Introduction
Oral cancer is the 3rd most common cancer type in
developing countries and the 6th most common worldwide (1). Oral squamous cell carcinoma (OSCC),
which is the representative form of oral cancer, shows the highest
incidence among all the head and neck cancers, and account for ~3%
of the newly diagnosed cancer cases. Despite the corresponding
diagnostic and clinical treatment strategies to prevent and treat
OSCC, the survival rate of OSCC patients has not been significantly
improved and is still below 50% (2).
In the past, clinical studies on OSCC have generally restricted at
the physiological and biochemical level, such as detecting the
activity of basal cells in OSCC, computing the rate and frequency
of proliferation and metastasis. However, the underlying molecular
mechanism of OSCC tumor formation remained unclear. In order to
improve the prognosis of OSCC patients and the treatment efficacy,
the molecular mechanism of OSCC tumor formation and the
identification of effective biological indicators and therapeutic
targets are urgently needed.
Long non-coding RNA (lncRNA) is a class of newly
discovered non-protein coding RNA transcripts >200 nt. Studies
have shown that lncRNAs have important biological functions in a
variety of physiological processes, such as carcinogenesis, cell
proliferation, differentiation, apoptosis and cancer cell
metastasis. Therefore, lncRNAs are often used as a biological
indicator of cancer to study the molecular mechanism of
tumorigenesis (3,4). In addition, lncRNAs have diverse
functions, that is, lncRNAs can regulate various cellular processes
at transcriptional, post-transcription, translational and
epigenetic level in the forms of signal molecules, scaffolds and
instructor (5,6). Maternally expressed gene 3 (MEG3),
located in human chromosome 14q32.3 with a length of about 1.6 kb
(7), is a lncRNA encoded by a
maternally imprinted gene, MEG3 and the paternally imprinted gene
DLK form the footprint. Studies have shown that the expression of
the MEG3 was downregulated in a variety of human tumor cells, such
as neuroblastoma, meningioma, bladder cancer, gastric cancer and
glioma (8). In addition, due to the
methylation of the promoter and differentially methylated DNA
fragments, the expression of MEG3 can not be detected in some tumor
tissues, such as human functionless adenomas (9). Moreover, MEG3 is a tumor suppressor gene
that can inhibit the development of a tumor in a variety of cell
lines, such as HCT116, HeLa and U87MG cells (10,11).
However, the expression pattern and biological function of MEG3 in
OSCC have not been reported yet.
The WNT/β-catenin signaling pathway is one of the
classical pathways in the process of cell signal transduction, and
the phosphorylation of β-catenin plays a key role in this signal
transduction. The interaction of WNT protein and its receptor on
the cell membrane can inhibit the phosphorylation of β-catenin
induced by GSK-3β, which in turn leads to the accumulation of
β-catenin in cytoplasm, and thus the accumulated β-catenin will
enter the nucleus to interacts with TCF/LEF transcription factor to
activate the expression of downstream target gene, so as to affect
cell proliferation, differentiation and tumorigenesis (12). Studies have shown that small RNA
molecules can regulate carcinogenesis by regulating WNT/β-catenin
signaling pathway (4). This study
investigates how lncRNA MEG3 inhibits the growth and metastasis of
OSCC by regulating WNT/β-catenin signaling pathway and to provide
potential molecular target for the treatment of OSCC.
Materials and methods
Subjects of study
Eighty-three OSCC tumor samples, as well as the
corresponding normal tissues (from the unaffected area of 2 cm from
the tumor) were collected during the period January 2011 to January
2016, and the patients signed an informed consent form.
Additionally, the study was approved by the Ethics Committee of
Xuzhou Stomatological Hospital. The sample consisted of 51 males
and 32 females with an average age of 55.5±6.76 years. High tumor
differentiation was found in 43 cases and middle to low
differentiation was found in 40 cases. Furthermore, 72 cases had
the tumor in tongue and 11 cases in gums, or other parts. None of
the patients received chemotherapy and radiation therapy before
surgery. All the samples were checked and the tumor and normal
tissues were confirmed by three pathologists and samples were
stored at −80°C.
Cells and reagents
Human OSCC cell lines SCC15 and Cal27 were
generously presented by the Department of Stomatology, Wuhan
University (Wuhan, China). Dulbecco's modified Eagle's medium
(DMEM) and fetal bovine serum (FBS) (both from Gibco, Grand Island,
NY, USA); TRIzol reagent, PrimeScript® RT reagent kit
with gDNA Eraser and SYBR® Premix Ex Taq™ II (Takara
Biotechnology Co., Ltd., Dalian, China); RIPA lysate (Thermo Fisher
Scientific, Inc., Carlsbad, CA, USA); protease inhibitors and PVDF
membranes (Roche, Basel, Switzerland); rabbit polyclonal TCF-4
antibody (dilution, 1:500; cat. no. ab185736), rabbit monoclonal
cyclin D1 antibody (dilution, 1:500; cat. no. ab16663), rabbit
polyclonal GAPDH antiboody (dilution, 1:500; cat. no. ab37168) and
secondary goat anti-rabbit (HRP) IgG antibody (dilution, 1:5,000;
cat. no. ab6721) were all purchased from Abcam (Cambridge, MA,
USA). Cell Counting Kit-8 (CCK-8) (Dojindo, Kumamoto, Japan);
Lipofectamine 3000 (Invitrogen Life Technologies, New York, NY,
USA); Annexin V-FITC cell apoptosis detection kit (eBioscience, San
Diego, CA, USA); 5-aza-2′-deoxycytidine (5-aza-CdR; Sigma, New
York, NY, USA); TOP/FOP-Flash luciferase reporter vector
(Millipore, Billerica, MA, USA); Dual-Luciferase assay kit
(Promega, Madison, WI, USA); primers (Sangon, Shanghai, China).
