Introduction
Gliomas are neuroepithelial tumors and malignant
gliomas are the most common type of primary tumors of the central
nervous system (1). The prognosis is
worse and the degree of malignancy is increased in tumors with a
higher grade (1). Usually, the
treatment for a glioma is surgery, complemented with radiotherapy
and chemotherapy (2). Although, in
recent years, the microsurgery technique, chemotherapy,
immunotherapy, gene therapy, molecular targeted therapy and
biological technology have progressed, the overall prognosis of
patients with glioma has not improved (2). The mortality and recurrence rates are
high, with the 5-year survival rate for the primary type of glioma,
glioblastoma, being 5.1% and the one-year recurrence rate being
~70% (3). The main reasons for the
high mortality of patients with glioma are resistance to treatment
and local recurrence, which have been difficult problems to
overcome in the treatment of gliomas (2).
Previous studies have revealed that cancer stem
cells (CSCs) or CSC-like cells exist in a variety of tumors, and
indicated that they serve an important role in the occurrence,
development, invasion, metastasis and recurrence of tumors
(4,5).
The cluster of differentiation (CD)133 protein has been reported as
a cell surface marker to human embryonic neural stem cells and
cancer stem cells (6–9), and CD133 positive cells present stem
cell properties when isolated from human brain tumors (6,7,10). In gliomas, glioma stem cells (GSCs)
have also been reported to serve a role in the progression of
glioma (11), which provides a novel
theory, and may provide a breakthrough point for studying the
mechanism of glioma treatment and improving the therapeutic
effect.
The proliferation of normal stem cells is strictly
regulated by the microenvironment, where cancer stem cells are also
likely to specifically grow and exist (12). Internal tumors are often in hypoxic
states during the growth process (13). Studies have demonstrated that a
hypoxic microenvironment serves an important role in the occurrence
and development of a tumor, changing the phenotype of tumor cells
and increasing the malignant degree through hypoxic stress
(14). The presence of hypoxia can
promote tumor cell proliferation, promote angiogenesis, inhibit
stem cell differentiation, increase resistance to radiotherapy and
chemotherapy, and increase tumor invasion and metastasis (11,15–20).
Cancer stem cells express several stem cell markers (21). Pluripotent stem cells are regulated by
a series of transcription factors, including Oct3/4 and
transcription factor SOX-2 (Sox2) (22,23).
Studies demonstrated that changes in neurogenic locus notch homolog
protein (Notch)1 signaling are associated with a number of human
cancers and Notch1 has been speculated to be a prognostic marker
for tumors (24–28). The present study established a hypoxic
model in human U87 glioma cells, observed the effect of hypoxia on
the expression of Oct3/4, Sox2 and Notch1 at mRNA and protein
levels, and assessed the influence of the hypoxic environment on
human U87 glioma cell physiology.
Materials and methods
Cell culture
The human glioma cell line, U87, was purchased from
the Cell Bank of Shanghai Institute of Cell Biology, Chinese
Academy of Sciences (Shanghai, China) and cultured in Dulbecco's
modified Eagle's medium (DMEM) with 10% fetal bovine serum (both
Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA), 100 U/ml
penicillin and 100 U/ml streptomycin at 37°C in the incubator with
5% CO2 and 95% humidity. The medium was changed every 72
h. Previously, this widely used cell line was reported to be
different from the original U87MG Uppsala cell line, but it is
likely that the cells line is a bona fide human glioblastoma cell
line of unknown origin (29). The
authors of the current study posit that the possible difference
between these two cell lines should not affect the conclusions of
the present study.
Hypoxic cell culture
U87 cells were cultured in DMEM containing fetal
bovine serum, penicillin and streptomycin at 37°C in the closed
incubation system (Xvivo system 300C; BioSpherix, Ltd., Parish, NY,
USA) with 1% O2, 94% N2, 5% CO2
and 95% humidity. Cells were assessed following 2, 6, 12, 24, 48
and 72 h of culturing.
Cell viability
A total of 100 ml U87 cells at a density of
1.5×105 cells/ml were plated into each well of a 96-well
plate and cultured under hypoxia with blank controls set up in the
middle and at the edge of the 96-well plate. Cells were incubated
for 2, 6, 12, 24, 48, and 72 h, with five replicates for each time
point. Following incubation, the medium was replaced with fresh
DMEM containing 20 ml MTT (5 mg/ml; Sigma-Aldrich; Merck KGaA,
Darmstadt, Germany) and incubated further for 4 h. Subsequently,
150 ul/well DMSO was added into the 96-well plate, followed by
agitating the plate for 10 min. Absorbance was measured by a
microplate reader at 490 nm. The growth curves of U87 cells under
low oxygen or normal conditions (37°C with 5% CO2) were
drawn using the duration of the cells cultures and the mean value
of absorbance of cells.
Apoptosis assay
A total of 2×106 U87 cells were suspended
with DMEM containing 10% fetal bovine serum (both from Gibco), 100
U/ml penicillin and 100 U/ml streptomycin, and inoculated in the
culture flask and cultured in the hypoxic environment or at normal
conditions at 37°C for 48 h. Following culturing, the single cell
suspension was fixed with 70% ethanol at 4°C overnight. Then cells
were collected by centrifugation at 300 × g for 5 min, washed three
times with PBS, resuspended with Annexin V-fluorescein
isothiocyanate (FITC) and propidium iodide (PI) staining solutions
using the FITC Annexin V Apoptosis Detection kit (BD Biosciences,
Franklin Lakes, NJ, USA), according to the manufacturer's
protocols. The cells were subsequently mixed gently and incubated
for 15 min at room temperature in the dark. Stained cells were
analyzed using the BD FACSCalibur™ analyzer with FlowJo
7.6 software (BD Biosciences), which used FL1 and FL2 channels to
produce scatter charts with PI and FITC as the parameters. The
experiment was repeated three times.
