Introduction
Gastric cancer is a common malignancy worldwide that
affects >1% of the worldwide population (1). A large number of patients with gastric
cancer are diagnosed with inoperable tumors or experience high
recurrence rates following curative resection (2). An unhealthy diet, cigarette smoking and
alcohol consumption have been identified as risk factors for
gastric cancer and reduced exposure to risk factors significantly
reduces the incidence of the disease (3–5). It has
been revealed genetic alterations as risk factors for gastric
cancer (3–6); however, the limited number of oncogenes
and tumor suppressors currently identified do not sufficiently
explain the complex pathogenesis of gastric cancer (6).
Previous studies have shown that long non-coding
RNAs (lncRNAs), transcripts >200 nucleotides in length, are not
transcriptional ‘noise’, but serve important roles in growth and
developmental processes (7,8). lncRNAs regulate gene expression at
multiple levels, and altered expression of lncRNAs may result in
dysregulated gene expression, thereby contributing to the
occurrence of diseases, such as cancer (9,10).
Therefore, characterization of the functions of lncRNAs may provide
insights into disease prevention and treatment. However, the
function of the majority of predicted or isolated lncRNAs remains
unclear. PMS1 homolog 2 mismatch repair system component pseudogene
2 (PMS2L2) is a recently identified lncRNA that exhibits protective
effects in chondrocytes during lipopolysaccharide-induced
inflammation (11). Preliminary deep
sequencing data revealed that PMS2L2 expression was downregulated
in gastric adenocarcinoma (GA) tissues in comparison with adjacent
non-tumor tissues and was inversely associated with microRNA
(miR/miRNA)-25 expression levels (data not shown). A previous study
revealed that miR-25 expression was upregulated in GA tissues
compared with adjacent non-tumor tissues and promoted the in
vitro proliferation, invasion and migration of GA cells
(12). Furthermore, high levels of
miR-25 predicted poor prognosis of patients with GA. Therefore, the
present study investigated the role of PMS2L2 in GA and its
potential interactions with miR-25.
Materials and methods
Specimens and patients
Tissue specimens, including GA and healthy adjacent
non-cancerous tissues (within 2 cm of the tumor margin), were
obtained from 72 patients with GA at The Second Hospital of
Shandong University between May 2010 and May 2015.
Histopathological examination revealed that the cancer cell content
in all adjacent non-cancerous tissues was <1%. The patients
included 38 males and 34 females (age range, 33–68 years; mean age,
52.2±8.3 years). The inclusion criteria were as follows: i) New GA
cases diagnosed by histopathological examination; ii) no previous
history of malignancies; and iii) no familial history of
malignancies. The exclusion criteria were as follows: i) Patients
transferred from other hospitals; ii) patients who received
treatment within 3 months prior to admission; and iii) patients
with co-morbidities. Based on the criteria established by the
American Joint Committee on Cancer (13), there were 14, 16, 28 and 14 patients
with stage I–IV cancer, respectively. The present study was
approved by the Ethics Committee of The Second Hospital of Shandong
University and all patients provided written informed consent.
Follow-up
A 5-year follow-up study was performed to monitor
the survival rate of the patients. Patients visited the outpatients
department or were monitored remotely over the telephone every
month. Patients who succumbed to other clinical disorders,
including heart disease, or accidents, including traffic accidents,
were excluded from the study. The aforementioned 72 patients
completed the follow-up study.
Cell lines and cell culture
The human GA cell line AGS (Sigma-Adrich; Merck
KGaA) was used to perform all in vitro experiments. The
normal gastric cell line Hs 738.St/Int (cat. no. CRL-7869™;
American Type Culture Collection) served as healthy control cells.
Cells were cultured in F-12K medium (Sigma-Adrich; Merck KGaA)
supplemented with 10% fetal bovine serum (FBS; Sigma-Adrich; Merck
KGaA) and maintained at 37°C and 5% CO2.
