Introduction
Lung cancer is the most frequently diagnosed cancer
and was reported in 2015 as the leading cause of cancer-associated
mortalities globally, as its incidence and mortality rate have been
increasing in numerous countries, including China (1). Of the lung cancer instances, ~80% are
non-small cell lung cancer types (NSCLC), which are clinically and
pathologically different from SCLC types (2). Treatment options for lung cancer include
surgery, radiation therapy, chemotherapy and targeted therapy.
Therapeutic modalities depend on a number of factors, including the
type and stage of cancer (3). Despite
ongoing therapeutic efforts, patients with lung cancer have a poor
prognosis with an arithmetic average 5-year survival rate of 15%
(4). This is primarily due to
inadequate knowledge regarding tumor progression and its associated
molecular alterations, which delay diagnosis (5); therefore, improvements in molecular
genetics diagnosis and prediction of prognosis for targeted
treatments and clinical decisions are required.
microRNAs (miRNAs) are a class of non-coding single
stranded RNA molecules of ~22 nucleotides, which are encoded by
endogenous genes (6). miRNAs are
important regulators of gene expression in plants and animals
(7). Recent studies have determined
that miRNAs are associated with the formation and suppression of
tumors (8–10). Changes in miRNA expression may serve
an essential role in tumorigenesis and cancer inhibition. A number
of miRNAs act as tumor suppressors, while others stimulate tumor
growth. For example, there is reduced miR-143 expression in
patients with colorectal cancer (11), miR-15-a and miR-16-1 are reduced in
patients with B cell chronic lymphocytic leukemia (12), precursor miR-155 is highly expressed
in Burkitt lymphoma, and the miR-17/92 cluster has been determined
to be highly expressed in lung cancer, particularly in patients
with SCLC (13,14). Additionally, miR-608 regulates
apoptosis in human lung cancer via the regulation of AKT
serine/threonine kinase (Akt)2, and miR-99a suppresses the invasion
and migration of NSCLC cells (15).
miR-101-3p is a member of the miR-101 family, and a recent study
indicated that it has tumor suppressor effects in patients with
NSCLC (16). However, to the best of
our knowledge, a limited number of studies have addressed the
association between miR-101-3p and adjuvant chemotherapy in
patients with NSCLC (17–19).
In the present study, the miR-101-3p expression was
investigated using the Gene Expression Omnibus (GEO) database and
the expression levels of miR-101-3p in NSCLC tissues was evaluated
using quantitative polymerase chain reaction (qPCR). Additionally,
the association between miR-101-3p and prognosis following adjuvant
chemotherapy was investigated in patients with NSCLC.
Materials and methods
Compliance with ethical standards
The present study was approved by the Ethics
Committee of Shanghai Tenth People's Hospital, Tongji University
School of Medicine (approval no. SHSY-IEC-pap-15-18; Shanghai,
China). Each participant provided signed informed consent prior to
participate in the present study. Patients or their legal
surrogates provided signed informed consent for the surgical
procedures. All specimens were handled and anonymized according to
ethical and legal standards.
miRNA expression in NSCLC from the GEO
database
The expression levels of miRNAs were assessed in
NSCLC tissues and normal tissue samples from the GEO database
(http://www.ncbi.nlm.nih.gov/geo/)
(20–23) using the following keywords: ‘Homo
sapiens’; ‘NSCLC’ and ‘miRNA’. All datasets used the Illumina or
Agilent Array platform to detect signals. For quality control,
exclusion criteria for probes were as follows: i) had a low bead
count of <3 in at least 5% of samples and ii) indicated a
detection-P>0.05 in at least 5% of samples (20). The raw data set GSE61741 (21) was downloaded, which provided the
peripheral blood miRNA profiles from 94 healthy controls and 73
patients with lung cancer. Additionally, GSE24709 (22) (including 19 healthy controls and 28
patients with lung cancer) and GSE56036 (23) (including 29 health controls and 23
patients with NSCLC) were downloaded in the GEO datasets to
identify differentially expressed miRNAs in NSCLC samples and
adjacent non-tumor tissues. Fold change (FC ≥2) and P<0.05
served as basic screening parameters. Hierarchical clustering was
performed using the multiple experiment viewer 4.7.1 software
programs (http://www.tm4.org/).
Clinical specimens
A total of 327 lung cancer tissues were collected
from 206 male patients and 121 female patients with lung cancer who
underwent surgery in Shanghai Tenth People's Hospital of the Tongji
University School of Medicine (Shanghai, China) between January
2004 and December 2016. Inclusion criteria consisted of the
following: ≤75 years with histologically proven NSCLC; no severe
major organ dysfunction; World Health Organization (WHO)
performance status of 0 or 1 and no prior cancer chemotherapy.
Exclusion criteria consisted of the following: Age ≥76; severe
major organ dysfunction; WHO performance status of >1 or prior
cancer chemotherapy. Clinical information of the patients were
recorded, including sex, age, smoking history, the diameter and
differentiation of the tumor, lymph node metastasis, stage of
Tumor-Node-Metastasis (TNM), histological grade, degree of invasion
of the lung membrane, degree of vascular invasion, whether
chemotherapy had been administered, overall survival (OS) rate,
disease-free survival (DFS) rate and miR-101-3p expression status.
Each participant provided signed informed consent prior to
participation in the present study. Patients or their legal
surrogates provided signed informed consent for the surgical
procedures.
