Introduction
Tumor metastasis is the principal reason for poor
survival in patients with cancer (1), and its treatment and prevention are
particularly challenging (2). Lung
cancer is one of the most frequently diagnosed cancer types, and is
the leading cause of mortality among males, and the second leading
cause among females (3). A
considerable number of patients with lung cancer are diagnosed at
the late stages of the disease, and with existing tumor metastasis,
their prognosis is poor (3). The
majority of lung cancer studies have focused on non-small cell lung
cancer (NSCLC) (4). SCLC is an
aggressive, highly metastasizing and frequently lethal type of lung
cancer, yet it accounts for only ~15% of all cases of lung cancer
(5). At present, studies on SCLC are
limited in number.
Through its role in epithelial-mesenchymal
transition (EMT), activation of the transforming growth factor β
(TGF-β) pathway is associated with metastasis in a number of
malignancies (6). The tumor
suppressive and oncogenic roles of TGF-β have been extensively
investigated in different types of lung cancer (7,8). TGF-β
signaling inhibits cell proliferation at the early stages of tumor
initiation (7,8). By contrast, activation of TGF-β
signaling promotes tumor metastasis at the late stages of cancer
development. The actions of TGF-β may be mediated by interactions
with long non-coding RNAs (lncRNAs) (9), a group of ncRNAs composed of >200
nucleotides (10). A growing body of
literature has illustrated that lncRNAs are critical in human
diseases (10). lncRNA-neighboring
enhancer of FOXA2 (NEF) is a recently identified tumor suppressor
in hepatocellular carcinoma, but with unknown functionality in
other cancer types (11). In the
present study, lncRNA-NEF inhibited the migration and invasion of
SCLC cells, possibly as a result of TGF-β pathway inhibition.
Materials and methods
Patients
A total of 134 patients with SCLC were diagnosed and
treated in The First People's Hospital of Tianmen City (Tianmen,
Hubei, China) between May 2015 and January 2018. Among those
patients, 46 were included in the present study according to
pre-determined exclusion and inclusion criteria. Inclusion criteria
were as follows: i) Patients diagnosed using fine-needle lung
biopsies; ii) patients diagnosed and treated for the first time;
and iii) patients willing to participate. Exclusion criteria were
as follows: i) Patients with other malignancies; ii) patients with
chronic lung diseases; iii) patients >70 years of age; and iv)
patients who had previously received treatment for lung cancer. The
patient group included 29 males and 17 females (age range, 23–68
years; mean age, 45.5±5.6 years), and all patients were between
cancer stages IIIA and IVB (as defined by The American Joint
Committee on Cancer) (12). A total
of 98 individuals concurrently received lung biopsies in The First
People's Hospital of Tianmen City to detect potential lung lesions;
32 of these individuals tested negative for lung lesions, and were
recruited as the control group consisting of 18 males and 14
females (age range, 26–69 years; mean age, 46.6±6.1 years). There
were no significant differences in age and sex between the two
groups. The present study was approved by the Ethics Committee of
The First People's Hospital of Tianmen City, and all patients
provided written informed consent.
Lung biopsies and preparation of
plasma
Lung biopsies of all subjects were obtained from the
specimen library of The First People's Hospital of Tianmen City.
Blood (10 ml) was extracted in the morning prior to breakfast from
the elbow vein of each subject. Blood was used to prepare plasma
through centrifugation at 1,200 × g for 15 min at room
temperature.
Cell lines, culture and
transfection
The human SCLC cell lines, SHP-77 [American Type
Culture Collection (ATCC) CRL-2195™] and NCI-H69 [H69]
(ATCC HTB-119™) were purchased from the ATCC (Manassas,
VA, USA). The cells were cultured with ATCC-formulated RPMI-1640
Medium (cat. no. 30-2001; ATCC) containing 10% fetal bovine serum
(FBS; cat. no. 30–2020; ATCC) at 37°C with 5% CO2.
Full-length lncRNA-NEF DNA (11) was
amplified using polymerase chain reaction (PCR) and inserted into
the pIRSE2 vector (Clontech, Laboratories, Inc., Mountainview, CA,
USA) to establish an lncRNA-NEF expression vector. Cells were
cultured to 80–90% confluence and 5×105 cells were
transfected with 10 mM vector using Lipofectamine 2000®
reagent (cat. no. 11668-019; Invitrogen; Thermo Fisher Scientific,
Inc., Waltham, MA, USA). Untransfected cells and cells transfected
with empty vector were used as controls and further experiments
were carried out at 24 h following transfection.
