Introduction
Breast cancer is one of the leading causes of
cancer-associated mortality in females worldwide (1). Breast cancer mortality is largely due to
metastasis, which begins with the invasion and migration of cells
into the surrounding tissues and vasculature (2). To date, the molecular mechanisms
underlying breast cancer metastasis are poorly understood.
Carcinoembryonic antigen (CEA)-related cell adhesion molecule 1
(CEACAM1), formerly termed BGP, C-CAM or CD66a, is a member of the
CEA family, which belongs to the immunoglobulin superfamily
(3). CEACAM1 is a multifunctional
molecule with a broad distribution in human epithelia, endothelia
and hematopoietic cells (4). A
growing body of evidence suggests that CEACAM1 expression is
altered in numerous types of cancer. Downregulation of CEACAM1
expression has been reported in various types of malignancies,
including colorectal cancer (5),
hepatocellular cancer (6), renal
carcinoma (7) and prostate cancer
(8). Based on these findings, a tumor
suppressive role for CEACAM1 was postulated (9,10). Several
studies have reported a decreased expression of CEACAM1 in breast
cancer (11–13). In addition, CEACAM1 was revealed to
have an essential role in human mammary gland morphogenesis
(14). Notably, CEACAM1-4S, a major
isoform of CEAMCA1, can revert breast cancer cells to a normal
morphogenic phenotype (15). These
findings indicate that CEACAM1 may perform a pivotal role in breast
cancer initiation and progression. However, knowledge on the
functions of CEACAM1 in breast cancer remains limited.
The present study aimed to reveal the functions of
CEACAM1 in breast cancer. First, the pattern of expression of
CEACAM1 was observed, and its association with cancer metastasis
was evaluated. As CEACAM1-4S is the main CEACAM1 isoforms and acts
as the main modulator in cellular and molecular regulation in human
breast epithelia (15), subsequent
experiments concentrated on CEACAM1-4S to investigate the effect of
CEACAM1-4S on the invasive behavior of breast cancer cells.
Materials and methods
Patients and specimens
A total of 62 breast cancer tissue samples were
collected from patients who underwent surgical resection at
Shanghai Jiao Tong University Affiliated Sixth People's Hospital
(Shanghai, China) between October 2013 and February 2015. All
patients were female, with a mean age of 56.6 years (range, 26–78
years). None of the enrolled patients had undergone chemotherapy or
radiotherapy prior to surgery. The histological types were
evaluated according to the World Health Organization criteria
(16). The clinical information of
the patients was obtained from medical records. All patients
provided informed consent prior to involvement in the present
study. The present study was approved by the Ethics Committee of
Shanghai Jiao Tong University in accordance with The Declaration of
Helsinki.
Cell lines
Human breast cancer cell lines with different
metastatic potential (BT549, Hs578T, T-47D, MCF7, MDA-MB-231 and
MDA-MB-468) were purchased from the Institute of Biochemistry and
Cell Biology at the Chinese Academy of Science (Shanghai, China).
MCF10A, a non-invasive human immortal mammary epithelial cell line
derived from a patient with fibrocystic breast disease, was
obtained from the American Type Culture Collection (Manassas, VA,
USA). All cell lines were cultured according to the supplier's
protocol. BT-549 and T-47D cells were cultured in RPMI-1640 medium
(Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA), Hs578T
and MCF7 cells were cultured in Dulbecco's modified eagle's medium
(Gibco; Thermo Fisher Scientific, Inc.), and MDA-MB-231 and
MDA-MB-468 cells were cultured in Leibovitz's L-15 medium (Gibco;
Thermo Fisher Scientific, Inc.). MCF10A cells was cultured in
mammary epithelial growth media (Lonza Group, Ltd., Basel,
Switzerland). All cell lines were cultured at 37°C in a humidified
atmosphere containing 95% air and 5% CO2, and media were
supplemented with 10% fetal bovine serum (Gibco; Thermo Fisher
Scientific, Inc.).
Immunohistochemistry (IHC)
Immunohistochemical staining for CEACAM1 was
performed on breast cancer tissue specimens. The tissue samples
were sliced 4-µm thick and fixed in 4% (w/v) paraformaldehyde in
0.1 M PBS overnight at 4°C, dehydrated in increasing concentrations
of ethanol, and then embedded in paraffin. Briefly, the
paraffin-embedded tissue sections were de-waxed, rehydrated in
descending concentrations of ethanol, and retrieved in a water bath
at 95°C, followed by incubation overnight at 4°C with primary
anti-CEACAM1 antibody (1:75; ab49510; Abcam, Cambridge, UK) in a
humidified chamber. Next, the sections were incubated with a
biotinylated secondary antibody (1:100; K4061; Dako; Agilent
Technologies, Inc., Santa Clara, CA, USA) for 1 h at room
temperature. Subsequent to rinsing in PBS, the
streptavidin-peroxidase complex reagent [StrepABComplex/horseradish
peroxidase (HRP) Duet; Dako; Agilent Technologies, Inc.] was added.
Finally, sections were visualized with 3,3′-diaminobenzidine,
followed by hematoxylin counterstaining for 5 min at room
temperature. For negative controls, the primary antibody was
omitted. The images were obtained using a microscope (Nikon
Eclipse80i; Nikon Corporation, Tokyo, Japan) at ×200 magnification.
The intensity of CEACAM1 staining was quantitatively evaluated
using Image Pro-Plus 6.0 software (Media Cybernetics, Inc.,
Rockville, MD, USA).
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Quantitative real-time polymerase chain reaction
(qPCR) was performed using SYBR® Green Premix Ex Taq™
(Tli RNaseH Plus) kit (RR420A; Takara Bio Inc., Shiga, Japan). The
cells were collected, and total RNA was isolated using TRIzol
reagent (Invitrogen; Thermo Fisher Scientific, Inc.). The synthesis
of complementary DNA was performed with the PrimeScript RT reagent
kit (Takara Biotechnology, Shiga, Japan), according to the
manufacturer's protocol. qPCR was performed on a thermal cycler
ABI7500 (Applied Biosystems; Thermo Fisher Scientific, Inc.) using
specific primers for CEACAM1-4S (NCBI GenBank accession no.
NM_00104912) (forward, 5′-AAACCAGAGTCTCCCGTCCT-3′; reverse,
5′-TGGAGTGGTCCTGAGCTGCCG-3′). The amplification conditions were as
follows: 95°C for 30 sec, 95°C for 5 sec and 64°C for 34 sec, for
40 cycles. In addition, melting curve was generated to validate the
specificity of the analysis. For quantitation of CEACAM1-4S mRNA,
the 2−ΔΔCq
(−ΔCq=CqGAPDH-CqCEACAM1) method (17) was employed. The relative amount of
CEACAM1-4S mRNA was calculated by normalization to the housekeeping
gene GAPDH. The primer sequences for GAPDH were forward,
5′-GCACCGTCAAGGCTGAGAAC-3′ and reverse,
5′-ATGGTGGTGAAGACGCCAGT-3′.