Soft agar colony formation assay
Agar (BD Biosciences) plates were constructed with
0.5% agar forming the bottom layer and 0.3% agar forming the top
layer. The plate contained a 1×104 cells/well single
cell suspension and was cultured in hypoxic conditions or normal
conditions for 2–3 weeks for colony formation. The colonies were
subsequently stained by adding 0.2 ml/well of 1 mg/ml nitro blue
tetrazolium chloride (ThermoFisher Scientific, Inc.) dissolved in
PBS and incubated overnight at 37°C. Using a light microscope
(magnification, 100×), colonies with a diameter >75 µm or
colonies that contained >50 cells were counted. The rate of
colony formation was calculated with the following formula: Colony
formation rate (%)=number of colonies/number of inoculated cells
×100.
Flow cytometry analysis and sorting of
CD133+ cells
Cells in the logarithmic growth phase were cultured
under hypoxic conditions or normal conditions for 24 h at 37°C.
They were subsequently washed three times with PBS, collected by
centrifugation at 300 × g for 5 min at room temperature, and
resuspended and dissociated into single cell suspension in the
Pre-Sort Buffer (BD FACS™) by repeatedly pipetting. The
cells were then centrifuged at 300 × g for 5 min at room
temperature and the supernatant was discarded. The cells were
washed with 8 ml PBS, blocked with 2 ml FcR Blocking Reagent
(Miltenyi Biotec GmbH, Bergisch Gladbach, Germany) for 10 min at
4°C and incubated with 2 ml CD133/2 (293C3)-PE antibodies
(dilution, 1:1,000; cat. no. 130-090-853; Miltenyi Biotec GmbH) for
10 min at 4°C in the dark. The cells were washed once with 2 ml PBS
and then the proportion of CD133+ cells were detected using the BD
FACSCalibur™ analyzer with FlowJo 7.6 software.
Reverse transcription-quantitative
polymerase chain reaction analysis
Cells in the logarithmic growth phase at an
appropriate density were seeded in culture flasks and cultured for
24 h, then cultured in normoxic or hypoxic environments. Total RNA
was extracted using the TRIzol reagent kit (Invitrogen; Thermo
Fisher Scientific Inc.) and reverse transcribed into cDNA using the
PrimeScript RT reagent kit (Takara Biotechnology Co., Ltd., Dalian,
China) according to the manufacturer's protocol. cDNA was used as a
template for Oct3/4, Sox2, Notch1 and GAPDH gene amplification;
GAPDH was used as the reference gene. Primer sequences for genes
and reference gene are provided in Table
I. The qPCR was run with SYBR Green (Takara Biotechnology Co.,
Ltd.) on 7500 fast Real-Time PCR system (Applied Biosystems; Thermo
Fisher Scientific Inc) and the thermocycling conditions were as
follows: Denaturation at 95°C for 5 min, and then 35 cycles of
denaturation at 94°C for 20 sec, annealing at 60°C for 20 sec and
extension at 72°C for 40 sec. The results were quantified with the
2−ΔΔCq method (30).
 | Table I.The nucleotide sequence of primers
for reverse transcription-quantitative polymerase chain
reaction. |
Table I.
The nucleotide sequence of primers
for reverse transcription-quantitative polymerase chain
reaction.
| Gene | Primer sequence
(5′-3′) | Product length
(bp) |
|---|
| Oct3/4 | F:
ACATGTGTAAGCTGCGGCC | 297 |
|
| R:
GTTGTGCATAGTCGCTGCTTG |
|
| Transcription
factor SOX-2 | F:
TTGCTGCCTCTTTAAGACTAGGA | 75 |
|
| R:
CTGGGGCTCAAACTTCTCTC |
|
| Neurogenic locus
notch homolog protein 1 | F:
CACTGTGGGCGGGTCC | 85 |
|
| R:
GTTGTATTGGTTCGGCACCAT |
|
| GAPDH | F:
CCTCAAGATCATCAGCAATGC | 101 |
|
| R:
TGGTCATGAGTCCTTCCACG |
|
Western blot analysis
Cells in the logarithmic growth phase at an
appropriate density were seeded in culture flasks and cultured for
24 h, then cultured in normoxic or hypoxic environments. Total
protein was extracted with lysis buffer (pH 7.5) containing the
following: 50 mM Tris-HCl; 100 mM NaCl; 1% TritonX-100; 1 mM EDTA;
2 mM sodium vanadate, and protease inhibitor. Protein concentration
was determined using the bicinchoninic acid method. Protein (40
ug/lane) was loaded onto 12% SDS-PAGE gel to separate the proteins.
The semi-dry method was used to transfer protein onto a
polyvinylidene difluoride membrane (PVDF; 0.45 µm; EMD Millipore,
Billerica, MA, USA). The PVDF membrane was incubated with 5% skim
milk in TBST for 90 min at room temperature, followed by primary
antibodies against Oct3/4 (cat. no. sc-9081), Sox2 (cat. no.
sc-20088), Notch1 (cat. no. sc-9170) and GAPDH (cat. no. sc-25778)
at a dilution of 1:1,000 (all from Santa Cruz Biotechnology Inc) at
4°C overnight. The membrane was then washed with TBST three times
and incubated with horseradish peroxidase-conjugated goat
anti-rabbit immunoglobulin G secondary antibodies (dilution,
1:1,000; cat. no. sc-2004; Santa Cruz Biotechnology, Inc.) at 37°C
for 2 h. Finally, the membrane was washed with TBST three times,
developed with Clarity™ Western Electrochemiluminescence Substrate
(Bio-Rad Laboratories, Inc., Hercules, CA, USA) and detected by the
ChemiDoc™ XRS + System with Image Lab™ Software (170–8265; Bio-Rad
Laboratories, Inc.). GAPDH was used as the reference protein. The
experiment was repeated three times.