Reverse transcription-quantitative PCR
(RT-qPCR)
TRIzol® reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) was used to extract total RNA from lung
biopsy tissues, plasma and in vitro cultured cells. For
TGF-β1 (Sigma-Aldrich, USA) treatment, cells were treated with
exogenous TGF-β1 at 5, 10, 20 and 50 ng/ml for 24 at 37°C after
transfection before RNA extractions. The SuperScript IV Reverse
Transcriptase kit (Thermo Fisher Scientific, Inc.) was used to
synthesize cDNA, and SYBR® Green Real-Time PCR Master
mix (Thermo Fisher Scientific, Inc.) was used to conduct PCR using
an ABI PRISM 7500 sequence detection system (Applied Biosystems,
Rockford, IL, USA). The thermocycling conditions were as follows:
80 sec at 95°C, followed by 40 cycles of 22 sec at 95°C and 40 sec
at 58°C. Primers used in the PCR were as follows: lncRNA-NEF
forward, 5′-CTGCCGTCTTAAACCAACCC-3′ and reverse,
5′-GCCCAAACAGCTCCTCAATT-3′; and β-actin forward,
5′-GACCTCTATGCCAACACAGT-3′ and reverse, 5′-AGTACTTGCGCTCAGGAGGA-3′.
Data normalization was performed using the 2−ΔΔcq method
(13), and the experiment was
performed in triplicate.
Cell migration and invasion
assays
Following transfection, an lncRNA-NEF expression
rate of >200% was confirmed using RT-qPCR. Following
transfection, for TGF-β1 treatment, cells were treated with 10
ng/ml exogenous TGF-β1 (Sigma-Aldrich; Merck KGaA, Darmstadt,
Germany) for 24 h at 37°C prior to use. Cell migration and invasion
were detected using Transwell cell migration and invasion kits (BD
Biosciences, San Jose, CA, USA). Cell suspensions were prepared
using RPMI-1640 medium (non-serum) to a final concentration of
5×104 cell/ml. For the migration assay, 5×103
cells in 0.1 ml cell suspension were added to the upper Transwell
chamber, while the lower chamber was filled with RPMI containing
20% FBS. Cells were cultured for 6 h and the membranes were
subsequently stained with 0.5% crystal violet (Sigma-Aldrich; Merck
KGaA) at room temperature for 20 min. The invasion assay was
performed in the same manner, but the upper chamber was pre-coated
with Matrigel (cat. no. 356234; EMD Millipore, Billerica, MA, USA)
prior to the addition of the cells. Stained cells were counted
under an optical microscope (Olympus Corporation, Tokyo,
Japan).
Western blotting
Cell lysis buffer (Clontech, Laboratories, Inc.,)
was used to extract protein from in vitro-cultured cells,
and the protein concentration was determined using a bicinchoninic
acid assay. Protein samples were denatured and 20 µg protein per
lane was separated using SDS-PAGE on a 10% gel. Following transfer,
PVDF membranes (Bio-Rad, Laboratories, Inc., Hercules, CA, USA)
were blocked with 5% skimmed milk at room temperature for 2 h, and
incubated with rabbit anti-human primary antibodies against TGF-β1
(1:2,000; cat. no. ab92486, Abcam, Cambridge, UK) and GAPDH
(1:1,000; cat. no. ab9485, Abcam) overnight at 4°C. Subsequent to
washing with PBS in triplicate at room temperature for 15 min per
time, membranes were further incubated with goat anti-rabbit
IgG-HRP secondary antibody (1:1,000; cat. no. MBS435036;
MyBioSource, San Diego, CA, USA) at room temperature for 2 h.
ECL™ Prime Western Blotting System (ECL; Sigma-Aldrich;
Merck KGaA) was used to develop the blots, and the relative
expression level of TGF-β1 was normalized to GAPDH using Image J
1.51 software (National Institutes of Health, Bethesda, MD,
USA).
