Introduction
Glioblastoma multiforme (GBM) is the most common
histological subtype of high-grade glioma and the most prevalent
primary brain tumour in adults. The prevalence of GBM is
approximately 3–4 cases per 100,000 (1). GBM was previously considered an
incurable tumour, with a median survival of ~15 months (2). Isocitrate dehydrogenase (IDH)1/2
mutations have been described in ~12% of patients with GBM and are
associated with an improved long-term survival (3). However, the risk factors influencing the
prognosis of patients with GBM are still unclear.
GBM is characterized by tissue hypoxia, which is
known to mediate expression of the oxygen-regulated transcription
factor, hypoxia inducible factor (HIF)-1. It is established that
the hypoxic microenvironment of cancer tissue is closely associated
with tumour growth and development, and a poor prognosis (4,5). Numerous
studies have reported that hypoxia-associated markers, including
vascular endothelial growth factor and osteopontin, are correlated
with a poor prognosis for patients with GBM (6,7). Kaelin
also revealed that the extent of HIF-1α expression is associated
with cancer aggressiveness, resistance to radiation and
chemotherapy, and poor prognosis (8).
In addition, HIF-1α has been proposed as a prognostic marker to
monitor the development of GBM. It is therefore crucial to
understand the expression of HIF-1α in patients with GBM and to
determine its association with patient prognosis.
Caveolin-1 (CAV1) is a plasma membrane organizing
protein, the expression of which is increased in various types of
cancer (9). Previous studies have
demonstrated that CAV1 is upregulated by HIF-1α (10,11), and
that HIF-dependent upregulation of CAV1 enhances the oncogenic
potential of tumour cells by increasing proliferative, migratory
and invasive cell capacities (11).
This suggests that there may be an association between CAV1 and HIF
in vitro. Kannan et al reported that CAV1 promotes
gastric cancer progression in vivo by upregulating
epithelial to mesenchymal transition under hypoxic conditions
(12). It has also been proposed that
hypoxia-induced CAV1 drives tumourigenesis and metastasis in
hepatocellular carcinoma (9);
however, the expression of CAV1 and HIF-1α, and their implication
in IDH-wild type GBM are still unknown.
The present study aimed to examine the expression
levels of HIF-1α and CAV1 in IDH-wild type tissues from patients
with GBM compared to adjacent healthy tissues. In addition, the
association between HIF-1α and CAV1 expression levels and the
clinicopathological characteristics of patients with IDH-wild type
GBM, including sex, age, weight, methylation of the
O6-methylguanine-DNA methyltransferase (MGMT) promoter and
prognosis, were assessed.
Materials and methods
Patients and samples
A total of 42 patients diagnosed with IDH-wild type
GBM were recruited to the study between June 2012 and June 2014 at
the Department of Neurological Surgery, The First Affiliated
Hospital of Sun Yat-sen University (Guangzhou, China). Patients
were aged 26–76 years, and comprised 17 men and 25 women. Samples
were obtained by surgical procedure from patients who had not
received chemotherapy or radiotherapy prior to surgical tumour
resection. All tumour samples and adjacent samples were obtained
from the total resection by microscopy, and tissues were obtained
from surgical specimens and immediately snap-frozen in liquid
nitrogen until RNA and protein extraction. 42 tumour samples were
prepared as archival paraffin blocks. The histopathological
diagnosis of the 42 patients was confirmed by pathologists
according to the 2016 World Health Organization grading system
(malignancy scale) for central nervous system tumours (13).
A total of 17 matched pairs of fresh IDH-wild type
GBM and adjacent healthy tissue samples (non-tumour, N-T) from the
42 samples were also obtained to determine mRNA and protein
expression levels of HIF-1α and CAV1. Haematoxylin-eosin staining
analysis of frozen sections were used to confirm that the tumour
tissues were composed of >70% cancer cells without necrosis, and
no cancerous lesions were present in the healthy tissues The
staining protocol was as follows: The sections were stained with
Harris's hematoxylin for 7 min, and washed in water for 10 min and
following differentiation in 1% acid alcohol for 30 sec, the slides
were dipped in lithium carbonate for bluing for 5 min and were
stained with eosin for 15 sec. Then dehydrated with 75% alcohol,
95% alcohol and 100% alcohol, 5 min each, cleared in xylene and
mounted. All these steps were performed at room temperature. The
study was approved by The Medical Ethical Committee of The First
Affiliated Hospital of Sun Yat-sen University. Written informed
consent was obtained from all patients for use of their clinical
specimens.
