Introduction
In 2018, liver cancer ranked 6th in terms of
incidence (841,000 new cases) and 4th in terms of overall
mortality, with a mortality rate of 782,000 worldwide. Based on the
histological classification of primary liver cancer, it can be
divided into hepatocellular carcinoma (HCC; 75–85%), intrahepatic
cholangiocarcinoma (10–15%), and other rare types (1). Although numerous treatment approaches
are available for HCC, including liver transplantation, surgery,
radiofrequency ablation, radioembolization, trans-arterial
chemo-embolization and targeted therapy (2), improving the long-term survival for
patients with HCC remains a challenge. Therefore, more effective
molecular targets are urgently required in HCC.
microRNAs (miRNAs/miRs) constitute a class of small
noncoding RNAs, which regulate target gene expression mainly at the
post-transcription level (3).
Previous studies have reported that miRNAs are closely related to a
number of cellular processes (CCs), including cell cycle
regulation, inflammation, differentiation, migration and apoptosis,
thereby performing functional roles in the occurrence and
progression of various cancer types (4). A number of previous studies have shown
that miRNAs can act as biomarkers and therapeutic agents to improve
the diagnosis, and therapy, of cancer (5–7). Despite
these previous findings, it is important to identify potential
novel molecular targets.
Recently, miR-100-5p has been identified as a
biomarker in multiple types of cancer. For instance, miR-100-5p has
been shown to inhibit autophagy and induce apoptosis in colorectal
cancer cells by targeting autophagy protein 5 (8). Additionally, miR-100-5p enhances the
chemosensitivity of breast cancer by targeting HCLS1-associated
protein X-1 (9). Moreover,
miR-100-5p suppresses tumor migration and invasion by targeting
IGF1R in patients with nasopharyngeal carcinoma (10). miR-100-5p could also inhibit the
growth and metastasis of gastric cancer by targeting zinc finger
and BTB domain containing protein 7A (11). To date, few targets of miR-100-5p,
such as polo like kinase 1, mammalian target of rapamycin (mTOR),
Ras-related protein Rac1 (RAC1), isoprenylcystein carboxyl
methyltransferase (ICMT) and insulin like growth factor 1 receptor
(IGF1R), have been identified in HCC, which may be associated with
tumor progression, as well as poor prognosis, due to the
downregulation of miR-100-5p expression in patients with HCC
(12–15). However, the previous studies were
limited by small sample sizes and only several target genes of
miR-100-5p have been identified in HCC. The clinical value of
miR-100-5p in HCC and its molecular mechanisms are not fully
understood.
In the present study, to further explore the
clinical significance of miR-100-5p in HCC, the differential
expression of miR-100-5p between HCC tissues and non-cancerous
tissues was examined using meta-analysis based on the big data from
The Cancer Genome Atlas (TCGA), The Gene Expression Omnibus (GEO)
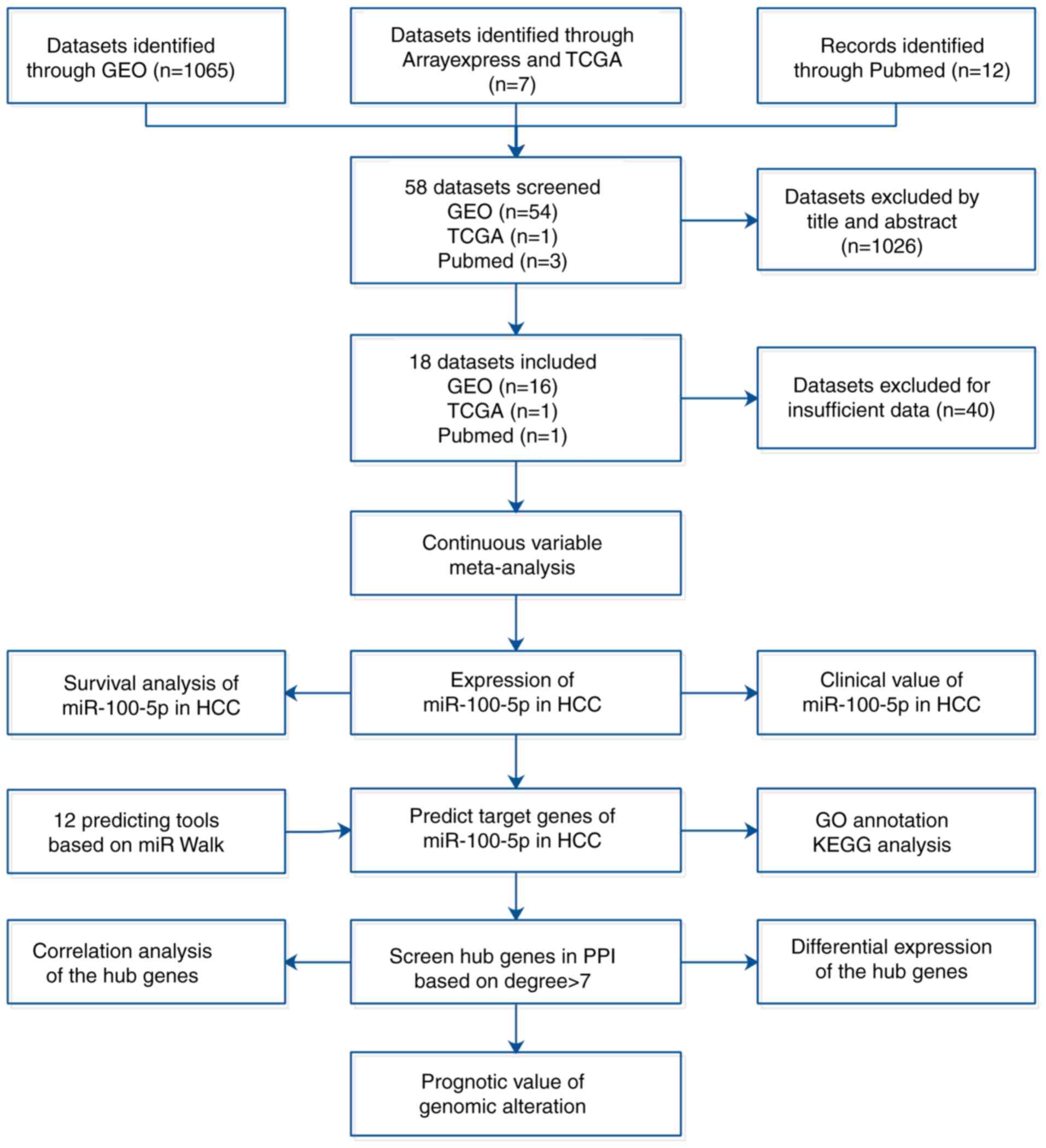
database and relevant literature (Fig.
1). The present study also attempted to reveal the
clinicopathological role and prognostic value of miR-100-5p using
TCGA. Bioinformatics analysis was carried out to investigate the
underlying mechanism of miR-100-5p in HCC, which may provide novel
therapeutic targets for HCC.
Materials and methods
Meta-analysis based on TCGA, GEO and
relevant literature
Datasets about HCC were downloaded from the TCGA
database (16) and were used to
determine the expression level of miR-100-5p. Subsequently, the
expression value of miR-100-5p was selected and converted using a
log2 transformation. The GEO database (ncbi.nlm.nih.gov/gds/) was used to screen for HCC
datasets with the keywords ‘(hepatocellular OR liver OR hepatoma OR
hepatic) AND (carcinoma OR cancer* OR adenocarcinoma OR tumor OR
malignan* OR neoplas*) AND (MicroRNA OR miRNA OR ‘Micro RNA’ OR mir
OR ‘Small Temporal RNA’ OR ‘non-coding RNA’ OR ncRNA OR ‘small
RNA’)’ until 7th January 2019. The inclusion criteria were as
follow: i) A least 3 HCC and noncancerous tissue samples, including
paracarcinoma tissue and normal liver tissue in the microarray; ii)
the expression of miR-100-5p was determined and calculated; and
iii) profiling obtained from Homo sapiens. miRNA-sequencing
(seq) data were excluded according to the following criteria: i)
The miRNA-seq did not meet the inclusion criteria; ii) the
miRNA-seq did not include the expression of miR-100-5p; iii) the
miRNA-seq only provided HCC tissues without noncancerous tissues;
and iv) microarrays used cell line samples. For additional
information about miR-100-5p in HCC, PubMed and ArrayExpress were
searched until 7th January 2019 using the following search
strategy: (Carcinoma OR cancer OR tumour OR tumor OR adenocarcinoma
OR neoplas* OR malignan*) AND (HCC OR hepatocellular OR hepatic OR
liver) AND (microRNA-100 OR miR-100 OR miRNA-100 OR miR100 OR
miRNA100 OR microRNA100 OR ‘miR 100’ OR ‘miRNA 100’ OR ‘microRNA
100’ OR miR-100-5p OR miRNA-100-5p OR microRNA-100-5p). The
inclusion criteria were as aforementioned. The number of samples,
mean and standard deviation between groups were obtained for a
comprehensive meta-analysis. Fixed effects, random effects models,
heterogeneity and sensitivity analysis were applied in the
meta-analysis using Stata SE12.0 software (Stata Corp LLC).
Publication bias was determined using Egger's plot and Begg's
funnel (17).
Exploring the clinical value of
miR-100-5p and its hub genes
The differential expression of miR-100-5p was
computed between HCC tissues and noncancerous tissues using
Student's unpaired t-test. The association between miR-100-5p
levels and clinicopathological features were then evaluated based
on the TCGA dataset using a Student's unpaired t-test. Patients
with HCC were divided into high and low expression groups according
to the median expression level. The prognostic value of miR-100-5p
and its hub genes in HCC patients (survival time >90 days) was
estimated using Kaplan-Meier analysis and log rank tests. Analysis
was carried out using GraphPad Prism 7.04 (GraphPad Software,
Inc.). P<0.05 was considered to indicate a significant
difference.
Predicting target genes of
miR-100-5p
miRWalk 2.0 (http://zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk2/)
provides an intuitive interface for generating predicted and
validated miRNA-binding sites for known genes in humans, mouse,
rat, dog and cow (18), and was used
to predict the target gees of miR-100-5p. The prospective target
genes of miR-100-5p were obtained from 12 different databases
(miRWalk2.0, miRanda-rel2010 (19),
DIANA-microTv5.0 (20), miRNAMap2.0
(21), miRBridge4.0 (22), miRMap4.0 (23), PicTar2 (24), PITA2007 (25), miRDB4.0 (26), RNAhybrid2.1 (27), RNA22v2 (28) and TargetScan6.1 (29). The target genes of miR-100-5p that
appeared in ≥4 different databases were selected for further
analysis.
GO and KEGG clustering analysis
DAVID 6.7 (https://david-d.ncifcrf.gov/), an online open platform
that disseminates biologically abundant information across a
comprehensive analysis of large gene lists, was used for clustering
analysis (30). Potential target
genes were searched in the annotated portal DAVID. Based on the
DAVID database, enrichment annotation was performed, using GO
(geneontology.org/) and KEGG (genome.jp/kegg/). Statistically significant GO and
KEGG terms (P<0.05) were selected.
Construction of a protein-protein
interaction (PPI) network
Cytoscape 3.7 (31)
was used to visualize the interaction networks of hub gene products
in HCC. A PPI network was constructed using the StringApp plugin,
which enabled Cytoscape 3.7 to link to the STRING database
(https://string-db.org/). Thereafter, the gene
symbols with a degree >7 were selected from the PPI network,
with the aim of identifying the most likely hub genes of miR-100-5p
in patients with HCC.
Correlation analysis
Gene expression data (log2
transformation) in HCC were downloaded from the TCGA database. In
order to reveal the association between miR-100-5p and its
candidate genes, Spearman's correlation analysis was performed, and
linear regression plots were constructed using GraphPad Prism
7.04.
Detection of the expression of hub
genes
The differential expression of the hub genes was
evaluated in patients with HCC based on the Roessler's (GSE14520)
(32) and Chen's dataset (GSE3500)
(33) from the GEO database.
Differences in the expression of the hub genes were visualized
using boxplots. In addition, the protein expression of hub genes
was verified using The Human Protein Atlas (proteinatlas.org), which provides a large number of
antibody-based images (34).
Detecting alterations in hub
genes
The cBio Cancer Genomics Portal (http://cbioportal.org), a public resource that
provides a convenient interface for accessing the multidimensional
cancer genomics and clinical data based on multiple platforms
(35), was used to detect
alterations in hub genes and to investigate the association between
alterations and outcomes for patient.
Determining alternative splicing of
hub genes
TCGA SpliceSeq (http://bioinformatics.mdanderson.org/TCGASpliceSeq) is
a database that enables investigators to explore the alternative
splicing events of various tumors based on TCGA data (36). In general, seven alternative splicing
events were classified, including alternate donor (AD) sites,
alternate terminators (ATs), alternate promoters (APs), alternate
acceptor (AA) sites, exon skipping (ES), retained introns (RIs) and
mutually exclusive exons (MEs) (37). Accordingly, the splicing events of
the hub genes in HCC were extracted from TCGA SpliceSeq. Percent
spliced in (PSI) values, an intuitive ratio for quantifying
splicing events (38), were then
compared between HCC and the normal tissues with Student's unpaired
t-test using GraphPad Prism 7.04.
