Introduction
Prostate cancer is one of the most frequent causes
of cancer-associated mortality and diagnosed malignancies in the
male population. In 2012, 238,590 new patients were diagnosed with
prostate cancer and 29,720 mortalities were attributed to the
disease in the USA (1). In
early-stage prostate cancer, patients may be cured using radical
prostatectomy, whereas it remains a challenge to cure patients with
metastatic disease. In the early stages, metastatic prostate cancer
is sensitive to androgen ablation therapy; however, the majority of
cases progress to an androgen-independent stage (2), which is a primary cause for prostate
cancer-associated mortality. Although the American Food and Drug
Administration have approved several therapeutic agents for the
treatment of metastatic prostate cancer, these therapies have not
been proven to prolong the survival of patients beyond two years
from the start of treatment.
In the majority of cancer types, apoptosis is
inhibited in tumor tissue and the activation of apoptosis signaling
pathways is a major underlying mechanism by which cytotoxic drugs
are able to kill tumor cells (3).
Signal transducer and activator of transcription 6 (STAT6) is a
member of the STAT family of latent transcription factors and is
able to be stimulated using cytokines, hormones and growth factors
that are hypothesized to serve an important role in
androgen-independence (4). In a
previous study (5), the knockdown of
STAT6 in prostate cancer cells promoted apoptosis and it was
therefore hypothesized that that STAT6 may be a potential
therapeutic target for prostate cancer. However, the underlying
mechanisms are yet to be elucidated.
The 3′ untranslated region (3′UTR) of target
transcripts is bound by microRNAs (miRNAs), which regulate gene
expression via inhibition of translation or promotion of mRNA
degradation. miRNAs are involved in numerous biological processes,
including apoptosis. A particular association has been established
between the aberrant expression levels of miRNAs and the
development, progression and chemotherapy resistance of tumors. The
expression of a number of miRNAs has been identified to be
upregulated in prostate cancer, including miRNA (miR)-21, miR-125b
and miR-221/222 (6). By contrast,
miR-7 (7), miR-143/145 (8) and miR-146a (9) have been identified as tumor suppressors.
miR-135a is an androgen-regulated miRNA, which is associated with
prostate cancer cell migration and invasion. Analyses from the Gene
Series (GSE40026) dataset demonstrated that miR-135a has reduced
expression levels, particularly in androgen-independent cells
(10). In the present study, miR-135a
functioned as a tumor suppressor in androgen receptor-negative
prostate cancer cells by promoting apoptosis via direct targeting
of STAT6 mRNA. This inhibited the protein expression and
phosphorylation of STAT6 and limited its DNA binding ability, which
affected its function as a transcriptional factor. Additionally,
low levels of STAT6 in prostate cancer tissue were associated with
an increased overall survival rate and a reduced biochemical
relapse rate in the ONCOMINE® database.
Materials and methods
Retrieval of the ONCOMINE®
database and GSE40026 and GSE21032 datasets
Microarray datasets for prostate cancer were
retrieved from the ONCOMINE® Cancer Profiling Database
(www.oncomine.org) to investigate STAT6 expression
levels including Wallace, Grasso and Lapointe datasets. The
co-expression data of miR-135a and STAT6 in prostate cancer cell
lines was retrieved from the GSE40026 (10) and GSE21032 (11) datasets in the Gene Expression Omnibus
database.
Gene Set Enrichment Analysis (GSEA)
with STAT6 expression levels
The Cancer Genome Atlas (TCGA) and Memorial
Sloan-Kettering Cancer Center (MSKCC) prostate cancer databases
were used to generate STAT6 positively- and negatively-associated
gene sets. The Pearson score was calculated for STAT6-correlated
genes and >+0.3 was recognized as positively correlated while
<-0.3 was considered negatively correlated. The pre-ranked gene
sets were deposited into GSEA (12).
Cell culture and transfection
The DU145 and PC3 human prostate cancer cell lines
were obtained from the American Type Culture Collection (ATCC) and
grown according to the ATCC protocol. Briefly, the cell lines were
cultured in Dulbecco's modified Eagle's medium or F12K medium
supplemented respectively with 50 U/ml penicillin, 50 mg/ml
streptomycin (Gibco; Thermo Fisher Scientific, Inc., Waltham, MA,
USA) and 10% fetal bovine serum (FBS; Hangzhou Sijiqing Biological
Engineering Materials Co., Ltd., Hangzhou, China) at 37°C in an
atmosphere of 5% CO2. Transfections with miR-135a and
negative control (NC) mimics (Shanghai GenePharma Co., Ltd.,
Shanghai, China) were performed at a concentration of 40 nM
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
Hoechst33342 staining
The DU145 and PC3 cells were cultured on slides in a
six-well plate. Following transfection for 48 h, the cells were
washed with PBS and stained with Hoechst33342 (1 mg/ml; Beyotime
Institute of Biotechnology, Haimen, China) for 5 min at 37°C.