Statistical analysis
GraphPad Prism 6 (GraphPad Software, Inc., La Jolla,
CA, USA) was used for all statistical analyses. Gene expression,
cell migration and invasion data are expressed as the mean ±
standard deviation, and compared using the unpaired t-test (between
two groups), or one-way analysis of variance followed by least
significant difference test (among multiple groups). Associations
between the plasma levels of lncRNA-NEF and the clinicopathological
data of patients were analyzed using the χ2 test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Expression of lncRNA-NEF is
downregulated in patients with SCLC compared with healthy
controls
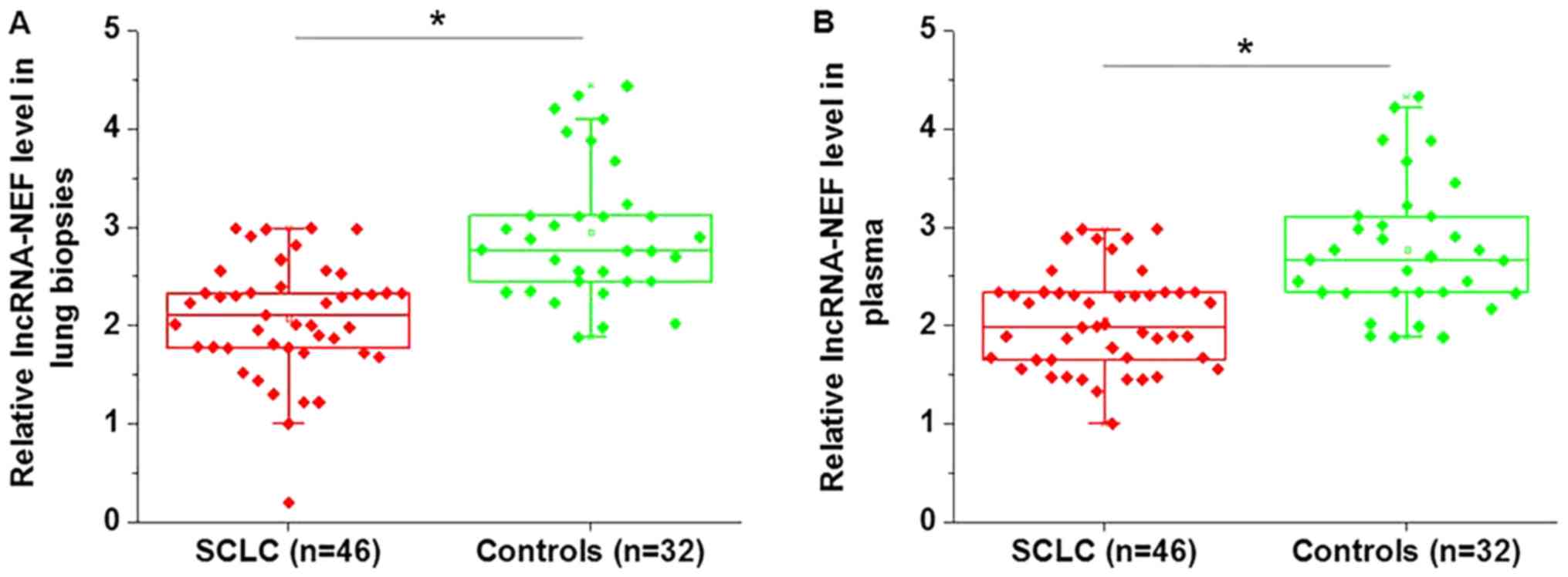
Expression level of lncRNA-NEF was detected in the
lung tissue and plasma of patients with SCLC, and in healthy
controls. As illustrated in Fig. 1,
the expression levels of lncRNA-NEF were significantly lower in the
lung tissue (Fig. 1A, P<0.05) and
plasma (Fig. 1B, P<0.05) of
patients with SCLC, compared with those of the healthy
controls.
Downregulation of lncRNA-NEF
distinguishes patients with SCLC from healthy controls
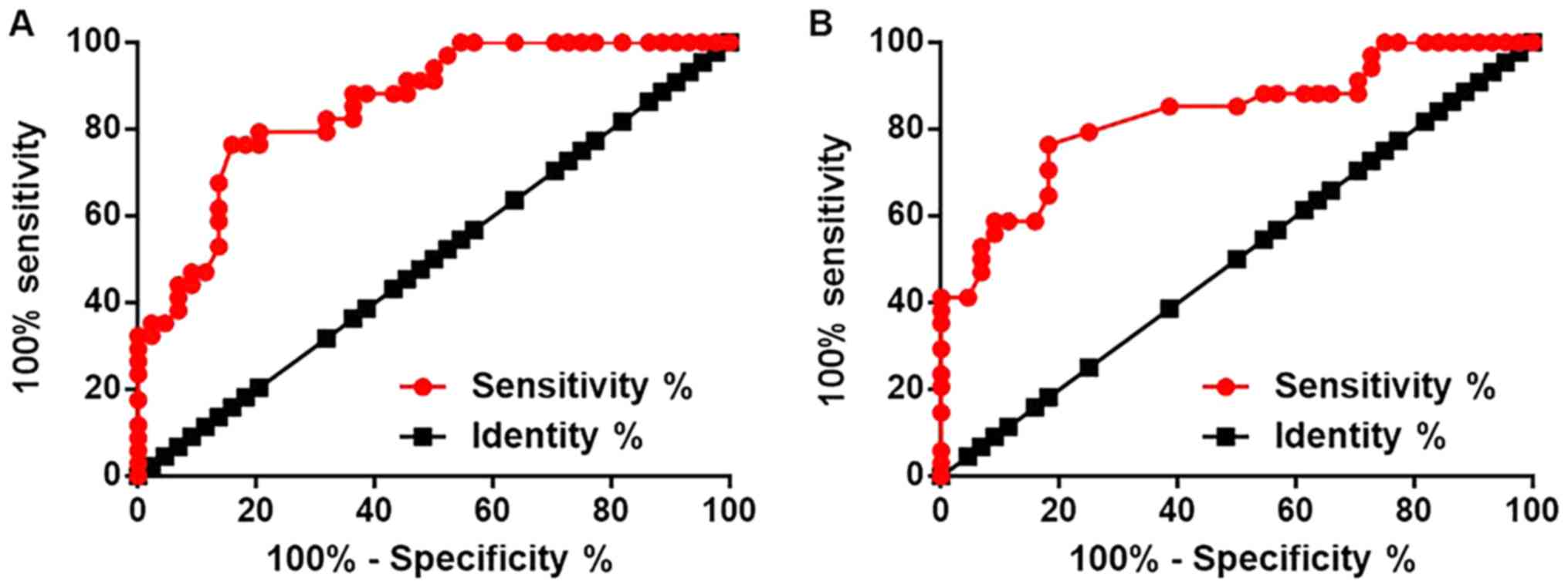
Receiver operating characteristic curve analysis was
performed to evaluate the diagnostic value of lncRNA-NEF in SCLC.
