Introduction
Bladder cancer is a malignancy of the bladder
mucosa; it is the most common malignancy of the urinary system. Its
incidence ranks top among all urogenital tumors in China, and ranks
second in Western countries only to prostate cancer (1–5). Bladder
urothelial carcinoma is diagnosed mainly via cystoscopy (6,7).
miRNAs are a class of evolutionarily conserved
non-coding small molecule RNAs that function by regulating gene
expression at the translational level; the specific biological
targets of most miRNAs remain to be discovered (8,9). miRNAs
have been found to regulate cell growth and tissue differentiation,
and thus play roles during development and can be involved in the
establishment of diseases such as cancer (10). A series of studies have shown that
microRNAs play roles in cell growth and apoptosis, hemocytes
differentiation, homeobox gene regulation, neuronal polarity,
insulin secretion, cerebral morphogenesis, cardiac development and
postembryonic development (11–14).
Active research focuses on the association between miRNAs and
different tumor development.
This study was designed in order to analyze the
information contained in The Cancer Genome Atlas (TCGA) for bladder
urothelial carcinoma patients, so as to identify independent
prognostic miRNAs that should be useful as reference markers for
the diagnosis of the cancer.
Materials and methods
The miRNA data for bladder urothelial carcinoma
patients were downloaded from TCGA in the USA. The data involved
cases from 1991 to 2013. Overall the tissue specimen data of 418
patients diagnosed with bladder urothelial carcinoma as well as the
data for 19 cases of para-carcinoma tissue specimens were obtained
from TCGA. To study the association between miRNA expression and
clinical features in patients with bladder urothelial carcinoma in
an accurate manner, we selected the patient data according to the
following criteria: The age of the patient was available, the
survival time was longer than 1 month and pathology had confirmed
the diagnosis of bladder urothelial carcinoma. After exclusion, a
total of 399 cases of bladder urothelial carcinoma and 19 cases
with para-carcinoma tissue specimens were included in this study.
The available clinical features included age, sex, race, the
American Joint Committee on Cancer (AJCC) TNM staging system
classification, the survival time and the survival status.
The miRNA expression data for the samples of 399
bladder urothelial carcinoma patients and 19 para-carcinoma
specimens were included in the analysis, and a total of 1,581
miRNAs were detected in tissue specimens by using the Illumina
HiSeq platform and the Illumina Genome Analyzer platform. To screen
out the differentially expressed miRNAs, we set the screening
criteria as log fold-change >2 and the associated P<0.01
miRNAs that were upregulated and downregulated in the tissue
specimens of bladder urothelial carcinoma patients were screened
out, and the expression of miRNA was normalized.
In addition, we transformed the miRNA expression
levels into log2, and screened out the miRNAs and clinical features
that were associated with the survival of bladder urothelial
carcinoma using the univariate Cox proportional hazards regression.
Then, we used the multivariate Cox proportional hazards regression
to screen out the independent prognostic factors of bladder
urothelial carcinoma. Furthermore, we performed a Kaplan-Meier (KM)
survival curve analysis on the independent prognostic miRNAs for
bladder urothelial carcinoma, divided the miRNA levels into
high-expression and low-expression groups with the median as the
cut-off point, and processed data using the survival package of the
R software, version 2.14.1 (R development Core Team, Vienna
Austria, 2009) (15).
TargetScan was used for target prediction on the
independent prognostic miRNAs for bladder urothelial carcinoma.
After excluding repeated target genes, cytoscape was used
performing a GO enrichment analysis on target genes (p<0.05) to
screen out ten functions with the closest relationship to target
genes. Finally, KOBAS was used for KEGG pathway enrichment analysis
on the target genes to screen out ten pathways with the closest
association to target genes.