Following a second wash with PBS, images of the cells were captured
and counted using immunofluorescence microscopy (Olympus
Corporation, Tokyo, Japan).
Quantification of apoptosis using
fluorescence-activated cell sorting (FACS)
To determine the levels of apoptosis following
treatment with miR-135a or NC, an Annexin-V (AV) and propidium
iodide (PI; Beyotime Institute of Biotechnology) stain was
performed. Briefly, DU145 and PC3 cells were harvested 72 h
post-transfection, washed in PBS and suspended in binding buffer
(Beyotime Institute of Biotechnology) and cells were serum-deprived
for 24 h prior to FACS. Following this, the cells were stained with
AV and PI (50 µM) and incubated at room temperature for 15 min. A
total of 10,000 events were recorded using a flow cytometer (BD
Biosciences, Franklin Lakes, NJ, USA) and the proportion of
apoptotic cells was analyzed.
Western blotting
DU145 and PC3 cells were transfected with miR-135a
or NC after 72 h prior to being washed with cold PBS and lysed in
RIPA buffer (Beyotime Institute of Biotechnology) on ice. The
protein concentration was determined by bicinchoninic acid protein
assay kit (Beyotime Institute of Biotechnology). The western blot
protocol was performed as described in previous studies (13,14).
Proteins were probed with anti-STAT6 (catalog no. 9362, dilution,
1:500), anti-phosphorylated (p)-STAT6 (Tyr641) (catalog no. 9361,
dilution, 1:500) and anti-GAPDH (catalog no. 2118, dilution,
1:2,000) (Cell Signaling Technology, Inc., Danvers, MA, USA). The
expression of p-STAT6 was induced by IL-4 (100 ng/ml).
Electrophoretic mobility shift assay
(EMSA)
An EMSA was conducted to evaluate the DNA-binding
activity of STAT6 in miR-135a-transfected or NC-transfected cells.
Briefly, following transfection of DU145 and PC3 cells for 72 h,
nuclear proteins from each sample were extracted with Nuclear
Extraction kit (Merck KGaA, Darmstadt, Germany) and subjected to
EMSA following the manufacturer's standard protocol using the
LightShift® Chemiluminescent EMSA kit (Thermo Fisher
Scientific, Inc.). The STAT6 target probe was synthesized with a
3′-biotin modification (Invitrogen; Thermo Fisher Scientific, Inc.)
and the sequence was 5′-acg ttg aat TTCTAAGAA aga ggg a-3′, in
which the binding region is underlined.
Luciferase assay
The 3′UTR of STAT6 was amplified using polymerase
chain reaction and the primer sequences were as follows: Sense,
5′-CCGACGCGTTCTTTTCTGTTGCCCCATC-3′; and antisense,
5′-CCCAAGCTTGGACCGCAGCACCTATCT-3′. The fragment was cloned
downstream of the luciferase gene in a pMIR-REPORT™ Luciferase
miRNA Expression Reporter vector (Ambion®; Thermo Fisher
Scientific, Inc.) The pMIR-Luciferase NC vector or gene-specific
vector was co-transfected with miR-135a or NC into DU145 and PC3
cells. pRL Renilla luciferase control reporter vector (Promega
Corporation, Madison, WI, USA) was also transfected with each
condition as a reference control with Lipofectamine®
2000 (Invitrogen; Thermo Fisher Scientific, Inc.). A total of 72 h
after transfection, the firefly luciferase and Renilla luciferase
activities were measured using the Dual-Luciferase®
Reporter assay system (Promega Corporation).
Bioinformatics and statistical
analysis
The online miRNA databases TargetScan 5.1 (15) and RNAhybrid (16) were used to determine the predicted
miR-135a target genes. Statistical analyses were performed using
SPSS software version 16.0 (SPSS, Inc., Chicago, IL, USA). The
aforementioned experiments were repeated three times. Data were
presented as the mean ± standard deviation. Group means were
compared using the Student's t-test. Survival data was analyzed
using the Kaplan-Meier estimate and log-rank P was calculated.
P<0.05 was considered to indicate a statistically significant
difference.
Results
STAT6 expression levels and its
prognostic role in prostate cancer in the ONCOMINE®
prostate database and co-expression of miR-135a and STAT6 in
prostate cancer cell lines
The ONCOMINE® prostate cancer database is
an established database containing several high-quality datasets.
