Introduction
Osteosarcoma primarily impacts children, juveniles
and adults from an early age (1).
Although the worldwide incidence rate of this rare disease is only
3–4 cases per million, osteosarcoma is the most common form of bone
cancer (2). Various chemotherapeutic
and radiation treatments have been developed over the past two
decades; however, the survival rate is still low. Almost 50% of
patients succumb to pulmonary metastasis in the terminal stages of
the disease (3). Elucidating the
underlying molecular mechanisms of osteosarcoma metastasis is
therefore a desirable research outcome.
The epithelial-mesenchymal transition (EMT) is
involved in the process of invasion and metastasis in many cancers,
including osteosarcoma (4,5). Zinc finger E-box binding homeobox 1
(ZEB1), is a key protein that functions to regulate the phenotype
of EMT during cancer progression (6).
ZEB1 may promote prostatic cancer, and overexpression of ZEB1 leads
to the promotion of lung cancer cell metastasis (7,8).
Additionally, the overexpression of ZEB1 has been demonstrated to
be associated with the development, carcinogenesis, invasion and
metastasis of osteosarcoma (9).
Long non-coding RNAs (lncRNAs) are RNA molecules
that have over 200 nucleotides and possess the potential for
protein-coding (10). LncRNAs are
involved in regulating cellular functions and the progression of
various types of cancer (11).
Several studies have demonstrated that lncRNAs serve a critical
function in numerous cellular processes by competing with RNAs to
regulate microRNAs (12–14). The myocardial infarction associated
transcript (MIAT) may be expressed in postmitotic retinal precursor
cells and mitotic progenitors (15).
With regards to human malignancies, MIAT upregulation has been
observed in conditions including neuroendocrine prostatic and
gastric cancer (16,17). However, the role of MIAT in the
regulation of osteosarcoma remains unresolved.
Materials and methods
Tissue samples
Patients with osteosarcoma provided six tissue
samples and were operated on at The First Affiliated Hospital of
Harbin Medical University, China. Samples were snap frozen at −80°C
until RNA extraction. Written informed consent was obtained from
all patients. The study was approved by the Research Ethics
Committee at Harbin Medical University (Harbin, China).
Cell culture and transfection
The Chinese Cell Bank of the Chinese Academy of
Sciences (Shanghai, China) provided the osteoblast cell lines hFOB
(OB3) and osteosarcoma cell lines Saos-2 and MG-63. Dulbecco's
modified Eagle's medium (DMEM; HyClone; GE Healthcare, Chicago, IL,
USA) was used as cell medium at 37°C, with 10% heat-inactivated
fetal bovine serum (FBS; Biological Industries, Kibbutz Beit
Haemek, Israel) and with 95% air and 5% CO2. siRNAs
targeting MIAT (forward, 5′-GGACGTTCACAACCACACTG-3′ and reverse,
5′-TCCCACTTTGGCATTCTAGG-3′) were designed by Guangzhou RiboBio Co.,
Ltd. (Guangzhou, China). Knockdown and overexpression of miR-150-5p
were obtained from Invitrogen (Thermo Fisher Scientific, Inc.,
Waltham, MA, USA). The sequences were as follows: Human miR-150-5p,
5′-UCUCCCAACCCUUGUACCAGUG-3′ and 29-O-methyl modified miR-150
inhibitor, 5′-CACUGGUACAAGGGUUGGGAGA-3′. Cell transfections were
performed using X-tremeGENE siRNA Transfection Reagent (Roche
Diagnostics, Indianapolis, IN, USA) according to the manufacturer's
protocol. siNC (50 µl DMEM was mixed with 20 pmol siNC;
GCACCTTGAGTGAATGTCAGGGACTCCCTGATGATGTGA; Guangzhou RiboBio Co.,
Ltd.) was defined as the negative control.
RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
TRIzol® reagent (Life Technologies;
Thermo Fisher Scientific, Inc.) was used to extract RNA according
to the manufacturers protocol. Invitrogen (Thermo Fisher
Scientific, Inc.) provided the PCR primers. A NanoDrop
Spectrophotometer (NanoDrop Technologies, Wilmington, DE) was used
to measure the concentration of extracted RNA. A TaqMan®
miRNA reverse transcription kit (Applied Biosystems, Foster City,
CA, USA) synthesized cDNA via RT in 5 ng of total RNA to find
miR-150-5p levels. The 2−ΔΔCq method was used to
determine the expression levels of miR-150-5p (18). MIAT expression was divided into high
and low groups using RT-qPCR, using the median value as the cut-off
to differentiate between the high and low groups. Bioinformatics
analysis was used (MicroRNA, Starbase version 2.0) to determine the
potential complementarity between MIAT and miRNAs.
In the synthesis kit of cDNA, RNA synthesized the
cDNAs via specific gene primers. (Invitrogen; Thermo Fisher
Scientific, Inc.) to determine ZEB1 mRNA expression. qPCR was
performed using a SYBR Green Real-Time PCR Master Mix kit (Toyobo
Life Science, Osaka, Japan) according to the manufacturer's
protocol, and the ABI 7500 Sequence Detection System (Life
Technologies; Thermo Fisher Scientific, Inc.). In a total reaction
volume of 20 µl, amplification was performed with 1 µl reverse
primer, 1 µl forward primer, 10 µl SYBR Master Mix, 6 µl diethyl
pyrocarbonate and 2 µl cDNA. The conditions for the reaction were
as follows: 72°C for 45 sec, 60°C for 15 sec and 40 cycles of 95°C
for 15 sec. The internal control was GAPDH. Expression levels of
ZEB1 were determined in relation to GAPDH by the 2−ΔΔCq
method (18). Table I presents the primer sequences.
 | Table I.Primers used for RT-PCR. |
Table I.
