Introduction
Human non-small cell lung cancer (NSCLC) is a
malignant tumor with a high morbidity, and is the most common cause
of cancer-associated mortality worldwide (1). With improvements in living standards and
public health education, the morbidity of NSCLS has been reduced in
recent years, but the morbidity for female patients with NSCLC
remains the second highest among malignant tumors (2). Furthermore, the incidence of NSCLC is
increasing for women. Therapeutic approaches for NSCLC are
improving, but the prognosis and survival rate of NSCLC are
unsatisfactory, which may be due to the lack of early diagnosis
methods and treatment approaches for NSCLC, in addition to the
complications associated with the disease (3). Therefore, it is necessary to explore the
potential molecular mechanisms underlying the occurrence of NSCLC,
in addition to novel approaches for the early diagnosis and
treatment of NSCLC.
In recent years, the molecular diagnosis and
targeted therapy of NSCLC have provided a new approach for the
management of NSCLC (4). The use of
individual genetic expression profiling and clinical staging of
NSCLC in providing prognosis information has gradually become a
focus for NSCLC research (5). The
occurrence of NSCLC is associated with mutations, gene
amplifications and epigenetic changes in tumor-associated genes
(6). Epigenetic changes do not cause
changes to the gene sequence, but can induce changes in the
expression of tumor-associated genes, resulting in tumorigenesis
and progression (6). Epigenetic
research primarily focuses on the methylation of DNA, chromatin
rearrangement and RNA editing changes. Additionally, miRNAs have
become the focus of numerous studies in the field of cancer
(7). It has been demonstrated that
changes in miRNA expression are implicated in the occurrence of
malignancies, at least to an extent (8).
miRNAs are small, non-coding RNAs that arise from
transcriptional pri-miRNA via the activity of specific processing
enzymes, including DROSHA and DICER (9). miRNA pairs with the mRNA of a target
gene, resulting in mRNA degradation, or initiating gene silencing
by preventing translation. Through interactions with a negative
regulatory sequence in the 3′-non-coding region of specific mRNA
targets, miRNA regulation is involved in a wide variety of normal
and abnormal cell behaviors (10).
miRNAs have potential as tumor biomarkers (11). Although the functions of a number of
miRNAs have yet to be elucidated, miRNAs have been implicated in a
wide range of physiological and developmental processes (10).
The present study aimed to explore the role of
microRNA-133a signaling in the regulation of NSCLC development; to
the best of our knowledge, the present study is the first to
investigate this area of NSCLC research.
Materials and methods
Tissue samples
The present study was approved by the Institutional
Medical Ethics Committee of The First Affiliated Hospital of the
General Hospital of the Chinese People's Liberation Army (Beijing,
China), and all participants (n=8; all male; 63±6 years old)
provided informed, written consent. Human NSCLC and adjacent normal
lung tissue samples (n=8) were obtained from The First Affiliated
Hospital. NSCLC and adjacent normal tissues (>5 cm from tumor)
were pathologically validated by a pathologist. None of the
patients had received chemotherapy or radiotherapy prior to
surgery.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was prepared from the tissues or cells
using TRIzol reagent (Bio Scientific; PerkinElmer, Inc., Waltham,
MA, USA), and 1–2 ng total RNA was used for reverse transcription
to cDNA using One-Step RT-PCR Kit (Takara Biotechnology Co., Ltd.,
Dalian, China). RT-qPCR was performed using SYBR® Green
Real-Time PCR Master mix (Takara Biotechnology Co., Ltd., Dalian,
China) by the Stratagene Mx3000P Real-Time PCR system (Agilent
Technologies, Inc., Santa Clara, CA, USA). The specific primer
pairs used are as follows: miR-133a, forward,
5′-TGCTTTGCTAGAGCTGGTAAAATG-3 and reverse,
5-AGCTACAGCTGGTTGAAGGG-3′; U6, forward, 5-CTCGCTTCGGCAGCACA-3 and
reverse, 5-AACGCTTCACGAATTTGCGT-3′. The thermocycling conditions
were as follows: 95°C for 5 min followed by 40 amplification cycles
at 95°C for 15 sec and 60°C for 30 sec. The 2−ΔΔCq
method (12) was used to calculate
the relative change in RNA expression as a ratio.
Immunohistochemistry
Immunohistochemistry was performed to analyze EGFR
expression. Sections (thickness, 4 µm) were prepared from
paraffin-embedded NSCLC and adjacent tissue samples. Sections were
prepared using citrate buffer (pH=6.0, Sangon Biotech Co., Ltd.
Shanghai, China) for 10 min at 95°C and developed with 3%
H2O2 in PBS for 10 min at room temperature.
Sections were blocked with 5% BSA (Sangon Biotech Co., Ltd.) +0.1%
TriX100 in PBS for 1 h at room temperature. Tissues were stained
with anti-EGFR antibodies (cat no. sc-514995; 1:100 dilution; Santa
Cruz Biotechnology, Inc., Dallas, TX, USA) overnight at 4°C.