RNA extraction and detection of
miR-101-3p expression by qPCR
RNA, including miRNAs, from NSCLC and normal tissue
samples, were extracted using TRIzol® reagent (Thermo
Fisher Scientific, Inc., Waltham, MA, USA), according to the
manufacturer's protocols and an optimized protocol (13). RNA concentration was measured using
NanoDrop ND-1000 (Thermo Fisher Scientific, Inc.) and the quality
was assessed using electrophoresis in 1.5% denaturing agarose gels
and viewed on a Kodak Gel Logic 2200 imaging system (Kodak,
Rochester, NY, USA). TaqMan probe-based qPCR was carried out using
a commercial kit (cat. no., A25576; Applied Biosystems; Thermo
Fisher Scientific, Inc.), according to the protocol of the
manufacturer (24). RT reactions were
performed using AMV Reverse Transcriptase (Takara Biotechnology
Co., Ltd., Dalian, China) and qPCR was performed using a standard
TaqMan PCR kit protocol on the Applied Biosystems 7900HT Sequence
Detection system (Thermo Fisher Scientific, Inc.), according to the
manufacturer's protocols. qPCR for miR-101-3p was executed using
the TaqMan® universal PCR kit (Thermo Fisher Scientific,
Inc.), according to the manufacturer's protocols. Thermocycling
conditions were as follows: Initial denaturation at 94°C for 10
min, followed by 35 cycles of 94°C for 30 sec, 60°C for 30 sec and
72°C for 30 sec, with a final extension at 72°C for 10 min. Each
reaction was independently tested in duplicate at a minimum of
three times. The following primers were used: miR-101-3p, forward,
5′-GGTCACTAAGGCGGT-3′ and reverse, 5′-CAGTCGTTGCGTCGGAGT-3′; U6,
forward, 5′-CTGGTTAGTACTTGGACGGGAGAC-3′ and reverse,
5′-GTGCAGGGTCCGAGGT-3′. U6 was used as the endogenous control and
the 2−ΔΔCq method was used to analyze expression levels
(25).
Statistical analysis
Expression levels of miR-101-3p were summarized and
presented as the mean ± standard deviation. All statistical
analyses were performed with IBM SPSS statistics software version
20.0 for Windows (IBM Corp., Armonk, NY, USA). The paired Student's
t-test was used to determine the difference between two groups of
data. The χ2 test was used to evaluate the differences
among groups. Kaplan-Meier estimator curves and the log-rank test
were performed to analyze the OS or DFS of patients with NSCLC.
Hierarchical clustering was performed using the multiple experiment