Results
The data from a total of 399 patients with bladder
urothelial carcinoma were included in the analysis. The data for
the expression of 1,581 miRNAs and a variety of clinical features
can be seen in Table I. In addition,
we found the up and downregulated miRNAs in bladder urothelial
carcinoma. Among them, the ten most upregulated miRNAs were
hsa-mir-210, hsa-mir-96, hsa-mir-18a, hsa-mir-183, hsa-mir-130b,
hsa-mir-141, hsa-mir-33a, hsa-mir-345, hsa-mir-182 and hsa-mir-301a
(Table II), whereas, the most
downregulated ones were hsa-mir-143, hsa-mir-1298, hsa-mir-139,
hsa-mir-1-2, hsa-mir-195, hsa-mir-1-1, hsa-mir-133a-1, hsa-mir-30a,
hsa-mir-133a-2 and hsa-mir-133b (Table
III).
 | Table I.Clinical features of bladder
urothelial carcinoma. |
Table I.
Clinical features of bladder
urothelial carcinoma.
| Characteristics | Data (TCGA) |
|---|
| Age |
|
| ≤65 | 160 |
|
>65 | 239 |
| Sex |
|
|
Female | 105 |
| Male | 294 |
| Race |
|
|
White | 318 |
|
Asian | 40 |
| Black or
African American | 23 |
| NA | 18 |
| Pathology stage |
|
| Stage
I | 2 |
| Stage
II | 125 |
| Stage
III | 137 |
| Stage
IV | 133 |
| NA | 2 |
| T stage |
|
| T0 | 1 |
| T1 | 3 |
| T2 | 114 |
| T3 | 191 |
| T4 | 58 |
| TX | 1 |
| NA | 31 |
| N stage |
|
| N0 | 229 |
| N1 | 46 |
| N2 | 76 |
| N3 | 7 |
| NA | 5 |
| NX | 36 |
| M stage |
|
| M0 | 188 |
| M1 | 10 |
| MA | 198 |
| MX | 3 |
| Survival state |
|
|
Survive | 223 |
| Dead | 176 |
 | Table II.The top ten miRNA of upregulated
expression. |
Table II.
The top ten miRNA of upregulated
expression.
| miRNA | logFC | logCPM | P-value | FDR |
|---|
| hsa-mir-210 | 4.803108853 | 9.8642203 | 3.45E-18 | 3.63E-16 |
| hsa-mir-96 | 3.410662815 | 5.143223988 | 1.15E-16 | 1.07E-14 |
| hsa-mir-18a | 2.988898835 | 5.68100551 | 6.61E-16 | 5.50E-14 |
| hsa-mir-183 | 2.943927749 | 13.56270017 | 2.06E-14 | 1.25E-12 |
| hsa-mir-130b | 2.303867052 | 5.74901106 | 2.72E-14 | 1.59E-12 |
| hsa-mir-141 | 2.494450708 | 11.09284497 | 1.42E-11 | 5.49E-10 |
| hsa-mir-33a | 2.22270633 | 4.907692995 | 7.44E-11 | 2.67E-09 |
| hsa-mir-345 | 2.673695186 | 4.694451091 | 1.23E-10 | 4.23E-09 |
| hsa-mir-182 | 2.254097092 | 14.41365672 | 1.34E-10 | 4.49E-09 |
| hsa-mir-301a | 2.088008339 | 4.476181049 | 1.76E-10 | 5.79E-09 |
 | Table III.The top ten miRNA of downregulated
expression. |
Table III.
The top ten miRNA of downregulated
expression.