Tissues were washed with PBS with 0.1% Tween-20 for 5 min 5 times
and stained with horseradish peroxidase conjugated goat anti-rabbit
IgG (1:100, sc-2039; Santa Cruz Biotechnology, Inc.) for 1 h at
room temperature and developed with 3,3′-diaminobenzidine
tetrahydrochloride (Fuzhou Maixin Biotech Co., Ltd.). The slides
were stained with DAB and counterstained with Mayer's hematoxylin
(Beyotime Institute of Biotechnology, Inc., Haimen, China) for 1
h.
Cells lines and reagents
The H358 human NSCLC cell line was purchased from
the the Cell Bank of Type Culture Collection of the Chinese Academy
of Sciences (Shanghai, China) and maintained in RPMI-1640 (Thermo
Fisher Scientific, Inc., Waltham, MA, USA) supplemented with 10%
fetal bovine serum (Gibco; Thermo Fisher Scientific, Inc.) with 100
units/ml streptomycin and 100 units/ml penicillin (Thermo Fisher
Scientific, Inc.) at 37°C in a humidified 5% CO2
atmosphere until 75% confluent.
Transfection and lentiviral
transduction
miRNA-133a mimics (forward,
5′-ACAATGTTTGCTAGAGCTG-3′ and reverse, 5′-GCTGTAGCTATGCATTGA-3′)
and negative control (forward, 5′-CCCCCCCCCCCC-3′ and
5′-CCCCCCCCCCCCCCC-3′) plasmids were transfected into H358 cells
using Lipofectamine 2000 transfection reagent (Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
Subsequent to transfection for 24 or 48 h, transfected cells were
used in further procedures.
MTT assay
The proliferation of transfected H358 cells was
assayed at 24, 48 h using a MTT assay kit (50 µg/ml; Sigma-Aldrich;
Merck KGaA, Darmstadt, Germany) according to the manufacturer's
protocol. Dimethyl sulfoxide (150 µl; Sigma-Aldrich; Merck KGaA)
was used to dissolve the formazan crystals, and the absorbance at
492 nm was measured using a Model 550 microplate reader (Bio-Rad
Laboratories, Inc., Hercules, CA, USA).
Apoptosis assay
H358 cells transfected with miRNA-133a mimics and
negative plasmids was assayed 24, 48 h later using an Annexin
V-fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis
assay. Annexin V-FITC/PI (BD Biosciences, San Jose, CA, USA) was
added and incubated for 10 min at room temperature in the dark. A
flow cytometer (C6; BD Biosciences, San Jose, CA, USA) was used to
examine the rate of apoptosis using FlowJo software (version 7.6.1;
FlowJo LLC, Ashland, OR, USA).
Caspase-3 activity assay
Transfected H358 cells were assayed at 24, 48 h
using a Caspase-3 activity ELISA kit (cat no. C1116; Beyotime
Institute of Biotechnology) for 1 h at 37°C according to the
manufacturer's protocol. Caspase-3 activity was measured using a
Model 550 microplate reader (Bio-Rad Laboratories, Inc.) to measure
absorbance at 405 nm.
Western blot analysis
Transfected H358 cells were assayed at 24, 48 h
using a western blot analysis. The cells were lysed with a RIPA
buffer (Beyotime Institute of Biotechnology) containing a protease
inhibitor cocktail (Sigma-Aldrich; Merck KGaA). The protein
concentration was determinate by the Coomassie blue method. A total
of 50 µg protein per lane was separated with 8–15% SDS-PAGE and
blotted onto a polyvinylidene difluoride membrane (EMD Millipore,
Billerica, MA, USA). Subsequent to blocking with 5% non-fat milk in
Tris-buffered saline with 0.1% Tween-20 for 1 h at 37°C, blots were
incubated with antibodies against caspase-3 (cat no. sc-98785),
EGFR (cat no. sc-514995), phosphorylated (p)-AKT (cat no.
sc-7985-R), phosphorylation-ERK (cat no. sc-23759-R), Bax (cat no.
sc-6236) and GAPDH (cat no. sc-25778) (all dilution, 1:500; Santa
Cruz Biotechnology, Inc.) at 4°C overnight, followed by incubation
with anti-mouse and anti-rabbit horseradish peroxidase-conjugated
secondary antibodies for 1 h (cat nos. sc-2005 and sc-2004;
dilution, 1:5,000; Santa Cruz Biotechnology, Inc.) at 37°C. Blots
were developed with an Amersham ECL Western Blotting Detection kit
(GE Healthcare Life Sciences, Shanghai, China) and analyzed using
Image Lab v.4.62 (Bio-Rad Laboratories, Inc.).
Statistical analysis
All values were expressed as the mean ± standard
deviation using SPSS 17.0 (SPSS, Inc., Chicago, IL, USA).
Comparisons between the means of two groups were performed using
Student's t-test, and with one-way analysis of variance followed by
Tukey's post hoc test for multiple groups. P<0.05 was considered
to indicate a statistically significant difference.