RNA extraction and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
Total RNA from the IDH-wild type GBM and non-tumour
tissue samples was extracted using TRIzol® reagent
(Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA)
according to the manufacturer's protocol. RNA concentration and
quality were assessed spectrophotometrically at 260 and 280 nm.
RT-qPCR was performed using an ABI PRISM® 7500 Sequence
Detection system (Applied Biosystems; Thermo Fisher Scientific,
Inc.). Reverse transcription was carried out using the TaKaRa
PrimeScript™ RT reagent Kit with gDNA Eraser (Perfect Real Time)
(RR047A, TaKaRa Biotechnology, Inc., Otsu, Japan) according to the
manufacturer's protocol. The primer sequences used for the
reactions were designed as follows (7): CAV1, forward 5′-TACTGGTTTTACCGCTTGCT-3′,
reverse 5′-ACGGCTGATGCACTGAATC-3′; HIF-1α, forward
5′-GTGGATTACCACAGCTGA-3′, reverse 5′-GCTCAGTTAACTTGATCCA-3′; and
18S, forward 5′-CCTGGATACCGCAGCTAGGA-3′ and reverse
5′-GCGGCGCAATACGAATGCCCC-3′. According to the qPCR kit
(SYBR® Premix Ex Taq™, RR430A, TaKaRa
Biotechnology, Inc.) according to the manufacturer's protocol, the
qPCR protocol used was as follows: Denaturation at 95°C for 5 min,
followed by 40 cycles of denaturation at 95°C for 15 sec and
annealing for 32 sec at 60°C. The housekeeping gene 18S was used as
the reference gene, and the 2−ΔΔCq method was used to
analysis the CAV1 and HIF-1α Expression. Expression data were
normalized to the geometric mean of the housekeeping gene 18S
(14).
Western blotting
A total of 17 matched pairs of cancerous tissues and
healthy tissues were homogenized in 50 mM Tris (pH 7.5), 100 mM
NaCl, 1 mM EDTA, 0.5% NP40, 0.5% Triton X-100, 2.5 mM sodium
orthovanadate, 10 µM protease inhibitor cocktail, and 1 mM
phenylmethylsulfonyl fluoride. Protein determination was performed
using a Pierce™ BCA Protein assay kit (Thermo Fisher Scientific
Inc.) to quantify the total protein following extraction, and 20 µg
protein were loaded per lane and electrophoretically separated on a
9% SDS polyacrylamide gel and transferred to polyvinylidene
fluoride membranes (EMD Millipore, Billerica, MA, USA). The
membranes were blocked with 10% BSA (Thermo Fisher Scientific,
Inc.) in PBST, at room temperature for 1 h, then were incubated at
4°C overnight with anti-human HIF-1α rabbit monoclonal antibody
(cat no. GTX127309, 1:1,000; GeneTex Inc., Irvine, CA, USA) and
CAV1 rabbit monoclonal antibody (cat no. ab192869, 1:1,000; Abcam,
Cambridge, USA). HIF-1α and CAV1 expressions levels were detected
with horseradish peroxidase (HRP)-conjugated goat anti-rabbit
immunoglobulin G secondary antibody following incubation at room
temperature for 1 h (cat no. 4050-05; 1:5,000; SouthernBiotech,
Birmingham, AL, USA). Anti-GAPDH mouse monoclonal antibody (cat no.
2118; 1:10,000; Cell Signaling Technology, Inc., Danvers, MA, USA)
was used as a loading control. Protein bands were eventually
visualized with an automatic chemiluminescence imaging method and
the results were analyzed by imaging analysis software (Version
5200, Tanon Science and Technology Co., Ltd., Shanghai, China).
Immunohistochemistry
Immunohistochemical staining was carried out on
formalin-fixed, paraffin-embedded sections (4 µm) deparaffinized in
xylene at room temperature for 10 min twice, rehydrated in an
ethanol series (absolute alcohol for 5 min twice, 95% alcohol for 2
min and 70% alcohol for 2 min) at room temperature and rinsed in
PBS. Antigen retrieval was then performed in a microwave with 10 mM
citrate buffer for 10 min (pH 6.0). Immunohistochemical staining
was carried out using the EnVision™ kit (Dako; Agilent
Technologies, Inc., Santa Clara, CA, USA) according to the
manufacturer's protocol. Endogenous peroxidase activity was
quenched by treatment with 3% hydrogen peroxide at room temperature
for 15 min. The sections were then incubated with primary
anti-rabbit antibody against HIF-1α (cat no. GTX127309; 1:500;
GeneTex Inc.) and CAV1 (cat no. ab192869; 1:500; Abcam) overnight
at 4°C. Subsequently, the tissue sections were sequentially
incubated with ready-to-use HRP immunoglobulin (EnVision kit) at
room temperature for 30 min and developed with 3,3-diaminobenzidine
as a chromogen substrate. The nuclei were eventually counterstained
with Meyer's haematoxylin at room temperature for 2–5 min. Images
were captured using an inverted microscope (×40 and ×400
magnification) (Nikon eclipse E100).