Cell culture and transfection
SCC15 and Cal27 cells were cultured in DMEM (10%
FBS) until the cells adhered; pcDNA3.1-MEG3 overexpression vector
and the si-MEG3 interference vector and its corresponding control
vector pcDNA3.1-Ctrl were constructed, si-Ctrl were transfected
into cells using liposomes. After 48 h, the cells were collected
for subsequent experiments and siRNA was synthesized by GenePharma
(Shanghai, China). The sequences are listed in Table I.
 | Table I.The sequences of siRNA and primers
used in RT-qPCR. |
Table I.
The sequences of siRNA and primers
used in RT-qPCR.
| Primers/siRNA | Sequences |
|---|
| si-MEG3-3 |
5′-CCCUCUUGCUUGUCUUACUTT-3′ |
| MEG3-F |
5′-GCCCTAGGGGAGTGACTACA-3′ |
| MEG3-R |
5′-ACTCGGGACATACCTGCTCT-3′ |
| ACTB-F |
5′-CAGGGCGTGATGGTGGGCA-3′ |
| ACTB-R |
5′-CAAACATCATCTGGGTCATCTTCTC-3′ |
Real-time fluorescence quantitative
PCR (RT-qPCR) analysis
RT-qPCR was used to quantify the relative expression
levels of MEG3 in tissues and cells. Total RNA was extracted using
TRIzol reagent according to the instructions of manuals and cDNA
was synthesized by reverse transcription using
PrimeScript® RT reagent kit with gDNA Eraser. Moreover,
the PCR reaction system was prepared using SYBR® Premix
Ex Taq™ II and PCR reaction was performed using CFX-96 Real-Time
PCR (Bio-Rad, New York, NY, USA). Finally, the data were analyzed
by 2−ΔΔCt method with β-actin used as endogenous
control. All primers are listed in Table
I.
Western blot analysis
The transfected cells were washed twice with
phosphate-buffered saline (PBS) and lysed on ice in RIPA lysates
containing protease inhibitors for 30 min. After that, the protein
was quantified by using BCA method. The protein (50 µg) was
subjected to SDS-PAGE electrophoresis to separate the protein. Then
the protein was transferred to SDS-PAGE by an electric method. The
membrane was blocked by TBST buffer containing 5% skimmed milk for
1 h at room temperature. Then primary rabbit polyclonal TCF-4
antibody (dilution, 1:500; cat. no. ab185736) and rabbit monoclonal
cyclin D1 antibody (dilution, 1:500; cat. no. ab16663) were added
and incubated overnight at 4°C. After washing, secondary goat
anti-rabbit (HRP) IgG antibody (dilution, 1:5,000; cat. no. ab6721)
was added and incubated for 2 h, and after incubation, the signal
was detected by chemiluminescence method.
Methylation treatment
SCC15 and Cal27 (2.5×105) cells were
seeded in a 6-well culture plate with fresh DMEM (containing 0, 5
and 10 µM of 5-aza-CdR) that was changed daily. After incubation
for 5 days, RT-qPCR was used to detect the relative expression of
MEG3.
Cell proliferation and apoptosis
analysis
According to the kit manual instructions, cell
proliferation assay was performed using the CCK-8. The transfected
cells (5×103) were seeded in 96-well plates with 90 µl
medium containing 10% FBS in each well. Then the CCK-8 solution (10
µl/well) was added at 0, 1, 2, 3 and 4 days, after incubation at
37°C for 2 h, and the number of cells in each well was calculated
by measuring the absorbance at 450 nm. Apoptosis analysis was
performed at 48 h after transfection. Cells were harvested from
each well, and Annexin V-FITC Apoptosis Detection kit and BD
FACSCalibur Flow Cell Analyzer (BD Biosciences, New York, NY, USA)
were used to calculate the rate of apoptosis.
Cell migration assay
The artificial basement membrane was added to
24-well plates and DMEM containing transfected SCC15 and Cal27
cells (3×104) was added to the upper chamber.
Furthermore, the lower chamber was filled with DMEM containing 20%
FBS. After incubation for 24 h, the membrane was fixed with 4%
methanol and rinsed with 0.1% crystal violet three times. Finally,
the membrane with the colored cells on the surface was randomly
divided into 5 regions for cell migration analysis.
Analysis of the activity of WNT signal
pathway
TOP-Flash and FOP-Flash luciferase reporter vector
and pcDNA3.1-MEG3 or si-MEG3 vector were co-transfected into OSCC
cells. After incubation for 48 h, luciferase activity was measured
by dual-luciferase assay kit.
Statistical analysis
Statistical analysis was performed using SPSS 17.0
statistical software (SPSS, Inc., Chicago, IL, USA). All
experiments were repeated three times. The data were expressed as
mean ± SD. The two-tailed t-test was applied. P<0.05 was
considered to indicate a statistically significant difference.
