Introduction
Globally, as of 2017, ~500,000 women were, and
~288,000 patients died from cervical cancer (1). In addition, there are ~135,000 new
cases in China each year, accounting for 1/3 of the global
incidence rate (2,3). Cervical cancer is highly malignant,
progresses quickly and has a poor prognosis. Furthermore, its
occurrence is affected by genetic and environmental factors
(4). The main treatment is
radiotherapy, chemotherapy and surgery; therefore, it is important
to develop novel drugs, as well as diagnostic and prognostic
markers (5).
Traditional Chinese Medicine (TCM) serves an
increasingly important role in the comprehensive treatment of
cervical cancer, and may be a new direction for the prevention and
treatment of this disease in the future (6). Qian et al (7) reported that Erhuang powder had
significant clinical effects in the treatment of high-risk human
papillomavirus (HPV) infection with mild cervical epithelial
neoplasia and moderate cervical epithelial neoplasia. At the same
time, the serum concentrations of IFN-γ and TNF-α in patients with
cervical cancer were increased, and IL-4 and IL-10 concentrations
were reduced. Furthermore, Chen et al (8) revealed that the Houttuynia
cordata decoction could effectively increase the HPV negative
rate of cervical intraepithelial neoplasia.
Sarcandra Herb (SH) is an integral component of
Castanea mollissima named grass coral in China, with an apparent
therapeutic effect (9), has become a
highly researched Chinese herbal medicine due to its significant
clinical effects, and its main method of use includes Zhong Jie
Feng injection and grass coral lozenges (10). Several monomers in SH, showing
significant research value, have been analyzed and screened using
network pharmacology. Researchers in China have previously reported
that the caffeic acid, 3,4-dihydroxyphenethyl ester monomer,
isolated and extracted from SH, could selectively activate the
phosphorylation of p38 and ERK1/2, in the MAPK signaling pathway,
in cervical cancer cells, and could promote apoptosis and autophagy
(11). In addition, Trusov et
al (12) examined the effect of
the natural flavonoid, quercetin (Que), found in SH, on tumor
growth inhibition and the prolongation of the survival time in C-57
mice. These authors identified that Que had an inhibitory rate of
20.7–60.1% on the solid tumors from the mouse liver cancer cell
line, S180, and a 36.8% survival time extension rate for the S180
ascites model.
The aim of the present study was to investigate the
antitumor activity of SH to identify the target that plays a role
in the biological functions of cervical cancer cells, and analyze
the positive and negative regulatory effects.
Materials and methods
Reagents and antibodies
Que was purchased from Sigma Aldrich (Merck KGaA).
The following primary antibodies were used in the present study:
phosphorylated (p)-EGFR (Tyr1068) (1:1,000; cat. no. 3777; Cell
Signaling Technology, Inc.), p-MEK (1:1,000; cat. no. 2338; Cell
Signaling Technology, Inc.), p-cell division cycle (Cdc) 42
(1:1,000; cat. no. 2461; Cell Signaling Technology, Inc.), EGFR
(1:1,000; cat. no. 4267; Cell Signaling Technology, Inc.), MEK
(1:1,000; cat. no. 9122; Cell Signaling Technology, Inc.), p-44/42
MAPK (ERK1/2) (1:1,000; cat. no. 4370; Cell Signaling Technology,
Inc.), caspase-3 (1:1,000; cat. no. 9664; Cell Signaling
Technology, Inc.), GAPDH (1:3,000; cat. no. 5174; Cell Signaling
Technology, Inc.), β-tubulin (1:3,000; cat. no. 10094-1-AP;
ProteinTech Group, Inc.), cdc42 (1:1,000; cat. no. 10155-1-AP;
ProteinTech Group, Inc.), Bax (1:1,000; cat. no. 50599-2-lg;
ProteinTech Group, Inc.) and Bcl-2 (1:1,000; cat. no. 12789-1-AP;
ProteinTech Group, Inc.). The secondary antibodies, HRP-conjugated
goat-anti-rabbit (cat. no. L3012) and goat-anti-mouse (cat. no.