The levels of HIF-1α and CAV1 immunostaining were
evaluated independently by two pathologists who did not know the
survival outcomes of the participants. Positive CAV1 protein
expression was defined as a diffuse brown staining in the cell
membrane, whereas positive HIF-1α immunoreactivity was observed in
the cell nuclei.
The immunostaining levels were evaluated by
immunoreactive score (IRS). Staining intensity was scored as
follows: No staining at all (score 0), faint staining (score 1),
moderate staining (score 2) and strong staining (score 3). The
protein distribution was defined as the percentage accounting for
the whole area in the section: 0% (score 0), 1–25% (score 1),
26–50% (score 2), 51–75% (score 3) and 76–100% (score 4). Total
scores were calculated by combining the staining intensity
evaluation and the staining distribution. The scores were
independently evaluated by two researchers who had to reach an
agreement. If divergences appeared, a third researcher participated
in the evaluation to obtain the final score.
The median value of CAV1 IRS for all samples was 4.
The CAV1 protein expression levels were therefore further analysed
and IRS values were classified as low (IRS value <4) or high
(IRS value ≥4). The median value of HIF-1α IRS for all samples was
5. The HIF-1α protein expression levels were likewise further
analysed, and IRS values were categorized as low (IRS value <5)
and high (IRS value ≥5).
Statistical analysis
Statistical calculations were performed using SPSS