Results
The expression of MEG3 is
downregulated in OSCC cells
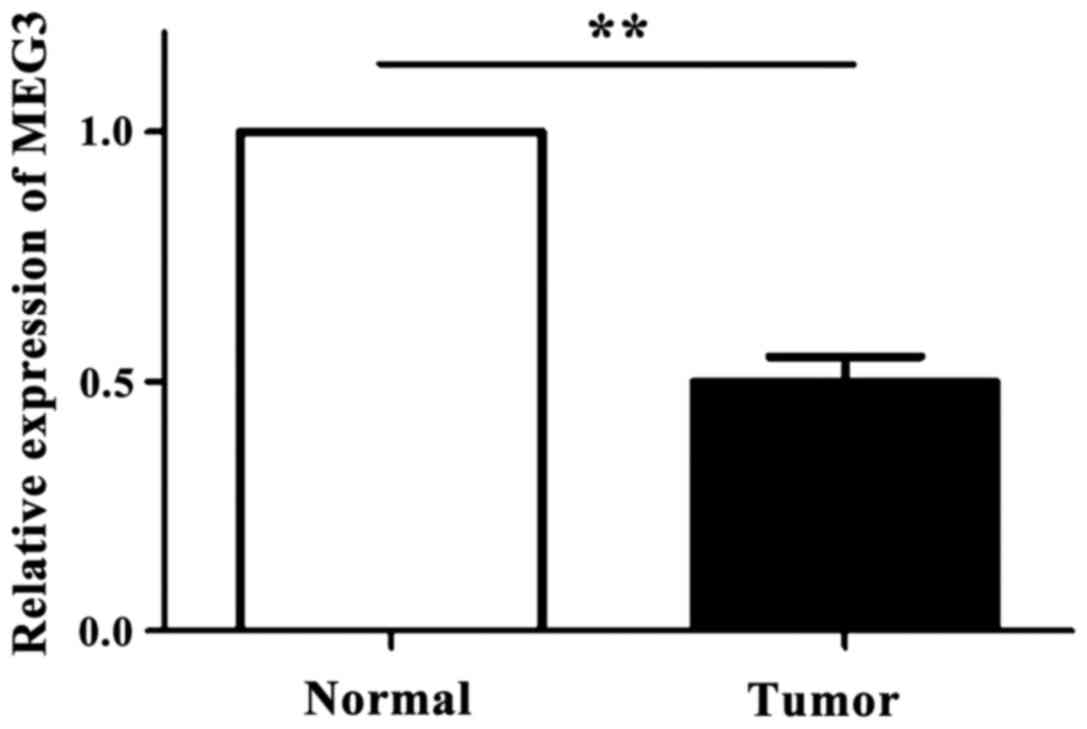
As shown in Fig. 1,
the expression levels of MEG3 in 83 OSCC tissues and the
corresponding normal tissues were detected by RT-qPCR technique. It
was found that the expression level of MEG3 in OSCC tissue was
significantly downregulated compared with normal oral tissue
(p<0.05).
The expression of MEG3 is regulated by
DNA methylation
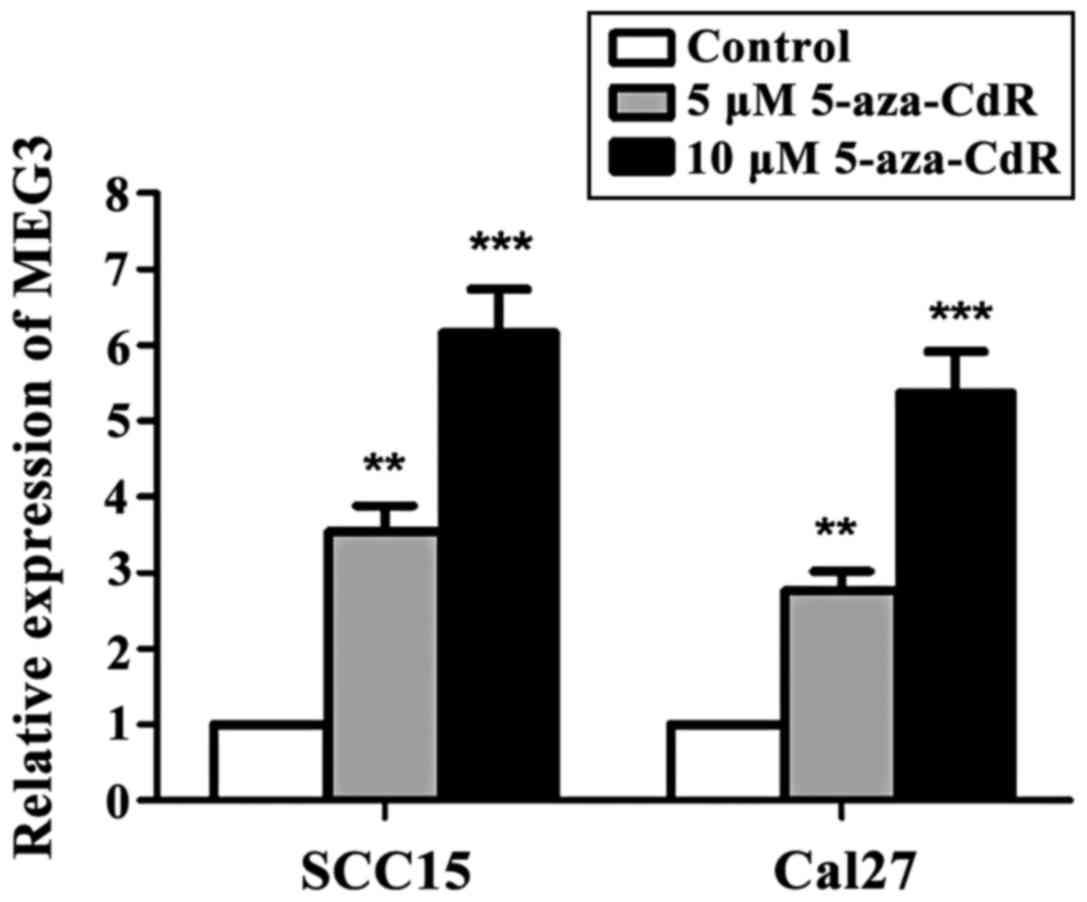
SCC15 and Cal27 cells were treated with different
concentrations of 5-aza-CdR. The cells without 5-aza-CdR treatment
were used as the control group. The results showed that compared
with the control group, the expression of MEG3 in OSCC cells was
significantly increased with the increase in the concentration of
5-aza-CdR (p<0.05) (Fig. 2).