Transient transfections
The PMS2L2 expression vector was constructed by
inserting full length PMS2L2 cDNA into a pcDNA3.1 vector (Sangon
Biotech Co., Ltd.). Negative control (NC) miRNA
(5′-CUAGUCGUGUAUACAGUGUGA-3′) and the miR-25 mimic
(5′-CAUUGCACUUGUCUCGGUCUGA-3′) were purchased from Sigma-Aldrich,
Merck KGaA. All cell transfections were performed using
Lipofectamine® 2000 (Thermo Fisher Scientific, Inc.) and
10 nM vector and 35 nM miRNA with all procedures performed
following the manufacturer's instructions. Subsequent experiments
were performed 24 h post-transfection. Negative controls included
cells transfected with an empty vector or NC miRNA and controls
included untransfected cells.
Reverse-transcription quantitiative
polymerse chain reaction (RT-qPCR)
Total RNA was extracted from GA and healthy adjacent
non-cancerous tissues as well as the AGS and Hs 738.St/Int cell
lines using RNAzol reagent (Sigma-Aldrich; Merck KGaA). Total RNA
was reverse transcribed into cDNA using AMV Reverse Transcriptase
XL (Clontech Laboratories, Inc.). The expression of PMS2L2 was
analyzed by qPCR using the DyNAmo Flash SYBR Green qPCR kit (Thermo
Fisher Scientific, Inc.) and 18S rRNA as an endogenous control.
The miRNA isolation kit (cat. no. RMI050; Geneaid
Biotech Ltd.) was used to extract miRNA from AGS and Hs 738.St/Int
cell lines. The TaqMan MicroRNA Reverse Transcription kit (Thermo
Fisher Scientific, Inc.) was used to perform reverse transcription.
The expression of miR-25 was analyzed by qPCR using the TaqMan
Real-Time PCR Master mix (Thermo Fisher Scientific, Inc.) and U6 as
an endogenous control.
Primer sequences were: PMS2L2 forward,
5′-AGTCTAAGCACTGCGGTGAA-3′ and reverse, 5′-GGTAGAAATGGTGACATCAT-3′;
18S rRNA forward, 5′-CTACCACATCCAAGGAAGCA-3′ and reverse,
5′-TTTTTCGTCACTACCTCCCCG-3′. The forward primer sequence of miR-25
was: 5′-CATTGCACTTGTCTCGGTC-3′. Universal reverse primers and U6
forward primer were from the kit. The thermocycling conditions
were: 95°C for 1 min, then 40 cycles of 95°C for 10 sec and 60°C
for 50 sec. All qPCR reactions were performed in triplicate and the
2−ΔΔCq method (14) was
used to quanitfy expression levels.
Cell migration and invasion
AGS and Hs 738.St/Int cells were harvested 24 h
post-transfection. A single cell suspension was prepared using
serum-free F-12K medium and the final cell density was adjusted to
3×104 cells/ml. A total of 0.1 ml cell suspension was
plated in the upper chamber of Corning Transwell Cell Culture Plate
Insert (8.0 µm pore; Corning, Inc.). The lower chamber was filled
with F-12K medium (supplemented with 20% FBS). For invasion assays,
the Transwell membranes were coated with Matrigel (EMD Millipore)
for 6 h at 37°C. Following incubation at 37°C for 2.5 h, cells were
fixed in ice-cold 70% ethanol for 20 min at 4°C. The migratory
cells were then stained with 0.5% crystal violet (Sigma-Aldrich;
Merck KGaA) for 16 min at room temperature. Stained cells were
observed under an optical microscope and cells of five randomly
selected visual fields were counted (magnification, ×40). The
control groups (untransfected cells) were set to 100% and all other
groups were normalized to the control groups.
Statistical analysis
Data are presented as the mean ± standard deviation
values calculated from experiments performed in triplicate.
GraphPad prism 6 (GraphPad Software, Inc.) was used to perform all
statistical analysis. Differences between GA tissues and healthy
adjacent tissues were analyzed by the paired t-test. Differences
among cell transfection groups or clinical stages were analyzed by
the one-way ANOVA followed by Tukey's post hoc test. Associations
between PMS2L2 and miR-25 were analyzed by linear regression.