Cell transfection
Plasmids containing the CEACAM1-4S gene and green
fluorescent protein (GFP) reporter gene were constructed by OriGene
Technologies, Inc. (Rockville, MD, USA) as previously described by
Oliveira-Ferrer et al (18).
Empty-vector (PS100071; OriGene Technologies, Inc.) was used as a
control. Cultured breast cancer BT549 or Hs578T cells were seeded
in 6-well plates at 3×105 cells/well and transfected
with 1.25 µg plasmid DNA using Opti-MEM Reduced Serum medium
(Invitrogen; Thermo Fisher Scientific, Inc.) and Lipofectamine Plus
reagent (Invitrogen; Thermo Fisher Scientific, Inc.) according to
the manufacturer's protocol. The transfection efficiency was
monitored using the GFP reporter gene. The result of the
GFP-positive cells counting revealed transfection efficiency
ranging between 50 and 75%. At 6 h post-transfection, the cells
were washed and incubated in complete growth medium (RPMI-1640
medium for BT549 and Dulbecco's modified Eagle's medium for
Hs578T). After 48 h, the cells were lysed for western blot analysis
or used in subsequent experiments.
Western blot analysis
The cells were harvested using a curet and
centrifuged at 3,500 × g for 10 min at 4°C and then lysed in
ice-cold radioimmunoprecipitation assay lysis buffer (P0013B;
Beyotime Institute of Biotechnology, Shanghai, China). Equal
amounts of protein (30 µg) were separated by SDS-PAGE (10% gel) and
subsequently transferred to a polyvinylidene difluoride membrane.
Subsequent to blocking with 5% skimmed milk at room temperature for
1 h, the membranes were incubated at 4°C overnight with primary
antibodies, including anti-CEACAM1 (1:1,000; ab49510; Abcam),
anti-N-cadherin (1:500; sc8424; Santa Cruz Biotechnology, Inc.,
Dallas, TX, USA), anti-E-cadherin (1:500; sc8426; Santa Cruz
Biotechnology, Inc.), anti-vimentin (1:1,000; #3932; Cell Signaling
Technology, Inc., Danvers, MA, USA), anti-signal transducer and
activator of transcription 3 (STAT3; 1:1,000; #9139; Cell Signaling
Technology, Inc.), anti-Smad2/3 (1:1,000; #8685; Cell Signaling
Technology, Inc.), anti-phospho-STAT3 (Tyr705; 1:1,000; #4113; Cell
Signaling Technology, Inc.), anti-phospho-Smad2 (Ser465/467;
1:1,000; #3108; Cell Signaling Technology, Inc.), anti-matrix
metalloproteinase (MMP)2 (1:1,000; #4022; Cell Signaling
Technology, Inc.), anti-MMP9 (1:1,000; #3852; Cell Signaling
Technology, Inc.), anti-tissue inhibitor of metalloproteinase
(TIMP)1 (1:1,000; #8946; Cell Signaling Technology, Inc.),
anti-TIMP2 (1:1,000; #5738; Cell Signaling Technology, Inc.) and
anti-GAPDH (1:1,000; ab9484; Abcam), followed by incubation at room
temperature for 2 h with HRP-conjugated polyclonal secondary
antibody (1:5,000; ab6789/ab6721; Abcam). All western blots were
visualized using the enhanced plus chemiluminescence assay kit (EMD
Millipore, Billerica, MA, USA), according to the manufacturer's
protocol.
Transwell invasion assay
Cell invasion was investigated using Matrigel
invasion chambers with a pore size of 8 mm (Costar; Corning Life
Sciences, Cambridge, MA, USA). Briefly, BT549 or Hs578T cells
(4×104 cells per chamber) in serum-free medium were
seeded in the upper chamber, and 10% fetal bovine serum (FBS)
(Gibco; Thermo Fisher Scientific, Inc.) was used as a
chemoattractant in the bottom well. After incubation for 24 h at
37°C, the non-invasive cells on the upper surface of the membrane
were removed with a cotton swab, and the invasive cells on the
bottom side were fixed in 100% methanol at room temperature for 5
min, stained with 1% crystal violet at room temperature for 10 min
and counted using a microscope (Nikon Eclipse80i; Nikon
Corporation) under ×100 magnification with five fields of view per
sample.
Wound healing assay
Wound healing assays were performed to investigate
the cell migration. BT549 or Hs578T cells were cultured on 6-well
plates (3×105 cells/well) until 100% confluence was
reached. The cultures were then scratched to form a wound line
using a pipette tip. Subsequent to washing with PBS, the cells were
cultured in media containing 1% FBS for an additional 24 h. The
wound area was observed by inverted phase contrast microscopy
(magnification, ×200; Nikon Corporation). The migration area was
calculated by subtracting the unhealed area at 24 h from the
initial wound area.
Immunofluorescent staining (IF)
For immunofluorescence, BT549 or Hs578T cells were
seeded on coverslips (2×104 cells/well; 24-well plates).
The next day, the cells were fixed with 100% cold methanol at room
temperature for 10 min and subsequently blocked with 1% bovine
serum albumin (ST023; Beyotime Institute of Biotechnology) in PBS
at room temperature for 1 h, followed by overnight incubation at
4°C with the primary antibodies: Anti-E-cadherin (1:100; sc8426;
Santa Cruz Biotechnology, Inc.) and anti-vimentin (1:200; #3932;
Cell Signaling Technology, Inc.). Following washing using PBS three
times, the cells were incubated at room temperature for 1 h with
Alexa Fluor 594-conjugated secondary antibody (016580084; Jackson
ImmunoResearch Laboratories, Inc., West Grove, PA, USA) at a
dilution of 1:400. The nuclei were stained with DAPI (Invitrogen;
Thermo Fisher Scientific, Inc.) at room temperature for 10 min in
the dark. Images were captured using an inverted fluorescence
microscope (magnification, ×200; Nikon Corporation). For negative
controls, the primary antibody incubation step was omitted. The
Image Pro-Plus version 6.0 software (Media Cybernetics, Inc.) was
used for imaging analysis.
Statistical analysis
All data are expressed as the mean ± standard
deviation and represent at least three experiments unless stated
otherwise. The statistical significance of the differences between
two different groups was determined with Student's t-test or
nonparametric Mann-Whitney test using SPSS19.0 statistics software
(IBM SPSS, Armonk, NY, USA). P<0.05 was considered to indicate a
statistically significant difference.
