Introduction
Intrahepatic cholangiocarcinoma (ICC), the second
most common intrahepatic primary tumor after hepatocellular
carcinoma (HCC), exhibits a highly invasive nature and frequently
metastasizes to the lymph nodes (1).
Although progress has been made in the diagnosis and treatment of
ICC, the long-term prognosis of patients with ICC remains
unsatisfactory (2–4). At present, the treatment for this
disease is curative resection, which is associated with a low
5-year survival rate (between 25 and 35%) and a high recurrence
rate (67.9%) (5,6). Furthermore, there have been few
effective diagnostic and prognostic molecular markers reported for
ICC. A comprehensive understanding of the mechanisms of ICC
metastasis will contribute to the identification of novel diagnosis
and therapeutic targets.
MicroRNAs (miRNAs/miRs) are 18-25 bp, non-coding,
single-strand RNAs that regulate gene expression at the
post-transcriptional level by binding to the 3′ untranslated region
(3′UTR) of target mRNAs (7,8). miRNAs are involved in a variety of basic
physiological and pathological processes, including cell
proliferation, differentiation, metabolism and apoptosis (9,10).
Previous studies have demonstrated that miRNAs are frequently
downregulated or dysregulated in a range of types of malignancy,
including ICC, breast cancer and colorectal cancer, potentially
acting as oncogenes or tumor suppressor genes (9–11).
Emerging evidence has demonstrated that the aberrant expression of
miRNA may serve key roles in the initiation and progression of ICC
and HCC (11–13). However, the exact mechanisms by which
miRNAs regulate the metastasis of ICC are poorly characterized.
The present study aimed to investigate the effects
of miR-26b-5p expression in ICC. miR-26b-5p was identified as
downregulated in ICC, particularly in ICC with lymphatic metastasis
(mICC) and in invasive subpopulations of the RBE and HCCC-9810 cell
lines (iRBE and iHCCC-9810). It was identified that miR-26b-5p
mimics inhibited the proliferation, migration and invasion of RBE
and HCCC-9810 cells in vitro. Furthermore, S100
calcium-binding protein A7 (S100A7), which is implicated in
tumorigenesis, was demonstrated to be a direct target of
miR-26b-5p. Results of the present study revealed that miR-26b-5p
hindered the proliferation, migration and invasive abilities of ICC
cells by inhibiting S100A7 expression.
Materials and methods
Clinical samples
A total of 20 ICC histopathologically verified tumor
samples, together with their matched adjacent non-tumor tissues and
an additional 40 tumor samples, including 20 mICC samples and 20
samples without detected metastasis, were obtained from patients
that underwent resective surgery for ICC or mICC (age range, 35–85
years, mean ± standard deviation (SD), 67±11; 25 female and 35 male
samples; 31 cases of stage I–II, 29 cases of stage III according to
the grading standards from Chinese Society of Liver Cancer
(14) at the Eastern Hepatobiliary
Surgical Hospital between September 2012 and May 2014. The tissues
were snap frozen in liquid nitrogen immediately subsequent to
surgical removal and stored at −70°C until required. The use of all
specimens in the present study was approved by the Institutional
Ethics Committee of the Second Military Medical University
(Shanghai, China). All patients signed informed consent forms for
the use of their data and surgical specimens in the present
study.
Cell lines and growth conditions
Human ICC RBE and HCCC-9810 cell lines were
purchased from Thermo Fisher Scientific, Inc. (Waltham, MA, USA).
All cells were cultured in Dulbecco's modified Eagle's medium
(DMEM; Invitrogen; Thermo Fisher Scientific, Inc.) supplemented
with 10% fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific,
Inc.), 100 IU/ml penicillin (Gibco; Thermo; Fisher Scientific,
Inc.) and 100 µg/ml streptomycin (Gibco; Thermo Fisher Scientific,
Inc.) in a humidified 5% CO2 cell incubator at 37°C.
Establishment of invasive cell
lines
Highly invasive subpopulations from the RBE and
HCCC-9810 cell lines were established with 10 rounds of selection
from a Matrigel-coated Transwell assay. All Matrigel and
supplements were purchased from BD Biosciences (Franklin Lakes, NJ,
USA). In brief, the inserts were coated with 50 ml of 1 mg/ml
Matrigel matrix and 1×105 cells in 200 ml serum-free
DMEM were seeded in the Transwell upper chambers. A total of 600 ml
DMEM with 10% fetal bovine serum was added to the lower chamber.
Following incubation for 12–72 h at 37°C, the cells that migrated
through the Matrigel membranes and attached to the lower surface
were further passaged in subsequent sub-clone cultures. The
selection of ICC cells was repeated 10 times to establish highly
invasive sub-lines. These were designated as iRBE and iHCCC-9810
cells.
RNA isolation and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA was extracted from cultured cells and
tissues using TRIzol RNA isolation reagent (Invitrogen; Thermo
Fisher Scientific, Inc.). For the reverse transcription, all
reagents were purchased from Applied Biosystems (Thermo Fisher
Scientific, Inc.). Mature miRNAs were reverse-transcribed from
total RNA using specific miRNA RT-primers from the TaqMan Reverse
MicroRNA Transcription kit (Applied Biosystems; Thermo Fisher
Scientific, Inc.). The temperature protocol for RT was as follows:
16°C for 30 min, 42°C for 30 min and 85°C for 2 min, and finally
4°C hold. Following this, pre-amplification step was conducted
according to the protocol of TaqMan Reverse MicroRNA Transcription
kit (Applied Biosystems; Thermo Fisher Scientific, Inc.). Following
the RT process, the pre-amplification process was performed. The
temperature cycles were as follows: 95°C for 10 min, 55°C for 2 min
and 72°C for 2 min, then 12 cycles of 95°C for 15 sec, 60°C for 4
min and 99.9°C for 10 min, and then 4°C hold.