Statistical analysis
The data are presented as mean ± standard deviation
and analyzed using SPSS 18.0 software (SPSS, Inc., Chicago, IL,
USA). A two-sample t-test or a one-way analysis of variance with
the Tukey's Honest Significant Difference post-hoc test were used
for statistical analysis. P<0.05 indicated that the difference
between groups was statistically significant.
Results
Hypoxia increases U87 cell
viability
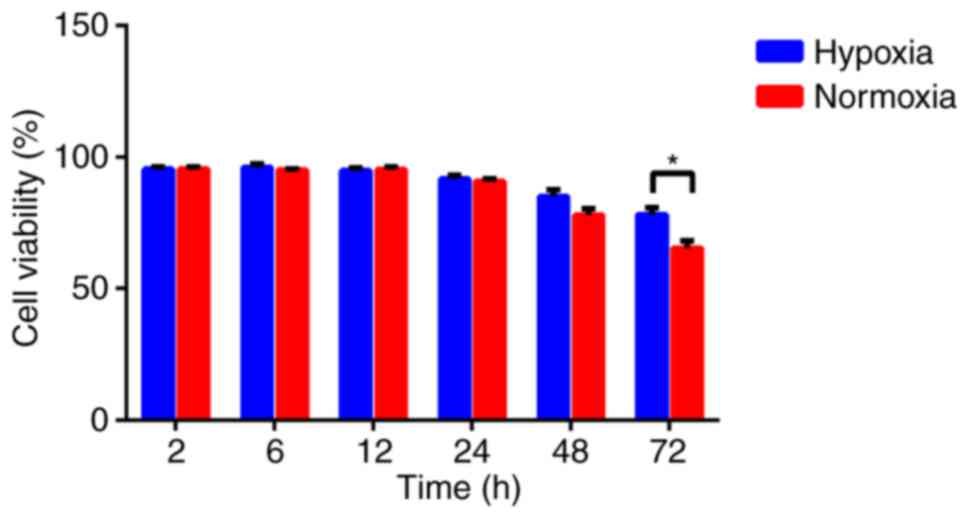
To assess the effect of hypoxia on U87 cell
viability, cells were cultured under hypoxic and normal conditions,
and their viability were analyzed by the MTT assay. Hypoxic U87
cells exhibited a higher viability rate compared with the cells
cultured under normal conditions when exposed to hypoxia for 48 and
72 h. Although the increased viability of hypoxic cells was slight,
the difference between the two groups at 72 h was statistically
significant (P<0.05; Fig. 1).
Hypoxia decreases U87 cell
apoptosis
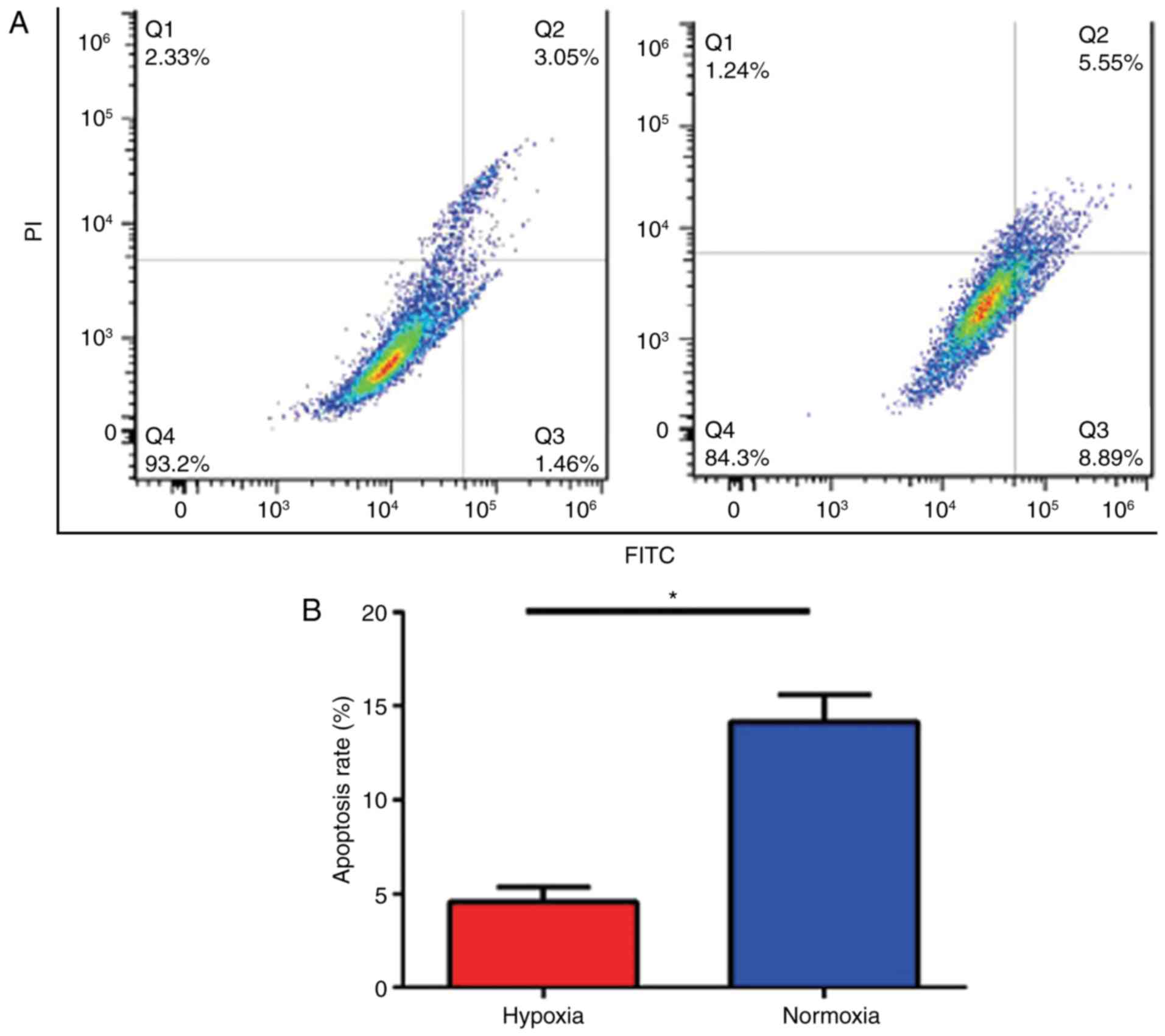
To assess if hypoxia induced apoptosis in U87 cells,
cultured cells were analyzed using flow cytometry. The data
demonstrated that the apoptosis rate in U87 cells cultured under
hypoxic conditions was decreased compared with that in cells in
normal conditions (Fig. 2A). The
greatest difference in the apoptosis rate in U87 cells cultured
under hypoxic or normoxic conditions was 48 h post treatment
(4.55±0.46 and 14.15±0.84%, respectively); the difference in the
apoptosis rate was statistically significant (P<0.05; Fig. 2B).
