Introduction
Lung cancer is the leading cause of cancer-related
deaths in men and the second leading cause of cancer death in women
worldwide and presents a serious problem to global health (1). It was estimated that 1.8 million new
lung cancer cases and 1.6 million lung cancer related deaths
occurred in 2012 worldwide, accounting for approximately 19% of all
cancer deaths (2). In recent decades,
despite of great research effort in diagnosis and treatment for
lung cancer, progress in the treatment is still slow (3). Non-small cell lung cancer (NSCLC) is the
major lung cancer, accounts for approximately 85% of lung cancer.
Lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC)
are the major two pathological subtypes of NSCLC and in many ways
they are different, such as originate, biological patterns and
molecular characteristics (4–6). Related studies on NSCLC should be
analyzed separately according to histological type.
Interleukin-6 (IL-6) signaling through IL-6 receptor
(IL-6R) regulates cell growth and differentiation and plays an
important role in the immune response (7). This signaling pathway can also promote
tumor growth which has both pro-inflammatory and anti-inflammatory
effect (8). Evidence has shown that
the IL-6 signaling pathway contribute to the pathogenesis of NSCLC
(9). Several studies have revealed
the role of IL-6 in NSCLC and suggested that it promotes tumor
growth and survival (10–12). Increased expression of IL-6R in human
LUSC-derived cells (HARA-B) has been shown in vitro
(13) and in a murine model of brain
metastasis (14). Tocilizumab is an
anti-IL-6R antibody, upon application, the stimulated growth of
HARA-B cells was significantly inhibited and when it was injected
to the animal model, the volume of metastatic focus was
significantly smaller (14). Studies
also have demonstrated that blockade of IL-6R can significantly
suppress the proliferation of NSCLC cell and reduce the mRNA levels
of IL-6R (15,16). Meanwhile, tocilizumab also seems to be
effective for lung cancer cachexia (17,18).
However, there was no study investigating the prognostic effect of
IL-6R on lung cancer, which is addressed by our current study.
Materials and methods
Search strategy
We searched public databases such as The Cancer
Genome Atlas (TCGA) and Gene Expression Ominibus (GEO database;
last update by July 03, 2017) using the keywords ‘Lung cancer’. The
search strategy is designed as follows: The study type was set as
‘expression profiling by array. The entry type was set as
‘datasets’. The sample size of all selected datasets should be
greater or equal to 100. The organism was homo sapiens. Database
searching was carried out by two researchers independently.
Data extraction and quality
assessment
Data from all eligible datasets were abstracted
independently by two authors, using information recorded as
follows: First author's surname, publication year, origin of
population, sample number, tumor stage, follow-up period and clinic
outcome. We separate those microarray datasets into LUAD and LUSC.
HRs and 95% CIs were evaluated by Cox's proportional hazards
model.
The quality of all eligible studies was assessed
according to the Newcastle-Ottawa Quality Assessment Scale (NOS) by
two researchers independently (19).
The quality scores span from 0 to 9, and higher the score is,
higher the quality is.
Statistical analysis
For those public microarray data, gene expression
was represented by metric variables. We use Cutoff Finder
(http://molpath.charite.de/cutoff) to
determine a cutoff point and stratify patients into two groups
(20). The range of IL-6R mRNA values
for each data and the corresponding cutoff values were listed in
Table I. HRs and 95% CIs were
calculated to measure the effective prognostic value of expression
of IL-6R mRNA in LUAD and LUSC patients. Pooled HRs were carried
out using STATA software package (version 12.0; Stata Corp LP,
College Station, TX, USA). All P-values were obtained upon two
tailed analysis.
 | Table I.Cut-off value of IL-6R. |
Table I.
Cut-off value of IL-6R.
| Datasets | Minimum | P25 | Median | Mean | P75 | Maximum | Cut-off value |
|---|
| LUAD |
|
|
|
|
|
|
|
|
GSE14814 | 4.211 | 4.779 | 5.027 | 5.134 | 5.394 | 6.611 | 4.607 |
|
GSE30219 | 5.127 | 6.343 | 6.908 | 7.019 | 7.686 | 9.247 | 7.648 |
|
GSE37745 | 3.970 | 7.232 | 7.698 | 7.697 | 8.272 | 10.200 | 7.789 |
|
GSE42127 | 3.670 | 4.475 | 4.930 | 4.945 | 5.405 | 7.680 | 5.515 |
|
GSE50081 | 5.049 | 7.281 | 7.809 | 7.823 | 8.405 | 10.049 | 7.675 |
|
GSE68465 | 31.955 | 246.235 | 361.102 | 437.587 | 543.700 | 1940.570 | 393.500 |
|
GSE68571 | −41.850 | 17.188 | 52.725 | 69.762 | 101.213 | 297.650 | 44.020 |
|
TCGA | 4.360 | 8.364 | 9.037 | 8.970 | 9.274 | 11.608 | 9.125 |
| LUSC |
|
|
|
|
|
|
|
|
GSE14814 | 4.324 | 4.644 | 4.912 | 4.909 | 5.076 | 5.871 | 5.072 |
|
GSE30219 | 4.232 | 5.430 | 6.004 | 5.946 | 6.453 | 7.988 | 5.741 |
|
GSE37745 | 4.523 | 6.395 | 6.992 | 6.851 | 7.458 | 8.770 | 7.637 |
|
GSE42127 | 2.830 | 4.100 | 4.390 | 4.456 | 4.730 | 6.260 | 4.115 |
|
GSE50081 | 5.475 | 6.681 | 7.144 | 7.125 | 7.421 | 9.009 | 6.678 |
|
TCGA | 3.751 | 7.456 | 8.286 | 8.171 | 8.879 | 10.982 | 9.134 |
In The Cancer Genome Atlas (TCGA) lung
adenocarcinoma dataset and TCGA lung squamous cell carcinoma
dataset, there were gene expression data in both tumor and normal
tissues. We used paired test to compare the differences in IL-6R
mRNA expression between tumor and adjacent normal tissues.
For each dataset, we calculated the correlation
coefficient between IL-6R and the remaining genes, and then matched
the coefficients in all datasets. Genes with absolute correlation
coefficient which were greater than 0.4 in half or more
publications were extracted. 193 genes in LUAD and 101 genes in
LUSC were included in subsequent analysis (Table II).
 | Table II.Gene list in LUAD and LUSC. |
Table II.
Gene list in LUAD and LUSC.