For lncRNA-NEF expression in lung biopsies, the area under the
curve (AUC) was 0.8549, with a standard error of 0.04144 and a 95%
confidence interval of 0.7737–0.9362 (Fig. 2A). For lncRNA-NEF expression in
plasma, the AUC was 0.8309, with a standard error of 0.04755 and a
95% confidence interval of 0.7377–0.9241 (Fig. 2B).
Expression levels of lncRNA-NEF are
associated with distant tumor metastasis, but not tumor size
Patients were divided into high and low expression
groups according to the median relative expression level of
lncRNA-NEF in lung biopsies (2.14) and plasma (2.02), respectively.
Associations between plasma levels of lncRNA-NEF and the
clinicopathological data of patients were analyzed using the
χ2 test. The results illustrated that expression levels
of lncRNA-NEF in lung biopsies (Table
I) and plasma (Table II) were
significantly associated with distant tumor metastasis (P<0.05),
but not age, sex, smoking status, alcohol consumption or tumor
size.
 | Table I.Associations between the expression
levels of long non-coding RNA-neighboring enhancer of FOXA2 in lung
biopsy tissues and the clinicopathological characteristics of
patients with small cell lung cancer. |
Table I.
Associations between the expression
levels of long non-coding RNA-neighboring enhancer of FOXA2 in lung
biopsy tissues and the clinicopathological characteristics of
patients with small cell lung cancer.
| Variable | Cases, n | High expression,
n | Low expression,
n | χ2
value | P-value |
|---|
| Sex |
|
|
| 0.84 | 0.36 |
| Male | 29 | 13 | 16 |
|
|
|
Female | 17 | 10 | 7 |
|
|
| Age, years |
|
|
| 0.79 | 0.37 |
| ≥45 | 25 | 11 | 14 |
|
|
|
<45 | 21 | 12 | 9 |
|
|
| Tumor size, cm |
|
|
|
|
|
|
>7 | 16 | 7 | 9 | 0.47 | 0.79 |
| 5–7 | 18 | 10 | 8 |
|
|
|
<5 | 12 | 6 | 6 |
|
|
| Distant tumor
metastasis |
|
|
| 5.66 | 0.02 |
| Yes | 26 | 9 | 17 |
|
|
| No | 20 | 14 | 6 |
|
|
| Smoking status |
|
|
| 0.35 | 0.55 |
|
Positive | 22 | 10 | 12 |
|
|
|
Negative | 24 | 13 | 11 |
|
|
| Alcohol
consumption |
|
|
| 0.81 | 0.37 |
| Yes | 19 | 8 | 11 |
|
|
| No | 27 | 15 | 12 |
|
|
 | Table II.Associations between the plasma
expression levels of long non-coding RNA-neighboring enhancer of
FOXA2 in and the clinicopathological characteristics of patients
with small cell lung cancer. |
Table II.
Associations between the plasma
expression levels of long non-coding RNA-neighboring enhancer of
FOXA2 in and the clinicopathological characteristics of patients
with small cell lung cancer.
