|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cohen PA, Jhingran A, Oaknin A and Denny
L: Cervical cancer. Lancet. 393:169–182. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Cai C, Rajaram M, Zhou X, Liu Q, Marchica
J, Li J and Powers RS: Activation of multiple cancer pathways and
tumor maintenance function of the 3q amplified oncogene FNDC3B.
Cell Cycle. 11:1773–1781. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Smith RA, Andrews KS, Brooks D, Fedewa SA,
Manassaram-Baptiste D, Saslow D, Brawley OW and Wender RC: Cancer
screening in the United States, 2018: A review of current American
Cancer Society guidelines and current issues in cancer screening.
CA Cancer J Clin. 68:297–316. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Choi YJ and Park JS: Clinical significance
of human papillomavirus genotyping. J Gynecol Oncol. 27:e212016.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Tominaga K, Kondo C, Johmura Y, Nishizuka
M and Imagawa M: The novel gene fad104, containing a fibronectin
type III domain, has a significant role in adipogenesis. FEBS Lett.
577:49–54. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Nishizuka M, Kishimoto K, Kato A, Ikawa M,
Okabe M, Sato R, Niida H, Nakanishi M, Osada S and Imagawa M:
Disruption of the novel gene fad104 causes rapid postnatal death
and attenuation of cell proliferation, adhesion, spreading and
migration. Exp Cell Res. 315:809–819. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lin CH, Lin YW, Chen YC, Liao CC, Jou YS,
Hsu MT and Chen CF: FNDC3B promotes cell migration and tumor
metastasis in hepatocellular carcinoma. Oncotarget. 7:49498–49508.
2016.PubMed/NCBI
|
|
10
|
Noell G, Faner R and Agusti A: From
systems biology to P4 medicine: Applications in respiratory