Following the RT and pre-amplification process, qPCR
was performed using the TaqMan miRNA assay kit primers (Thermo
Fisher Scientific, Inc.) with the TaqMan Universal PCR Master mix
(Thermo Fisher Scientific, Inc.) and analyzed using an ABI 7500
system (Applied Biosystems; Thermo Fisher Scientific, Inc.). SYBR
Premix Ex Taq (Perfect Real Time; Takara Biotechnology Co., Ltd.)
was used for qPCR. The temperature protocol for qPCR was as
follows: 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec
and 60°C for 1 min. To normalize mRNA expression, β-actin was used
as the endogenous internal control with the 2−ΔΔCq
method. The primers for S100A7 were as follows: Forwards,
5′-AACTTCCTTAGTGCCTGTG-3′; reverse, 5′-TGGTAGTCTGTGGCTATGTC-3′. The
primers for β-actin were: Forward, 5′-AGAGCTACGAGCTGCCTGAC-3′;
reverse, 5′-AGCACTGTGTTGGCGTACAG-3′. For all figures of RT-qPCR
data, values on the y-axis are equal to 2ΔΔCq (15). Gene expression data were obtained in
triplicate in three independent experiments.
Cell transfection
miRNA vectors were purchased from Shanghai
GenePharma Co., Ltd. (Shanghai, China), including an miR-26b-5p
expression vector (catalog no. mh40312; Applied Biological
Materials Inc., Richmond, BC, Canada), a miR-26b-5p control vector
(cat. no. mm30393; Applied Biological Materials Inc., Richmond, BC,
Canada), miR-26b-5p inhibitor (catalog no. MIN0000083; Qiagen China
Co., Ltd., Shanghai, China) and miR-26b-5p mimic (catalog no.
MSY0000083; Qiagen China Co., Ltd.). cDNA for the miR-26b-5p
precursor (pre-miR-26b-5p) was synthesized by Shanghai GenePharma
Co., Ltd. based the sequence information for harmiR-26b-5p
(MI0000083) from the miRBase database (www.mirbase.org/cgi-bin/mature.pl?mature_acc=MIMAT0000083),
which was sub-cloned into a lentiviral expression vector by
Shanghai Sunbio Biotechnology Co., Ltd. (Shanghai, China).
The S100A7 cDNA was purchased from Sino Biological,
Inc. (Beijing, China) and packaged into a lentiviral expression
vector by Shanghai Sunbio Biotechnology Co., Ltd. S100A7 Pre-design
Chimera RNAi was purchased from Abnova Co. Ltd. (Taoyuan, Taiwan).
The wild-type 3′UTR for human S100A7 was sub-cloned into a
firefly/Renilla Dual-Luciferase reporter vector by
GeneCopoeia, Inc. (Rockville, MD, USA). A point mutation was
produced within the predicted miR-26b-5p interaction region of the
S100A7 3′UTR fragments using a Fusion Site Directed Mutagenesis kit
from Thermo Fisher Scientific, Inc. The mutated and wild-type 3′UTR
region of human S100A7 were each cloned into the 3′UTR position of
the previously constructed S100A7 expression vector. The
aforementioned vectors (1 µg/ml) were transfected into ICC cells
using Lipofectamine 2000 (Invitrogen; Thermo Fisher Scientific,
Inc.) according to the manufacturer's protocol.
Determination of cell
proliferation
The transfected cells were seeded on 96-well plates
at a density of 1×104 cells/well. The plates were
incubated at 37°C for 48 h, and MTT solution (20 µl, 5 mg/ml) was
added to each well (to a total volume of 250 µl). Following
incubation for 3 h at 37°C, the MTT solution was removed and the
formazan precipitates were dissolved in 200 µl dimethyl sulfoxide
and incubated for 30 min at 37°C. The absorbance was measured at
540 nm. The experiment was performed in triplicate.
Migration and invasion assays
For the Transwell migration assays, 1×104
cells were plated in the top chamber containing a non-coated
membrane (24-well insert; 8 mm pore size; BD Biosciences). For the
invasion assays, 2×105 cells were plated in the top
chamber containing a Matrigel-coated membrane (24-well insert; 8 mm
pore size; BD Biosciences) with serum free DMEM. DMEM supplemented
with 10% (v/v) FBS was used as a chemoattractant in the lower
chamber. At 16 h, the non-migrated/non-invading cells were removed
from the upper sides of the Transwell membrane filter inserts using
cotton-tipped swabs. The migrated/invaded cells on the lower sides
of the inserts were stained with Giemsa (3% working solution) for
30 min at 37°C and the cells were counted using a light microscope
(CKX41; Olympus Corporation, Tokyo, Japan). And the experiment was
repeated for 3 times.
Luciferase reporter assay
For the prediction of miRNA targets, TargetScan
(heep://www.targetscan.org) and miRanda
(http://www.microrna.org) were used. For the
luciferase reporter assay, RBE cells were respectively transfected
with wild-type and mutant firefly/Renilla Dual-Luciferase
reporter vectors, as previously described, and subsequently
transduced with the miR-26b-5p mimic or inhibitor. The cells were
collected following incubation for 48 h. A dual-luciferase reporter
assay system (Promega Corporation, Madison, WI, USA) was used to
measure Renilla luciferase activity, according to the
manufacturer's protocol. The luciferase reporter assay was repeated
three times.
Statistical analysis
The quantitative data are expressed as the mean ±
SD. GraphPad Prism 5 software (GraphPad Software, Inc., La Jolla,
CA, USA) was used for statistical analysis. Comparisons between two
or more groups were subjected to a two-tailed Student's t-test or
one-way analysis of variance followed by least significant
difference test. P<0.05 was considered to indicate a
statistically significant difference.