Results
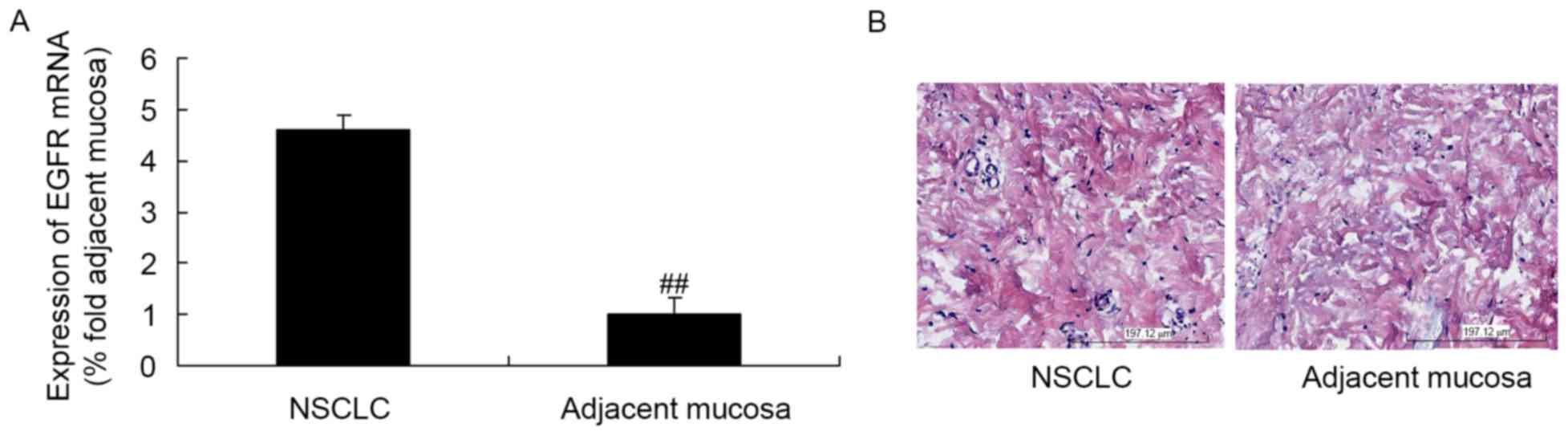
EGFR expression in NSCLC tissues and
adjacent mucosae
As presented in Fig.
1A, the expression of EGFR mRNA in NSCLC tissues was higher
than that in the adjacent mucosal tissues. Additionally, EGFR
protein expression in the NSCLC tissues was higher than that in the
adjacent mucosae (Fig. 1B).
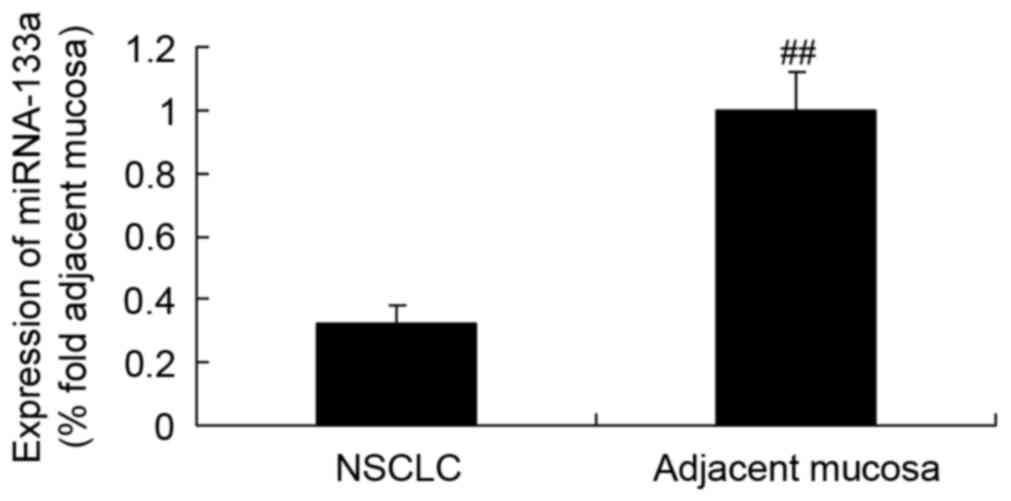
miRNA-133a expression in NSCLC tissues
and adjacent mucosae
The expression of miRNA-133a in NSCLC tissue was
initially detected by RT-qPCR, which revealed that it was markedly
lower than that in the adjacent mucosal tissue (Fig. 2).
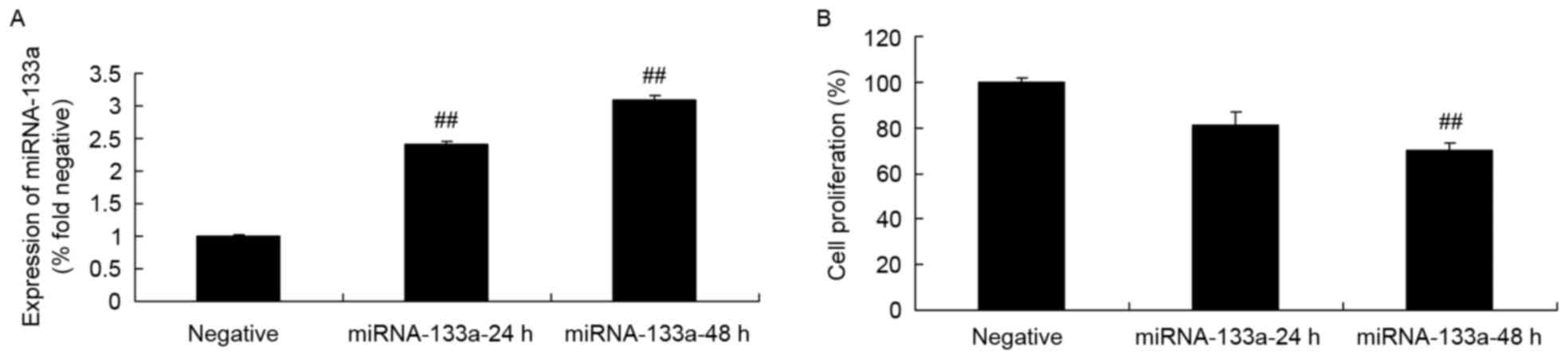
NSCLC cell growth rate following
miRNA-133a upregulation
To test the hypothesis that miRNA-133a may be
involved in the alternative regulation of NSCLC growth, H358 cells
were transfected with miRNA-133a mimics, and the expression of
miRNA-133a was assessed via RT-qPCR. The miRNA-133a mimics
significantly increased miRNA-133a expression in H358 cells,
compared with in the control group (Fig.