Patients were grouped into high (n=33) and low (n=39) PMS2L2 level
groups based on Youden's index (cut-off value=2.14). Survival
curves were plotted using the Kaplan-Meier method and compared by
the log-rank test. P<0.05 was used to indicate a statistically
significant difference.
Results
PMS2L2 is downregulated in GA
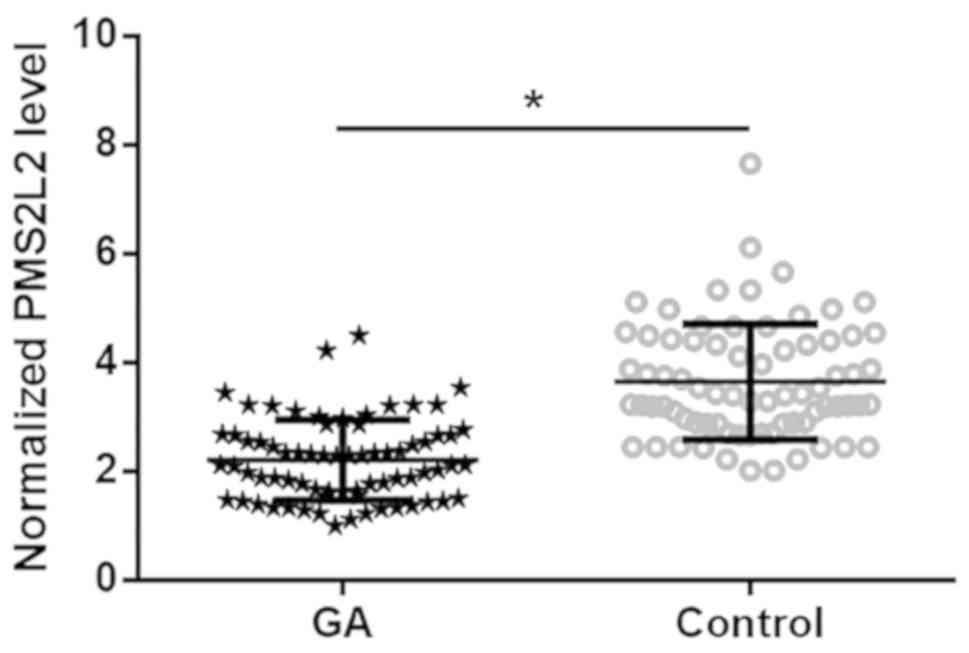
PMS2L2 expression in tissues was detected by
RT-qPCR. The GA tissue with the lowest expression level was set to
‘1’, and all other samples were normalized to this sample.
Expression data were analyzed by the paired t-test. PMS2L2 was
significantly downregulated in GA tissues compared with healthy
adjacent tissues (Fig. 1;
P<0.05).
PMS2L2 is not affected by clinical
stage but is associated with survival rate
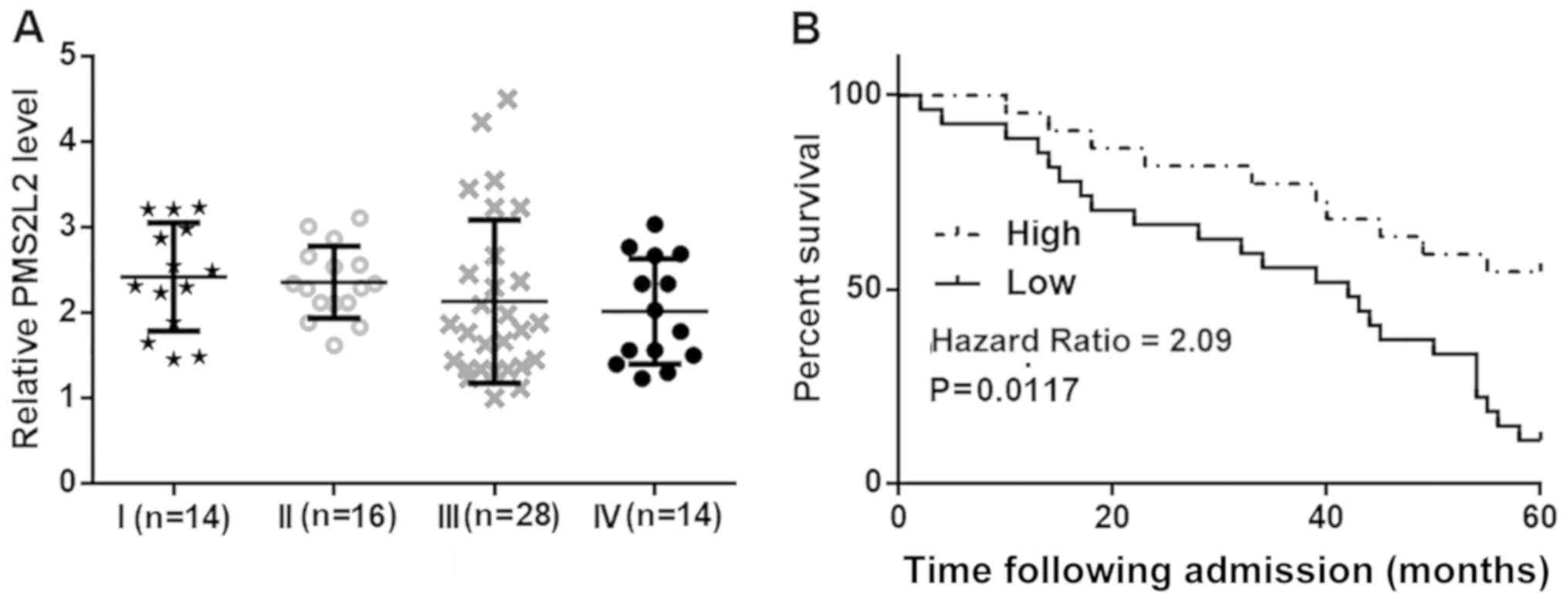
PMS2L2 expression in GA tissues obtained from
patients with different clinical stages was compared by the one-way
ANOVA and Tukey's post hoc test. There were no significant