MEG3 inhibits OSCC cell
proliferation
MEG3 overexpression vector pcDNA-MEG3 and the
interfering vector si-MEG3 in the observation group, and the
corresponding control vectors pcDNA-Ctrl and si-Ctrl were
transfected into SCC15 and Cal27 cells, respectively. Then the
CCK-8 was used to analyze the cell proliferation at 1, 2, 3 and 4
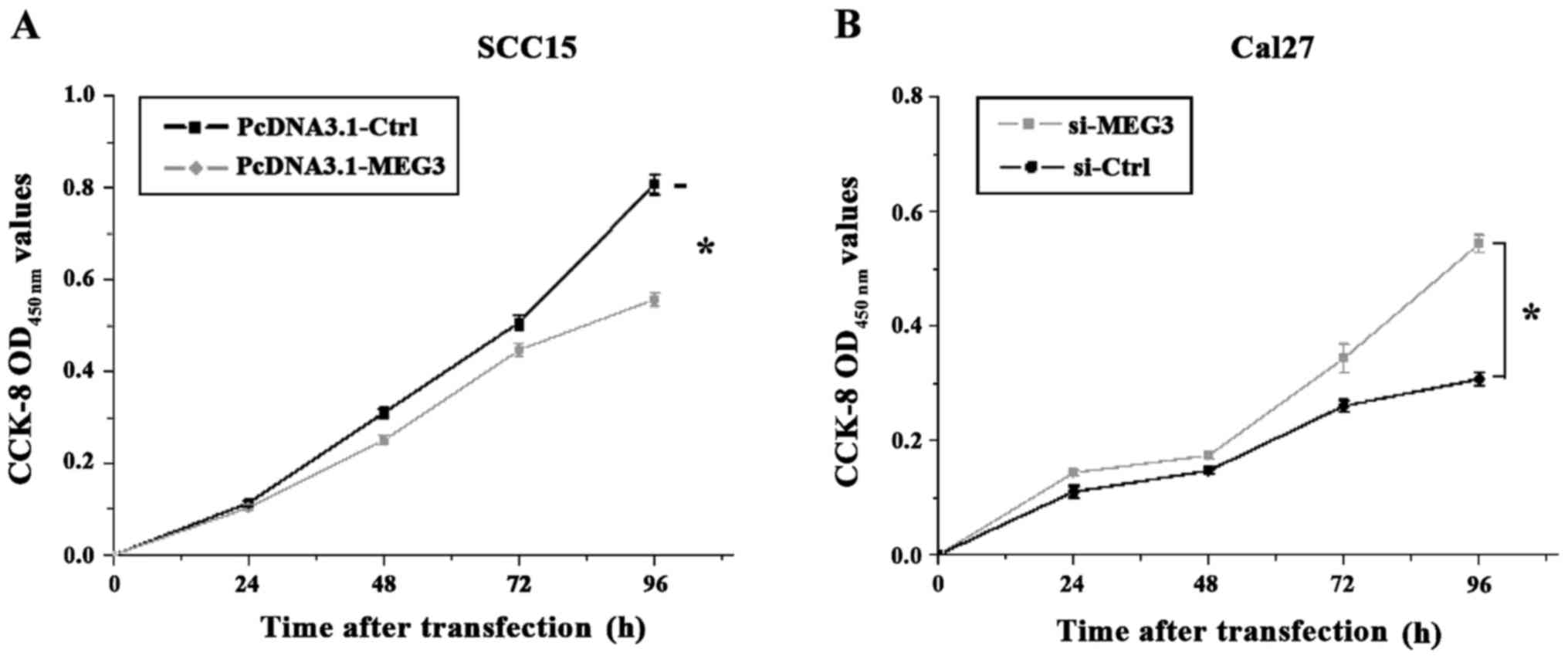
days after transfection. As shown in Fig.
3, the OD value measured at OD450 nm showed that the
proliferation of the SCC15 cells with MEG3 overexpression was
significantly inhibited compared to the control group (p<0.05).
Furthermore, the proliferation rate of the Cal27 cells with reduced
MEG3 expression was significantly increased compared to the control
group (p<0.05).