Results
CEACAM1 expression is inversely
associated with metastasis in patients with breast cancer
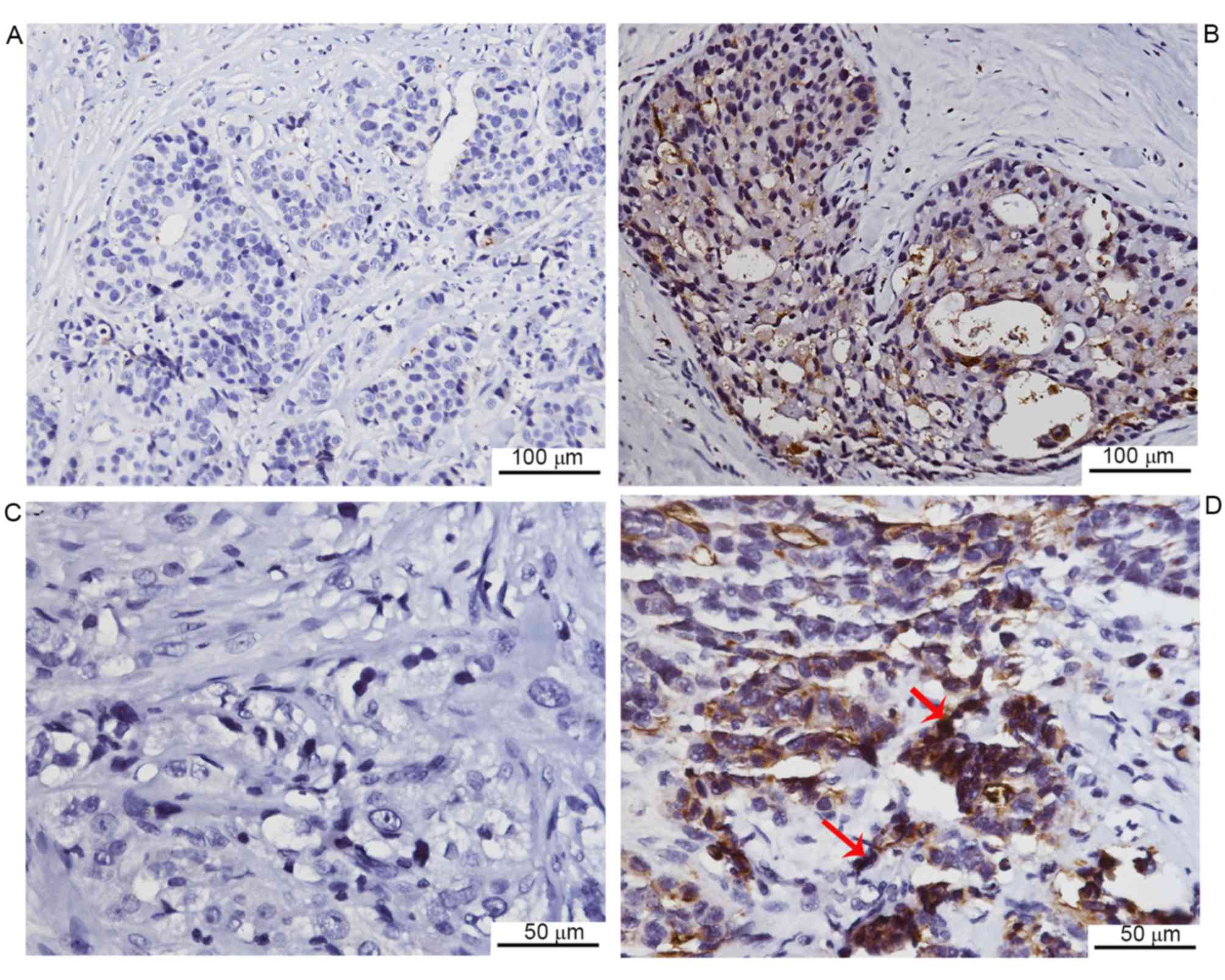
To investigate CEACAM1 expression in breast cancer
tissues from patients, immunohistochemical staining was performed.
Negative CEACAM1 expression was observed in 13 of 62 cases, while
the remaining cases exhibited positive expression (Fig. 1). Furthermore, the CEACAM1 expression
was quantitatively assessed, and the association between CEACAM1
expression levels and clinical characteristics was analyzed. It was
revealed that CEACAM1 staining was markedly lower in the lymph node
metastasis group compared with the non-metastasis group. In
addition, the CEACAM1 expression was markedly lower in invasive
duct cancer cases compared with cancer cases of other histological
types, including lobular, mucinous and tubular types of cancer. No
significant association was observed between CEACAM1 expression and
other clinical parameters, including age, histological grade and
tumor size (Table I). The data
indicated that CEACAM1 expression is negatively associated with
lymph node metastasis in breast cancer.
 | Table I.Association between clinical
parameters of the study population and CEACAM1 expression. |
Table I.
Association between clinical
parameters of the study population and CEACAM1 expression.