Results
Differences in the expression of
miR-100-5p between groups
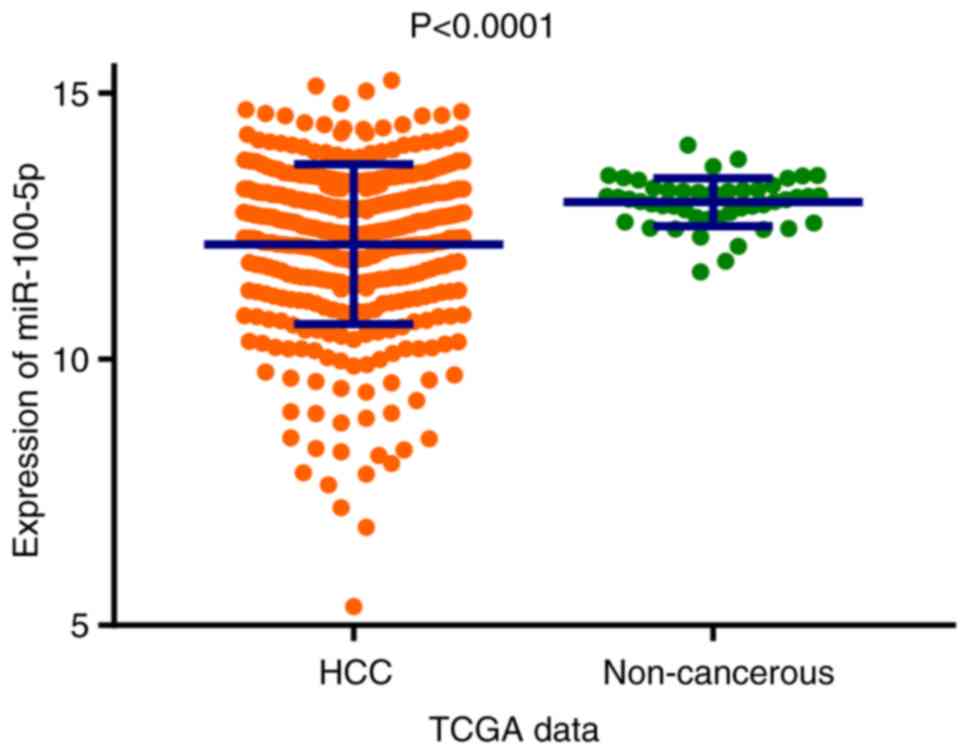
The scatter plot presented in Fig. 2 demonstrates that miR-100-5p
expression was significantly reduced in the HCC group compared with
the non-cancerous group based on TCGA data (P<0.0001). In total,
there were 16 microarray datasets, one TCGA study and one published
article (Table I), these included a
total of 1,258 samples (806 HCC and 452 non-cancerous tissues) for
the comprehensive meta-analysis (Fig.
3). The fixed effects model indicated that the pooled standard
mean difference (SMD) was −0.90 (95% CI, −1.04 to −0.77;
I2, 66.0%; P<0.001), while the pooled SMD was −1.00
(95% CI, −1.28 to −0.72; I2, 66.0%; P<0.001) in the
random effects model. Influence analysis revealed that two
microarrays (GSE22058 and GSE21362), the TCGA data and one article
may lead to heterogeneity. After removing the two microarrays
(GSE22058 and GSE21362), the TCGA data and the article, the pooled
SMD was −0.94 (95% CI, −1.14 to −0.74; I2, 35.2%;
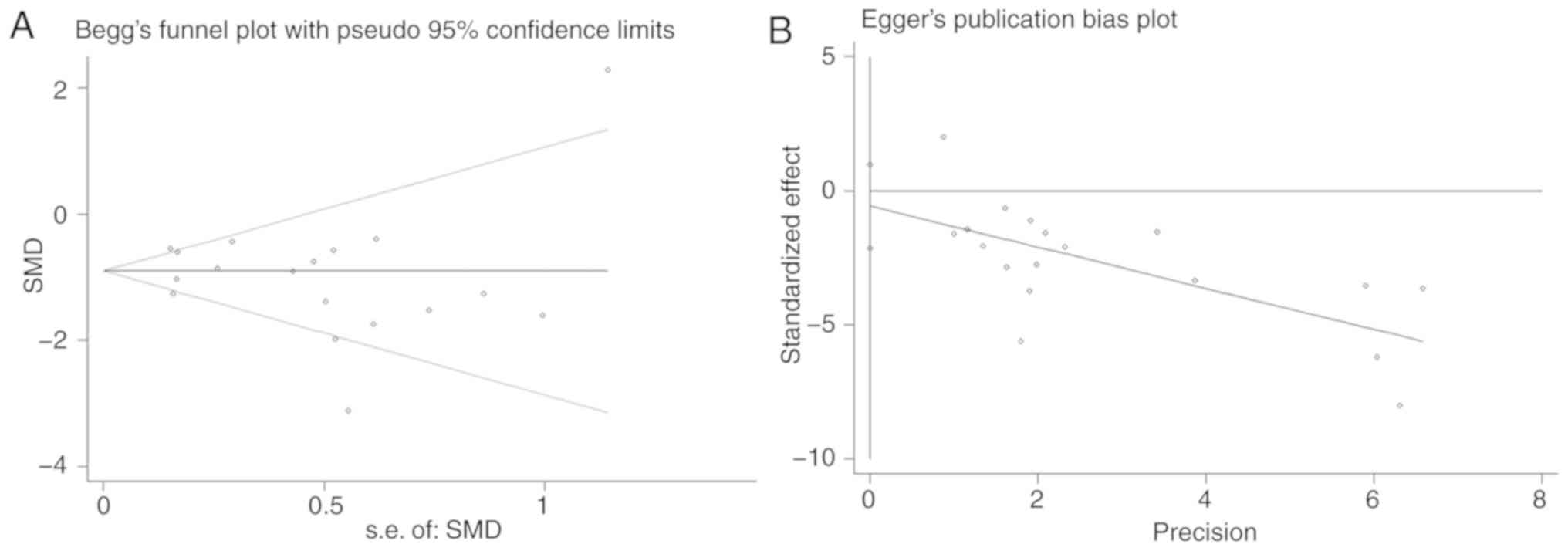
P=0.093). It was determined that there was no publication bias by
performing Begg's test (P=0.705) and Egger's test (P=0.443;
Fig. 4). The meta-analysis indicated
that the expression of miR-100-5p was significantly decreased in
HCC tissue compared with non-cancerous tissue.
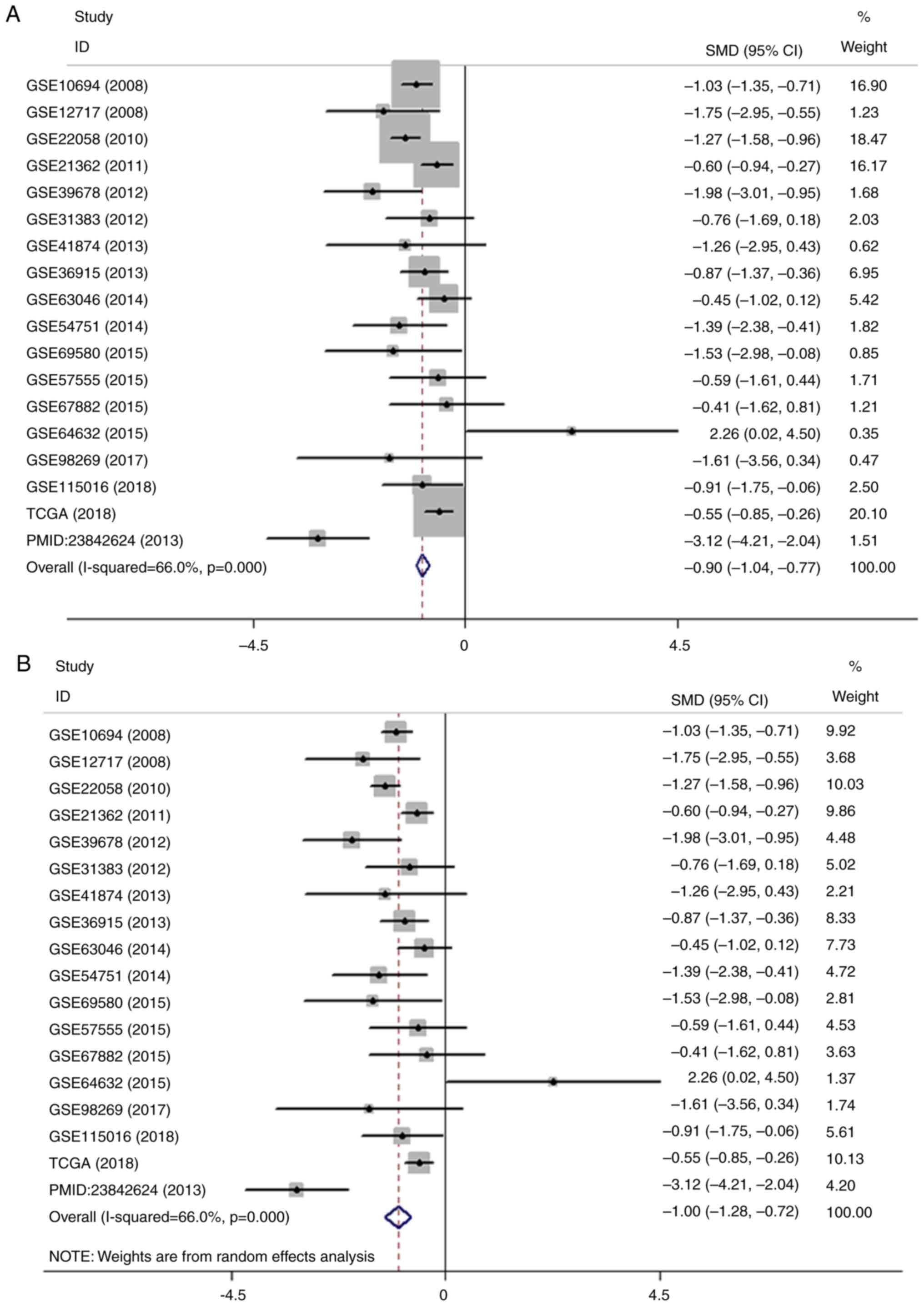 | Figure 3.Meta-analysis of the differential
expression of miR-100-5p in HCC and non-cancerous tissues. (A)
Forest plot of miR-100-5p expression datasets from Gene Expression
Omnibus, TCGA and an article. The fixed effects model indicated
that the pooled SMD was −0.90 (95% CI, −1.04 to −0.77;
I2, 66.0%; P<0.001. (B) Forest plot of the pooled SMD
of miR-100-5p was −1.00 (95% CI, −1.28 to −0.72; I2,
66.0%; P<0.001) using the random effects model. Meta-analysis of
the differential expression of miR-100-5p in HCC and non-cancerous
tissues. (C) Sensitivity analysis of the included studies. (D)
Forest plot with the data leading to heterogeneity removed. The
pooled SMD was −0.94 (95% CI, −1.14 to −0.74; I2, 35.2;
P=0.093). miR, microRNA; HCC, hepatocellular carcinoma; CGA, The
Cancer Genome Atlas; SMD, standard mean difference; CI, confidence
interval. |
 | Table I.Characteristics of the studies
included in the meta-analysis. |
Table I.
Characteristics of the studies
included in the meta-analysis.
| Author, year | Country | Series | Platform | HCC samples | Non-HCC
samples | (Refs.) |
|---|
| Li et al,
2008 | China | GSE10694 | GPL6542 | 78 | 88 | (89) |
| Su et al,
2009 | China | GSE12717 | GPL7274 | 10 | 6 | (90) |
| Burchard et
al, 2010 | USA | GSE22058 | GPL10457 | 96 | 96 | (91) |
| Sato et al,
2011 | Japan | GSE21362 | GPL10312 | 73 | 73 | (92) |
| Noh et al,
2013 | South Korea | GSE39678 | GPL15852 | 16 | 8 | (93) |
| Wang et al,
2012 | USA | GSE31383 | GPL10122 | 9 | 10 | (94) |
| Morita et
al, 2016 | Japan | GSE41874 | GPL7722 | 3 | 4 | (95) |
| Shih et al,
2012 | Taiwan | GSE36915 | GPL8179 | 68 | 21 | (96) |
| Wojcicka et
al, 2014 | Poland | GSE63046 | GPL11154 | 24 | 24 | (97) |
| Shen et al,
2014 | USA | GSE54751 | GPL18262 | 10 | 10 | (98) |
| Lou et al,
2019 | Taiwan | GSE69580 | GPL10850 | 5 | 5 | (99) |
| Murakami et
al, 2015 | Japan | GSE57555 | GPL18044 | 5 | 16 | (100) |
| Ghosh et al,
2016 | India | GSE67882 | GPL10850 | 4 | 8 | (101) |
| Peng et al,
2015 | USA | GSE64632 | GPL18116 | 3 | 3 | (102) |
| Zhang et al,
2017 | China | GSE98269 | GPL20712 | 3 | 3 | (103) |
| Shi et al,
2018 | China | GSE115016 | GPL21572 | 12 | 12 | (104) |
| TCGA, 2018 | N/A | N/A | N/A | 372 | 50 | – |
| Chen et al,
2013 | China | N/A | N/A | 15 | 15 | (12) |
Clinical value of miR-100-5p in HCC
with TCGA data
The association between the expression of miR-100-5p
and the clinicopathological features of patients with HCC was
investigated. The clinicopathological parameters of the patients
with HCC are presented in Table II.
