Introduction
Estrogen receptor (ER), which is encoded in the
estrogen receptor 1 (ESR1) gene, belongs to the nuclear
hormone receptor family (1) and the
ESR1 gene is located on chromosome 6 (6q25.1) and includes 8
exons (2). ER is expressed in over
60% of breast cancers (3) and
consists of two activation function domains, AF1/2, a DNA binding
domain and a hinge domain, and a ligand-binding domain (LBD)
(4). ER functions as a
ligand-dependent transcription factor, and ligand binding to the
LBD leads to activation of gene transcription, resulting in breast
cancer progression (5,6). Adjuvant endocrine therapy (ET)
inhibiting ER-induced breast cancer progression reduces local
recurrence and mortality in patients with ER-positive early breast
cancer (3,7). Similarly, ET with aromatase inhibitors
(AIs), selective estrogen receptor modulators (SERMs) and selective
estrogen receptor degrader (SERD) serves an important role in the
treatment of patients with ER-positive metastatic breast cancer
(MBC) (1). However, a number of
patients with ER-positive MBC have intrinsic ET resistance or
acquire resistance following response to ET, and eventually almost
all patients with MBC develop resistance to ET (8). ET resistance mechanisms include
upregulation of the steroid receptor coactivator-3, human epidermal
growth factor 2 (HER2) or nuclear factor κB and activator protein 1
(9–11). Cyclin D1 gene amplification
frequently occurs in ER-positive breast cancer and overexpression
of Cyclin D1 leads to ET resistance in ER-positive breast cancer
(12,13). A number of previous studies indicated
that LBD mutations (i.e., Tyr537Ser and Asp538Gly) in ESR1
gene were induced by long-term ET and resulted in acquired ET
resistance independently of estrogen levels, as well as poor
outcomes in patients with MBC (14–18).
Knowledge of the associations between the development of
ESR1 mutation and patient clinicopathological
characteristics may guide the decision-making process of MBC
treatment, including ET. The aim of the present study was to
evaluate the association between the development of ESR1
mutation and the clinicopathologic characteristics of patients with
MBC.
Materials and methods
Clinical samples
Patients with MBC who had received treatment for
primary ER-positive breast cancer and were followed-up at Keio
University Hospital between January 2012 and December 2015 were
enrolled in this study; during that period, 24 biopsy samples from
the metastatic sites were available for analysis. The inclusion and
exclusion criteria were as follows: Metastatic samples with ≥100×
sequence coverage on next generation sequence (NGS) were included,
while samples <100× sequence coverage were excluded from the
present study. Metastatic samples were evaluated by NGS, with a
level of sequence coverage of >100-fold, to detect the LBD
mutations of the ESR1 gene and validate these ER mutations
in patients with MBC. The 24 metastatic site samples were assessed,
and two were excluded from NGS due to low sequence coverage. Thus,
a total of 22 metastatic samples were evaluated in the present
study. The Allred score was used to assess the receptor status at
metastatic sites (19). The clinical
data, including the ET administered and clinical outcomes, were
obtained from all patients. The study protocol and the opt-out
informed consent procedure were approved by the Ethics Review Board
of Keio University Hospital and conformed to the Declaration of
Helsinki. Informed consent was acquired from the patients by
opt-out procedure prior to the beginning of the study.
DNA extraction
Using the biopsy samples from the MBC patients,
unstained 10-µm thick formalin-fixed paraffin embedded (FFPE)
blocks were obtained. Tumor tissue was collected from the two
blocks and placed in 1.5 ml microtubes. Genomic DNA was extracted
using a NucleoSpin DNA FFPE XS isolation kit (Takara Bio, Inc.).
DNA was quantified by Qubit 3.0 Fluorometer using the Qubit dsDNA
HS Assay kit (Thermo Fisher Scientific, Inc.), according to the
manufacturer's instructions, and adjusted to a final concentration
of 20 ng/µl.