viewer 4.7.1 software programs: (http://www.tm4.org/mev/), according to the
manufacturer's protocols. Univariate and multivariate Cox
proportional hazards regression models were used to investigate the
multiple characteristics associated with the prognosis of patients
with NSCLC. P<0.05 was considered to indicate a statistically
significant difference.
Results
Expression of miR-101-3p using the GEO
database and clustering analysis
Firstly, miR-101-3p expression analysis was
performed using data from the GEO database. GSE24709 included the
miRNA profile in peripheral blood samples from patients with lung
diseases and healthy controls. The present analysis demonstrated
that miR-101-3p expression in NSCLC tissues was significantly
reduced, compared with non-tumor tissue (FC, 1.5; P<0.00001;
Fig. 1A and B). The miR-101-3p
expression was validated in NSCLC tissues in two datasets (GSE56036
and GSE61741) and indicated that miR-101-3p levels were
significantly reduced in NSCLC tissues, compared with normal
tissues (P=0.005, Fig. 1C; P=0.009,
Fig. 1D).
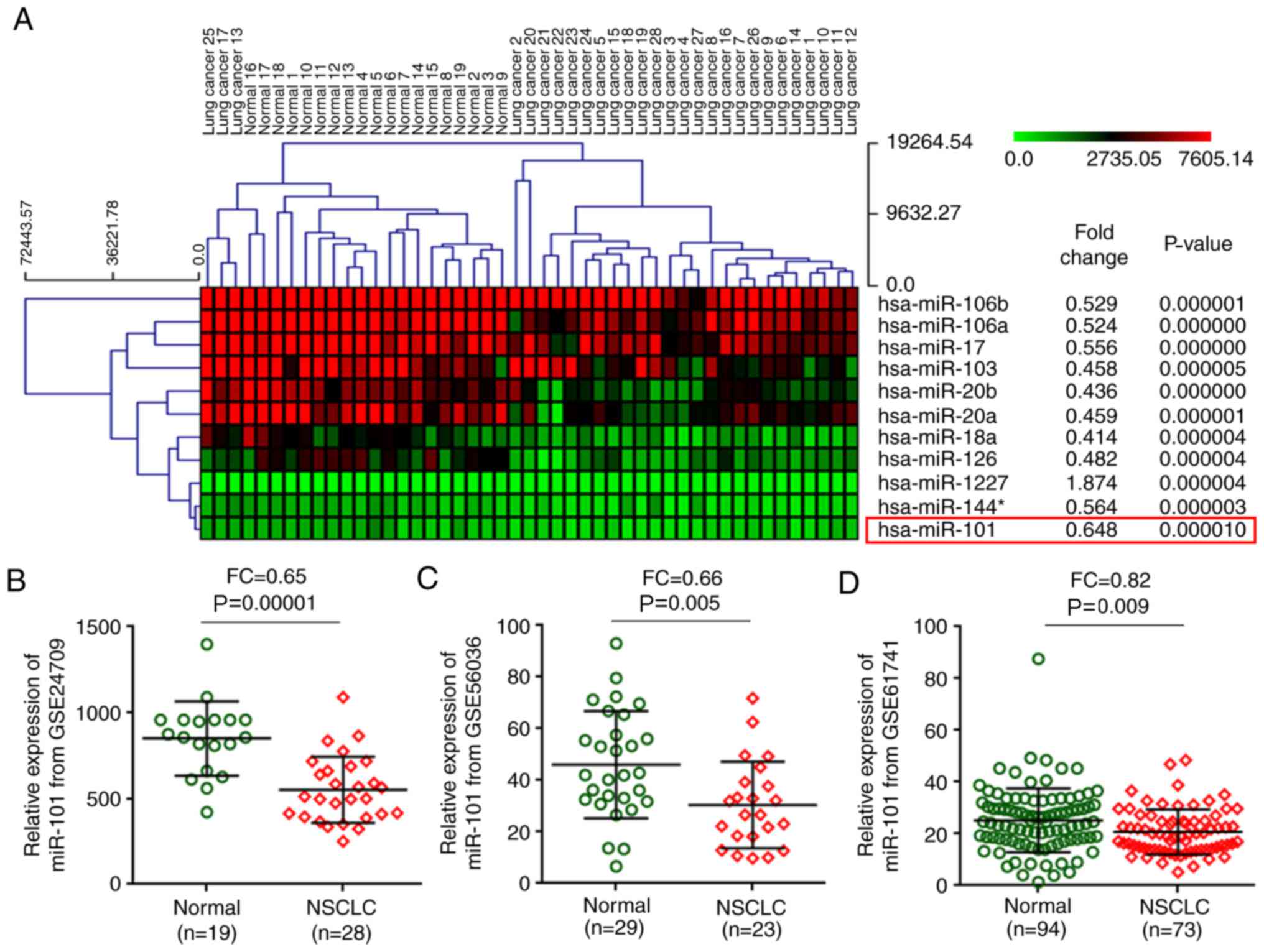 | Figure 1.Analysis of miR-101-3p expression in
NSCLC using the GEO database. (A) Data from the GEO dataset
(GSE24709) was clustered using the multiple experiment viewer 4.7.1
software. (B) The miR-101-3P expression level in normal tissues,
compared with NSCLC tissues, from the GEO database (GSE24709). (C)
The miR-101-3p expression level in normal tissues, compared with
NSCLC tissues, from the GEO database (GSE56036). (D) The miR-101-3p
expression level in normal tissues, compared with NSCLC tissues,
from the GEO database (GSE61741). miR, microRNA; NSCLC, non-small
cell lung cancer; GEO, Gene Expression Omnibus; FC, fold
change. |
miR-101-3p expression in NSCLC and
adjacent non-tumor tissues
Expression of miR-101-3p was evaluated in NSCLC
samples (n=327) and compared with adjacent non-tumor tissue (n=42)
by qPCR. The present results indicated that miR-101-3p expression
levels were significantly reduced in NSCLC tissues, compared with
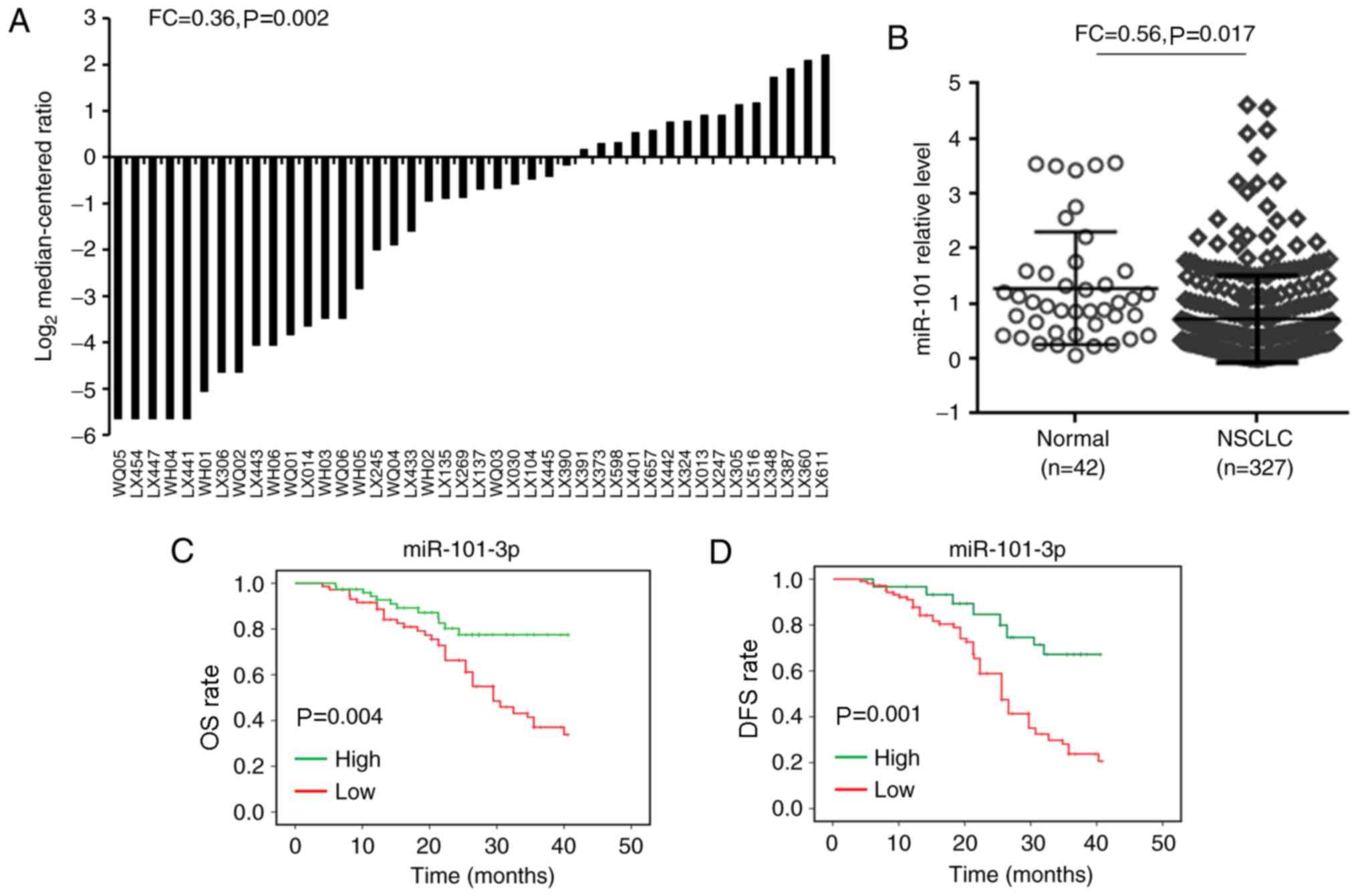
adjacent non-tumor tissues (FC, 0.36; P=0.002; Fig. 2A). The level of miR-101-3p expression
was 0.72±0.04 in 327 NSCLC tissues, which was significantly reduced
compared with normal tissues (n=42; 1.28±0.16) (FC, 0.56; P=0.117;
Fig. 2B).