Hypoxia increases the colony formation
ability of U87 cells
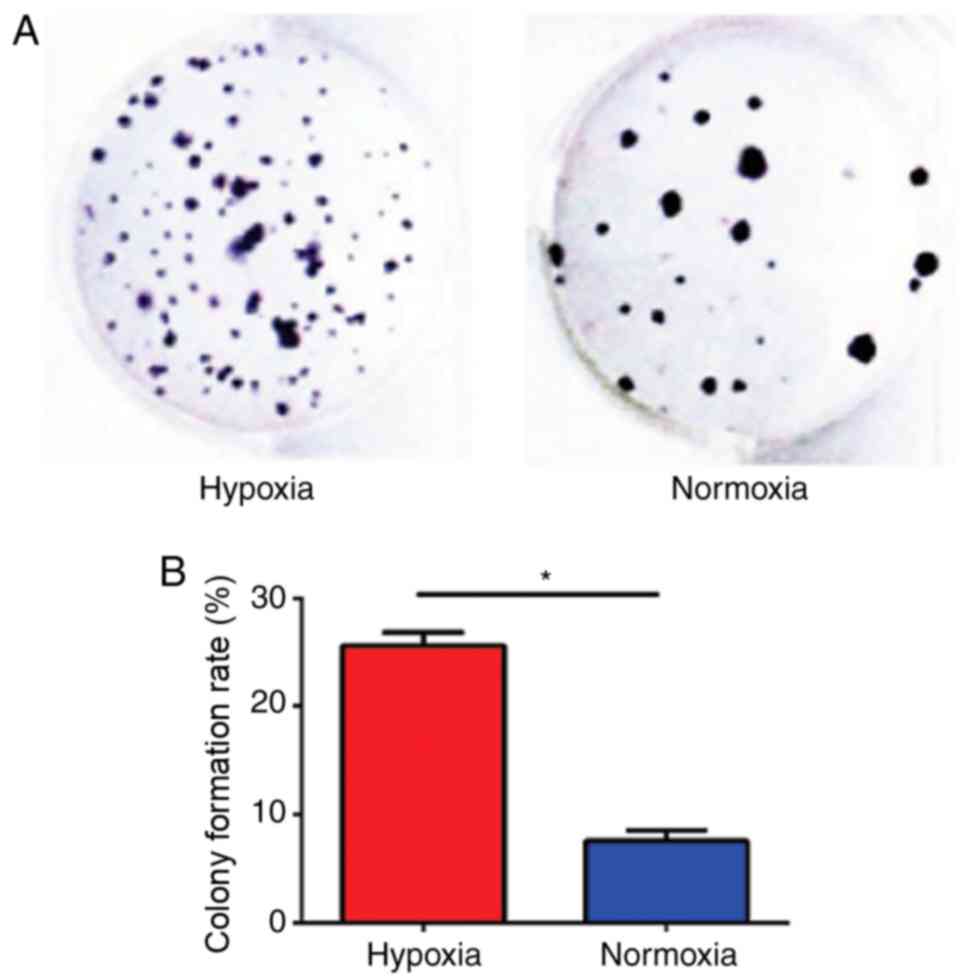
To assess the proliferation of U87 cells under
hypoxic conditions, a colony formation assay was performed in U87
cells (Fig. 3A). U87 cells cultured
under hypoxic conditions possessed a significantly greater clone
formation ability (25.38±1.46%) compared with U87 cells cultured
under normal conditions (7.25±1.56%; P<0.05; Fig. 3B).