| Cancer | Gene names |
|---|
| LUAD | ABCC3, ADCY9,
AKAP13, ALDH2, ALDH5A1, ALOX15B, APLP2, APOH, AQP3, ARHGAP31,
ASF1B, ASPM, ATP13A4, AURKB, BFAR, BIRC5, BUB1, BUB1B, C16orf89,
C1orf116, C5orf46, CC2, CCNB1, CCNB2, CCND3, CD302, CD81, CDC20,
CDC25A, CDC25C, CDC45, CDC6, CDCA2, CDCA3, CDCA5, CDCA8, CDK1,
CDKL2, CDKN3, CDT1, CEBPA, CENPM, CENPN, CENPW, CEP55, CHEK1, CISH,
CITED2, CKS2, CLIC3, CLIC5, COL4A3, COL4A4, CPM, CRY2, CTDSPL,
CTSH, CYP4B1, DEPDC1, DEPDC1B, DJC9, DLC1, DLGAP5, DYNLL1, ECHDC2,
ESPL1, EVPL, EXO1, FAM184A, FAM64A, FAM83D, FANCG, FANCI, FBP1,
FBXO5, FCGRT, FEN1, FMO5, FNIP2, FOLR1, FOXM1, GALNT11, GAPDH,
GGT1, GGTLC1, GINS1, GPD1L, GPR116, GTSE1, H2AFX, HIP1, HJURP, HLF,
HNF1B, ICAM4, ICAM5, IL6R, INMT, KIAA0101, KIF11, KIF18B, KIF20A,
KIF23, KIF2C, KIF4A, KIFC1, KNTC1, LOC101930052, LRRK2, MAD2L1,
MALL, MAMDC2, MELK, MGLL, MGRN1, MIR3658, MIR6883, MKI67, MLLT11,
MTHFD2, MUC1, NBEAL1, NCAPG, NCAPH, NDC80, NEK2, NPC2, NRM, NUF2,
NUSAP1, OCLN, OIP5, PARM1, PARPBP, PBK, PC, PCDP1, PEBP4, PI4KB,
PLA2G1B, PLK1, PLLP, PMVK, POLE2, PON3, PRC1, PRR15L, PSA, PSRC1,
PTS, PTTG1, RAD51AP1, RAD54L, RFC4, RRM2, RSE1, RSEH2A, SELENBP1,
SFTA1P, SFTPD, SKA1, SKA3, SLC22A3, SLC34A2, SLC41A1, SNX30, SPAG5,
SPAG9, SPC25, ST3GAL5, STAT6, STIL, SUSD2, SYNE1, TLR2, TLR5,
TMEM163, TMEM243, TMPRSS2, TPX2, TRIP13, TROAP, TTK, TUBA1C, TUBB,
TYMS, UBE2C, UBE2T, UHRF1, USP13, ZBTB4, ZNF385B, ZWINT |
| LUSC | ADAP2, ADCY9,
ANXA11, APOBR, ARHGEF3, BTK, BUB1B, C1RL, CCR1, CD37, CD4, CD52,
CD93, CD97, CEBPD, COQ3, CORO1A, CRTAP, CSE1L, CTSH, DENND2D, DHX9,
DOK2, DUT, ECT2, EPAS1, EZH2, FBXO5, FGR, FHOD1, FOS, GIMAP6,
HDAC2, HLA-DPB1, HLA-DQB1, IER2, IFITM1, IL2RB, IL6R, ITGAL, JUNB,
KIF15, KLF10, LCP2, LRP10, LSM4, LST1, MAST3, MCM6, MOSPD1, MSH6,
MSN, MVP, MYO1F, P2RY13, PAGR1, PDCD5, PDE4A, PLEKHO2, PLIN3,
POLA1, PPP1CC, PPP1R16B, PRKCB, PTAFR, PTK2B, PTPN6, RAB27A, RAB31,
RAD51AP1, RAD54B, RCN2, RHOG, RPL22, S1PR1, SASH3, SIGLEC1, SLA,
SLCO2B1, THEMIS2, TMPO, TNFRSF1A, TNFRSF1B, TPP1, TRANK1, TRPV2,
TYROBP, VASP, VWF, ZFYVE26, ARHGAP30, ARHGAP9, C1orf162, CARD6,
CXCL16, GIMAP1, LOC100129034, LRRCC1, SARNP, TNFAIP8L2, ZNF664 |
Functional enrichment analysis of genes whose
expressions are significantly correlated that of IL-6R was
performed to allow the identification of biological processes or
functions associated with IL-6R expression. In this study, the
Database for Annotation, Visualization and Integrated Discovery
(DAVID) was used to analyze gene enrichment and pathway analysis to
explore the biological processes of gene enrichment (https://david.ncifcrf.gov/summary.jsp).
Results
Study characteristics
A total of 7 related studies were identified from
the GEO database (GSE14814 (21),
GSE30219 (22), GSE37745 (23), GSE42127 (24), GSE50081 (25), GSE68465 (26), GSE68571 (27)). Including TCGA lung adenocarcinoma and
TCGA lung squamous cell carcinoma datasets, 9 datasets were
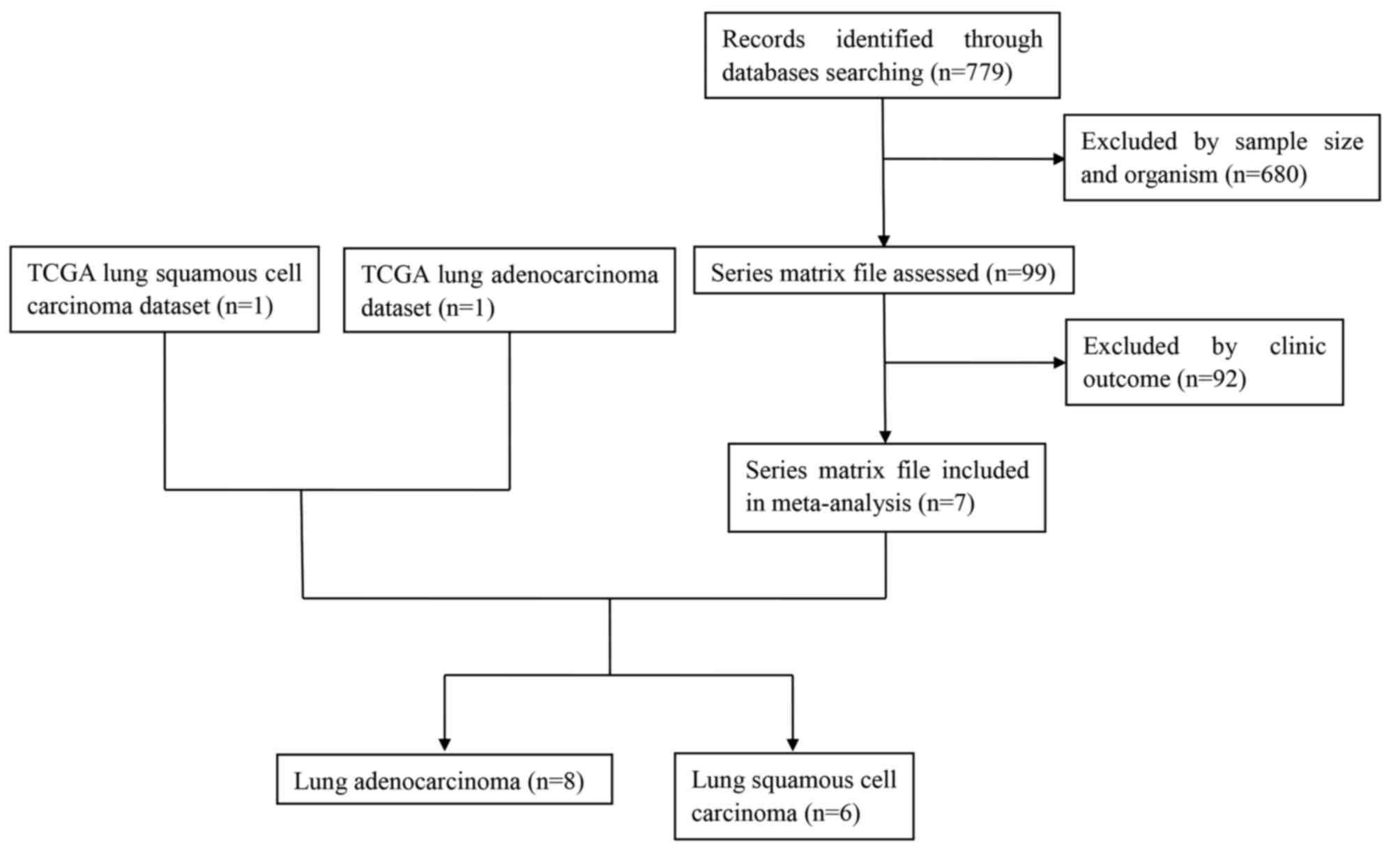
included in our analysis. In the initial screening, a total of 779
potentially relevant datasets from the GEO database were selected
for keyword retrieval. A total of 680 datasets were retrieved after
screening sample size and organism. After examination of summary
and the clinic outcome of those data, a total of 7 microarray
datasets from the GEO database that met the inclusion criteria were
included in the present study. Finally, 8 datasets and 6 datasets
for adenocarcinoma and squamous cell carcinoma respectively were
further analyzed (Fig. 1). Table III showed the baseline
characteristics of all included studies. Date of 1,536 LUAD and 739
LUSC patients from Canada, France, UK and USA were included in this
analysis.
 | Table III.Baseline characteristics of
microarray data. |
Table III.