Association between clinical
characteristics and miR-101-3p expression
Additionally, miR-101-3p expression was analyzed in
NSCLC samples based on various clinical characteristics, including
age, sex, smoking history, lymph node metastasis, tumor
differentiation, histology, TNM stage, invasion of lung membrane,
vascular invasion and tumor diameter. Univariate analysis
demonstrated that miR-101-3p expression was significantly
associated with lymph node metastasis (P=0.08), tumor diameter
(P=0.019) and TNM stage (P=0.036) in all patients with NSCLC
(Table I). However, no significant
association was determined between miR-101-3p expression in NSCLC
samples and age, sex, smoking history, tumor differentiation,
histology, invasion of the lung membrane or vascular invasion
(P>0.05; Table I).
 | Table I.Association between miR-101
expression and clinical characteristics. |
Table I.
Association between miR-101
expression and clinical characteristics.
| Factor | No. of
patients | miR-101 expression
(mean ± SD) | P-value |
|---|
| Age (years) |
|
| 0.668 |
|
≥60 | 167 | 0.703±0.021 |
|
|
<60 | 160 | 0.728±0.059 |
|
| Sex |
|
| 0.239 |
|
Male | 206 | 0.732±0.045 |
|
|
Female | 121 | 0.699±0.091 |
|
| Smoking
history |
|
| 0.078 |
|
Smoked | 136 | 0.682±0.028 |
|
| Never
smoked | 119 | 0.735±0.071 |
|
|
Unknown | 72 | 0.728±0.031 |
|
| Lymph node
metastasis |
|
| 0.008a |
|
Positive | 105 | 0.632±0.084 |
|
|
Negative | 166 | 0.753±0.054 |
|
|
Unknown | 56 | 0.721±0.029 |
|
| Tumor
differentiation |
|
| 0.097 |
|
Poorly | 136 | 0.691±0.063 |
|
|
Moderately | 111 | 0.714±0.036 |
|
|
Well | 80 | 0.722±0.089 |
|
| Histology |
|
| 0.604 |
|
Adenocarcinoma | 185 | 0.705±0.039 |
|
|
Squamous cell carcinoma | 142 | 0.711±0.057 |
|
| TNM stage (38) |
|
| 0.036a |
|
III–IV | 158 | 0.681±0.067 |
|
|
I–II | 169 | 0.736±0.055 |
|
| Invasion of lung
membrane |
|
| 0.063 |
|
Positive | 109 | 0.686±0.029 |
|
|
Negative | 168 | 0.725±0.013 |
|
|
Unknown | 50 | 0.721±0.066 |
|
| Vascular
invasion |
|
| 0.269 |
|
Positive | 65 | 0.692±0.057 |
|
|
Negative | 189 | 0.721±0.033 |
|
|
Unknown | 73 | 0.708±0.037 |
|
| Tumor diameter
(cm) |
|
| 0.019a |
| ≥5 | 186 | 0.677±0.054 |
|
|
<5 | 141 | 0.743±0.065 |
|
Univariate analysis of prognosis based
on various clinical characteristics in patients with NSCLC
To ascertain whether the prognosis of patients with
NSCLC was influenced by age, sex, smoking history, lymph node
metastasis, tumor differentiation, histology, TNM stage, invasion
of lung membrane, vascular invasion, tumor diameter or miR-101-3p
expression, univariate analysis with the Kaplan-Meier estimator
method was performed. The results demonstrated that lymph node
metastasis (P=0.042), TNM stage (P=0.018), tumor diameter (P=0.013)
and miR-101-3p expression were significantly associated with OS
(P=0.004) and DFS (P=0.001) (Table
II; Fig 2).
 | Table II.Univariate analysis of OS and DFS
based on patients stratified by clinical characteristics. |
Table II.
Univariate analysis of OS and DFS
based on patients stratified by clinical characteristics.
|
|
| OS time | Progression-free
survival time |
|---|
|
|
|
|
|
|---|
| Factor | No. of
patients | Months (mean) | 95% CI (mean) | P-value (log-rank
test) | Months (mean) | 95% CI (mean) | P-value (log-rank
test) |
|---|
| Age (years) |
|
|
|
|
|
| 0.331 |
|
≥60 | 167 | 28.065 | 26.945–30.253 | 0.289 | 28.069 | 26.889–30.453 |
|
|
<60 | 160 | 29.331 | 28.314–31.846 |
| 29.564 | 28.651–31.754 |
|
| Sex |
|
|
|
|
|
| 0.822 |
|
Male | 206 | 29.652 | 28.773–31.325 | 0.806 | 30.594 | 28.797–31.393 |
|
|
Female | 121 | 31.068 | 28.962–32.618 |
| 31.056 | 28.845–32.648 |
|
| Smoking
history |
|
|
|
|
|
| 0.115 |
|
Smoked | 136 | 27.849 | 26.152–29.086 | 0.109 | 27.654 | 26.465–29.754 |
|
| Never
smoked | 119 | 29.659 | 27.854–30.554 |
| 29.435 | 27.784–30.058 |
|
|
Unknown | 72 | 28.381 | 26.989–31.062 |
| 28.675 | 27.095–31.365 |
|
| Lymph node
metastasis |
|
|
|
|
|
| 0.041a |
|
Positive | 105 | 26.731 | 24.377–29.501 | 0.042a | 26.456 | 24.943–29.063 |
|
|
Negative | 166 | 31.229 | 28.815–33.056 |
| 31.854 | 28.326–33.064 |
|
|
Unknown | 56 | 28.056 | 26.053–30.434 |
| 28.06 | 26.366–30.582 |
|
| Tumor
differentiation (38) |
|
|
|
|
|
| 0.274 |
|
Poorly | 136 | 28.851 | 26.826–31.058 | 0.246 | 28.821 | 26.487–31.348 |
|
|
Moderately | 111 | 29.348 | 27.854–31.854 |
| 29.815 | 27.145–31.487 |
|
|
Well | 80 | 31.857 | 29.581–32.783 |
| 31.825 | 29.747–33.045 |
|
| Histology (38) |
|
|
|
|
|
| 0.867 |
|
Adenocarcinoma | 185 | 29.744 | 26.451–31.845 | 0.865 | 28.745 | 26.787–31.165 |
|
|
Squamous cell carcinoma | 142 | 29.043 | 25.624–32.434 |
| 29.257 | 25.474–32.345 |
|
| TNM stage (38) |
|
|
|
|
|
| 0.012a |
|
III–IV | 158 | 26.748 | 24.285–30.647 | 0.018a | 26.778 | 24.548–30.487 |
|
|
I–II | 169 | 31.049 | 28.834–34.415 |