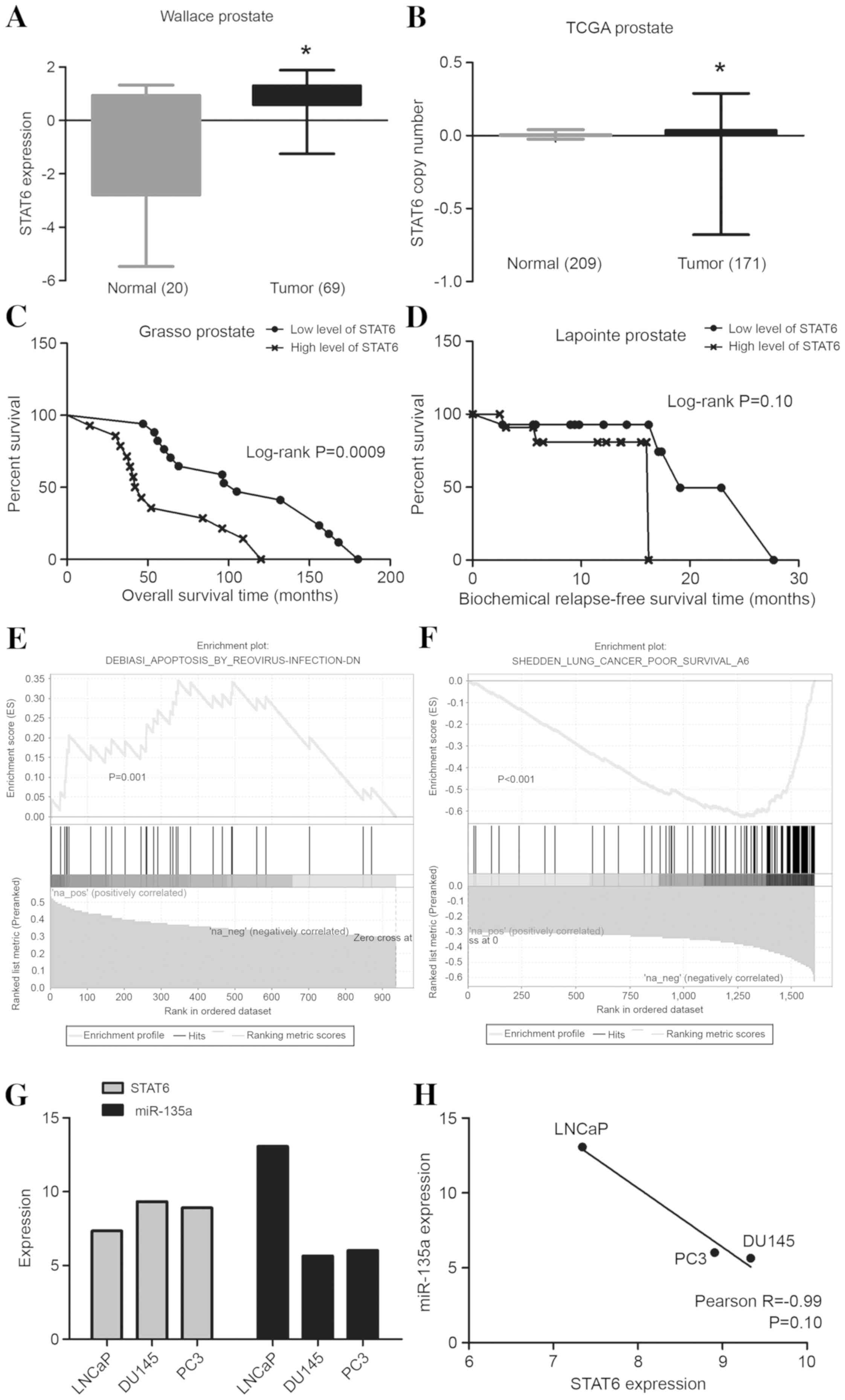
In the Wallace prostate cancer dataset, STAT6 expression levels in
tumor tissues were higher compared with those in normal prostate
gland tissues (P<0.0001; Fig. 1A)
and tumors exhibited a higher STAT6 copy number in the TCGA dataset
(P=0.0002; Fig. 1B). The Grasso and
Lapointe prostate cancer datasets contain data on the overall
survival time of patients with metastatic cancer and the
biochemical relapse-free survival time, which are important events
in the progression of prostate cancer. An increased level of STAT6
expression was associated with a significantly reduced overall
survival rate in Grasso's study (P=0.0009; Fig. 1C) and also an increased biochemical
relapse rate (P=0.1000; Fig. 1D)
following radical prostatectomy in Lapointe's study, but this was
not statistically significant. Additionally, in the TCGA and MSKCC
databases, STAT6-correlated genes were involved in cell apoptosis
signaling pathways (P=0.001; Fig. 1E)
and cancer survival (P<0.001; Fig.
1F).
