|
1
|
GBD 2013 Mortality and Causes of Death
Collaborators: Global, regional, and national levels of age-sex
specific all-cause and cause-specific mortality for 240 causes of
death, 1990–2013: A systematic analysis for the Global Burden of
Disease Study. Lancet. 385:117–171. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Landgren O, Kyle RA, Pfeiffer RM, Katzmann
JA, Caporaso NE, Hayes RB, Dispenzieri A, Kumar S, Clark RJ, Baris
D, et al: Monoclonal gammopathy of undetermined significance (MGUS)
consistently precedes multiple myeloma: A prospective study. Blood.
113:5412–5417. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Berenson JR, Matous J, Swift RA, Mapes R,
Morrison B and Yeh HS: A phase I/II study of arsenic
trioxide/bortezomib/ascorbic acid combination therapy for the
treatment of relapsed or refractory multiple myeloma. Clin Cancer
Res. 13:1762–1768. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Agarwal A and Ghobrial IM: Monoclonal
gammopathy of undetermined significance and smoldering multiple
myeloma: A review of the current understanding of epidemiology,
biology, risk stratification, and management of myeloma precursor
disease. Clin Cancer Res. 19:985–994. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Landgren O: Monoclonal gammopathy of
undetermined significance and smoldering multiple myeloma:
Biological insights and early treatment strategies. Hematology Am
Soc Hematol Educ Program. 2013:478–487. 2013.PubMed/NCBI
|
|
6
|
Kubiczkova L, Kryukov F, Slaby O,
Dementyeva E, Jarkovsky J, Nekvindova J, Radova L, Greslikova H,
Kuglik P, Vetesnikova E, et al: Circulating serum microRNAs as
novel diagnostic and prognostic biomarkers for multiple myeloma and
monoclonal gammopathy of undetermined significance. Haematologica.
99:511–518. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Shvartsur A, Chatterjee D, Chen H,
Berenson J, Garban H and Bonavida B: A bioinformatics approach
revealed a differential expression of RKIP-related genes in
pre-multiple myeloma and multiple myeloma: Clinical implications.
Blood. 126:53092015.
|
|
8
|
Amend S, Liang C, Serie D, Vachon CM, Lu
L, Vij R, Colditz GA, Weilbaecher KN and Tomasson MH: Deletion of
samsn1 underlies genetic susceptibility to Monoclonal Gammopathy of
undetermined significance (MGUS) in mice. Blood. 122:3972013.
|
|
9
|
Kuehl WM and Bergsagel PL: Molecular
pathogenesis of multiple myeloma and its premalignant precursor. J
Clin Invest. 122:3456–3463. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bergsagel PL, Kuehl WM, Zhan F, Sawyer J,
Barlogie B and Jr SJ: Cyclin D dysregulation: An early and unifying
pathogenic event in multiple myeloma. Blood. 106:296–303. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Annunziata CM, Davis RE, Demchenko Y,
Bellamy W, Gabrea A, Zhan F, Lenz G, Hanamura I, Wright G, Xiao W,
et al: Frequent engagement of the classical and alternative
NF-kappaB pathways by diverse genetic abnormalities in multiple
myeloma. Cancer Cell. 12:115–130. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
McNee G, Eales KL, Wei W, Williams DS,
Barkhuizen A, Bartlett DB, Essex S, Anandram S, Filer A, Moss PA,
et al: Citrullination of histone H3 drives IL-6 production by bone
marrow mesenchymal stem cells in MGUS and multiple myeloma.
Leukemia. 31:373–381. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Clough E and Barrett T: The gene
expression omnibus database. Methods Mol Biol. 1418:93–110. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
MacDonald JW: Huex10sttranscriptcluster.
db: Affymetrix huex10 annotation data (chip
huex10sttranscriptcluster). R package. version 8.2.0. 2017.
|
|
15
|
Gentleman R: Annotate: Annotation for
microarrays. R package. version 1.54.0. 2017.