|
| CEACAM1 expression
(IOD) |
|---|
|
|
|
|---|
| Clinical
parameters | Number of
samples | Median | Range | P-value |
|---|
| Histological
grade |
|
|
|
|
| 1 | 12 | 12,489 | 0–69,744 | 0.164 |
| 2 | 25 | 11,805 | 0–58,974 |
|
| 3 | 25 | 4,853 | 0–59,528 |
|
| Histological
type |
|
|
|
|
|
Invasive duct carcinoma | 42 | 6,400 | 0–59,528 | 0.038 |
|
Others | 20 | 27,957 | 0–69,744 |
|
| Tumor size |
|
|
|
|
| T1 | 25 | 9,347 | 0–58,974 | 0.363 |
| T2 | 33 | 8,707 | 0–69,744 |
|
| T3 | 4 | 2,513 | 0–32,286 |
|
| Lymph node
metastasis |
|
|
|
|
| No | 39 | 19,248 | 0–69,744 | 0.017 |
|
Yes | 23 | 5,025 | 0–59,528 |
|
| Age, years |
|
|
|
|
|
<50 | 15 | 7,884 | 0–59,528 | 0.710 |
|
≥50 | 47 | 8,707 | 0–69,744 |
|
| Estrogen receptor
status |
|
|
|
|
|
Negative | 26 | 11,321 | 0–55,320 | 0.414 |
|
Positive | 36 | 6,400 | 0–69,744 |
|
| Progesterone
receptor status |
|
|
|
|
|
Negative | 33 | 10,202 | 0–69,744 | 0.960 |
|
Positive | 29 | 7,040 | 0–55,320 |
|
| Her2 status |
|
|
|
|
|
Negative | 25 | 7,040 | 0–69,744 | 0.834 |
|
Positive | 37 | 8,707 | 0–59,528 |
|
| Ki-67 expression,
% |
|
|
|
|
|
<20 | 24 | 10,078 | 0–69,744 | 0.270 |
|
≥20 | 38 | 6,127 | 0–59,528 |
|
CEACAM1-4S mRNA expression is
associated with decreased metastatic potential of human breast
cancer cell lines
To confirm immunohistochemical findings, a panel of
breast cancer cell lines with different metastatic potential was
analyzed for CEACAM1 mRNA expression levels by qPCR. Although 12
isoforms of CEACAM1 have been described in humans (19), it is well-accepted that CEACAM1-4S, a
short cytoplasmic isoform of CEACAM1, predominates in the human
breast epithelium (15,20). Therefore, the following experiments
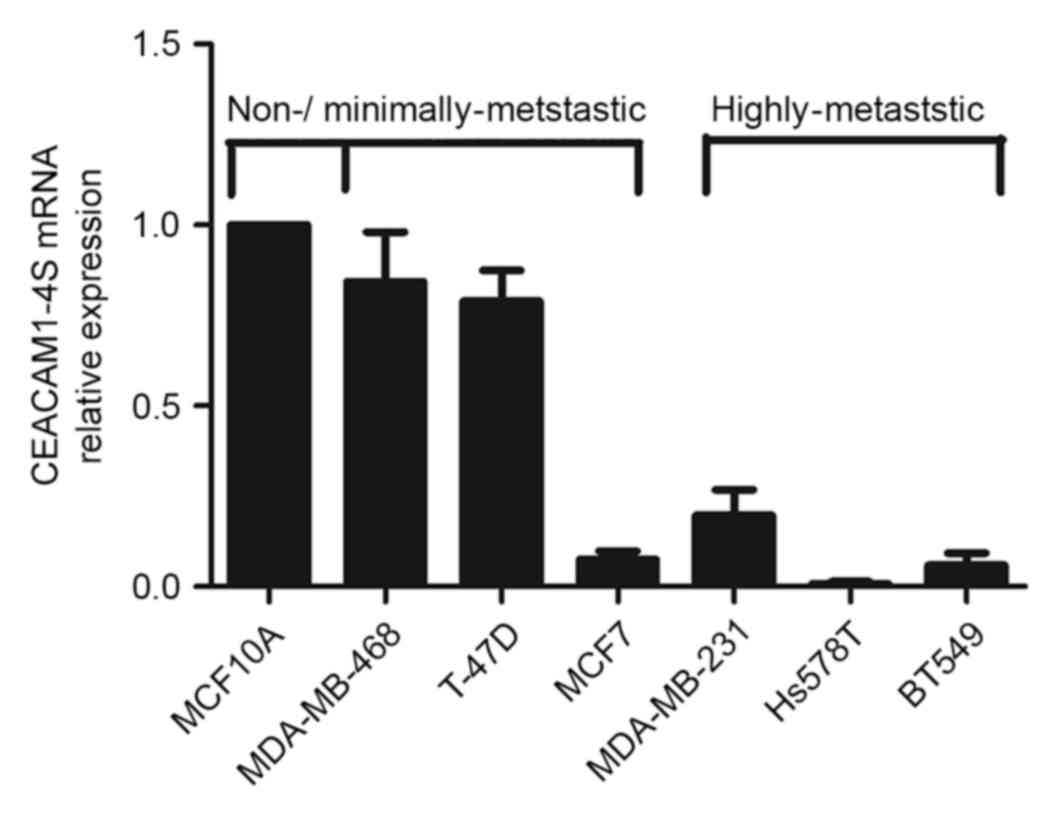
focused on CEACAM1-4S as a representative of CEACAM1. As shown in
Fig. 2, there was markedly decreased
expression of CEACAM1-4S mRNA in the highly metastatic Hs578T,
BT549 and MDA-MB-231 cells compared with the minimally-metastatic
MDA-MB-468 and T47D cells as well as the non-invasive breast
epithelial MCF10A cells. When normalized to MCF10A, the levels of
CEACAM1-4S mRNA in the highly metastatic cell lines exhibited a
mean expression 4–5-fold lower compared with the expression in
minimally metastatic cell lines, with the exception of MCF7
cells.
It was also noted that MCF7, a less aggressive cell
line compared with the other cell lines (21), expressed a low level of CEACAM1-4S,
indicating that CEACAM1-4S may not have a critical role in
regulation of metastatic potential of MCF7 cells. These data