| 32.091 | 28.674–34.358 |
|
| Invasion of lung
membrane |
|
|
|
|
|
| 0.067 |
|
Positive | 109 | 27.995 | 25.453–29.454 | 0.086 | 27.123 | 25.748–29.486 |
|
|
Negative | 168 | 31.412 | 28.542–32.966 |
| 31.545 | 28.364–32.684 |
|
|
Unknown | 50 | 28.095 | 26.354–30.888 |
| 28.157 | 26.387–30.054 |
|
| Vascular
invasion |
|
|
|
|
|
| 0.066 |
|
Positive | 65 | 27.849 | 25.446–29.354 | 0.072 | 27.048 | 25.487–29.954 |
|
|
Negative | 189 | 31.069 | 26.878–33.147 |
| 31.157 | 26.748–33.444 |
|
|
Unknown | 73 | 29.534 | 27.956–31.259 |
| 29.348 | 27.248–31.187 |
|
| Tumor diameter
(cm) |
|
|
|
|
|
| 0.009a |
| ≥5 | 186 | 25.386 | 23.685–27.913 | 0.013a | 24.886 | 23.065–26.648 |
|
|
<5 | 141 | 31.553 | 27.456–33.546 |
| 30.661 | 28.446–32.596 |
|
| miR-101 expression
(median) |
|
|
|
|
|
| 0.001a |
|
Low | 163 | 24.154 | 22.314–26.043 | 0.004a | 24.365 | 22.625–26.443 |
|
|
High | 164 | 32.317 | 28.414–34.916 |
| 33.053 | 28.456–34.994 |
|
Univariate Cox regression analysis revealed that
lymph-node metastasis [hazard ratio (HR), 1.743; 95% confidence
interval (CI), 1.191–2.421; P=0.032], TNM stage (HR, 1.562; 95% CI,
1.124–1.962; P=0.036), tumor diameter (HR, 2.125; 95% CI,
1.563–3.346; P=0.005), chemotherapy (HR, 0.778; 95% CI,
0.469–0.968; P=0.026) and miR-101 expression (HR, 0.687; 95% CI,
0.498–0.952; P=0.003) were also positively significantly associated
with a poor prognosis. No significant associations were observed
for age, sex, smoking history, tumor differentiation, histology,
invasion of lung membrane or vascular invasion (Table III).
 | Table III.Cox regression model analysis for
prognosis based on various clinical characteristics in patients
with NSCLC. |
Table III.
Cox regression model analysis for
prognosis based on various clinical characteristics in patients
with NSCLC.
|
| miR-101 univariate
analysis | miR-101
multivariate analysis |
|---|
|
|
|
|
|---|
| Factor | HR | 95% CI | P-value | HR | 95% CI | P-value |
|---|
| Age | 1.153 | 0.681–1.554 | 0.093 | 1.176 | 0.699–1.568 | 0.087 |
| Sex | 0.887 | 0.525–1.115 | 0.801 | 0.867 | 0.504–1.089 | 0.542 |
| Smoking
history | 1.196 | 0.723–1.587 | 0.087 | 1.198 | 0.825–1.589 | 0.061 |
| Lymph-node
metastasis | 1.743 | 1.191–2.421 | 0.032a | 1.924 | 1.386–3.405 | 0.014a |
| Tumor
differentiation | 1.154 | 0.961–1.224 | 0.224 | 1.168 | 0.979–1.231 | 0.201 |
| Histology | 0.835 | 0.698–1.142 | 0.324 | 0.831 | 0.624–1.104 | 0.265 |
| TNM stage | 1.562 | 1.124–1.962 | 0.036a | 1.967 | 1.544–2.325 | 0.018a |
| Invasion of lung
membrane | 1.164 | 1.006–1.453 | 0.089 | 1.196 | 1.075–1.469 | 0.066 |
| Vascular
invasion | 1.169 | 1.021–1.468 | 0.088 | 1.182 | 1.069–1.494 | 0.059 |
| Tumor diameter | 2.125 | 1.563–3.346 | 0.005a | 2.869 | 2.025–3.396 | 0.002a |
| Chemotherapy | 0.778 | 0.469–0.968 | 0.026a | 0.559 | 0.374–0.786 | 0.006a |
| miR-101
expression | 0.687 | 0.498–0.952 | 0.003a |
|
|
|
Cox regression model analysis of
prognosis based on various clinical characteristics in patients
with NSCLC
To determine whether miR-101-3p expression levels,
in combination with lymph-node metastasis, TNM stage or tumor
diameter had prognostic value, multivariate analysis with a Cox
regression model was used (Table
II). This analysis also indicated that lymph node metastasis
(HR, 1.924; 95% CI, 1.386–3.405; P=0.014), TNM stage (HR, 1.967;
95% CI, 1.544–2.325; P=0.018) and tumor diameter (HR, 2.869; 95%
CI, 2.025–3.396; P=0.002) were significantly associated with
reduced prognosis (Table III). This
analysis initially included all of the parameters that were
predictive of OS in the univariate analysis of the entire study
group as presented in Table II (age,
sex, smoking history, lymph-node metastasis, tumor differentiation,
histology, vascular invasion, tumor diameter and invasion of the
lung membrane).
Prediction of OS and DFS for patients
with NSCLC based on chemotherapy alone or chemotherapy and
miR-101-3p expression
There was a statistically significant association
between chemotherapy and OS (27.624±3.858 vs. 31.457±2.924,
respectively; P=0.012) and DFS (26.985±2.247 vs. 31.004±3.357,
respectively; P=0.005) in patients with NSCLC. Chemotherapy is the
primary adjuvant treatment for the majority of patients with NSCLC
undergoing surgery. In the present study, it was determined that
adjuvant chemotherapy and high expression of miR-101-3p increased
the OS and DFS of patients (32.738±3.574 and 31.946±3.789,
respectively) compared with non-therapeutic patients with low
expression of miR-101-3p (25.352±2.568 and 25.004±2.876,
respectively) (Table IV). These data
indicated that patients with NSCLC with a high expression of
miR-101-3p may have an increased benefit from chemotherapy.
 | Table IV.OS and DFS of patients with NSCLC
based on chemotherapy alone or chemotherapy and miR-101
expression. |
Table IV.