Significance was determined for the following clinicopathological
parameters: Age (P=0.049), tumor stage (P=0.022), tumor node
metastasis (TNM) stage (P=0.020), tumor grade (P<0.001),
non-alcoholic fatty liver disease (NAFLD; P<0.001) and
α-fetoprotein (AFP) level (P<0.001). However, there was no
statistical significance in liver cirrhosis, sex, lymph node stage,
metastasis stage, Child-Pugh classification grade, hepatitis B
virus or hepatitis C virus infection and alcohol consumption.
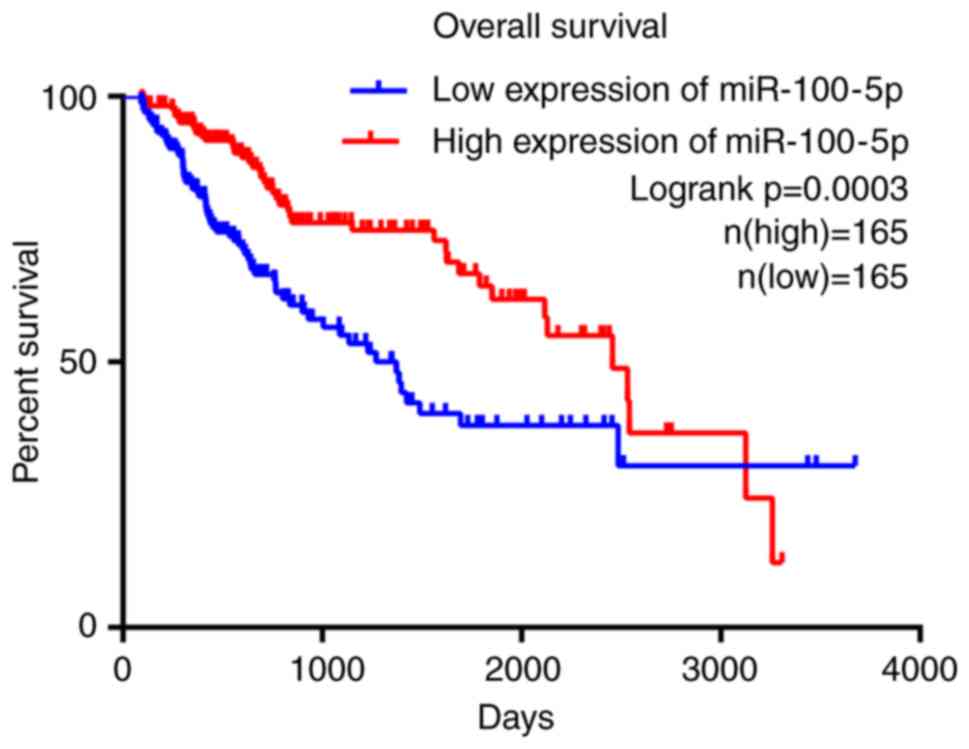
Furthermore, survival analysis suggested that patients with HCC and
lower expression levels of miR-100-5p exhibited significantly less
favorable OS times (P=0.0003) compared with patients with higher
expression levels of miR-100-5p (Fig.
5). The median survival of patients with high and low
miR-100-5p expression were 2456 and 1372 days, respectively,
indicating that miR-100-5p overexpression is associated with a
better clinical outcome for patients with HCC.
 | Table II.Association of miR-100-5p expression
and clinicopathological parameters in HCC. |
Table II.
Association of miR-100-5p expression
and clinicopathological parameters in HCC.
| Parameters | n | miR-100-5p
expression (2−ΔCq), mean ± SD | t | P-value |
|---|
| Sex |
|
|
|
|
|
Male | 253 | 12.2301±1.4228 | 1.330 | 0.185 |
|
Female | 119 | 11.9950±1.6634 |
|
|
| Age, years |
|
|
|
|
|
<65 | 222 | 12.0354±1.5341 | −1.979 | 0.049 |
|
≥65 | 149 | 12.3490±1.4388 |
|
|
| T |
|
|
|
|
|
T1+T2 | 276 | 12.2602±1.4053 | 2.313 | 0.022 |
|
T3+T4 | 93 | 11.8000±1.7363 |
|
|
| N |
|
|
|
|
|
Yes | 4 | 10.9563±2.2066 | 1.575 | 0.116 |
| No | 254 | 12.1331±1.4717 |
|
|
| M |
|
|
|
|
|
Yes | 4 | 12.1561±1.1603 | −0.092 | 0.927 |
| No | 269 | 12.0859±1.5263 |
|
|
| Stage |
|
|
|
|
|
I+II | 258 | 12.2622±1.4063 | 2.355 | 0.020 |
|
III+IV | 90 | 11.7777±1.7662 |
|
|
| Grade |
|
|
|
|
|
G1+G2 | 231 | 12.4402±1.4008 | 4.964 | <0.001 |
|
G3+G4 | 137 | 11.6580±1.5585 |
|
|
| Child-Pugh |
|
|
|
|
| A | 220 | 12.2815±1.4836 | −0.548 | 0.584 |
|
B+C | 22 | 12.4645±1.6079 |
|
|
| HBV/HCV |
|
|
|
|
|
Yes | 156 | 12.2373±1.4439 | −0.530 | 0.596 |
| No | 197 | 12.1527±1.5251 |
|
|
| Alcohol |
|
|
|
|
|
Yes | 117 | 12.2012±1.4718 | 0.098 | 0.922 |
| No | 236 | 12.1846±1.4994 |
|
|
| NAFLD |
|
|
|
|
|
Yes | 19 | 12.9891±0.6046 | 5.229 | <0.001 |
| No | 334 | 12.1447±1.5110 |
|
|
| AFP (ng/ml) |
|
|
|
|
|
<400 | 216 | 12.3754±1.4072 | 4.071 | <0.001 |
|
≥400 | 65 | 11.3984±1.7744 |
|
|
| Cirrhosis |
|
|
|
|
|
Yes | 80 | 12.3822±1.3630 | −0.568 | 0.571 |
| No | 135 | 12.2631±1.5524 |
|
|
Potential target genes of
miR-100-5p
The target genes of miR-100-5p were identified using
the following 12 target gene prediction platforms: miRWalk,
miRanda, miRNAMap, Microt4, miRBridge, PicTar, PITA, miRMap, miRDB,
RNAhybrid, RNA22 and TargetScan. In total, 447 candidate hub genes
(Table SI) for miR-100-5p were
predicted by at least 4 of the 12 online platforms used in the
present study.
GO annotation and KEGG pathway
GO and KEGG annotation of the 447 potential hub
genes of miR-100-5p was conducted using the online platform DAVID.
GO annotation revealed that a total of 99 terms were significantly
enriched (P<0.05) through the candidate genes identified. KEGG
analysis identified four signaling pathways that were significantly
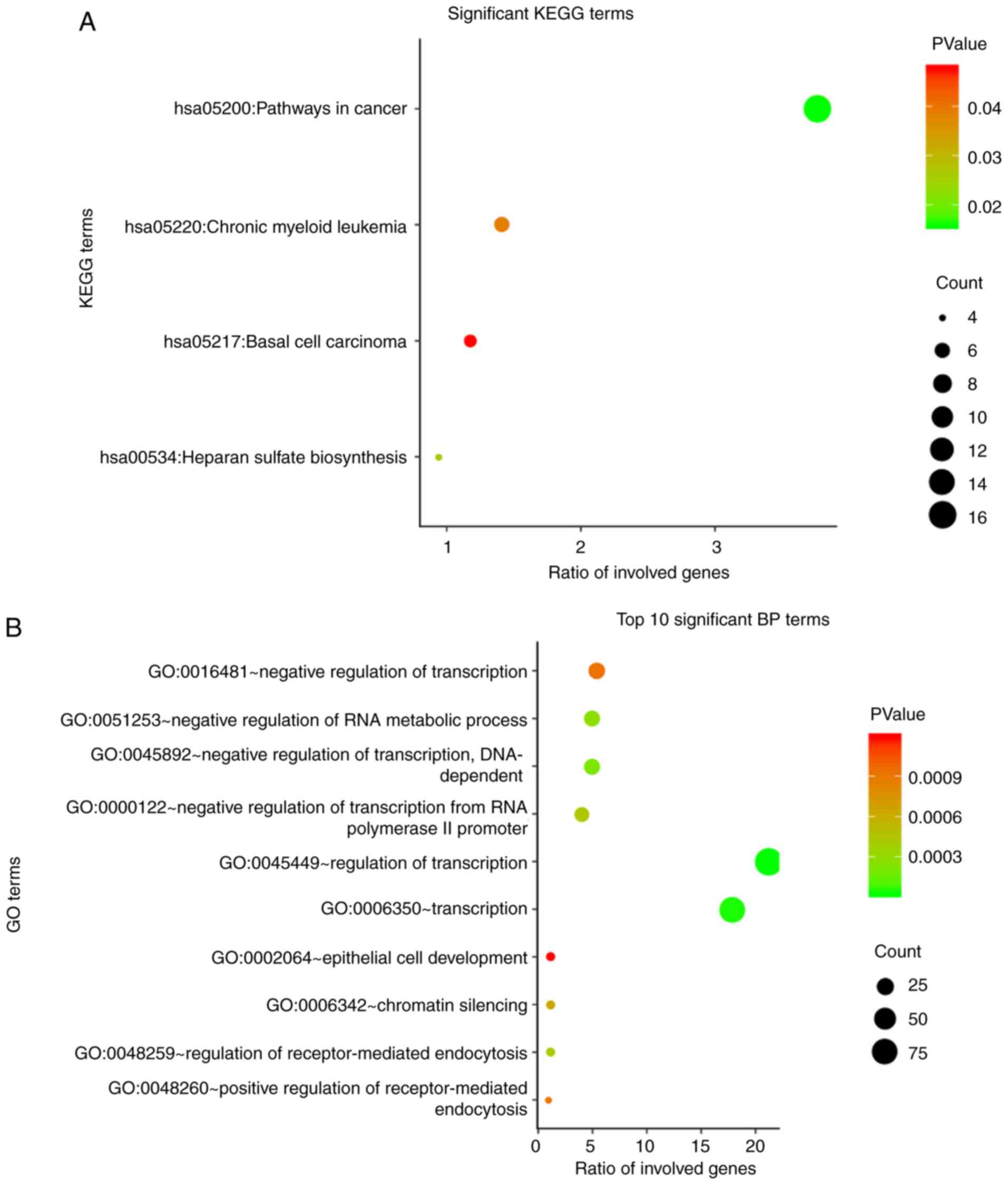
enriched. The top 10 enriched GO terms and the significant KEGG
pathways, which may contribute to targeted therapy for patients
with HCC, are presented in Fig. 6.
GO annotation showed that the three most significant terms for
biological processes (BPs) were ‘regulation of transcription’,
‘transcription’, and ‘negative regulation of transcription,
DNA-dependent’ (Fig. 6B). For
cellular components (CCs), the candidate genes most significantly
enriched were ‘chromatin remodeling complex’, ‘membrane fraction’
and ‘plasma membrane part’ (Fig.
6C). In molecular functions (MFs), ‘transcription regulator
activity’, ‘DNA binding’ and ‘ion binding’ were the terms most
commonly associated with the target genes (Fig. 6D). The two most significant KEGG
pathways identified were involved in cancer and heparan sulfate
biosynthesis (Fig. 6A).