medicine. Eur Respir Rev. 27(pii): 1701102018. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lin M, Ye M, Zhou J, Wang ZP and Zhu X:
Recent advances on the molecular mechanism of cervical
carcinogenesis based on systems biology technologies. Comput Struct
Biotechnol J. 17:241–250. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
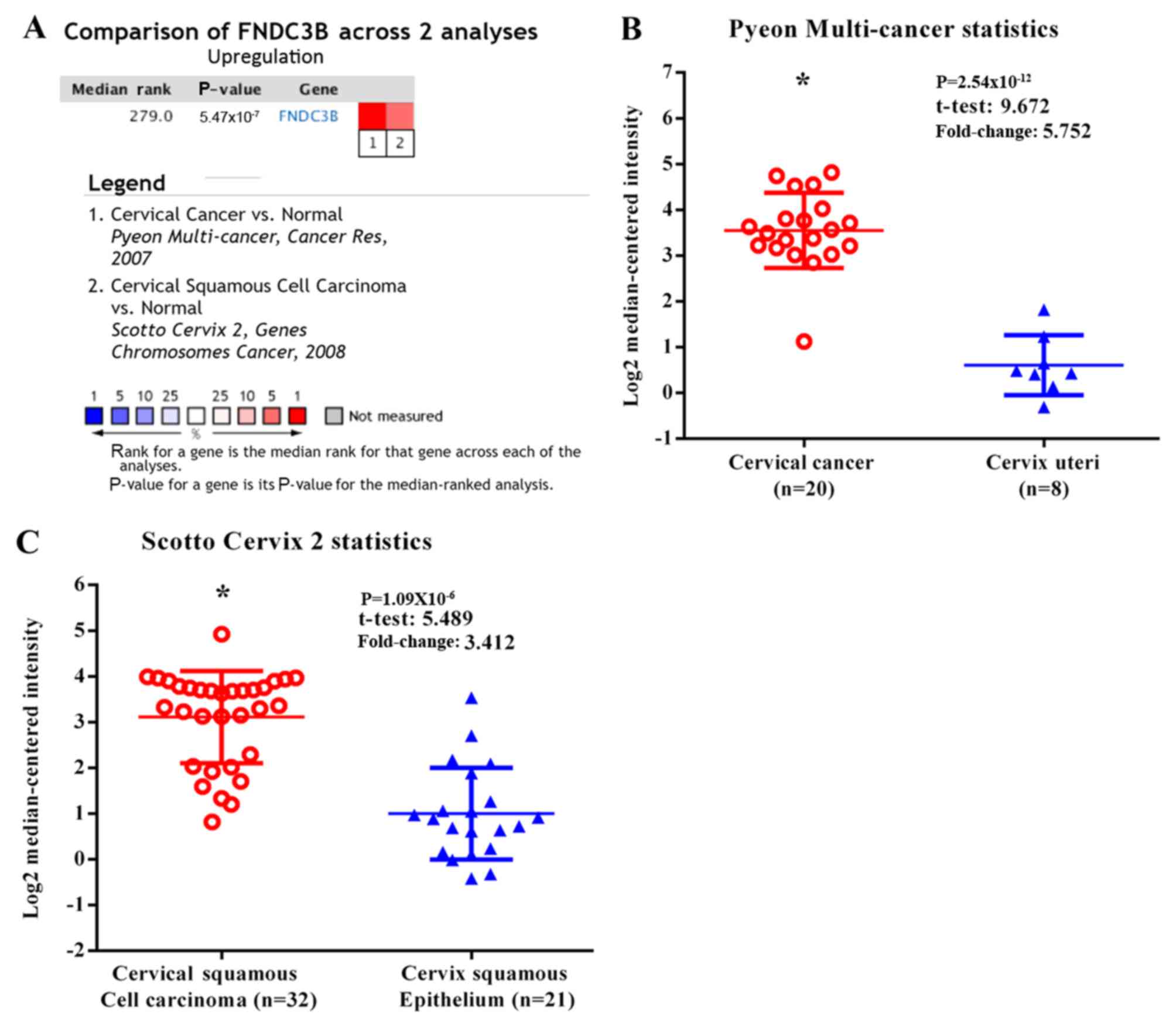
Rhodes DR, Yu J, Shanker K, Deshpande N,
Varambally R, Ghosh D, Barrette T, Pandey A and Chinnaiyan AM:
ONCOMINE: A cancer microarray database and integrated data-mining
platform. Neoplasia. 6:1–6. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Liu Y, Cui S, Li W, Zhao Y, Yan X and Xu
J: PAX3 is a biomarker and prognostic factor in melanoma: Database
mining. Oncol Lett. 17:4985–4993. 2019.PubMed/NCBI
|
|
14
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
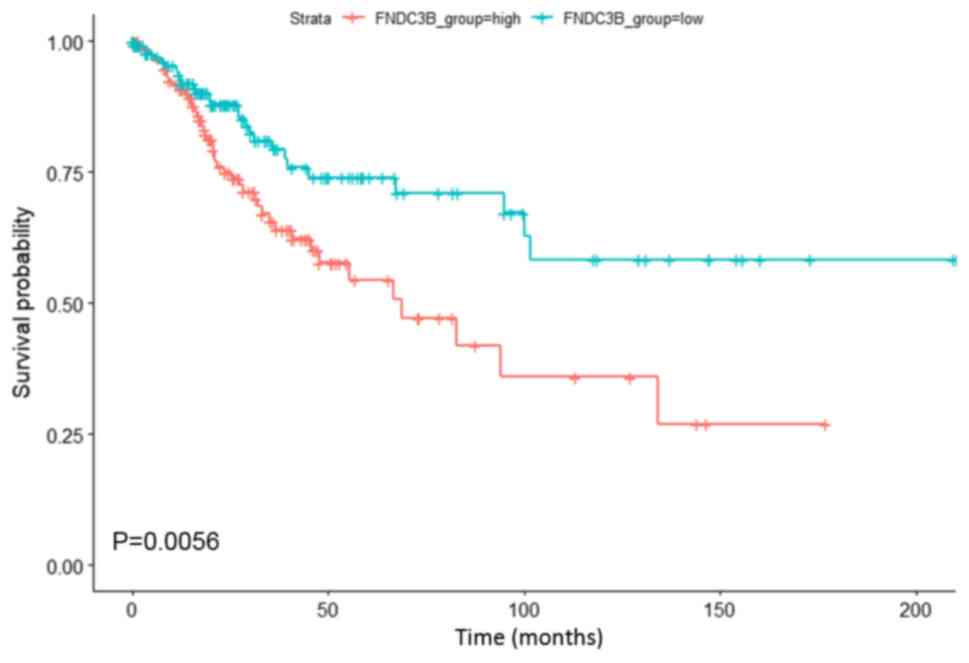
Therneau T: A Package for survival