differences in PMS2L2 levels among these groups of patients
(Fig. 2A). Subsequently, patients
were grouped into high (n=33) and low (n=39) PMS2L2 expression
level groups based on Youden's index. As shown in Fig. 2B, patients with low levels of PMS2L2
exhibited a significantly shorter survival rate compared with
patients with high expression (P=0.0117).
miR-25 is upregulated in GA and
inversely associated with PMS2L2
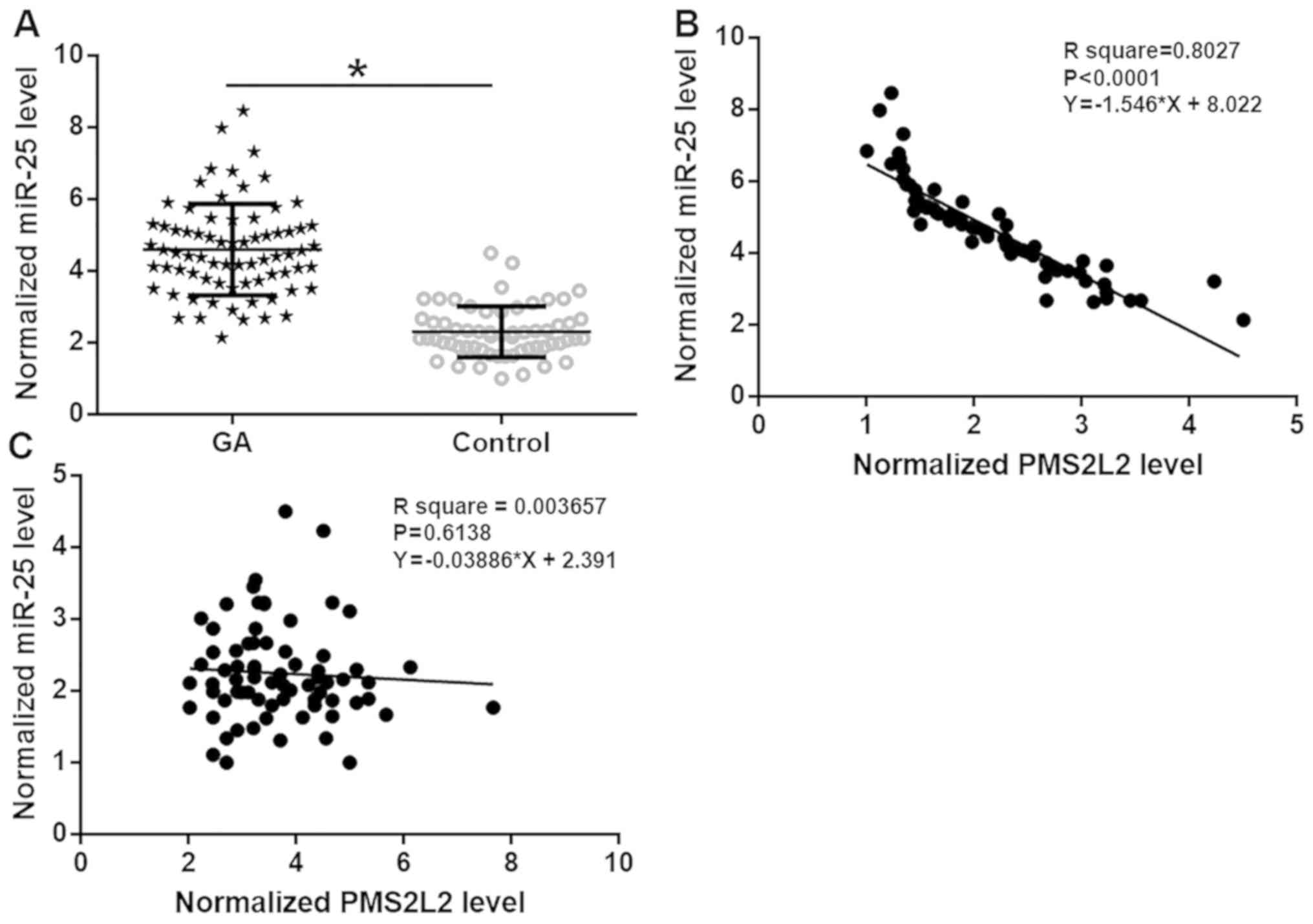
miR-25 expression in tissues was detected by
RT-qPCR. The GA tissue with the lowest expression level was set to
‘1’, and all other samples were normalized to this sample.
Expression data were analyzed by the paired t-test. miR-25 was
significantly upregulated in GA tissues compared with healthy
adjacent tissues (Fig. 3A;
P<0.05). Associations between PMS2L2 and miR-25 were analyzed by
linear regression. PMS2L2 and miR-25 expression levels were
significantly inversely associated in GA tissues (Fig. 3B, R2=0.8207, P<0.0001)
but not in healthy adjacent tissues (Fig. 3C R2=0.003657,
P=0.6138).
PMS2L2 downregulates miR-25 to
regulate the migration and invasion of GA cells but not normal
gastric tissue cells
A PMS2L2 overexpression vector and miR-25 mimic were
transfected into AGS and Hs 738.St/Int cells, and compared with the
NC and control cells, PMS2L2 (Fig.
4A) and miR-25 (Fig. 4B)
expression levels were significantly upregulated 24 h
post-transfection (both P<0.05). In addition, overexpression of
PMS2L2 downregulated the expression of miR-25 in AGS cells but not
in Hs 738.St/Int (Fig. 4A;
P<0.05), while miR-25 overexpression did not significantly
affect PMS2L2 expression in both cell lines (Fig. 4B, both P>0.05). Transwell
migration and invasion assays revealed that PMS2L2 overexpression
inhibited migration (Fig. 4C) and
invasion (Fig. 4D) of AGS cells
(both P<0.05) but not Hs 738.St/Int cells. Moreover, miR-25
overexpression partially rescued the decreased migration and
invasion of AGS cells caused by PMS2L2 overexpression.