OS and DFS of patients with NSCLC
based on chemotherapy alone or chemotherapy and miR-101
expression.
|
|
| OS | DFS |
|---|
|
|
|
|
|
|---|
|
Characteristics | No. of
patients | Mean ± SD | 95% CI | P-value | Mean ± SD | 95% CI | P-value |
|---|
| Chemotherapy |
|
|
| 0.012 |
|
| 0.005 |
|
Yes | 231 | 31.457±2.924 | 28.728–34.632 |
| 31.004±3.357 | 28.244–33.148 |
|
| No | 96 | 27.624±3.858 | 25.741–29.478 |
| 26.985±2.247 | 25.311–28.634 |
|
| Chemotherapy &
miR-101 expression |
|
|
| 0.001 |
|
| <0.001 |
| P &
H | 65 | 32.738±3.574 | 30.469–34.774 |
| 31.946±3.789 | 28.158–33.564 |
|
| N &
L | 87 | 25.352±2.568 | 22.914–27.353 |
| 25.004±2.876 | 23.043–26.965 |
|
Discussion
Lung cancer is the most common type of malignant
tumor and was reported in 2015 as the leading cause of
cancer-associated mortality worldwide (26). Drug resistance has remained the
primary factor influencing prognosis, therefore the treatment of
advanced and metastatic NSCLC remains a notable challenge (27). The recognition of novel biomarkers for
prediction of patient response to chemotherapy and prognosis is
essential for improving OS and DFS in patients with NSCLC.
As molecular biomarkers, miRNAs serve significant
roles in the selection of therapeutic schedules. Their functions as
tumor suppressors and oncogenes in human cancer have been
previously reported (28). In lung
cancer, miRNAs exhibit alterations in expression that predict
survival and relapse (29). miR-101
inhibits ovarian cancer cell invasion and proliferation by
downregulating the expression of suppression of cytokine signaling
2 (30), and also has implications in
suppressing the spread of a number of tumor types, including
chondrosarcoma (31), thyroid cancer
(32), breast cancer (33) and hepatocellular carcinoma (34). It has been reported that miR-101
inhibits cell proliferation and invasion of lung cancer by
regulating cyclooxygenase-2 (35).
Additionally, low expression of miR-101 in lung cancer has been
demonstrated to inhibit the invasion of lung cancer by regulating
its target gene enhancer of zerte 2 polycomb repressive complex 2
subunit (36).
miR-101-3p is a member of the miR-101 family and
inhibits the metastasis-associated lung adenocarcinoma transcript 1
(MALAT-1)-induced activation of the phosphoinositide 3-kinase/Akt
signaling pathway, resulting in the inhibition of NSCLC growth and
metastasis (16). It is notable that
miR-101-3p expression in NSCLC cells was significantly reduced,
whereas MALAT-1 expression was significantly increased.
Furthermore, high expression of miR-101-3p inhibits the
proliferation, migration and invasion of NSCLC (37). In the present study, significantly
reduced levels of miR-101-3p was observed in patients with NSCLC,
compared with healthy controls. The level of miR-101-3p expression
in NSCLC was determined to be associated with OS and DFS,
indicating that miR-101-3p may serve as a prognostic biomarker in
patients with NSCLC. The analysis of a number of clinical factors,
including tumor diameter, TNM stage and lymph node metastasis,
demonstrated associations with OS and DFS. Notably, as an
independent parameter, miR-101-3p expression levels in NSCLC also
affected prognosis and survival.
miR-101-3p is a valuable prognostic predictor and
therapeutic target of clinical chemotherapy. The present analysis
determined that a high level of expression of miR-101-3p in
patients with NSCLC who had undergone routine chemotherapy was
positively associated with OS and DFS rates. This indicated that
patients with NSCLC with high levels of miR-101-3p expression may
have improved benefit from chemotherapy. In brief, miR-101-3p was
significantly downregulated in NSCLC, which increased the OS and
DFS rates of patients receiving adjuvant chemotherapy. Based on
these results, whether the miR-101-3p expression level may serve as
a biomarker for chemotherapy use in patients with NSCLC should be
investigated, and therefore may be a valuable and promising
biomarker for this disease. However, the molecular and
pathophysiological mechanisms of miR-101-3p in NSCLC are not fully
understood, and will be assessed in subsequent studies. The present
study had a number of limitations. For example, this is a
retrospective study and only one marker was addressed. There are
other markers or associated pathways that must be studied. In the
future, multicenter studies regarding miR-101-3p in NSCLC should be
performed.
In conclusion, the present data confirmed that with
adjuvant chemotherapeutic treatment the median OS and DFS rate of
patients improved. The analyses demonstrated that low expression of
miR-101-3p, together with adjuvant chemotherapy, notably improved
the OS and DFS of patients with NSCLC. The use of miR-101-3p as a
specific and sensitive biomarker may be suitable for prediction of
therapeutic responses in patients with advanced NSCLC, which may
result in a superior level of personalized therapy. Therefore,
miR-101-3p may be considered as a potential biomarker for
chemosensitivity in the tumors of patients with NSCLC.
Acknowledgements
We would like to thank the experimental support of
Central Laboratory for Medical Research, Shanghai Tenth People's
Hospital (Shanghai, China).
Funding
The present study was partially supported by grants
from the National Natural Science Foundation of China (grant nos.
81472202, 81772932, 81201535, 81302065, 81301993, 81702243,
81260345 and 81372175), Shanghai Natural Science Foundation (grant
nos. 12ZR1436000 and 16ZR1428900), Shanghai Municipal Commission of
Health and Family Planning (grant nos. 201540228 and 201440398),
Program of Shanghai Subject Chief Scientist (grant no.