MEG3 promotes apoptosis of OSCC
cells
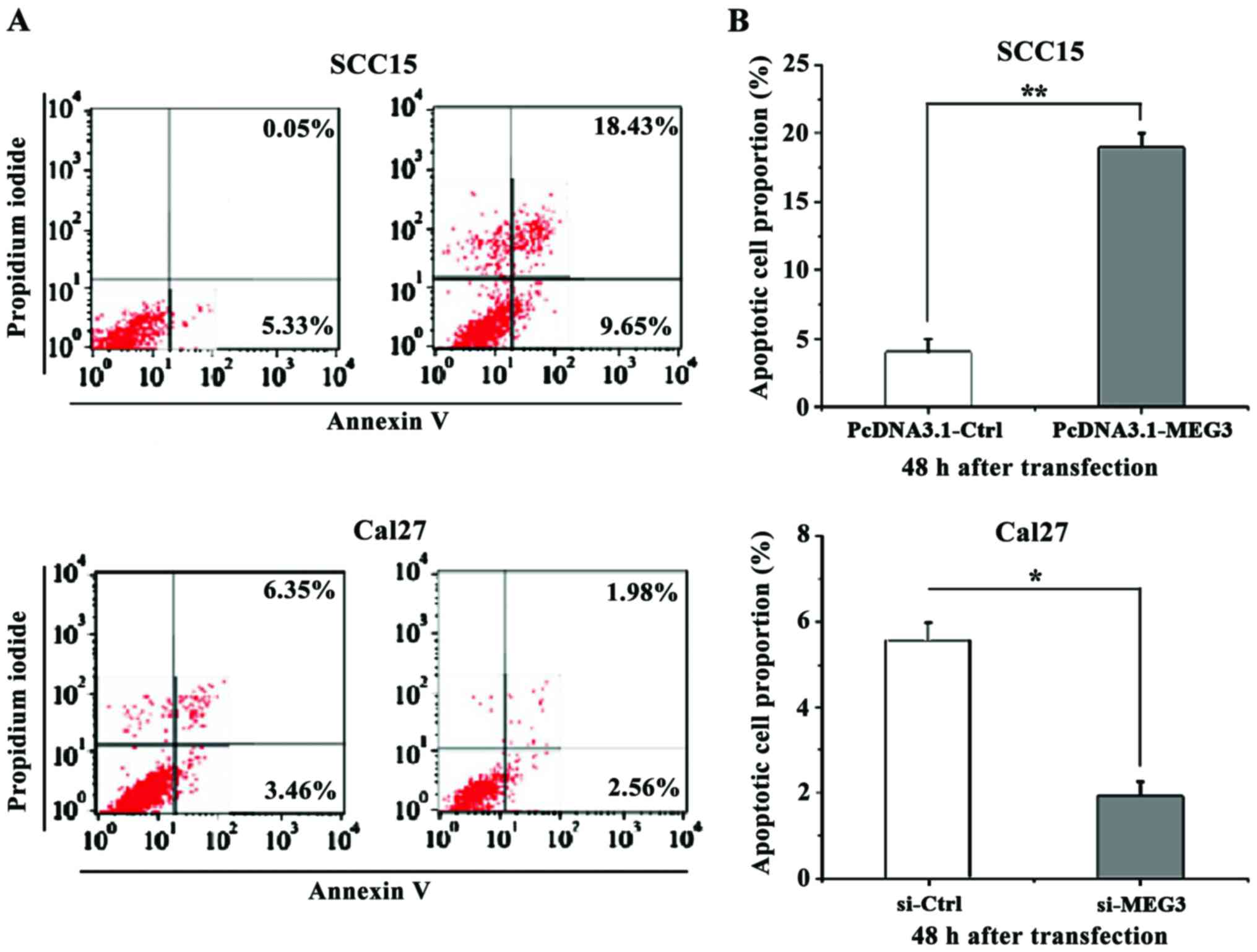
For this, the apoptotic assay was performed at 48 h
after transfection. Then Annexin V-FITC Apoptosis Detection kit and
the Flow Cell Analyzer were used to detect the apoptotic cells, and
the proportion of the apoptotic cells was calculated. The results
showed that MEG3 overexpression in Cal27 cells had significantly
increased the apoptosis rate of OSCC cells compared to the control
group, while the reduced expression level of MEG3 in Cal27 cells
had significantly decreased the apoptosis rate of OSCC cells
compared to the control group (p<0.05) (Fig. 4).