L3032) IgG were purchased from Signalway Antibody LLC. The small
molecule inhibitors, aftinib, PD98059, U0126, ML141 for EGFR, MEK,
ERK and Cdc42 were purchased from MedChemExpress.
Cell culture
The human cervical cancer cell lines (HeLa and SiHa)
were purchased from Shanghai Zhongqiao Xinzhou Biological
Technology Co., Ltd., and were cultured in RPIM-1640 medium
(Biological Industries), supplemented with 10% FBS (Biological
Industries), 100 units of penicillin and 100 µg/ml streptomycin at
37°C in a humidified incubator with 5% CO2.
Public data collection
Prediction and screening of the anti-cervical cancer
effects and disease targets of SH ingredients from TCM systems
pharmacology database and analysis platform (TCMSP), GeneCards and
OMIM databases were conducted. The chemical compositions and genes
associated with SH in 3 separate databases were searched: i) TCMSP
(http://ibts.hkbu.edu.hk/LSP/tcmsp.php), oral
bioavailability was set to >30% and drug-likeness was set to
>0.18, to obtain the chemical composition of SH; ii) the
molecular structure was confirmed using PubChem (https://pubchem.ncbi.nlm.nih.gov/), the targets
associated with the effective components of SH could also be
exported in batches in TCMSP; and iii) the UniProt database
(http://www.uniprot.org/uniprot/) ID
number was downloaded to convert the entire documents using Perl
codes, then, the association between drug and disease was
evaluated.
Screening of key words (cervical cancer) was
performed to obtain related targets in the GeneCards (http://www.genecards.org/) and OMIM databases
(http://www.ncbi.nlm.nih.gov/omim) by
eliminating duplicate and pseudo-positive genes. These were then
matched with potential targets for the anti-cervical cancer effect
and the common targets were presented as a Venn diagram. The
Protein Analysis by Evolutionary Relationship database (PANTHER;
v11; http://pantherdb.org) contains
comprehensive information on the evolution and function of
protein-coding genes from 104 complete sequenced genomes. The
PANTHER software tool allows users to classify new protein
sequences and analyze gene lists obtained from large-scale genome
experiments. The Search Tool for the Retrieval of Interacting
Genes/Proteins database (https://string.db.org/) contains known and predicted
protein-protein interactions. Thus, the protein interactions in
tab-separated values format were obtained. Cytoscape3.7.1
(http://www.cytoscape.org/) software was
used to generate the Zhong Jie Feng-Target network diagram. Nodes-1
and −2, and the combined score information in the files were
inputted into Cytoscape to create the interaction network; the
style option from the statistics tool
(Cytoscape>Tools>Network>Analyzer>Analysis>Generate)
was used to apply the node size and color settings to reflect the
combination degree. R software (v.3.7.1; http://cran.r-project.org/bin/windows/base/)
(13) was used for bioinformatics
analysis.
Staining using
5-Ethynyl-2′-deoxyuridine (EdU)
The detailed steps of the experiment were conducted
according to the manufacturer's instructions (Guangzhou RiboBio
Co., Ltd.). Approximately, 10,000 cells/per well in logarithmic
growth phase were seeded in a 48-well plate, cultured to the normal
growth stage, then Que (40 µM) was added after 8 h. EdU solution
was diluted in complete medium at a ratio of 1,000:1 in 200 µl,
then added to each well and the cells were incubated for 2 h, then
washed with PBS twice, for 5 min each time. Subsequently, 100 µl
PBS, containing 4% paraformaldehyde, was added to each well, then
the cells were incubated at room temperature for 30 min and 200 µl
2 mg/ml glycine solution was added. The cells were incubated for 5
min on a shaker, following which 200 µl PBS was added to wash the
cells, then 200 µl 0.5% TritonX-100 was added to each well and
incubated at room temperature on a shaker for 10 min. Next, the
cells were washed with PBS once for 5 min, then 100 µl 1X Apollo
staining reaction solution was added to each well, and the cells
were incubated at room temperature for 30 min then the solution was
discarded. Subsequently, the cells were washed with 200 µl 0.5%
TritonX-100 twice for 10 min each time, then 200 µl 1X Hoechst
33342 reaction solution (diluted with deionized water at ratio
100:1) was added to each well and the cells were incubated at room
temperature for 30 min in a shaker. Lastly, the cells were washed
with 200 µl PBS twice and the images were captured using a
fluorescence microscope (×40; Axio Scope.A1; Zeiss AG). The red
signal that was observed indicated a positive signal and lower
levels indicated that the cell proliferation ability was lower.