| Variable | Cases, n | High expression,
n | Low expression,
n | χ2
value | P-value |
|---|
| Sex |
|
|
|
|
|
|
Male | 29 | 14 | 15 | 0.10 | 0.79 |
|
Female | 17 | 9 | 8 |
|
|
| Age, years |
|
|
|
|
|
|
≥45 | 25 | 10 | 15 | 2.19 | 0.14 |
|
<45 | 21 | 13 | 8 |
|
|
| Tumor size, cm |
|
|
|
|
|
|
>7 | 16 | 7 | 9 | 0.58 | 0.75 |
|
5–7 | 18 | 9 | 9 |
|
|
|
<5 | 12 | 7 | 5 |
|
|
| Distant tumor
metastasis |
|
|
|
|
|
|
Yes | 26 | 9 | 17 | 5.66 | 0.02 |
| No | 20 | 14 | 6 |
|
|
| Smoking status |
|
|
|
|
|
|
Positive | 22 | 9 | 13 | 1.39 | 0.24 |
|
Negative | 24 | 14 | 10 |
|
|
| Alcohol
consumption |
|
|
|
|
|
|
Yes | 19 | 8 | 11 | 0.81 | 0.37 |
| No | 27 | 15 | 12 |
|
|
lncRNA-NEF overexpression inhibits and
exogenous TGF-β1 promotes the migration and invasion of SCLC
cells
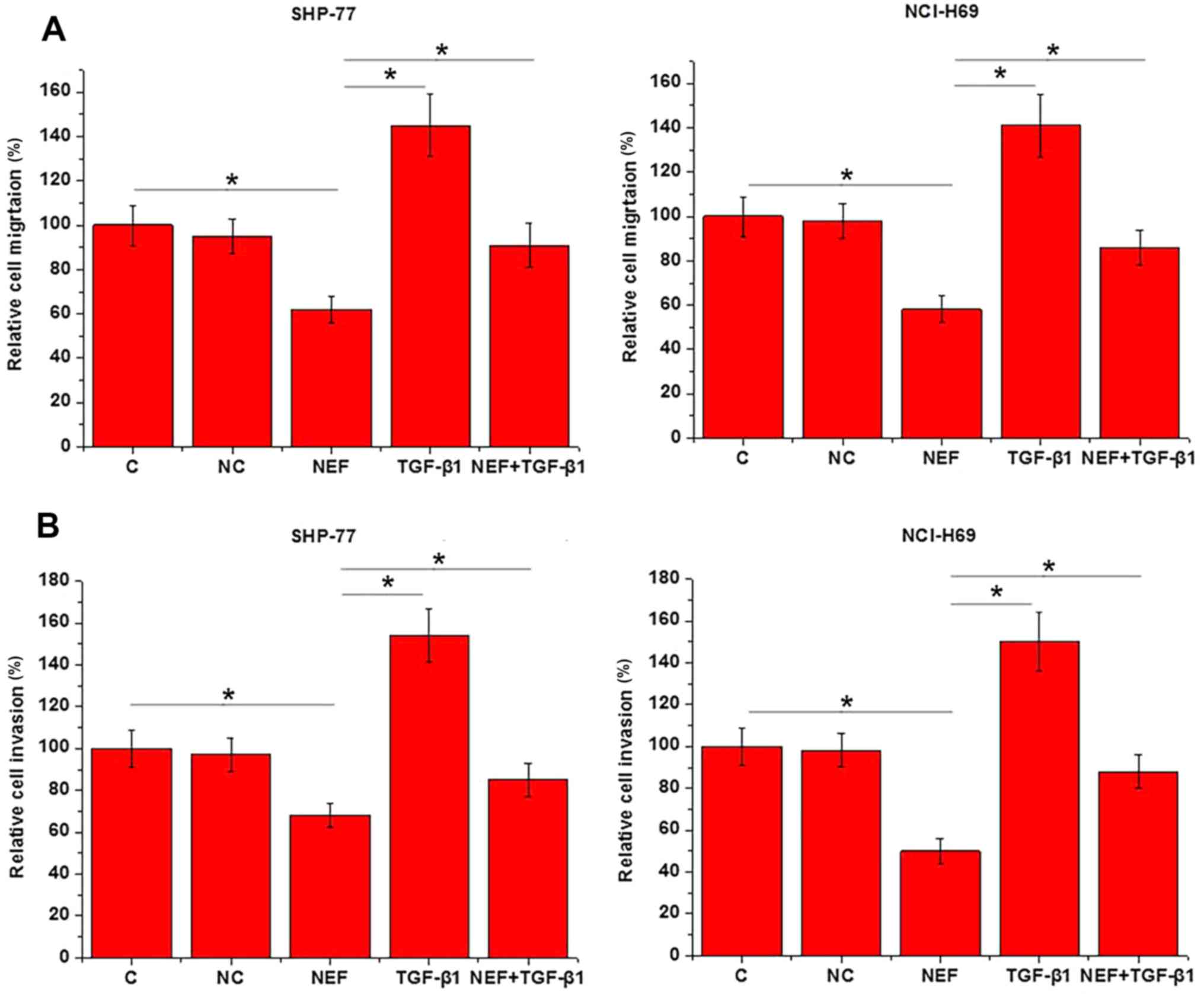
The aforementioned results indicated the involvement
of lncRNA-NEF in SCLC metastasis. To further support these
findings, lncRNA-NEF expression vectors were transfected into
SHP-77 and NCI-H69 cell lines, and cell migration and invasion were
assessed using Transwell migration and invasion assays,
respectively. Compared with the control and negative control cells,
the cells overexpressing lncRNA-NEF demonstrated a significant
reduction in migration (Fig. 3A) and
invasion (P<0.05; Fig. 3B). By
contrast, treatment with 10 ng/ml exogenous TGF-β1 significantly
promoted cell migration and invasion, and reduced the inhibitory
effects of lncRNA-NEF overexpression on cell migration and invasion
(P<0.05).
lncRNA-NEF is a potential upstream
inhibitor of TGF-β1 in SCLC
To further investigate the interactions between
TGF-β1 and lncRNA-NEF, the expression level of TGF-β1 following
lncRNA-NEF overexpression was detected using western blotting. As
displayed in Fig. 4A, overexpression
of lncRNA-NEF was achieved in two SCLC cell lines (P<0.05).