3A). Furthermore, upregulated miRNA-133a significantly
suppressed H358 cell growth, compared with that of the control
group (Fig. 3B).
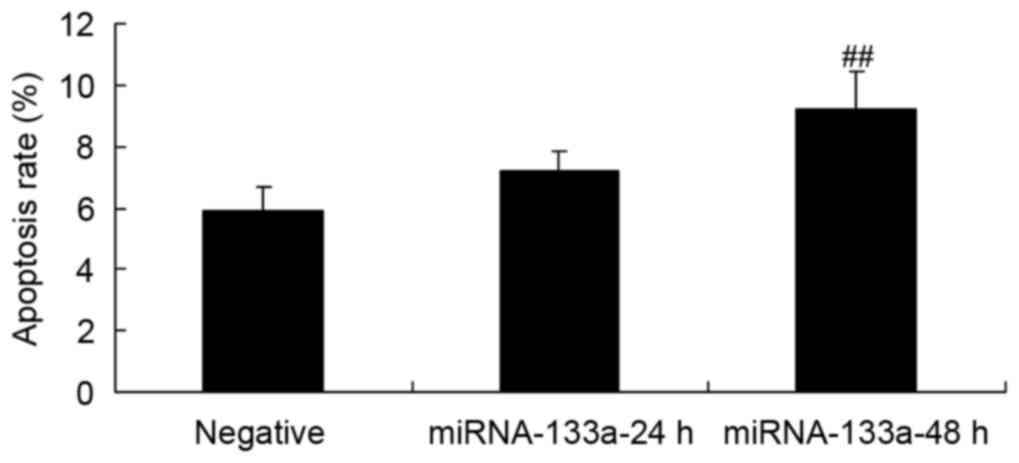
NSCLC apoptosis following miRNA-133a
upregulation
NSCLC apoptosis was evaluated using an Annexin
V-FITC/PI apoptosis assay, in order to assess the effect of
upregulated miRNA-133a. The results showed that increased
miRNA-133a significantly induced H358 cell apoptosis, as compared
with the control group (Fig. 4).
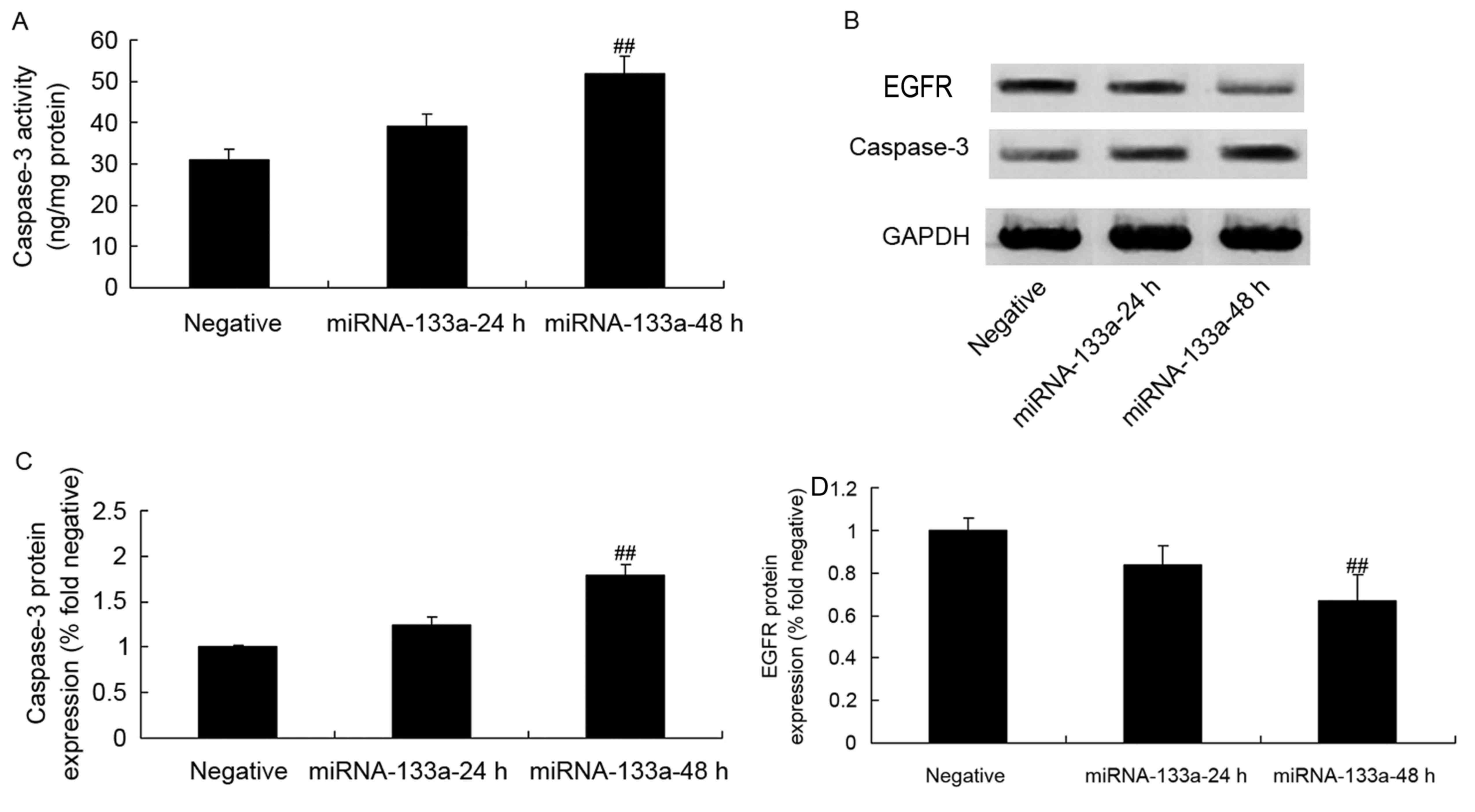
Caspase-3 activity, protein expression
and EGFR protein expression in NSCLC following miRNA-133a
upregulation
To determine the levels of caspase-3 activity,
protein expression and EGFR protein expression in NSCLC following
the upregulation of miRNA-133a, a caspase-3 ELISA and western blot