20.0 (IBM Corp., Armonk, NY, USA) and GraphPad Prism v5.0 (GraphPad
Software, Inc., La Jolla, CA, USA). Data were expressed as the mean
± standard error of the mean. Differences in CAV1 and HIF-1α
expression between the two groups were compared by paired t-test.
The protein and mRNA levels of CAV1 and HIF-1α in the tissues of
patients with GBM were categorized as low expression or high
expression, according to their mean value. A χ2 test was
applied to determine the association between CAV1 expression and
the clinicopathological parameters of GBM. The Kaplan-Meier method
and log-rank test were used to evaluate and compare the prognosis
of patients with GBM. The HIF-1α IRS of each tissue was compared
with CAV1 IRS using Spearman's correlation coefficients (r) and
curve-estimation analysis. P<0.05 was considered to indicate a
statistically significant difference.
Results
HIF-1α and CAV1 are overexpressed in
IDH-wild type GBM
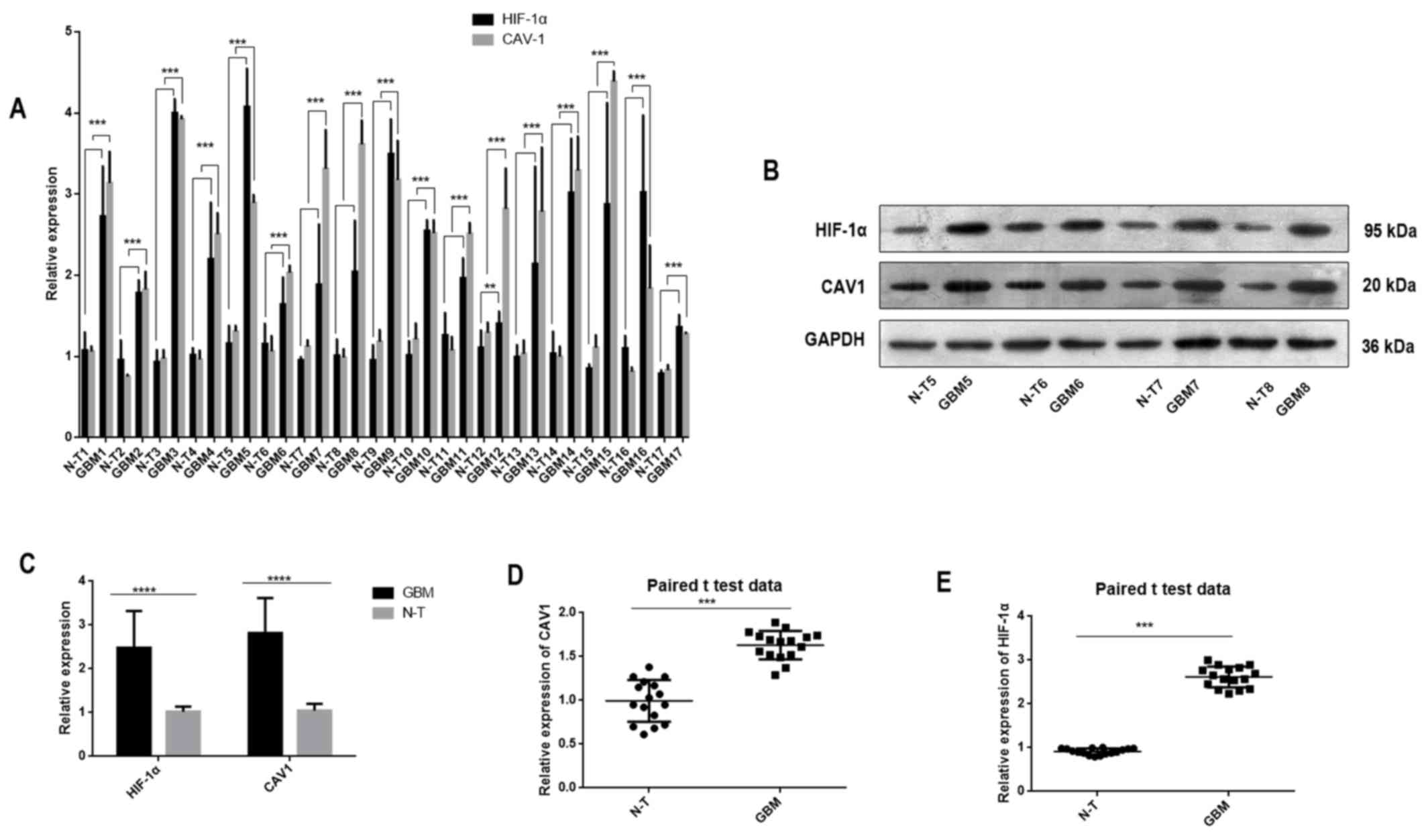
In the present study, RT-qPCR and western blotting
were performed to determine the mRNA and protein expression levels,
respectively, of HIF-1α and CAV1 in non-tumour and IDH-wild type
GBM (Fig. 1). The data revealed that
HIF-1α and CAV1 expression levels were significantly higher in
cancerous tissues than in non-tumour tissues.
Upregulation of CAV1 is highly
correlated with HIF-1α expression
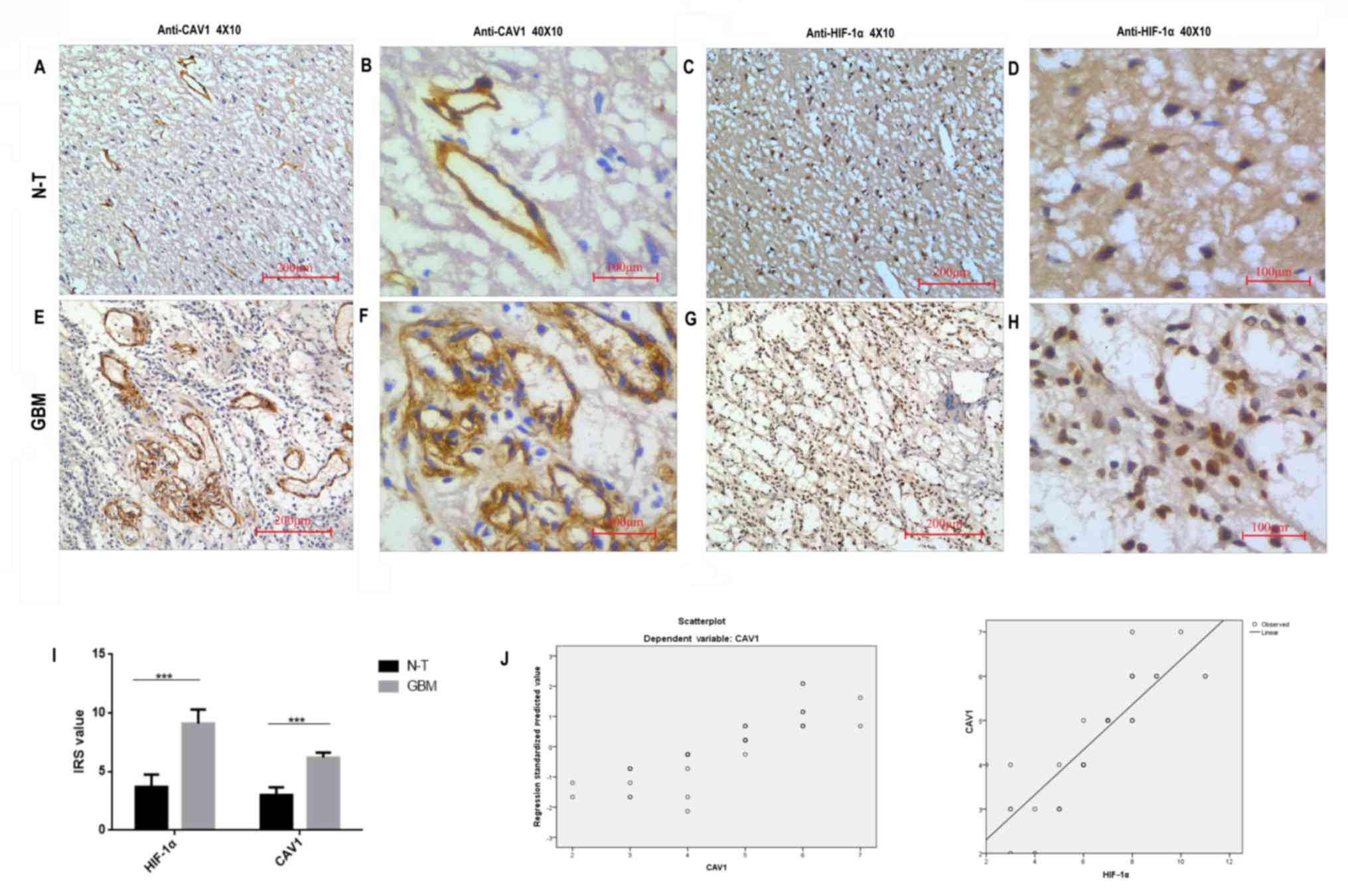
To further understand the cellular localization of
the CAV1 and HIF-1α proteins, immunohistochemistry was performed on