MTT assay
Logarithmic growth phase cells were seeded in a
96-well plate, in triplicate, at a density of 5×103/well
(the edge holes were filled with sterile PBS). Different
concentrations of Que (0, 20, 40, 60, 80, 100, 120, 160 and 200 µM)
were used to treat the cells for 24, 48 and 72 h, MTT solution (0.5
mg/ml; 0.05% MTT) was added to each well and the culture was
continued for 4 h. Then, 150 µl dimethyl sulfoxide was added and
the plate was placed on a shaker at low speed for 10 min to fully
dissolve the crystals. The absorbance was measured at 490 nm and
the IC50 was calculated using SPSS software (v20.0; IBM
Corp.).
The inhibitors, afatinib (2 µM) and U0126 (10 µM)
were also used to measure cell viability and added to the cells 2 h
before the cells were then treated with Que for 24 h.
Detection of cell cycle
The cells were treated with Que (120 µM) for 8 h,
then digested and resuspended using 1 ml pre-chilled 70% alcohol,
administered drop by drop. After fixing overnight at −20°C, the
samples were centrifuged at 3,000 × g for 5 min at 4°C to remove
the ethanol. Then, 5 µl RNase was added and the samples were
incubated in a 37°C water bath for 30 min, following which 100 µl
1X binding buffer and 5 µl PI staining solution (Becton, Dickinson
and Company) were added into each sample and incubated at room
temperature for 15 min in the dark. Subsequently, the cell cycle
was detected using a flow cytometer (BD FACS Calibur; Becton,
Dickinson and Company) and analyzed using FlowJo software (v10;
Tree Star, Inc.).
Wound healing assay
The HeLa and SiHa cells were seeded into a 6-well
culture plate with horizontal straight lines at the bottom, at a
density of 5×106 cells/ml per hole. The cells were
gently scratched according to the middle line using a pipette tip
after 24 h. Then, the plate was washed with PBS to remove cell
debris, then cultured again for 24 and 36 h. At the beginning, the
degree of confluence at 0 h time point on both sides of the wound
was checked using a microscope (×40; Axio Scope.A1; Zeiss AG) and
was 80–100%. Images of all the groups at 0, 24 and 36 h were
captured, and the wound healing difference after Que administration
at the indicated concentration (0 and 80 µM) in serum-free
RPMI-1640 medium (Biological Industries) were detected. The cell
scratch area was used as an index to evaluate the degree of wound
healing and was analyzed using ImageJ software (v1.8.0.112;
National Institutes of Health).
Transwell assay
For the migration assay, the HeLa and SiHa cells, in
the log-phase, at a density of 5×104 cells/ml, were
added to the upper chamber, in a 200 µl suspension diluted with 1%
FBS-containing RPMI-1640 medium (Biological Industries). Next, 600
µl RPMI-1640 culture medium, containing 10% serum, was added to the
lower chamber for a 24 h incubation. Each group had 2 replicates.
For the invasion assay, Matrigel was dissolved overnight at 4°C and
diluted with basic culture medium at a volume ratio of 1:7. Then,
20 µl Matrigel was placed on the Transwell chamber in a sterile
24-well plate at 37°C for 30 min. The subsequent methods used were
performed according to the same protocol as the cell migration
assay. After the Matrigel solidified, the non-migrated/invaded
cells on the membrane were removed using a cotton swab and PBS. The
cells were fixed with 4% paraformaldehyde solution at room
temperature for 10 min, then stained with crystal violet at room
temperature for 5 min. The number of migrated and invaded cells,
and the average value for each sample were observed and calculated
using a light microscope (×40; Axiolab 5; Zeiss AG).