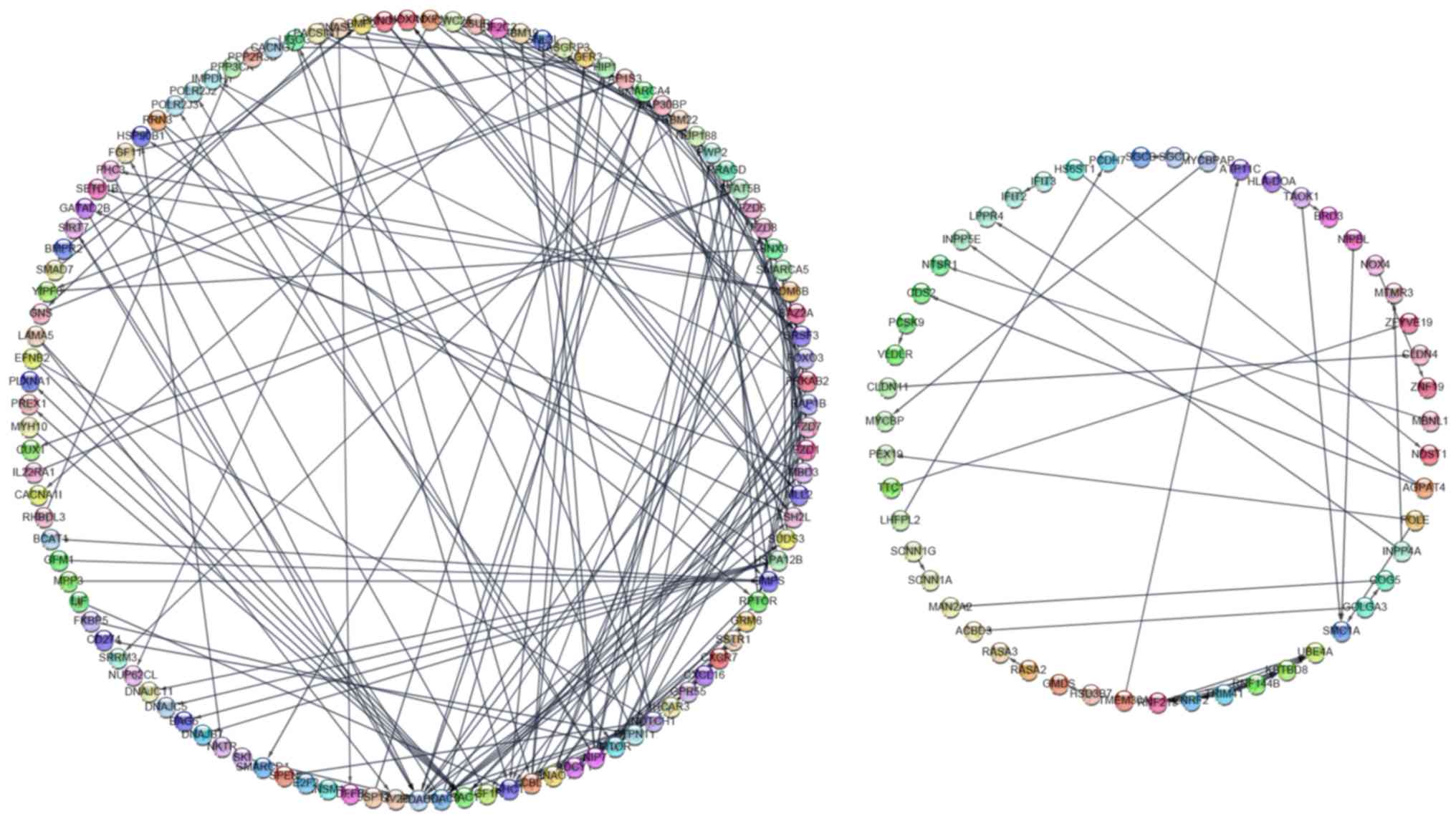
PPI network analysis
A PPI network was constructed, consisting of 158
nodes and 229 lines, by inputting a total of 447 potential target
genes into String App (Fig. 7). Each
protein may interact with multiple proteins, which may form the
underlying molecular regulatory mechanism of miR-100-5p in HCC. In
total, the following 6 hub genes were identified based on a degree
>7 in the PPI network: Histone deacetylase (HDAC)2, HDAC3, SHC
transforming protein 1 (SHC1), RAC1, IGF1R and E3-unbiqitin protein
ligase CBL (CBL). The six key genes may exert an important function
in the regulatory mechanism of miR-100-5p in HCC.
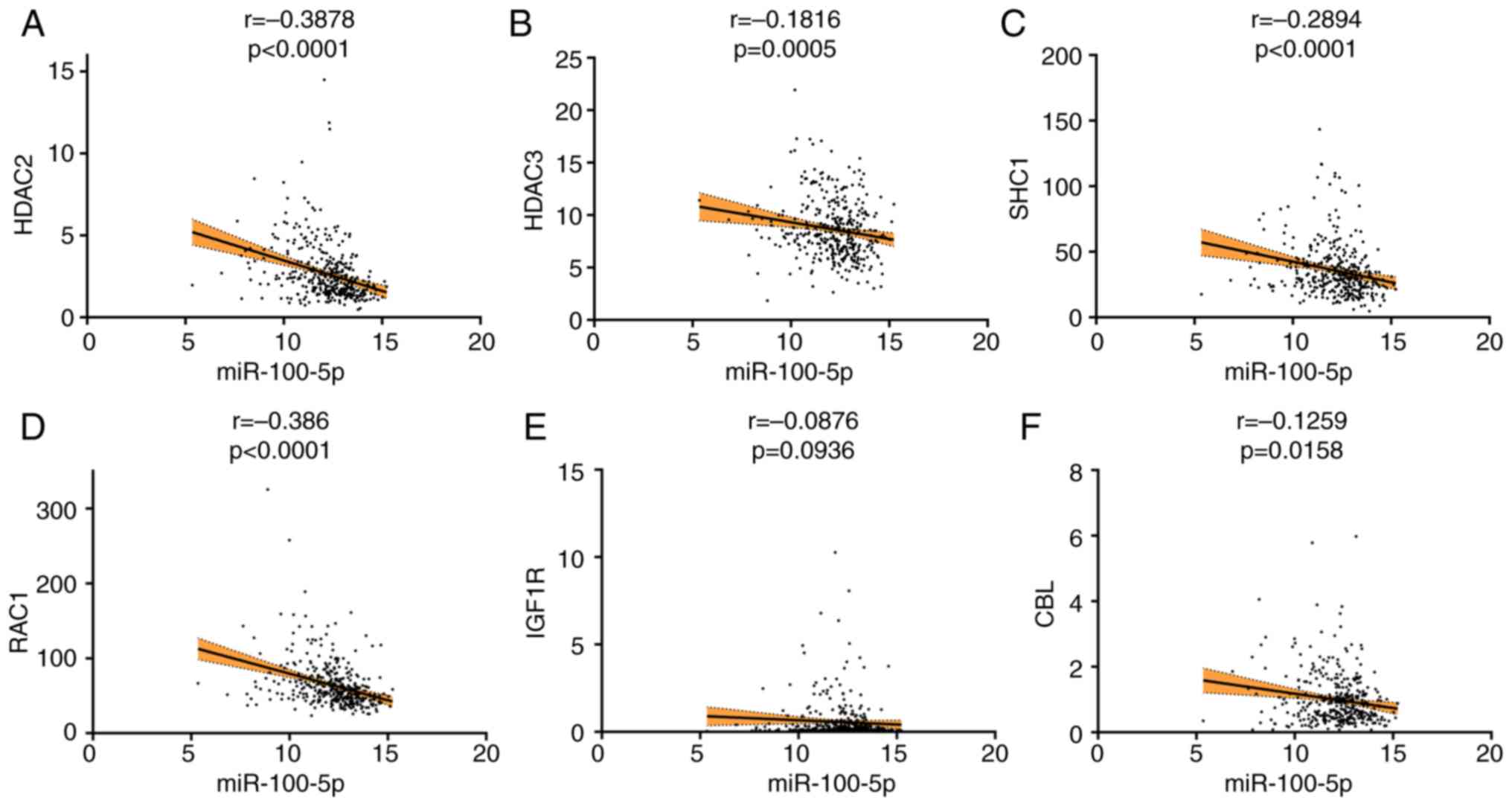
Spearman correlation analysis
Spearman correlation analysis revealed that HDAC2
(r= −0.3878; P<0.0001), HDAC3 (r= −0.1816; P=0.0005), SHC1 (r=
−0.2894; P<0.0001), RAC1 (r= −0.386; P<0.0001) and CBL (r=
−0.1259; P=0.0158) were inversely associated with miR-100-5p
expression. IGF1R (r= −0.0876; P=0.0936) exhibited a trend towards
negative correlation with miR-100-5p expression, although this was
not statistically significant (Fig.
8).
Protein levels of hub genes
The protein levels of HDAC2 (antibody, CAB005054),
HDAC3 (antibody, CAB005583), SHC1 (antibody, CAB016305), RAC1
(antibody, CAB035994), IGF1R (antibody, CAB010268) and CBL
(antibody, HPA027956) in HCC and noncancerous tissue were obtained
from the Human Protein Atlas database. As presented in Fig. 9, HDAC2, HDAC3, SHC1 and IGF1R
exhibited high staining and strong intensity in HCC. RAC1 and CBL
exhibited medium staining and an average intensity in HCC. However,
in normal liver tissue, IGF1R and SHC1 exhibited moderate and low
staining, respectively. The protein expression levels of HDAC2,
HDAC3, RAC1 and CBL were below the limit of detection in normal
liver tissue.
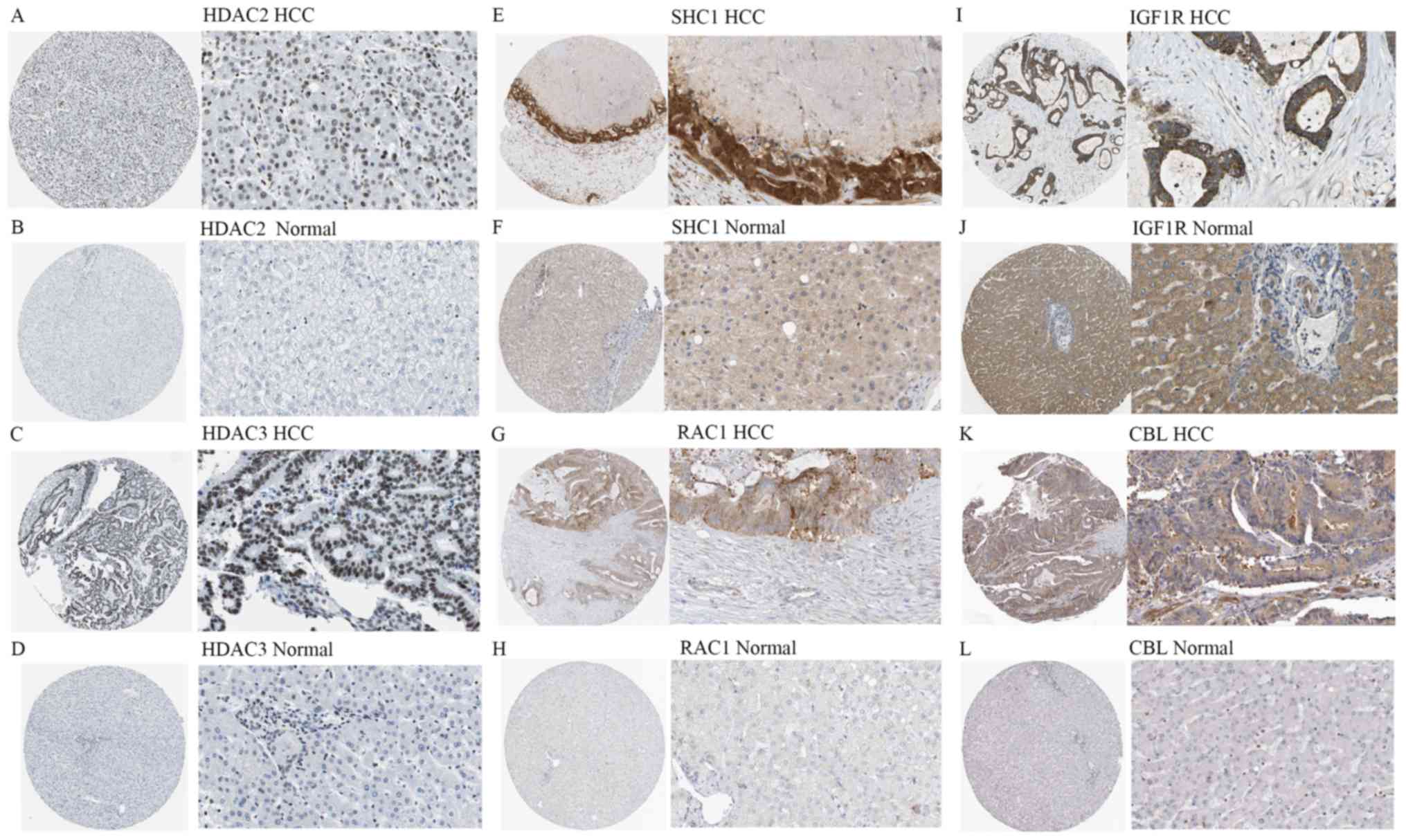 | Figure 9.Immunohistochemical staining
(magnification, ×40 or ×400) for protein expression of the 6 hub
genes of miR-100-5p. (A) High staining and strong intensity of
HDAC2 in HCC (antibody, CAB005054). (B) Low expression of HDAC2 in
normal liver tissue (antibody, CAB005054). (C) High staining and
strong intensity of HDAC3 in HCC (antibody, CAB005583). (D) Low
expression of HDAC3 in normal liver tissue (antibody, CAB005583).