|
|
16
|
R Development Core Team: R: A language and
environment for statistical computingR Foundation for Statistical
Computing. Vienna, Austria: 2017
|
|
17
|
Smyth GK; Gentleman R, Carey V, Dudoit S,
Irizarry R and Huber W: Limma: Linear models for microarray
dataBioinformatics and Computational Biology Solutions Using R and
Bioconductor. Springer; New York: pp. 397–420. 2005, View Article : Google Scholar
|
|
18
|
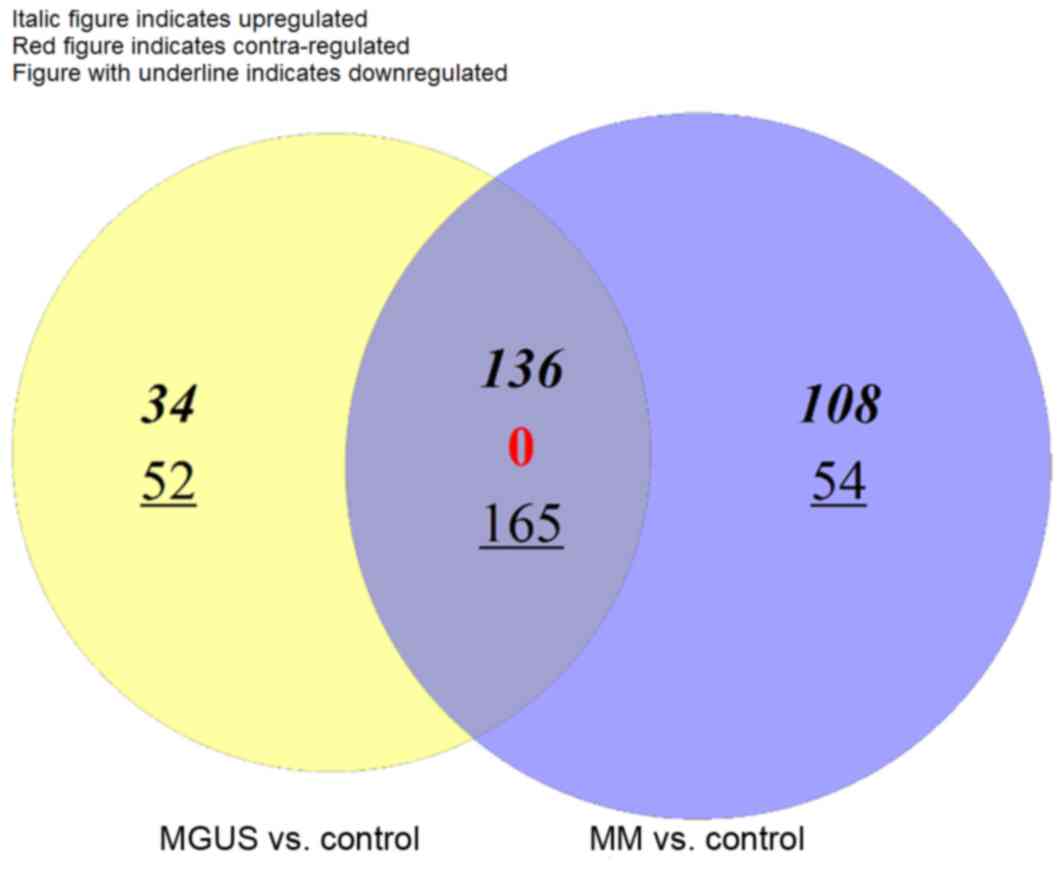
Chen H and Boutros PC: VennDiagram: A
package for the generation of highly-customizable Venn and Euler
diagrams in R. Bmc Bioinformatics. 12:352011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kanehisa M, Goto S, Sato Y, Furumichi M