To investigate the co-expression of miR-135a and
STAT6 in DU145, PC3 and LNCaP prostate cancer cell lines,
microarray data from the GSE40026 and GSE21032 datasets were
evaluated and the results revealed that miR-135a was underexpressed
and STAT6 was overexpressed in DU145 and PC3 cells compared with
LNCaP cells. This indicated an inverse correlation between miR-135a
and STAT6 expression in these cell lines (P=0.10; Fig. 1G and H).
miR-135a increases apoptosis in
prostate cancer cells
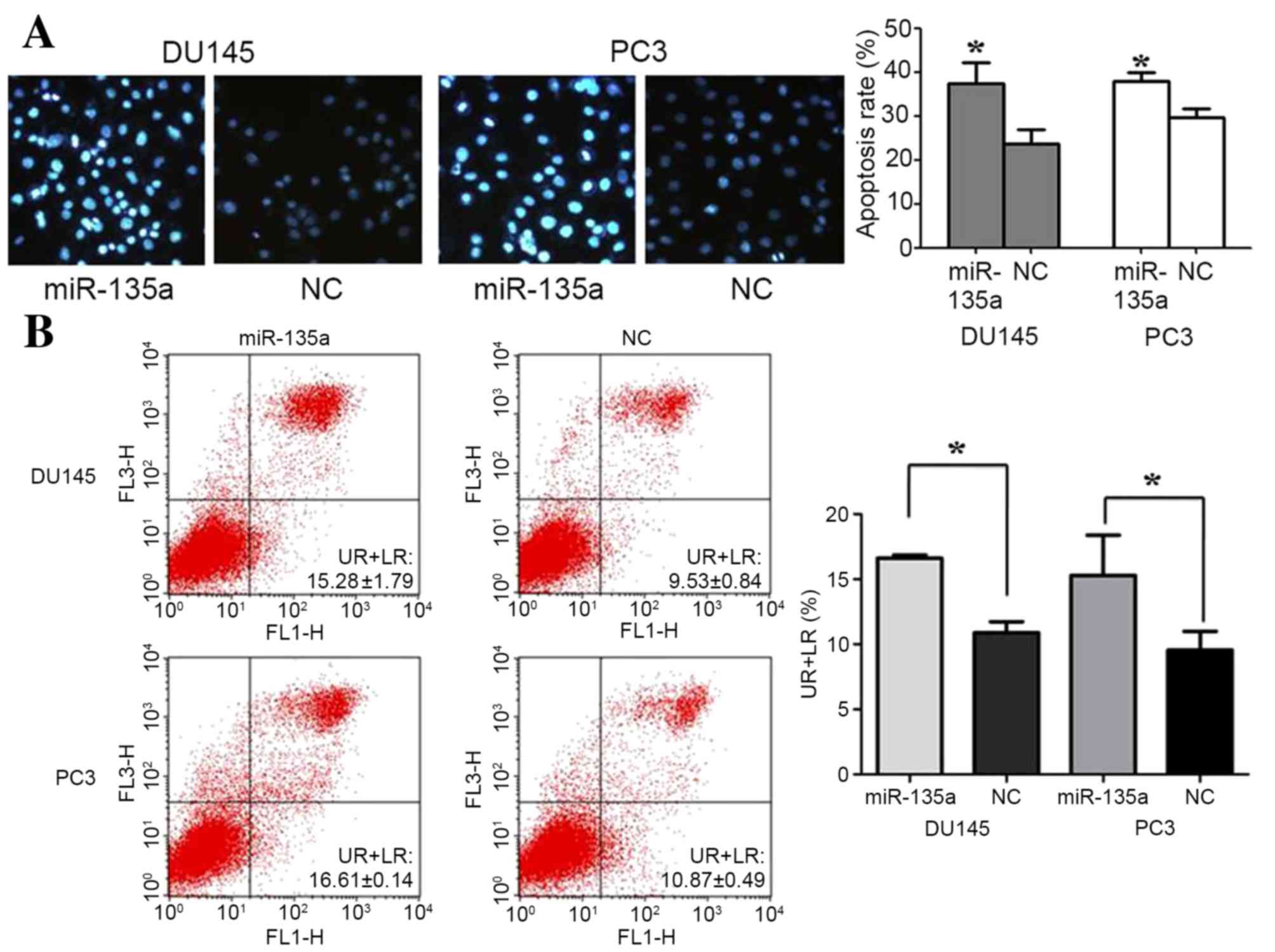
To determine whether the overexpression of miR-135a
affects the levels of apoptosis in prostate cancer cells, Hoechst
staining and FACS analyses were performed. Notably, overexpression
of miR-135a in DU145 and PC3 cells led to an increase in the
percentage of Hoechst-positive cells (37.3±2.7 vs. 23.67±1.9% in
DU145 cells and 38.0±1.2 vs. 29.7±1.2% in PC3 cells; P=0.0144 in
DU145 and P=0.0075 in PC3; Fig. 2A).