(E) High staining and strong intensity of SHC1 in HCC (antibody,
CAB016305). (F) Low expression of SHC1 in normal liver tissue
(antibody, CAB016305). (G) Medium staining and average intensity of
RAC1 in HCC (antibody, CAB035994). (H) Low expression of RAC1 in
normal liver tissue (antibody, CAB035994). (I) High staining and
strong intensity of IGF1R in HCC (antibody, CAB010268). (J) Medium
staining of IGF1R in normal liver tissue (antibody, CAB010268). (K)
Medium staining and average intensity of CBL in HCC (antibody,
HPA027956). (L) Low expression of CBL expression in normal liver
tissue (antibody, HPA027956). HCC, hepatocellular carcinoma; miR,
microRNA; HDAC, histone deacetylate; SHC1, SHC-transforming protein
1; RAC1, Ras-related protein Rac1; IGF1R, insulin like growth
factor 1 receptor; CBL, E3 ubiquitin-protein ligase CBL. |
Expression and prognostic significance
of hub genes
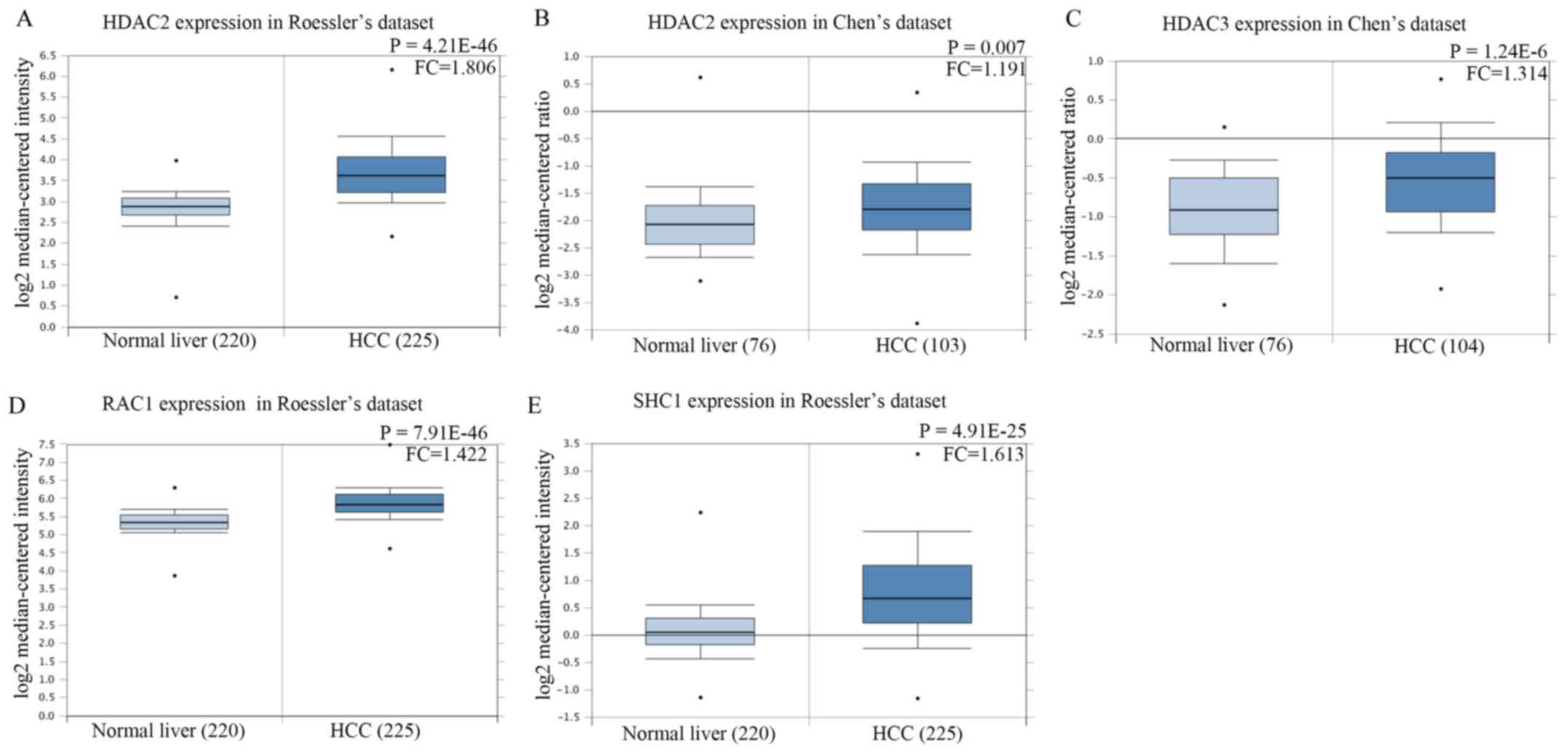
Of the 6 hub genes, 4 genes (HDAC2, HDAC3, SHC1 and
RAC1) exhibited overexpression according to Roessler's dataset and
Chen's dataset from the GEO database. As shown in Fig. 10A and B, the expression levels of
HDAC2 in both Roessler's dataset [P=4.21×10−46; fold
change (FC), 1.806] and Chen's dataset (P=0.007; FC, 1.191) were
significantly upregulated in HCC tissue compared with the normal
liver tissue. The expression of HDAC3 in Chen's dataset was
significantly higher in HCC tissues compared with the normal liver
tissue (Fig. 10C;
P=1.24×10−6; FC, 1.314). The expression of RAC1
(P=7.91×10−46; FC, 1.422) and SHC1
(P=4.91×10−25; FC, 1.613) in Roessler's dataset were
significantly increased in HCC tissues compared with the normal
liver tissue (Fig. 10D and E). As
miR-100-5p is downregulated in HCC, the genes with increased
expression in HCC are potential target genes of miR-100-5p. As
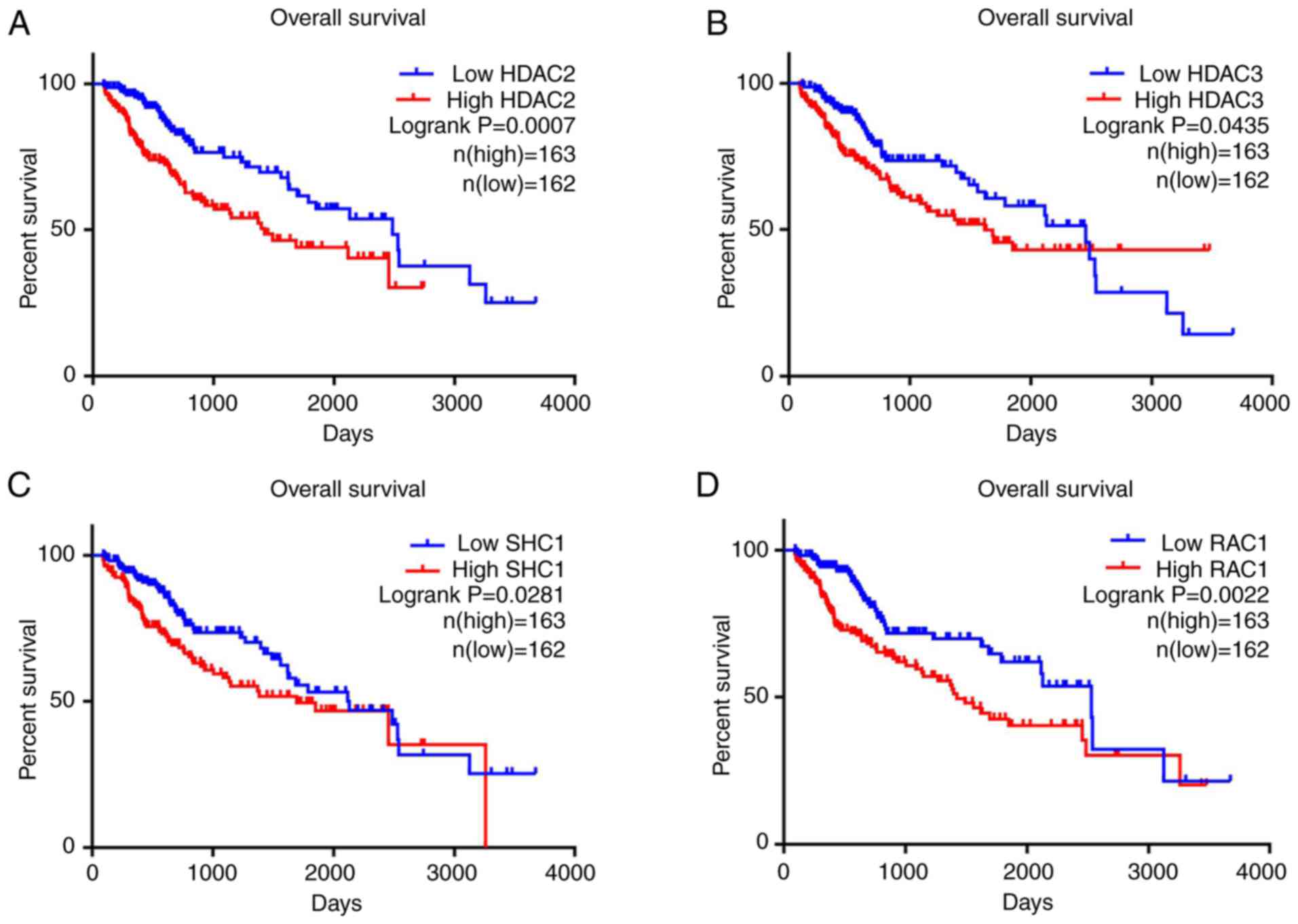
presented in Fig. 11, it was found
that higher expression levels of HDAC2 (HR, 1.910; 95% CI,
1.309–2.787; P=0.0007), HDAC3 (HR, 1.474; 95% CI, 1.012–2.146;
P=0.0435), SHC1 (HR, 1.52; 95% CI, 1.043–2.215; P=0.0281) and RAC1
(HR, 1.817; 95% CI, 1.248–2.645; P=0.0022) were significantly
associated with worse OS.
Alterations of hub genes
Gene alteration analysis of the 6 hub genes in the
360 TCGA patients were analyzed using the cBioPortal database, this
analysis indicated that the main types of gene alteration were mRNA
upregulation and amplification. In total 7, 11, 21, 12, 11 and 6%
of cases had genetic alterations in HDAC2, HDAC3, SHC1, RAC1, IGF1R
and CBL, respectively (Fig. S1).
The alterations in HDAC2 (log rank P=0.0213), SHC1 (log rank
P=6.153×10−3) and RAC1 (log rank P=0.0144) were
associated with a poorer OS in patients with HCC from the TCGA
dataset, while the alterations that occurred in HDAC3, IGF1R and
CBL were not significantly associated with OS (Fig. S2). The changes in HDAC2 (log rank
P=6.488×10−3), SHC1 (log rank P=0.0347) and IGF1R (log
rank P=6.991×10−3) were associated with unfavorable
disease-free survival (DFS) in patients with HCC from the TCGA
dataset, whereas alterations in HDAC3, RAC1 and CBL were not
significantly associated with DFS (Fig.
S3).
Alternative splicing of hub genes
In total, the splicing events of 4 genes (HDAC2,
HDAC3, SHC1 and RAC1) were analyzed. To define a splicing event
accurately, each splicing event was named with a unique code in the
present study. For example, for the code SHC1-7856-AA, SHC1 is the
gene symbol, 7856 denotes the order number of the splicing event in
the dataset and AA indicates the type of splicing. The results
showed that alternative splicing event SHC1-7856-AA was
significantly decreased in HCC (P<0.0001), while RAC1-78720-ES
was significantly increased (P=0.0253; Fig. S4). However, no statistically
significant alternative splicing events were identified in HDAC2
and HDAC3 (P>0.05). These findings may facilitate the
understanding of the potential molecular mechanisms of the target
genes.
Discussion
HCC is the most common type of primary liver cancer
(1); however, predicting the outcome
for patients remains a challenge. Identifying prospective
biomarkers and understanding the underlying mechanisms of HCC may
provide novel therapeutic, prognostic and monitoring strategies for
HCC. The present study aimed to identify hub genes and signaling
pathways regulated by miR-100-5p, and to elucidate the role of this
miRNA in HCC. It has been reported that miR-100-5p functions in
diverse human cancers. For example, previous studies reported that
miR-100-5p could reverse cisplatin resistance in breast cancer,
lung cancer and ovarian cancer (9,39–41).
Previous studies have also reported that miR-100 may influence the
metastatic potential of various cancers, including prostate cancer,
gastric cancer and nasopharyngeal carcinoma (10,11,42,43). It
has been proposed that high levels of miR-100-5p are associated
with a longer survival time in patients with various types of
cancer, including glioblastoma and epithelial ovarian cancer
(44,45). In two studies by Zhou et al
(13,14), miR-100-5p was found to target mTOR
and block the mTOR-p70S6K signaling pathway in order to
downregulate the protein level of angiopoietin 2, thus abrogating
the vessels that encapsulated tumor cluster-dependent metastasis of
hepatoma cells. The decreased expression of miR-100-5p can enhance
ICMT-RAC1 signaling and promote the metastasis of HCC cells
(13,14). Furthermore, Ge et al (15) reported that miR-100-5p may reduce
mTOR and IGF-1R levels by promoting the autophagy of hepatocellular
carcinoma cells. A study by Petrelli et al (46) revealed that dysregulation of
miR-100-5p was likely to be an early event in the development of
hepatocarcinogenesis. However, the biological functions and
mechanisms of miR-100-5p expression in HCC are still not completely
understood.