| miRNA | logFC | logCPM | P-value | FDR |
|---|
| hsa-mir-143 | −3.599914903 | 17.8945552 | 3.72E-46 | 5.88E-43 |
| hsa-mir-1298 | −4.526050745 | 1.187271104 | 1.13E-39 | 8.92E-37 |
| hsa-mir-139 | −2.621941214 | 6.101969894 | 9.29E-39 | 4.89E-36 |
| hsa-mir-1-2 | −3.736158674 | 5.508819647 | 4.22E-34 | 1.67E-31 |
| hsa-mir-195 | −2.432143312 | 4.989973255 | 6.44E-33 | 2.04E-30 |
| hsa-mir-1-1 | −3.693697839 | 5.417492946 | 1.41E-32 | 3.73E-30 |
| hsa-mir-133a-1 | −3.558543939 | 5.92141089 | 2.24E-28 | 5.06E-26 |
| hsa-mir-30a | −2.228925868 | 13.94817286 | 6.07E-27 | 1.20E-24 |
| hsa-mir-133a-2 | −3.515469057 | 5.736507 | 2.23E-26 | 3.92E-24 |
| hsa-mir-133b | −3.88838679 | 4.46304315 | 3.14E-26 | 4.97E-24 |
The univariate Cox proportional risk regression
shown in Table I demonstrates the
clinical features of pathology N stage (p=1.01E-07), pathology M
stage (p=0.0002), pathology T stage (p=0.0005) and patient's age
(p=0.007) were all closely associated with the survival length for