Target amplification and
sequencing
For target amplification of the mutation hotspot in
the LBD of the ESR1 gene, PCR of the extracted genomic DNA
was performed using custom primers and ligating Illumina read1 and
read2 sequences (Illumina, Inc.). The second PCR primer pairs were
used to ligate the Illumina adaptor and index sequence to the first
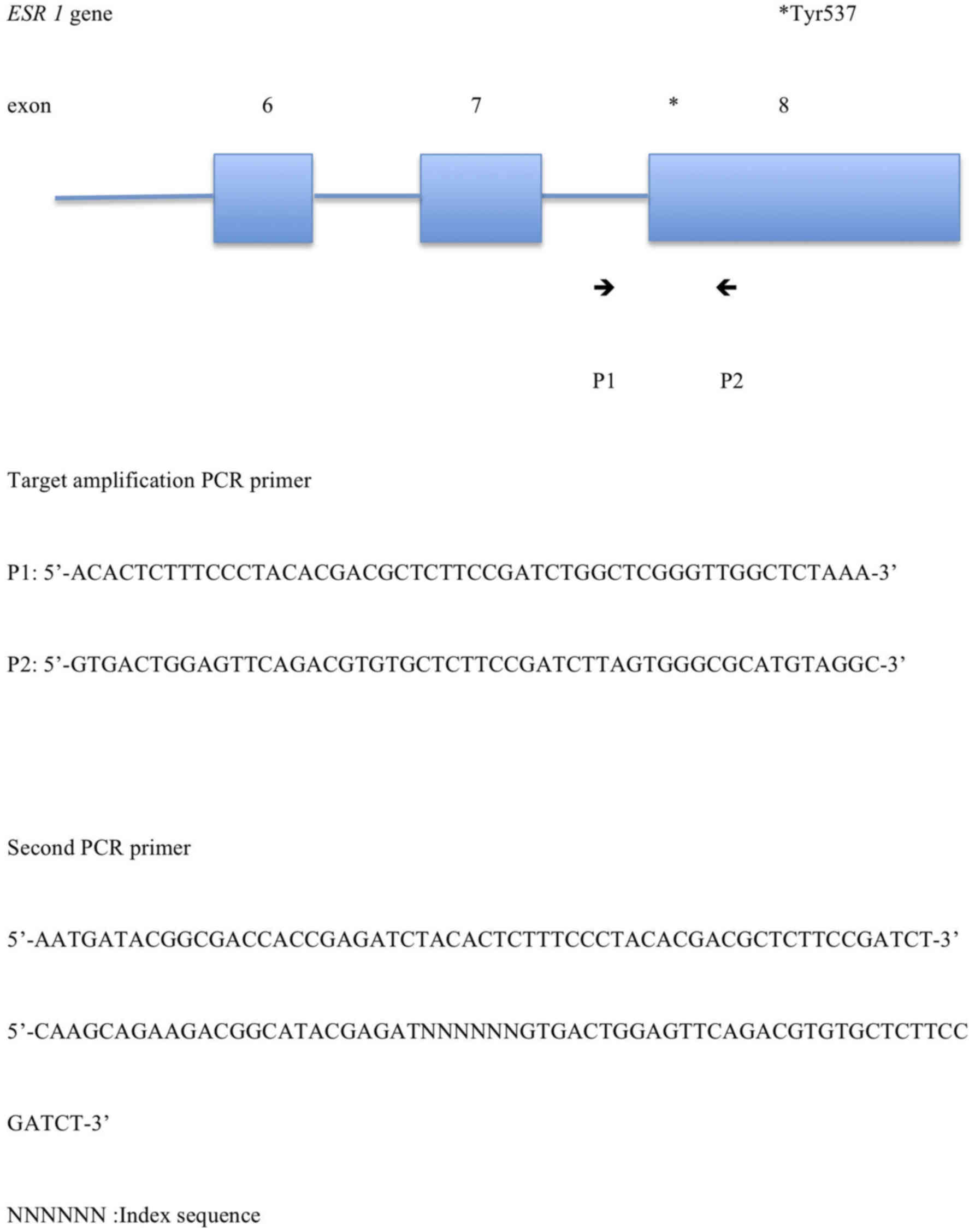
PCR products. The target amplification PCR primer sequences were as
follows: Forward,
5′-ACACTCTTTCCCTACACGACGCTCTTCCGATCTGGCTCGGGTTGGCTCTAAA-3′ and
reverse,
5′-GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTTAGTGGGCGCATGTAGGC-3′. The
second PCR primer sequences were as follows: Forward,
5′-AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT-3′
and reverse,
5′-CAAGCAGAAGACGGCATACGAGATNNNNNNGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT-3′
(NNN NNN: Index sequence). These sequences refer to ESR1.
Fig. 1 demonstrates the target
amplification PCR primer pairs and the second PCR primer pairs. In
a 10-µl reaction buffer that contained 5 mM deoxynucleotide
triphosphate mix, 0.25 µM of each custom made primer and 0.2 µl
Herculase II Fusion DNA polymerase (Agilent Technologies, Inc.), 20
ng of the genomic DNA underwent amplification for 20 cycles of 10
sec at 98°C, 30 sec at 55°C and 30 sec at 72°C. The amplicon, which
was 1 µl of the PCR product diluted 10 times, was marked in a
second PCR with molecular indices for Illumina Miseq, using TrueSeq
DNA HT Sample Kits (Illumina, Inc.). The second amplification was
performed as aforementioned.
DNA libraries were formed from the second PCR
products, which were purified using the Agencourt AMPure XP
reagent, according to the manufacturer's instructions (Beckman
Coulter, Inc.) and quantified by the method described above. The
library was sequenced on the Miseq instrument on the paired-end
mode with the Miseq Reagent kit v3 (Illumina, Inc.) according to
the manufacturer's instructions. The sequence data were mapped to
the reference human genome (hg19) using BWA aligner [version
0.7.16a-r1181; (20)], SAMtools
[version 1.6; (21)] and Picard
(http://sourceforge.net/projects/picard). Local
alignment and quality score calibration were performed according to
the Genome Analysis Toolkit (GATK) best practice (22). Single nucleotide variants were called
using the ‘HaplotypeCaller’ tool in GATK. All variants were
annotated using snpEff (23) and
reviewed by the Integrative Genomics Viewer (24). Variants were filtered using dbSNP_138
(http://www.ncbi.nlm.nih.gov/projects/SNP). GRCh37.75
in the Ensembl genome browser (http://www.ensembl.org) was used as the reference
genome of annotation.