CD133 expression is increased in U87
cells cultured in a hypoxic environment
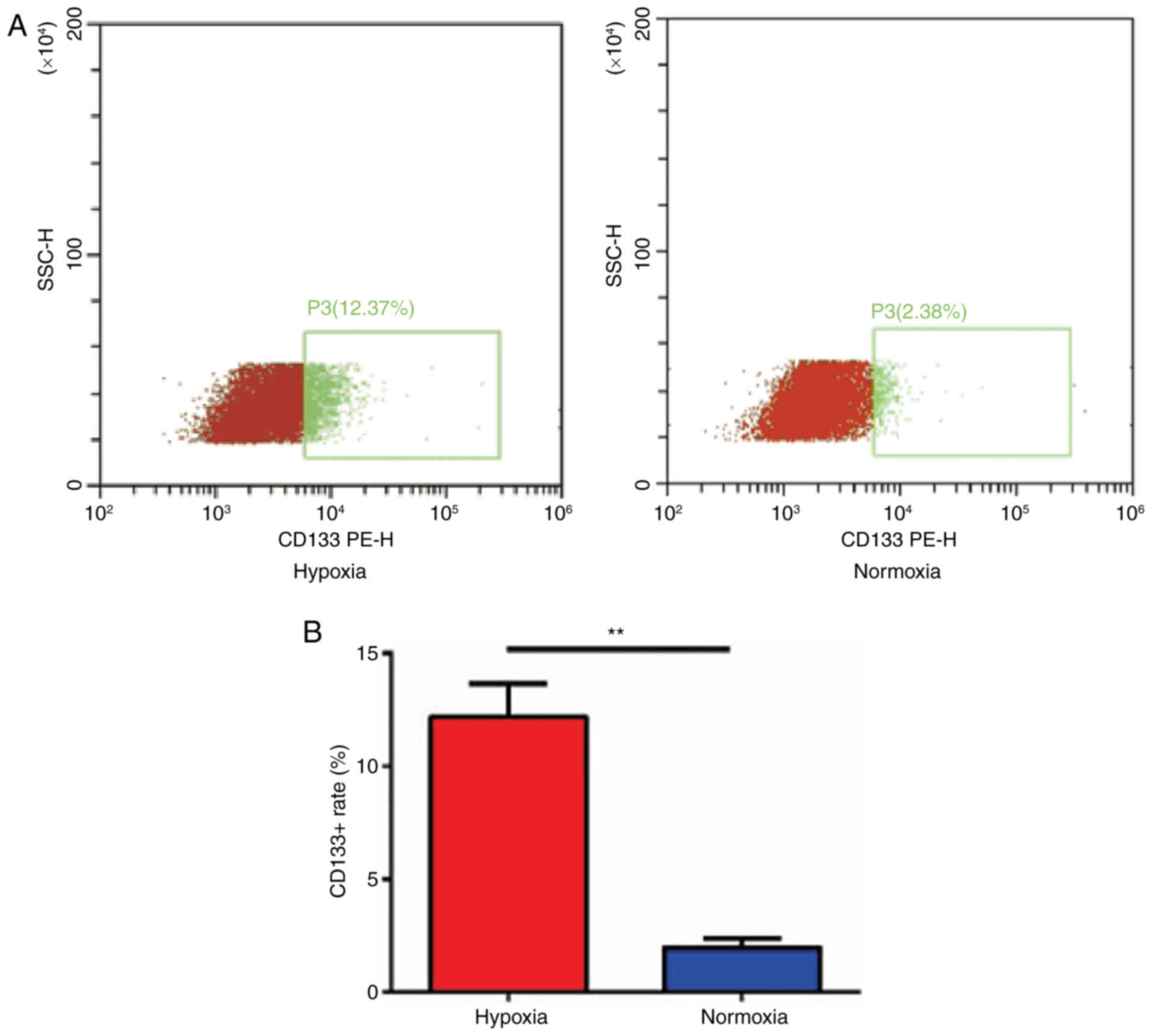
To assess if CD133 expression can be induced under
hypoxic conditions, the expression of CD133 in U87 cells was
detected using flow cytometry (Fig.
4A). The data demonstrated that the expression of CD133 in U87
cells cultured in a hypoxic environment was significantly higher
(12.18±0.86%) compared with those cultured in a normal environment
(1.96±0.23%; P<0.01; Fig. 4B).
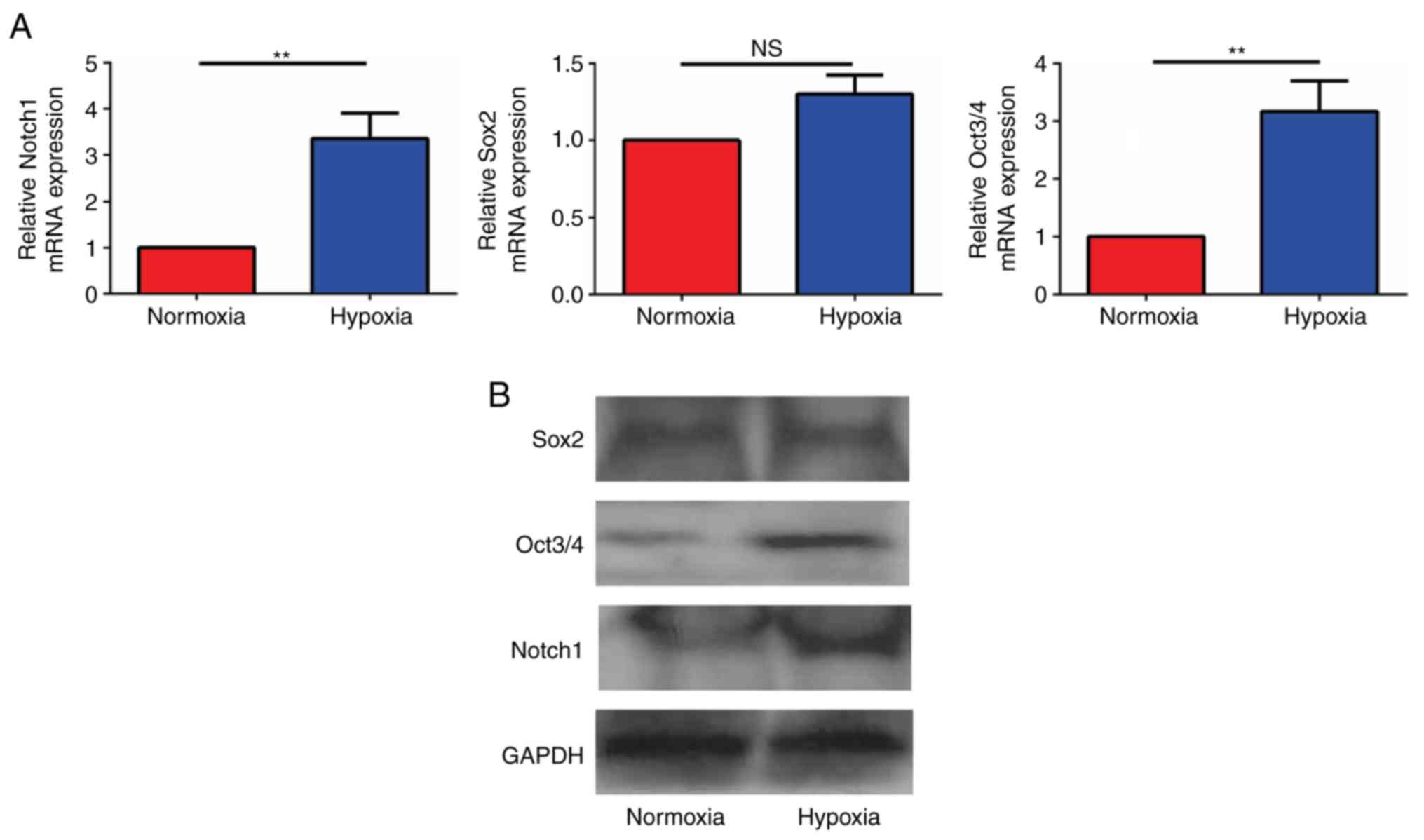
Hypoxia increases the mRNA expression
of genes in the Notch signaling pathway
To detect mRNA expression of genes in the Notch
signaling pathway in glioma cells, U87 cells cultured in hypoxia
and normoxia for 24 h were collected, and the mRNA expression
levels were detected by RT-qPCR. When the mRNA expression levels in
the hypoxia group were compared with that of the normoxia group,
the ratios were revealed to be 3.12±0.23 for Notch1, for 1.23±0.35
Sox2 and 3.17±0.30 for Oct3/4. The results demonstrated that the
Notch1 and Oct3/4 mRNA levels in the hypoxia group were
significantly increased compared with those in the normoxia group
and that the difference was statistically significant (P<0.05;
Fig. 5A). No significant difference
in the expression of the stem cell associated gene, Sox2, was
identified between the two groups.