analysis were conducted. As depicted in Fig. 5, there were significant increases in
caspase-3 activity, protein expression and EGFR protein expression
in H358 cells following miRNA-133a elevation, as compared with
those in the control group.
p-AKT, p-ERK and Bax protein
expression in NSCLC following miRNA-133a upregulation
To investigate the association between p-AKT, p-ERK
and Bax and miRNA-133a upregulation in H358 cells, p-AKT, p-ERK and
Bax protein expressions was analyzed via western blotting. As
presented in Fig. 6, miRNA-133a
upregulation significantly suppressed p-AKT, p-ERK and Bax protein
expression in H358 cells, as compared with in the control
group.
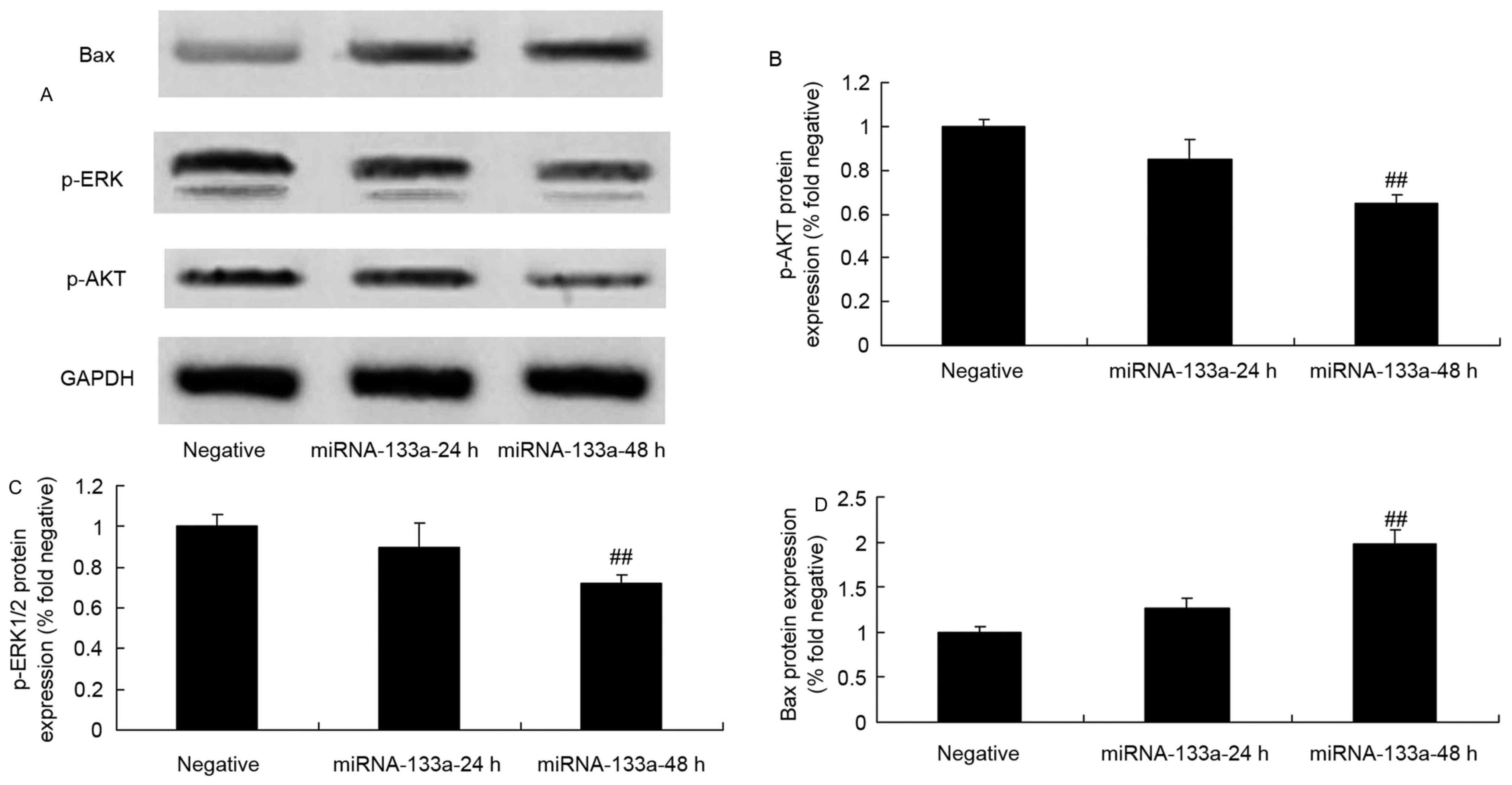 | Figure 6.p-AKT, p-ERK and Bax protein
expression in NSCLC cells following the upregulation of miRNA-133a.