Results
Candidate miRNA selection
In order to identify novel biomarkers for ICC, 5
candidate miRNAs (miR-101-3p, miR-147a, miR-196-5p, miR-93-3p and
miR-26b-5p), which have been associated with other types of cancer
and diseases (16–20), were quantified in RBE, iRBE, HCCC-9810
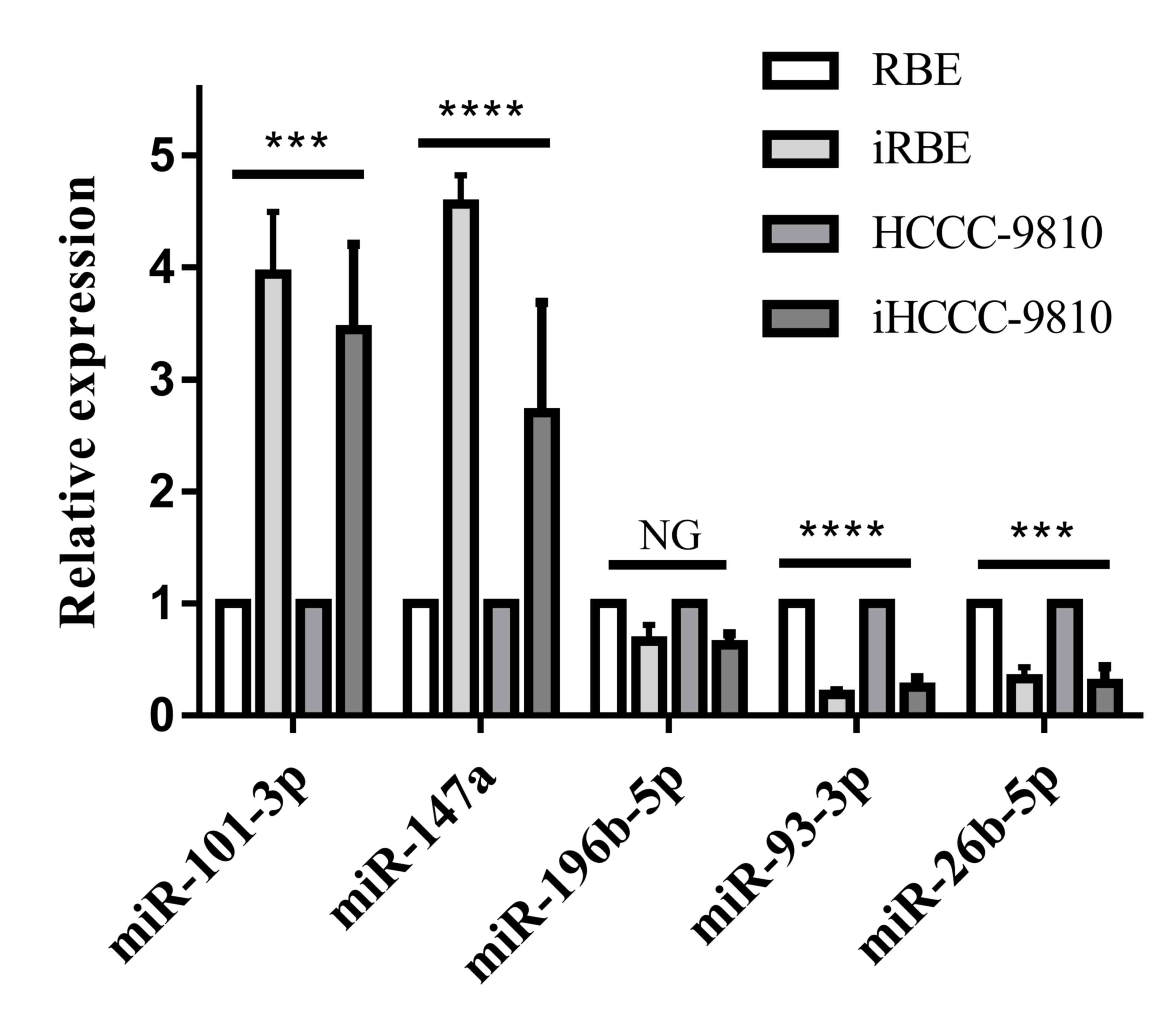
and iHCCC-9810 cells with RT-qPCR. As presented in Fig. 1, the levels of miR-101-3p and miR-147a
increased significantly (P<0.001) in the invasive cell
subpopulations compared with the normal cell lines, whereas the
level of miR-93-3p and miR-26b-5p was significantly decreased
(P<0.001). There was no alteration in the level of miR-196-5p.
Therefore, the levels of miR-101-3p, miR-147a, miR-93-3p and
miR-26b-5p were selected for quantification in human ICC
tissue.
miR-26-5p is downregulated in ICC
tumors
To explore the roles of miRNAs in human ICC
development, the expression levels of the four previously named
candidate miRNAs were detected in 20 samples of ICC tumor and
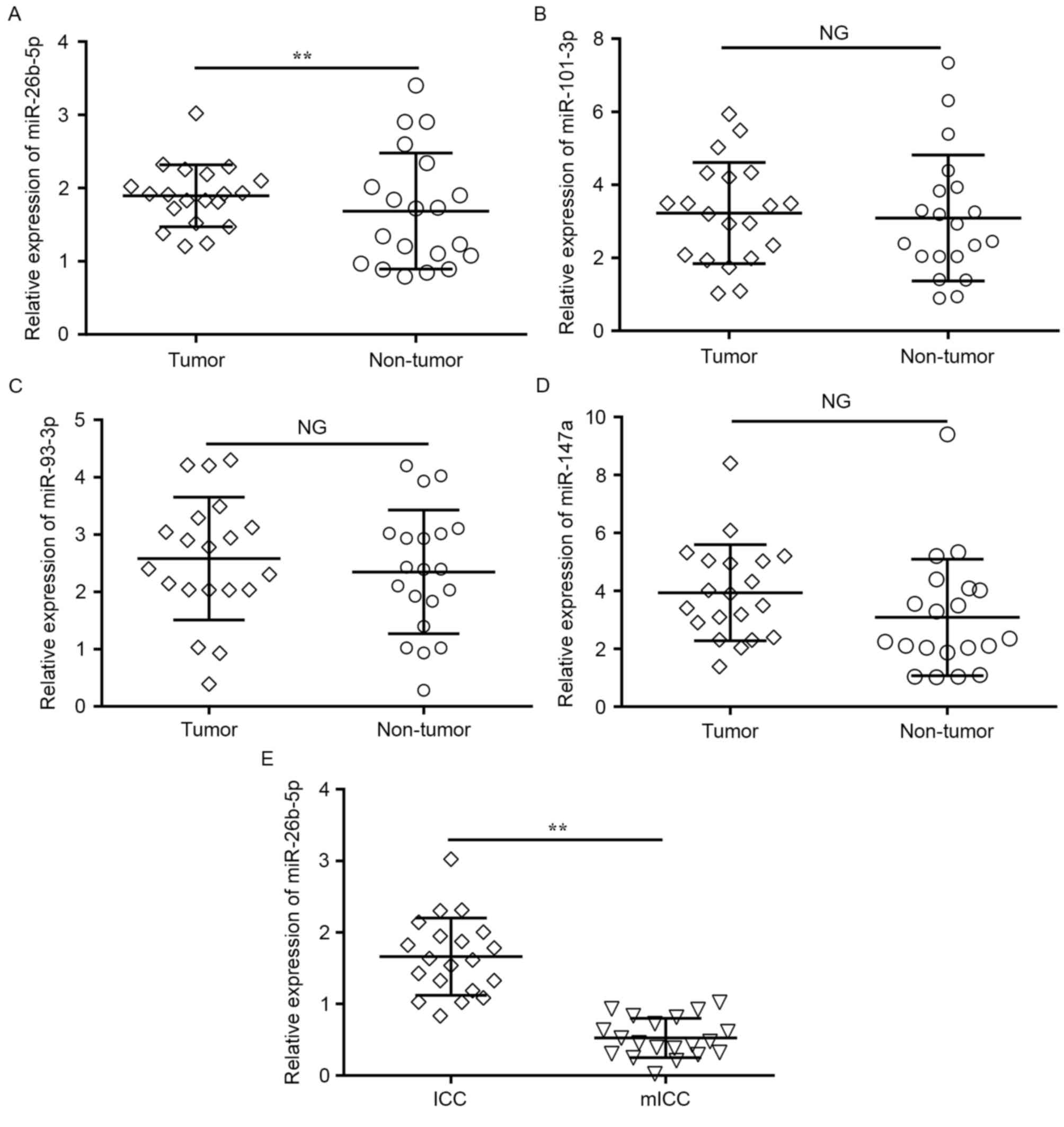
matched adjacent normal tissues with RT-qPCR. Only the expression
of miR-26b-5p was significantly downregulated in the tumor tissues
compared with the adjacent normal tissues (P=0.009; Fig. 2A), whereas no change was observed in
the expression of miR-101-3p, miR-147a and miR-93-3p (Fig. 2B-D).