Hypoxia increases the protein
expression of genes in the Notch signaling pathway
To detect the protein expression of genes in the
Notch signaling pathway in glioma cells, U87 cells cultured in
hypoxia and normoxia environments for 24 h were collected, and the
proteins were detected by western blot analysis. Notch1, Oct3/4 and
Sox2 were identified in all groups, and the expression of Notch1
and Oct3/4 were higher in hypoxic cells compared with normoxic
cells, while hypoxia did not increase Sox2 expression (Fig. 5B).
Discussion
Glioma is the most common type of primary tumor of
the central nervous system (1) and
recurrence, which is associated with the unlimited proliferation
and invasive growth of tumor cells, is the greatest hindrance to
successful treatment (2). Although
differentiated tumor cells constituted the majority of the cell
killed with the administration of current cancer treatments, a
small amount of CSCs are resistant to the treatment (4,5). GSCs
serve a very important role in the occurrence, development,
invasion and recurrence of gliomas (6–9). GSCs have
been demonstrated to cause tumors, maintain tumor growth, and are
the root of tumor recurrence following surgery (6–8). In
addition, there is a close association between tumor angiogenesis
and resistance to chemotherapy or radiotherapy (10,11).
Therefore, GSCs are a potential target for glioma therapy (31). Identifying a specific surface marker
of CSCs is the key to further study of the occurrence, metastasis,
recurrence and prognosis of the tumor. Multiple studies have
revealed that CD133 is a specific surface marker of stem cells and
CSCs (32–34). A hypoxic microenvironment is one of
the important features of solid tumors as well as gliomas (35). Hypoxia can promote cell proliferation
and angiogenesis in tumors, inhibit stem cell differentiation,
increase tumor resistance to chemotherapy and radiotherapy, and
make tumors more invasive (36).
The growth rate of tumors is usually greater than
that of the blood vessels, which generates a hypoxic state inside
the tumor (35). The hypoxic
microenvironment has been revealed to be associated with the growth
and metastasis of tumors (36). The
results of the current study demonstrated that cell proliferation
under hypoxia (1% O2) for >24 h was slightly higher
compared with that under normoxia (20% O2), which
suggested that the cell proliferation ability was increased
following hypoxia induction. This paradoxical phenomenon may be
attributed to changes in multiple signaling pathways that
facilitate the de-differentiation and the acquisition of stem
cell-like features in differentiated cancer cells (37). The ability of colony formation was
also increased by hypoxia, which suggested that the
microenvironment of hypoxia could promote stem cell characteristics
in tumor cells. At the same time, it was demonstrated that CD133
expression in U87 cells increased 4–6-fold following hypoxia
treatment, which is consistent with a previously reported 3–5-fold
induction of CD133 positive fractions in glioblastoma-derived cells
under hypoxia (38).
In embryonic stem cells, Oct3/4, Sox2 and other
transcription factors may form a transcriptional network that
regulates a large number of genes associated with differentiation
of embryonic stem cell (39). Subtle
changes in Oct3/4 protein expression have great influences on the
differentiation of embryonic stem cells (40). Studies have demonstrated that Oct3/4
was only expressed in pluripotent cells, and that it maintained the
pluripotency of embryonic stem cells, inhibited embryonic stem cell
apoptosis and maintained their proliferative potential through the
signal transduction pathway (40,41). The
results of the current study supported the role of Oct3/4 in the
proliferation of undifferentiated cells and stem cells that may be
involved in tumor growth.
The Notch1 signaling pathway serves a major role in
the maintenance of stem cell proliferation (42). In malignant tumors, the Notch1
signaling pathway is activated when the cells are transformed to
malignant cells (43,44). The results of the current study
demonstrated that the expression of Notch1 in U87 cells was
affected by the hypoxia microenvironment. The Notch signaling
pathway serves an important role in the self-renewal, proliferation
and differentiation of neural stem cells or neural precursor cells
(42). In addition, the Notch
signaling pathway was also revealed to be highly activated in
glioma, hematological and intestinal tumor, and other types of
tumor, which suggested that the pathway can regulate the
self-renewal and differentiation of CSCs, and serve an important
role in malignant tumor formation and development (43–45). A
previous study demonstrated that the Notch signaling pathway serves
an essential role in hypoxia-mediated stem cell maintenance
(37). Politi et al (44) revealed that the Notch signaling
pathway is involved in the carcinogenesis of breast cancer,
maintains the malignant phenotype of the mutant cells and is highly
expressed in transplanted breast cancer tissue in nude mice. Jiang
et al (45) demonstrated that
compared with normal brain tissue, the expression of Notch1 was
significantly higher in glioma tissues and increased with the grade
of the glioma. Zhang et al (46) revealed that the activation of the
Notch1 signaling pathway can stimulate the activation of RAC-α
serine/threonine-protein kinase, activate the β-catenin and nuclear
factor NF-κ-B signaling pathway, and promote the proliferation and
invasion of glioma.