indicated a negative association between CEACAM1-4S mRNA expression
and breast cancer metastatic potential. Considering the IHC results
from clinical samples, it was hypothesized that CEACAM1-4S may
serve an important role in the breast cancer metastasis.
CEACAM1-4S suppresses invasion and
migration in human breast cancer cells
To verify the aforementioned hypothesis, a
CEACAM1-4S expression vector or an empty vector were transfected
into Hs578T and BT549 cells, two highly metastatic cell lines with
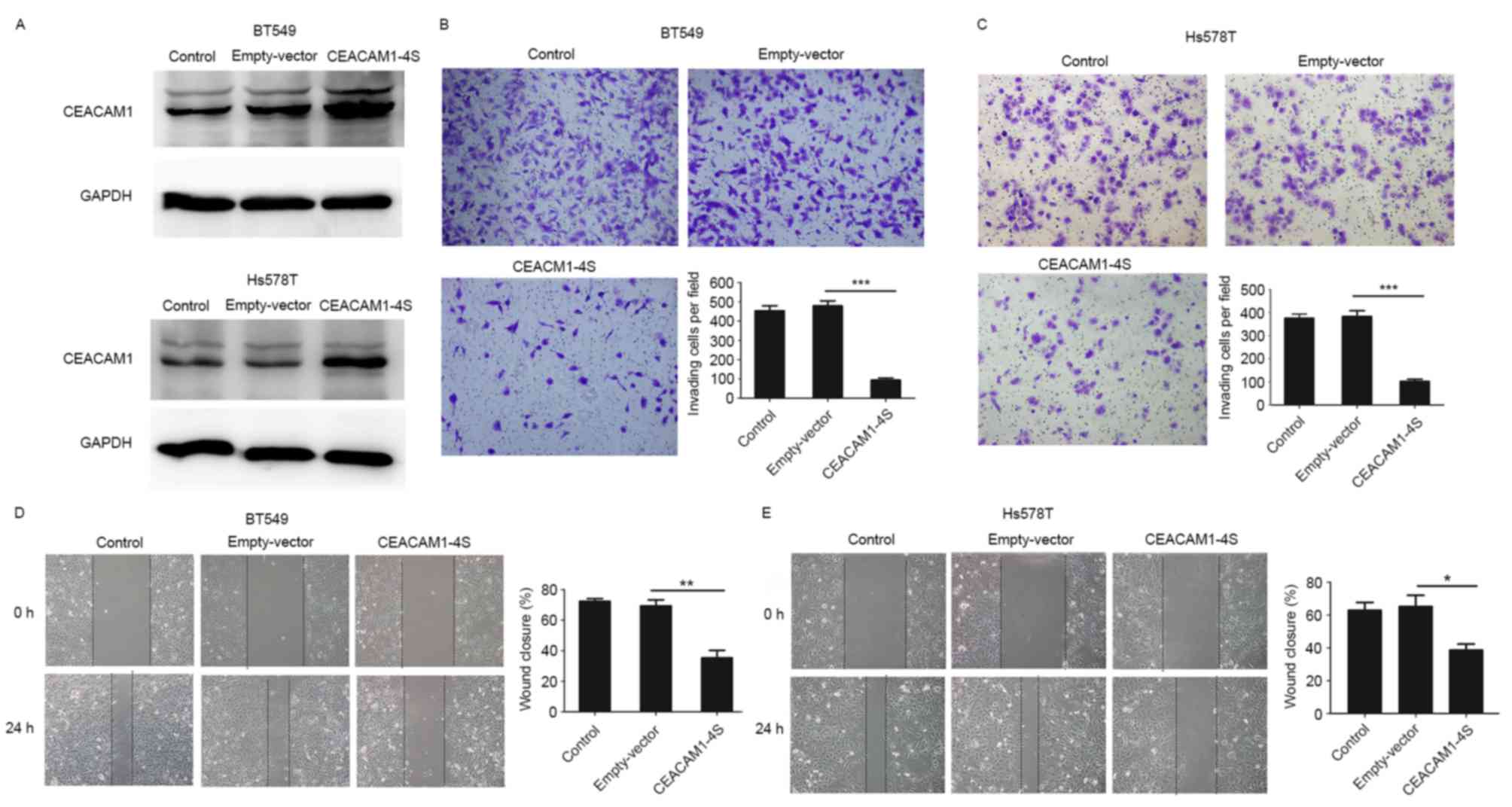
low levels of endogenous CEACAM1-4S expression (21). The transfection efficiency was
evidenced by western blot analysis (Fig.
3A), and Transwell invasion and wound healing assays were then
conducted. As shown in Fig. 3B-E,
when compared with wild-type or empty-vector controls,
CEACAM1-4S-tansfected cells exhibited significantly decreased
abilities to invade through the Matrigel-coated membranes and to
migrate into the wounded areas. For cells overexpressing
CEACAM1-4S, the number of invading cells was a mean 4–5-fold lower
compared with controls, and the motilities decreased by >40%. In
addition, it was revealed that CEACAM1-4S overexpression had little
effect on the proliferation of these two cell lines (data not
shown), indicating that all the changes observed in cell invasion
and migration were not due to cell proliferation. Therefore,
results indicated that CEACAM1-4S inhibits invasion and migration
of breast cancer cell.
MMP2 and ITMP2 are involved in the
inhibitory effects of CEACAM1-4S on breast cancer invasion and
migration
Next, it was investigated whether CEACAM1-4S
suppresses breast cancer cell invasion and migration. One possible
explanation is that CEACAM1 induces secondary activation of the
well-characterized MMPs/TIMPs system that performs a prominent role
in cancer invasion (22). Therefore,
four main members of the MMP/TIMP family were analyzed by western
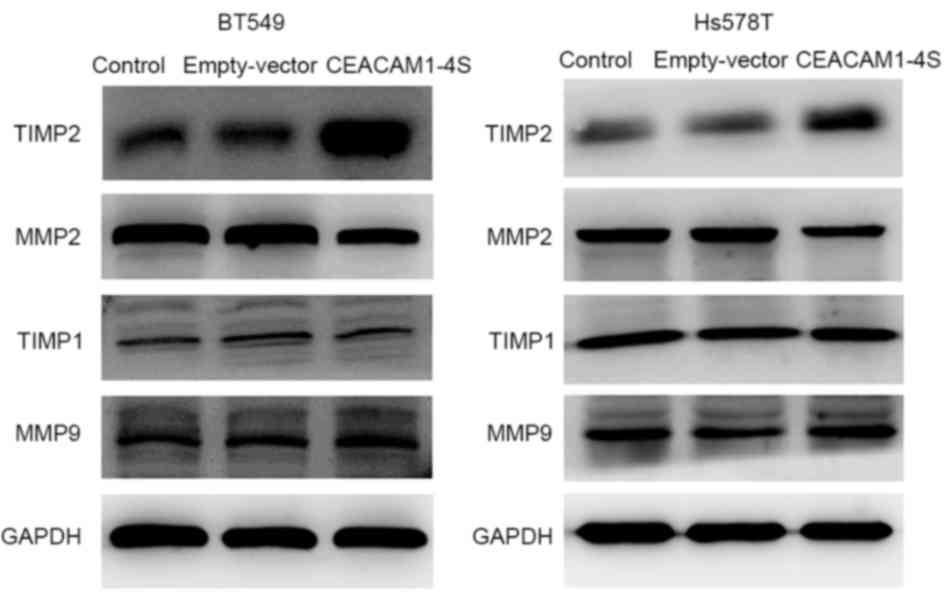
blot analysis in the present study. The results demonstrated that,
compared with corresponding controls, the levels of TIMP2
expression were markedly increased in CEACAM1-4S-transfected BT549
and Hs578T cells, while the levels of MMP2 expression were markedly
lower (Fig. 4). In addition, the
results revealed that MMP9 and TIMP1 expression were not markedly
changed following CEACAM1-4S overexpression in the two cell lines.
The data suggested that MMP2/TIMP2, but not MMP1/TIMP1, are
involved in CEACAM1-4S-mediated inhibition of cell invasion and
migration.