Baseline characteristics of
microarray data.
| Datasets | Author | Year | Country | Median duration,
months (range) | Sample size | Stage | Outcome
measures | Quality score | Journal | IF | (Refs.) |
|---|
| LUAD |
|
|
|
|
|
|
|
|
|
|
|
| GSE14814 | Zhu CQ | 2010 | Canada | 60.84
(0.36–109.08) | 70 | I/II (41/29) | OS | 8 | Journal of clinical
oncology | 24.008 | (21) |
| GSE30219 | Rousseaux S | 2013 | France | 68.00 (0–221) | 85 | NR | OS | 7 | Science
translational medicine | 16.796 | (22) |
| GSE37745 | Botling J | 2013 | UK | 47.80
(0.20–187.79) | 106 | I/II/III/IV
(70/19/13/4) | OS | 8 | Clinical cancer
research | 9.619 | (23) |
| GSE42127 | Tang H | 2013 | USA | 45.60 (0–132) | 133 | I/II/III/IV
(89/22/20/1) | OS | 7 | Clinical cancer
research | 9.619 | (24) |
| GSE50081 | Der SD | 2014 | Canada | 52.44
(1.08–130.56) | 129 | I/II (93/36) | OS | 6 | Journal of thoracic
oncology | 6.595 | (25) |
| GSE68465 | Shedden K | 2008 | USA | 47.00
(0.03–204.00) | 442 | NR | OS | 9 | Nature
medicine | 29.886 | (26) |
| GSE68571 | Beer DG | 2002 | USA | 29.50
(1.50–110.60) | 86 | I/III (67/19) | OS | 8 | Nature
medicine | 29.886 | (27) |
| TCGA | TCGA | 2015 | USA | 15.35
(0.00–223.96) | 485 | I/II/III/IV
(262/118/79/25) | OS | 8 | – | – |
|
| LUSC |
|
|
|
|
|
|
|
|
|
|
|
| GSE14814 | Zhu CQ | 2010 | Canada | 73.56
(8.28–111.48) | 52 | I/II (25/27) | OS | 8 | Journal of clinical
oncology | 24.008 | (21) |
| GSE30219 | Rousseaux S | 2013 | France | 59.00 (1–256) | 61 | NR | OS | 7 | Science
translational medicine | 16.796 | (22) |
| GSE37745 | Botling J | 2013 | UK | 36.36
(0.20–179.80) | 66 | I/II/III
(40/15/11) | OS | 8 | Clinical cancer
research | 9.619 | (23) |
| GSE42127 | Tang H | 2013 | USA | 48.00
(0.00–128.40) | 43 | I/II/III
(23/10/10) | OS | 7 | Clinical cancer
research | 9.619 | (24) |
| GSE50081 | Der SD | 2014 | Canada | 62.16
(5.76–144.48) | 43 | I/II (27/16) | OS | 6 | Journal of thoracic
oncology | 6.595 | (25) |
| TCGA | TCGA | 2015 | USA | 17.27
(0.03–174.08) | 474 | I/II/III/IV
(232148/84/7) | OS | 8 | – | – |
|
A quality assessment of the eligible datasets
included in this meta-analysis has been performed according to
Newcastle-Ottawa Quality Assessment Scale (NOS). The quality score
span was from 6 to 9 and the mean score was 7.63 for LUAD and the
quality score span was from 6 to 8 and the mean score was 7.33 for
LUSC. The impact factors of the journals where the studies were
published were of high caliber (Table
III). Thus, all of those studies were included in following
analysis.
Overall survival
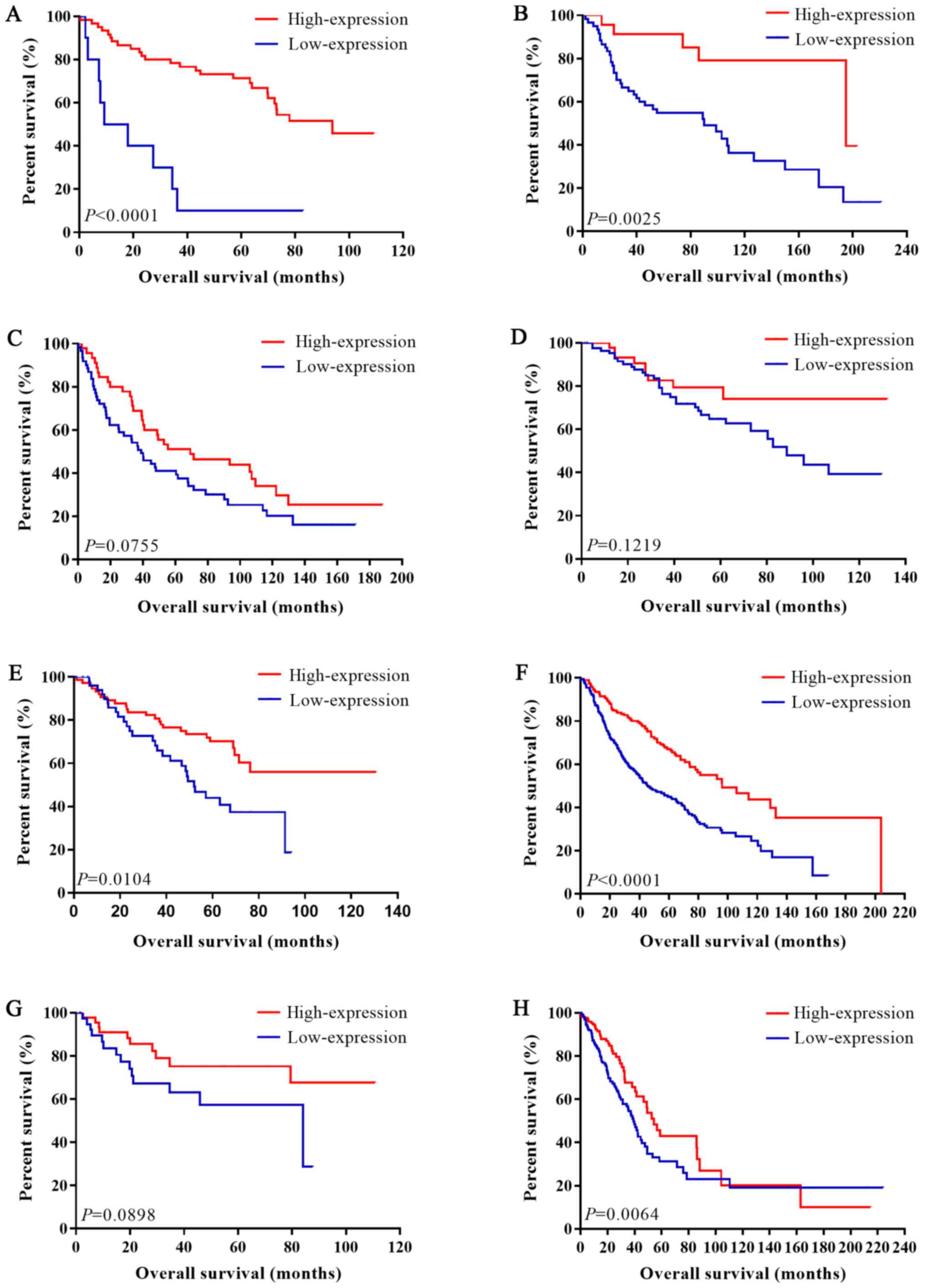
Univariate analysis and multivariate analysis were
respectively carried out for each dataset. P-values, HRs and 95%
CIs of IL-6R mRNA in each article for LUAD and LUSC were shown in
Table IV and Figs. 2 and 3.
 | Table IV.HR of interleukin-6R mRNA for OS. |
Table IV.