To further examine this result, the effects of miR-135a on the
levels of apoptosis were investigated using FACS analysis. DU145
and PC3 cells were transfected with miR-135a or NC and cells were
serum-deprived for 24 h prior to FACS. Compared with NC-transfected
cells, the proportion of apoptotic cells following transfection
with miR-135a was significantly increased (15.28±1.79 vs.
9.53±0.84% in DU145 cells and 16.61±0.14 vs. 10.87±0.49% in PC3
cells; P=0.0073 in DU145 and P<0.0001 in PC3; Fig. 2B).
STAT6 is a direct target of
miR-135a
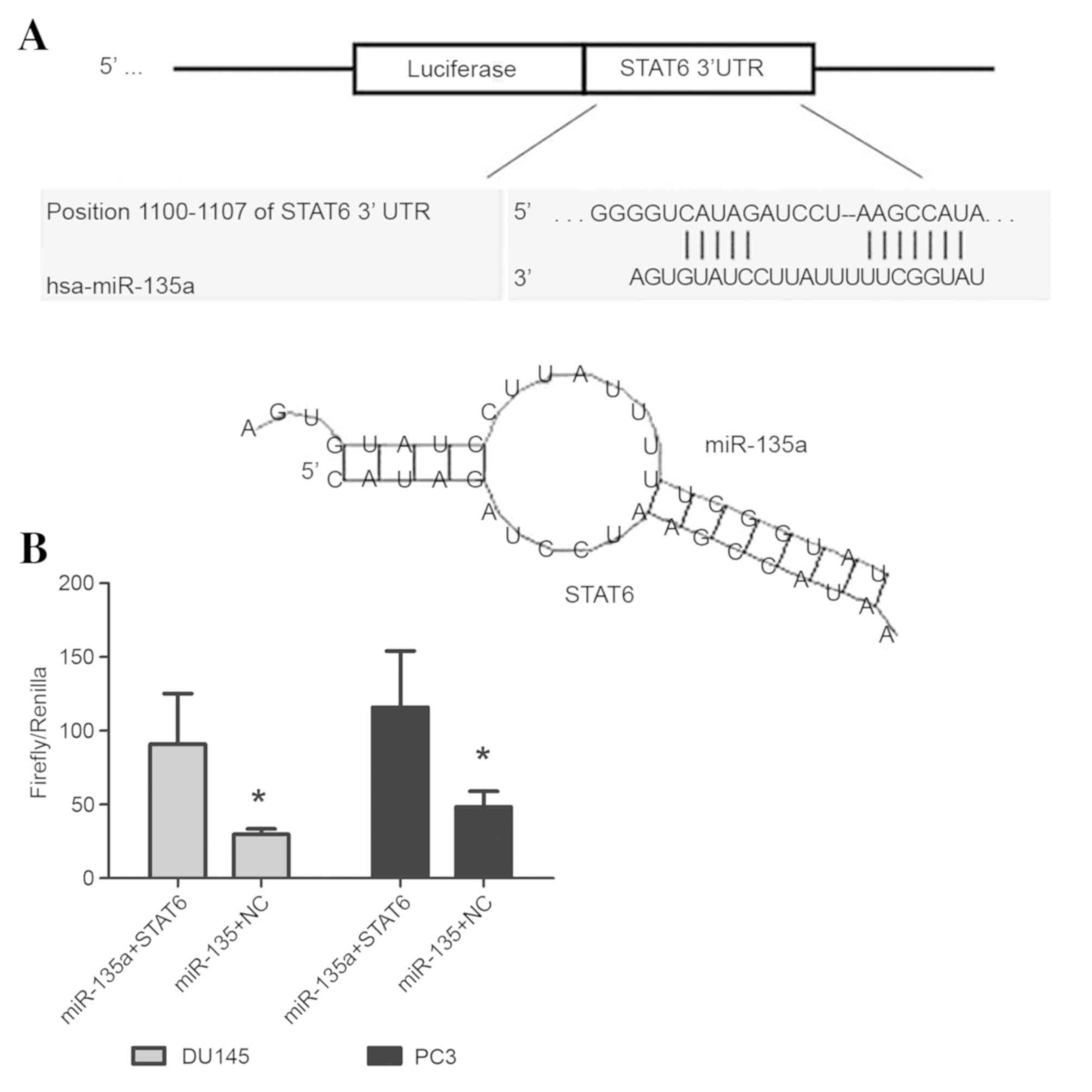
It is established that miRNAs exert their function
by directly binding to target genes. Therefore, STAT6 was selected
using TargetScan 5.1 (15) and
RNAhybrid (8). To investigate whether
STAT6 is a direct target of miR-135a, the predicted binding site
(1,100-1,107 bp) of miR-135a was cloned into the STAT6 3′UTR vector
(Fig. 3A). The luciferase reporter
expression was significantly reduced following co-transfection with
miR-135a compared with the NC group (Fig.