Primers used for RT-PCR.
| Name | Sequence (5′-3′) | Length (bp) |
|---|
| GAPDH forward |
AAGAAGGTGGTGAAGCAGGC | 20 |
| GAPDH reverse |
TCCACCACCCTGTTGCTGTA | 20 |
| miR-150-5p
forward |
GTCTCCCAACCCTTGTAC | 18 |
| miR-150-5p
reverse |
TATCCAGTGCGTGTCGTG | 18 |
| ZEB1 forward |
FGCCAATAAGCAAACGATTCTG | 22 |
| ZEB1 reverse |
TTTGGCTGGATCACTTTCAAG | 21 |
| U6 forward |
CTCGCTTCGGCAGCACATATACT | 23 |
| U6 reverse |
ACGCTTCACGAATTTGCGTGTC | 22 |
Western blot analysis
Cells were washed using PBS three times at room
temperature and lysed using radioimmunoprecipitation assay buffer
with 1% protease inhibitor (Sigma-Aldrich; Merck KGaA, Darmstadt,
Germany). The protein concentration was measured by Nanodrop.
Proteins (50 µg) were separated by SDS-PAGE (10% gel). Protein
transfer was then conducted onto nitrocellulose membranes. Blocking
was performed using 0.1% Tween 20 and 5% nonfat milk (BD
Biosciences, Franklin Lakes, NJ, USA) in TBS for 2 h at room
temperature. Samples were incubated with the proper primary
antibodies (cat. no. ab203829; 1:1,000; rabbit; Abcam, Cambridge,
MA, USA) at 4°C overnight with gentle agitation followed by
staining with the fluorochrome-labeled secondary antibody (cat. no.
A10235; 1:8,000; rabbit anti-mouse; Alexa Fluor 800; LI-COR
Biosciences, Lincoln, NE, USA) at room temperature for 1 h. The
Odyssey fluorescent scanning system (LI-COR Biosciences) detected
immunoreactivity and Image Studio software 4.0 (LI-COR Biosciences)
was used to examine captured images. The loading control was
β-actin.
Cell proliferation assay
The cell counting kit-8 kits (CCK-8) were used
according to the manufacturer's instructions. MG63 and Saos-2 cells
were seeded in 96-well plates at 1×104 cells/well for 24
h. CCK-8 solution (10 µl) was added to each well, and the cells
were incubated at 37°C for 2 h. Absorbance at 450 nm was measured
using a microplate reader. The assay was conducted in
triplicate.
Wound healing assays
Osteosarcoma cells (MG63 and Saos-2 cells) were
seeded in six-well plates and allowed to reach 80–90% confluence. A
wound line was drawn across the surface of the plates using a
200-µl sterile plastic tip. PBS was used to wash the plates. Images
were captured 24 h post wound infliction. Each test was conducted
in triplicate.
Transwell assays
Transwell filters (8 µm pore size; BD Biosciences)
were placed on a 24-well plate containing DMEM/F12 (Hyclone: GE
Healthcare). The medium in the upper membrane was serum free, and
in the lower chamber contained 10% FBS (Biological Industries,
Beit-Haemek, Israel).
MG63 and Saos-2 cells were suspended in DMEM/F12 at
a cell density of 2.5×105 cells/ml for 24 h. After 24 h,
cells present on the top of the membrane were cleared using a
cotton swab. Cells present on the bottom portion of the membrane
were fixed using 4% paraformaldehyde in PBS for 10 min and stained
using crystal violet at room temperature for 15 min. Cell invasion
was quantified as the average number of cells from 3 inserts
present on the bottom portion of the membrane.
Statistical analysis
All data are expressed as the mean ± standard error
of the mean. SPSS software (version 13.0; SPSS, Inc., Chicago, IL,
USA) was used to analyze all data. Statistical comparison of two
groups was performed using a Student's t-test. One-way analysis of
variance was also used to compare ≥2 groups. P<0.05 was
considered to indicate a statistically significant difference.
Results
MIAT levels are elevated in
osteosarcoma
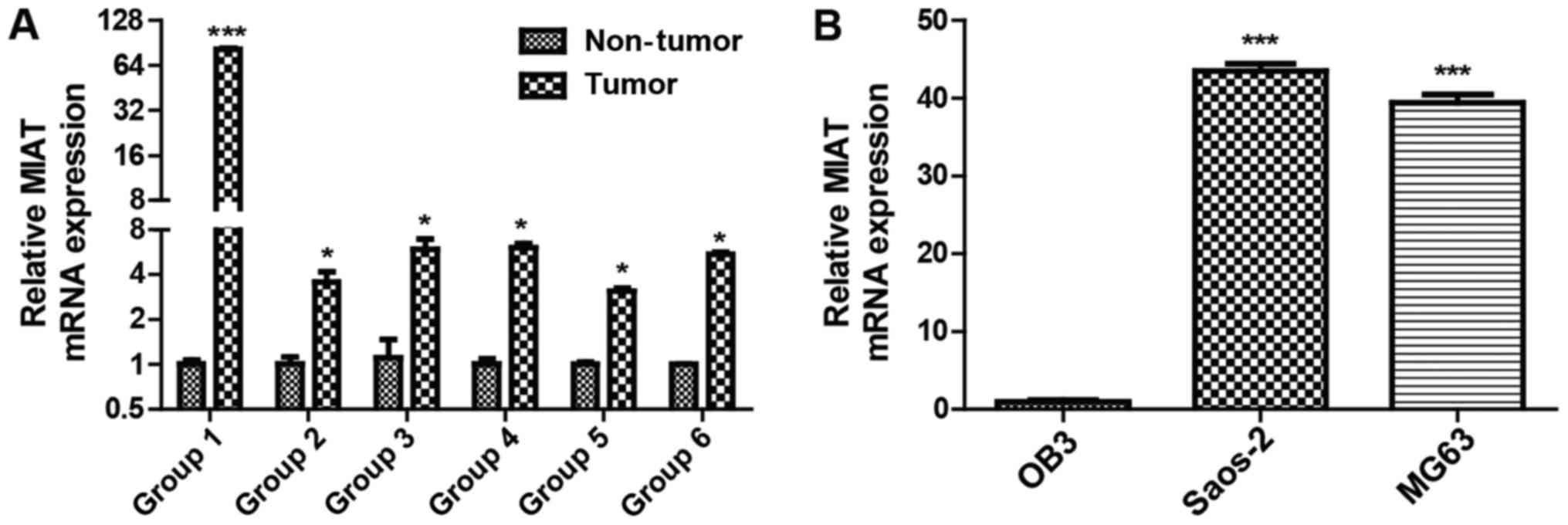
The expression levels of the lncRNA MIAT 6 were
investigated in groups of osteosarcoma tissues using RT-qPCR. MIAT
levels were decreased in adjacent non-tumor tissues compared with
osteosarcoma tissues (Fig. 1A). The
lncRNA MIAT expression levels were also investigated in MG63,
Saos-2 and OB3 cells and it was identified that MIAT expression was
lower in the OB3 cell line than in Saos-2 and MG63 cells (Fig. 1B). These results demonstrate that MIAT
levels are potentially associated with osteosarcoma.