HR of interleukin-6R mRNA for OS.
|
| Univariate
analysis | Multivariate
analysis |
|---|
|
|
|
|
|---|
| Datasets | HR | LCI | UCI |
P-valuea | HR | LCI | UCI |
P-valueb |
|---|
| LUAD |
|
|
|
|
|
|
|
|
|
GSE14814c | 0.201 | 0.092 | 0.443 |
<0.001 | 0.190 | 0.076 | 0.474 |
<0.001 |
|
GSE30219d | 0.237 | 0.093 | 0.603 | 0.003 | 0.215 | 0.081 | 0.566 | 0.002 |
|
GSE37745c | 0.659 | 0.416 | 1.044 | 0.076 | 0.589 | 0.303 | 1.147 | 0.120 |
|
GSE42127c | 0.559 | 0.267 | 1.168 | 0.122 | 0.693 | 0.319 | 1.505 | 0.354 |
|
GSE50081e | 0.493 | 0.287 | 0.846 | 0.010 | 0.656 | 0.370 | 1.163 | 0.149 |
|
GSE68465f | 0.492 | 0.375 | 0.646 |
<0.001 | 0.566 | 0.426 | 0.752 |
<0.0001 |
|
GSE68571g | 0.492 | 0.217 | 1.117 | 0.089 | 0.602 | 0.256 | 1.418 | 0.246 |
|
TCGAh | 0.617 | 0.436 | 0.873 | 0.006 | 0.515 | 0.344 | 0.772 | 0.001 |
| LUSC |
|
|
|
|
|
|
|
|
|
GSE14814i | 1.701 | 0.669 | 4.323 | 0.264 | 1.589 | 0.597 | 4.227 | 0.354 |
|
GSE30219d | 1.440 | 0.748 | 2.775 | 0.275 | 1.559 | 0.797 | 3.050 | 0.194 |
|
GSE37745c | 2.599 | 1.344 | 5.029 | 0.005 | 4.071 | 1.436 | 11.537 | 0.008 |
|
GSE42127c | 9.906 | 1.321 | 74.297 | 0.026 | 8.577 | 1.084 | 67.868 | 0.042 |
|
GSE50081j | 0.442 | 0.149 | 1.312 | 0.141 | 0.387 | 0.119 | 1.257 | 0.114 |
|
TCGAh | 1.375 | 0.954 | 1.981 | 0.088 | 1.308 | 0.899 | 1.903 | 0.161 |
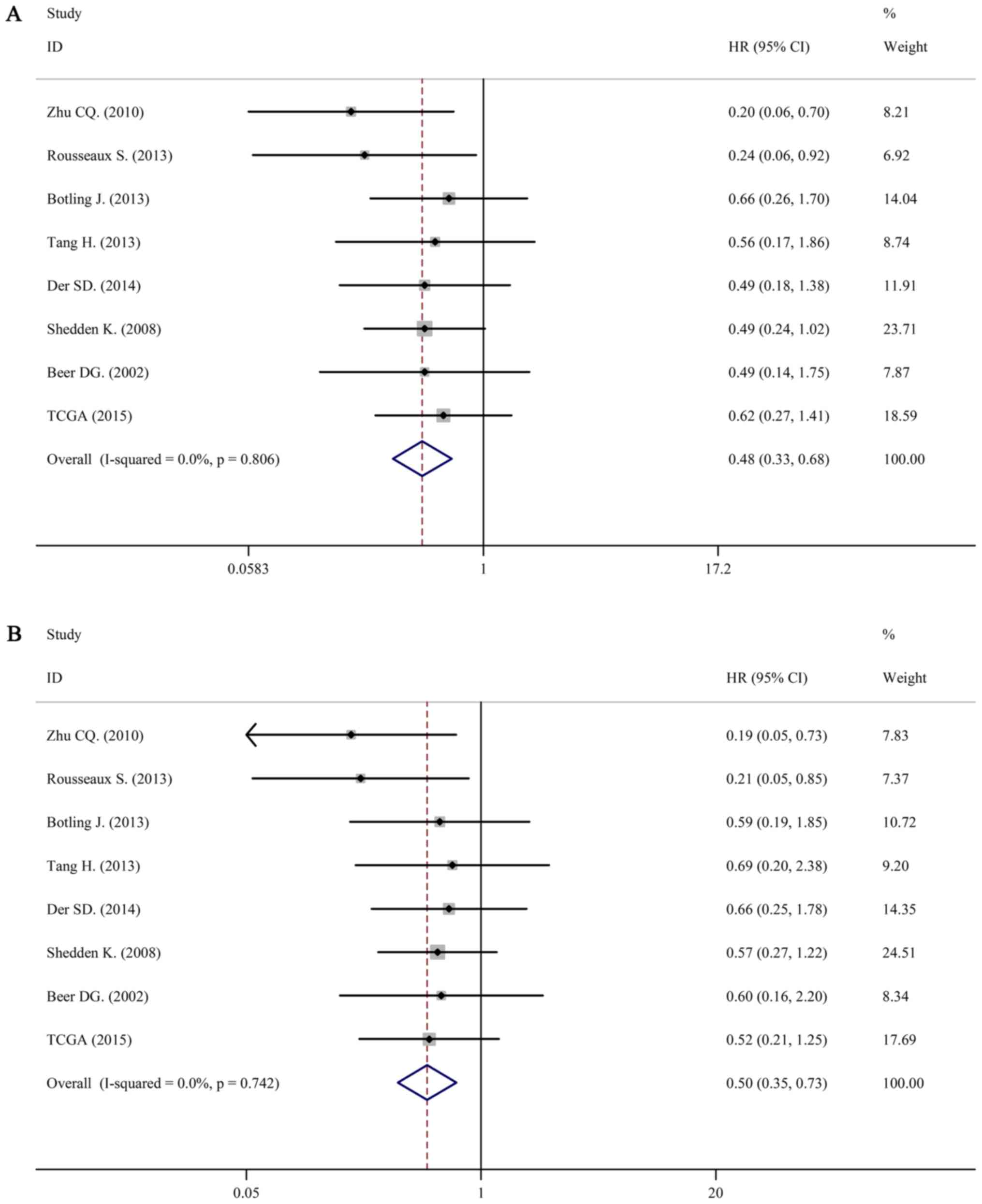
For LUAD, there was no obvious statistical
heterogeneity in all of those datasets both in univariate survival
analysis and multivariate survival analysis (I2=0.0%,
P=0.806; I2=0.0%, P=0.742), a fixed-effects model was
used to calculate the pool HRs. Our analysis demonstrated that a