Statistical analysis
Statistical analyses were performed with R commander
(version 2.4-1) based on R (version 3.3.3; http://cran.r-project.org/) and with EZR, which is a
modified version of the R commander (25). Among the continuous variables, age
followed a normal distribution as determined by the Shapiro-Wilk
test; thus, it was presented as the mean ± standard deviation and
analyzed using the independent two-sample Student's t-test. The
other continuous variables were expressed as the median (range) and
evaluated by the Wilcoxon rank sum test. Categorical variables were
analyzed by the Fisher's exact test. The correlation between two
continuous variables was evaluated by calculating Spearman's rank
correlation coefficient. P<0.05 was considered to indicate a
statistically significant difference.
Results
Baseline characteristics and detection
of ESR1 mutations in patients with MBC
The baseline characteristics of the 22 patients with
MBC are presented in Table I. The
mean age of the patients at the start of treatment was 54 years.
The TNM stage of primary breast cancer was I in 7 (32%) patients,
IIA in 9 (41%) patients, IIB in 4 (18) patients, IIIB in 1 (5%) patient and IV
in 1 (5%) patient. The histologic type of primary lesion was
invasive ductal carcinoma in 21 (95%) patients and invasive lobular
carcinoma in 1 (5%) patient. The Progesterone receptor (PgR) and
HER2 status of the primary tumor was positive in 21 (95%) and four
(18%) patients, respectively; HER2 status was unknown in one
case.
 | Table I.The baseline characteristics of 22
patients with metastatic breast cancer. |
Table I.
The baseline characteristics of 22
patients with metastatic breast cancer.
| Variable | Number of patients
(%) |
|---|
| Age at the start of
treatment, years | 54±13 |
| TNM stage of
primary breast cancer |
|
| I | 7 (32) |
|
IIA | 9 (41) |
|
IIB | 4 (18) |
|
IIIA | 0 (0) |
|
IIIB | 1 (5) |
|
IIIC | 0 (0) |
| IV | 1 (5) |
| Histological
type |
|
|
Invasive ductal | 21 (95) |
|
Invasive lobular | 1 (5) |
| PgR status of the
primary tumor |
|
|
Positive | 21 (95) |
|
Negative | 1 (5) |
| HER2 status of the
primary tumor |
|
|
Positive | 4 (18) |
|
Negative | 17 (77) |
|
Unknown | 1 (5) |
As presented in Table
II, ESR1 mutation at the metastatic site was observed in
14 (64%) of the 22 patients. A total of four ESR1 mutations
were identified including 1610A>C, 1626G>T, 1607T>G and
1642C>T, which led to amino acid mutations Tyr537Ser in 10 (45%)
patients, Glu542Asp in 2 (9%) patients, Leu536Arg in 1 (5%) patient
and Arg548Cys in 1 (5%) patient, respectively. The range of allele
mutation frequency was 2–99%.
 | Table II.Cases of metastatic breast cancer
with estrogen receptor 1 mutations (n=14). |
Table II.
Cases of metastatic breast cancer
with estrogen receptor 1 mutations (n=14).
| Case no. | Mutation | Amino acid
change | Mutation frequency,
% |
|---|
| 1 | 1610A>C | Tyr537Ser | 96 |
| 2 | 1610A>C | Tyr537Ser | 99 |
| 3 | 1610A>C | Tyr537Ser | 79 |
| 4 | 1610A>C | Tyr537Ser | 15 |
| 5 | 1610A>C | Tyr537Ser | 87 |
| 6 | 1610A>C | Tyr537Ser | 22 |
| 7 | 1610A>C | Tyr537Ser | 36 |
| 8 | 1642C>T | Arg548Cys | 2 |
| 9 | 1610A>C | Tyr537Ser | 18 |
| 10 | 1610A>C | Tyr537Ser | 58 |
| 11 | 1610A>C | Tyr537Ser | 21 |
| 12 | 1607T>G | Leu536Arg | 68 |
| 13 | 1626G>T | Glu542Asp | 15 |
| 14 | 1626G>T | Glu542Asp | 20 |
ESR1 mutation and the clinical
characteristics of patients with MBC
The ER Allred score of the primary tumor was
positive by immunohistochemical staining in all 22 patients prior
to this study (19). Of the 22
samples, 21 and 17 were positive for ER and PgR, respectively
(unavailable in 1 sample). The metastatic biopsy site was the liver
in 7 (32%) patients, skin in 6 (27%) patients, lymph node in 4
(18%) patients, lung in 3 (14%) patients, bone in 1 (5%) patient
and muscle in 1 (5%) patient.
Among the 22 patients with MBC, including one with
stage IV cancer, 19 patients received at least one ET agent prior
to the metastatic site biopsy; ESR1 mutation was detected in
13 patients, but it was not detected in 6 patients. Of the 21
patients with MBC (excluding the patient with stage IV MBC), 16
patients received ET in an adjuvant setting, which was complete in
10 cases and incomplete in 6 cases. An incomplete adjuvant setting
was defined as recurrence of breast cancer within five years of
treatment. Among the 16 patients who received ET in the adjuvant
setting, ESR1 mutation was detected in 2 of 6 patients who
received AIs, in 8 of 9 patients who received SERMs and in 1
patient who received both AI and SERM. Between the recurrence and
the biopsy of the metastatic site, 7 of 8 patients who received AIs
developed ESR1 mutation. ESR1 mutation was identified
in all 5 patients who received SERM in the adjuvant setting
followed by AI in the metastatic setting, as well as in 2 of 3
patients who did not receive SERM in adjuvant setting followed by
AI in the metastatic setting. In addition, 7 of 8 patients who had
no ESR1 mutation did not receive any ET for metastasis. The
patient with stage IV MBC, who was administered AIs until the
biopsy of the metastatic site, developed ESR1 mutation.