3B).
miR-135a negatively regulated the
protein expression and the function of STAT6
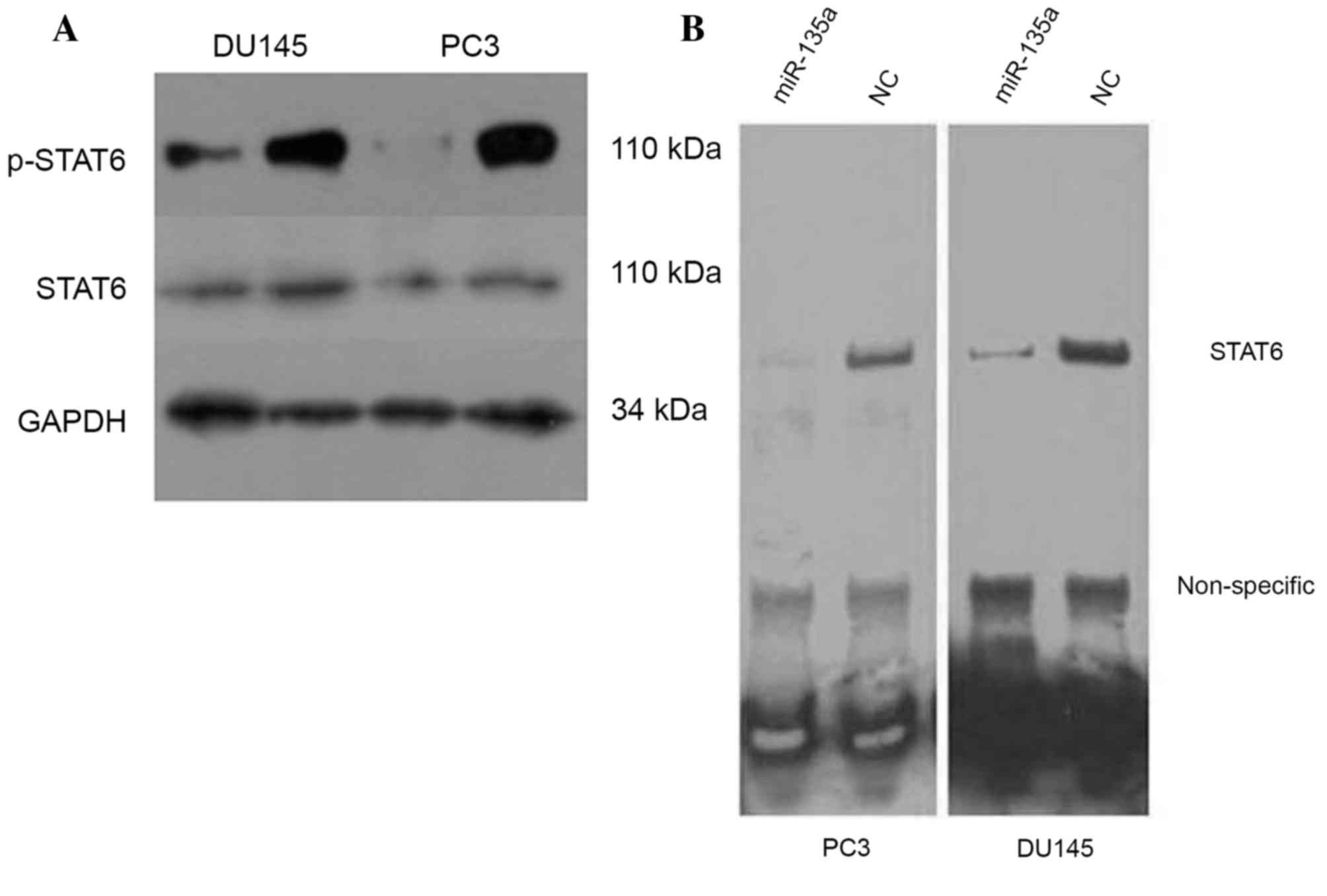
Western blot analyses for STAT6 expression in the
cells transfected with miR-135a or NC demonstrated that endogenous
STAT6 protein levels were downregulated in miR-135a-transfected
prostate cancer cells (Fig. 4A).
STAT6 is able to function as a transcriptional factor following its
phosphorylation (17), therefore
p-STAT6 expression levels and DNA-binding ability were analyzed
using western blotting and the EMSA assay. The results revealed
that the expression levels of p-STAT6 were also reduced by miR-135a
transfection following treatment with interleukin-4 (IL-4; 100
ng/ml). Additionally, EMSA demonstrated that the STAT6 DNA-binding
activity in prostate cancer cells was inhibited via the
overexpression of miR-135a (Fig.
4B).