p-AKT, p-ERK and Bax expression in NSCLC tissues was evaluated via
(A) western blot that was quantified for (B) p-AKT, (C) p-ERK and
(D) Bax expression, following the upregulation of miRNA-133a.
##P<0.01, compared with the NSCLC group. NSCLC,
non-small cell lung cancer; miRNA, microRNA; p-, phosphorylated;
ERK, phosphorylated extracellular signal-regulated kinase; Bax,
Bcl-2-associated X; AKT, protein kinase B. |
Discussion
NSCLC is the most common cause of cancer-associated
mortality. Although methods for the diagnosis and treatment of
NSCLC have improved at a constant rate, the prognosis for patients
with NSCLC remains poor (13).
Clinical staging of NSCLC via pathological analysis is the
recommended method for evaluating NSCLC prognosis and selecting the
optimal treatment approach (14).
With an increasing number of in-depth studies of the effects of
miRNA in tumors and other diseases, miRNA expression changes in
NSCLC are gradually becoming a hotspot in this area of research.
When classifying the subtype of a tumor, the miRNA expression
profile of the tumor tissue can be more accurate than using mRNA
(15). Studies have indicated that
miRNAs are abnormally expressed in NSCLC, but there is no consensus
on the miRNAs involved (8,15). A wide range of different alterations
to miRNA expression have been identified (15). Potentially, miRNA analysis can be
integrated into staging methods for the diagnosis and prognosis of
NSCLC, and potentially allow individualized NSCLC treatment
(16). In the present study, the
expression of miRNA-133a in NSCLC tissue was markedly lower than
that of the adjacent mucosa tissue. Furthermore, upregulating
miRNA-133a significantly suppressed cell growth, induced apoptosis,
and increase caspase-3 activity and protein expression in H358
cells. These data support that miRNA-133 is a tumor suppressor in
lung cancer.
A total of 90% the identified EGFR gene mutations in
NSCLC are located in exons 19–21 (17). EGFR can be expressed in epithelial,
mesenchymal and neurogenic tissues and it serves a important role
in regulating hyperplasia, growth and the differentiation of normal
cells (17). EGFR also induces the
growth of tumor cells, vascularization, tumor metastasis and
apoptosis resistance (18). Once a
ligand interacts with the EGFR-N extracellular domain, an EGFR
homo- or heterodimer can be formed, resulting in the
phosphorylation of intracellular tyrosine residues and the
activation of downstream signal pathways, including the
RAS/RAF/ERK/MAPK pathway, the P13K/AKT pathway and the STAT3/5
signal transduction pathways. The result is the expression of genes
that promote tumorigenic cell behavior, including proliferation,
invasion, metastasis, angiogenesis and dysplasia (18). The dimer formed upon the EGFR-N
extracellular binding domain and ligand binding is the requirement
for the phosphorylation of EGFR tyrosine residues and the
activation of downstream signaling (19). In the present study, it was identified
that upregulating miRNA-133a significantly suppressed the EGFR
expression of H358 cells. Cui et al (20) identified that microRNA-133a regulated
the proliferation of breast cancer through the EGFR/Akt signaling
pathway. Song et al (21)
demonstrated that miR-133a suppressed cervical cancer growth
through the Akt and ERK signaling pathways. The data presented in
the present study suggest that EGFR serves an important role in
miRNA-133-induced lung cancer suppression.