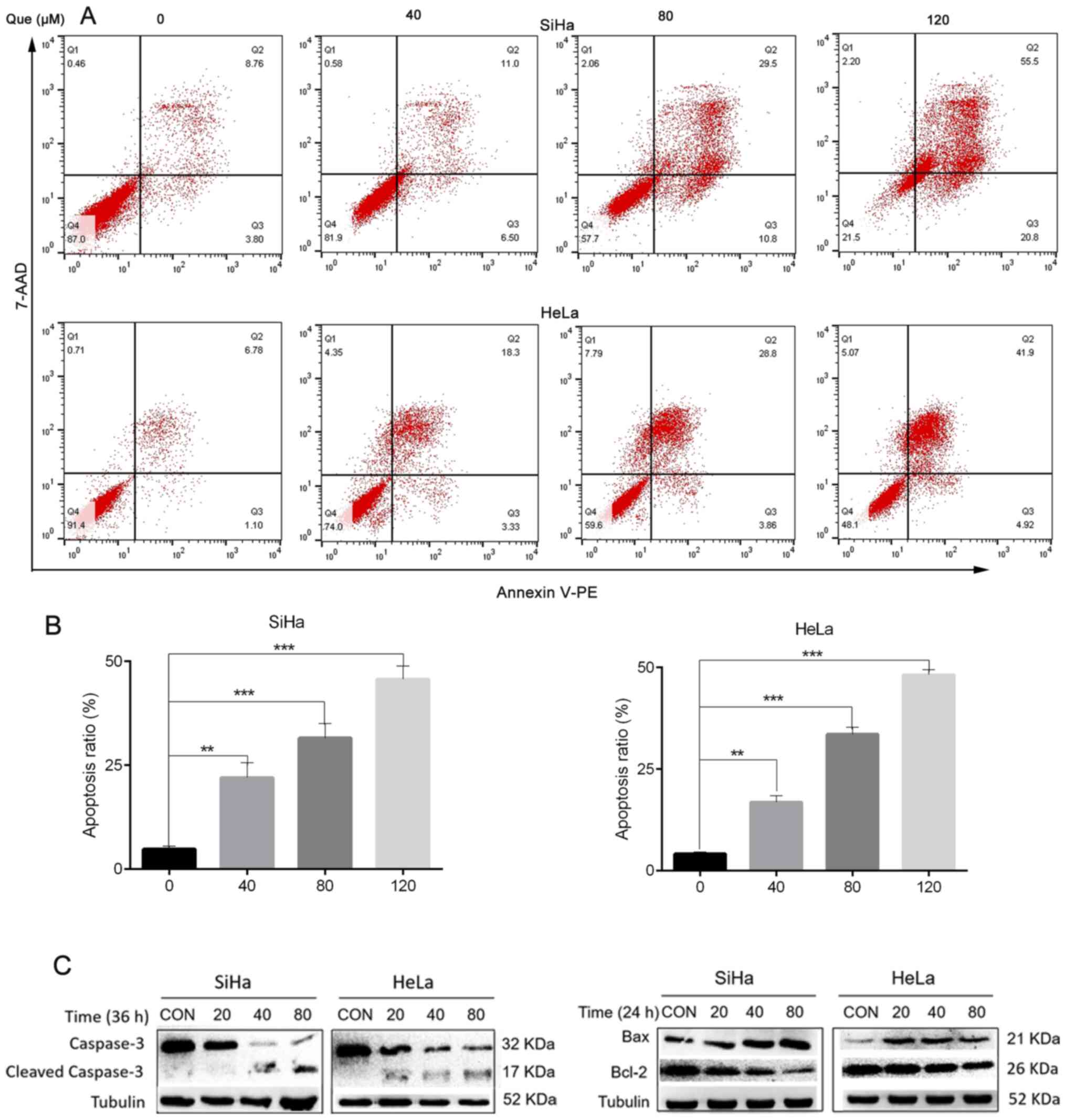
Annexin V-PE/7-AAD double-staining
assay
Cell apoptosis was detected using a double staining
Annexin V-PE/7-AAD (BD Biosciences) method. The cells were treated
with Que (0, 40, 80 and 120 µM) for 24 h, then digested with
trypsin, without EDTA and rinsed with sterile 1X binding buffer,
diluted with PBS. The cells were subsequently incubated at room
temperature for 10 min in the dark with 5 µl Annexin V-PE and 5 µl
7-AAD. FACS flow cytometry (BD FACS Calibur; Becton, Dickinson and
Company) was performed to detect apoptosis and the data were
analyzed using FlowJo software (v10; FlowJo LLC). The number of
cells in the upper right and the lower right quadrants of the flow
cytometry plots represent the proportion of late and early
apoptotic cells respectively, and the sum of the two represents the
proportion of cervical cancer cells undergoing apoptosis.