MIAT knockdown inhibits the
proliferation and invasion of osteosarcoma cells
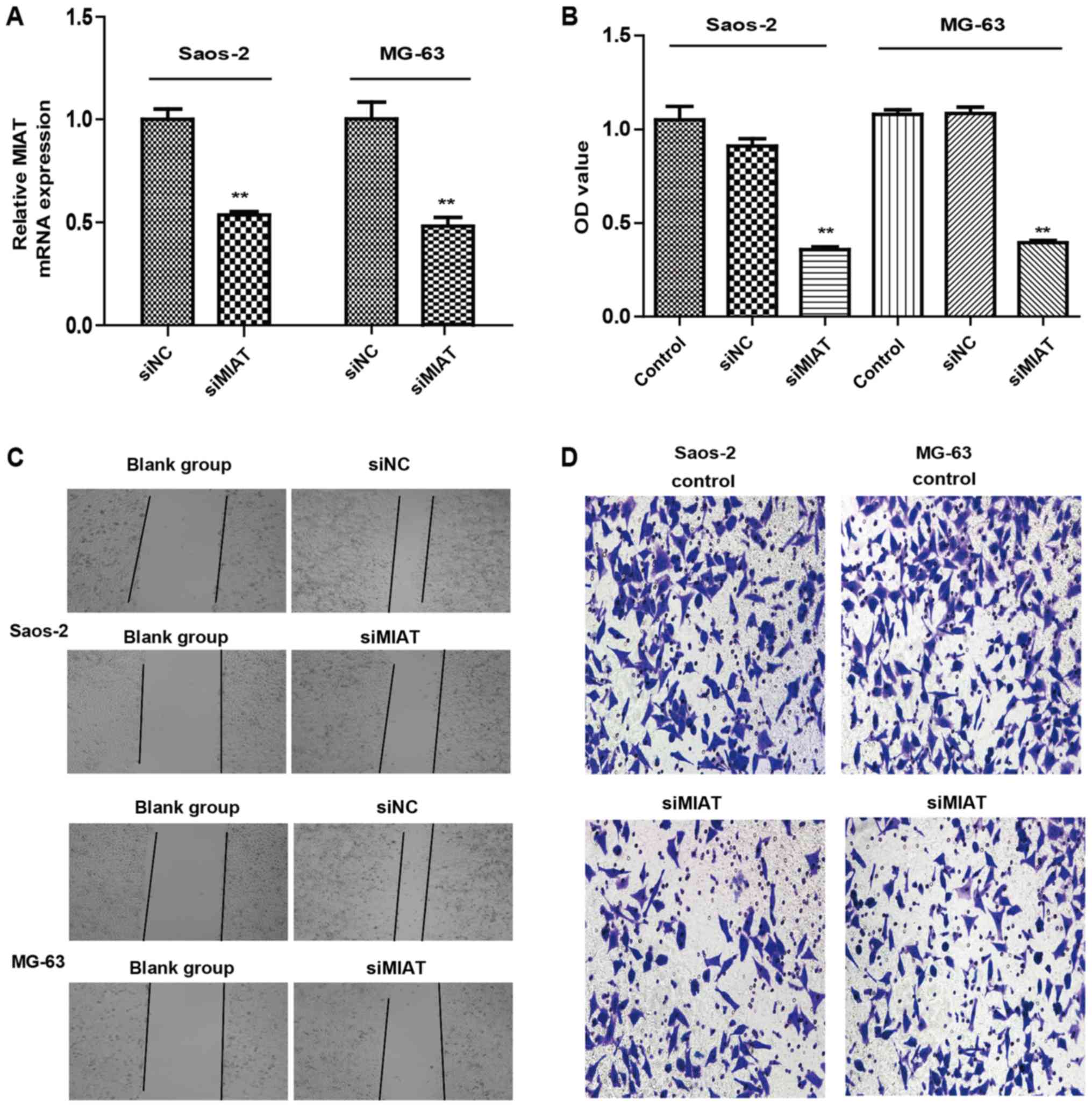
The function of MIAT in the proliferation and
invasion of osteosarcoma cells was examined. Transfection of MG63
and Saos-2 cells with MIAT siRNAs was demonstrated to lower MIAT
expression in comparison to control cells (Fig. 2A). CCK8 results indicated that
knockdown of MIAT reduced MG63 and Saos-2 cell proliferation
compared to cells transfected with siRNAs (Fig. 2B). MIAT knockdown also limited the
proliferation and invasion of MG63 and Saos-2 cells (Fig. 2C and D) compared to cells transfected
with siRNAs. Taken together, these data reveal that MIAT may
promote proliferation and invasion of osteosarcoma cells in
vitro.