Discussion
Elucidating the precise molecular mechanisms
underlying tumorigenesis is important for the development of
improved therapy strategies for patients with prostate cancer,
particularly for patients with metastases. Previous studies have
suggested that the dysregulation of miRNA is involved in the
formation of metastases, including miR-221/-222 (18) and miR-146a (14). The preliminary bioinformatic analyses
in the current study focused on miR-135a. The expression levels of
miR-135a in cancer are controversial, as it has been reported to be
upregulated and downregulated in distinct tumors and may function
as a tumor-inducer in colorectal (19) and breast cancer (20) and a tumor-suppressor in renal-cell
carcinoma (21) and gastric cancer
(22). Additionally, the
overexpression of miR-135a sensitized lung cancer cells to
cisplatin (23), whereas suppression
of miR-135a sensitized drug-resistant cell lines to
paclitaxel-induced cell death (24).
Prostate cancer has a distinct progression and the underlying
mechanisms are yet to be elucidated. The aberrant expression,
subsequent function and underlying mechanisms of miR-135a in
prostate cancer are yet to be established. In the present study,
STAT6 inhibition via miR-135a promoted apoptosis in prostate cancer
cells and it was demonstrated that STAT6 is a constitutively
expressed survival factor in several human prostate cancer
datasets, and may a direct functional target of miR-135a.
Prior to activation, STAT family proteins are
cytosolic and, following activation, translocate into the nucleus
and function as transcription factors. Sustained STAT activity has
been described in a number of tumor types (25). STAT6 serves an important role in
exerting IL-4-mediated biological responses. Following stimulation
by IL-4, STAT6 is phosphorylated via receptor-associated kinases
and translocated into the cell nucleus (25). Das et al (5) demonstrated that STAT6 expression levels
were significantly correlated with higher histological grades of
prostate cancer and with tumor size. Downregulation of STAT6 using
siRNA promoted the induction of apoptosis and a reduction in cell
viability in DU145 cells. Due to the established roles of STAT6 in
prostate cancer, the present study investigated the role of STAT6
and its association with miR-135a. Concordant with previous
studies, the results provided novel data to support the hypothesis
that the downregulation of STAT6 in prostate cancer may occur as a
result of the overexpression of the miRNA molecule miR-135a, which
may indicate a regulatory mechanism that is able to drive apoptosis
in prostate cancer cells.
In conclusion, miR-135a was identified as an
important miRNA that may regulate the levels of apoptosis in
prostate cancer cells. It was hypothesized that the miR-135a-STAT6
association may be a novel target for prognostic and therapeutic
advances in prostate cancer.
Acknowledgements
Not applicable.
Funding
The present study was supported by the National
Natural Science Foundation of China (grant no. 81202034), the
Science Foundation of Ministry of Education of China (grant no.
20120092120071) and the Natural Science Foundation of Jiangsu
Province (grant nos. BK2012336 and BL2013032), Nanjing City (grant
no. 201201053) and the Southeast University (grant no.
3290002402).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
BX and MC participated in study design and drafted
the manuscript. XL and YZ participated in collection of data and
manuscript preparation. CL and XH performed the statistical
analysis and participated in the critical revision of the
manuscript. SC, WZ and LZ assisted in data analysis, interpretation
of the data and critically revised the manuscript for important
intellectual content. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Feldman BJ and Feldman D: The development
of androgen-independent prostate cancer. Nat Rev Cancer. 1:34–45.
2001. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Debatin KM: Apoptosis pathways in cancer
and cancer therapy. Cancer Immunol Immunother. 53:153–159. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Tran C, Ouk S, Clegg NJ, Chen Y, Watson
PA, Arora V, Wongvipat J, Smith-Jones PM, Yoo D, Kwon A, et al:
Development of a second-generation antiandrogen for treatment of
advanced prostate cancer. Science. 324:787–790. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Das S, Roth CP, Wasson LM and Vishwanatha
JK: Signal transducer and activator of transcription-6 (STAT6) is a
constitutively expressed survival factor in human prostate cancer.
Prostate. 67:1550–1564. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kotb S, Mosharafa A, Essawi M, Hassan H,
Meshref A and Morsy A: Circulating miRNAs 21 and 221 as biomarkers
for early diagnosis of prostate cancer. Tumour Biol.