analysis in S. R package version 2.38. https://CRAN.R-project.org/package=survival2015
|
|
17
|
Terry MT and Grambsch PM: Modeling
survival data: Extending the cox model. Springer; New York:
2000
|
|
18
|
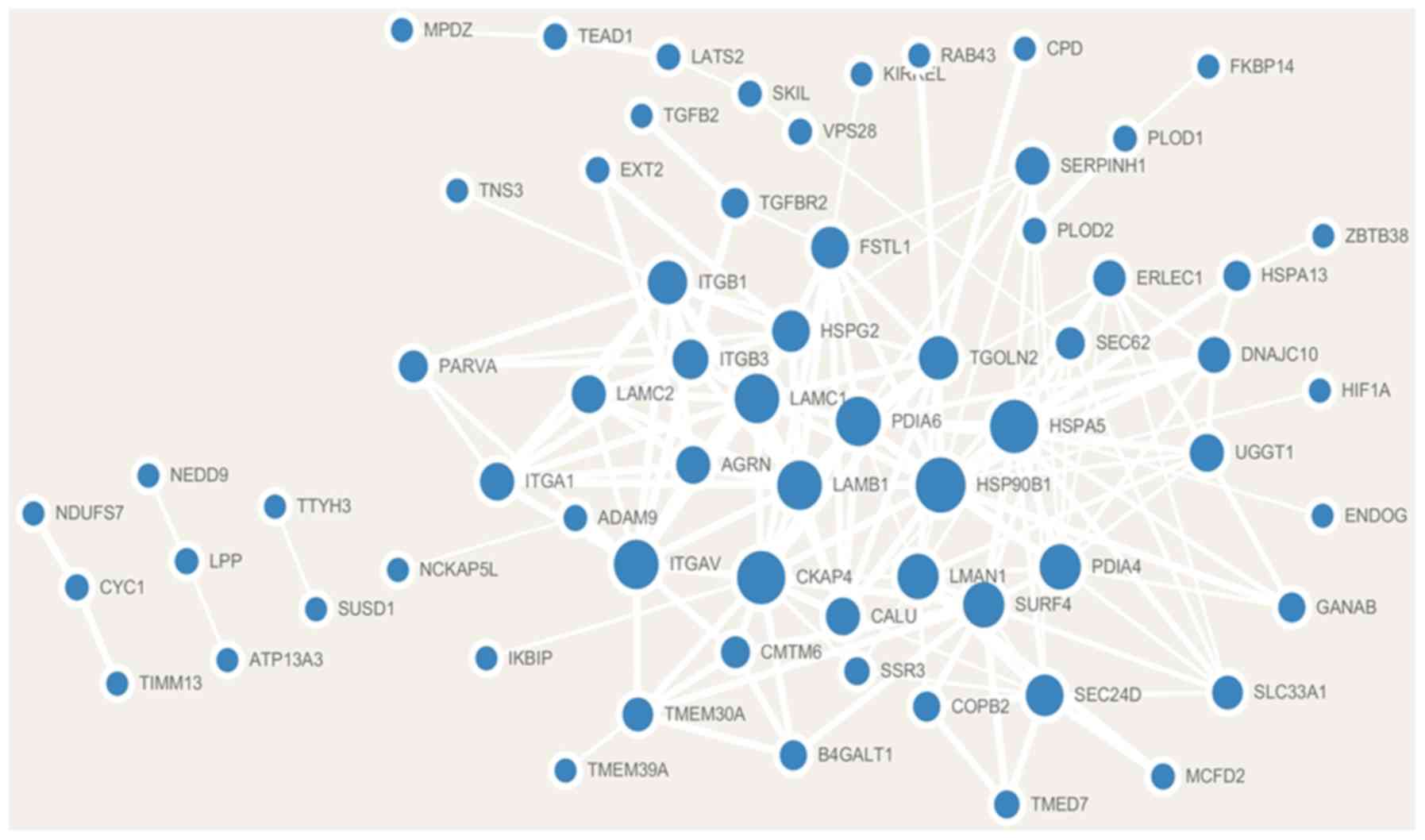
Szklarczyk D, Gable AL, Lyon D, Junge A,
Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork
P, et al: STRING v11: Protein-protein association networks with
increased coverage, supporting functional discovery in genome-wide
experimental datasets. Nucleic Acids Res. 47:D607–D613. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
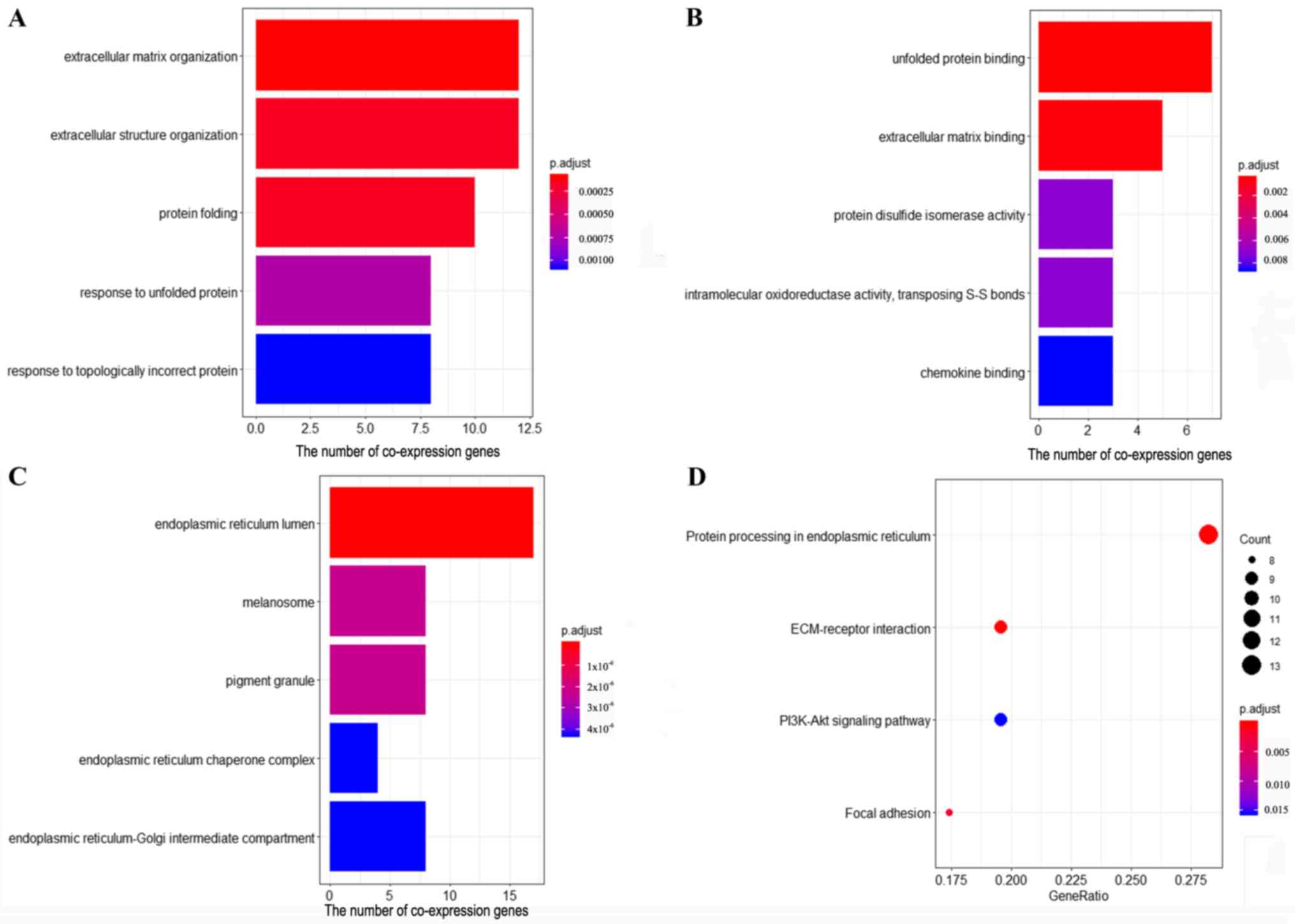
Yu G, Wang LG, Han Y and He QY:
ClusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Uhlen M, Zhang C, Lee S, Sjöstedt E,
Fagerberg L, Bidkhori G, Benfeitas R, Arif M, Liu Z, Edfors F, et
al: A pathology atlas of the human cancer transcriptome. Science.