MIAT promotes ZEB1 expression in
osteosarcoma
ZEB1 is important for the proliferation and invasion
of osteosarcoma cells (19). Thus, it
was investigated whether MIAT affects ZEB1 expression in
osteosarcoma cells. First, the association between MIAT and the
ZEB1 expression levels was evaluated in 6 samples of tumor-adjacent
tissue and osteosarcoma samples by western blotting and
RT-qPCR.
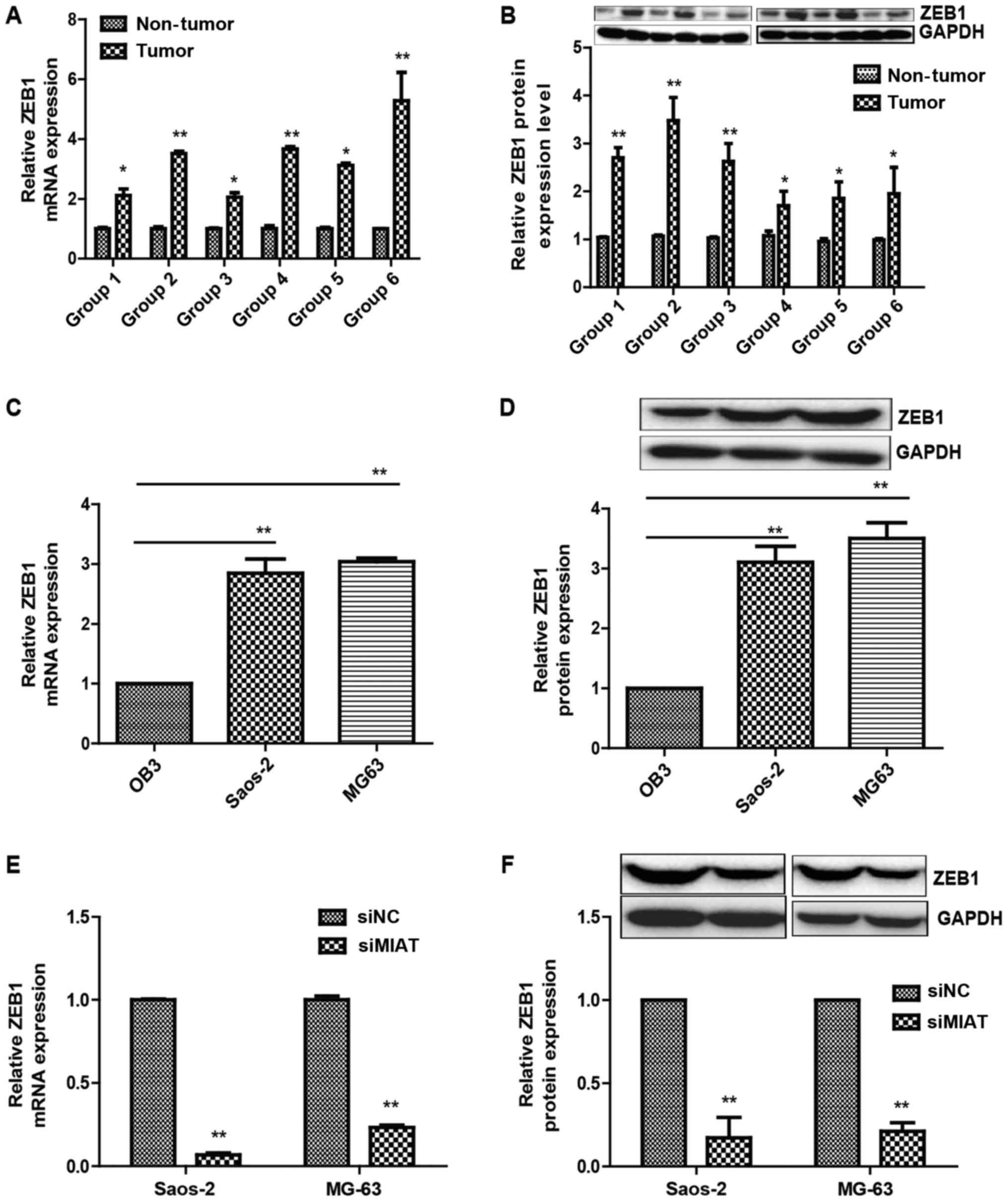
It was identified that ZEB1 expression was notably
increased in the high MIAT osteosarcoma tissue group compared with
that in the low MIAT group (Fig. 3A and
B). It was additionally noted that ZEB1 expression levels were
decreased in OB3 cells compared with MG63 and Saos-2 cells
(Fig. 3C and D). Furthermore, ZEB1
expression levels were evaluated in Saos-2 and MG63 cells
transfected with MIAT siRNAs or siNC (negative control) and it was
identified that MIAT knockdown by siRNAs resulted in lower ZEB1
expression compared to cells transfected with control siRNAs
(Fig. 3E and F).