bladder urothelial carcinoma. As for the miRNAs, related to the
bladder urothelial carcinoma survival, a total of 19 miRNAs stood
out, of which the three most closely related ones were hsa-mir-99a
(p=0.0001), hsa-let-7c (p=0.0004) and hsa-mir-100 (p=0.0005).
Next, a multivariate Cox proportional risk
regression showed the pathology N stage (p=0.00007), hsa-mir-518b
(p=0.02), the patient's age (p=0.03), hsa-mir-192 (p=0.04) and
hsa-mir-7705 (p=0.04) were all independent prognostic factors for
bladder urothelial carcinoma (Table
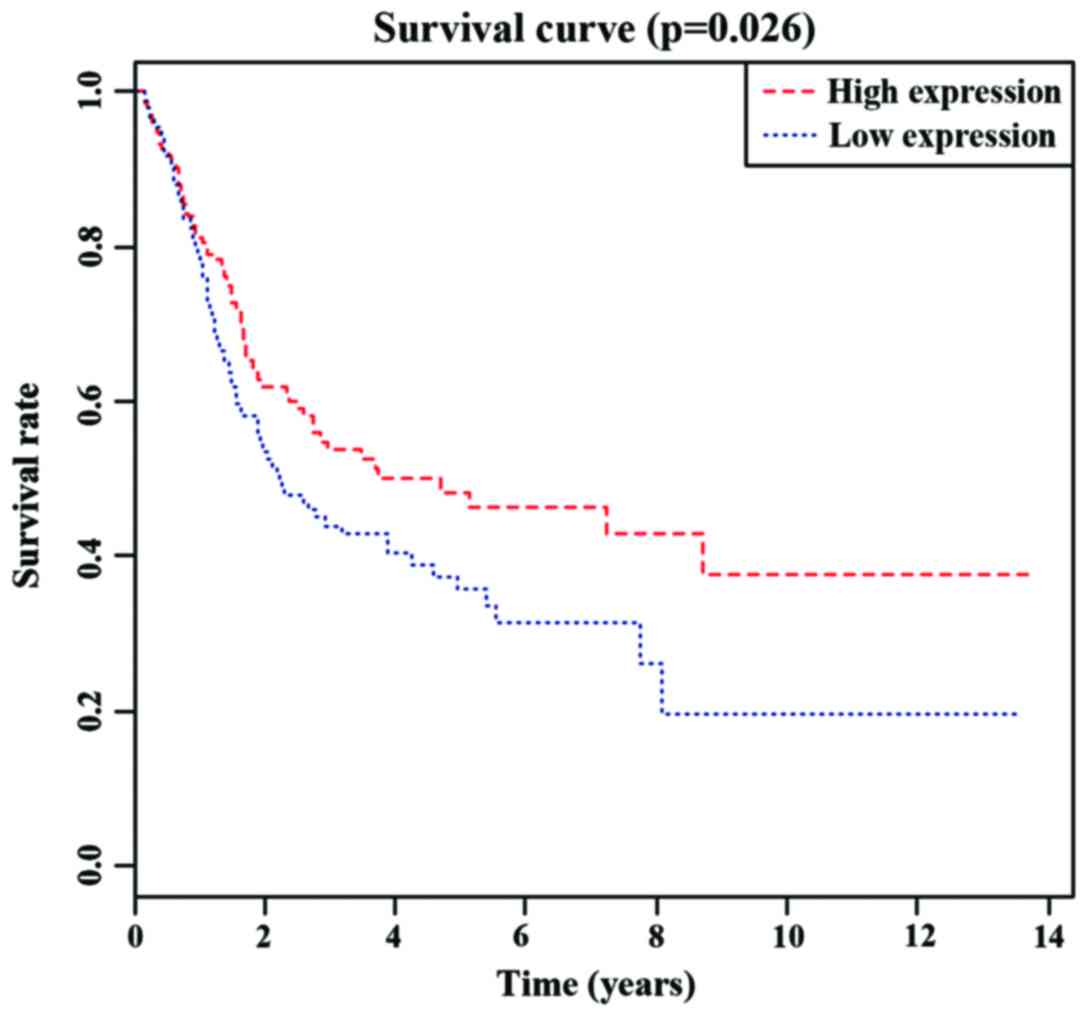
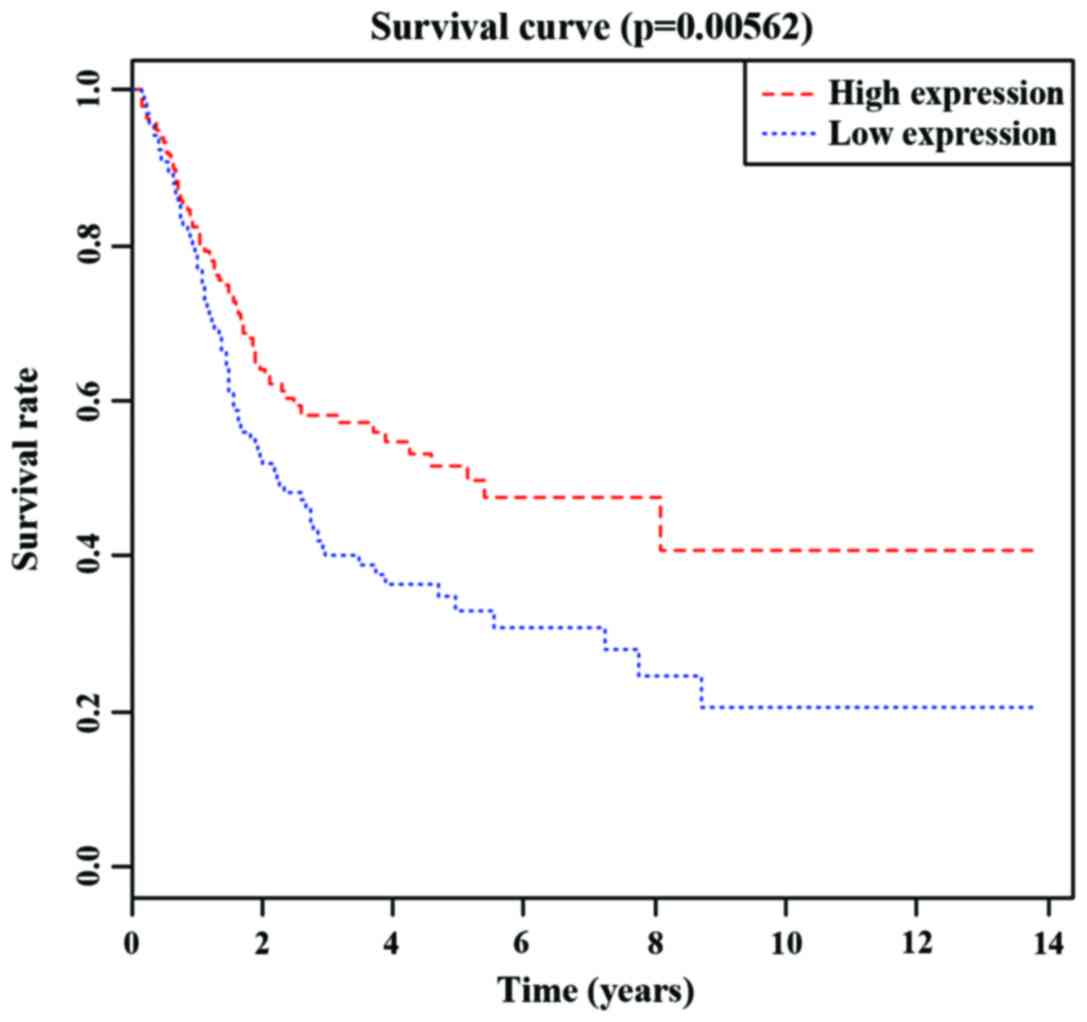
IV). We performed KM survival curve analyses on the three
independent prognostic miRNAs. As shown in Figs. 1–3,
there were significant differences in the survival time between the
high- and low-expression levels of hsa-mir-7705 and hsa-mir-192,
while the survival time between the high- and low-expression levels
of hsa-mir-518b did not differ significantly.