To the best of our knowledge, only 3 studies
(12,13,46) have
reported that the expression of miR-100-5p is decreased in HCC,
with 2 of these 3 studies (12,13)
indicating that low expression of miR-100-5p is associated with
clinicopathological features in patients with HCC. However,
Varnholt et al (47) reported
that the level of miR-100-5p was significantly increased in HCC
tissues compared with normal tissues. Similarly, the upregulation
of miR-100-5p in plasma or serum samples was demonstrated by Wang
et al (48). As
aforementioned, the expression of miR-100-5p in patents with HCC
remains controversial. Therefore, the present study conducted a
meta-analysis to understand whether miR-100-5p expression is
decreased in HCC samples compared with its expression in non-HCC
samples. In the present study, it was found that miR-100-5p was
downregulated in HCC based on a total of 1,258 samples from TCGA,
GEO and relevant articles. Reduced expression of miR-100-5p was
associated with poorer OS and worse clinical parameters compared
with normal miR-100-5p expression. Although the nature of the
present study was similar to a study conducted by Chen et al
(12), the findings of the present
study were novel and provided further information as follows: i)
The present study detected differences in the expression of
miR-100-5p with a larger sample size (1,258 samples) based on the
meta-analysis; ii) the present study suggested that miR-100-5p may
be used to predict the progression of HCC due to the close
associated between miR-100-5p expression and clinicopathological
features, including age, tumor stage, TNM stage, tumor grade, NAFL
and AFP level; and iii) the results of the current study suggested
that miR-100-5p may serve as a reliable biomarker, with higher
expression of miR-100-5p indicating a better OS. Further
investigations focusing on the association between miR-100-5p and
HCC are required. In the present study bioinformatics analysis was
performed to identify the potential molecular mechanisms of
miR-100-5p in HCC. For BPs, the top five related GO terms were
‘regulation of transcription’, ‘transcription’, ‘negative
regulation of transcription’, ‘regulation of receptor-mediated
endocytosis’ and ‘DNA-dependent, negative regulation of RNA
metabolic process’. For CCs, the top five statistically significant
terms were ‘chromatin remodeling complex’, ‘membrane fraction’,
‘plasma membrane part’, ‘insoluble fraction’ and ‘nucleoplasm’. For
MFs, the top 5 terms were ‘transcription regulator activity’,
‘transcription factor activity’, ‘DNA binding’, ‘ion binding’ and
‘metal ion binding’. These results suggest the target genes
primarily functioned in transcriptional regulation by binding to
chromatin, DNA or other biological molecules, thus affecting the
occurrence and development of HCC. For the KEGG pathway analysis,
the two most statistically significant pathways were ‘pathways in
cancer’ and ‘heparan sulfate biosynthesis’. Heparan sulfate is a
linear polysaccharide that modulates numerous biological processes,
including cell growth, angiogenesis and adhesion (49–51). A
study by Cassinelli et al (52) found that the heparanase/heparan
sulfate proteoglycan axis may be a potential novel therapeutic
target in sarcomas. Ling et al (53) suggested that blocking heparan sulfate
function in the heparin-binding domains of fibroblast growth factor
receptor 1 inhibited growth of cancer cells. Dudás et al
(54) showed that heparin and
heparan sulfate may interfere with the function of topotecan in
liver and liver cancer. Given these previous findings, it was
hypothesized that miR-100-5p may exert a regulatory function on the
biosynthesis of heparan sulfate, which may provide potential
candidate targets for drug development in HCC. However, this should
be verified in the future with experimental studies.
Additionally, 6 possible candidate genes, HDAC2,
HDAC3, SHC1, RAC1, IGF1R and CBL, were identified by constructing a
PPI network and were selected as they had a degree >7. The
expression of HDAC2, HDAC3, SHC1 and RAC1 were significantly
increased in HCC tissue compared with non-cancerous tissue.
Survival analysis showed that the upregulation of HDAC2, HDAC3,
SHC1 and RAC1 were associated with poorer outcome for patients with
HCC.
The proteins encoded for by HDAC2 and HDAC3 belonged
to the histone deacetylase family (55,56).
Previous studies have reported that these genes are involved in
various BPs, such as cell cycle progression and transcriptional
regulation (57–59). HDAC2 and HDAC3 have been found to be
closely associated with the occurrence and survival rate in various
cancer types, including colon cancer, breast cancer and HCC
(60–64). The aforementioned studies also
support the findings of the present study in which high expression
of HDAC2 and HDAC3 was found to indicate a poorer prognosis for
patients with HCC. To the best of our knowledge, the association
between miR-100-5p and the two genes, HDAC2 and HDAC3, has not
previously been identified. In the present study it was reported
that miR-100-5p is closely linked with HDAC2 and HDAC3, however,
experiments should be performed to support these findings.
The gene encoding SHC1 encodes three main isoforms,
including p66Shc, p52Shc and p46Shc. The p66Shc isoform may
participate in modulating lifetime and the effects of reactive
oxygen species (65). The other two
isoforms, p52Shc and p46Shc, may be involved in the transforming
activity of oncogenic tyrosine kinases (66,67).
Recently, various studies have found that this gene is closely
related to a variety of cancer types. In one previous study, Zhang
et al (68) found that p66Shc
was highly expressed in colon cancer tissue, and knockdown of
p66Shc in HCT8 cells reduced proliferation and accelerated
apoptosis. A study by Yukimasa et al (69) also suggested that increased
expression of p46Shc and p52Shc may be related to the occurrence of
gastric cancer. In addition, Muniyan et al (70) reported that p66Shc was highly
expressed in ovarian cancer cells compared with noncancerous cells,
and that it may regulate the proliferation of human ovarian cancer
cells. Furthermore, a previous study also identified that elevated
expression of p66Shc was associated with the advance and metastasis
of prostate cancer (71). At
present, to the best of our knowledge, studies focusing on the
regulatory mechanism of SHC1 in HCC are scarce. Yoshida et
al (72) reported that the
upregulation of SHC1 was closely associated with shorter survival
in patients with HCC patients. In the present study, it was found
that the expression of SHC1 was significantly increased in HCC
tissue compared with non-cancerous tissue, and was closely
associated with the outcome of patients with HCC. In the enriched
GO terms, SHC1 was found to be strongly associated with ‘regulation
of cell proliferation’, ‘positive regulation of cellular
biosynthetic process’ and ‘positive regulation of macromolecule
biosynthetic process’. Therefore, it was hypothesized that
miR-100-5p may target SHC1 to regulate cell proliferation and
biosynthetic processes, thus contributing to the progression of
HCC. However it is necessary to conduct experiments in vivo
and in vitro to validate this hypothesis.
RAC1 is important for a variety of cellular
functions, including proliferation, adhesion, motility, migration
and metastasis of tumor cells (73).
Recently, Cheng et al (74)
reported that RAC1 was upregulated in hypopharyngeal squamous cell
carcinoma (HSCC) tissues, and that the silencing of RAC1 could
reduce the invasion and migration abilities of HSCC cells.
Consistently, the upregulation of RAC1 has been reported to be
associated with poorer outcomes for patients with breast cancer
(75). RAC1 inhibition may prevent
metastasis and augment chemotherapy in gastric adenocarcinoma cells
by blocking epithelial to mesenchymal transition (EMT) and cancer
stem cell phenotypes (76). A
previous study found that downregulation of miR-100-5p enhanced
ICMT/RAC1 signaling and promoted the progression of HCC cells
(13). In the present study, it was
found that the expression of RAC1 was significantly higher in HCC
tissue compared with normal tissue, and was inversely related to
the expression of miR-100-5p. According to bioinformatics analysis,
RAC1 was enriched in various BPs, such as ‘regulation of
receptor-mediated endocytosis’, ‘cell morphogenesis’, ‘cellular
component morphogenesis’ and ‘epithelial cell differentiation’.
These BPs play important roles in the growth, migration and
differentiation of tumor cells, thereby contributing to the
occurrence and development of cancer (77–80).
Additionally, the data in the present study revealed that the
upregulation of RAC1 was associated with poorer OS in patients with
HCC. Therefore, it was hypothesized that the miR-100-5p/RAC1
signaling pathway may be important in the metastasis of HCC, and
hence, may influence prognosis of patients with HCC. This indicates
that interfering with the miR-100-5p/RAC1 signaling pathway may
provide a novel treatment strategy for HCC. Nevertheless, further
studies are required to support this hypothesis.
A number of previous studies have suggested that
genetic alterations may affect the occurrence and progression of
tumor (81–83). Jonckheere et al (84) reported that K-RAS mutation was the
earliest genetic alteration occurring in pancreatic ductal
adenocarcinoma, which could alter the expression of miRNA. In the
present study, alterations in 6 key hub genes (HDAC2, HDAC3, SHC1,
RAC1, IGF1R and CBL) were identified. The main alterations
identified in the hub genes were amplification and mRNA
upregulation. Alterations in HDAC2, SHC1, RAC1 and IGF1R were
closely associated with a poorer outcome for patients with HCC. It
is anticipated that these genetic alterations may be associated
with the downregulation of miR-100-5p, thus playing an important
role in the progression of HCC. However, further studies are
required.
Alternative splicing is a common mechanism for gene
regulation in humans, and it plays an important role in
tumorigenesis, progression and therapy (85,86).
Generally, miRNAs work by binding to the 3′untranslated regions
(UTRs) of their target genes (87).
However, other functions of miRNAs are being identified. For
example, Teplyuk et al (88)
revealed that miR-10b can modulate the splicing events of its
target genes by binding to 5′UTRs in intracranial glioblastoma
(88). In the present study, it was
found that the splicing events SHC1-7856-AA and RAC1-78720-ES were
significantly decreased and increased in HCC compared with normal
tissue, respectively, which may indicate that miR-100-5p is
associated with alternative splicing of SHC1 and RAC1. Overall, the
mechanism of how miR-100-5p influences its target genes warrants
further study.
Although the results of the present study suggest
that miR-100-5p may function as a tumor suppressor and be a
valuable prognostic biomarker for HCC, it should be emphasized that
several limitations exist in the present study. As only one
previous study (48) involving serum
samples could be found, no meta-analysis on circulating miRNAs
could be performed. The current study therefore was unable to
provide a minimally invasive method and early prognostic indicator
for patients with HCC. The sample population for the clinical
parameters was small, despite being larger than previous studies,
which was likely to limit the stability of results. In order to
comprehensively explore the potential value of miR-100-5p, this
study was extended to investigate numerous non-cancerous samples
(16). However, a subgroup analysis
is needed to reveal more accurate conclusions in future studies. In
addition, even though predictions were performed on an array of
candidate genes, only two genes (RAC1 and IGF1R) have been
experimentally verified (13,15).
More candidate genes must be verified with in vivo and in
vitro experiments. The nature of the present study was itself a
limitation and a prospective cohort study is expected to be
conducted to verify the functions of miR-100-5p in HCC.
In conclusion, miR-100-5p was significantly
downregulated in HCC based on the meta-analysis performed in the
present study. The downregulation of miR-100-5p was found to be
closely associated with a poorer prognosis and poorer clinical
characteristics, including NAFLD, in patients with HCC, which
indicated that miR-100-5p may function in the development and
progression of HCC through multiple biological mechanisms. The
overexpression of the 4 potential target genes (HDAC2, HDAC3, SHC1
and RAC1) of miR-100-5p were associated with poorer survival for
patients with HCC. Therefore, miR-100-5p and these potential target
genes may provide novel therapeutic strategies and biomarkers for
the diagnosis and prognosis of patients with HCC.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
QLH, HXJ and SYQ designed the study and wrote the
manuscript. QLH, LT and HJN performed the preprocessing of the data
and the analysis. HXJ and QLH critically revised the manuscript.
All authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
HCC
|
hepatocellular carcinoma
|
|
TCGA
|
The Cancer Genome Atlas
|
|
GEO
|
Gene Expression Omnibus
|
|
GO
|
Gene Ontology
|
|
KEGG
|
Kyoto Encyclopedia of Genes and
Genomes
|
|
PPI
|
protein-protein interaction
network
|
|
NAFLD
|
non-alcoholic fatty liver disease
|
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kudo M: Systemic therapy for
hepatocellular carcinoma: 2017 update. Oncology. 93 (Suppl
1):S135–S146. 2017. View Article : Google Scholar
|
|
3
|
Aguilar C, Mano M and Eulalio A: MicroRNAs
at the Host-bacteria interface: Host defense or bacterial offense.
Trends Microbiol. 27:206–218. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Mollaei H, Safaralizadeh R and Rostami Z:
MicroRNA replacement therapy in cancer. J Cell Physiol.
234:12369–12384. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Li TT, Gao X, Gao L, Gan BL, Xie ZC, Zeng
JJ and Chen G: Role of upregulated miR-136-5p in lung
adenocarcinoma: A study of 1242 samples utilizing bioinformatics
analysis. Pathol Res Pract. 214:750–766. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Gao L, Li SH, Tian YX, Zhu QQ, Chen G,
Pang YY and Hu XH: Role of downregulated miR-133a-3p expression in
bladder cancer: A bioinformatics study. Onco Targets Ther.