In addition, to determine whether miR-26b-5p
expression was associated with ICC metastasis, miR-26b-5p
expression level was compared between tumor tissues from 20
patients with mICC and 20 patients without detected metastases.
RT-qPCR revealed that the miR-26b-5p expression levels were
significantly lower in metastatic tumors compared with
non-metastatic tumors (P=0.005; Fig.
2E). These results suggested that miR-26b-5p was significantly
downregulated in ICC, and that the decreased expression of
miR-26b-5p was associated with the metastatic behavior of ICC
tumors and cells.
miR-26b-5p regulates ICC cell
proliferation, migration and invasion
To determine the functional significance of the
aberrant expression of miR-26b-5p in ICC, RBE and iRBE cells were
transfected with an miR-26b-5p mimic or inhibitor and the viability
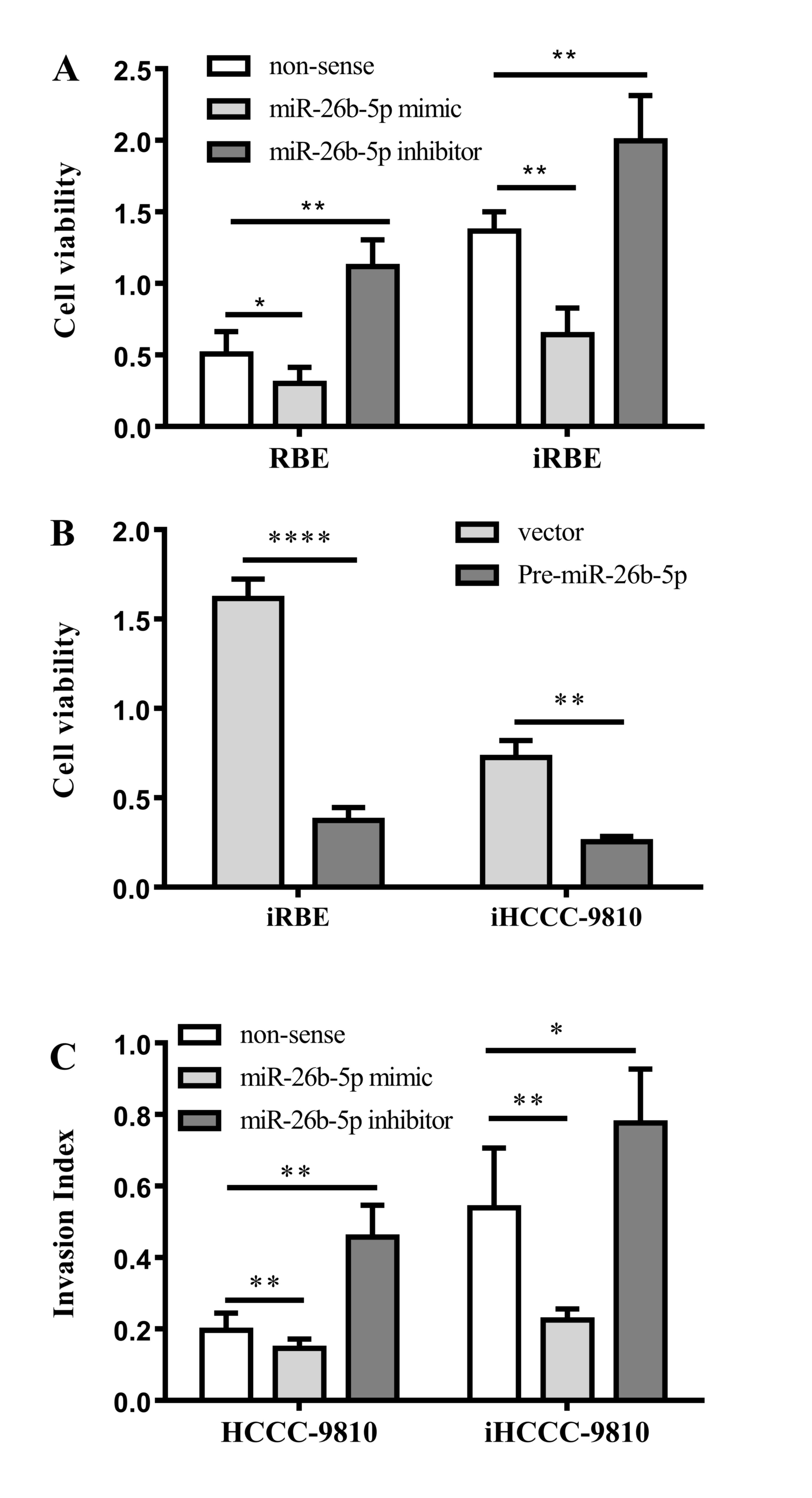
of the cells was examined with an MTT assay. As presented in
Fig. 3A, compared with RBE cells, the
viability in all the iRBE groups was increased. Transfection with
an miR-26b-5p mimic reduced the rate of proliferation in the RBE
and iRBE cells (P=0.0249 and P=0.0042, respectively), whereas
transfection with an miR-26b-5p inhibitor increased the rate of
proliferation in RBE and iRBE cells (P=0.0054 and P=0.0084,
respectively). This data suggested that miR-26b-5p expression
significantly inhibited cell proliferation in the two cell
lines.