These results are presented in Table
III.
 | Table III.ESR1 mutations and the
clinical data of 22 patients with metastatic breast cancer. |
Table III.
ESR1 mutations and the
clinical data of 22 patients with metastatic breast cancer.
| Case no. | Age,
yearsa | Mutations | Allred of
E/Pb | Biopsy site | Adjuvant ET
(months, status) | ET after recurrence
to biopsy |
|---|
| 1 | 74 | Yes | 7/0 | Skin | AI (21,
incomplete) | None |
| 2 | 48 | Yes | 8/8 | Liver | SERM (60,
complete) | AI, SERD, AI |
| 3 | 51 | Yes | 7/7 | Skin | AI (7), SERM (53,
complete) | None |
| 4 | 35 | Yes | 8/8 | LN | None | None |
| 5 | 30 | Yes | 7/5 | LN | None | AI, SERM, AI,
SERD |
| 6 | 56 | Yes | Unavailable | Bone | AI (60,
complete) | AI |
| 7 | 54 | Yes | 8/7 | Skin | SERM (60,
complete) | None |
| 8 | 47 | Yes | 7/0 | Liver | SERM (22,
incomplete) | AI, SERD |
| 9 | 54 | Yes | 8/6 | Skin | SERM (60,
complete) | AI |
| 10 | 39 | Yes | 8/8 | Liver | SERM (24,
incomplete) | AI, AI |
| 11 | 51 | Yes | 8/8 | Liver | SERM (24,
complete) | None |
| 12 | 57 | Yes | 8/7 | Liver | SERM (60,
complete) | AI |
| 13 | 42 | Yes | 8/5 | Skin | SERM (37,
incomplete) | None |
| 14 | 46 | Yes | 8/8 | Liver | None | AI, AI |
| 15 | 74 | No | 8/8 | Skin | AI (60,
complete) | None |
| 16 | 75 | No | 8/8 | Muscle | None | None |
| 17 | 58 | No | 8/6 | LN | AI (60,
complete) | None |
| 18 | 39 | No | 8/0 | LN | SERM (23,
incomplete) | None |
| 19 | 59 | No | 8/8 | Lung | None | AI, SERM |
| 20 | 63 | No | 7/7 | Lung | AI (58,
incomplete) | None |
| 21 | 60 | No | 8/0 | Liver | AI (60,
complete) | None |
| 22 | 74 | No | 8/5 | Lung | None | None |
Association between ESR1 mutation and
clinicopathological characteristics in 22 patients with MBC
Considering the total period between the beginning
of treatment and the biopsy of the metastatic site in 22 patients
with MBC, the SERM intake period was significantly longer in
patients with ESR1 mutation compared with that in patients
with wild-type ESR1 (26 vs. 0 months; P=0.01). The total
interval of treatment with AI and AI/ SERM was not significantly
different between patients with ESR1 mutation and those
without ESR1 mutation (P=0.92 and P=0.13, respectively;
Table IV).
 | Table IV.Association between estrogen receptor
1 gene mutation and clinicopathologic data in 22 patients with
metastatic breast cancer. |
Table IV.
Association between estrogen receptor
1 gene mutation and clinicopathologic data in 22 patients with
metastatic breast cancer.
| A, According to the
total duration from the beginning of treatment to biopsy of the
metastatic site (n=22) |
|---|
|
|---|
| Variable | Mutation (+) | Mutation (−) | P-value |
|---|
| AI | 15 (0–83) | 31 (0–60) | 0.92 |
| SERM | 26 (0–60) | 0 (0–23) | 0.01 |
| AI + SERM | 60 (0–143) | 41 (0–60) | 0.13 |
|
| B, According to
the total duration from recurrence to biopsy of the metastatic site
(n=21) |
|
|
Variable | Mutation
(+) | Mutation
(−) | P-value |
|
| AI | 5 (0–83) | 0 (0–3) | 0.04 |
| SERM | 0 (0–27) | 0 (0–7) | 0.83 |
| AI + SERM | 5 (0–83) | 0 (0–10) | 0.05 |
|
| C, According to
clinicopathological characteristics (n=22) |
|
|
Variable | Mutation
(+) | Mutation
(−) | P-value |
|
| Age at the start of
treatment, years | 49±11 | 63±12 | 0.01 |
| Primary TNM
stage |
|
|
|
| I | 4 | 3 |
|
|
IIA | 5 | 4 |
|
|
IIB | 3 | 1 |
|
|
IIIA | 0 | 0 |
|
|
IIIB | 1 | 0 |
|
|
IIIC | 0 | 0 |
|
| IV | 1 | 0 |
|
| Histological
type |
|
|
|
|
Invasive ductal | 13 | 8 |
|
|
Invasive lobular | 1 | 0 | >0.99 |
| PgR status of the
primary tumor |
|
|
|
|
Positive | 13 | 8 |
|
|
Negative | 1 | 0 | >0.99 |
| HER2 status of the
primary tumor |
|
|
|
|
Positive | 2 | 2 |
|
|
Negative | 11 | 6 | 0.62 |
| Total number of
administered ETa | 1.5 (0–3) | 1 (0–1) | 0.06 |
| The number of ET
from recurrence to biopsy | 1 (0–3) | 0 (0–2) | 0.10 |
| Spearman's rank
correlation | Coefficient |
|
| Age at the start of
treatment vs. | −0.45 | 0.03 |
| The total duration
of SERM until biopsy of the metastatic site |
|
|
|
Considering the treatment period from recurrence to
biopsy of the metastatic site in 21 patients with MBC, after
excluding the stage IV case, the AI intake period was significantly
longer in patients with ESR1 mutation than in those without
ESR1 mutation (5 vs. 0 months; P=0.04; Table IV). However, there were no
significant differences in the treatment period with SERM or AI/
SERM between patients with and without ESR1 mutation (P=0.83
and P=0.05, respectively).