10:3667–3683. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
He D, Yue Z, Li G, Chen L, Feng H and Sun
J: Low serum levels of miR-101 are associated with poor prognosis
of colorectal cancer patients after curative resection. Med Sci
Monit. 24:7475–7481. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zheng Y, Tan K and Huang H: Long noncoding
RNA HAGLROS regulates apoptosis and autophagy in colorectal cancer
cells via sponging miR-100 to target ATG5 expression. J Cell
Biochem. 120:3922–3933. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wu G, Zhou W, Pan X, Sun Y, Xu H, Shi P,
Li J, Gao L and Tian X: miR-100 reverses cisplatin resistance in
breast cancer by suppressing HAX-1. Cell Physiol Biochem.
47:2077–2087. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sun X, Liu X, Wang Y, Yang S, Chen Y and
Yuan T: miR-100 inhibits the migration and invasion of
nasopharyngeal carcinoma by targeting IGF1R. Oncol Lett.
15:8333–8338. 2018.PubMed/NCBI
|
|
11
|
Shi DB, Wang YW, Xing AY, Gao JW, Zhang H,
Guo XY and Gao P: C/EBPα-induced miR-100 expression suppresses
tumor metastasis and growth by targeting ZBTB7A in gastric cancer.
Cancer Lett. 369:376–385. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen P, Zhao X and Ma L: Downregulation of
microRNA-100 correlates with tumor progression and poor prognosis
in hepatocellular carcinoma. Mol Cell Biochem. 383:49–58. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhou HC, Fang JH, Luo X, Zhang L, Yang J,
Zhang C and Zhuang SM: Downregulation of microRNA-100 enhances the
ICMT-Rac1 signaling and promotes metastasis of hepatocellular
carcinoma cells. Oncotarget. 5:12177–12188. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhou HC, Fang JH, Shang LR, Zhang ZJ, Sang
Y, Xu L, Yuan Y, Chen MS, Zheng L, Zhang Y and Zhuang SM: MicroRNAs
miR-125b and miR-100 suppress metastasis of hepatocellular
carcinoma by disrupting the formation of vessels that encapsulate
tumour clusters. J Pathol. 240:450–460. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ge YY, Shi Q, Zheng ZY, Gong J, Zeng C,
Yang J and Zhuang SM: MicroRNA-100 promotes the autophagy of
hepatocellular carcinoma cells by inhibiting the expression of mTOR
and IGF-1R. Oncotarget. 5:6218–6228. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Pan WY, Zeng JH, Wen DY, Wang JY, Wang PP,
Chen G and Feng ZB: Oncogenic value of microRNA-15b-5p in
hepatocellular carcinoma and a bioinformatics investigation. Oncol
Lett. 17:1695–1713. 2019.PubMed/NCBI
|
|
17
|
Zhang S, Gao Y and Huang J: Interleukin-8
Gene-251 A/T (rs4073) polymorphism and coronary artery disease
risk: A meta-analysis. Med Sci Monit. 25:1645–1655. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhang X, Xin G and Sun D: Serum exosomal
miR-328, miR-575, miR-134 and miR-671-5p as potential biomarkers
for the diagnosis of Kawasaki disease and the prediction of
therapeutic outcomes of intravenous immunoglobulin therapy. Exp
Ther Med. 16:2420–2432. 2018.PubMed/NCBI
|
|
19
|
Betel D, Koppal A, Agius P, Sander C and
Leslie C: Comprehensive modeling of microRNA targets predicts
functional non-conserved and non-canonical sites. Genome Biol.
11:R902010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Paraskevopoulou MD, Georgakilas G,
Kostoulas N, Vlachos IS, Vergoulis T, Reczko M, Filippidis C,
Dalamagas T and Hatzigeorgiou AG: DIANA-microT web server v5.0:
Service integration into miRNA functional analysis workflows.
Nucleic Acids Res. 41:W169–W173. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hsu SD, Chu CH, Tsou AP, Chen SJ, Chen HC,
Hsu PW, Wong YH, Chen YH, Chen GH and Huang HD: miRNAMap 2.0:
Genomic maps of microRNAs in metazoan genomes. Nucleic Acids Res 36
(Database Issue). D165–D169. 2008.
|
|
22
|
Tsang JS, Ebert MS and van Oudenaarden A:
Genome-wide dissection of microRNA functions and cotargeting
networks using gene set signatures. Mol Cell. 38:140–153. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Vejnar CE, Blum M and Zdobnov EM: miRmap
web: Comprehensive microRNA target prediction online. Nucleic Acids
Res. 41:W165–W168. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Anders G, Mackowiak SD, Jens M, Maaskola
J, Kuntzagk A, Rajewsky N, Landthaler M and Dieterich C: doRiNA: A
database of RNA interactions in post-transcriptional regulation.
Nucleic Acids Res 40 (Database Issue). D180–D186. 2012. View Article : Google Scholar
|
|
25
|
Kertesz M, Iovino N, Unnerstall U, Gaul U
and Segal E: The role of site accessibility in microRNA target
recognition. Nat Genet. 39:1278–1284. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang X and El Naqa IM: Prediction of both
conserved and nonconserved microRNA targets in animals.
Bioinformatics. 24:325–332. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Rehmsmeier M, Steffen P, Hochsmann M and
Giegerich R: Fast and effective prediction of microRNA/target
duplexes. RNA. 10:1507–1517. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Loher P and Rigoutsos I: Interactive
exploration of RNA22 microRNA target predictions. Bioinformatics.
28:3322–3323. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Friedman RC, Farh KK, Burge CB and Bartel
DP: Most mammalian mRNAs are conserved targets of microRNAs. Genome
Res. 19:92–105. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Calderón-González KG, Hernández-Monge J,
Herrera-Aguirre ME and Luna-Arias JP: Bioinformatics tools for
proteomics data interpretation. Adv Exp Med Biol. 919:281–341.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Szklarczyk D, Morris JH, Cook H, Kuhn M,
Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, et al:
The STRING database in 2017: Quality-controlled protein-protein
association networks, made broadly accessible. Nucleic Acids Res.
45:D362–D368. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Roessler S, Jia HL, Budhu A, Forgues M, Ye
QH, Lee JS, Thorgeirsson SS, Sun Z, Tang ZY, Qin LX and Wang XW: A
unique metastasis gene signature enables prediction of tumor
relapse in early-stage hepatocellular carcinoma patients. Cancer
Res. 70:10202–10212. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Chen X, Cheung ST, So S, Fan ST, Barry C,
Higgins J, Lai KM, Ji J, Dudoit S, Ng IO, et al: Gene expression
patterns in human liver cancers. Mol Biol Cell. 13:1929–1939. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Uhlen M, Zhang C, Lee S, Sjöstedt E,
Fagerberg L, Bidkhori G, Benfeitas R, Arif M, Liu Z, Edfors F, et
al: A pathology atlas of the human cancer transcriptome. Science.
357(pii): eaan25072017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Jiao XD, Qin BD, You P, Cai J and Zang YS:
The prognostic value of TP53 and its correlation with EGFR mutation
in advanced non-small cell lung cancer, an analysis based on
cBioPortal data base. Lung Cancer. 123:70–75. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Ryan M, Wong WC, Brown R, Akbani R, Su X,
Broom B, Melott J and Weinstein J: TCGASpliceSeq a compendium of
alternative mRNA splicing in cancer. Nucleic Acids Res.
44:D1018–D1022. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhang R, Lin P, Yang X, He RQ, Wu HY, Dang
YW, Gu YY, Peng ZG, Feng ZB and Chen G: Survival associated
alternative splicing events in diffuse large B-cell lymphoma. Am J
Transl Res. 10:2636–2647. 2018.PubMed/NCBI
|
|
38
|
Li Y, Sun N, Lu Z, Sun S, Huang J, Chen Z
and He J: Prognostic alternative mRNA splicing signature in
non-small cell lung cancer. Cancer Lett. 393:40–51. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Xiao F, Bai Y, Chen Z, Li Y, Luo L, Huang
J, Yang J, Liao H and Guo L: Downregulation of HOXA1 gene affects
small cell lung cancer cell survival and chemoresistance under the
regulation of miR-100. Eur J Cancer. 50:1541–1554. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Guo P, Xiong X, Zhang S and Peng D:
miR-100 resensitizes resistant epithelial ovarian cancer to
cisplatin. Oncol Rep. 36:3552–3558. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Qin X, Yu S, Zhou L, Shi M, Hu Y, Xu X,
Shen B, Liu S, Yan D and Feng J: Cisplatin-resistant lung cancer
cell-derived exosomes increase cisplatin resistance of recipient
cells in exosomal miR-100-5p-dependent manner. Int J Nanomedicine.
12:3721–3733. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Wang M, Ren D, Guo W, Wang Z, Huang S, Du
H, Song L and Peng X: Loss of miR-100 enhances migration, invasion,
epithelial-mesenchymal transition and stemness properties in
prostate cancer cells through targeting Argonaute 2. Int J Oncol.
45:362–372. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Leite KR, Sousa-Canavez JM, Reis ST,
Tomiyama AH, Camara-Lopes LH, Sañudo A, Antunes AA and Srougi M:
Change in expression of miR-let7c, miR-100, and miR-218 from high
grade localized prostate cancer to metastasis. Urol Oncol.
29:265–269. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Zhang H, Wang J, Wang Z, Ruan C, Wang L
and Guo H: Serum miR-100 is a potential biomarker for detection and
outcome prediction of glioblastoma patients. Cancer Biomark.
24:43–49. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Azizmohammadi S, Azizmohammadi S, Safari
A, Kosari N, Kaghazian M, Yahaghi E and Seifoleslami M: The role
and expression of miR-100 and miR-203 profile as prognostic markers
in epithelial ovarian cancer. Am J Transl Res. 8:2403–2410.
2016.PubMed/NCBI
|
|
46
|
Petrelli A, Perra A, Schernhuber K,
Cargnelutti M, Salvi A, Migliore C, Ghiso E, Benetti A, Barlati S,
Ledda-Columbano GM, et al: Sequential analysis of multistage
hepatocarcinogenesis reveals that miR-100 and PLK1 dysregulation is
an early event maintained along tumor progression. Oncogene.
31:4517–4526. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Varnholt H, Drebber U, Schulze F,
Wedemeyer I, Schirmacher P, Dienes HP and Odenthal M: MicroRNA gene
expression profile of hepatitis C virus-associated hepatocellular
carcinoma. Hepatology. 47:1223–1232. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Wang Y, Gao Y, Shi W, Zhai D, Rao Q, Jia
X, Liu J, Jiao X and Du Z: Profiles of differential expression of
circulating microRNAs in hepatitis B virus-positive small
hepatocellular carcinoma. Cancer Biomark. 15:171–180. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Tsai CT, Zulueta MML and Hung SC:
Synthetic heparin and heparan sulfate: Probes in defining
biological functions. Curr Opin Chem Biol. 40:152–159. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Li JP and Kusche-Gullberg M: Heparan
Sulfate: Biosynthesis, structure, and function. Int Rev Cell Mol
Biol. 325:215–273. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Weiss RJ, Esko JD and Tor Y: Targeting
heparin and heparan sulfate protein interactions. Org Biomol Chem.
15:5656–5668. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Cassinelli G, Zaffaroni N and Lanzi C: The
heparanase/heparan sulfate proteoglycan axis: A potential new
therapeutic target in sarcomas. Cancer Lett. 382:245–254. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Ling L, Tan SK, Goh TH, Cheung E, Nurcombe
V, van Wijnen AJ and Cool SM: Targeting the heparin-binding domain
of fibroblast growth factor receptor 1 as a potential cancer
therapy. Mol Cancer. 14:1362015. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Dudás J, Bocsi J, Fullár A, Baghy K, Füle
T, Kudaibergenova S and Kovalszky I: Heparin and liver heparan
sulfate can rescue hepatoma cells from topotecan action. Biomed Res
Int. 2014:7657942014. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Gao J, Wang Y, Li W, Zhang J, Che Y, Cui
X, Sun B and Zhao G: Loss of histone deacetylase 2 inhibits
oxidative stress induced by high glucose via the HO-1/SIRT1 pathway
in endothelial progenitor cells. Gene. 678:1–7. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Li Y, Zhou M, Lv X, Song L, Zhang D, He Y,
Wang M, Zhao X, Yuan X, Shi G and Wang D: Reduced activity of HDAC3
and increased acetylation of histones H3 in peripheral blood
mononuclear cells of patients with rheumatoid arthritis. J Immunol
Res. 2018:73135152018. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Li S, Wang F, Qu Y, Chen X, Gao M, Yang J,
Zhang D, Zhang N, Li W and Liu H: HDAC2 regulates cell
proliferation, cell cycle progression and cell apoptosis in
esophageal squamous cell carcinoma EC9706 cells. Oncol Lett.
13:403–409. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Peng Z, Zhou W, Zhang C, Liu H and Zhang
Y: Curcumol controls choriocarcinoma stem-like cells self-renewal
via repression of DNA Methyltransferase (DNMT)- and histone
deacetylase (HDAC)-mediated epigenetic regulation. Med Sci Monit.