In the present study, it was demonstrated that in
the hypoxic environment, Notch1 and its downstream target gene,
Oct3/4, were markedly increased at mRNA and protein levels, while
there appeared to be little change in Sox2 expression. These
results indicated that hypoxia could promote the proliferation of
GSCs and maintain the characteristics of stem cells by activating
Notch1 and Oct3/4 or maintaining Notch1 activation.
Acknowledgements
Not applicable.
Funding
The present study was supported by Hubei Natural
Science Foundation Project (grant no. 2016CFB575; Wuhan, China) and
Wuhan University Renmin Hospital Guidance Fund (grant no.
RMYD2018M09; Wuhan, China).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
FZ designed the project, performed the assays and
analyzed the data regarding U87 cell viability, cell apoptosis, CD
133 expression, and the Notch signaling pathway, and was a major
contributor in writing the manuscript. HC performed the assays and
analyzed the data regarding the U87 cell viability, colony
formation, and Notch signaling pathway detection. ZZ, TY and GW
performed the assays and analyzed the data regarding the U87 cell
apoptosis, CD133 expression and Notch signaling pathway. QZ and SD
contributed to the data acquisition and analysis regarding to the
Notch signaling pathway detection. YZ was a major contributor to
the project design, data analysis and manuscript editing. All
authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Louis DN, Perry A, Reifenberger G, von
Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD,
Kleihues P and Ellison DW: The 2016 world health organization
classification of tumors of the central nervous system: A summary.
Acta Neuropathol. 131:803–820. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Davis ME: Glioblastoma: Overview of
disease and treatment. Clin J Oncol Nurs. 20 Suppl 5:S2–S8. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ostrom QT, Gittleman H, Fulop J, Liu M,
Blanda R, Kromer C, Wolinsky Y, Kruchko C and Barnholtz-Sloan JS:
Central brain tumor registry of the united States (CBTRUS)
statistical report: Primary brain and central nervous system tumors
diagnosed in the united states in 2008–2012. Neuro Oncol. 17 Suppl
4:iv1–iv62. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Clarke MF, Dick JE, Dirks PB, Eaves CJ,
Jamieson CH, Jones DL, Visvader J, Weissman IL and Wahl GM: Cancer
stem cells-perspectives on current status and future directions:
AACR Workshop on cancer stem cells. Cancer Res. 66:9339–9344. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Soltanian S and Matin MM: Cancer stem
cells and cancer therapy. Tumour Biol. 32:425–440. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Singh SK, Clarke ID, Terasaki M, Bonn VE,
Hawkins C, Squire J and Dirks PB: Identification of a cancer stem
cell in human brain tumors. Cancer Res. 63:5821–5828.
2003.PubMed/NCBI
|
|
7
|
Singh SK, Hawkins C, Clarke ID, Squire JA,
Bayani J, Hide T, Henkelman RM, Cusimano MD and Dirks PB:
Identification of human brain tumour initiating cells. Nature.
432:396–401. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Uchida N, Buck DW, He D, Reitsma MJ, Masek
M, Phan TV, Tsukamoto AS, Gage FH and Weissman IL: Direct isolation
of human central nervous system stem cells. Proc Natl Acad Sci USA.
97:14720–14725. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Galli R, Binda E, Orfanelli U, Cipelletti
B, Gritti A, De Vitis S, Fiocco R, Foroni C, Dimeco F and Vescovi
A: Isolation and characterization of tumorigenic, stem-like neural
precursors from human glioblastoma. Cancer Res. 64:7011–7021. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Vescovi AL, Galli R and Reynolds BA: Brain
tumour stem cells. Nat Rev Cancer. 6:425–436. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bao S, Wu Q, McLendon RE, Hao Y, Shi Q,
Hjelmeland AB, Dewhirst MW, Bigner DD and Rich JN: Glioma stem
cells promote radioresistance by preferential activation of the DNA
damage response. Nature. 444:756–760. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Costa FF, Seftor EA, Bischof JM,
Kirschmann DA, Strizzi L, Arndt K, Bonaldo Mde F, Soares MB and
Hendrix MJ: Epigenetically reprogramming metastatic tumor cells
with an embryonic microenvironment. Epigenomics. 1:387–398. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Eales KL, Hollinshead KE and Tennant DA:
Hypoxia and metabolic adaptation of cancer cells. Oncogenesis.
5:e1902016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Jensen RL: Brain tumor hypoxia:
Tumorigenesis, angiogenesis, imaging, pseudoprogression, and as a
therapeutic target. J Neurooncol. 92:317–335. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Birner P, Piribauer M, Fischer I,
Gatterbauer B, Marosi C, Ambros PF, Ambros IM, Bredel M, Oberhuber
G, Rössler K, et al: Vascular patterns in glioblastoma influence
clinical outcome and associate with variable expression of
angiogenic proteins: Evidence for distinct angiogenic subtypes.
Brain Pathol. 13:133–143. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chi JT, Wang Z, Nuyten DS, Rodriguez EH,
Schaner ME, Salim A, Wang Y, Kristensen GB, Helland A,
Børresen-Dale AL, et al: Gene expression programs in response to
hypoxia: Cell type specificity and prognostic significance in human
cancers. PLoS Med. 3:e472006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sathornsumetee S, Cao Y, Marcello JE,
Herndon JE II, McLendon RE, Desjardins A, Friedman HS, Dewhirst MW,
Vredenburgh JJ and Rich JN: Tumor angiogenic and hypoxic profiles
predict radiographic response and survival in malignant astrocytoma
patients treated with bevacizumab and irinotecan. J Clin Oncol.