To confirm this conclusion, pre-miR-26b-5p was
transfected into iRBE and iHCCC-9810 cells, and the viability of
the cells was examined. The data revealed that transfection with
pre-miR-26b-5p significantly inhibited the cell viability compared
with transfection with a non-sense control (P<0.0001 and
P=0.0013, respectively; Fig. 3B),
which is in accord with the previous data.
In addition to cell proliferation, the effects of
miR-26b-5p on ICC cell migration and invasion were also explored.
Cells were transfected with an miR-26b-5p mimic or inhibitor, and
their migration and invasion abilities were measured with a
Transwell assay. As shown in Fig. 3C,
compared with the non-sense control group, miR-26b-5p mimics
reduced the invasion ability in HCCC-9810 and iHCCC-9810 cells
(P=0.0106 and P=0.0051, respectively); the miR-26b-5p inhibitor,
however, increased the cell invasion ability of the cells (P=0.0028
and P=0.0162, respectively). Taken together, miR-26b-5p inhibits
the proliferation, migration and invasion of human ICC cells.
miR-26b-5p suppresses ICC migration
and invasion by directly targeting S100A7
miRNAs exert their function by regulating the
expression of downstream target genes (21–23). In
order to identify the underlying mechanisms by which miR-26-5p
suppresses ICC invasion, the target prediction software TargetScan
was used to search for predicted direct target genes of miR-26b-5p.
An initial screening identified S100A7 as a potential direct target
of miR-26b-5p, which is a calcium-binding protein previously
implicated in cell invasion in a number of types of cancer
(24,25). In order to confirm this prediction,
S100A7 expression was compared between normal and invasive cell
lines. S100A7 was upregulated in iHCCC-9810 and iRBE compared with
HCCC-9810 and RBE cells (P=0.0002 and P=0.0024; Fig. 4A). Furthermore, S100A7 expression
levels were tested in ICC and mICC tumor samples, and the result
indicated that, in line with the result from cell lines, S100A7
expression in mICC tumors was significantly higher compared with
that in ICC samples (P=0.0003; Fig.
4B), indicating that S100A7 expression is inversely associated
with miR-26b-5p.
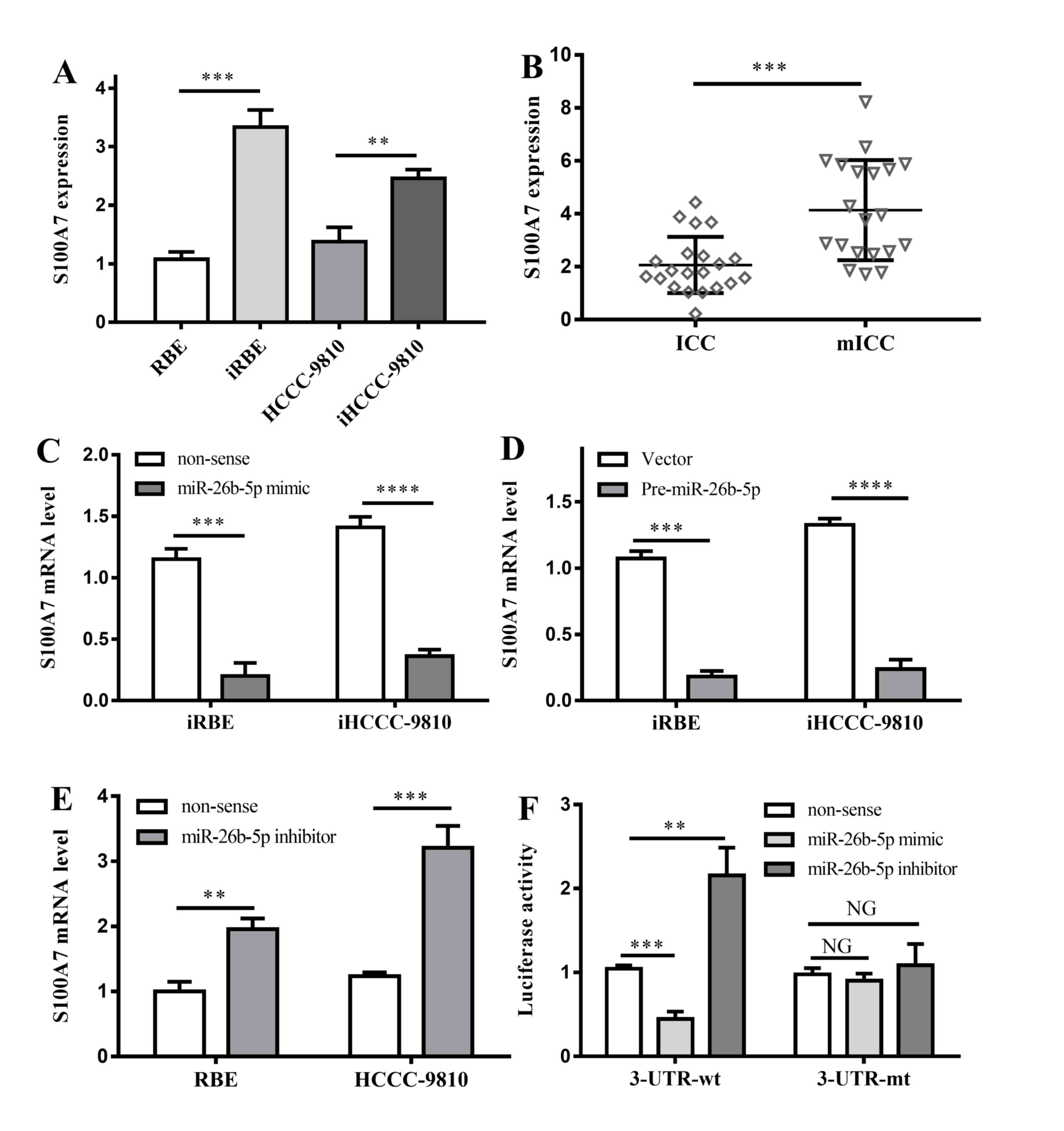 | Figure 4.Interactions between S100A7 and
miR-26b-5p. (A) S100A7 expression in 4 different kinds of cell
lines. **P<0.01, ***P<0.001 vs. non-invasive cell lines,
respectively; (B) S100A7 expression in ICC and mICC tumor tissues.