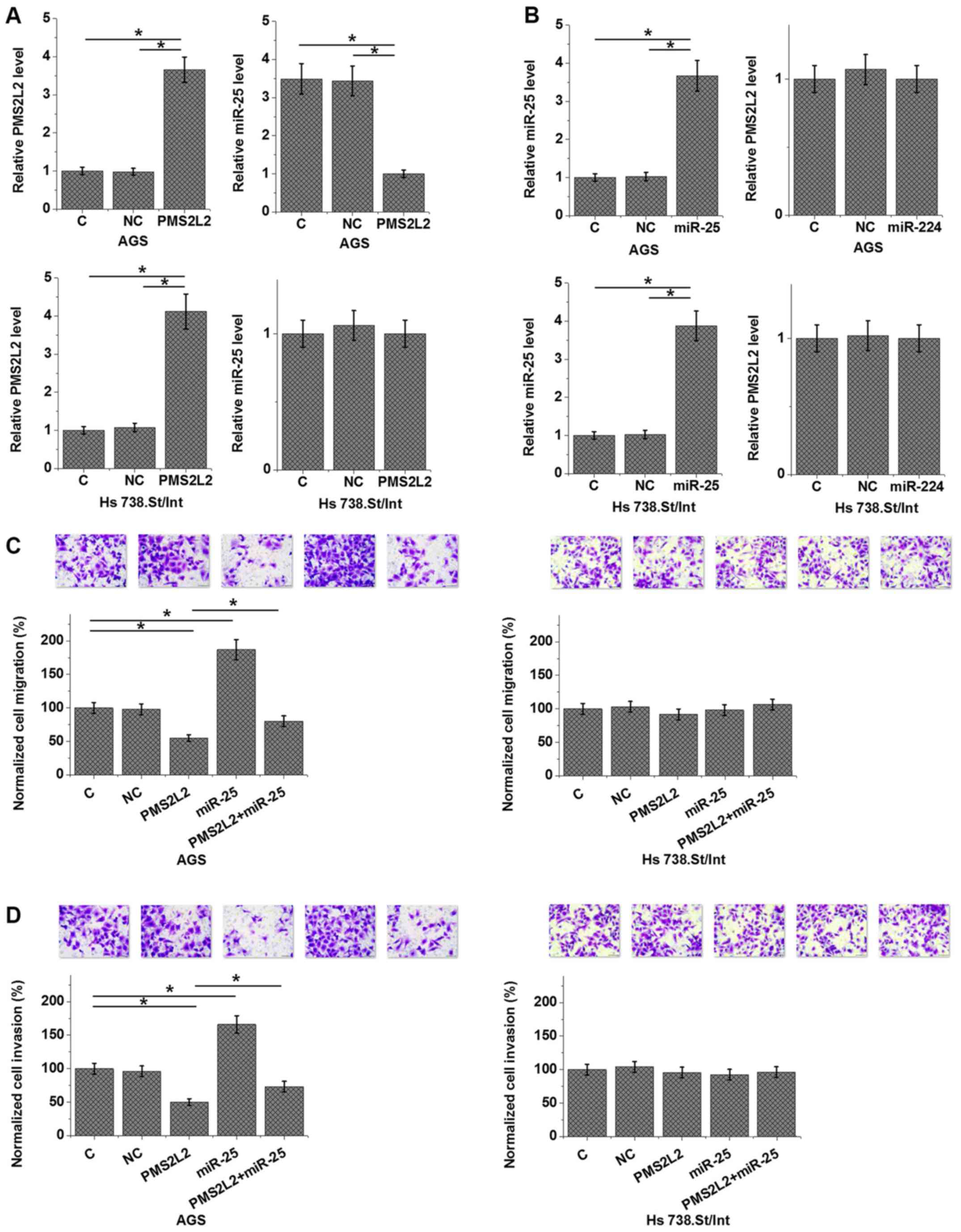 | Figure 4.PMS2L2 downregulated miR-25 to
regulate the migration and invasion of gastric adenocarcinoma
cells, but not normal gastric tissue cells. Compared with the C and
NC cells, cells transfected with a PMS2L2 overexpression vector and
an miR-25 mimic exhibited significantly upregulated (A) PMS2L2 and
(B) miR-25 levels 24 h post-transfection. Furthermore,
overexpression of PMS2L2 downregulated the expression of miR-25 in
AGS cells but not in Hs 738.St/Int, while miR-25 overexpression did
not significantly affect PMS2L2 expression in both cell lines.