 | Table IV.Univariate and multivariate
analyses. |
Table IV.
Univariate and multivariate
analyses.
|
| Univariate
analysis | Multivariate
analysis |
|---|
|
|
|
|
|---|
| Variables | HR (95% CI) | P-value | HR (95% CI) | P-value |
|---|
| Pathology N
stage | 0.87 | 0.0000001 | 0.78 | 0.00007 |
| hsa-mir-518b | 0.08 | 0.04 | 0.1 | 0.02 |
| Age | 0.49 | 0.007 | 0.42 | 0.03 |
| hsa-mir-192 | −0.17 | 0.01 | −0.17 | 0.04 |
| hsa-mir-7705 | −0.22 | 0.04 | −0.24 | 0.04 |
Finally, we used TargetScan to predict the target
genes of three independent prognostic miRNAs, and then used
Cytoscape to perform a GO enrichment analysis on all target genes.
As shown in Table V, the three most
significant functions found were transcription regulator activity,
regulation of transcription from RNA polymerase II promoter and
regulation of transcription. The KEGG pathway analysis revealed
that the most closely related pathway was Cytokine-cytokine
receptor interaction (Table VI).
 | Table V.The analysis of GO enrichment. |
Table V.
The analysis of GO enrichment.
| GO-ID | P-value | Corr P-value | Description |
|---|
| 30528 | 1.28E-10 | 1.86E-07 | Transcription
regulator activity |
| 6357 | 4.58E-07 | 2.29E-04 | Regulation of
transcription from RNA polymerase II promoter |
| 45449 | 7.35E-07 | 2.29E-04 | Regulation of
transcription |
| 16564 | 7.93E-07 | 2.29E-04 | Transcription
repressor activity |
| 3676 | 8.48E-07 | 2.29E-04 | Nucleic acid
binding |
| 19219 | 9.42E-07 | 2.29E-04 | Regulation of
nucleobase, nucleoside, nucleotide and nucleic acid metabolic
process |
| 51171 | 1.18E-06 | 2.45E-04 | Regulation of
nitrogen compound metabolic process |
| 6355 | 1.92E-06 | 3.49E-04 | Regulation of
transcription, DNA-dependent |
| 5634 | 2.71E-06 | 4.39E-04 | Nucleus |
| 51252 | 3.24E-06 | 4.62E-04 | Regulation of RNA
metabolic process |
 | Table VI.The analysis of KEGG enrichment. |
Table VI.
The analysis of KEGG enrichment.