tissues. The immunohistochemical staining patterns of HIF-1α and
CAV1 in the 42 patients indicated that HIF-1α was mainly localized
in the nucleus of the tumour cells, whereas CAV1 was located in the
cell membrane or cytoplasm of the tumour cells. The representative
images of immunohistochemical staining are presented in Fig. 2A-H. According to statistical analysis,
the expression levels of HIF-1α and CAV1 protein in GBM tissues
were significantly higher (P<0.001) than in non-tumour tissues.
In addition, the association between HIF-1α and CAV1 expression was
assessed by nonparametric Spearman's rank test and logistic
regression analysis (Fig. 2J). The
results demonstrated that the upregulation of CAV1 was positively
correlated with HIF-1α expression (P<0.01; r=0.765).
High levels of HIF-1α and CAV1 are
associated with poor prognosis for patients with IDH-wild type
GBM
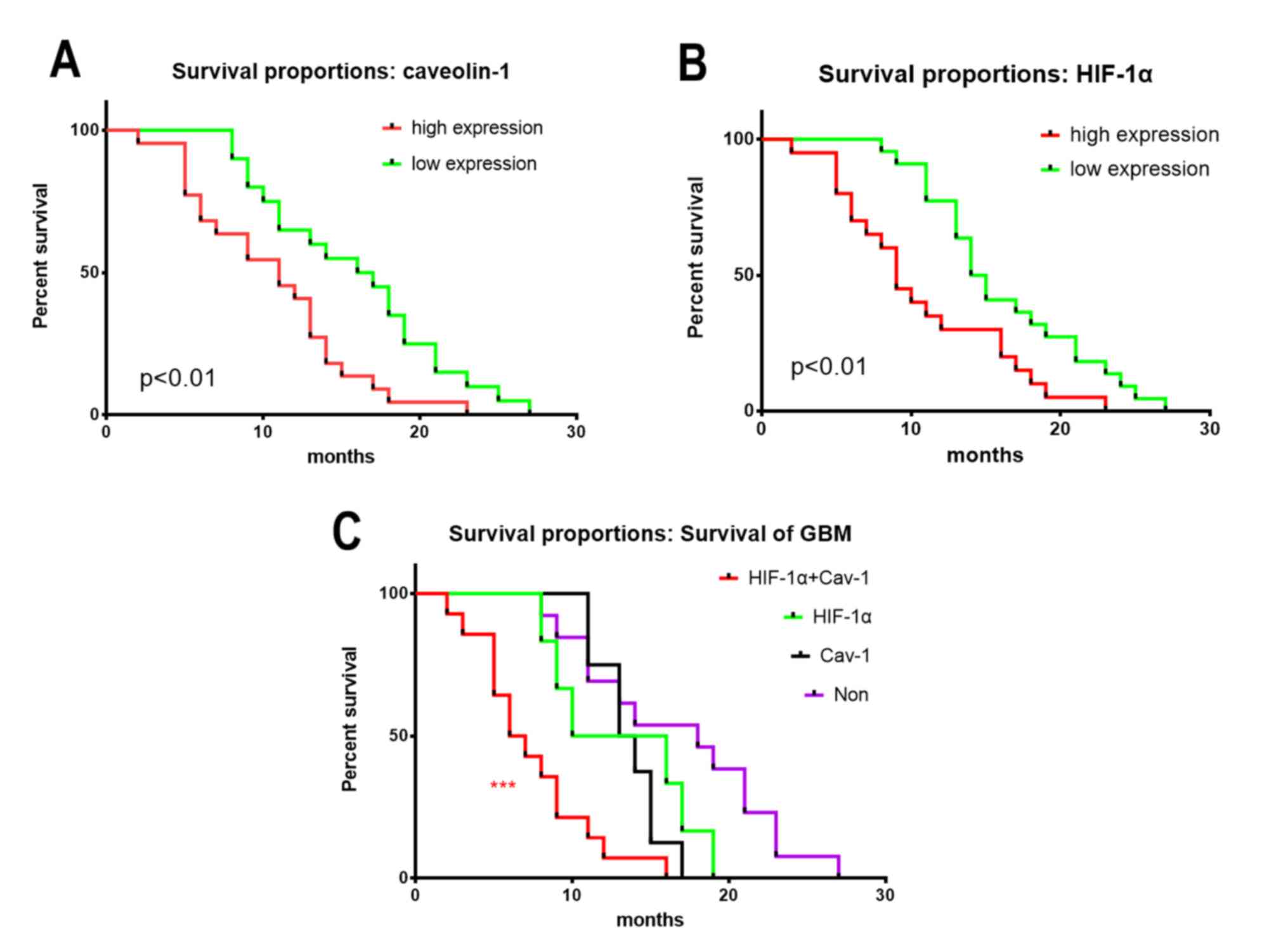
The association between HIF-1α and CAV1 expression
and the survival time of patients with GBM was further analysed,
high HIF-1α or CAV1 expression associated with larger size and
worse prognosis (Table I). The
results revealed that patients with IDH-wild type GBM and high
HIF-1α or CAV1 expression had a poorer prognosis compared to
patients with low levels of HIF-1α or CAV1. In addition, patients
with higher expression of HIF-1α and CAV1 had the poorest prognosis
(Fig. 3).
 | Table I.Clinicopathological parameters of 42
patients with IDH-wild type GBM. |