***P<0.001 vs. ICC tumor tissues. (C) The effects of miR-26b-5p
mimic on invasive cell lines. ***P<0.001, ****P<0.0001 vs.
non-sense controls. (D) The effects of pre-miR-26b-5p on invasive
cell lines. ***P<0.001, ****P<0.0001 vs. non-sense controls.
(E) The effects of miR-26b-5p inhibitor on RBE and HCCC-9810 cell
lines. **P<0.01, ***P<0.001 vs. non-sense controls. (F)
Effects of miR-26b-5p mimic and inhibitor on 3′UTR region of
S100A7. NG, P>0.05; **P<0.01; ***P<0.001; all vs.
non-sense control. ICC, intrahepatic cholangiocarcinoma; mICC, ICC
with lymphatic metastasis; 3′UTR, 3′ untranslated region; S100A7,
S100 calcium-binding protein A7; miR, microRNA; iRBE, invasive
subpopulation of RBE cells; iHCCC-9810, invasive subpopulation of
HCCC-9810 cells. |
To investigate the interaction between S100A7 and
miR-26b-5p, an miR-26b-5p mimic was transfected into iRBE and
iHCCC-9810 cells, in which miR-26b-5p expression was low, and the
S100A7 expression was measured. The data revealed that, compared
with the control group, transfection with an miR-26b-5p mimic
significantly lowered the expression of S100A7 (P=0.0002 and
P<0.0001; Fig. 4C). In addition,
transfection with pre-miR-26b-5p also inhibited S100A7 expression
in the two invasive cell lines (P=0.0001 and P<0.0001; Fig. 4D).
Additionally, an miR-26b-5p inhibitor was
transfected into RBE and HCCC-9810 cells, in which the expression
of miR-26b-5p was normal. The data demonstrated that following the
transfection with the miR-26b-5p inhibitor, S100A7 in the two cell
lines was significantly overexpressed (P=0.0019 and P=0.0007,
respectively; Fig. 4E). The results
verified that miR-26b-5p was associated with the inhibition of
S100A expression.
To determine whether miR-26b-5p binds directly to
the 3′UTR region of S100A7 mRNA, a dual-luciferase reporter assay
was performed. Transfection with an miR-26b-5p mimic markedly
reduced the luciferase activity of the 3′UTR-wt S100A7, whereas an
increase in the luciferase activity of the 3′UTR-wt S100A7 was
observed following the transfection of an miR-26b-5p inhibitor
(P=0.0004 and P=0.0038, respectively; Fig. 4F). However, no significant difference
in the luciferase activity of 3′UTR-mt S100A7 was observed
following the transfection with the miR-26b-5p mimic or inhibitor
(Fig. 4F).
It was also investigated whether the interaction
between miR-26b-5p and S100A7 led to alterations in the migration
and invasion ability of ICC cells. The non-sense control, the
miR-26b-5p mimic, the miR-26b-5p mimic with the 3′UTR-wt S100A7
vector or the miR-26b-5p mimic with the 3′UTR-wt S100A7 vector were
transfected into iRBE and iHCCC-9810 cells, and their invasive
ability was assessed with a Transwell invasion assay. The results
demonstrated that the invasion ability of cells transfected with
the miR-26b-5p mimic and 3′UTR-wt vector was not altered compared
with cells transfected with the miR-26b-5p mimic alone, whereas
transfection with the miR-26b-5p mimic and the 3′UTR-mt vector
increased the invasion ability (iRBE, P=0.0026; iHCCC-9810,
P=0.0054) and returned it to similar levels as the non-sense
control cells (Fig. 5A).