miR-150-5p is a downstream target of
MIAT
To identify the potential downstream miRNA targets
of MIAT and its interactions in osteosarcoma, bioinformatics
analysis was used (MicroRNA, Starbase version 2.0) to determine the
potential complementarity between MIAT and miRNAs. Bioinformatics
predictions revealed that the MIAT sequence has four putative miRNA
binding sites, including sites for miR-29a-3p, miR-29b-3p,
miR-29c-3p, and miR-150-5p-5p. Yan et al (20) previously reported that miR-150-5p-5p
focuses on MIAT in endothelial cells, and another study revealed
that miR-150-5p suppresses ZEB1 in epithelial ovarian cancer
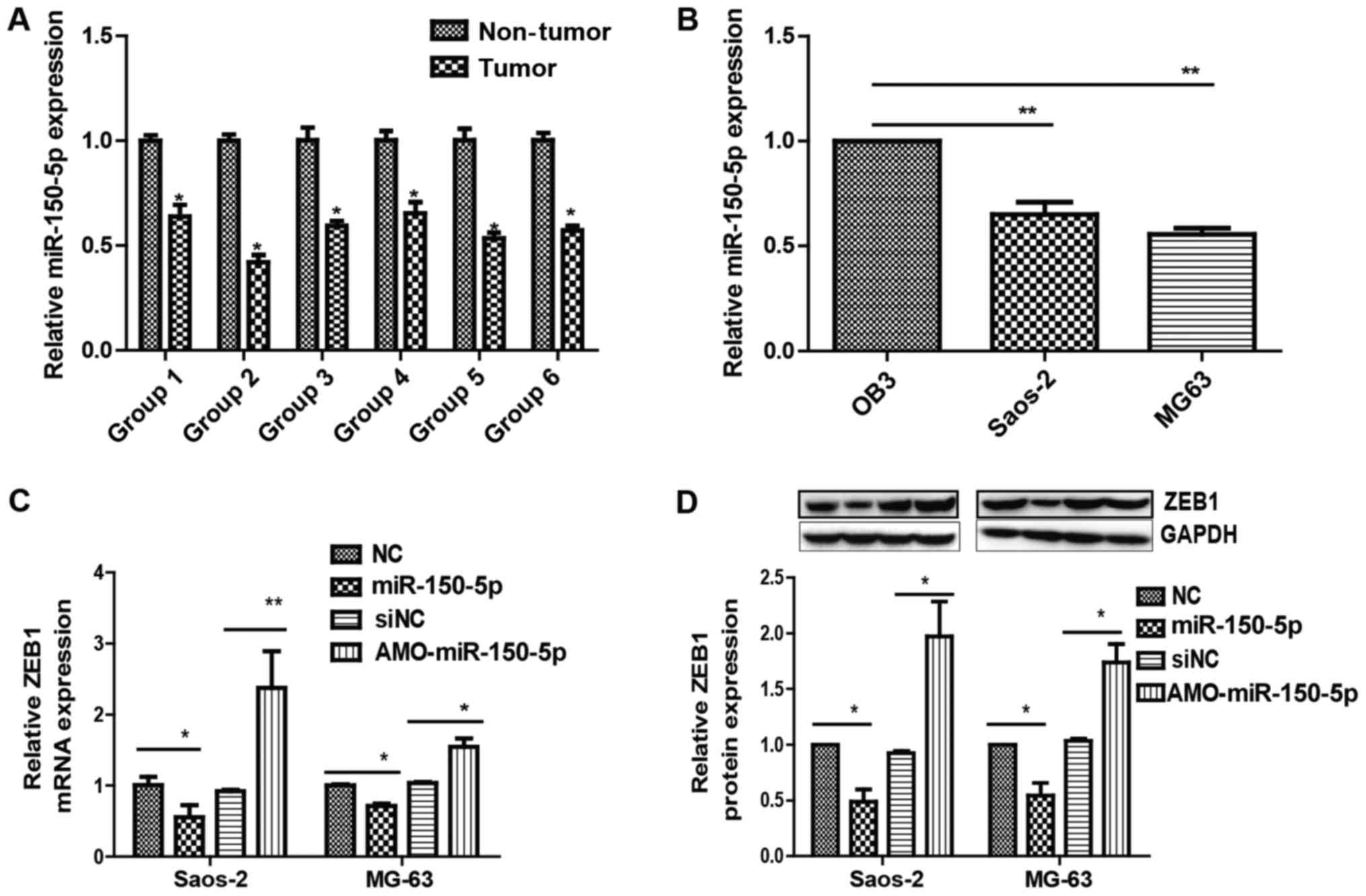
(21). It may be observed that
miR-150-5p levels were higher in adjacent non-tumor tissues than in
osteosarcoma tissues (Fig. 4A) and
lower in MG63 and Saos-2 cells than OB3 cells (Fig. 4B).