higher expression of IL-6R mRNA was significantly associated with
better overall survival (OS) (pooled HR=0.50; 95% CI: 0.33–0.68 in
univariate analysis; pooled HR=0.50; 95% CI: 0.35–0.73 in
multivariate analysis). The forest plots of study-specific HRs for
OS were presented in Fig. 4.
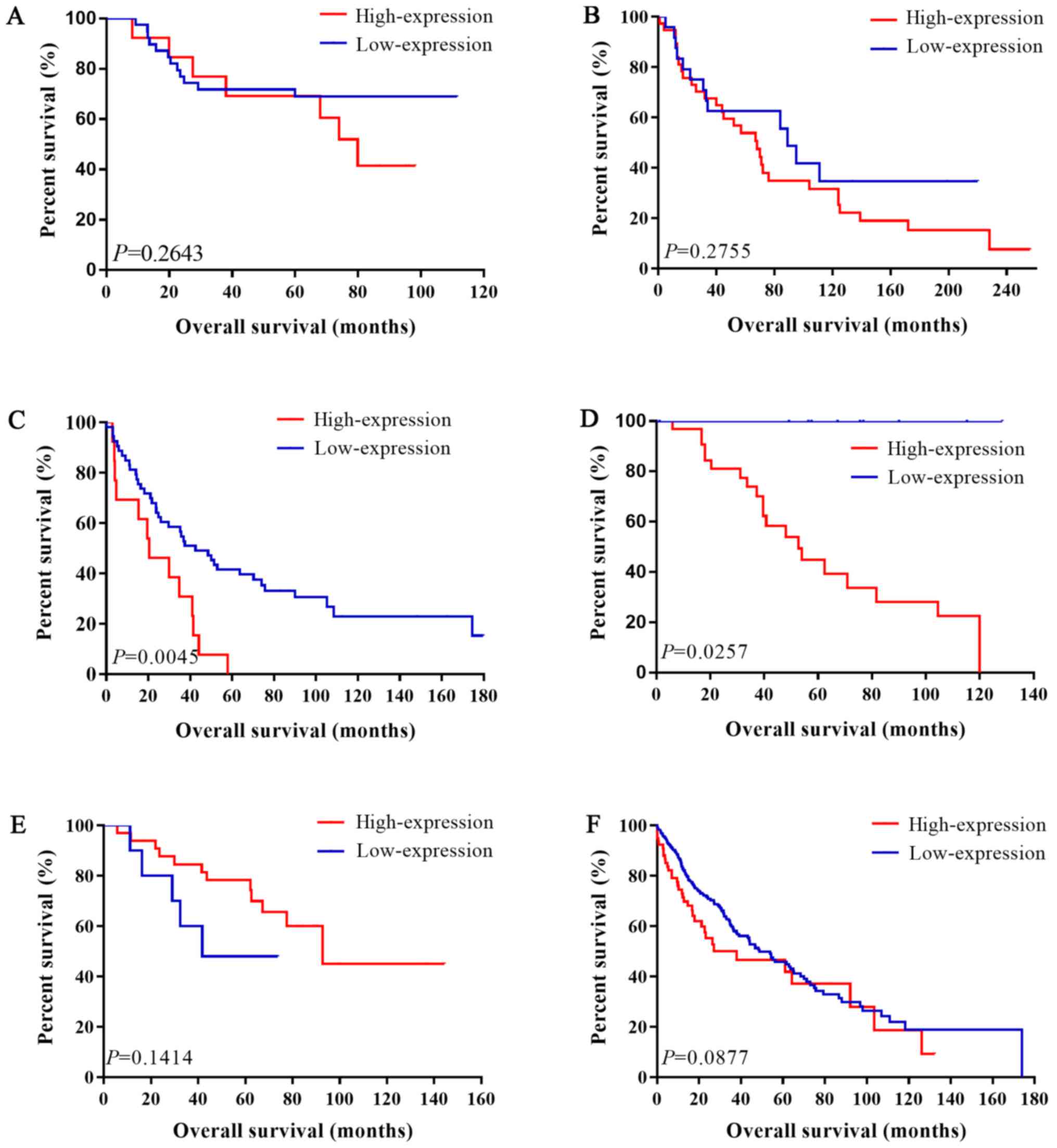
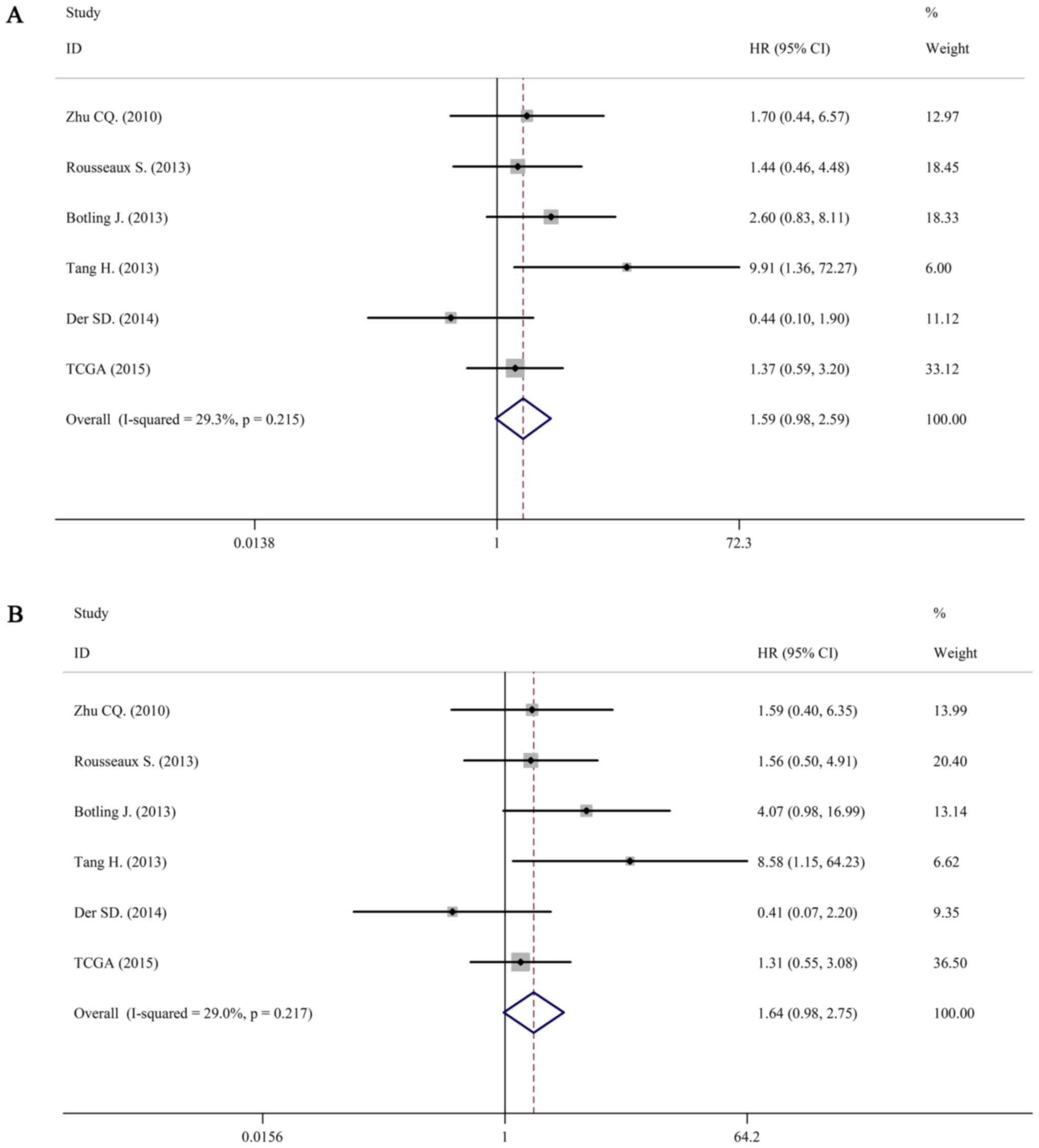
For LUSC, we still used a fixed-effects model to
pool the HRs (I2=29.3%, P=0.215; I2=36.5%,
P=0.217). Interestingly, there was no association between IL-6 mRNA
and OS in patients with LUSC (pooled HR=1.59; 95% CI: 0.98–2.59 in
univariate analysis; pooled HR=1.64; 95% CI: 0.98–2.75 in
multivariate analysis). The forest plots of study-specific HRs for
OS were presented in Fig. 5.
Correlation between IL-6 and IL-6R,
IL-6R and IL-6ST
The IL-6 receptor is a protein complex consisting of
an alpha chain, IL-6R, and IL-6 signal transducer (IL-6ST).
Relationship between the mRNA expression of IL-6 and IL-6R, and
between IL-6R and IL-6ST were all analyzed in those datasets. We
then performed a meta-analysis based on Fisher's z transformation.
Interestingly, IL-6 and IL-6R were negatively correlated in LUAD
(pooled r=−0.199, P<0.001), while they were positively
correlated in LUSC (pooled r=0.288, P=0.001). The correlation
coefficient between IL-6R and IL-6ST in LUAD was similar with
correlation coefficient in LUSC (pooled r=0.331, P<0.001 in LUAD
and pooled r=0.334, P<0.001 in LUSC; Table V).
 | Table V.Correlation between IL-6 and IL-6R,
IL-6R and IL-6ST. |
Table V.
Correlation between IL-6 and IL-6R,
IL-6R and IL-6ST.
|
| IL-6 vs. IL-6R | IL-6R vs.