| Term | Database | ID | P-value | Corrected
P-value |
|---|
| Cytokine-cytokine
receptor interaction | KEGG PATHWAY | hsa04060 | 0.000784246 | 0.025901739 |
| TGF-β signaling
pathway | KEGG PATHWAY | hsa04350 | 0.001564908 | 0.025901739 |
| PI3K-Akt signaling
pathway | KEGG PATHWAY | hsa04151 | 0.001619992 | 0.025901739 |
| mRNA surveillance
pathway | KEGG PATHWAY | hsa03015 | 0.001865223 | 0.025901739 |
| NF-κB signaling
pathway | KEGG PATHWAY | hsa04064 | 0.00190454 | 0.025901739 |
| Sphingolipid
signaling pathway | KEGG PATHWAY | hsa04071 | 0.003163201 | 0.035849606 |
| Apoptosis | KEGG PATHWAY | hsa04210 | 0.004187233 | 0.040675974 |
| Transcriptional
misregulation in cancer | KEGG PATHWAY | hsa05202 | 0.00677533 | 0.049549952 |
| Herpes simplex
infection | KEGG PATHWAY | hsa05168 | 0.007212651 | 0.049549952 |
| Chemokine signaling
pathway | KEGG PATHWAY | hsa04062 | 0.007286758 | 0.049549952 |
Discussion
Bladder cancer is the most common malignancy of the
urinary system. Studies have found that miRNAs can regulate the
expression of tens to hundreds of genes, and that miRNAs play a hub
role in tumorigenesis and progression as regulatory molecules in
the processes of gene expression and protein translation (16,17).
Although these endogenous miRNAs only account for 2% of the total
human transcripts, they regulate the expression of >30% of the
genes in the whole human genome. Therefore, research on the
function of miRNAs and their association to the development of
cancer is of great significance. This study identified independent
prognostic factors of bladder urothelial carcinoma by studying the
expression data of miRNAs.
We extracted the miRNA expression data of bladder
urothelial carcinoma from the free and open TCGA. After screening,
we listed the ten most significantly upregulated and downregulated
miRNAs in the specimens, of which the most pronouncedly upregulated
ones were hsa-mir-210, hsa-mir-96 and hsa-mir-18a. The level of
hsa-miR-210 has been associated with hypoxia in head and neck
tumors and other indicators, and a high hsa-miR-210 expression is
closely related to disease recurrence and overall survival time
(18). In addition, another study
showed that hsa-miR-210 is an independent prognostic marker for
breast cancer, and that hsa-miR-210 overexpression is induced by
hypoxia in a HIF-1α- and VHL-dependent manner (19). Then, the miRNA hsa-miR-96 can
upregulate the levels of IRS1 and MAP4K1 to affect bladder cancer
cell growth and has been signaled as a promising diagnostic marker
for human bladder urothelial carcinomas (20). Interestingly, hsa-mir-18a has also
been shown to be involved in many cancer processes. A study
demonstrated that hsa-mir-18a was able to prevent estrogen
receptor-α expression and promote hepatoma cell proliferation
(21).
In addition, we screened out the three best
independent prognostic miRNAs for bladder urothelial carcinoma by
constructing a multivariate Cox regression model, and obtained
hsa-mir-518b (p=0.02), hsa-mir-192 (p=0.04) and hsa-mir-7705
(p=0.04). Furthermore, by plotting the KM survival curve, we found
that the survival time of patients expressing high hsa-mir-7705 and
hsa-mir-192 levels were significantly shorter than those of
patients expressing low levels. In addition, both hsa-mir-7705 and
hsa-mir-192 were upregulated in the bladder urothelial carcinoma
samples. A study in mice showed that together with elevated
transforming growth factor-β (TGF-β) and Col1a2 levels, the levels
of miR-192 are significantly higher in the glomeruli of diabetic
mice when compared to the levels in non-diabetic controls, these
finding suggest that miRNAs in the kidney play a role
downregulating E-box repressors to control the TGF-β-induced Col1a2
expression (22).
Finally, we predicted target genes for the three
independent prognostic miRNAs, and then subjected those target
genes to GO enrichment analysis using Cytoscape. The transcription
regulator activity was the most closely associated enrichment area,
while the other findings included transcription from RNA polymerase
II promoter and regulation of transcription. The KEGG pathway
results pointed to three associated pathways including
Cytokine-cytokine receptor interactions, TGF-β, and PI3K-Akt
signaling pathways. According to Kaminska et al TGF-β
signaling inhibitors may be promising novel, potential antitumor
therapeutics as they decrease glioma viability and invasion in
animal models (23). In addition,
other researchers have shown possible initiation of pathological
changes in osteoarthritis by high concentrations of active TGF-β1
in subchondral bone, and a potential approach for managing this
disease may be to suppress TGF-β1 (24). Finally, the PI3K-Akt signaling pathway
has also been studied, and it is thought that phosphoinositide
3-kinases (PI3Ks) are involved in a variety of tumor processes
(25).