Transwell assays revealed that PMS2L2 overexpression inhibited the
(C) migration and (D) invasion of AGS cells but not Hs 738.St/Int
cells. Moreover, miR-25 overexpression partially rescued the
decreased migration and invasion of AGS cells caused by PMS2L2
overexpression. *P<0.05, as indicated. PMS2L2, PMS1 homolog 2
mismatch repair system component pseudogene 2; miR, microRNA; C,
control; NC, negative control. |
Discussion
The present study investigated the role of PMS2L2 in
GA. PMS2L2 was downregulated in GA tissues compared with healthy
adjacent tissues and had prognostic value. Moreover, the results
suggested that PMS2L2 may inhibit the migration and invasion of GA
cells by downregulating miR-25, which is an oncogenic miRNA in this
disease (12).
The prognosis and outlook of patients with GA is
worse in China compared with developed countries (15). It has been revealed that lncRNAs
serve important roles in GA (16).
For example, lncRNA ncRuPAR inhibited protease-activated receptor-1
to suppress the development of GA (16). Therefore, the altered expression of
lncRNAs may be associated with the survival time of patients with
GA. The present study revealed that PMS2L2 expression was
downregulated in GA tissues compared with healthy adjacent tissues.
Interestingly, the level PMS2L2 expression in GA tissues was not
significantly affected by the clinical stage of GA. Therefore,
PMS2L2 is likely to be involved in the overall process of GA rather
than specific stages. Furthermore, low expression of PMS2L2 in GA
was closely associated with poor survival rate. Therefore, PMS2L2
expression may be used to predict the survival of patients with GA
and aid the development of individualized treatment.
Preliminary deep sequencing data revealed that
PMS2L2 and miR-25 were inversely associated in GA tissues (data not
shown). miR-25 is a well-characterized oncogenic microRNA and
participates in several aspects of cancer development, including
the regulation of cancer cell behavior and the responses of cancer
cells to chemotherapy (17–19). The results of the present study
suggested that PMS2L2 is likely to be an upstream inhibitor of
miR-25 in GA cells, and the modulation of miR-25 by PMS2L2 is
involved in the regulation of migration and invasion of GA cells
but not normal gastric tissues cells. Therefore, PMS2L2 may serve
as a potential therapeutic target for GA. It is known that lncRNAs
can inhibit the function of miRNAs by serving as sponges (20,21). In
effect, PMS2L2 sponges miR-203 and serves a protective role in
chondrocytes protect against apoptosis (11). However, PMS2L2 does not sponge miR-25
as an appropriate binding site has not been identified (data not
shown). In the present study, PMS2L2 and miR-25 were not
significantly associated in healthy adjacent tissues. In addition,
PMS2L2 overexpression did not affect miR-25 expression in normal
gastric cancer cells. Therefore, the interaction between PMS2L2 and
miR-25 is likely indirect or GA-specific. Future studies are
required to identify the disease-related factors that mediate the
interaction between PMS2L2 and miR-25.