IL-6ST |
|---|
|
|
|
|
|---|
| Datasets | r | P-value | r | P-value |
|---|
| LUAD |
|
|
|
|
|
GSE14814 | −0.345 |
0.004 | 0.141 |
0.245 |
|
GSE30219 | −0.515 |
<0.001 | 0.506 |
<0.001 |
|
GSE37745 | −0.107 |
0.272 | 0.294 |
0.002 |
|
GSE42127 |
0.041 |
0.642 | 0.243 |
0.005 |
|
GSE50081 | −0.207 |
0.019 | 0.363 |
<0.001 |
|
GSE68465 | −0.166 |
0.001 | 0.271 |
<0.001 |
|
GSE68571 | −0.194 |
0.073 | 0.191 |
0.079 |
|
TCGA | −0.165 |
<0.001 | 0.527 |
<0.001 |
| Pooled
r | −0.199 |
<0.001 | 0.331 |
<0.001 |
| LUSC |
|
|
|
|
|
GSE14814 |
0.395 |
0.004 | 0.073 |
0.606 |
|
GSE30219 |
0.036 |
0.782 | 0.469 |
0.000 |
|
GSE37745 |
0.417 |
0.001 | 0.369 |
0.002 |
|
GSE42127 |
0.610 |
<0.001 | 0.272 |
0.077 |
|
GSE50081 |
0.069 |
0.661 | 0.155 |
0.320 |
|
TCGA |
0.178 |
<0.001 | 0.465 |
<0.001 |
| Pooled
r |
0.288 |
0.001 | 0.334 |
<0.001 |
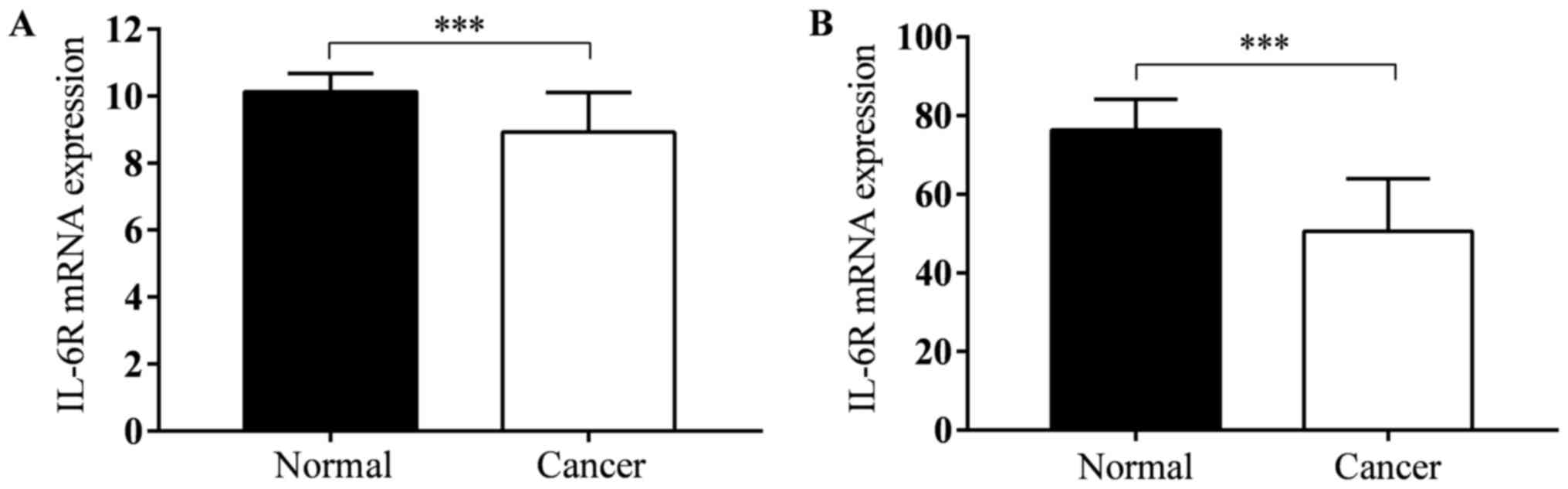
IL-6R mRNA expression in tumor tissues
and adjusted normal tissues
TCGA lung adenocarcinoma and TCGA lung squamous cell
carcinoma datasets contains gene expression data both in tumor
tissues and normal tissues. There were 57 and 51 pairs in LUAD and
LUSC. In both two types of lung cancer, IL-6R mRNA expression level
in tumor tissues was less than normal tissues (P<0.001; Fig. 6).
Biological processes and pathway
analysis
Functional enrichment analysis was performed on
IL-6R and the most related genes (all those genes were shown in
Table II). Table VI lists the top five biological
processes and pathways enriched in IL-6R correlated genes in LUAD.
One of the most significant biological processes is cell division
(GO:0051301, P=4.038E-20). Results also showed that those genes
enriched in mitotic division. GO:0007067, mitotic nuclear division,
GO:0007059, chromosome segregation, GO:0000070, mitotic sister
chromatid segregation and GO:0045143, homologous chromosome
segregation.
 | Table VI.Top 5 biological processes and
pathway of interleukin-6R and associated genes in lung
adenocarcinoma. |
Table VI.
Top 5 biological processes and
pathway of interleukin-6R and associated genes in lung
adenocarcinoma.
| Analysis | ID | Biological
processes | P-value | Genes |
|---|
| Biological
processes | GO:0051301 | Cell division |
4.038×10−20 | CDK1, CDC6, PSRC1,
TPX2, BIRC5, PTTG1, UBE2C, CDC25C, CDC25A, CCNB1, FAM83D, SPC25,
NCAPH, CCNB2, CCND3, ZWINT, CDCA2, CKS2, SKA3, SKA1, CDCA5, ASPM,
CDCA3 |
|
| GO:0007067 | Mitotic nuclear
division |
3.084×10−14 | CENPN, CDK1, NUF2,
PTTG1, CDC25C, CDC25A, FAM83D, SPC25, CCNB2, PLK1, ZWINT, CDCA2,
SKA3, CENPW, SKA1, ASPM |
|
| GO:0007059 | Chromosome
segregation |
1.529×10−09 | SPC25, CENPN,
KIF11, OIP5, NEK2, HJURP, SKA3, CENPW, BIRC5, SKA1 |
|
| GO:0000070 | Mitotic sister
chromatid segregation |
1.186×10−06 | CDCA8, PLK1, NEK2,
SPAG5, KIF18B, ESPL1 |
|
| GO:0045143 | Homologous
chromosome |
5.275×10−06 | PLK1, ESPL1,
PTTG1 |
| Pathway | bta04110 | Cell cycle |
8.584×10−15 | CDK1, CDC6, TTK,
CDC20, ESPL1, CHEK1, PTTG1, CDC25C, CDC25A, CCNB1, CDC45, CCNB2,
MAD2L1, CCND3, PLK1, BUB1, BUB1B |
|
| bta04114 | Oocyte meiosis |
2.573×10−07 | CDK1, MAD2L1,
LADCY9, PLK1, BUB1, FBXO5, ESPL1, CDC20, PTTG1, CDC25C |
|
| bta04914 |
Progesterone-mediated oocyte
maturation |
3.845×10−06 | CCNB1, CDK1,
MAD2L1, CCNB2, LADCY9, PLK1, BUB1, CDC25C, CDC25A |
|
| bta04115 | p53 signaling
pathway | <0.001 | CCNB1, CDK1, CCNB2,
CCND3, RRM2, CHEK1, GTSE1 |
|
| bta05166 | HTLV-I
infection | 0.008 | MAD2L1, CCND3,
POLE2, LADCY9, BUB1B, CDC20, CHEK1, PTTG1 |
The most important pathway in LUAD is cell cycle
(bta04110, P=8.584E-15). As with the results of biological
processes, pathway analysis also shows that these genes are
involved in cell division, eg. oocyte meiosis (bta04114,
P=2.573E-07), progesterone-mediated oocyte maturation (bta04914,
P=3.845E-06). The other two important pathways are p53 signaling
pathway (bta04115, P<0.001) and HTLV-I infection (bta05166,
P=0.008).
Table VII lists the
top five biological processes and pathways enriched in LUSC. The
most significant biological processes are regulation of cell
proliferation (GO:0042127, P=0.001) and positive regulation of
osteoclast differentiation (GO:0045672, P=0.003). Result also
showed that those genes are involved in immune response
(GO:0006955, P=0.003). The other two biological processes are
trans-membrane receptor protein tyrosine kinase signaling pathway
(GO:0007169, P=0.006) and integrin-mediated signaling pathway
(GO:0007229, P=0.006).
 | Table VII.Top 5 biological processes and
pathway of interleukin-6R and associated genes in lung squamous
cell carcinoma. |
Table VII.