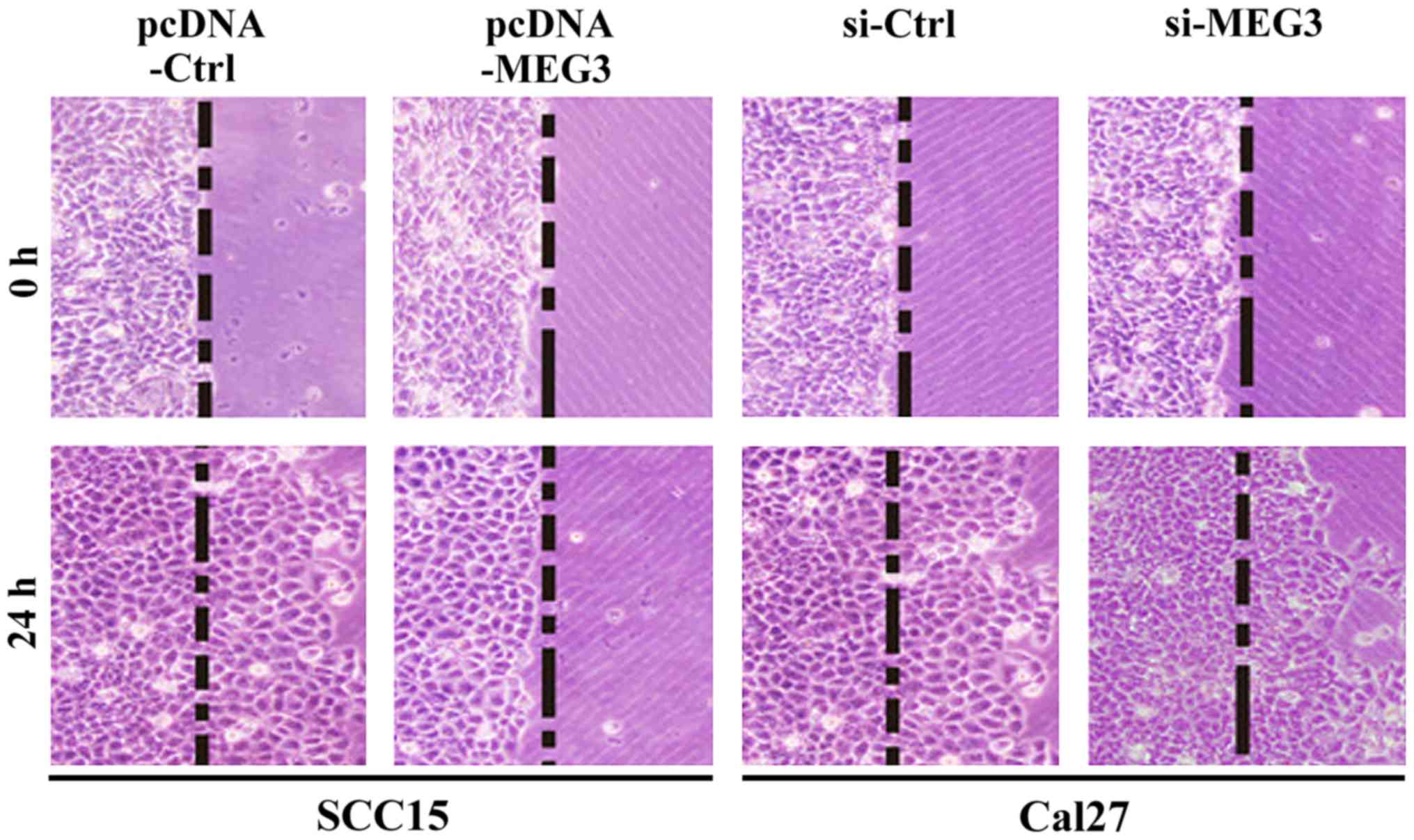
MEG3 inhibits OSCC cell migration
As seen in Fig. 5 the
transfected OSCC cells were stained after incubation on the
artificial basement membrane for 24 h. The results further showed
that the cell migration of SCC15 cells with MEG3 overexpression was
inhibited, whereas the cells in control group showed a certain
degree of migration. The MEG3 silencing in Cal27 cells did not
affect the cell migration, whereas the cells in control group
showed a certain degree of migration. The results indicated that
the expression of MEG3 can inhibit OSCC cell migration.
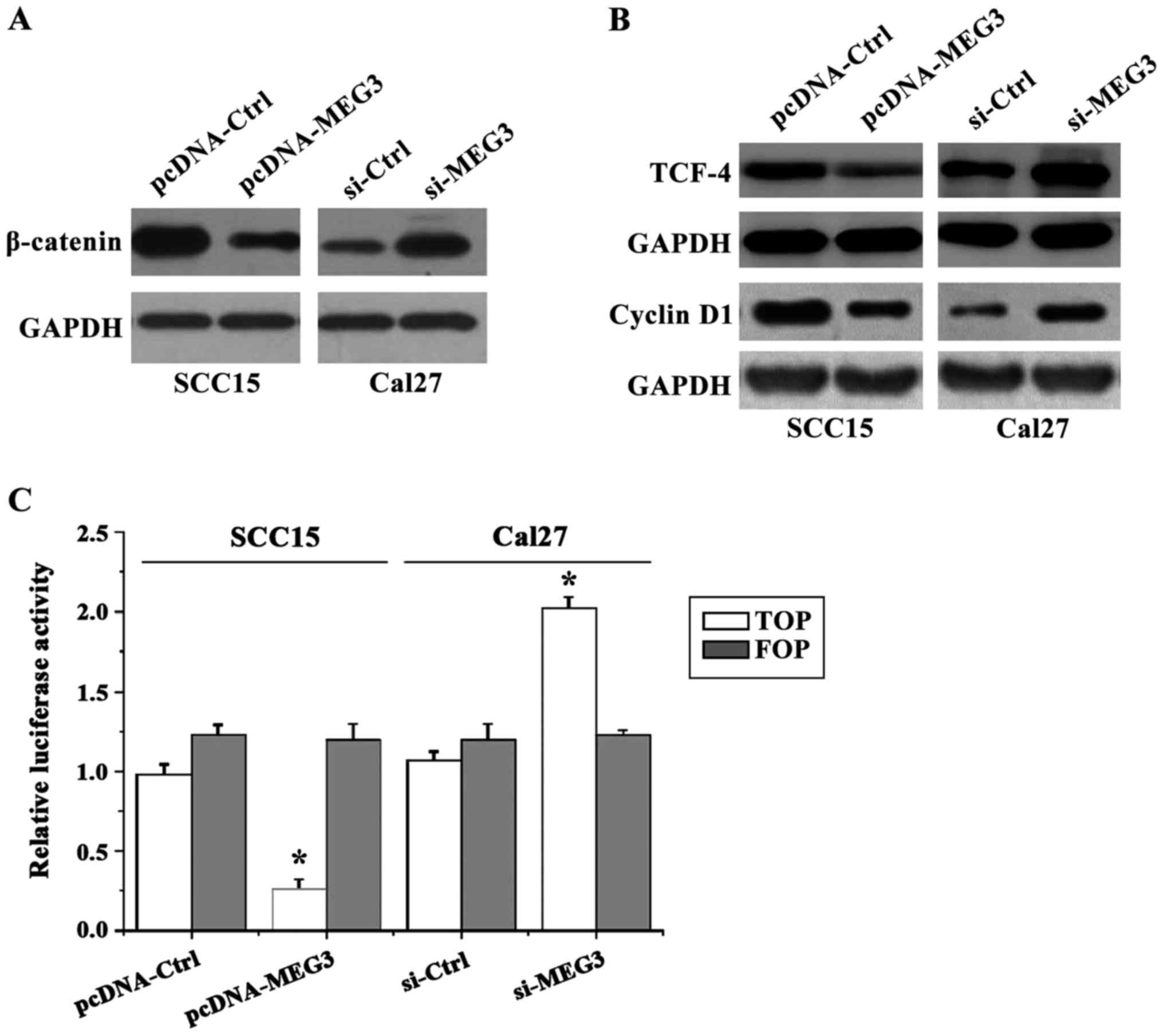
MEG3 regulates the activity of
WNT/β-catenin signaling pathway in OSCC cells
In order to analyze the expression of related
protein and the activity of WNT signal pathway in SCC15 and Cal27
cells after co-transfection of pcDNA3.1-MEG3 or si-MEG3, western
blot analysis and transfection of TOP/FOP flash reporter vector
have been used. The results showed (Fig.
6) that the expression level of WNT pathway core protein
β-catenin in SCC15 cells with MEG3 overexpression was significantly
lower than that in control group, whereas the expression level of
β-catenin in Cal27 cells transfected with si-MEG3 was significantly
higher than that in control group. Since the expression of
β-catenin is directly related to the expression of T-cell
factor/lymphoid enhancer factor (TCF/LEF) superfamily and the
downstream target gene cyclin D1 (cyclin D1), therefore, the
western blot analysis was used to detect the expression levels of
TCF-4 and cyclin D1 protein. The results showed that the expression
patterns of TCF-4 and cyclin D1 protein were similar to that of
β-catenin, which supported our hypothesis. In addition, the results
of TOP/FOP flash showed that the activity of TOP luciferase in
SCC15 cells with MEG3 overexpression was significantly decreased,
that is, the transcription activity of TCF/LEF was decreased
compared to the control group. On the other hand, the activity of
TOP luciferase in Cal27 cells with MEG3 silencing was significantly