35:12613–12617. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chang YL, Zhou PJ, Wei L, Li W, Ji Z, Fang
YX and Gao WQ: MicroRNA-7 inhibits the stemness of prostate cancer
stem-like cells and tumorigenesis by repressing KLF4/PI3K/Akt/p21
pathway. Oncotarget. 6:24017–24031. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kojima S, Enokida H, Yoshino H, Itesako T,
Chiyomaru T, Kinoshita T, Fuse M, Nishikawa R, Goto Y, Naya Y, et
al: The tumor-suppressive microRNA-143/145 cluster inhibits cell
migration and invasion by targeting GOLM1 in prostate cancer. J Hum
Genet. 59:78–87. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sun Q, Zhao X, Liu X, Wang Y, Huang J,
Jiang B, Chen Q and Yu J: miR-146a functions as a tumor suppressor
in prostate cancer by targeting Rac1. Prostate. 74:1613–1621. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kobayashi N, Uemura H, Nagahama K, Okudela
K, Furuya M, Ino Y, Ito Y, Hirano H, Inayama Y, Aoki I, et al:
Identification of miR-30d as a novel prognostic maker of prostate
cancer. Oncotarget. 3:1455–1471. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Taylor BS, Schultz N, Hieronymus H,
Gopalan A, Xiao Y, Carver BS, Arora VK, Kaushik P, Cerami E, Reva
B, et al: Integrative genomic profiling of human prostate cancer.
Cancer Cell. 18:11–22. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Mootha VK, Lindgren CM, Eriksson KF,
Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E,
Ridderstråle M, Laurila E, et al: PGC-1alpha-responsive genes
involved in oxidative phosphorylation are coordinately
downregulated in human diabetes. Nat Genet. 34:267–273. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Xu B, Niu X, Zhang X, Tao J, Wu D, Wang Z,
Li P, Zhang W, Wu H, Feng N, et al: miR-143 decreases prostate
cancer cells proliferation and migration and enhances their
sensitivity to docetaxel through suppression of KRAS. Mol Cell
Biochem. 350:207–213. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Xu B, Wang N, Wang X, Tong N, Shao N, Tao
J, Li P, Niu X, Feng N, Zhang L, et al: MiR-146a suppresses tumor
growth and progression by targeting EGFR pathway and in a
p-ERK-dependent manner in castration-resistant prostate cancer.
Prostate. 72:1171–1178. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Garcia DM, Baek D, Shin C, Bell GW,
Grimson A and Bartel DP: Weak seed-pairing stability and high
target-site abundance decrease the proficiency of lsy-6 and other
microRNAs. Nat Struct Mol Biol. 18:1139–1146. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Krüger J and Rehmsmeier M: RNAhybrid:
microRNA target prediction easy, fast and flexible. Nucleic Acids
Res. 34:(Web Server Issue). W451–W454. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Rahal OM, Wolfe AR, Mandal PK, Larson R,
Tin S, Jimenez C, Zhang D, Horton J, Reuben JM, McMurray JS and
Woodward WA: Blocking interleukin (IL)4- and IL13-mediated
phosphorylation of STAT6 (Tyr641) decreases M2 polarization of
macrophages and protects against macrophage-mediated
radioresistance of inflammatory breast cancer. Int J Radiat Oncol
Biol Phys. 100:1034–1043. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Sun T, Wang X, He HH, Sweeney CJ, Liu SX,
Brown M, Balk S, Lee GS and Kantoff PW: MiR-221 promotes the
development of androgen independence in prostate cancer cells via
downregulation of HECTD2 and RAB1A. Oncogene. 33:2790–2800. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhou W, Li X, Liu F, Xiao Z, He M, Shen S
and Liu S: MiR-135a promotes growth and invasion of colorectal
cancer via metastasis suppressor 1 in vitro. Acta Biochim Biophys
Sin (Shanghai). 44:838–846. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen Y, Zhang J, Wang H, Zhao J, Xu C, Du
Y, Luo X, Zheng F, Liu R, Zhang H and Ma D: miRNA-135a promotes
breast cancer cell migration and invasion by targeting HOXA10. BMC
Cancer. 12:1112012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yamada Y, Hidaka H, Seki N, Yoshino H,
Yamasaki T, Itesako T, Nakagawa M and Enokida H: Tumor-suppressive
microRNA-135a inhibits cancer cell proliferation by targeting the
c-MYC oncogene in renal cell carcinoma. Cancer Sci. 104:304–312.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wu H, Huang M, Cao P, Wang T, Shu Y and
Liu P: MiR-135a targets JAK2 and inhibits gastric cancer cell
proliferation. Cancer Biol Ther. 13:281–288. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhou L, Qiu T, Xu J, Wang T, Wang J, Zhou
X, Huang Z, Zhu W, Shu Y and Liu P: miR-135a/b modulate cisplatin
resistance of human lung cancer cell line by targeting MCL1. Pathol
Oncol Res. 19:677–683. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Holleman A, Chung I, Olsen RR, Kwak B,
Mizokami A, Saijo N, Parissenti A, Duan Z, Voest EE and Zetter BR:
miR-135a contributes to paclitaxel resistance in tumor cells both
in vitro and in vivo. Oncogene. 30:4386–4398. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Watson CJ: Stat transcription factors in
mammary gland development and tumorigenesis. J Mammary Gland Biol
Neoplasia. 6:115–127. 2001. View Article : Google Scholar : PubMed/NCBI
|