Top 5 biological processes and
pathway of interleukin-6R and associated genes in lung squamous
cell carcinoma.
| Analysis | ID | Biological
processes | P-value | Genes |
|---|
| Biological
processes | GO:0042127 | Regulation of cell
proliferation | 0.001 | TNFRSF1A, TNFRSF1B,
FGR, PTK2B, JUNB, BTK |
|
| GO:0045672 | Positive regulation
of osteoclast differentiation | 0.003 | FOS, KLF10,
CCR1 |
|
| GO:0006955 | Immune
response | 0.003 | TNFRSF1A, TNFRSF1B,
CCR1, CD4, CTSH, LCP2 |
|
| GO:0007169 | Trans-membrane
receptor protein tyrosine kinase signaling pathway | 0.006 | DOK2, FGR, LCP2,
BTK |
|
| GO:0007229 | Integrin-mediated
signaling pathway | 0.006 | ITGAL, FGR, PTK2B,
TYROBP |
| Pathway | ptr04650 | Natural killer cell
mediated cytotoxicity | <0.001 | PTPN6, ITGAL,
PTK2B, PRKCB, LCP2, TYROBP |
|
| ptr04380 | Osteoclast
differentiation | 0.002 | FOS, TNFRSF1A,
JUNB, LCP2, BTK, TYROBP |
|
| ptr04611 | Platelet
activation | 0.011 | VWF, LADCY9,
PPP1CC, LCP2, BTK |
|
| ptr04060 | Cytokine-cytokine
receptor interaction | 0.015 | TNFRSF1A, TNFRSF1B,
IL2RB, CCR1, CXCL16, IL6R |
|
| ptr05166 | HTLV-I
infection | 0.028 | FOS, ITGAL,
TNFRSF1A, IL2RB, LADCY9, BUB1B |
The most important pathway in LUSC is natural killer
cell mediated cytotoxicity (ptr04650, P<0.001). And the others
are osteoclast differentiation (ptr04380, P=0.002), platelet
activation (ptr04611, P=0.011), cytokine-cytokine receptor
interaction (ptr04060, P=0.015) and HTLV-I infection (ptr05166,
P=0.028).
Discussion
Lung cancer is a serious threat to public health in
the world. Cytokines play important roles in tumorigenesis as well
as immune surveillance of lung cancer. LUAD and LUSC are two main
pathological subtypes of NSCLC. To study the association of various
cytokines and their receptors with clinical parameters of NSCLC is
an important step for further mechanistic investigations and
provides insight into new therapeutic targets. Our study revealed
that higher expression levels of IL-6R mRNA in tumor tissues were
positively associated with better overall survival in LUAD. These
data suggest an antitumoral role of IL-6R signaling.
Recent works have demonstrated that IL-6 signaling
pathway plays an important role in the immune response (28). Several studies have demonstrated the
pro-tumor effect of IL-6 in NSCLC. Our study revealed a predictive
value of IL-6R mRNA expression in LUAD. It suggested that higher
IL-6R mRNA was associated with better survival. However, the
prognostic value of IL-6R was not shown in LUSC. It is speculated
that LUAD and LUSC arise from distinct cells based on the
histopathological appearance and gene expression signatures. It is
generally accepted that LUAD originates mainly from alveolar
epithelial cells and LUSC is possibly derived from basal cells
(29,30). LUAD and LUSC undergo distinct
developmental processes. Several articles also have showed
different result between LUAD and LUSC. One study showed that an
increased expression of the embryonic stem cells gene set was
associated with overall survival in LUAD. However, there was no
correlation in LUSC (31). Meanwhile,
other study found that the expression levels of PTN1 genes were
associated with survival in LUAD but not LUSC (32). The different results may mainly due to
its different cellular origins, developmental stages and tumor
microenvironment.
In LUAD, enrichment analysis of IL-6R and its most
relevant genes showed that the most significant biological
processes were cell division and mitotic division. That means those
genes mainly involved in cell cycle progression in LUAD. While in
LUSC, the most significant biological processes were regulation of
cell proliferation and several signaling pathway. Pathways analysis
revealed that those genes were involved in natural killer cell
mediated cytotoxicity and platelet activation, meaning that they
were mostly involved in tumor angiogenesis, invasion and
metastasis. Genes that mostly related with IL-6R and the most
significant biological processes were all different in LUAD and
LUSC. That means the tumor microenvironment in both cancers were
discriminate. This prompts us that studies on LUAD and LUSC should
be analyzed separately.
In our report, although there was no statistical
significance between IL-6R mRNA expression and OS in LUSC, the
lower 95% CI limit of HR (0.98) were very close to 1, showing a
trend that IL-6R may be a risk factor for LUSC. Considering the
small sample size in each study, further investigations with a
larger scale of samples are needed to confirm this result.
IL-6R is a part of the receptor for IL-6 which binds
to IL-6 with low affinity, but does not transduce a signal
(33). IL-6ST is necessary for this
signal activation. Correlation analysis showed that IL-6 and IL-6R
were negative correlated in LUAD while they were positive
correlated in LUSC, indicating that the higher the expression of
IL-6R, the lesser the expression of IL-6 in LUAD and the higher the
expression of IL-6 in LUSC. And the pooled correlation coefficient
of IL-6R and IL-6ST were positive both in LUAD and LUSC. However,
Brooks et al have found that IL-6R protein displayed a
positive correlation with IL-6 in LUAD (34). The correlation coefficient of IL-6 and
IL-6R in LUAD and LUSC were −0.199 and 0.288, respectively. Both of
them were less than 0.3, showing a weak correlation. In our report,
the correlation between IL-6 and IL-6R was calculated based on
public datasets, showing a possible phenomenon. The possible
biological process or interaction between these cytokines should be
verified at a tissue, cellular or molecular level by subsequent
experimental verification.
In TCGA dataset, we found that IL-6R mRNA expression
level in tumor tissues was less than normal tissues in both LUAD
and LUSC. Balabko et al have demonstrated the same trend,
and they also found that IL-6R mRNA was found significantly induced
in the tumoural region of LUAD as compared to LUSC (35). STAT3, a transcription factor
downstream of IL-6R, has also been found increased and
phosphorylated in LUAD while there was no phosphorylation in LUSC
(35). Considering the different
effect of IL-6R mRNA in LUAD and LUSC, we deduce that the
expression level of IL-6R mRNA can affect the expression level of
IL-6 and the activation of downstream pathways and can affect the
most important biological processes it involved in.
However, some details need to be further refined.
Firstly, this study included only 8 eligible datasets for LUAD and
6 studies for LUSC, which resulted in relatively insufficiency data
in the subgroup analyses. Secondly, sample size of LUSC in each
study was smaller than LUAD, further articles with a larger scale
of samples are needed to confirm the result. Thirdly, these results
were calculated based on public datasets and these results should
be verified using cells or tumor samples.
In conclusion, our results showed that mRNA levels
of IL-6R in LUAD was associated with better prognosis and can
potentially be used as a prognostic marker for this cancer. While
in LUSC, although there was no statistically significance between
IL-6R mRNA and OS in LUSC, taking the small sample size of each
dataset, the results should be regarded cautiously. Further
prospective studies available of pivotal parameters are needed to
verify the prognosis value of IL-6R in LUAD and LUSC patients.
Acknowledgements
Not applicable.
Funding
This study was supported by funding from the
National Science and Technology Support Program (grant no.
2015BAI12B12), the National Natural Science Foundation of China
(grant nos. 31570877 and 31570908), the Special Funds of Science
and Technology of the People's Livelihood Construction Condition of
Jiangsu Province (grant no. BL2014034), the Science and Technology
Bureau foundation application project of Changzhou (grant no.
CJ20159018), the Key R&D Project of Science and Technology
Department of Jiangsu Province (grant no. BE2015633) and the
Program of Jiangsu Engineering Research Center for Tumor
Immunotherapy (grant no. BM2014404).
Availability of data and materials
The datasets analyzed during the current study are
available in the Cancer Genome Atlas (cancergenome.nih.gov/) and the Gene Expression
Ominibus database (ncbi.nlm.nih.gov/gds/).
Authors' contributions
BL, JJ and YS designed the study and interpreted the
results. LL and CY collected the public datasets and reorganized
and cleaned the data. QC, CY and BX designed the experiments and
helped write the manuscript. QC and BX wrote and organized the
manuscript, with editorial input from YS, JJ and BL. All authors
read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2016. CA Cancer J Clin. 66:7–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Torre LA, Siegel RL and Jemal A: Lung
cancer statistics. Adv Exp Med Biol. 893:1–19. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Johnson DH, Schiller JH and Bunn PA Jr:
Recent clinical advances in lung cancer management. J Clin Oncol.
32:973–982. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Herbst RS, Heymach JV and Lippman SM: Lung
cancer. N Engl J Med. 359:1367–1380. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Travis WD, Brambilla E, Noguchi M,
Nicholson AG, Geisinger KR, Yatabe Y, Beer DG, Powell CA, Riely GJ,
Van Schil PE, et al: International association for the study of
lung cancer/American thoracic society/European respiratory society
international multidisciplinary classification of lung
adenocarcinoma. J Thorac Oncol. 6:244–285. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhan C, Yan L, Wang L, Sun Y, Wang X, Lin
Z, Zhang Y, Shi Y, Jiang W and Wang Q: Identification of
immunohistochemical markers for distinguishing lung adenocarcinoma
from squamous cell carcinoma. J Thorac Dis. 7:1398–1405.