increased. The results were consistent with the results of western
blot analysis (FOP-Flash with the mutation on the TCF binding site
was used as a negative control).
Discussion
In the past few decades, studies on the regulatory
role of MEG3 in specific tumorigenesis have shown that MEG3 is
highly expressed in the central nervous system and the expression
level is abnormally low in cancerous tissues (13), therefore, we speculated that MEG3 may
also play an important regulatory role in the tumorigenesis of
OSCC. In our study, we observed a decrease in the expression of
MEG3 in OSCC tissue, which was consistent with the results of
previous studies related to non-functional pituitary adenoma, such
as gliomas, and meningiomas (8). In
addition, epigenetic modifications induced tumor suppressor
silencing through methylation at transcriptional level has
attracted considerable attention in recent years (14). Recent studies showed that
hypermethylation of the MEG3 promoter can inhibit MEG3 expression
(15,16) and this finding has been confirmed for
a variety of tumors (9). Our results
also showed that the expression of MEG3 was increased with the
increase of 5-aza-CdR concentration, which was further confirmed
that MEG3 hypermethylation plays an important role in inhibiting
MEG3 expression in OSCC.
In order to investigate the effects of MEG3
expression on OSCC cell proliferation, metastasis and apoptosis,
pcDNA-MEG3 and si-MEG3 and their control vectors were transfected
into SCC15 and Cal27 cells, respectively. The results showed that
overexpression of MEG3 inhibited OSCC cell proliferation and
metastasis, but promoted apoptosis and vice versa (17). This suggests that MEG3 may act as a
tumor suppressor gene to inhibit OSCC.
The WNT/β-catenin signaling pathway, which regulates
a variety of cellular processes, such as cell proliferation,
invasion, and differentiation, is one of the classical pathways for
signal transduction. The function of WNT/β-catenin signaling
pathway has been extensively studied and reported. Recent studies
have shown that downregulation of MEG3 in lung cancer cells can
activate the WNT pathway (18).