Compared with the control and negative control cells, cells
overexpressing lncRNA-NEF exhibited significantly lower TGF-β1
expression levels (active homodimer; Fig. 4B; P<0.05). By contrast, treatment
with exogenous TGF-β1 at 5, 10, 20 and 50 ng/ml did not
significantly effect lncRNA-NEF expression (Fig. 4C; P>0.05).
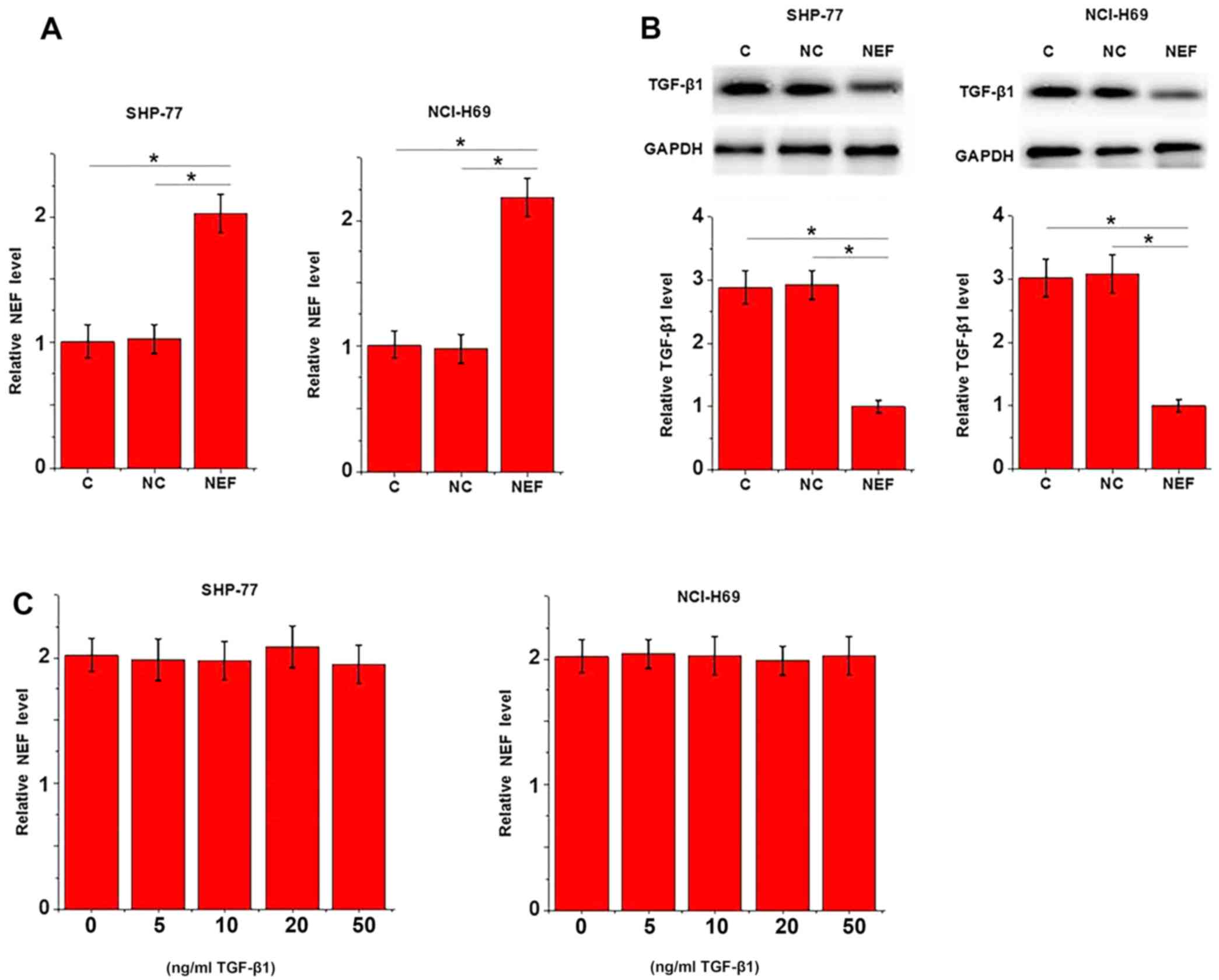 | Figure 4.Potential lncRNA-NEF-associated
inhibition of TGF-β1 in SCLC. (A) Compared with the control and
negative control, overexpression of lncRNA-NEF was achieved in
SHP-77 and NCI-H69 cells. (B) Western blot analysis revealed that
lncRNA-NEF overexpression significantly downregulated TGF-β1
expression. (C) By contrast, reverse transcription-quantitative
polymerase chain reaction revealed that treatment with exogenous
TGF-β1 (15, 10, 20 and 50 ng/ml) exhibited no significant effect on
lncRNA-NEF expression. The experiment was performed in triplicate.