To determine whether miR-150-5p targets MIAT,
miR-150-5p expression in Saos-2 and MG63 cells transfected with
MIAT-siRNA or siNC was examined. The results revealed that
miR-150-5p expression was visibly elevated in MG63 and Saos-2 cells
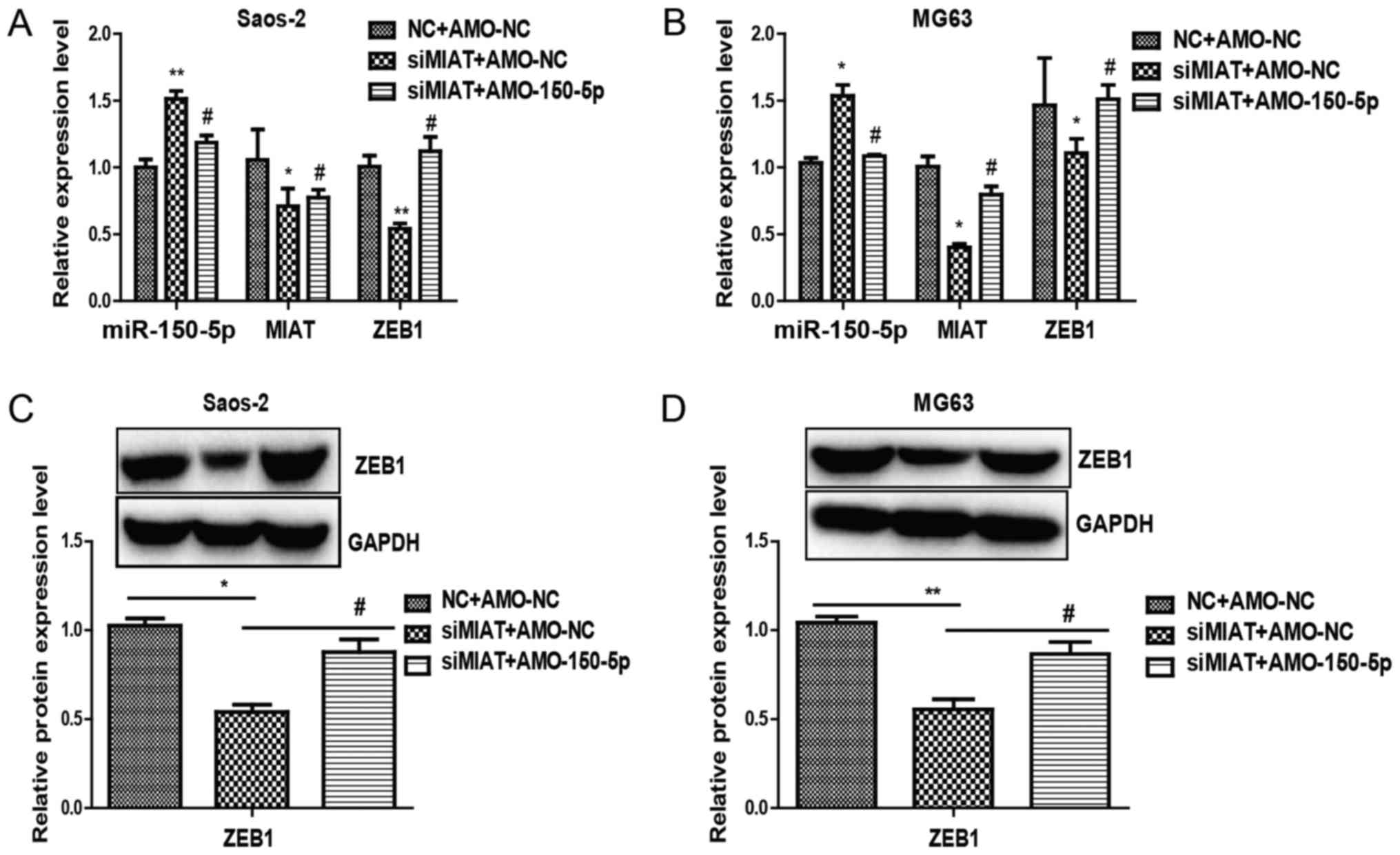
transfected with MIAT-siRNA compared to control siRNA (Fig. 5A and B). Rescue experiments were
subsequently performed by transfecting miR-150-5p in Saos-2 and
MG63 cells. Overexpressing miR-150-5p led to an increase in ZEB1
(Fig. 4C and D). Additionally,
inhibiting MIAT limited ZEB1 levels by inhibiting miR-150-5p
(Fig. 5A and B). Together, the
results demonstrate the importance of MIAT in regulating ZEB1, by
controlling miR-150-5p.
miR-150-5p reverses the effects of
MIAT in osteosarcoma cells
These results demonstrated that miR-150-5p is a
downstream target of MIAT. However, the function of miR-150-5p in
the MIAT-mediated influence on osteosarcoma cells remains unclear.
To determine whether MIAT may promote proliferation and invasion of
osteosarcoma cells via the MIAT-miR-150-5p-ZEB1 axis, further
experiments were carried out. RT-qPCR and western blotting revealed
that reduced ZEB1 expression by inhibition of MIAT could be largely
reversed by AMO-miR-150-5p (Fig. 5).
Together, these results suggest that miR-150-5p was able to change
the function of MIAT in osteosarcoma cells and that MIAT was able
to promote the proliferation and invasion of osteosarcoma cells via
the MIAT-miR-150-5p-ZEB1 axis.
Discussion
Osteosarcoma has been considered a common type of
basic malignancy of bone and is derived from the progenitor
mesenchymal cells of bone-forming cells (22). The morbidity associated with
osteosarcoma is high since early diagnosis is difficult, and
therapeutic solutions to osteosarcoma are lacking. Therefore, it is
important to identify new molecules associated with developing
osteosarcomas and develop novel targeted therapy strategies.
Previous studies have demonstrated the role of
lncRNAs (23) and the molecular
mechanisms through which lncRNAs affect human tumors (24–26).
However, the mechanism involving the lncRNA MIAT in osteosarcoma
has remained unknown. This research shows that highly overexpressed
MIAT in cell lines and osteosarcoma tissues may have a monogenic
role.
miR-150-5p was first considered as the main miRNA in
immune and hematopoietic cells (27).
Research has recently shown that specific cellular functions in
diverse tumors also involve miR-150-5p (28–30). A
previous study demonstrated that expression of miR-150-5p was
decreased in osteosarcoma cells compared to analogous human normal
osteoblasts and cells from normal tissues (31).
MIAT was identified as a target of miR-150-5p with
both MIAT and miR-150-5p having an inhibitory effect. These results
demonstrate that MIAT may improve tumor progression in osteosarcoma
since it can inhibit miR-150-5p and activate ZEB1. The present
study demonstrated that in osteosarcoma tissues, MIAT expression
was increased compared with the adjacent normal tissues. The
present study also identifies that MIAT is important in the
development and progression of osteosarcoma. However, the
mechanisms associated with MIAT-mediated gene expression in
tumorigenesis need to be clarified.