E and N-cadherin are implicated in
CEACAM1-4S-induced repression of cell invasion and migration
In an attempt to identify the involvement of other
regulatory pathways in the inhibition of cell invasion and
migration by CEACAM1-4S, the effect of CEACAM1-4S on the expression
of E- and N-cadherin, two crucial molecules for cancer cell
metastasis, was analyzed (23,24). As
expected, western blot analysis revealed that overexpression of
CEACAM1-4S markedly altered the expression of E- and N-cadherin. As
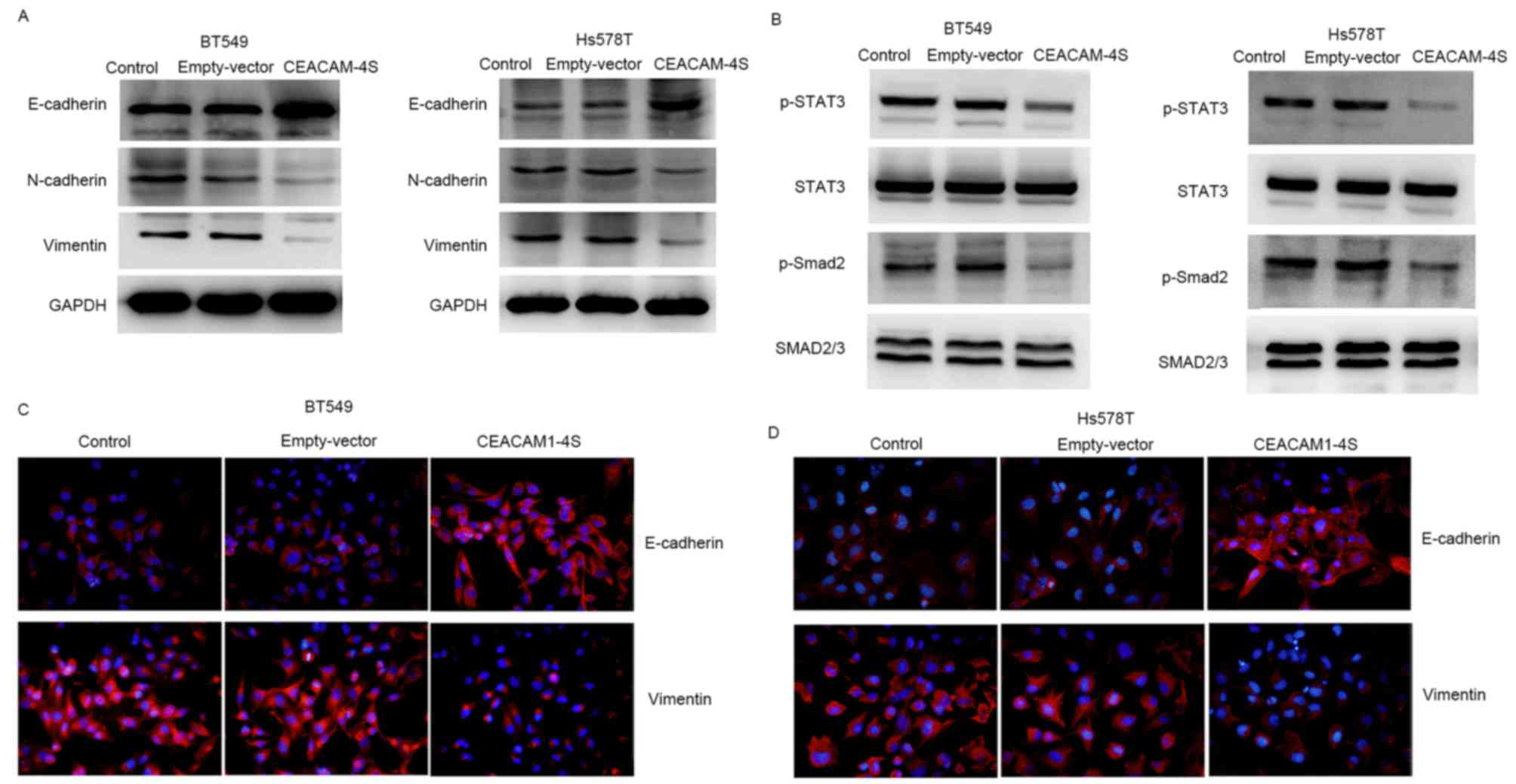
shown in Fig. 5A, CEACAM1-4S
overexpression in BT549 and Hs578T cells resulted in a substantial
increase in E-cadherin expression, whereas N-cadherin expression
was markedly decreased. Notably, E- and N-cadherin are widely
viewed as markers of epithelial-mesenchymal transition (EMT), an
important process during cancer metastasis (25,26). The
CEACAM1-4S-induced changes in E- and N-cadherin expression appeared
to be in parallel with EMT reversal. Therefore, the expression of
vimentin, another critical molecular marker for EMT (25,26), was
examined by western blot analysis. The results showed a marked
reduction of vimentin expression in CEACAM1-4S-transfected BT549
and Hs578T cells compared with the respective controls (Fig. 5A). The results were in accordance with
the aforementioned western blot analysis, showing a downregulation
of vimentin accompanied by upregulation of E-cadherin following
CEACAM1-4S overexpression in BT549 and Hs578T cells (Fig. 5A). Furthermore, two well-known signal
molecules associated with cancer metastasis and EMT, STAT3 and
Smad2 (25), were analyzed. It was
revealed that CEACAM1-4S overexpression markedly decreased the
phosphorylation of STAT3 and Smad2 but had no effect on the levels
of total protein levels (Fig. 5B).
Results of western blot analysis of vimentin and E-cadherin
expression in CECAM1-4S-transfected BT549 and Hs578T cells were
confirmed by IF staining (Fig. 5C and
D). Taken together, these data demonstrated that the
suppressive functions of CEACAM1-4S may be dependent on the
regulation of E and N-cadherin as well as EMT in breast cancer
cells.
Discussion
A previous study by the present authors reported
that CEACAM1 is downregulated in breast cancer (12). The present study aimed to address the
functional significance of CEACAM1 in breast cancer. First, CEACAM1
expression was examined in 62 tissue samples from patients with
breast cancer by IHC staining. The results indicated a negative
association between CEACAM1 expression and breast cancer
metastasis. It has been reported that the loss of CEACAM1
expression is associated with a poorer prognosis for patients with
breast cancer (11), and the findings
of the present study indicated that CEACAM1 may have a protective
role against breast cancer metastasis. To test this hypothesis, the
mRNA expression of CEACAM1-4S, the short cytoplasmic isoform of
CEACAM1 that predominates in human breast epithelium (15), was detected in a panel of six breast
cancer cell lines with different metastatic potentials. Consistent
with IHC results, the data revealed that CEACAM1-4S mRNA expression
was associated with the decreased metastatic potential of breast
cancer cell lines. The subsequent in vitro experiments
demonstrated that the overexpression of CEACAM1-4S significantly
inhibited the invasion and migration of BT549 and Hs578T cells.
Aberrant expression of CEACAM1 in breast cancer was observed a few
years ago (13), however, there is
limited information regarding the roles of CEACAM1 in the invasive
behavior of breast cancer. However, it has been demonstrated
previously that CEACAM1 is able to function as a tumor suppressor
that prevents tumorigenicity in a mice model of breast cancer
(27). The present study provides
evidence that the tumor suppressive activities of CEACAM1 extend to
the aggressive breast cancer cell phenotype including increased
migratory and invasive abilities.
CEACAM1 has a wide range of biological functions,
the majority of which are associated with the hallmarks of cancer,
including proliferation, apoptosis, immune evasion, inflammation
and angiogenesis (19,28). In 2004, Ebrahimnejad et al
(29) reported that CEACAM1 actively
contributes to tumor invasion and migration in melanoma.