04.01.13.059), The Fundamental Research Funds for the Central
Universities (grant no. 22120170212 and 22120170117), Clinical
Research Special Foundation of the Wu Jieping Medical Foundation
(grant no. 320.6750.14326) and Nantong Science and Technology
Project (grant no. yyz15026).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
HML, WWY, YSM, FY, JBL, and DF designed the study.
HML, WWY, FY, YSM, WTX, HWF, ZWL, LKH, WW, JJJ, ZYC, MXS, YCS, LC,
CYJ, GXL and DF performed the qPCR experiments. HML, WWY, FY, YSM,
WTX, LKH, WW, MXS, HQY, CZ, LC, CYJ, GXL, CYW and DF performed the
statistical analyses and interpreted the data. FY, YSM, ZWL, LKH,
WW, HML, JJJ, MXS, LC, CYJ, GXL, CYW, XJZ, JBL, and DF are involved
in patient recruitment. FY, YSM, ZWL, CYW, JBL, and DF contributed
to study materials and consumables. FY, YSM, ZWL, WTX and DF wrote
the manuscript. HML, WWY, YSM, WW and FY contributed equally to
this work. All authors agreed with the results and conclusions.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Shanghai Tenth People's Hospital, Tongji University
School of Medicine (approval no. SHSY-IEC-pap-15-18). Each
participant provided signed informed consent prior to participate
in the present study. Patients or their legal surrogates provided
signed informed consent for the surgical procedures.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Lu G, Fu D, Jia C, Chai L, Han Y, Liu J,
Wu T, Xie R, Chang Z, Yang H, et al: Reduced miR-105-1 levels are
associated with poor survival of patients with non-small cell lung
cancer. Oncol Lett. 14:7842–7848. 2017.PubMed/NCBI
|
|
2
|
Zhang T, Rong N, Chen J, Zou C, Jing H,
Zhu X and Zhang W: SIRT1 expression is associated with the
chemotherapy response and prognosis of patients with advanced
NSCLC. PLoS One. 8:e791622013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hou L, Luo P, Ma Y, Jia C, Yu F, Lv Z, Wu
C and Fu D: MicroRNA-125a-3p downregulation correlates with
tumorigenesis and poor prognosis in patients with non-small cell
lung cancer. Oncol Lett. 14:4441–4448. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hou LK, Ma YS, Han Y, Lu GX, Luo P, Chang
ZY, Xie RT, Yang HQ, Chai L, Cai MX, et al: Association of
microRNA-33a molecular signature with non-small cell lung cancer
diagnosis and prognosis after chemotherapy. PLoS One.
12:e01704312017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yu F, Liu JB, Wu ZJ, Xie WT, Zhong XJ, Hou
LK, Wu W, Lu HM, Jiang XH, Jiang JJ, et al: Tumor suppressive
microRNA-124a inhibits stemness and enhances gefitinib sensitivity
of non-small cell lung cancer cells by targeting ubiquitin-specific
protease 14. Cancer Lett. 427:74–84. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ma YS, Wu TM, Lv ZW, Lu GX, Cong XL, Xie
RT, Yang HQ, Chang ZY, Sun R, Chai L, et al: High expression of
miR-105-1 positively correlates with clinical prognosis of
hepatocellular carcinoma by targeting oncogene NCOA1. Oncotarget.
8:11896–11905. 2017.PubMed/NCBI
|
|
7
|
Xu C, Haque F, Jasinski DL, Binzel DW, Shu
D and Guo P: Favorable biodistribution, specific targeting and
conditional endosomal escape of RNA nanoparticles in cancer
therapy. Cancer Lett. 414:57–70. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Salzman DW, Nakamura K, Nallur S, Dookwah
MT, Metheetrairut C, Slack FJ and Weidhaas JB: miR-34 activity is
modulated through 5′-end phosphorylation in response to DNA damage.
Nat Commun. 7:109542016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Persson H, Søkilde R, Häkkinen J, Pirona
AC, Vallon-Christersson J, Kvist A, Mertens F, Borg Å, Mitelman F,
Höglund M and Rovira C: Frequent miRNA-convergent fusion gene
events in breast cancer. Nat Commun. 8:7882017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Xie RT, Cong XL, Zhong XM, Luo P, Yang HQ,
Lu GX, Luo P, Chang ZY, Sun R, Wu TM, et al: MicroRNA-33a
downregulation is associated with tumorigenesis and poor prognosis
in patients with hepatocellular carcinoma. Oncol Lett.
15:4571–4577. 2018.PubMed/NCBI
|
|
11
|
Mi Y, Zhang D, Jiang W, Weng J, Zhou C,
Huang K, Tang H, Yu Y, Liu X, Cui W, et al: miR-181a-5p promotes
the progression of gastric cancer via RASSF6-mediated MAPK
signalling activation. Cancer Lett. 389:11–22. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cutrona G, Matis S, Colombo M, Massucco C,
Baio G, Valdora F, Emionite L, Fabris S, Recchia AG, Gentile M, et
al: Effects of miRNA-15 and miRNA-16 expression replacement in
chronic lymphocytic leukemia: Implication for therapy. Leukemia.