Tumor cells commonly exhibit altered signal
transduction and imbalanced cell growth, differentiation and
apoptosis (22). PI3K/AKT signal
transduction is an important intracellular signal transduction
pathway that can induce the occurrence and development of numerous
types of tumor by affecting cell cycle control, cell survival,
metastasis, angiogenesis and chemotherapy resistance (23). Activation often occurs in the early
stages of oncogenesis and tumor progression. The degree of signal
pathway activation is an important indicator of the prognosis of
patients with tumors (24). In the
present study, the upregulation of miRNA-133a significantly
suppressed the p-AKT protein expression in H358 cells. Cui et
al (20) demonstrated that
microRNA-133a regulated the proliferation of breast cancer cells
through the EGFR/Akt signaling pathway. This is in accord with our
novel finding that the regulation of Akt serves a role in the
effect of miRNA-133a on lung cancer.
The ERK pathway is an important signal pathway in
the occurrence and development of NSCLC, and is a vital target of
anticancer treatment (25).
Transcription factor AP-2β overexpression has been associated with
the incidence of lung adenocarcinoma, which is associated with the
regulation of the ERK pathway. Mitogen-inducible gene 6 expression
downregulation inhibits NSCLC cell apoptosis, activates the ERK
pathway and upregulates the anti-apoptotic protein Bcl-2 (26). ERK activation mediated by Src can
promote the gefitinib resistance of NSCLC cells (26). Inhibition of the ERK pathway can
prevent NSCLC cells from undergoing epithelial-mesenchymal
transition and promote their sensitivity to EGFR inhibitors
(27). AKT and ERK pathways also
interact to promote NSCLC cell proliferation and survival.
Activating the AKT and ERK pathways simultaneously induces drug
resistance in NSCLC cells (28).
Mucin 1 mucoprotein may induce the ERK2 pathway to regulate cell
growth and differentiation (28). In
the present study, the upregulation of miRNA-133a significantly
inhibited p-ERK protein expression and induced the expression of
Bax protein in H358 cells. Song et al (21) demonstrated that miR-133a suppresses
cervical cancer growth through an effect on the AKT and ERK
signaling pathways.
In conclusion, the present study has demonstrated
that the upregulation of miRNA-133a significantly suppressed cell
growth, induced apoptosis, and increased caspase-3 activity and
protein expression via the EGFR/AKT/ERK signaling pathway in H358
NSCLC cells. Thus, targeting miRNA-133a may exhibit potential as a
novel therapeutic strategy against NSCLC through the EGFR/AKT/ERK
signaling pathway.
Acknowledgements
Not applicable.
Funding
Not applicable.
Availability of data and materials
The analyzed data sets generated during the study
are available from the corresponding author, on reasonable
request.
Authors' contributions
WZ designed the experiment. NNG, YNZ, SJL, SSL and
JQY performed the experiment. WZ and JQY analyzed the data. WZ
wrote the manuscript.
Ethics approval and consent to
participate
The present study was approved by the Institutional
Medical Ethics Committee of The First Affiliated Hospital of the
General Hospital of the Chinese People's Liberation Army, and all
participants provided informed, written consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Videtic GM, Hu C, Singh AK, Chang JY,
Parker W, Olivier KR, Schild SE, Komaki R, Urbanic JJ, Timmerman RD
and Choy H: A randomized phase 2 study comparing 2 stereotactic
body radiation therapy schedules for medically inoperable patients
with stage i peripheral non-small cell lung cancer: NRG oncology
RTOG 0915 (NCCTG N0927). Int J Radiat Oncol Biol Phys. 93:757–764.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Nie X, Cheng G, Ai B and Zhang S: The
tailored chemotherapy based on RRM1 immunohistochemical expression
in patients with advanced non-small cell lung cancer. Cancer
Biomark. 13:433–440. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Karp DD, Lee SJ, Keller SM, Wright GS,
Aisner S, Belinsky SA, Johnson DH, Johnston MR, Goodman G, Clamon
G, et al: Randomized, double-blind, placebo-controlled, phase III
chemoprevention trial of selenium supplementation in patients with
resected stage I non-small-cell lung cancer: ECOG 5597. J Clin
Oncol. 31:4179–4187. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wang R, Chen XF and Shu YQ: Prediction of
non-small cell lung cancer metastasis-associated microRNAs using
bioinformatics. Am J Cancer Res. 5:32–51. 2014.PubMed/NCBI
|
|
5
|
Sin TK, Wang F, Meng F, Wong SC, Cho WC,
Siu PM, Chan LW and Yung BY: Implications of MicroRNAs in the
treatment of gefitinib-resistant non-small cell lung cancer. Int J
Mol Sci. 17:2372016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chen F, Hou SK, Fan HJ and Liu YF:
MiR-15a-16 represses Cripto and inhibits NSCLC cell progression.
Mol Cell Biochem. 391:11–19. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wu N, Zhang C, Bai C, Han YP and Li Q:
MiR-4782-3p inhibited non-small cell lung cancer growth via USP14.