*P<0.05, as revealed by one-way analysis of variance, followed
by least significant difference test. lncRNA-NEF, long non-coding
RNA-neighboring enhancer of FOXA2; TGF-β1, transforming growth
factor β1; SCLC, small cell lung carcinoma; C, control; NC,
negative control. |
Discussion
lncRNA-NEF is a recently identified lncRNA with
characterized tumor suppressor activity in hepatocellular carcinoma
(11). The primary finding of the
present study was that lncRNA-NEF may also be a tumor suppressor in
SCLC. The results provided evidence that lncRNA-NEF was involved in
the regulation of SCLC tumor metastasis, and that this was
associated with the TGF-β pathway.
Altered expression levels of particular lncRNAs have
been observed in the development of different types of lung cancer,
including SCLC (14); lncRNA-taurine
upregulated gene 1 (TUG1) was upregulated in SCLC, and the
overexpression of TUG1 in cancer tissues and cells not only
promoted cell growth, but also reduced the chemoresistance of
cancer cells (15). Upregulation of
lncRNA-exocyst complex component 7 (EXOC7) has also been observed
in the cancer tissues of patients with SCLC (compared with
para-cancerous tissues), and increased expression levels of
lncRNA-EXOC7 provided diagnostic value (16). Additionally, lncRNA-NEF displayed a
downregulated expression pattern in hepatocellular carcinoma
(11). In the present study,
significantly lower expression levels of lncRNA-NEF were observed
in the lung tissue and plasma of patients with SCLC, compared with
those in the healthy controls, indicating a role for lncRNA-NEF as
a tumor suppressor in SCLC.
Differential expression of lncRNAs provides guidance
for the diagnosis of lung cancer (17,18). In
the present study, ROC curve analysis revealed that the
downregulation of lncRNA-NEF may be used to effectively distinguish
SCLC patients from healthy individuals. Furthermore, lncRNA-NEF
expression levels were not significantly associated with patients'
age, gender, alcohol consumption and smoking status, factors that
may potentially influence lncRNA expression. Therefore, lncRNA-NEF
may serve as a potential diagnostic marker for SCLC. However,
lncRNA-NEF is a newly identified lncRNA with a known expression
pattern in hepatocellular carcinoma only (11), thus the use of multiple biomarkers
may improve diagnostic specificity.
TGF-β signaling is a double-edged sword in cancer
biology (19), as it inhibits tumor
growth at the initiation of cancer, but promotes tumor metastasis
at the later stages of disease (7,20). The
involvement of TGF-β signaling in the regulation of lung cancer
metastasis has been reported in a number of subtypes, including
lung adenocarcinoma metastasis (7)
and NSCLC (21). In the present
study, TGF-β1 resulted in accelerated migration and invasion of
SCLC cells in vitro, suggesting that TGF-β1 is involved in
the metastasis of SCLC. It has been frequently observed that
expression of TGF-β1 can be regulated by lncRNAs during cancer
development (22,23). The present study suggested that
lncRNA-NEF is an upstream inhibitor of TGF-β1 in the regulation of
migration and invasion of cells in SCLC. However, whether this
regulatory role is direct or indirect remains unknown. Future
studies aim to identify the potential intermediates between
lncRNA-NEF and TGF-β1.
In conclusion, lncRNA-NEF is downregulated in SCLC
patients compared with healthy controls, supporting its role as a
tumor suppressor in this disease. lncRNA-NEF may promote
metastasis, but not tumor growth, in SCLC by interacting with
TGF-β1.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Author's contributions
LW and PW were responsible for the conception and
design of the study, and performed the experiments. LW analyzed and
interpreted the data. LW and PW then drafted the article and were
responsible for the revision of the manuscript.
Ethics approval and consent to
participate
The protocol of the present study was approved by
the Ethics Review Committee of The First People's Hospital of
Tianmen City (Tianmen, China), and all patients provided written
informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Heerboth S, Housman G, Leary M, Longacre
M, Byler S, Lapinska K, Willbanks A and Sarkar S: EMT and tumor
metastasis. Clin Transl Med. 4:62015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ghajar CM: Metastasis prevention by
targeting the dormant niche. Nat Rev Cancer. 15:238–247. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ettinger DS, Akerley W, Bepler G, Blum MG,
Chang A, Cheney RT, Chirieac LR, D'Amico TA, Demmy TL, Ganti AK, et
al: Non-small cell lung cancer. J Natl Compr Canc Netw. 8:740–801.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Tanoue LT, Tanner NT, Gould MK and
Silvestri GA: Lung cancer screening. Am J Respir Crit Care Med.