Table I.
Clinicopathological parameters of 42
patients with IDH-wild type GBM.
|
|
| CAV1 | HIF-1α |
|---|
|
|
|
|
|
|---|
| Variable | Number | Low expression | High expression | P-value | Low expression | High expression | P-value |
|---|
| Age |
| <60
years | 19 | 9 | 10 | 0.954 | 11 | 8 | 0.358 |
| ≥60
years | 23 | 12 | 11 |
| 9 | 14 |
|
| Sex |
| Male | 17 | 7 | 10 | 0.236 | 8 | 9 | 0.764 |
|
Female | 25 | 14 | 11 |
| 12 | 13 |
|
| KPS |
| ≤70 | 28 | 16 | 12 | 0.286 | 15 | 13 | 0.395 |
|
<70 | 14 | 6 | 8 |
| 5 | 9 |
|
| Size |
| <5
cm | 19 | 12 | 7 |
<0.0001a | 15 | 4 | 0.213 |
| ≥5
cm | 23 | 8 | 15 |
| 5 | 18 |
|
| MGMT |
|
Positive | 26 | 12 | 14 | 0.392 | 11 | 15 | 0.315 |
|
Negative | 16 | 10 | 6 |
| 9 | 7 |
|
| Survival |
| <13
m | 20 | 6 | 14 | 0.029a | 7 | 13 | 0.013a |
| ≥13
m | 22 | 16 | 6 |
| 13 | 9 |
|
Discussion
CAV1, an integral structural component of caveolae,
is a direct target of HIF-1. Numerous studies have recently
demonstrated that hypoxia-associated biological markers can be used
as predictors of treatment, metastasis and prognosis in various
types of cancer, including breast, lung and colorectal cancer
(15–17). The present study revealed that HIF-1α
and CAV1 were significantly upregulated in patients with IDH-wild
type GBM. In addition, the overexpression of HIF-1α and CAV1 was
markedly associated with poor survival rates in GBM. These results
suggested that HIF-1α and CAV1 may be potential markers for
treatment and prognosis in IDH-wild type GBM.