Subsequently, CEACAM1 has been reported to promote tumor
invasiveness in thyroid cancer (30).
However, these findings were contradictory to previously published
studies that identified CEACAM1 as a tumor suppressor (27,31,32), which
was also demonstrated in the current study. Therefore, the roles of
CEACAM1 in malignancies appear to be conflicting. One of the
reasons responsible for this paradox may be that CEACAM1 comprises
isoform diversity. Previous studies have demonstrated that
CEACAM1-4L promotes colon and hepatocellular cancer cell invasion
and migration, while CEACAM1-4S exhibits inhibitory effects.
Therefore, there are functional differences between CEACAM1
isoforms (33,34). Considering that CEACAM1-4S is the
physiological predominant isoform (15), the present study mimicked the scenario
in normal human breast cells by increasing CEACAM1-4S expression in
breast cancer cells. In accordance with the aforementioned studies
(33,34), results of the present study revealed
an inhibitory effect of CEACAM1-4S on cell invasion and migration
in breast cancer. Based on the present results and results of
previous studies (33,34), it is hypothesized that CEACAM1-4S acts
as a tumor suppressor, whereas CEACAM1-4L may be a tumor
stimulator. Combined with the findings that different malignancies
preferentially express distinct CEACAM1 isoforms (15,33), this
partly explains the paradoxical roles attributed to CEACAM1.
Additional extensive and intensive studies are required to confirm
the biological functions of CEACAM1 in cancer metastasis.
As cancer metastasis is a highly integrated process,
which is regulated by a large number of extracellular
matrix-associated enzymes, including MMPs and their natural
inhibitors, TIMPs (35), experiments
detecting changes of these enzymes were performed. High TIMP2 and
low MMP2 expression patterns were observed in
CEACAM1-4S-transfected breast cancer cells. Therefore, CEACAM1-4S
may exhibit the suppressive functions through up and downregulation
of the MMP2/TIMP2 balance. In addition, the present results
demonstrated that the expression of MMP9 and its natural inhibitor
TIMP1 were not markedly affected by CEACAM1-4S overexpression,
indicating that the MMP9/TIMP1 balance was not involved in the
regulation of cell invasion by CEACAM1-4S. Furthermore, the present
data revealed that cadherins may be downstream effectors of
CEACAM1-4S. It is well-documented that cadherins control the
balance between repression and promotion of cancer cell migration
and invasion (23,36–38).
Currently, E-cadherin and N-cadherin are the most comprehensively
investigated cadherins in cancer, and they balance each other with
E-cadherin frequently downregulated in cancer and acts as an
invasion suppressor, whereas N-cadherin is an invasion promoter
upregulated in the majority of cancer types (23,39). A
previous study demonstrated that N-cadherin has an important
promoting effect on CEACAM1-mediated migration in LoVo (human colon
cancer) cells (24). In the present
study, it was revealed that CEACAM1-4S not only regulates
N-cadherin, but also E-cadherin, and this regulation was in
parallel with EMT reversal. Additional findings that vimentin and
intracellular signals p-Smad2 and p-STAT3 were downregulated by
CEACAM1-4S further supported a reversal effect of CEACAM1-4S on EMT
in breast cancer cells. However, it should be noted that EMT is a
complicated process orchestrated by various biological molecules,
and whether CEACAM1-4S serves a crucial role in EMT reversal
requires further study.
In summary, the present study demonstrated that
CEACAM1-4S performs an inhibitory role in breast cancer cell
invasion and migration, possibly through regulating the balance
between MMP2/TIMP2 and E-/N-cadherins. In addition, evidence that
CEACAM1-4S may induce EMT reversal in breast cancer cells was
provided. Therefore, restoring CEACAM1-4S expression may provide a
novel avenue for therapy of patients with breast cancer.
Acknowledgements
The present study was supported by the National
Natural Science Foundation of China (grant nos. 81272479,
814024198, 81572821, 81502490 and 81502491), the Shanghai Committee
of Science and Technology (grant no. 14YF1412200), the Program of
Shanghai Leading Talents (grant no. 038) and the Program of
Shanghai Shen-Kang Hospital Development Center (grant no.
SHDC22014004).
Glossary
Abbreviations
Abbreviations:
|
CEACAM1
|
carcinoembryonic antigen-related cell
adhesion molecule 1
|
|
PCR
|
polymerase chain reaction
|
|
EMT
|
epithelial-mesenchymal transition
|
|
TIMP
|
tissue inhibitor of
metalloproteinase
|
|
MMP
|
matrix metalloproteinase
|
References
|
1
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Weigelt B, Peterse JL and van 't Veer LJ:
Breast cancer metastasis: Markers and models. Nat Rev Cancer.
5:591–602. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Gray-Owen SD and Blumberg RS: CEACAM1:
Contact-dependent control of immunity. Nat Rev Immunol. 6:433–446.
2006. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Prall F, Nollau P, Neumaier M, Haubeck HD,
Drzeniek Z, Helmchen U, Löning T and Wagener C: CD66a (BGP), an
adhesion molecule of the carcinoembryonic antigen family, is
expressed in epithelium, endothelium and myeloid cells in a wide
range of normal human tissues. J Histochem Cytochem. 44:35–41.
1996. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Neumaier M, Paululat S, Chan A, Matthaes P
and Wagener C: Biliary glycoprotein, a potential human cell
adhesion molecule, is down-regulated in colorectal carcinomas. Proc
Natl Acad Sci USA. 90:10744–10748. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Tanaka K, Hinoda Y, Takahashi H, Sakamoto
H, Nakajima Y and Imai K: Decreased expression of biliary
glycoprotein in hepatocellular carcinomas. Int J Cancer. 74:15–19.
1997. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Kammerer R, Riesenberg R, Weiler C,
Lohrmann J, Schleypen J and Zimmermann W: The tumour suppressor
gene CEACAM1 is completely but reversibly downregulated in renal
cell carcinoma. J Pathol. 204:258–267. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Busch C, Hanssen TA, Wagener C and OBrink
B: Down-regulation of CEACAM1 in human prostate cancer: Correlation
with loss of cell polarity, increased proliferation rate and
Gleason grade 3 to 4 transition. Hum Pathol. 33:290–298. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Nittka S, Gunther J, Ebisch C,
Erbersdobler A and Neumaier M: The human tumor suppressor CEACAM1
modulates apoptosis and is implicated in early colorectal
tumorigenesis. Oncogene. 23:9306–9313. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Estrera VT, Chen DT, Luo W, Hixson DC and
Lin SH: Signal transduction by the CEACAM1 tumor suppressor.