357(pii): eaan25072017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Pyeon D, Newton MA, Lambert PF, den Boon
JA, Sengupta S, Marsit CJ, Woodworth CD, Connor JP, Haugen TH,
Smith EM, et al: Fundamental differences in cell cycle deregulation
in human papillomavirus-positive and human papillomavirus-negative
head/neck and cervical cancers. Cancer Res. 67:4605–4619. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Scotto L, Narayan G, Nandula SV,
Arias-Pulido H, Subramaniyam S, Schneider A, Kaufmann AM, Wright
JD, Pothuri B, Mansukhani M and Murty VV: Identification of copy
number gain and overexpressed genes on chromosome arm 20q by an
integrative genomic approach in cervical cancer: Potential role in
progression. Genes Chromosomes Cancer. 47:755–765. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Vitale A and Denecke J: The endoplasmic
reticulum-gateway of the secretory pathway. Plant Cell. 11:615–628.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Jain BP: An overview of unfolded protein
response signaling and its role in cancer. Cancer Biother
Radiopharm. 32:275–281. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang M and Kaufman RJ: The impact of the
endoplasmic reticulum protein-folding environment on cancer
development. Nat Rev Cancer. 14:581–597. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wang M, Law ME, Castellano RK and Law BK:
The unfolded protein response as a target for anticancer
therapeutics. Crit Rev Oncol Hematol. 127:66–79. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Taguchi Y, Horiuchi Y, Kano F and Murata
M: Novel prosurvival function of Yip1A in human cervical cancer
cells: Constitutive activation of the IRE1 and PERK pathways of the
unfolded protein response. Cell Death Dis. 8:e27182017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Cubillos-Ruiz JR, Bettigole SE and
Glimcher LH: Tumorigenic and immunosuppressive effects of
endoplasmic reticulum stress in cancer. Cell. 168:692–706. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Walter P and Ron D: The unfolded protein
response: From stress pathway to homeostatic regulation. Science.
334:1081–1086. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang M and Kaufman RJ: Protein misfolding
in the endoplasmic reticulum as a conduit to human disease. Nature.
529:326–335. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tashiro E: Screening and identification of
inhibitors of endoplasmic reticulum stress-induced activation of
the IRE1a-XBP1 branch. J Antibiot (Tokyo). Aug 9–2019.(Epub ahead
of print). View Article : Google Scholar
|
|
33
|
Maurel M, McGrath EP, Mnich K, Healy S,
Chevet E and Samali A: Controlling the unfolded protein
response-mediated life and death decisions in cancer. Semin Cancer
Biol. 33:57–66. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Jain S, Wheeler JR, Walters RW, Agrawal A,
Barsic A and Parker R: ATPase-modulated stress granules contain a
diverse proteome and substructure. Cell. 164:487–498. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Namkoong S, Ho A, Woo YM, Kwak H and Lee
JH: Systematic characterization of stress-induced RNA granulation.
Mol Cell. 70:175–187 e8. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kedersha N, Ivanov P and Anderson P:
Stress granules and cell signaling: More than just a passing phase?
Trends Biochem Sci. 38:494–506. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Castello A, Fischer B, Eichelbaum K, Horos
R, Beckmann BM, Strein C, Davey NE, Humphreys DT, Preiss T,
Steinmetz LM, et al: Insights into RNA biology from an atlas of
mammalian mRNA-binding proteins. Cell. 149:1393–1406. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
McAuliffe PF, Meric-Bernstam F, Mills GB
and Gonzalez-Angulo AM: Deciphering the role of PI3K/Akt/mTOR
pathway in breast cancer biology and pathogenesis. Clin Breast
Cancer. 10 (Suppl 3):S59–S65. 2010. View Article : Google Scholar : PubMed/NCBI
|