At present, the markers available to predict
prognosis of IDH-wild type GBM patients are insufficient. Patients
with IDH-wild type GBM usually have low survival rates. Although
hypoxia is usually present in the tumour environment, it represents
an uncertain prognostic marker in various types of cancer. It has
been described that HIF-1α, an important transcription factor,
regulates numerous biological functions and activates critical
genes involved in angiogenesis, migration, invasion and metastasis
(18,19). Cai et al (20) reported that HIF-1α is a biomarker
useful for the identification of subgroups of patients with a poor
prognosis, and for the early detection of subclinical metastasis.
As previously mentioned, hypoxia is a critical feature of the
glioma microenvironment and is associated with a poor prognosis and
resistance to most therapies (21);
however, only a few studies have investigated the expression of
HIF-1α in GBM. Further investigations are therefore required to
understand the role of HIF-1α expression in GBM. In the present
study, HIF-1α expression was upregulated in GBM tissues at the mRNA
and protein levels, compared to adjacent healthy tissues, which
confirmed previous findings. In addition, the overexpression of
HIF-1α in GBM was significantly associated with poor prognosis.
The role of HIF-1α in cancer pathogenesis and
prognosis is complex. A recent study reported that hypoxia
regulates membrane protein endocytosis through CAV1 in cancer cells
(21). CAV1 has also been implicated
in tumour development and described as a negative regulator of
endocytosis (22,23). It has been demonstrated that the lipid
raft-associated CAV1 negatively regulates the uptake of exosomes
derived from GBM cells via extracellular signal-regulated kinases
1/2-heat shock protein 27 signalling (24). The present study detected higher CAV1
expression in GBM tissues compared to healthy tissues. This result
was similar to previous findings. Bourseau-Guilmain et al
(21) observed significantly enhanced
expression of CAV1 in hypoxic regions of tumours from patients with
GBM. To further analyse the association between HIF-1α and CAV1
expression, the present study confirmed that there was a
significant correlation between both proteins. In addition, CAV1
was positively associated with poor prognosis for patients with
GBM. Therefore, high expression levels of HIF-1α and CAV1 may be
associated with shorter survival rate; however, the underlying
molecular mechanisms of action of HIF-1α and CAV1 in GBM require
further investigation.
In conclusion, the present study investigated HIF-1α
and CAV1 expression in IDH-wild type GBM and healthy tissues. High
expression levels of HIF-1α and CAV1 in patients with IDH-wild type
GBM were demonstrated. Furthermore, poor patient prognosis was
associated with high expression levels of HIF-1α and CAV1. These
findings suggested that HIF-1α and CAV1 expression levels may aid
in the identification of patients with a poor prognosis that
require more aggressive treatment.
Acknowledgements
Not applicable.
Funding
The present study was supported by a grant from The
Technology Project of Huangpu District Guangzhou City (grant no.
201611).
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
WLC, XC and XBW performed the staining and western
blotting. JSW, XLW and CFX performed the data analysis. CFX oversaw
the study, and CXL conceived the study and made substantial
intellectual contributions. CFX and CXL designed this article,
helped to analyse the data and revised the final manuscript. All
authors read and approved the final manuscript.
Ethics approval and consent to
participate
The study was approved by The Medical Ethics
Committee of The First Affiliated Hospital of Sun Yat-sen
University. Written informed consent was obtained from all patients
for use of their tissues.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Kaya V, Yıldırım M, Yazıcı G, Yalçın AY,
Orhan N and Güzel A: Prognostic significance of indicators of
systemic inflammatory responses in glioblastoma multiforme
patients. Asian Pac J Cancer Prev. 18:3287–3291. 2017.PubMed/NCBI
|
|
2
|
Koshy M, Villano JL, Dolecek TA, Howard A,
Mahmood U, Chmura SJ, Weichselbaum RR and McCarthy BJ: Improved
survival time trends for glioblastoma multiforme using the SEER 17
population-based registries. J Neurooncol. 107:207–212. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Waitkus M, Diplas B and Yan H: Isocitrate
dehydrogenase mutations in gliomas. Neuro Oncol. 18:16–26. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Jun JC, Rathore A, Younas H, Gilkes D and
Polotsky VY: Hypoxia-inducible factors and cancer. Curr Sleep Med
Rep. 3:1–10. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Qin J, Liu Y, Lu Y, Liu M, Li M, Li J and