In conclusion, the present study revealed that
PMS2L2 was downregulated in GA and PMS2L2 may downregulate miR-25
to inhibit the migration and invasion of GA cells.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
JB and BL designed the experiments. JB and GL
performed experiments. ZZ collected and analyzed data. BL drafted
the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of The Second Hospital of Shandong University and all
patients provided written informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Hartgrink HH, Jansen EPM, van Grieken NC
and van de Velde CJ: Gastric cancer. Lancet. 374:477–490. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Van Cutsem E, Sagaert X, Topal B,
Haustermans K and Prenen H: Gastric cancer. Lancet. 388:2654–2664.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Fang X, Wei J, He X, An P, Wang H, Jiang
L, Shao D, Liang H, Li Y, Wang F and Min J: Landscape of dietary
factors associated with risk of gastric cancer: A systematic review
and dose-response meta-analysis of prospective cohort studies. Eur
J Cancer. 51:2820–2832. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Praud D, Rota M, Pelucchi C, Bertuccio P,
Rosso T, Galeone C, Zhang ZF, Matsuo K, Ito H, Hu J, et al:
Cigarette smoking and gastric cancer in the Stomach Cancer Pooling
(StoP) Project. Eur J Cancer Prev. 27:124–133. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ma K, Baloch Z, He TT and Xia X: Alcohol
consumption and gastric cancer risk: A meta-analysis. Med Sci
Monit. 23:238–246. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mocellin S, Verdi D, Pooley KA and Nitti
D: Genetic variation and gastric cancer risk: A field synopsis and
meta-analysis. Gut. 64:1209–1219. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Engreitz JM, Ollikainen N and Guttman M:
Long non-coding RNAs: Spatial amplifiers that control nuclear
structure and gene expression. Nat Rev Mol Cell Biol. 17:756–770.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Fatica A and Bozzoni I: Long non-coding
RNAs: New players in cell differentiation and development. Nat Rev
Genet. 15:7–21. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shi X, Sun M, Liu H, Yao Y and Song Y:
Long non-coding RNAs: A new frontier in the study of human
diseases. Cancer Lett. 339:159–166. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li J, Xuan Z and Liu C: Long non-coding
RNAs and complex human diseases. Int J Mol Sci. 14:18790–18808.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li X, Yu M, Chen L, Sun T, Wang H, Zhao L
and Zhao Q: LncRNA PMS2L2 protects ATDC5 chondrocytes against
lipopolysaccharide-induced inflammatory injury by sponging miR-203.
Life Sci. 217:283–292. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Li BS, Zuo QF, Zhao YL, Xiao B, Zhuang Y,
Mao XH, Wu C, Yang SM, Zeng H, Zou QM and Guo G: MicroRNA-25
promotes gastric cancer migration, invasion and proliferation by
directly targeting transducer of ERBB2, 1 and correlates with poor
survival. Oncogene. 34:2556–2665. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yuan SQ, Chen YT and Huang ZP: Equipping
the 8th edition American Joint Committee on Cancer Staging for
Gastric Cancer with the 15-node minimum: A population-based study
using recursive partitioning analysis. J Gastrointest Surg. 21:1–8.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Strong VE, Wu A, Selby LV, Gonen M, Hsu M,
Song KY, Park CH, Coit DG, Ji JF and Brennan MF: Differences in
gastric cancer survival between the US and China. J Surg Oncol.
112:31–37. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Liu L, Yan B, Yang Z, Zhang X, Gu Q and
Yue X: ncRuPAR inhibits gastric cancer progression by
down-regulating protease-activated receptor-1. Tumour Biol.
35:7821–7829. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Deng T, Yuan Y, Zhang C, Zhang C, Yao W,
Wang C, Liu R and Ba Y: Identification of circulating miR-25 as a
potential biomarker for pancreatic cancer diagnosis. Cell Physiol
Biochem. 39:1716–1722. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Feng X, Jiang J, Shi S, Xie H, Zhou L and
Zheng S: Knockdown of miR-25 increases the sensitivity of liver
cancer stem cells to TRAIL-induced apoptosis via PTEN/PI3K/Akt/Bad
signaling pathway. Int J Oncol. 49:2600–2610. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wu T, Chen W, Kong D, Li X, Lu H, Liu S,
Wang J, Du L, Kong Q, Huang X and Lu Z: miR-25 targets the
modulator of apoptosis 1 gene in lung cancer. Carcinogenesis.
36:925–935. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Liang WC, Fu WM, Wong CW, Wang Y, Wang WM,
Hu GX, Zhang L, Xiao LJ, Wan DC, Zhang JF and Waye MM: The lncRNA
H19 promotes epithelial to mesenchymal transition by functioning as
miRNA sponges in colorectal cancer. Oncotarget. 6:22513–22525.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wu XS, Wang F, Li HF, Hu YP, Jiang L,
Zhang F, Li ML, Wang XA, Jin YP, Zhang YJ and Lu W: LncRNA-PAGBC
acts as a microRNA sponge and promotes gallbladder tumorigenesis.
EMBO Rep. 18:1837–1853. 2017. View Article : Google Scholar : PubMed/NCBI
|