Cell Physiol Biochem. 33:457–467. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lin R, Chen L, Chen G, Hu C, Jiang S,
Sevilla J, Wan Y, Sampson JH, Zhu B and Li QJ: Targeting miR-23a in
CD8+ cytotoxic T lymphocytes prevents tumor-dependent
immunosuppression. J Clin Invest. 124:5352–5367. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li N, Zhang F, Li S and Zhou S: Epigenetic
silencing of MicroRNA-503 regulates FANCA expression in non-small
cell lung cancer cell. Biochem Biophys Res Commun. 444:611–616.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang X, Ling C, Bai Y and Zhao J:
MicroRNA-206 is associated with invasion and metastasis of lung
cancer. Anat Rec (Hoboken). 294:88–92. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Aghanoori MR, Mirzaei B and Tavallaei M:
MiRNA molecular profiles in human medical conditions: Connecting
lung cancer and lung development phenomena. Asian Pac J Cancer
Prev. 15:9557–9565. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Dy GK, Hillman SL, Rowland KM Jr, Molina
JR, Steen PD, Wender DB, Nair S, Mandrekar S, Schild SE and Adjei
AA: North Central Cancer Treatment Group Study N0326: A front-line
window of opportunity phase 2 study of sorafenib in patients with
advanced nonsmall cell lung cancer: North central cancer treatment
group study N0326. Cancer. 116:5686–5693. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Mita AC, Papadopoulos K, de Jonge MJ,
Schwartz G, Verweij J, Mita MM, Ricart A, Chu QS, Tolcher AW, Wood
L, et al: Erlotinib ‘dosing-to-rash’: A phase II intrapatient dose
escalation and pharmacologic study of erlotinib in previously
treated advanced non-small cell lung cancer. Br J Cancer.
105:938–944. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Roa WH, Kim JO, Razzak R, Du H, Guo L,
Singh R, Gazala S, Ghosh S, Wong E, Joy AA, et al: Sputum microRNA
profiling: A novel approach for the early detection of non-small
cell lung cancer. Clin Invest Med. 35:E2712012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhang W, Winder T, Ning Y, Pohl A, Yang D,
Kahn M, Lurje G, Labonte MJ, Wilson PM, Gordon MA, et al: A let-7
microRNA-binding site polymorphism in 3′-untranslated region of
KRAS gene predicts response in wild-type KRAS patients with
metastatic colorectal cancer treated with cetuximab monotherapy.
Ann Oncol. 22:104–109. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Karachaliou N, Rosell R, Morales-Espinosa
D and Viteri S: Systemic treatment in EGFR-ALK NSCLC patients:
Second line therapy and beyond. Expert Rev Anticancer Ther.
14:807–815. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Janmaat ML, Rodriguez JA, Gallegos-Ruiz M,
Kruyt FA and Giaccone G: Enhanced cytotoxicity induced by gefitinib
and specific inhibitors of the Ras or phosphatidyl inositol-3
kinase pathways in non-small cell lung cancer cells. Int J Cancer.
118:209–214. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Raben D, Helfrich B and Bunn PA Jr:
Targeted therapies for non-small-cell lung cancer: Biology,
rationale, and preclinical results from a radiation oncology
perspective. Int J Radiat Oncol Biol Phys. 59 2 Suppl:S27–S38.
2004. View Article : Google Scholar
|
|
20
|
Cui W, Zhang S, Shan C, Zhou L and Zhou Z:
microRNA-133a regulates the cell cycle and proliferation of breast
cancer cells by targeting epidermal growth factor receptor through
the EGFR/Akt signaling pathway. FEBS J. 280:3962–3974. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Song X, Shi B, Huang K and Zhang W:
miR-133a inhibits cervical cancer growth by targeting EGFR. Oncol
Rep. 34:1573–1580. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhen Y, Li D, Li W, Yao W, Wu A, Huang J,
Gu H, Huang Y, Wang Y, Wu J, et al: Reduced PDCD4 expression
promotes cell growth through PI3K/Akt signaling in non-small cell
lung cancer. Oncol Res. 23:61–68. 2016. View Article : Google Scholar
|
|
23
|
Cheng H, Shcherba M, Pendurti G, Liang Y,
Piperdi B and Perez-Soler R: Targeting the PI3K/AKT/mTOR pathway:
Potential for lung cancer treatment. Lung Cancer Manag. 3:67–75.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Dong M, Yang G, Liu H, Liu X, Lin S, Sun D
and Wang Y: Aged black garlic extract inhibits HT29 colon cancer
cell growth via the PI3K/Akt signaling pathway. Biomed Rep.
2:250–254. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang H, Wu C, Wan S, Zhang H, Zhou S and
Liu G: Shikonin attenuates lung cancer cell adhesion to
extracellular matrix and metastasis by inhibiting integrin β1
expression and the ERK1/2 signaling pathway. Toxicology.
308:104–112. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zinn RL, Gardner EE, Marchionni L, Murphy
SC, Dobromilskaya I, Hann CL and Rudin CM: ERK phosphorylation is
predictive of resistance to IGF-1R inhibition in small cell lung
cancer. Mol Cancer Ther. 12:1131–1139. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhao S, Wu J, Zheng F, Tang Q, Yang L, Li
L, Wu W and Hann SS: β-elemene inhibited expression of DNA
methyltransferase 1 through activation of ERK1/2 and AMPKα
signalling pathways in human lung cancer cells: the role of Sp1. J
Cell Mol Med. 19:630–641. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gao J, Zhao Y, Lv Y, Chen Y, Wei B, Tian
J, Yang Z, Kong F, Pang J, Liu J and Shi H: Mirk/Dyrk1B mediates
G0/G1 to S phase cell cycle progression and cell survival involving
MAPK/ERK signaling in human cancer cells. Cancer Cell Int.
13:22013. View Article : Google Scholar : PubMed/NCBI
|