The age at the time of treatment initiation was
significantly lower in patients with mutant ESR1 compared
with that in patients without mutant ESR1 (49±11 vs. 63±12
years; P=0.01; Table IV). There
were no significant differences in the primary TNM stage,
histologic type, PgR and HER2 status of primary tumor between the
two patient groups. The number of administered ETs tended to be
higher in patients with ESR1 mutation compared with that in
those without ESR1 mutation, but the difference was not
significant (1.5 vs. 0, respectively; P=0.06; Table IV). The number of ETs administered
to the patients from recurrence to biopsy was not significantly
different between the two patient groups (1 vs. 0; P=0.10). The age
at the time of the initiating treatment was associated with the
SERM intake period (Spearman's rank correlation coefficient, −0.45;
P=0.03; Table IV).
Discussion
In the present study on a cohort of patients with
MBC, the presence of LBD mutations in the ESR1 gene was
detected by targeted NGS, and its association with patient
clinicopathologic characteristics was assessed. The frequency of
ESR1 mutations in metastatic samples was 64% in this study
and varied between 13 and 55% among published studies that also
used NGS (14–16,26).
Compared with previous studies, the present study had a higher
frequency of ESR1 mutations and a lower number of
administered ETs. This study demonstrated that there was no
association between the number of administered ET and the
development of ESR1 mutation. Two previous studies have
demonstrated an association between ET exposure and the prevalence
of mutated ESR1 (26,27), although the number of previously
administered ET was not clarified.
In the current study, the ESR1 mutations
Leu536Arg, Tyr537Ser, Glu542Asp and Arg548Cys were detected. The
Tyr537Ser and Arg548Cys mutations have been demonstrated to induce
estrogen-independent activity of the ER, leading to ET resistance
(15,28). In addition, bioinformatics analysis
has indicated that an Arg548Cys mutation in the ER is deleterious
(29). Similarly, amino acid
mutations in Leu536 have been reported to increase the
estrogen-independent activity of the ER (30), and Toy et al (15) identified the Leu536Arg mutation in
the ER, but they did not investigate its function. The impact of
the Glu542Asp alteration on the ER function remains unknown and
further investigation is needed (31). Therefore, the detected ER mutations
in this study, with the exception of Glu542Asp, may induce a
ligand-independent ER activation resulting in ET resistance. It is
important to determine the changes of ESR1 alterations
between primary and metastatic tumor; however, in this study, the
ESR1 mutations of the primary lesion were not investigated
because primary breast cancers have very rare ESR1 mutation,
which has been reported in previous studies, including The Cancer
Atlas data (32–34).
A number of previous studies on patients with MBC
have demonstrated that compared with wild-type ESR1, ESR1
mutation led to worse progression-free survival (PFS) and overall
survival (OS) (35,36). However, in the present study,
ESR1 mutation had no adverse impact on the outcomes of
patients with MBC (data not shown). The small cohort used in the
present study limited the statistical power to assess the impact on
the outcomes of patients with MBC.
This may be due to the Glu542Asp mutation, which was
detected in 2 patients with MBC in this study, having no negative
effect on the patients' outcomes. The present study revealed that
prolonged AI treatment for metastasis had a significant impact on
the development of ESR1 mutation and that patients with MBC
who received AIs in an adjuvant setting exhibited low rates of
ESR1 mutation. A number of previous studies reported that
mutated ESR1 rarely occurred during adjuvant therapy with
AI, but its prevalence was high during recurrence treatment with AI
(17,35). The results of the present study
appeared to support these studies. Previous studies have also
demonstrated the superior effects of fulvestrant on the PFS and OS
compared with those of anastrozole in patients with
endocrine-sensitive MBC (37,38).
Furthermore, the addition of palbociclib, a CDK4/6 inhibitor, to
letrozole or fulvestrant improved the PFS in patients with MBC
(39,40). Of note, it was reported that
palbociclib combined with fulvestrant improved the PFS irrespective
of the ESR1 mutation status in patients with MBC (41) and that palbociclib plus letrozole did
not prevent the development of ESR1 mutation in a small
cohort of patients with MBC who received the combination treatment
(42). These results suggest that
the assessment of dynamic changes of the ESR1 mutation
status using minimally invasive procedures such as liquid biopsy in
patients with MBC who receive CDK4/6 inhibitors may be of
importance for investigating acquired resistance to these
drugs.