26:271–278. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Semenza GL: Targeting HIF-1 for cancer
therapy. Nat Rev Cancer. 3:721–732. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Semenza GL: Intratumoral hypoxia,
radiation resistance, and HIF-1. Cancer Cell. 5:405–406. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Vaupel P and Mayer A: Hypoxia in cancer:
Significance and impact on clinical outcome. Cancer Metastasis Rev.
26:225–239. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ludwig K and Kornblum HI: Molecular
markers in glioma. J Neurooncol. 134:505–512. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Gonzalez F and Huangfu D: Mechanisms
underlying the formation of induced pluripotent stem cells. Wiley
Interdiscip Rev Dev Biol. 5:39–65. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Takahashi K and Yamanaka S: Induction of
pluripotent stem cells from mouse embryonic and adult fibroblast
cultures by defined factors. Cell. 126:663–673. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Fazio C and Ricciardiello L: Inflammation
and notch signaling: A crosstalk with opposite effects on
tumorigenesis. Cell Death Dis. 7:e25152016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Jundt F, Pröbsting KS, Anagnostopoulos I,
Muehlinghaus G, Chatterjee M, Mathas S, Bargou RC, Manz R, Stein H
and Dörken B: Jagged1-induced Notch signaling drives proliferation
of multiple myeloma cells. Blood. 103:3511–3515. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Weijzen S, Rizzo P, Braid M, Vaishnav R,
Jonkheer SM, Zlobin A, Osborne BA, Gottipati S, Aster JC, Hahn WC,
et al: Activation of Notch-1 signaling maintains the neoplastic
phenotype in human ras-transformed cells. Nat Med. 8:979–986. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Balint K, Xiao M, Pinnix CC, Soma A, Veres
I, Juhasz I, Brown EJ, Capobianco AJ, Herlyn M and Liu ZJ:
Activation of Notch1 signaling is required for
beta-catenin-mediated human primary melanoma progression. J Clin
Invest. 115:3166–3176. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
28
|
Amaya-Chanaga CI and Rassenti LZ:
Biomarkers in chronic lymphocytic leukemia: Clinical applications
and prognostic markers. Best Pract Res Clin Haematol. 29:79–89.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Allen M, Bjerke M, Edlund H, Nelander S
and Westermark B: Origin of the U87MG glioma cell line: Good news
and bad news. Sci Transl Med. 8:354re32016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Evers P, Lee PP, DeMarco J, Agazaryan N,
Sayre JW, Selch M and Pajonk F: Irradiation of the potential cancer
stem cell niches in the adult brain improves progression-free
survival of patients with malignant glioma. BMC Cancer. 10:3842010.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Yin AH, Miraglia S, Zanjani ED,
Almeida-Porada G, Ogawa M, Leary AG, Olweus J, Kearney J and Buck
DW: AC133, a novel marker for human hematopoietic stem and
progenitor cells. Blood. 90:5002–5012. 1997.PubMed/NCBI
|
|
33
|
Wu Y and Wu PY: CD133 as a marker for
cancer stem cells: Progresses and concerns. Stem Cells Dev.
18:1127–1134. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Li Z: CD133: A stem cell biomarker and
beyond. Exp Hematol Oncol. 2:172013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Denko NC: Hypoxia, HIF1 and glucose
metabolism in the solid tumour. Nat Rev Cancer. 8:705–713. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Muz B, de la Puente P, Azab F and Azab AK:
The role of hypoxia in cancer progression, angiogenesis,
metastasis, and resistance to therapy. Hypoxia (Auckl). 3:83–92.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gustafsson MV, Zheng X, Pereira T, Gradin
K, Jin S, Lundkvist J, Ruas JL, Poellinger L, Lendahl U and
Bondesson M: Hypoxia requires notch signaling to maintain the
undifferentiated cell state. Dev Cell. 9:617–628. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Bar EE, Lin A, Mahairaki V, Matsui W and
Eberhart CG: Eberhart, hypoxia increases the expression of
stem-cell markers and promotes clonogenicity in glioblastoma
neurospheres. Am J Pathol. 177:1491–1502. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Rizzino A: Concise review: The Sox2-Oct4
connection: Critical players in a much larger interdependent
network integrated at multiple levels. Stem Cells. 31:1033–1039.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Niwa H, Miyazaki J and Smith AG:
Quantitative expression of Oct-3/4 defines differentiation,
dedifferentiation or self-renewal of ES cells. Nat Genet.
24:372–376. 2000. View
Article : Google Scholar : PubMed/NCBI
|
|
41
|
Tai MH, Chang CC, Kiupel M, Webster JD,
Olson LK and Trosko JE: Oct4 expression in adult human stem cells:
Evidence in support of the stem cell theory of carcinogenesis.
Carcinogenesis. 26:495–502. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Liu J, Sato C, Cerletti M and Wagers A:
Notch signaling in the regulation of stem cell self-renewal and
differentiation. Curr Top Dev Biol. 92:367–409. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Li JL and Harris AL: Notch signaling from
tumor cells: A new mechanism of angiogenesis. Cancer Cell. 8:1–3.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Politi K, Feirt N and Kitajewski J: Notch
in mammary gland development and breast cancer. Semin Cancer Biol.
14:341–347. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Jiang L, Wu J, Chen Q, Hu X, Li W and Hu
G: Notch1 expression is upregulated in glioma and is associated
with tumor progression. J Clin Neurosci. 18:387–390. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Zhang X, Chen T, Zhang J, Mao Q, Li S,
Xiong W, Qiu Y, Xie Q and Ge J: Notch1 promotes glioma cell
migration and invasion by stimulating β-catenin and NF-κB signaling
via AKT activation. Cancer Sci. 103:181–190. 2012. View Article : Google Scholar : PubMed/NCBI
|