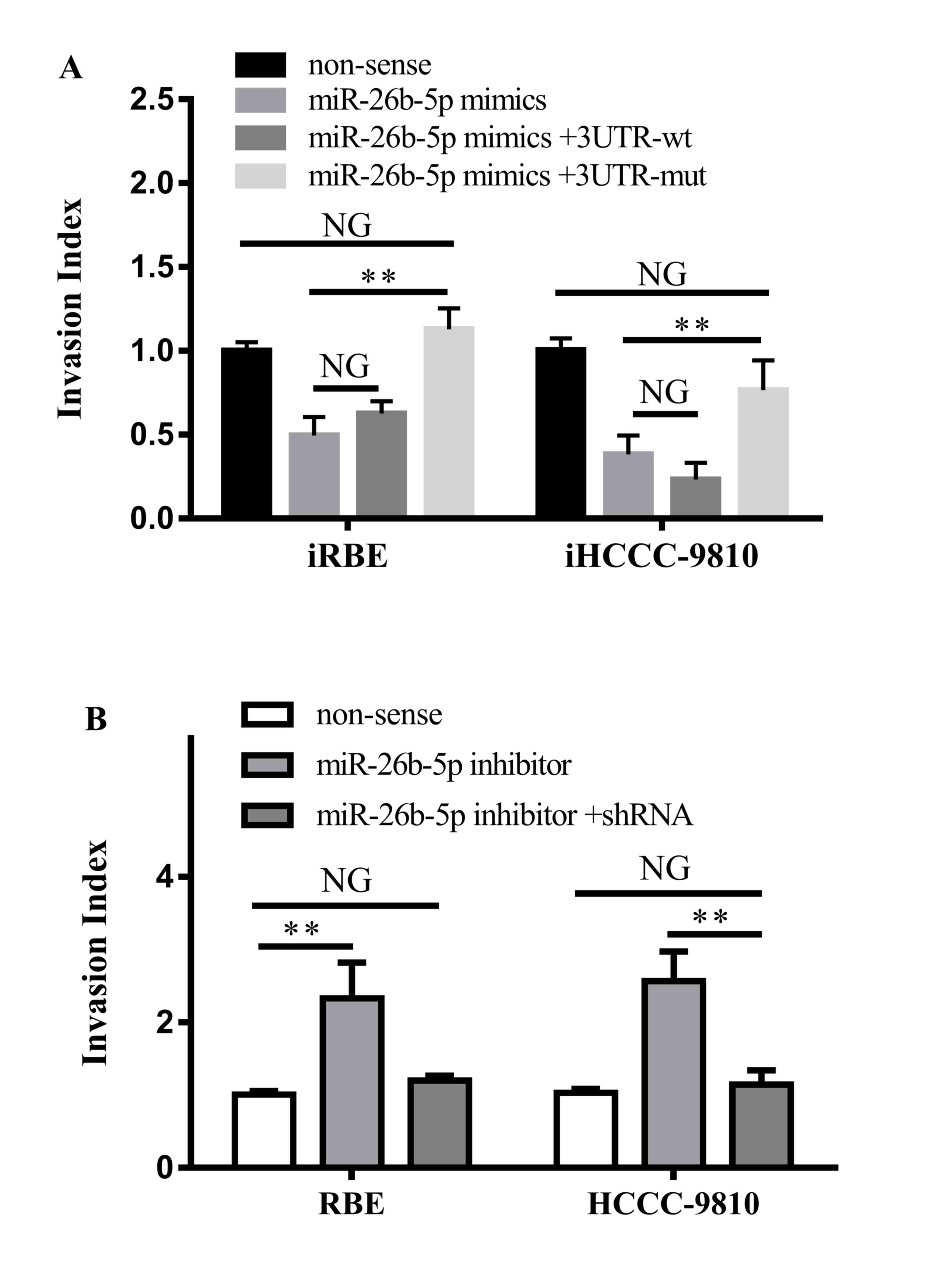 | Figure 5.Effects of the interaction between
S100A7 and miR-26b-5p on cell invasion. (A) Invasiveness of iRBE or
iHCCC-9810 cells following transfection with a non-sense control,
or an miR-25b-5p mimic with a 3′UTR wt or mt S100A7. (B)
Invasiveness of RBE and HCCC-9810 cells following transfection with
a non-sense control, an miR-26b-5p inhibitor or the inhibitor and
shRNA against S100A7. S100A7, S100 calcium-binding protein A7; miR,
microRNA; iRBE, invasive subpopulation of RBE cells; iHCCC-9810,
invasive subpopulation of HCCC-9810 cells; 3′UTR, 3′ untranslated
region; wt, wild-type; mt, mutant; shRNA, short hairpin RNA. NG,
P>0.05; **P<0.01; vs. non-sense controls respectively. |
Furthermore, the miR-26b-5p inhibitor or the
miR-26b-5p inhibitor with a small hairpin RNA against S100A7 were
transfected into RBE and HCCC-9810 cells. It was revealed that
transfection with the miR-26b-5p inhibitor increased the invasion
ability of the two cells (P=0.0097 and P=0.0027, respectively),
whereas transfection with the inhibitor and the shRNA against
S100A7 did not change the invasiveness of the cells (Fig. 5B). Therefore, the results demonstrated
that miR-26b-5p suppressed the invasive ability of ICC cells by
directly targeting the 3′UTR region of S100A7 mRNA.
Discussion
A number of studies have demonstrated that miRNAs
regulate the expression of oncogenes and tumor suppressors, which
may be associated with the initiation and development of HCC and
ICC (11,14,17). The
expression levels of five miRNAs potentially associated with cancer
development were investigated in ICC cells, and four candidate
miRNAs were selected on the basis of their differential expression
in invasive cell subpopulations. These candidate miRNAs were
further examined in 20 paired ICC and matched tumor-adjacent
tissues with RT-qPCR. The data revealed that only the expression of
miR-26b-5p in ICC was significantly lower than in matched
tumor-adjacent tissues. The expression of miR-26b-5p was also
significantly reduced in mICC tissue compared with non-metastatic
ICC tumors. The abnormal express of miR-26b-5p has been
demonstrated in types of cancer, including pulmonary and cancer
(26,27). However, to the best of our knowledge,
the present study was the first to identify that miR-26b-5p
exhibits a function in HCC.
Gain- and loss-of-function experiments demonstrated
that an miR-26b-5p mimic inhibited the proliferation, migration and
invasion of RBE and HCCC-9810 cells, whereas an miR-26b-5p
inhibitor led an increase in RBE and HCCC-9810 cell proliferation,
migration and invasion. These results indicated that miR-26b-5p
inhibited the proliferation, migration and invasion of ICC cells.
TargetScan was utilized to predict the target genes of miR-26b-5p,
and it was predicted that S100A7 may be a direct target. In order
to confirm this prediction, the expression level of S100A7 was
determined in RBE, iRBE, HCCC-9810 and iHCCC-9810 cells, and in
non-metastatic ICC and mICC tumors. The data revealed that S100A7
was upregulated in the two invasive cells and mICC tumors compared
with that in cell lines and non-metastatic ICC tumors.
Subsequently, the effect of the miR-26b-5p mimic or inhibitor on
S100A7 expression was determined, and S100A7 expression was
demonstrated to be inversely associated with miR-26b-5p expression
in ICC. Furthermore, the effect of the interaction between
miR-26b-5p and S100A7 on the invasive ability of the cells was
investigated. These results demonstrated that the miR-26b-5p mimic
decreased the invasive ability of the invasive cell sub-populations
by downregulating S100A7 expression, whereas the miR-26b-5p
inhibitor increased the invasion ability of the RBE and HCCC-9810
cells by upregulating S100A7 expression. Therefore, to the best of
our knowledge, it was demonstrated for the first time that
miR-26b-5p inhibits cell migration and invasion by suppressing
S100A7 expression in ICC.
In summary, the present study revealed that
miR-26b-5p was downregulated in ICC tissues and cells, particularly
in invasive ICC subpopulations and tumors from patients with mICC.