The results of the present study demonstrated that
the total period of SERM treatment was associated with the
emergence of ESR1 mutation. To the best of our knowledge,
the effects of SERM on ESR1 mutation have not been fully
clarified to date. Among patients with MBC who received tamoxifen
alone, ESR1 mutation was not detected in all 22 patients in
a study by Schiavon et al (17), but it was detected in 4 out of 11
patients in a study by Takeshita et al (43). The present study demonstrated that
the frequency rate of ESR1 alteration in patients who
received adjuvant-SERM followed by metastatic AI treatment was
higher compared with that in patients who received no adjuvant-SERM
followed by metastatic AI. The association of SERM with ESR1
mutation that was identified in this study may be explained by the
finding that most of the patients with mutated ESR who
received AIs for metastasis had been administered SERMs in an
adjuvant setting. Administration of SERM in an adjuvant setting may
have been used as a compelling indication for the use of AI for
metastasis; therefore, SERM for adjuvant setting, followed by AI
for metastasis, may increase the frequency of ESR1
mutations. However, this result needs to be clarified and verified
in a future study.
The present study revealed that age at the time of
treatment initiation for breast cancer was significantly associated
with the development of ESR1 mutation and the total duration
of SERM treatment. These associations may be due to the
premenopausal status of the majority of patients with MBC who
received SERM in an adjuvant setting and AI in a metastatic
setting, leading to the subsequent development of ESR1
mutation. To the best of our knowledge, no previous studies have
demonstrated an association between age at the time of treatment
initiation for breast cancer and the occurrence of ESR1 gene
mutation.
This study had several limitations, including the
retrospective design using a small cohort from a single institute.
In addition, the effects of SERD on ESR1 mutations were not
analyzed, as only three patients with MBC received SERD, and no
multivariate analysis was performed due to the small cohort. The
small cohort of this study limited the statistical power to assess
the association between ESR1 mutation and
clinicopathological features in patients with MBC.
In conclusion, the results of the present study
demonstrated that SERM in an adjuvant setting followed by AI for
metastasis may increase the frequency of ESR1 mutation, and
that age at the time of treatment onset for breast cancer may be
significantly associated with the development of ESR1
mutation. Further studies are needed to confirm and validate these
findings.
Acknowledgements
The authors would like to thank Mr. Kazuhiro Miyao
at the Department of Surgery, Keio University School of Medicine
(Tokyo, Japan) for his excellent technical assistance and helpful
insights on DNA extraction and next generation sequencing.
Funding
The present study was supported by a grant-in-aid
from the Japan Society for the Promotion Science (grant no.
16K10474 to MT).
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
KT, TH, TS, MT and YK conceived and designed this
study. HM, AN and TY performed the experiments and collected
clinical data from the patients. KT analyzed data and drafted the
manuscript. All authors revised the manuscript and approved the
final version.
Ethics approval and consent to
participate
The study protocol, including the opt-out informed
consent procedure, was approved by the Ethics Review Board of Keio
University Hospital (approval no. 20150439) and conformed to
Declaration of Helsinki.
Patient consent for publication
Not applicable.
Competing interests
TH and YK received lecture fee and research funding
from Pfizer Inc., Novartis Pharma K.K., and AstraZeneca. All other
authors confirm that they have no competing interests.
Glossary
Abbreviations
Abbreviations:
|
AI
|
aromatase inhibitor
|
|
ER
|
estrogen receptor
|
|
ET
|
endocrine therapy
|
|
LBD
|
ligand-binding domain
|
|
MBC
|
metastatic breast cancer
|
|
NGS
|
next generation sequencing
|
|
OS
|
overall survival
|
|
PFS
|
progression-free survival
|
|
PgR
|
progesterone receptor
|
|
SERD
|
selective estrogen receptor
degrader
|
|
SERM
|
selective estrogen receptor
modulator
|
References
|
1
|
Olefsky JM: Nuclear receptor minireview
series. J Biol Chem. 276:36863–36864. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Gianluca A, Flavia P and Paolo M: Update
on Mechanisms of Hormone Action: Focus on Metabolism, Growth and
Reproduction. Tech; Rijeka, Croatia: pp. 71–73. 2011
|
|
3
|
Tamoxifen for early breast cancer, . An
overview of the randomised trials. Early Breast Cancer Trialists'
Collaborative Group. Lancet. 351:1451–1467. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ruff M, Gangloff M, Wurtz JM and Moras D:
Estrogen receptor transcription and transactivation:
Structure-function relationship in DNA- and ligand-binding domains
of estrogen receptors. Breast Cancer Res. 2:353–359. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kojetin DJ, Burris TP, Jensen EV and Khan
SA: Implications of the binding of tamoxifen to the coactivator
recognition site of the estrogen receptor. Endocr Relat Cancer.
15:851–870. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Heldring N, Pike A, Andersson S, Matthews
J, Cheng G, Hartman J, Tujague M, Ström A, Treuter E, Warner M and
Gustafsson JA: Estrogen receptors: How do they signal and what are
their targets. Physiol Rev. 87:905–931. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Goldhirsch A, Glick JH, Gelber RD, Coates
AS and Senn HJ: Meeting highlights: International Consensus Panel
on the Treatment of Primary Breast Cancer. Seventh International
Conference on Adjuvant Therapy of Primary Breast Cancer. J Clin
Oncol. 19:3817–3827. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Pritchard KI: Endocrine therapy: Is the
first generation of targeted drugs the last? J Intern Med.