Western blot analysis
RIPA (Beijing Solarbio Science and Technology Co.,
Ltd.) was used to extract the total protein from the cells and the
BCA method was used for sample concentration determination. In
total, ~20 µg total protein was used for separation using 8–12%
SDS-PAGE, following which the separated protein was
electro-transferred to a PVDF membrane, and the primary antibody
was added and incubated overnight at 4°C after blocking with 5%
skimmed milk at room temperature for 1 h. The membranes were washed
3 times with 1X TBS-Tween-20 (at a ratio of 1,000:1), for 10 min
each time, then the membranes were incubated for 2 h with the
corresponding secondary antibody (1:3,000) at room temperature. The
expression of each target protein was detected using an ECL reagent
(EMD Millipore) and a multifunctional molecular gel imaging
system.
The following inhibitors, afatinib (2 µM), U0126 (10
µM), PD98059 (5 µM) and ML141 (10 µM) were added to the cells 2 h
before the cells were then treated with Que for 8 h at different
combinations, to also measure protein expression. Each experiment
was repeated at least 3 times and the representative results are
presented.
Statistical analysis
GraphPad Prism v8.0 software (GraphPad Software,
Inc.) was used to analyze the data. One-way ANOVA followed by
Tukey's post hoc test or unpaired Student's t-test were used to
evaluate the statistical differences between >2 or 2 groups,