Phosphorylation of serine 503 is required for growth-inhibitory
activity. J Biol Chem. 276:15547–15553. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang JL, Sun SZ, Qu X, Liu WJ, Wang YY, Lv
CX, Sun JZ and Ma R: Clinicopathological significance of CEACAM1
gene expression in breast cancer. Chin J Physiol. 54:332–338.
2011.PubMed/NCBI
|
|
12
|
Yang C, He P, Liu Y, He Y, Yang C, Du Y,
Zhou M, Wang W, Zhang G, Wu M and Gao F: Down-regulation of CEACAM1
in breast cancer. Acta Biochim Biophys Sin (Shanghai). 47:788–794.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Riethdorf L, Lisboa BW, Henkel U, Naumann
M, Wagener C and Loning T: Differential expression of CD66a (BGP),
a cell adhesion molecule of the carcinoembryonic antigen family, in
benign, premalignant and malignant lesions of the human mammary
gland. J Histochem Cytochem. 45:957–963. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Huang J, Hardy JD, Sun Y and Shively JE:
Essential role of biliary glycoprotein (CD66a) in morphogenesis of
the human mammary epithelial cell line MCF10F. J Cell Sci.
112:4193–4205. 1999.PubMed/NCBI
|
|
15
|
Kirshner J, Chen CJ, Liu P, Huang J and
Shively JE: CEACAM1-4S, a cell-cell adhesion molecule, mediates
apoptosis and reverts mammary carcinoma cells to a normal
morphogenic phenotype in a 3D culture. Proc Natl Acad Sci USA.
100:521–526. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tavassoli FA and Devilee P: World Health
Organization classification of tumors Pathology and genetics of
tumors of the breast and female genital organs. Lyon: IARC Press;
2003
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Oliveira-Ferrer L, Tilki D, Ziegeler G,
Hauschild J, Loges S, Irmak S, Kilic E, Huland H, Friedrich M and
Ergün S: Dual role of carcinoembryonic antigen-related cell
adhesion molecule 1 in angiogenesis and invasion of human urinary
bladder cancer. Cancer Res. 64:8932–8. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Beauchemin N and Arabzadeh A:
Carcinoembryonic antigen-related cell adhesion molecules (CEACAMs)
in cancer progression and metastasis. Cancer Metastasis Rev.
32:643–671. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kirshner J, Hardy J, Wilczynski S and
Shively JE: Cell-cell adhesion molecule CEACAM1 is expressed in
normal breast and milk and associates with beta1 integrin a 3D
model of morphogenesis. J Mol Histol. 35:287–299. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Chakrabarti R, Hwang J, Blanco Andres M,
Wei Y, Lukačišin M, Romano RA, Smalley K, Liu S, Yang Q, Ibrahim T,
et al: Elf5 inhibits the epithelial -mesenchymal transition in
mammary gland development and breast cancer metastasis by
transcriptionally repressing Snail2. Nat Cell Biol. 14:1212–1222.
2012. View
Article : Google Scholar : PubMed/NCBI
|
|
22
|
Said AH, Raufman JP and Xie G: The role of
matrix metalloproteinases in colorectal cancer. Cancers. 6:366–375.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Derycke LD and Bracke ME: N-cadherin in
the spotlight of cell-cell adhesion, differentiation,
embryogenesis, invasion and signalling. Int J Dev Biol. 48:463–476.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu J, Di G, Wu CT, Hu X and Duan H:
CEACAM1 inhibits cell-matrix adhesion and promotes cell migration
through regulating the expression of N-cadherin. Biochem Biophys
Res Commun. 430:598–603. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lamouille S, Xu J and Derynck R: Molecular
mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell
Biol. 15:178–196. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hugo H, Ackland ML, Blick T, Lawrence MG,
Clements JA, Williams ED and Thompson EW: Epithelial-mesenchymal
and mesenchymal-epithelial transitions in carcinoma progression. J
Cell Physiol. 213:374–383. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Luo WP, Wood CG, Earley K, Hung MC and Lin
SH: Suppression of tumorigenicity of breast cancer cells by an
epithelial cell adhesion molecule (C-CAM1): The adhesion and growth
suppression are mediated by different domains. Oncogene.
14:1697–1704. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Obrink B: Is CEACAM1 a lymphangiogenic
switch? Blood. 110:4137–4138. 2007. View Article : Google Scholar
|
|
29
|
Ebrahimnejad A, Streichert T, Nollau P,
Horst AK, Wagener C, Bamberger AM and Brümmer J: CEACAM1 enhances
invasion and migration of melanocytic and melanoma cells. Am J
Pathol. 165:1781–1787. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Liu W, Wei W, Winer D, Bamberger AM,
Bamberger C, Wagener C, Ezzat S and Asa SL: CEACAM1 impedes thyroid
cancer growth but promotes invasiveness: A putative mechanism for
early metastases. Oncogene. 26:2747–2758. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Luo W, Tapolsky M, Earley K, Wood CG,
Wilson DR, Logothetis CJ and Lin SH: Tumor-suppressive activity of
CD66a in prostate cancer. Cancer Gene Ther. 6:313–321. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Volpert O, Luo W, Liu TJ, Estrera VT,
Logothetis C and Lin SH: Inhibition of prostate tumor angiogenesis
by the tumor suppressor CEACAM1. J Biol Chem. 277:35696–35702.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ieda J, Yokoyama S, Tamura K, Takifuji K,
Hotta T, Matsuda K, Oku Y, Nasu T, Kiriyama S, Yamamoto N, et al:
Re-expression of CEACAM1 long cytoplasmic domain isoform is
associated with invasion and migration of colorectal cancer. Int J
Cancer. 129:1351–1361. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Kiriyama S, Yokoyama S, Ueno M, Hayami S,
Ieda J, Yamamoto N, Yamaguchi S, Mitani Y, Nakamura Y, Tani M, et
al: CEACAM1 long cytoplasmic domain isoform is associated with
invasion and recurrence of hepatocellular carcinoma. Ann Surg
Oncol. 21 Suppl 4:S505–S514. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Stamenkovic I: Matrix metalloproteinases
in tumor invasion and metastasis. Semin Cancer Biol. 10:415–433.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Cavallaro U and Christofori G: Cell
adhesion and signaling by cadherins and Ig-CAMs in cancer. Nat Rev
Cancer. 4:118–132. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wheelock MJ, Shintani Y, Maeda M, Fukumoto
Y and Johnson KR: Cadherin switching. J Cell Sci. 121:727–735.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Theveneau E and Mayor R: Cadherins in
collective cell migration of mesenchymal cells. Curr Opin Cell
Biol. 24:677–684. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Nakada M, Nakada S, Demuth T, Tran NL,
Hoelzinger DB and Berens ME: Molecular targets of glioma invasion.
Cell Mol Life Sci. 64:458–478. 2007. View Article : Google Scholar : PubMed/NCBI
|