274:144–152. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Osborne CK, Bardou V, Hopp TA, Chamness
GC, Hilsenbeck SG, Fuqua SA, Wong J, Allred DC, Clark GM and Schiff
R: Role of the estrogen receptor coactivator AIB1 (SRC-3) and
HER-2/neu in tamoxifen resistance in breast cancer. J Natl Cancer
Inst. 95:353–361. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Schiff R, Massarweh SA, Shou J, Bharwani
L, Mohsin SK and Osborne CK: Cross-talk between estrogen receptor
and growth factor pathways as a molecular target for overcoming
endocrine resistance. Clin Cancer Res. 10 (Suppl):S331–S336. 2004.
View Article : Google Scholar
|
|
11
|
Zhou Y, Yau C, Gray JW, Chew K, Dairkee
SH, Moore DH, Eppenberger U, Eppenberger-Castori S and Benz CC:
Enhanced NF kappa B and AP-1 transcriptional activity associated
with antiestrogen resistant breast cancer. BMC Cancer. 7:592007.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Thangavel C, Dean JL, Ertel A, Knudsen KE,
Aldaz CM, Witkiewicz AK, Clarke R and Knudsen ES: Therapeutically
activating RB: Reestablishing cell cycle control in endocrine
therapy-resistant breast cancer. Endocr Relat Cancer. 18:333–345.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Witkiewicz AK and Knudsen ES:
Retinoblastoma tumor suppressor pathway in breast cancer:
Prognosis, precision medicine, and therapeutic interventions.
Breast Cancer Res. 16:2072014. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Robinson DR, Wu YM, Vats P, Su F, Lonigro
RJ, Cao X, Kalyana-Sundaram S, Wang R, Ning Y, Hodges L, et al:
Activating ESR1 mutations in hormone-resistant metastatic breast
cancer. Nat Genet. 45:1446–1451. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
15
|
Toy W, Shen Y, Won H, Green B, Sakr RA,
Will M, Li Z, Gala K, Fanning S, King TA, et al: ESR1
ligand-binding domain mutations in hormone-resistant breast cancer.
Nat Genet. 45:1439–1445. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Merenbakh-Lamin K, Ben-Baruch N, Yeheskel
A, Dvir A, Soussan-Gutman L, Jeselsohn R, Yelensky R, Brown M,
Miller VA, Sarid D, et al: D538G mutation in estrogen receptor-α: A
novel mechanism for acquired endocrine resistance in breast cancer.
Cancer Res. 73:6856–6864. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Schiavon G, Hrebien S, Garcia-Murillas I,
Cutts RJ, Pearson A, Tarazona N, Fenwick K, Kozarewa I,
Lopez-Knowles E, Ribas R, et al: Analysis of ESR1 mutation in
circulating tumor DNA demonstrates evolution during therapy for
metastatic breast cancer. Sci Transl Med. 7:313ra1822015.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Reinert T, Saad ED, Barrios CH and Bines
J: Clinical implications of ESR1 mutations in hormone
receptor-positive advanced breast cancer. Front Oncol. 7:262017.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Allred DC, Harvey JM, Berardo M and Clark
GM: Prognostic and predictive factors in breast cancer by
immunohistochemical analysis. Mod Pathol. 11:155–168.
1998.PubMed/NCBI
|
|
20
|
Li H and Durbin R: Fast and accurate short
read alignment with Burrows-Wheeler transform. Bioinformatics.
25:1754–1760. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Li H, Handsaker B, Wysoker A, Fennell T,
Ruan J, Homer N, Marth G, Abecasis G and Durbin R; 1000 Genome
Project Data Processing Subgroup, : The Sequence Alignment/Map
format and SAMtools. Bioinformatics. 25:2078–2079. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
DePristo MA, Banks E, Poplin R, Garimella
KV, Maguire JR, Hartl C, Philippakis AA, del Angel G, Rivas MA,
Hanna M, et al: A framework for variation discovery and genotyping
using next-generation DNA sequencing data. Nat Genet. 43:491–498.
2011. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Cingolani P, Platts A, Wang le L, Coon M,
Nguyen T, Wang L, Land SJ, Lu X and Ruden DM: A program for
annotating and predicting the effects of single nucleotide
polymorphisms, SnpEff: SNPs in the genome of Drosophila
melanogaster strain w1118; iso-2; iso-3. Fly (Austin). 6:80–92.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Robinson JT, Thorvaldsdóttir H, Winckler
W, Guttman M, Lander ES, Getz G and Mesirov JP: Integrative
genomics viewer. Nat Biotechnol. 29:24–26. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kanda Y: Investigation of the freely
available easy-to-use software ‘EZR’ for medical statistics. Bone
Marrow Transplant. 48:452–458. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Jeselsohn R, Yelensky R, Buchwalter G,
Frampton G, Meric-Bernstam F, Gonzalez-Angulo AM, Ferrer-Lozano J,
Perez-Fidalgo JA, Cristofanilli M, Gómez H, et al: Emergence of
constitutively active estrogen receptor-α mutations in pretreated
advanced estrogen receptor-positive breast cancer. Clin Cancer Res.
20:1757–1767. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Takeshita T, Yamamoto Y, Yamamoto-Ibusuki
M, Tomiguchi M, Sueta A, Murakami K, Omoto Y and Iwase H:
Comparison of ESR1 mutations in tumor tissue and matched plasma
samples from metastatic breast cancer patients. Transl Oncol.