Even though this study identified three independent
prognostic miRNAs, we are aware of the study's limitations.
Firstly, all 399 cases of bladder urothelial carcinoma were from a
single database (TCGA), so verification with data from another
database is needed. On the same line, 399 cases are a rather small
group size; so larger sample verification is needed. Finally, PCR
validation assays will definitely enhance the reliability of our
findings.
In conclusion, our approach was successful in
identifying hsa-mir-7705, hsa-mir-192 and hsa-mir-518b as
independent prognostic factors for bladder urothelial
carcinoma.
References
|
1
|
Babjuk M, Burger M, Zigeuner R, Shariat
SF, van Rhijn BW, Compérat E, Sylvester RJ, Kaasinen E, Böhle A,
Palou Redorta J, et al: European Association of Urology: EAU
guidelines on non-muscle-invasive urothelial carcinoma of the
bladder: Update 2013. Eur Urol. 64:639–653. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cancer Genome Atlas Research Network:
Comprehensive molecular characterization of urothelial bladder
carcinoma. Nature. 507:315–322. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Sternberg CN, Skoneczna I, Kerst JM,
Albers P, Fossa SD, Agerbaek M, Dumez H, de Santis M, Théodore C,
Leahy MG, et al: European Organisation for Research and Treatment
of Cancer Genito-Urinary Cancers Group; Groupe d'Etude des Tumeurs
Urogénitales; National Cancer Research Institute Bladder Cancer
Study Group; National Cancer Institute of Canada Clinical Trials
Group; German Association of Urologic Oncology: Immediate versus
deferred chemotherapy after radical cystectomy in patients with
pT3-pT4 or N+ M0 urothelial carcinoma of the bladder (EORTC 30994):
An intergroup, open-label, randomised phase 3 trial. Lancet Oncol.
16:76–86. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sanford T, Porten S and Meng MV: Molecular
analysis of upper tract and bladder urothelial carcinoma: Results
from a microarray comparison. PLoS One. 10:e01371412015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Liu J, Cai M, Chen J, Liao Y, Mai S, Li Y,
Huang X, Liu Y, Zhang J, Kung H, et al: α4 contributes to bladder
urothelial carcinoma cell invasion and/or metastasis via regulation
of E-cadherin and is a predictor of outcome in bladder urothelial
carcinoma patients. Eur J Cancer. 50:840–851. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Sahin S, Resorlu B, Atar FA, Eksi M, Sener
NC and Tugcu V: Laparoscopic ureterolithotomy with concomitant
pyelolithotomy using flexible cystoscope. Urol J. 13:2833–2836.