In addition, it was revealed that miR-26b-5p inhibited the
proliferation, migration and invasion of ICC cells by targeting
S100A7. In conclusion, the present study demonstrated that
miR-26b-5p functions as a tumor suppressive miRNA, and inhibits the
invasive behavior of ICC by inhibiting S100A7.
Acknowledgements
The present study was supported by a grant from the
Natural Science Foundation of PRC (grant no. 81072026).
References
|
1
|
Bartella I and Dufour JF: Clinical
diagnosis and staging of intrahepatic cholangiocarcinoma. J
Gastrointestin Liver Dis. 24:481–489. 2015.PubMed/NCBI
|
|
2
|
Guo LH and Xu HX: Contrast-enhanced
ultrasound in the diagnosis of hepatocellular carcinoma and
intrahepatic cholangiocarcinoma: Controversy over the ASSLD
guideline. Biomed Res Int. 2015:3491722015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lafaro KJ, Cosgrove D, Geschwind JF, Kamel
I, Herman JM and Pawlik TM: Multidisciplinary care of patients with
intrahepatic cholangiocarcinoma: Updates in management.
Gastroenterol Res Pract. 2015:8608612015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Brandi G, Venturi M, Pantaleo MA and G;
GICO Ercolani: Cholangiocarcinoma: Current opinion on clinical
practice diagnostic and therapeutic algorithms: A review of the
literature and a long-standing experience of a referral center. Dig
Liver Dis. 48:231–241. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Maithel SK, Gamblin TC, Kamel I,
Corona-Villalobos CP, Thomas M and Pawlik TM: Multidisciplinary
approaches to intrahepatic cholangiocarcinoma. Cancer.
119:3929–3942. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hyder O, Hatzaras I, Sotiropoulos GC, Paul
A, Alexandrescu S, Marques H, Pulitano C, Barroso E, Clary BM,
Aldrighetti L, et al: Recurrence after operative management of
intrahepatic cholangiocarcinoma. Surgery. 153:811–818. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hawkins PG and Morris KV: RNA and
transcriptional modulation of gene expression. Cell Cycle.
7:602–607. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kunej T, Godnic I, Ferdin J, Horvat S,
Dovc P and Calin GA: Epigenetic regulation of microRNAs in cancer:
An integrated review of literature. Mutat Res. 717:77–84. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Haga H, Yan I, Takahashi K, Wood J and
Patel T: Emerging insights into the role of microRNAs in the
pathogenesis of cholangiocarcinoma. Gene Expr. 16:93–99. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Maemura K, Natsugoe S and Takao S:
Molecular mechanism of cholangiocarcinoma carcinogenesis. J
Hepatobiliary Pancreat Sci. 21:754–760. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Piontek K and Selaru FM: MicroRNAs in the
biology and diagnosis of cholangiocarcinoma. Semin Liver Dis.
35:55–62. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chinese Medical Association, . The updated
grading standards for primary hepatic cancerClinical Practice
Guidelines – Tumor. People's Medical Publishing House; China: pp.
325–326. 2001
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sheng Y, Li J, Zou C, Wang S, Cao Y, Zhang
J, Huang A and Tang H: Downregulation of miR-101-3p by hepatitis B
virus promotes proliferation and migration of hepatocellular
carcinoma cells by targeting Rab5a. Arch Virol. 159:2397–2410.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Augello C, Gianelli U, Savi F, Moro A,
Bonoldi E, Gambacorta M, Vaira V, Baldini L and Bosari S: MicroRNA
as potential biomarker in HCV-associated diffuse large B-cell
lymphoma. J Clin Pathol. 67:697–701. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ertekin S, Yildirim O, Dinc E, Ayaz L,
Fidanci SB and Tamer L: Evaluation of circulating miRNAs in wet
age-related macular degeneration. Mol Vis. 20:1057–1066.
2014.PubMed/NCBI
|
|
19
|
Kozubek J, Ma Z, Fleming E, Duggan T, Wu
R, Shin DG and Dadras SS: In-depth characterization of microRNA
transcriptome in melanoma. PLoS One. 8:e726992013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yan S, Wang J, Zhang W and Dai J:
Circulating MicroRNA profiles altered in mice after 28 days
exposure to perfluorooctanoic acid. Toxicol Lett. Oct 24–2013.(Epub
ahead of print).
|
|
21
|
Tessitore A, Cicciarelli G, Del Vecchio F,
Gaggiano A, Verzella D, Fischietti M, Vecchiotti D, Capece D,
Zazzeroni F and Alesse E: MicroRNAs in the DNA damage/repair
network and cancer. Int J Genomics. 2014:8202482014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Son DJ, Kumar S, Takabe W, Kim CW, Ni CW,
Alberts-Grill N, Jang IH, Kim S, Kim W, Won Kang S, et al: The
atypical mechanosensitive microRNA-712 derived from pre-ribosomal
RNA induces endothelial inflammation and atherosclerosis. Nat
Commun. 4:30002013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Sampath D, Liu C, Vasan K, Sulda M,
Puduvalli VK, Wierda WG and Keating MJ: Histone deacetylases
mediate the silencing of miR-15a, miR-16, and miR-29b in chronic
lymphocytic leukemia. Blood. 119:1162–1172. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Dey KK, Sarkar S, Pal I, Das S, Dey G,
Bharti R, Banik P, Ray JG, Maity S, Kulavi I and Mandal M: Erratum
to: Mechanistic attributes of S100A7 (psoriasin) in resistance of
anoikis resulting tumor progression in squamous cell carcinoma of
the oral cavity. Cancer Cell Int. 15:942015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Liu G, Wu Q, Liu G, Song X and Zhang J:
Psoriasin (S100A7) is a novel biomarker for lung squamous cell
carcinoma in humans. Cancer Cell Int. 15:182015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wu T, Chen W, Liu S, Lu H, Wang H, Kong D,
Huang X, Kong Q, Ning Y and Lu Z: Huaier suppresses proliferation
and induces apoptosis in human pulmonary cancer cells via
upregulation of miR-26b-5p. FEBS Lett. 588:2107–2114. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Armstrong DA, Green BB, Seigne JD, Schned
AR and Marsit CJ: MicroRNA molecular profiling from matched tumor
and bio-fluids in bladder cancer. Molecular Cancer. 14:1942015.
View Article : Google Scholar : PubMed/NCBI
|