Research has revealed that MIAT acts as a molecular
sponge by managing microRNAs in the progression of breast cancer
(32). It has been reported that
during specific cellular processes, lncRNAs are able to compete
with endogenous RNAs to manage microRNAs (33). MIAT may therefore be considered as an
endogenous miRNA which controls miR-150-5p-5p and manages its role.
Although several potential miRNA binding partners were identified,
the present study focused on miR-150-5p, as it has been
demonstrated to be important in numerous cancers, including lung
(34) and liver cancer (35). The present study revealed that
miR-150-5p levels were decreased in the osteosarcoma cell lines
MG63 and Saos-2 cells. An opposing association was observed between
miR-150-5p and MIAT levels in Saos-2 and MG63 cells, suggesting
that there may be an association between miR-150-5p and MIAT to
control osteosarcoma cell proliferation and invasion. In addition,
miR-150-5p expression decreased ZEB1 levels in Saos-2 cells and
MG63. Furthermore, the present data suggested that the MIAT
sequence had miR-150-5p sites for binding and implied that
miR-150-5p reduced MIAT level by directly binding MIAT. ZEB1 is a
master regulator of the EMT phenotype within the progression of
cancer (36). The present study
demonstrates that by the miR-150-5p/ZEB1 pathway, MIAT may induce
EMT phenotype in osteosarcoma cells.
To conclude, the present study revealed that MIAT
may be a biomarker for patients with osteosarcoma. It is
hypothesized that the MIAT-miR-150-5p-ZEB1 axis may be a potential
therapeutic target in osteosarcoma.
Acknowledgements
Not applicable.
Funding
This study was financially supported by Heilongjiang
provincial academy of medical sciences (grant no. 201618).
Availability of data and materials
The analyzed datasets generated during the study are
available from the corresponding author on reasonable request.
Authors' contributions
HJ performed the flow cytometric analysis and
drafted the manuscript. XJ performed cell culture and viral
preparation experiments. WC and FD contributed to statistical
analyses. ZY, YL and WW designed the study.
Ethics approval and consent to
participate
Written informed consent was obtained from all
patients. The study was approved by the Research Ethics Committee
at Harbin Medical University.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Liu C and Lin J: Long noncoding RNA
ZEB1-AS1 acts as an oncogene in osteosarcoma by epigenetically
activating ZEB1. Am J Transl Res. 8:4095–4105. 2016.PubMed/NCBI
|
|
2
|
Mirabello L, Troisi RJ and Savage SA:
Osteosarcoma incidence and survival rates from 1973 to 2004: Data
from the surveillance, epidemiology, and end results program.
Cancer. 115:1531–1543. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Daw NC, Chou AJ, Jaffe N, Rao BN, Billups
CA, Rodriguez-Galindo C, Meyers PA and Huh WW: Recurrent
osteosarcoma with a single pulmonary metastasis: A
multiinstitutional review. Br J Cancer. 112:278–282. 2012.
View Article : Google Scholar
|
|
4
|
Yu L, Liu S, Guo W, Zhang C, Zhang B, Yan
H and Wu Z: hTERT promoter activity identifies osteosarcoma cells
with increased EMT characteristics. Oncol Lett. 7:239–244. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Sung JY, Park SY, Kim JH, Kang HG, Yoon
JH, Na YS, Kim YN and Park BK: Interferon consensus
sequence-binding protein (ICSBP) promotes epithelial-to-mesenchymal
transition (EMT)-like phenomena, cell-motility, and invasion via
TGF-β signaling in U2OS cells. Cell Death Dis. 5:e12242014.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Al-Khalaf HH and Aboussekhra A:
MicroRNA-141 and microRNA-146b-5p inhibit the prometastatic
mesenchymal characteristics through the RNA-binding protein AUF1
targeting the transcription factor ZEB1 and the protein kinase AKT.
J Biol Chem. 289:31433–31447. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Putzke AP, Ventura AP, Bailey AM, Akture
C, Opoku-Ansah J, Celiktaş M, Hwang MS, Darling DS, Coleman IM,
Nelson PS, et al: Metastatic progression of prostate cancer and
e-cadherin regulation by zeb1 and SRC family kinases. Am J Pathol.
179:400–410. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Roy BC, Kohno T, Iwakawa R, Moriguchi T,
Kiyono T, Morishita K, Sanchez-Cespedes M, Akiyama T and Yokota J:
Involvement of LKB1 in epithelial-mesenchymal transition (EMT) of
human lung cancer cells. Lung Cancer. 70:136–145. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shen A, Zhang Y, Yang H, Xu R and Huang G:
Overexpression of ZEB1 relates to metastasis and invasion in
osteosarcoma. J Surg Oncol. 105:830–834. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Nagano T and Fraser P: No-nonsense
functions for long noncoding RNAs. Cell. 145:178–181. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen LL and Zhao JC: Functional analysis
of long noncoding RNAs in development and disease. Adv Exp Med
Biol. 825:129–158. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhu HY, Bai WD, Li C, Zheng Z, Guan H, Liu
JQ, Yang XK, Han SC, Gao JX, Wang HT and Hu DH: Knockdown of
lncRNA-ATB suppresses autocrine secretion of TGF-β2 by targeting
ZNF217 via miR-200c in keloid fibroblasts. Sci Rep. 6:247282016.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wu X, He X, Li S, Xu X, Chen X and Zhu H:
Long non-coding RNA ucoo2kmd.1 regulates CD44-dependent cell growth
by competing for miR-211-3p in colorectal cancer. PLoS One.