Therefore, in order to investigate the molecular mechanism of MEG3
regulated WNT pathway in OSCC cells, we detected and calculated the
level of WNT pathway's core protein β-catenin by semi-quantitative
analyses, and the activity of WNT pathway was detected using
TOP/FOP flash reporter vector. Interestingly, both experiments
showed that MEG3 levels were negatively correlated with the WNT
pathway, that is, MEG3 inhibited OSCC cell proliferation,
metastasis and promotes apoptosis by negatively regulating the WNT
pathway (19,20). In addition, our findings provide new
insights for future studies to investigate the tumor suppressive
effect of MEG3 through local regulation of WNT/β-catenin signaling
in OSCC cells.
In conclusion, our results suggest that MEG3 can
possibly be used as a new biomarker for the diagnosis and treatment
of the OSCC. It has also been found that DNA hypermethylation can
inhibit MEG3 expression. In addition, MEG3 can inhibit the activity
of OSCC WNT/β-catenin signaling pathway to inhibit tumor
development. Therefore, the full understanding of this newly
discovered functional mechanism of the MEG3 will be helpful for the
diagnosis and treatment of OSCC.
References
|
1
|
Rosebush MS, Rao SK, Samant S, Gu W,
Handorf CR, Pfeffer LM and Nosrat CA: Oral cancer: Enduring
characteristics and emerging trends. J Tenn Dent Assoc. 91:24–29.
2011.PubMed/NCBI
|
|
2
|
Brocklehurst PR, Baker SR and Speight PM:
Oral cancer screening: What have we learnt and what is there still
to achieve? Future Oncol. 6:299–304. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Gibb EA, Brown CJ and Lam WL: The
functional role of long non-coding RNA in human carcinomas. Mol
Cancer. 10:382011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Gutschner T and Diederichs S: The
hallmarks of cancer: A long non-coding RNA point of view. RNA Biol.
9:703–719. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Qiu MT, Hu JW, Yin R and Xu L: Long
noncoding RNA: An emerging paradigm of cancer research. Tumour
Biol. 34:613–620. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Eades G, Zhang YS, Li QL, Xia JX, Yao Y
and Zhou Q: Long non-coding RNAs in stem cells and cancer. World J
Clin Oncol. 5:134–141. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wylie AA, Murphy SK, Orton TC and Jirtle
RL: Novel imprinted DLK1/GTL2 domain on human chromosome 14
contains motifs that mimic those implicated in IGF2/H19 regulation.
Genome Res. 10:1711–1718. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhou Y, Zhang X and Klibanski A: MEG3
noncoding RNA: A tumor suppressor. J Mol Endocrinol. 48:R45–R53.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Balik V, Srovnal J, Sulla I, Kalita O,
Foltanova T, Vaverka M, Hrabalek L and Hajduch M: MEG3: A novel
long noncoding potentially tumour-suppressing RNA in meningiomas. J
Neurooncol. 112:1–8. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhou Y, Zhong Y, Wang Y, Zhang X, Batista
DL, Gejman R, Ansell PJ, Zhao J, Weng C and Klibanski A: Activation
of p53 by MEG3 non-coding RNA. J Biol Chem. 282:24731–24742. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang P, Ren Z and Sun P: Overexpression of
the long non-coding RNA MEG3 impairs in vitro glioma cell
proliferation. J Cell Biochem. 113:1868–1874. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
MacDonald BT, Tamai K and He X:
Wnt/beta-catenin signaling: Components, mechanisms, and diseases.
Dev Cell. 17:9–26. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Schmidt JV, Matteson PG, Jones BK, Guan XJ
and Tilghman SM: The Dlk1 and Gtl2 genes are linked
and reciprocally imprinted. Genes Dev. 14:1997–2002.
2000.PubMed/NCBI
|
|
14
|
Jones PA and Baylin SB: The fundamental
role of epigenetic events in cancer. Nat Rev Genet. 3:415–428.
2002.PubMed/NCBI
|
|
15
|
Modali SD, Parekh VI, Kebebew E and
Agarwal SK: Epigenetic regulation of the lncRNA MEG3 and its target
c-MET in pancreatic neuroendocrine tumors. Mol Endocrinol.
29:224–237. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Braconi C, Kogure T, Valeri N, Huang N,
Nuovo G, Costinean S, Negrini M, Miotto E, Croce CM and Patel T:
microRNA-29 can regulate expression of the long non-coding RNA gene
MEG3 in hepatocellular cancer. Oncogene. 30:4750–4756. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Xiao W, Chen X and He M: Inhibition of the
Jagged/Notch pathway inhibits retinoblastoma cell proliferation via
suppressing the PI3K/Akt, Src, p38MAPK and Wnt/β catenin signaling
pathways. Mol Med Rep. 10:453–458. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Xia Y, He Z, Liu B, Wang P and Chen Y:
Downregulation of Meg3 enhances cisplatin resistance of lung cancer
cells through activation of the WNT/β-catenin signaling pathway.
Mol Med Rep. 12:4530–4537. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Rowe MK and Chuang DM: Lithium
neuroprotection: Molecular mechanisms and clinical implications.
Expert Rev Mol Med. 6:1–18. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Binnerts ME, Kim KA, Bright JM, Patel SM,
Tran K, Zhou M, Leung JM, Liu Y, Lomas WE III, Dixon M, et al:
R-Spondin1 regulates Wnt signaling by inhibiting internalization of
LRP6. Proc Natl Acad Sci USA. 104:14700–14705. 2007. View Article : Google Scholar : PubMed/NCBI
|