and Tanabe M: KEGG for Integration and Interpretation of
large-scale molecular data sets. Nucleic Acids Res. 40:(Database
issue). D109–D114. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Dennis G, Sherman BT, Hosack DA, Yang J,
Gao W, Lane HC and Lempicki RA: DAVID: Database for annotation,
visualization, and integrated discovery. Genome Biol. 4:P32003.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
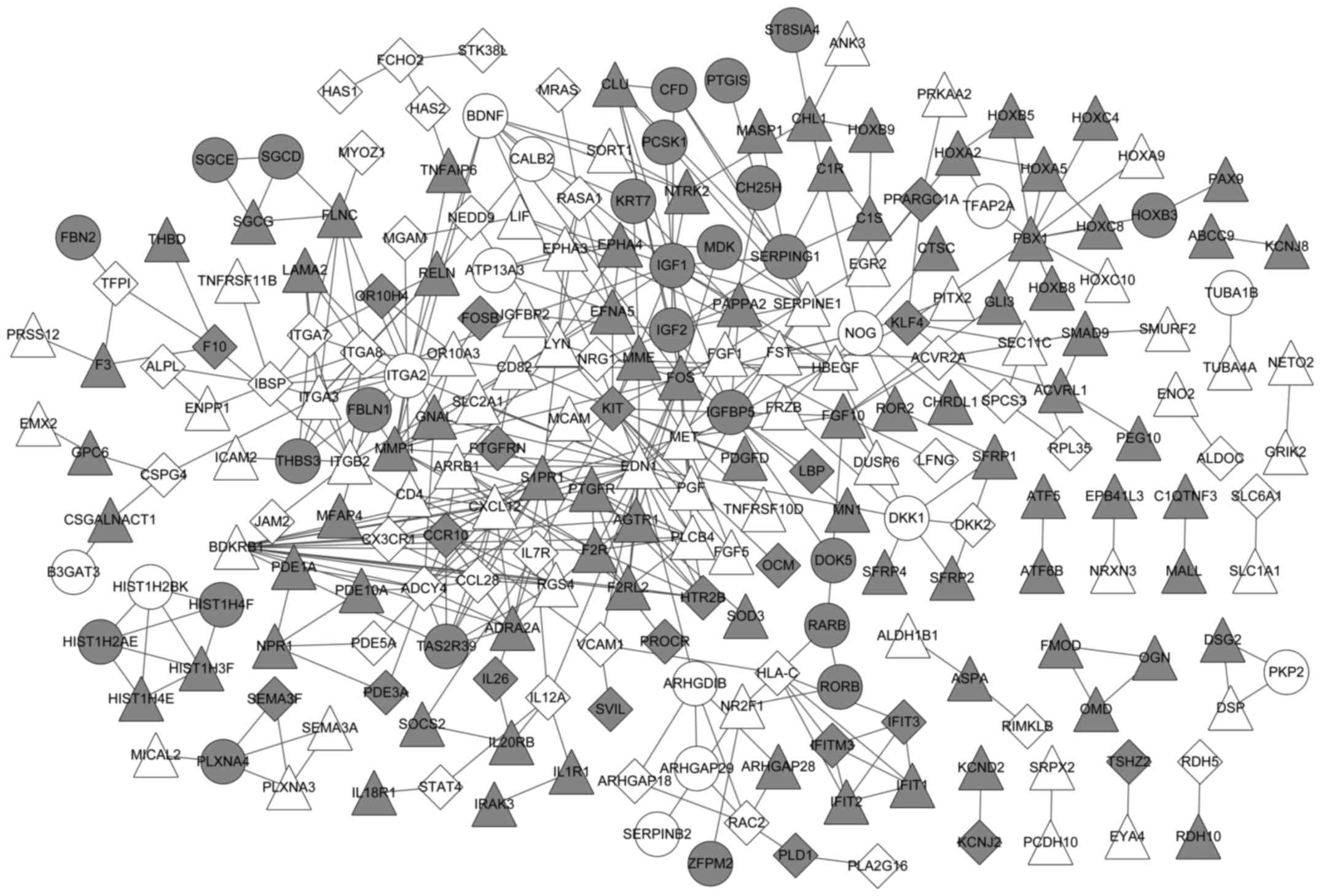
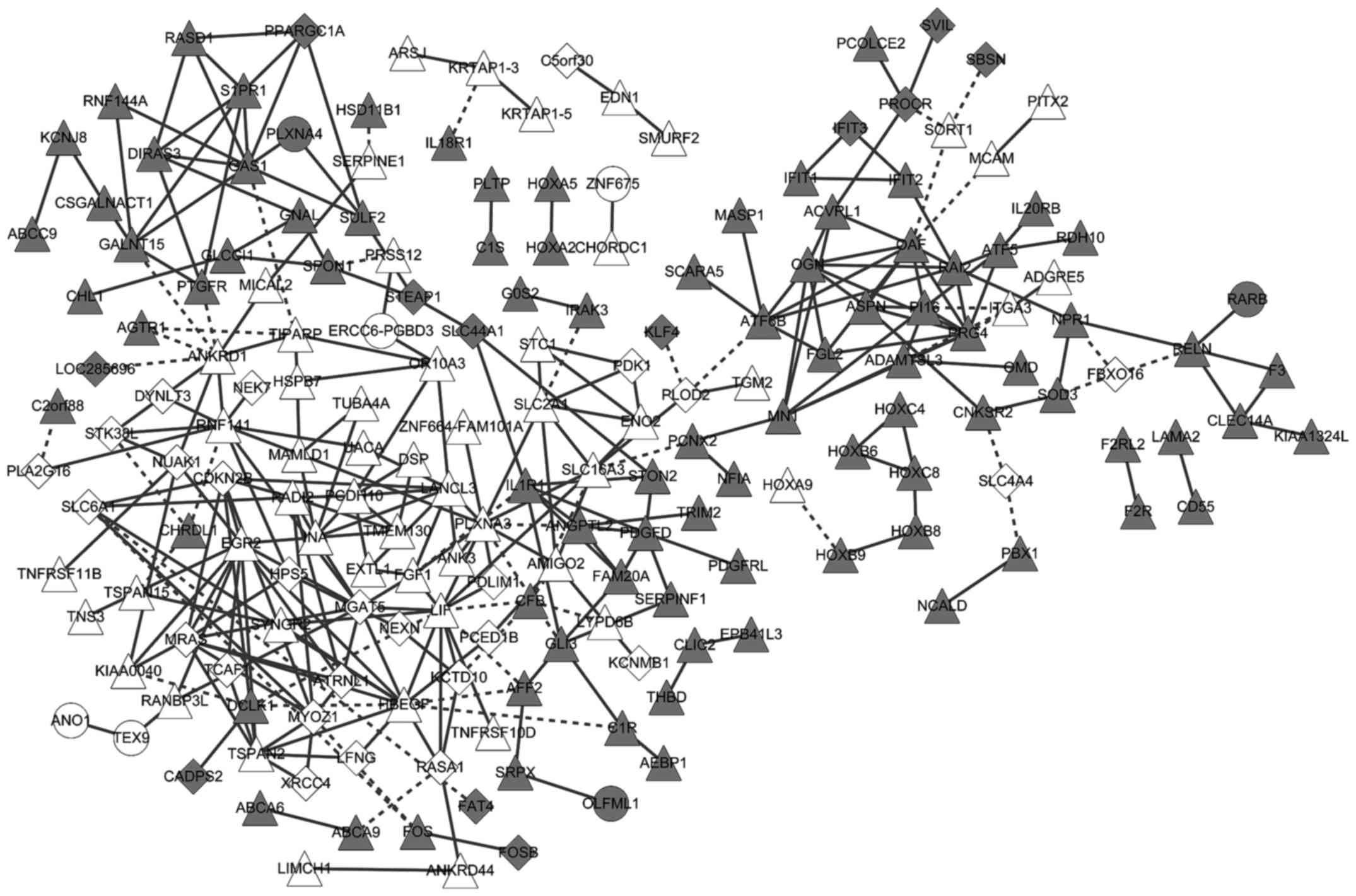
Jensen LJ, Kuhn M, Stark M, Chaffron S,
Creevey C, Muller J, Doerks T, Julien P, Roth A, Simonovic M, et
al: STRING 8-a global view on proteins and their functional
interactions in 630 organisms. Nucleic Acids Res. 37:(Database