191:19–33. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
David CJ, Huang YH, Chen M, Su J, Zou Y,
Bardeesy N, Iacobuzio-Donahue CA and Massagué J: TGF-β tumor
suppression through a lethal EMT. Cell. 164:1015–1030. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yu JR, Tai Y, Jin Y, Hammell MC, Wilkinson
JE, Roe JS, Vakoc CR and Van Aelst L: TGF-β/Smad signaling through
DOCK4 facilitates lung adenocarcinoma metastasis. Genes Dev.
29:250–261. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Yang H, Wang L, Zhao J, Chen Y, Lei Z, Liu
X, Xia W, Guo L and Zhang HT: TGF-β-activated SMAD3/4 complex
transcriptionally upregulates N-cadherin expression in non-small
cell lung cancer. Lung Cancer. 87:249–257. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li C, Wan L, Liu Z, Xu G, Wang S, Su Z,
Zhang Y, Zhang C, Liu X, Lei Z and Zhang HT: Long non-coding RNA
XIST promotes TGF-β-induced epithelial-mesenchymal transition by
regulating miR-367/141-ZEB2 axis in non-small-cell lung cancer.
Cancer Lett. 418:185–195. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Fatica A and Bozzoni I: Long non-coding
RNAs: New players in cell differentiation and development. Nat Rev
Genet. 15:7–21. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liang WC, Ren JL, Wong CW, Chan SO, Waye
MM, Fu WM and Zhang JF: LncRNA-NEF antagonized epithelial to
mesenchymal transition and cancer metastasis via cis-regulating
FOXA2 and inactivating Wnt/β-catenin signaling. Oncogene.
37:1445–1456. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rami-Porta R, Asamura H, Travis WD and
Rusch VW: Lung cancer-major changes in the American Joint Committee
on Cancer eighth edition cancer staging manual. CA Cancer J Clin.
67:138–155. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Peng Z, Zhang C and Duan C: Functions and
mechanisms of long noncoding RNAs in lung cancer. Onco Targets
Ther. 9:4411–4424. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Niu Y, Ma F, Huang W, Fang S, Li M, Wei T
and Guo L: Long non-coding RNA TUG1 is involved in cell growth and
chemoresistance of small cell lung cancer by regulating LIMK2b via
EZH2. Mol Cancer. 16:52017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zeng L, Yang R, Wang J and Liu Y: The
relationship between the expression of LncRNA EXOC7 and the
prognosis of patients with small cell lung cancer. Tumor.
37:269–274. 2017.
|
|
17
|
Wan L, Zhang L, Fan K and Wang JJ:
Diagnostic significance of circulating long noncoding RNA PCAT6 in
patients with non-small cell lung cancer. Onco Targets Ther.
10:5695–5702. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang HM, Lu JH, Chen WY and Gu AQ:
Upregulated lncRNA-UCA1 contributes to progression of lung cancer
and is closely related to clinical diagnosis as a predictive
biomarker in plasma. Int J Clin Exp Med. 8:11824–11830.
2015.PubMed/NCBI
|
|
19
|
Akhurst RJ and Derynck R: TGF-beta
signaling in cancer-a double-edged sword. Trends Cell Biol.
11:S44–S51. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Seoane J and Gomis RR: TGF-β family
signaling in tumor suppression and cancer progression. Cold Spring
Harb Perspect Biol. 9(pii): a0222772017. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang L, Yang H, Lei Z, Zhao J, Chen Y,
Chen P, Li C, Zeng Y, Liu Z, Liu X and Zhang HT: Repression of
TIF1γ by SOX2 promotes TGF-β-induced epithelial-mesenchymal
transition in non-small-cell lung cancer. Oncogene. 35:867–877.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang D, Qin H, Leng Y, Li X, Zhang L, Bai
D, Meng Y and Wang J: LncRNA MEG3 overexpression inhibits the
development of diabetic retinopathy by regulating TGF-β1 and VEGF.
Exp Ther Med. 16:2337–2342. 2018.PubMed/NCBI
|
|
23
|
Zhao B, Lu YL, Yang Y, Hu LB, Bai Y, Li
RQ, Zhang GY, Li J, Bi CW, Yang LB, et al: Overexpression of lncRNA
ANRIL promoted the proliferation and migration of prostate cancer
cells via regulating let-7a/TGF-β1/Smad signaling pathway. Cancer
Biomark. 21:613–620. 2018. View Article : Google Scholar : PubMed/NCBI
|