respectively. P<0.05 was considered to indicate a statistically
significant difference.
Results
Que in SH targets EGFR
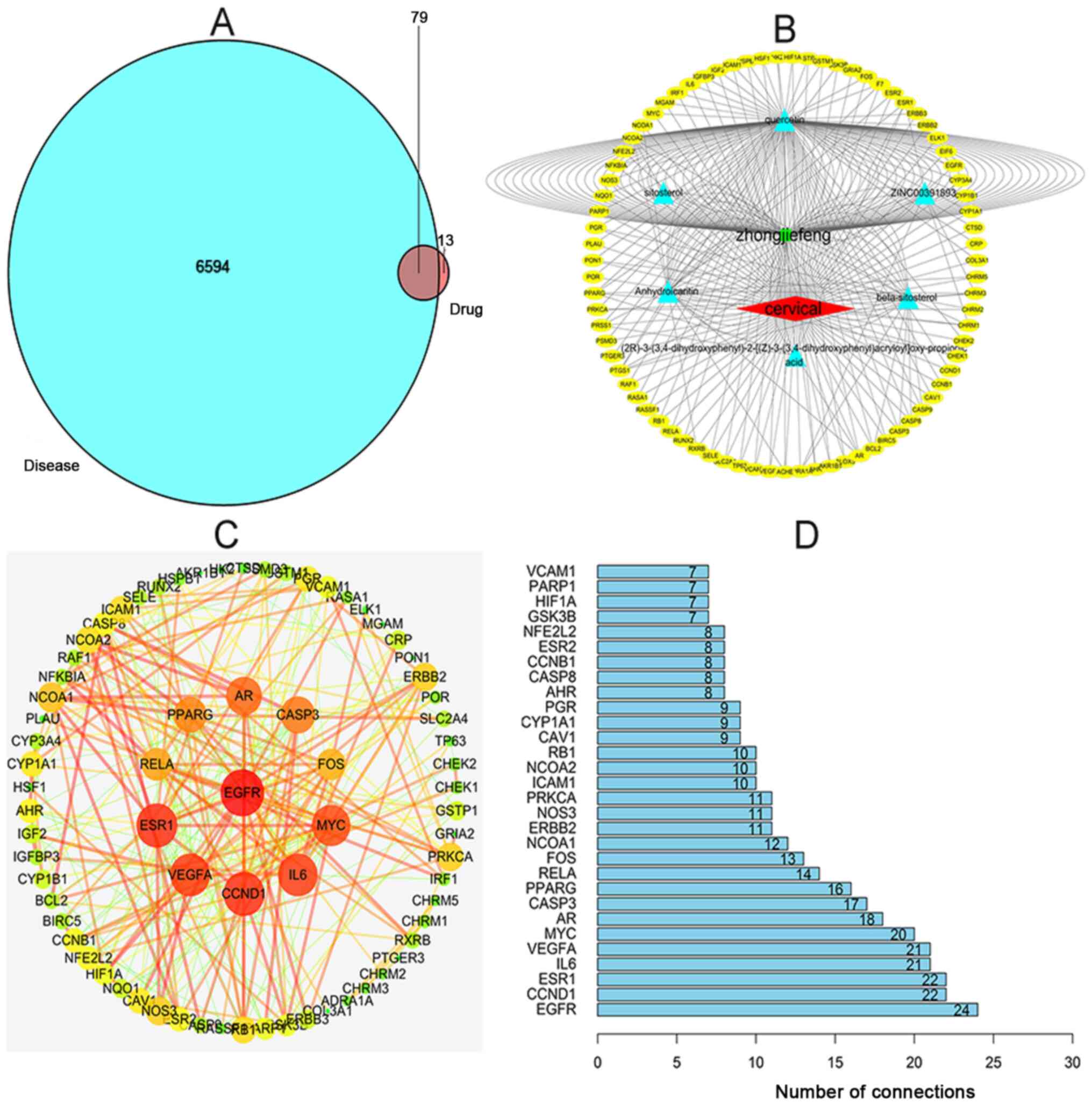
There were 92 targets predicted by TCMSP, for the
active ingredients of SH, and a total of 6,673 targets, associated
with cervical cancer, were searched in the GeneCards and OMIM
databases. In addition, the total number of drug-related targets
was 92, with 79 common targets following the intersection with
disease-related targets. The drug component-target-disease
relationship network diagram was established using R software
(Fig. 1A and B). The yellow ellipse
represents common targets of cervical cancer and the drug, while
the blue triangles represent the active ingredients of SH. The same
targets, corresponding to different effective ingredients, are
presented in Fig. 1B, which shows
the characteristics of the multi-components and multi-targets of
TCM. It can also be seen that Que has an intersection with almost
every target, which proves that it has strong research value
(Fig. 1B). In Fig. 1C, the color and thickness of the
lines represent the strength of the association and the final
functional protein network diagram is presented. The obtained
network revealed the node, EGFR as the hub gene, suggesting that it
may have important biological significance in SH-induced anticancer
effects in cervical cancer cell lines (Fig. 1C and D).