10:766–771. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Li B, Liu L, Fu X, Zhou WQ, Zou DT, Zhao
XY, Cai YN, Tu HB, Liu QC and Chen YY: A novel mutation of estrogen
receptor gene detected in girls with precocious puberty. Yi Chuan
Xue Bao. 32:1011–1017. 2005.PubMed/NCBI
|
|
29
|
Rebaï M and Rebaï A: In silico
characterization of functional SNP within the oestrogen receptor
gene. J Genet. 95:865–874. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zhao C, Koide A, Abrams J,
Deighton-Collins S, Martinez A, Schwartz JA, Koide S and Skafar DF:
Mutation of Leu-536 in human estrogen receptor-alpha alters the
coupling between ligand binding, transcription activation, and
receptor conformation. J Biol Chem. 278:27278–27286. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chung JH, Pavlick D, Hartmaier R, Schrock
AB, Young L, Forcier B, Ye P, Levin MK, Goldberg M, Burris H, et
al: Hybrid capture-based genomic profiling of circulating tumor DNA
from patients with estrogen receptor-positive metastatic breast
cancer. Ann Oncol. 28:2866–2873. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Karnik PS, Kulkarni S, Liu XP, Budd GT and
Bukowski RM: Estrogen receptor mutations in tamoxifen-resistant
breast cancer. Cancer Res. 54:349–353. 1994.PubMed/NCBI
|
|
33
|
Roodi N, Bailey LR, Kao WY, Verrier CS,
Yee CJ, Dupont WD and Parl FF: Estrogen receptor gene analysis in
estrogen receptor-positive and receptor-negative primary breast
cancer. J Natl Cancer Inst. 87:446–451. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Cancer Genome Atlas Network: Comprehensive
molecular portraits of human breast tumours. Nature. 490:61–70.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Chandarlapaty S, Chen D, He W, Sung P,
Samoila A, You D, Bhatt T, Patel P, Voi M, Gnant M, et al:
Prevalence of ESR1 mutations in cell-free DNA and outcomes in
metastatic breast cancer: A secondary analysis of the BOLERO-2
clinical trial. JAMA Oncol. 2:1310–1315. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Clatot F, Perdrix A, Augusto L, Beaussire
L, Delacour J, Calbrix C, Sefrioui D, Viailly PJ, Bubenheim M,
Moldovan C, et al: Kinetics, prognostic and predictive values of
ESR1 circulating mutations in metastatic breast cancer patients
progressing on aromatase inhibitor. Oncotarget. 7:74448–74459.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Robertson JFR, Bondarenko IM, Trishkina E,
Dvorkin M, Panasci L, Manikhas A, Shparyk Y, Cardona-Huerta S,
Cheung KL, Philco-Salas MJ, et al: Fulvestrant 500 mg versus
anastrozole 1 mg for hormone receptor-positive advanced breast
cancer (FALCON): An international, randomised, double-blind, phase
3 trial. Lancet. 388:2997–3005. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ellis MJ, Llombart-Cussac A, Feltl D,
Dewar JA, Jasiówka M, Hewson N, Rukazenkov Y and Robertson JF:
Fulvestrant 500 mg versus anastrozole 1 mg for the First-line
treatment of advanced breast cancer: Overall survival analysis from
the phase II FIRST study. J Clin Oncol. 33:3781–3787. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Finn RS, Crown JP, Lang I, Boer K,
Bondarenko IM, Kulyk SO, Ettl J, Patel R, Pinter T, Schmidt M, et
al: The cyclin-dependent kinase 4/6 inhibitor palbociclib in
combination with letrozole versus letrozole alone as first-line
treatment of oestrogen receptor-positive, HER2-negative, advanced
breast cancer (PALOMA-1/TRIO-18): A randomised phase 2 study.
Lancet Oncoly. 16:25–35. 2015. View Article : Google Scholar
|
|
40
|
Turner NC, Ro J, André F, Loi S, Verma S,
Iwata H, Harbeck N, Loibl S, Huang Bartlett C, Zhang K, et al:
Palbociclib in hormone-receptor-positive advanced breast cancer. N
Engl J Med. 373:209–219. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Fribbens C, O'Leary B, Kilburn L, Hrebien
S, Garcia-Murillas I, Beaney M, Cristofanilli M, Andre F, Loi S,
Loibl S, et al: Plasma ESR1 mutations and the treatment of estrogen
receptor-positive advanced breast cancer. J Clin Oncol.
34:2961–2968. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Gyanchandani R, Kota KJ, Jonnalagadda AR,
Minteer T, Knapick BA, Oesterreich S, Brufsky AM, Lee AV and
Puhalla SL: Detection of ESR1 mutations in circulating cell-free
DNA from patients with metastatic breast cancer treated with
palbociclib and letrozole. Oncotarget. 8:66901–66911. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Takeshita T, Yamamoto Y, Yamamoto-Ibusuki
M, Inao T, Sueta A, Fujiwara S, Omoto Y and Iwase H: Droplet
digital polymerase chain reaction assay for screening of ESR1
mutations in 325 breast cancer specimens. Transl Res.
166:540–553.e2. 2015. View Article : Google Scholar : PubMed/NCBI
|