Chiu E, Martino S, Paik S, Kaufman
PA, et al: Trastuzumab plus adjuvant chemotherapy for operable
HER2-positive breast cancer. N Engl J Med. 353:1673–1684. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Slamon D, Eiermann W, Robert N, Pienkowski
T, Martin M, Press M, Mackey J, Glaspy J, Chan A, Pawlicki M, et
al: Adjuvant trastuzumab in HER2-positive breast cancer. N Engl J
Med. 365:1273–1283. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Han X, Diao L, Xu Y, Xue W, Ouyang T, Li
J, Wang T, Fan Z, Fan T, Lin B and Xie Y: Association between the
HER2 Ile655Val polymorphism and response to trastuzumab in women
with operable primary breast cancer. Ann Oncol. 25:1158–1164. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tommasi S, Fedele V, Lacalamita R, Bruno
M, Schittulli F, Ginzinger D, Scott G, Eppenberger-Castori S,
Calistri D, Casadei S, et al: 655Val and 1170Pro ERBB2 SNPs in
familial breast cancer risk and BRCA1 alterations. Cell Oncol.
29:241–248. 2007.PubMed/NCBI
|
|
16
|
Fleishman SJ, Schlessinger J and Ben-Tal
N: A putative molecular-activation switch in the transmembrane
domain of erbB2. Proc Natl Acad Sci USA. 99:15937–15940. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Jorissen RN, Walker F, Pouliot N, Garrett
TP, Ward CW and Burgess AW: Epidermal growth factor receptor:
Mechanisms of activation and signalling. Exp Cell Res. 284:31–53.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhang X, Gureasko J, Shen K, Cole PA and
Kuriyan J: An allosteric mechanism for activation of the kinase
domain of epidermal growth factor receptor. Cell. 125:1137–1149.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Jo UH, Han SG, Seo JH, Park KH, Lee JW,
Lee HJ, Ryu JS and Kim YH: The genetic polymorphisms of HER-2 and
the risk of lung cancer in a Korean population. BMC cancer.
8:3592008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Benusiglio PR, Lesueur F, Luccarini C,
Conroy DM, Shah M, Easton DF, Day NE, Dunning AM, Pharoah PD and
Ponder BA: Common ERBB2 polymorphisms and risk of breast cancer in
a white British population: A case-control study. Breast Cancer
Res. 7:R204–R209. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
21
|
Han W, Kang D, Lee JE, Park IA, Choi JY,
Lee KM, Bae JY, Kim S, Shin ES, Lee JE, et al: A haplotype analysis
of HER-2 gene polymorphisms: Association with breast cancer risk,
HER-2 protein expression in the tumor, and disease recurrence in
Korea. Clin Cancer Res. 11:4775–4782. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Breyer JP, Sanders ME, Airey DC, Cai Q,
Yaspan BL, Schuyler PA, Dai Q, Boulos F, Olivares MG, Bradley KM,
et al: Heritable variation of ERBB2 and breast cancer risk. Cancer
Epidemiol Biomarkers Prev. 18:1252–1258. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Tong SY, Ha SY, Ki KD, Lee JM, Lee SK, Lee
KB, Kim MK, Cho CH and Kwon SY: The effects of obesity and HER-2
polymorphisms as risk factors for endometrial cancer in Korean
women. BJOG. 116:1046–1052. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Stanton SE, Ward MM, Christos P, Sanford
R, Lam C, Cobham MV, Donovan D, Scheff RJ, Cigler T, Moore A, et
al: Pro1170 Ala polymorphism in HER2-neu is associated with risk of
trastuzumab cardiotoxicity. BMC cancer. 15:2672015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wittekind C: 2010 TNM system: On the 7th
edition of TNM classification of malignant tumors. Pathologe.
31:331–332. 2010.(In German). View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wittekind C: Lymph nodes, tumour deposits,
and TNM: Are we getting better? 7th edition of UICC 2010 TNM
classification of malignant tumors. Strahlenther Onkol.
188:191–192. 2012.(In German). View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wolff AC, Hammond ME, Hicks DG, Dowsett M,
McShane LM, Allison KH, Allred DC, Bartlett JM, Bilous M,
Fitzgibbons P, et al: Recommendations for human epidermal growth
factor receptor 2 testing in breast cancer: American Society of
Clinical Oncology/College of American Pathologists clinical
practice guideline update. J Clin Oncol. 31:3997–4013. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Franklin MC, Carey KD, Vajdos FF, Leahy
DJ, de Vos AM and Sliwkowski MX: Insights into ErbB signaling from
the structure of the ErbB2-pertuzumab complex. Cancer Cell.
5:317–328. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Honegger AM, Kris RM, Ullrich A and
Schlessinger J: Evidence that autophosphorylation of solubilized
receptors for epidermal growth-factor is mediated by intermolecular
cross-phosphorylation. Proc Natl Acad Sci USA. 86:925–929. 1989.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Alvarez CV, Shon KJ, Miloso M and Beguinot
L: Structural requirements of the epidermal growth-factor receptor
for tyrosine phosphorylation of Eps8 and Eps15, substrates lacking
Src Sh2 homology domains. J Biol Chem. 270:16271–16276. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wood ER, Truesdale AT, McDonald OB, Yuan
D, Hassell A, Dickerson SH, Ellis B, Pennisi C, Horne E, Lackey K,
et al: A unique structure for epidermal growth factor receptor
bound to GW572016 (Lapatinib): Relationships among protein
conformation, inhibitor off-rate and receptor activity in tumor
cells. Cancer Res. 64:6652–6659. 2004. View Article : Google Scholar : PubMed/NCBI
|