issue). D412–D416. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Tang Y, Li M, Wang J, Pan Y and Wu FX:
CytoNCA: A cytoscape plugin for centrality analysis and evaluation
of protein interaction networks. Biosystems. 127:67–72. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Goh KI, Oh E, Kahng B and Kim D:
Betweenness centrality correlation in social networks. Phys Rev E
Stat Nonlin Soft Matter Phys. 67:0171012003. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Du Y, Gao C, Chen X, Hu Y, Sadiq R and
Deng Y: A new closeness centrality measure via effective distance
in complex networks. Chaos. 25:0331122015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Estrada E and Rodríguezvelázquez JA:
Subgraph centrality in complex networks. Phys Rev E Stat Nonlin
Soft Matter Phys. 71:0561032005. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Liang M, Zhang F, Jin G and Zhu J:
FastGCN: A GPU accelerated tool for fast gene co-expression
networks. PloS One. 10:e01167762015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
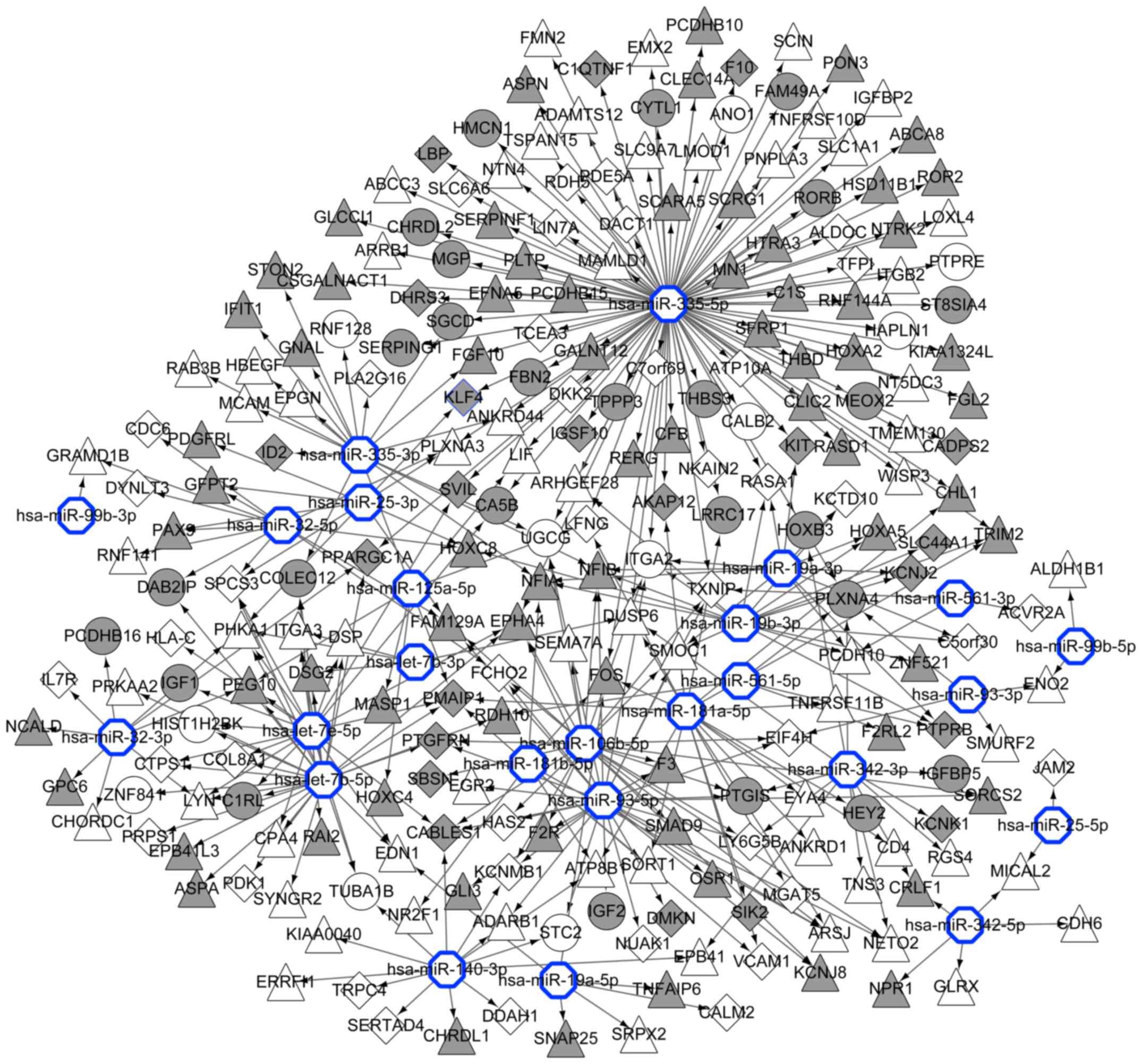
Jiang Q, Wang Y, Hao Y, Juan L, Teng M,
Zhang X, Li M, Wang G and Liu Y: miR2Disease: A manually curated
database for microRNA deregulation in human disease. Nucleic Acids
Res. 37:(Suppl 1). D98–D104. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Dweep H and Gretz N: miRWalk2.0: A
comprehensive atlas of microRNA-target interactions. Nat Methods.