11:e01512872016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang GQ, Wang Y, Xiong Y, Chen XC, Ma ML,
Cai R, Gao Y, Sun YM, Yang GS and Pang WJ: Sirt1 AS lncRNA
interacts with its mRNA to inhibit muscle formation by attenuating
function of miR-34a. Sci Rep. 6:218652016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ishii N, Ozaki K, Sato H, Mizuno H, Saito
S, Takahashi A, Miyamoto Y, Ikegawa S, Kamatani N, Hori M, et al:
Identification of a novel non-coding RNA, MIAT, that confers risk
of myocardial infarction. J Hum Genet. 51:1087–1099. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Crea F, Venalainen E, Ci X, Cheng H, Pikor
L, Parolia A, Xue H, Saidy Nur NR, Lin D, Lam W, et al: The role of
epigenetics and long noncoding RNA MIAT in neuroendocrine prostate
cancer. Epigenomics. 8:721–731. 2016. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sha M, Lin M, Wang J, Ye J, Xu J, Xu N and
Huang J: Long non-coding RNA MIAT promotes gastric cancer growth
and metastasis through regulation of miR-141/DDX5 pathway. J Exp
Clin Cancer Res. 37:582018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yan H, Zhang B, Fang C and Chen L: miR-340
alleviates chemoresistance of osteosarcoma cells by targeting ZEB1.
Anticancer Drugs. 29:440–448. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yan B, Yao J, Liu JY, Li XM, Wang XQ, Li
YJ, Tao ZF, Song YC, Chen Q and Jiang Q: LncRNA-MIAT regulates
microvascular dysfunction by functioning as a competing endogenous
RNA. Circ Res. 116:1143–1156. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Jin M, Yang Z, Ye W, Xu H and Hua X:
MicroRNA-150 predicts a favorable prognosis in patients with
epithelial ovarian cancer, and inhibits cell invasion and
metastasis by suppressing transcriptional repressor ZEB1. PLoS One.
9:e1039652014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Lulla RR, Costa FF, Bischof JM, Chou PM,
de F Bonaldo M, Vanin EF and Soares MB: Identification of
differentially expressed MicroRNAs in osteosarcoma. Sarcoma.
2011:7326902011. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Orom UA, Derrien T, Beringer M, Gumireddy
K, Gardini A, Bussotti G, Lai F, Zytnicki M, Notredame C, Huang Q,
et al: Long noncoding RNAs with enhancer-like function in human
cells. Cell. 143:46–58. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Barnhill LM, Williams RT, Cohen O, Kim Y,
Batova A, Mielke JA, Messer K, Pu M, Bao L, Yu AL and Diccianni MB:
High expression of CAI2, a 9p21-embedded long noncoding RNA,
contributes to advanced-stage neuroblastoma. Cancer Res.
74:3753–3763. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Nie FQ, Zhu Q, Xu TP, Zou YF, Xie M, Sun
M, Xia R and Lu KH: Long non-coding RNA MVIH indicates a poor
prognosis for non-small cell lung cancer and promotes cell
proliferation and invasion. Tumour Biol. 35:7587–7594. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhen L, Yun-Hui L, Hong-Yu D, Jun M and
Yi-Long Y: Long noncoding RNA NEAT1 promotes glioma pathogenesis by
regulating miR-449b-5p/c-Met axis. Tumour Biol. 37:673–683. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
He Y, Jiang X and Chen J: The role of
miR-150-5p in normal and malignant hematopoiesis. Oncogene.
33:3887–3893. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang X, Ren Y, Wang Z, Xiong X, Han S, Pan
W, Chen H, Zhou L, Zhou C, Yuan Q and Yang M: Down-regulation of 5S
rRNA by miR-150-5p and miR-383 enhances c-Myc-rpL11 interaction and
inhibits proliferation of esophageal squamous carcinoma cells. FEBS
Lett. 589:3989–3997. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Liu DZ, Zhang HY, Long XL, Zou SL, Zhang
XY, Han GY and Cui ZG: MIR-150-5P promotes prostate cancer stem
cell development via suppressing p27Kip1. Eur Rev Med Pharmacol
Sci. 19:4344–4352. 2015.PubMed/NCBI
|
|
30
|
Li J, Hu L, Tian C, Lu F, Wu J and Liu L:
microRNA-150 promotes cervical cancer cell growth and survival by
targeting FOXO4. BMC Mol Biol. 16:242015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Qu Y, Pan S, Kang M, Dong R and Zhao J:
MicroRNA-150 functions as a tumor suppressor in osteosarcoma by
targeting IGF2BP1. Tumor Biol. 37:5275–5284. 2016. View Article : Google Scholar
|
|
32
|
Luan T, Zhang X, Wang S, Wang S, Song Y,
Zhou S, Lin J, An W, Yuan W, Yang Y, et al: Long non-coding RNA
MIAT promotes breast cancer progression and functions as ceRNA to
regulate DUSP7 expression by sponging miR-155-5p. Oncotarget.
8:76153–76164. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Mercer TR, Dinger ME and Mattick JS: Long
non-coding RNAs: Insights into functions. Nat Rev Genet.
10:155–159. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
34
|
Cao M, Hou D, Liang H, Gong F, Wang Y, Yan
X, Jiang X, Wang C, Zhang J, Zen K, et al: miR-150-5p promotes the
proliferation and proliferation of lung cancer cells by targeting
SRC kinase signalling inhibitor 1. Eur J Cancer. 50:1013–1024.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li T, Xie J, Shen C, Cheng D, Shi Y, Wu Z,
Zhan Q, Deng X, Chen H, Shen B, et al: miR-150-5p-5p inhibits
hepatoma cell proliferation and invasion by targeting MMP14. PLoS
One. 9:e1155772014. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Preca BT, Bajdak K, Mock K, Sundararajan
V, Pfannstiel J, Maurer J, Wellner U, Hopt UT, Brummer T, Brabletz
S, et al: A self-enforcing CD44S/ZEB1 feedback loop maintains EMT
and stemness properties in cancer cells. Int J Cancer.
137:2566–2577. 2015. View Article : Google Scholar : PubMed/NCBI
|