2015.PubMed/NCBI
|
|
7
|
Kishimoto T, Akira S, Narazaki M and Taga
T: Interleukin-6 family of cytokines and gp130. Blood.
86:1243–1254. 1995.PubMed/NCBI
|
|
8
|
Hodge DR, Hurt EM and Farrar WL: The role
of IL-6 and STAT3 in inflammation and cancer. Eur J Cancer.
41:2502–2512. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Haura EB, Livingston S and Coppola D:
Autocrine interleukin-6/interleukin-6 receptor stimulation in
non-small-cell lung cancer. Clin Lung Cancer. 7:273–275. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yanagawa H, Sone S, Takahashi Y, Haku T,
Yano S, Shinohara T and Ogura T: Serum levels of interleukin 6 in
patients with lung cancer. Br J Cancer. 71:1095–1098. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Songür N, Kuru B, Kalkan F, Ozdilekcan C,
Cakmak H and Hizel N: Serum interleukin-6 levels correlate with
malnutrition and survival in patients with advanced non-small cell
lung cancer. Tumori. 90:196–200. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chang CH, Hsiao CF, Yeh YM, Chang GC, Tsai
YH, Chen YM, Huang MS, Chen HL, Li YJ, Yang PC, et al: Circulating
interleukin-6 level is a prognostic marker for survival in advanced
nonsmall cell lung cancer patients treated with chemotherapy. Int J
Cancer. 132:1977–1985. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Seike T, Fujita K, Yamakawa Y, Kido MA,
Takiguchi S, Teramoto N, Iguchi H and Noda M: Interaction between
lung cancer cells and astrocytes via specific inflammatory
cytokines in the microenvironment of brain metastasis. Clin Exp
Metastasis. 28:13–25. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Noda M, Yamakawa Y, Matsunaga N, Naoe S,
Jodoi T, Yamafuji M, Akimoto N, Teramoto N, Fujita K, Ohdo S and
Iguchi H: IL-6 receptor is a possible target against growth of
metastasized lung tumor cells in the brain. Int J Mol Sci.
14:515–526. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yi H, Cho HJ, Cho SM, Jo K, Park JA, Kim
NH, Amidon GL, Kim JS and Shin HC: Blockade of interleukin-6
receptor suppresses the proliferation of H460 lung cancer stem
cells. Int J Oncol. 41:310–316. 2012.PubMed/NCBI
|
|
16
|
Kim NH, Kim SK, Kim DS, Zhang D, Park JA,
Yi H, Kim JS and Shin HC: Anti-proliferative action of
IL-6R-targeted antibody tocilizumab for non-small cell lung cancer
cells. Oncol Lett. 9:2283–2288. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ando K, Takahashi F, Motojima S, Nakashima
K, Kaneko N, Hoshi K and Takahashi K: Possible role for
tocilizumab, an anti-interleukin-6 receptor antibody, in treating
cancer cachexia. J Clin Oncol. 31:e69–e72. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ando K, Takahashi F, Kato M, Kaneko N, Doi
T, Ohe Y, Koizumi F, Nishio K and Takahashi K: Tocilizumab, a
proposed therapy for the cachexia of Interleukin6-expressing lung
cancer. PLoS One. 9:e1024362014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Stang A: Critical evaluation of the
Newcastle-Ottawa scale for the assessment of the quality of
nonrandomized studies in meta-analyses. Eur J Epidemiol.
25:603–605. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Budczies J, Klauschen F, Sinn BV, Győrffy
B, Schmitt WD, Darb-Esfahani S and Denkert C: Cutoff Finder: A
comprehensive and straightforward Web application enabling rapid
biomarker cutoff optimization. PLoS One. 7:e518622012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhu CQ, Ding K, Strumpf D, Weir BA,
Meyerson M, Pennell N, Thomas RK, Naoki K, Ladd-Acosta C, Liu N, et
al: Prognostic and predictive gene signature for adjuvant
chemotherapy in resected non-small-cell lung cancer. J Clin Oncol.
28:4417–4424. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Rousseaux S, Debernardi A, Jacquiau B,
Vitte AL, Vesin A, Nagy-Mignotte H, Moro-Sibilot D, Brichon PY,
Lantuejoul S, Hainaut P, et al: Ectopic activation of germline and
placental genes identifies aggressive metastasis-prone lung
cancers. Sci Transl Med. 5:186ra662013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Botling J, Edlund K, Lohr M, Hellwig B,
Holmberg L, Lambe M, Berglund A, Ekman S, Bergqvist M, Pontén F, et
al: Biomarker discovery in non-small cell lung cancer: Integrating
gene expression profiling, meta-analysis, and tissue microarray
validation. Clin Cancer Res. 19:194–204. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tang H, Xiao G, Behrens C, Schiller J,
Allen J, Chow CW, Suraokar M, Corvalan A, Mao J, White MA, et al: A
12-gene set predicts survival benefits from adjuvant chemotherapy
in non-small cell lung cancer patients. Clin Cancer Res.
19:1577–1586. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Der SD, Sykes J, Pintilie M, Zhu CQ,
Strumpf D, Liu N, Jurisica I, Shepherd FA and Tsao MS: Validation
of a histology-independent prognostic gene signature for
early-stage, non-small-cell lung cancer including stage IA
patients. J Thorac Oncol. 9:59–64. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Director's Challenge Consortium for the
Molecular Classification of Lung Adenocarcinoma, . Shedden K,
Taylor JM, Enkemann SA, Tsao MS, Yeatman TJ, Gerald WL, Eschrich S,
Jurisica I, Giordano TJ, et al: Gene expression-based survival
prediction in lung adenocarcinoma: A multi-site, blinded validation
study. Nat Med. 14:822–827. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
27
|
Beer DG, Kardia SL, Huang CC, Giordano TJ,
Levin AM, Misek DE, Lin L, Chen G, Gharib TG, Thomas DG, et al:
Gene-expression profiles predict survival of patients with lung
adenocarcinoma. Nat Med. 8:816–824. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yu H and Jove R: The STATs of cancer-new
molecular targets come of age. Nat Rev Cancer. 4:97–105. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Han X, Li F, Fang Z, Gao Y, Li F, Fang R,
Yao S, Sun Y, Li L, Zhang W, et al: Transdifferentiation of lung
adenocarcinoma in mice with Lkb1 deficiency to squamous cell
carcinoma. Nat Commun. 5:32612014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Sutherland KD and Berns A: Cell of origin
of lung cancer. Mol Oncol. 4:397–403. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Hassan KA, Chen G, Kalemkerian GP, Wicha
MS and Beer DG: An embryonic stem cell-like signature identifies
poorly differentiated lung adenocarcinoma but not squamous cell
carcinoma. Clin Cancer Res. 15:6386–6390. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li P, Zhang L, Yu X, Tong R, Di X, Mao Y,
Gao Y, Zhang K, Feng L and Cheng S: Proliferation genes in lung
development associated with the prognosis of lung adenocarcinoma
but not squamous cell carcinoma. Cancer Sci. 109:308–316. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Montero-Julian FA: The soluble IL-6
receptors: Serum levels and biological function. Cell Mol Biol
(Noisy-le-grand). 47:583–597. 2001.PubMed/NCBI
|
|
34
|
Brooks GD, McLeod L, Alhayyani S, Miller
A, Russell PA, Ferlin W, Rose-John S, Ruwanpura S and Jenkins BJ:
IL6 trans-signaling promotes KRAS-driven lung carcinogenesis.
Cancer Res. 76:866–876. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Balabko L, Andreev K, Burmann N, Schubert
M, Mathews M, Trufa DI, Reppert S, Rau T, Schicht M, Sirbu H, et
al: Increased expression of the Th17-IL-6R/pSTAT3/BATF/RorγT-axis
in the tumoural region of adenocarcinoma as compared to squamous
cell carcinoma of the lung. Sci Rep. 4:73962014. View Article : Google Scholar : PubMed/NCBI
|