24:461–472. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Liu L, Lin W, Zhang Q, Cao W and Liu Z:
TGF-β induces miR-30d down-regulation and podocyte injury through
Smad2/3 and HDAC3-associated transcriptional repression. J Mol Med
(Berl). 94:291–300. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Mao QD, Zhang W, Zhao K, Cao B, Yuan H,
Wei LZ, Song MQ and Liu XS: MicroRNA-455 suppresses the oncogenic
function of HDAC2 in human colorectal cancer. Braz J Med Biol Res.
50:e61032017. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Yang Y, Zhang J, Wu T, Xu X, Cao G, Li H
and Chen X: Histone deacetylase 2 regulates the doxorubicin (Dox)
resistance of hepatocarcinoma cells and transcription of ABCB1.
Life Sci. 216:200–206. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Zhao H, Yu Z, Zhao L, He M, Ren J, Wu H,
Chen Q, Yao W and Wei M: HDAC2 overexpression is a poor prognostic
factor of breast cancer patients with increased multidrug
resistance-associated protein expression who received
anthracyclines therapy. Jpn J Clin Oncol. 46:893–902. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Cui Z, Xie M, Wu Z and Shi Y: Relationship
between histone deacetylase 3 (HDAC3) and breast cancer. Med Sci
Monit. 24:2456–2464. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Wu LM, Yang Z, Zhou L, Zhang F, Xie HY,
Feng XW, Wu J and Zheng SS: Identification of histone deacetylase 3
as a biomarker for tumor recurrence following liver transplantation
in HBV-associated hepatocellular carcinoma. PLoS One. 5:e144602010.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Lebiedzinska-Arciszewska M, Oparka M,
Vega-Naredo I, Karkucinska-Wieckowska A, Pinton P, Duszynski J and
Wieckowski MR: The interplay between p66Shc, reactive oxygen
species and cancer cell metabolism. Eur J Clin Invest. 45 (Suppl
1):S25–S31. 2015. View Article : Google Scholar
|
|
66
|
Wong N, Chan A, Lee SW, Lam E, To KF, Lai
PB, Li XN, Liew CT and Johnson PJ: Positional mapping for amplified
DNA sequences on 1q21-q22 in hepatocellular carcinoma indicates
candidate genes over-expression. J Hepatol. 38:298–306. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Kisielow M, Kleiner S, Nagasawa M, Faisal
A and Nagamine Y: Isoform-specific knockdown and expression of
adaptor protein ShcA using small interfering RNA. Biochem J.
363:1–5. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Zhang L, Zhu S, Shi X and Sha W: The
silence of p66(Shc) in HCT8 cells inhibits the viability via
PI3K/AKT/Mdm-2/p53 signaling pathway. Int J Clin Exp Pathol.
8:9097–9104. 2015.PubMed/NCBI
|
|
69
|
Yukimasa S, Masaki T, Yoshida S, Uchida N,
Watanabe S, Usuki H, Yoshiji H, Maeta T, Ebara K, Nakatsu T, et al:
Enhanced expression of p46 Shc in the nucleus and p52 Shc in the
cytoplasm of human gastric cancer. Int J Oncol. 26:905–911.
2005.PubMed/NCBI
|
|
70
|
Muniyan S, Chou YW, Tsai TJ, Thomes P,
Veeramani S, Benigno BB, Walker LD, McDonald JF, Khan SA, Lin FF,
et al: p66Shc longevity protein regulates the proliferation of
human ovarian cancer cells. Mol Carcinog. 54:618–631. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Rajendran M, Thomes P, Zhang L, Veeramani
S and Lin MF: p66Shc-a longevity redox protein in human prostate
cancer progression and metastasis: p66Shc in cancer progression and
metastasis. Cancer Metastasis Rev. 29:207–222. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Yoshida S, Kornek M, Ikenaga N, Schmelzle
M, Masuzaki R, Csizmadia E, Wu Y, Robson SC and Schuppan D:
Sublethal heat treatment promotes epithelial-mesenchymal transition
and enhances the malignant potential of hepatocellular carcinoma.
Hepatology. 58:1667–1680. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Bid HK, Roberts RD, Manchanda PK and
Houghton PJ: RAC1: An emerging therapeutic option for targeting
cancer angiogenesis and metastasis. Mol Cancer Ther. 12:1925–1934.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Cheng H, Wang W, Wang G, Wang A, Du L and
Lou W: Silencing ras-related C3 botulinum toxin substrate 1
inhibits growth and migration of hypopharyngeal squamous cell
carcinoma via the P38 mitogen-activated protein kinase signaling
pathway. Med Sci Monit. 24:768–781. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Liu B, Xiong J, Liu G, Wu J, Wen L, Zhang
Q and Zhang C: High expression of Rac1 is correlated with partial
reversed cell polarity and poor prognosis in invasive ductal
carcinoma of the breast. Tumour Biol. 39:10104283177109082017.
View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Yoon C, Cho SJ, Chang KK, Park DJ, Ryeom
SW and Yoon SS: Role of Rac1 pathway in epithelial-to-mesenchymal
transition and cancer stem-like cell phenotypes in gastric
adenocarcinoma. Mol Cancer Res. 15:1106–1116. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Niu X, Gao Z, Qi S, Su L, Yang N, Luan X,
Li J, Zhang Q, An Y and Zhang S: Macropinocytosis activated by
oncogenic Dbl enables specific targeted delivery of Tat/pDNA
nano-complexes into ovarian cancer cells. Int J Nanomedicine.
13:4895–4911. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Poudel KR, Roh-Johnson M, Su A, Ho T,
Mathsyaraja H, Anderson S, Grady WM, Moens CB, Conacci-Sorrell M,
Eisenman RN and Bai J: Competition between TIAM1 and membranes
balances endophilin A3 activity in cancer metastasis. Dev Cell.
45:738–752.e6. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Aspenström P: Activated Rho GTPases in
cancer-the beginning of a new paradigm. Int J Mol Sci. 19(pii):
E39492018. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Ching YP, Leong VY, Lee MF, Xu HT, Jin DY
and Ng IO: P21-activated protein kinase is overexpressed in
hepatocellular carcinoma and enhances cancer metastasis involving
c-Jun NH2-terminal kinase activation and paxillin phosphorylation.
Cancer Res. 67:3601–3608. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Zhu S, Jin J, Gokhale S, Lu AM, Shan H,
Feng J and Xie P: Genetic alterations of TRAF proteins in human
cancers. Front Immunol. 9:21112018. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Reder H, Wagner S, Gamerdinger U, Sandmann
S, Wuerdemann N, Braeuninger A, Dugas M, Gattenloehner S, Klussmann
JP and Wittekindt C: Genetic alterations in human
papillomavirus-associated oropharyngeal squamous cell carcinoma of
patients with treatment failure. Oral Oncol. 93:59–65. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Sajnani K, Islam F, Smith RA, Gopalan V
and Lam AK: Genetic alterations in Krebs cycle and its impact on
cancer pathogenesis. Biochimie. 135:164–172. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Jonckheere N, Vasseur R and Van Seuningen
I: The cornerstone K-RAS mutation in pancreatic adenocarcinoma:
From cell signaling network, target genes, biological processes to
therapeutic targeting. Crit Rev Oncol Hematol. 111:7–19. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Song X, Zeng Z, Wei H and Wang Z:
Alternative splicing in cancers: From aberrant regulation to new
therapeutics. Semin Cell Dev Biol. 75:13–22. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Climente-González H, Porta-Pardo E, Godzik
A and Eyras E: The functional impact of alternative splicing in
cancer. Cell Rep. 20:2215–2226. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Xie M, Dart DA, Owen S, Wen X, Ji J and
Jiang W: Insights into roles of the miR-1, −133 and −206 family in
gastric cancer (Review). Oncol Rep. 36:1191–1198. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Teplyuk NM, Uhlmann EJ, Gabriely G,
Volfovsky N, Wang Y, Teng J, Karmali P, Marcusson E, Peter M, Mohan
A, et al: Therapeutic potential of targeting microRNA-10b in
established intracranial glioblastoma: First steps toward the
clinic. EMBO Mol Med. 8:268–287. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Li W, Xie L, He X, Li J, Tu K, Wei L, Wu
J, Guo Y, Ma X, Zhang P, et al: Diagnostic and prognostic
implications of microRNAs in human hepatocellular carcinoma. Int J
Cancer. 123:1616–1622. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Su H, Yang JR, Xu T, Huang J, Xu L, Yuan Y
and Zhuang SM: MicroRNA-101, down-regulated in hepatocellular
carcinoma, promotes apoptosis and suppresses tumorigenicity. Cancer
Res. 69:1135–1142. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Burchard J, Zhang C, Liu AM, Poon RT, Lee
NP, Wong KF, Sham PC, Lam BY, Ferguson MD, Tokiwa G, et al:
microRNA-122 as a regulator of mitochondrial metabolic gene network
in hepatocellular carcinoma. Mol Sys Biol. 6:4022010. View Article : Google Scholar
|
|
92
|
Sato F, Hatano E, Kitamura K, Myomoto A,
Fujiwara T, Takizawa S, Tsuchiya S, Tsujimoto G, Uemoto S and
Shimizu K: MicroRNA profile predicts recurrence after resection in
patients with hepatocellular carcinoma within the Milan Criteria.
PLoS One. 6:e164352011. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Noh JH, Chang YG, Kim MG, Jung KH, Kim JK,
Bae HJ, Eun JW, Shen Q, Kim SJ, Kwon SH, et al: MiR-145 functions
as a tumor suppressor by directly targeting histone deacetylase 2
in liver cancer. Cancer Lett. 335:455–462. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Wang PR, Xu M, Toffanin S, Li Y, Llovet JM
and Russell DW: Induction of hepatocellular carcinoma by in vivo
gene targeting. Proc Natl Acad Sci USA. 109:11264–11269. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Morita K, Shirabe K, Taketomi A, Soejima
Y, Yoshizumi T, Uchiyama H, Ikegami T, Yamashita Y, Sugimachi K,
Harimoto N, et al: Relevance of microRNA-18a and microRNA-199a-5p
to hepatocellular carcinoma recurrence after living donor liver
transplantation. Liver Transpl. 22:665–676. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Shih TC, Tien YJ, Wen CJ, Yeh TS, Yu MC,
Huang CH, Lee YS, Yen TC and Hsieh SY: MicroRNA-214 downregulation
contributes to tumor angiogenesis by inducing secretion of the
hepatoma-derived growth factor in human hepatoma. J Hepatol.
57:584–591. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Wojcicka A, Swierniak M, Kornasiewicz O,
Gierlikowski W, Maciag M, Kolanowska M, Kotlarek M, Gornicka B,
Koperski L, Niewinski G, et al: Next generation sequencing reveals
microRNA isoforms in liver cirrhosis and hepatocellular carcinoma.
Int J Biochem Cell Biol. 53:208–217. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Shen J, Siegel AB, Remotti H, Wang Q and
Santella RM: Identifying microRNA panels specifically associated
with hepatocellular carcinoma and its different etiologies.
Hepatoma Res. 2:151–162. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Lou W, Liu J, Ding B, Chen D, Xu L, Ding
J, Jiang D, Zhou L, Zheng S and Fan W: Identification of potential
miRNA-mRNA regulatory network contributing to pathogenesis of
HBV-related HCC. J Transl Med. 17:72019. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Murakami Y, Kubo S, Tamori A, Itami S,
Kawamura E, Iwaisako K, Ikeda K, Kawada N, Ochiya T and Taguchi YH:
Comprehensive analysis of transcriptome and metabolome analysis in
Intrahepatic Cholangiocarcinoma and hepatocellular carcinoma. Sci
Rep. 5:162942015. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Ghosh A, Ghosh A, Datta S, Dasgupta D, Das
S, Ray S, Gupta S, Datta S, Chowdhury A, Chatterjee R, et al:
Hepatic miR-126 is a potential plasma biomarker for detection of
hepatitis B virus infected hepatocellular carcinoma. Int J Cancer.
138:2732–2744. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Peng H, Ishida M, Li L, Saito A, Kamiya A,
Hamilton JP, Fu R, Olaru AV, An F, Popescu I, et al: Pseudogene
INTS6P1 regulates its cognate gene INTS6 through competitive
binding of miR-17-5p in hepatocellular carcinoma. Oncotarget.
6:5666–5677. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Zhang Y, Wen DY, Zhang R, Huang JC, Lin P,
Ren FH, Wang X, He Y, Yang H, Chen G and Luo DZ: A preliminary
investigation of PVT1 on the effect and mechanisms of
hepatocellular carcinoma: Evidence from clinical data, a
meta-analysis of 840 cases and in vivo validation. Cell Physiol
Biochem. 47:2216–2232. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Shi J, Ye G, Zhao G, Wang X, Ye C,
Thammavong K, Xu J and Dong J: Coordinative control of G2/M phase
of the cell cycle by non-coding RNAs in hepatocellular carcinoma.
PeerJ. 6:e57872018. View Article : Google Scholar : PubMed/NCBI
|