2016.PubMed/NCBI
|
|
7
|
Mahmood N and Pasha T: A new technique to
prevent curling of guide wire in urinary bladder during J stent
insertion with flexible cystoscope. Can Urol Assoc J. 10:E34–E35.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Frampton GM, Fichtenholtz A, Otto GA, Wang
K, Downing SR, He J, Schnall-Levin M, White J, Sanford EM, An P, et
al: Development and validation of a clinical cancer genomic
profiling test based on massively parallel DNA sequencing. Nat
Biotechnol. 31:1023–1031. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
9
|
Köttgen A, Albrecht E, Teumer A, Vitart V,
Krumsiek J, Hundertmark C, Pistis G, Ruggiero D, O'Seaghdha CM,
Haller T, et al: LifeLines Cohort Study; CARDIoGRAM Consortium;
DIAGRAM Consortium; ICBP Consortium; MAGIC Consortium: Genome-wide
association analyses identify 18 new loci associated with serum
urate concentrations. Nat Genet. 45:145–154. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Nunez-Iglesias J, Liu CC, Morgan TE, Finch
CE and Zhou XJ: Joint genome-wide profiling of miRNA and mRNA
expression in Alzheimer's disease cortex reveals altered miRNA
regulation. PLoS One. 5:e88982010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Khorshid M, Hausser J, Zavolan M and van
Nimwegen E: A biophysical miRNA-mRNA interaction model infers
canonical and noncanonical targets. Nat Methods. 10:253–255. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Muniategui A, Pey J, Planes FJ and Rubio
A: Joint analysis of miRNA and mRNA expression data. Brief
Bioinform. 14:263–278. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Luo D, Wilson JM, Harvel N, Liu J, Pei L,
Huang S, Hawthorn L and Shi H: A systematic evaluation of miRNA:
mRNA interactions involved in the migration and invasion of breast
cancer cells. J Transl Med. 11:572013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gu P, Reid JG, Gao X, Shaw CA, Creighton
C, Tran PL, Zhou X, Drabek RB, Steffen DL, Hoang DM, et al: Novel
microRNA candidates and miRNA-mRNA pairs in embryonic stem (ES)
cells. PLoS One. 3:e25482008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Baayen RH: Analyzing Linguistic Data: A
Practical Introduction to Statistics Using R. Cambridge University
Press; New York: 2008, https://doi.org/10.1017/CBO9780511801686
View Article : Google Scholar
|
|
16
|
Liu B, Li J and Tsykin A: Discovery of
functional miRNA-mRNA regulatory modules with computational
methods. J Biomed Inform. 42:685–691. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Tonevitsky AG, Maltseva DV, Abbasi A,
Samatov TR, Sakharov DA, Shkurnikov MU, Lebedev AE, Galatenko VV,
Grigoriev AI and Northoff H: Dynamically regulated miRNA-mRNA
networks revealed by exercise. BMC Physiol. 13:92013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gee HE, Camps C, Buffa FM, Patiar S,
Winter SC, Betts G, Homer J, Corbridge R, Cox G, West CM, et al:
hsa-mir-210 is a marker of tumor hypoxia and a prognostic factor in
head and neck cancer. Cancer. 116:2148–2158. 2010.PubMed/NCBI
|
|
19
|
Camps C, Buffa FM, Colella S, Moore J,
Sotiriou C, Sheldon H, Harris AL, Gleadle JM and Ragoussis J:
hsa-miR-210 is induced by hypoxia and is an independent prognostic
factor in breast cancer. Clin Cancer Res. 14:1340–1348. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang Y, Luo H, Li Y, Chen T, Wu S and Yang
L: hsa-miR-96 up-regulates MAP4K1 and IRS1 and may function as a
promising diagnostic marker in human bladder urothelial carcinomas.
Mol Med Rep. 5:260–265. 2012.PubMed/NCBI
|
|
21
|
Liu WH, Yeh SH, Lu CC, Yu SL, Chen HY, Lin
CY, Chen DS and Chen PJ: MicroRNA-18a prevents estrogen receptor-α
expression, promoting proliferation of hepatocellular carcinoma
cells. Gastroenterology. 136:683–693. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kato M, Zhang J, Wang M, Lanting L, Yuan
H, Rossi JJ and Natarajan R: MicroRNA-192 in diabetic kidney
glomeruli and its function in TGF-β-induced collagen expression via
inhibition of E-box repressors. Proc Natl Acad Sci USA. 104:pp.
3432–3437. 2007; View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kaminska B, Kocyk M and Kijewska M: TGF
beta signaling and its role in glioma pathogenesis. Adv Exp Med
Biol. 986:171–187. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhen G, Wen C, Jia X, Li Y, Crane JL,
Mears SC, Askin FB, Frassica FJ, Chang W, Yao J, et al: Inhibition
of TGF-β signaling in mesenchymal stem cells of subchondral bone
attenuates osteoarthritis. Nat Med. 19:704–712. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Martini M, De Santis MC, Braccini L,
Gulluni F and Hirsch E: PI3K/AKT signaling pathway and cancer: An
updated review. Ann Med. 46:372–383. 2014. View Article : Google Scholar : PubMed/NCBI
|