31:1894–1904. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Linnstaedt SD, Gottwein E, Skalsky RL,
Luftig MA and Cullen BR: Virally induced cellular microRNA miR-155
plays a key role in B-cell immortalization by Epstein-Barr virus. J
Virol. 84:11670–11678. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yanaihara N, Caplen N, Bowman E, Seike M,
Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T, et
al: Unique microRNA molecular profiles in lung cancer diagnosis and
prognosis. Cancer Cell. 9:189–198. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Othman N and Nagoor NH: miR-608 regulates
apoptosis in human lung adenocarcinoma via regulation of AKT2. Int
J Oncol. 51:1757–1764. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhang X, He X, Liu Y, Zhang H, Chen H, Guo
S and Liang Y: MiR-101-3p inhibits the growth and metastasis of
non-small cell lung cancer through blocking PI3K/AKT signal pathway
by targeting MALAT-1. Biomed Pharmacother. 93:1065–1073. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang H, Wang L, Zhang G, Lu C, Chu H, Yang
R and Zhao G: MALAT1/miR-101-3p/MCL1 axis mediates cisplatin
resistance in lung cancer. Oncotarget. 9:7501–7512. 2017.PubMed/NCBI
|
|
18
|
Liu C, Fu H, Liu X, Lei Q, Zhang Y, She X,
Liu Q, Liu Q, Sun Y, Li G and Wu M: LINC00470 coordinates the
epigenetic regulation of ELFN2 to distract GBM cell autophagy. Mol
Ther. 26:2267–2281. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wakasugi H, Takahashi H, Niinuma T,
Kitajima H, Oikawa R, Matsumoto N, Takeba Y, Otsubo T, Takagi M,
Ariizumi Y, et al: Dysregulation of miRNA in chronic hepatitis B is
associated with hepatocellular carcinoma risk after nucleos(t)ide
analogue treatment. Cancer Lett. 434:91–100. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Cao L, Chen Y, Zhang M, Xu DQ, Liu Y, Liu
T, Liu SX and Wang P: Identification of hub genes and potential
molecular mechanisms in gastric cancer by integrated bioinformatics
analysis. PeerJ. 6:e51802018. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Keller A, Leidinger P, Vogel B, Backes C,
ElSharawy A, Galata V, Mueller SC, Marquart S, Schrauder MG, Strick
R, et al: miRNAs can be generally associated with human pathologies
as exemplified for miR-144. BMC Med. 12:2242014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Leidinger P, Brefort T, Backes C, Krapp M,
Galata V, Beier M, Kohlhaas J, Huwer H, Meese E and Keller A:
High-throughput qRT-PCR validation of blood microRNAs in non-small
cell lung cancer. Oncotarget. 7:4611–4623. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Fujita Y, Yagishita S, Hagiwara K,
Yoshioka Y, Kosaka N, Takeshita F, Fujiwara T, Tsuta K, Nokihara H,
Tamura T, et al: The clinical relevance of the
miR-197/CKS1B/STAT3-mediated PD-L1 network in chemoresistant
non-small-cell lung cancer. Mol Ther. 23:717–727. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee
DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, et al:
Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic
Acids Res. 33:e1792005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta (T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Liang Z, Kong R, He Z, Lin LY, Qin SS,
Chen CY, Xie ZQ, Yu F, Sun GQ, Li CG, et al: High expression of
miR-493-5p positively correlates with clinical prognosis of non
small cell lung cancer by targeting oncogene ITGB1. Oncotarget.
8:47389–47399. 2017.PubMed/NCBI
|
|
27
|
Zhang B, Fu D, Xu Q, Cong X, Wu C, Zhong
X, Ma Y, Lv Z, Chen F, Han L, et al: The senescence-associated
secretory phenotype is potentiated by feedforward regulatory
mechanisms involving Zscan4 and TAK1. Nat Commun. 9:17232018.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Huang Q, Zhang XW, Ma YS, Lu GX, Xie RT,
Yang HQ, Lv ZW, Zhong XM, Liu T, Huang SX, et al: Up-regulated
microRNA-299 corrected with poor prognosis of glioblastoma
multiforme patients by targeting ELL2. Jpn J Clin Oncol.
47:590–596. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chang TH, Tsai MF, Gow CH, Wu SG, Liu YN,
Chang YL, Yu SL, Tsai HC, Lin SW, Chen YW, et al: Upregulation of
microRNA-137 expression by Slug promotes tumor invasion and
metastasis of non-small cell lung cancer cells through suppression
of TFAP2C. Cancer Lett. 402:190–202. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zheng HB, Zheng XG and Liu BP: miRNA-101
inhibits ovarian cancer cells proliferation and invasion by
down-regulating expression of SOCS-2. Int J Clin Exp Med.
8:20263–20270. 2015.PubMed/NCBI
|
|
31
|
Tsai CH, Yang DY, Lin CY, Chen TM, Tang CH
and Huang YL: Sphingosine-1-phosphate suppresses chondrosarcoma
metastasis by upregulation of tissue inhibitor of metalloproteinase
3 through suppressing miR-101 expression. Mol Oncol. 11:1380–1398.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wang C, Lu S, Jiang J, Jia X, Dong X and
Bu P: Hsa-microRNA-101 suppresses migration and invasion by
targeting Rac1 in thyroid cancer cells. Oncol Lett. 8:1815–1821.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Li JT, Jia LT, Liu NN, Zhu XS, Liu QQ,
Wang XL, Yu F, Liu YL, Yang AG and Gao CF: MiRNA-101 inhibits
breast cancer growth and metastasis by targeting CX chemokine
receptor 7. Oncotarget. 6:30818–30830. 2015.PubMed/NCBI
|
|
34
|
Shaker O, Alhelf M, Morcos G and
Elsharkawy AL: miRNA-101-1 and miRNA-221 expressions and their
polymorphisms as biomarkers for early diagnosis of hepatocellular
carcinoma. Infect Genet Evol. 51:173–181. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Lv P, Zhang P, Li X and Chen Y: Micro
ribonucleic acid (RNA)-101 inhibits cell proliferation and invasion
of lung cancer by regulating cyclooxygenase-2. Thorac Cancer.
6:778–784. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Lei YM, Zu YF, Wang J, Bai S, Shi YF, Shi
R, Duan J, Cui D, Chen J, Xiang Y and Dong J:
Interleukin-1β-mediated suppression of microRNA-101 and
upregulation of enhancer of zeste homolog 2 is involved in
particle-induced lung cancer. Med Oncol. 32:3872015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wan X, Kong Z, Chu K, Yi C, Hu J, Qin R,
Zhao C, Fu F, Wu H, Li Y and Huang Y: Co-expression analysis
revealed PTCH1-3′UTR promoted cell migration and invasion by
activating miR-101-3p/SLC39A6 axis in non-small cell lung cancer:
Implicating the novel function of PTCH1. Oncotarget. 9:4798–4813.
2017.PubMed/NCBI
|
|
38
|
Edge SB and Compton CC: The American Joint
Committee on Cancer: The 7th edition of the AJCC cancer staging
manual and the future of TNM. Ann Surg Oncol. 17:1471–1474. 2010.
View Article : Google Scholar : PubMed/NCBI
|