Wu L: Hypoxia-inducible factor 1 alpha promotes cancer stem
cells-like properties in human ovarian cancer cells by upregulating
SIRT1 expression. Sci Rep. 7:105922017. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Said HM, Hagemann C, Staab A, Stojic J,
Kühnel S, Vince GH, Flentje M, Roosen K and Vordermark D:
Expression patterns of the hypoxia-related genes osteopontin, CA9,
erythropoietin, VEGF and HIF-1alpha in human glioma in vitro and in
vivo. Radiother Oncol. 83:398–405. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Agrawal R, Pandey P, Jha P, Dwivedi V,
Sarkar C and Kulshreshtha R: Hypoxic signature of microRNAs in
glioblastoma: Insights from small RNA deep sequencing. BMC
Genomics. 15:6862014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kaelin WG Jr: Molecular basis of the VHL
hereditary cancer syndrome. Nat Rev Cancer. 2:673–682. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Mao X, Wong SY, Tse EY, Ko FC, Tey SK,
Yeung YS, Man K, Lo RC, Ng IO and Yam JW: Mechanisms through which
hypoxia-induced caveolin-1 drives tumorigenesis and metastasis in
hepatocellular carcinoma. Cancer Res. 76:7242–7253. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Goetz JG, Minguet S, Navarro-Lérida I,
Lazcano JJ, Samaniego R, Calvo E, Tello M, Osteso-Ibáñez T,
Pellinen T, Echarri A, et al: Biomechanical remodeling of the
microenvironment by stromal caveolin-1 favors tumor invasion and
metastasis. Cell. 146:148–163. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang Y, Roche O, Xu C, Moriyama EH, Heir
P, Chung J, Roos FC, Chen Y, Finak G, Milosevic M, et al: Hypoxia
promotes ligand-independent EGF receptor signaling via
hypoxia-inducible factor-mediated upregulation of caveolin-1. Proc
Natl Acad Sci USA. 109:4892–4897. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kannan A, Krishnan A, Ali M, Subramaniam
S, Halagowder D and Sivasithamparam ND: Caveolin-1 promotes gastric
cancer progression by up-regulating epithelial to mesenchymal
transition by crosstalk of signalling mechanisms under hypoxic
condition. Eur J Cancer. 50:204–15. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hoshide R and Jandial R: 2016 world health
organization classification of central nervous system tumors: An
Era of molecular biology. World Neurosurg. 94:561–562. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Milani M, Venturini S, Bonardi S, Allevi
G, Strina C, Cappelletti MR, Corona SP, Aguggini S, Bottini A,
Berruti A, et al: Hypoxia-related biological markers as predictors
of epirubicin-based treatment responsiveness and resistance in
locally advanced breast cancer. Oncotarget. 8:78870–78881. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Khan MN, Haggag YA, Lane ME, McCarron PA
and Tambuwala MM: Polymeric nano-encapsulation of curcumin enhances
its anti-cancer activity in breast (MDA-MB231) and lung (A549)
cancer cells through reduction in expression of HIF-1α and nuclear
p65 (Rel A). Curr Drug Deliv. 15:286–295. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li W, Zong S, Shi Q, Li H, Xu J and Hou F:
Hypoxia-induced vasculogenic mimicry formation in human colorectal
cancer cells: Involvement of HIF-1a, claudin-4, and E-cadherin and
vimentin. Sci Rep. 6:375342016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Cavallo F, De Giovanni C, Nanni P, Forni G
and Lollini PL: 2011: The immune hallmarks of cancer. Cancer
Immunol Immunother. 60:319–26. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Cai FF, Xu C, Pan X, Cai L, Lin XY, Chen S
and Biskup E: Prognostic value of plasma levels of HIF-1a and
PGC-1a in breast cancer. Oncotarget. 7:77793–77806. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bourseau-Guilmain E, Menard JA, Lindqvist
E, Indira Chandran V, Christianson HC, Cerezo Magaña M, Lidfeldt J,
Marko-Varga G, Welinder C and Belting M: Hypoxia regulates global
membrane protein endocytosis through caveolin-1 in cancer cells.
Nat Commun. 7:113712016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Le PU, Guay G, Altschuler Y and Nabi IR:
Caveolin-1 is a negative regulator of caveolae-mediated endocytosis
to the endoplasmic reticulum. J Biol Chem. 277:3371–3379. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Shvets E, Ludwig A and Nichols BJ: News
from the caves: Update on the structure and function of caveolae.
Curr Opin Cell Biol. 29:99–106. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Svensson KJ, Christianson HC, Wittrup A,
Bourseau-Guilmain E, Lindqvist E, Svensson LM, Mörgelin M and
Belting M: Exosome uptake depends on ERK1/2-heat shock protein 27
signaling and lipid Raft-mediated endocytosis negatively regulated
by caveolin-1. J Biol Chem. 288:17713–17724. 2013. View Article : Google Scholar : PubMed/NCBI
|