12:6972015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Weiss BM, Abadie J, Verma P, Howard RS and
Kuehl WM: A monoclonal gammopathy precedes multiple myeloma in most
patients. Blood. 113:5418–5422. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Spendlove I, Ramage JM, Bradley R, Harris
C and Durrant LG: Complement decay accelerating factor (DAF)/CD55
in cancer. Cancer Immunol Immunother. 55:987–995. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Matsushita M, Thiel S, Jensenius JC, Terai
I and Fujita T: Proteolytic activities of two types of
mannose-binding lectin-associated serine protease. J Immunol.
165:2637–2642. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Markiewski MM, Nilsson B, Ekdahl KN,
Mollnes TE and Lambris JD: Complement and coagulation: Strangers or
partners in crime? Trends Immunol. 28:184–192. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Crowely MP, Quinn S, Coleman E, Eustace
JA, Gilligan OM and O'Shea SI: Differing coagulation profiles of
patients with monoclonal gammopathy of undetermined significance
and multiple myeloma. J Thromb Thrombolysis. 39:245–249. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
De Stefano V, Za T and Rossi E: Venous
thromboembolism in multiple myeloma. Semin Thromb Hemost.
40:338–347. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Kaelin WG Jr and Ratcliffe PJ: Oxygen
sensing by metazoans: The central role of the HIF hydroxylase
pathway. Mol Cell. 30:393–402. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Martin SK, Diamond P, Gronthos S, Peet DJ
and Zannettino AC: The emerging role of hypoxia, HIF-1 and HIF-2 in
multiple myeloma. Leukemia. 25:1533–1542. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Asosingh K, De Raeve H, de Ridder M,
Storme GA, Willems A, Van Riet I, Van Camp B and Vanderkerken K:
Role of the hypoxic bone marrow microenvironment in 5T2 MM murine
myeloma tumor progression. Haematologica. 90:810–817.
2005.PubMed/NCBI
|
|
38
|
Azab AK, Hu J, Quang P, Azab F,
Pitsillides C, Awwad R, Thompson B, Maiso P, Sun JD, Hart CP, et
al: Hypoxia promotes dissemination of multiple myeloma through
acquisition of epithelial to mesenchymal transition-like features.
Blood. 119:5782–5794. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Hu Y, Kirito K, Yoshida K, Mitsumori T,
Nakajima K, Nozaki Y, Hamanaka S, Nagashima T, Kunitama M, Sakoe K
and Komatsu N: Inhibition of hypoxia-inducible factor-1 function
enhances the sensitivity of multiple myeloma cells to melphalan.
Mol Cancer Ther. 8:2329–2338. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lee C, Oh JI, Park J, Choi JH, Bae EK, Lee
HJ, Jung WJ, Lee DS, Ahn KS and Yoon SS: TNF α mediated IL-6
secretion is regulated by JAK/STAT pathway but Not by MEK
phosphorylation and AKT phosphorylation in U266 multiple myeloma
cells. Biomed Res Int. 2013:5801352013. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Mitsiades CS, Mitsiades N, Poulaki V,
Schlossman R, Akiyama M, Chauhan D, Hideshima T, Treon SP, Munshi
NC, Richardson PG and Anderson KC: Activation of NF-kappaB and
upregulation of intracellular anti-apoptotic proteins via the
IGF-1/Akt signaling in human multiple myeloma cells: Therapeutic
implications. Oncogene. 21:5673–5683. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Elton TS, Oparil S, Taylor GR, Hicks PH,
Yang RH, Jin H and Chen YF: Normobaric hypoxia stimulates
endothelin-1 gene expression in the rat. Am J Physiol.
263:R1260–1264. 1992.PubMed/NCBI
|