Que reduces cell viability
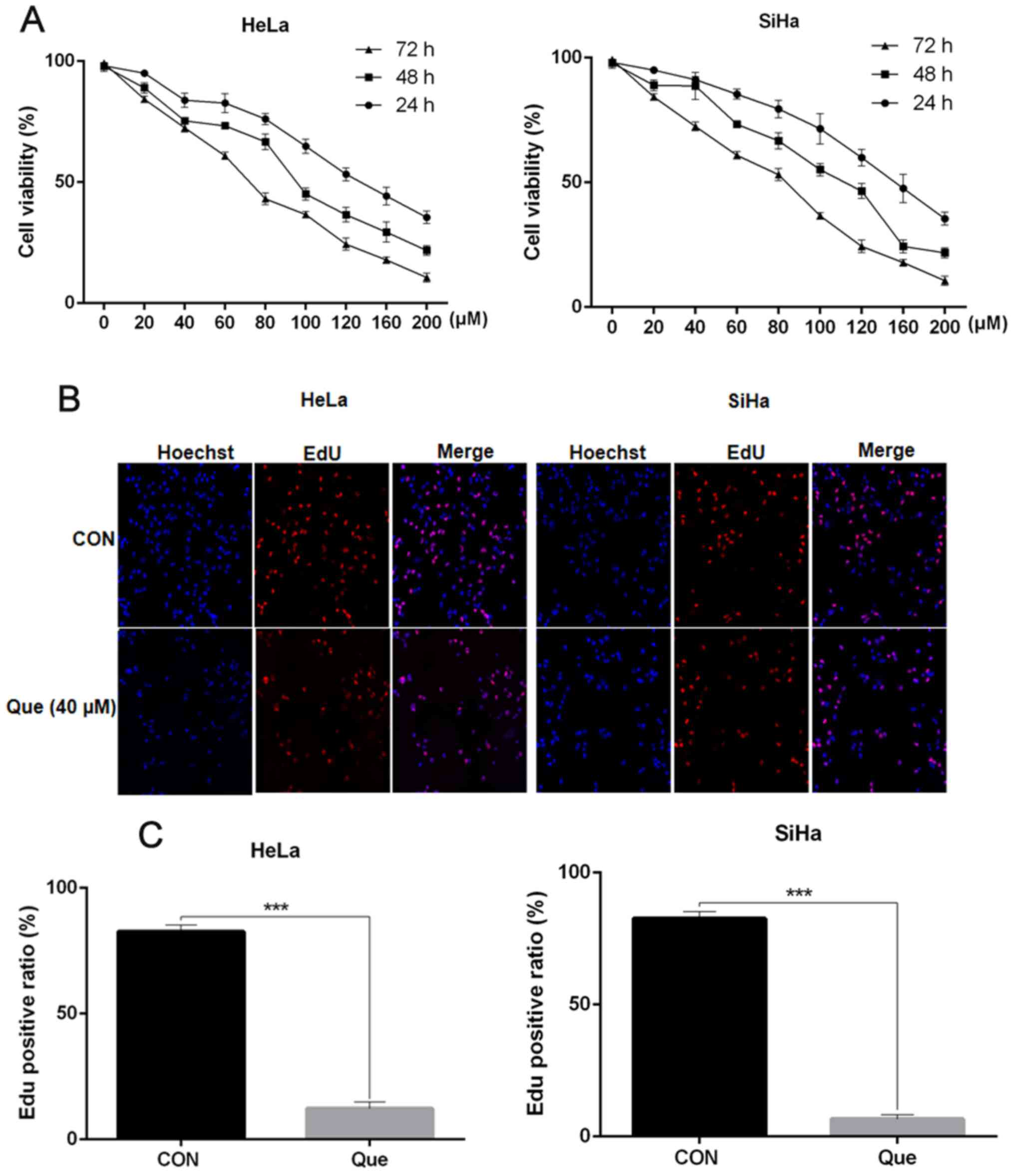
To determine the cytotoxic effect of Que on the
human cervical cancer cell lines, the cells were treated with
increasing concentrations of Que for 24, 48 and 72 h, then a MTT
assay was performed to measure cell viability. The IC50
of SiHa and HeLa cells at the three time points was 150.21, 112.06
and 83.3 µM, and 132.12, 95.25 and 70.12 µM, respectively (Fig. 2A). Consistently, the EdU staining
results also suggested that Que inhibited the proliferation and
division of the cervical cancer cells (Fig. 2B).
Que induces G2/M
arrest
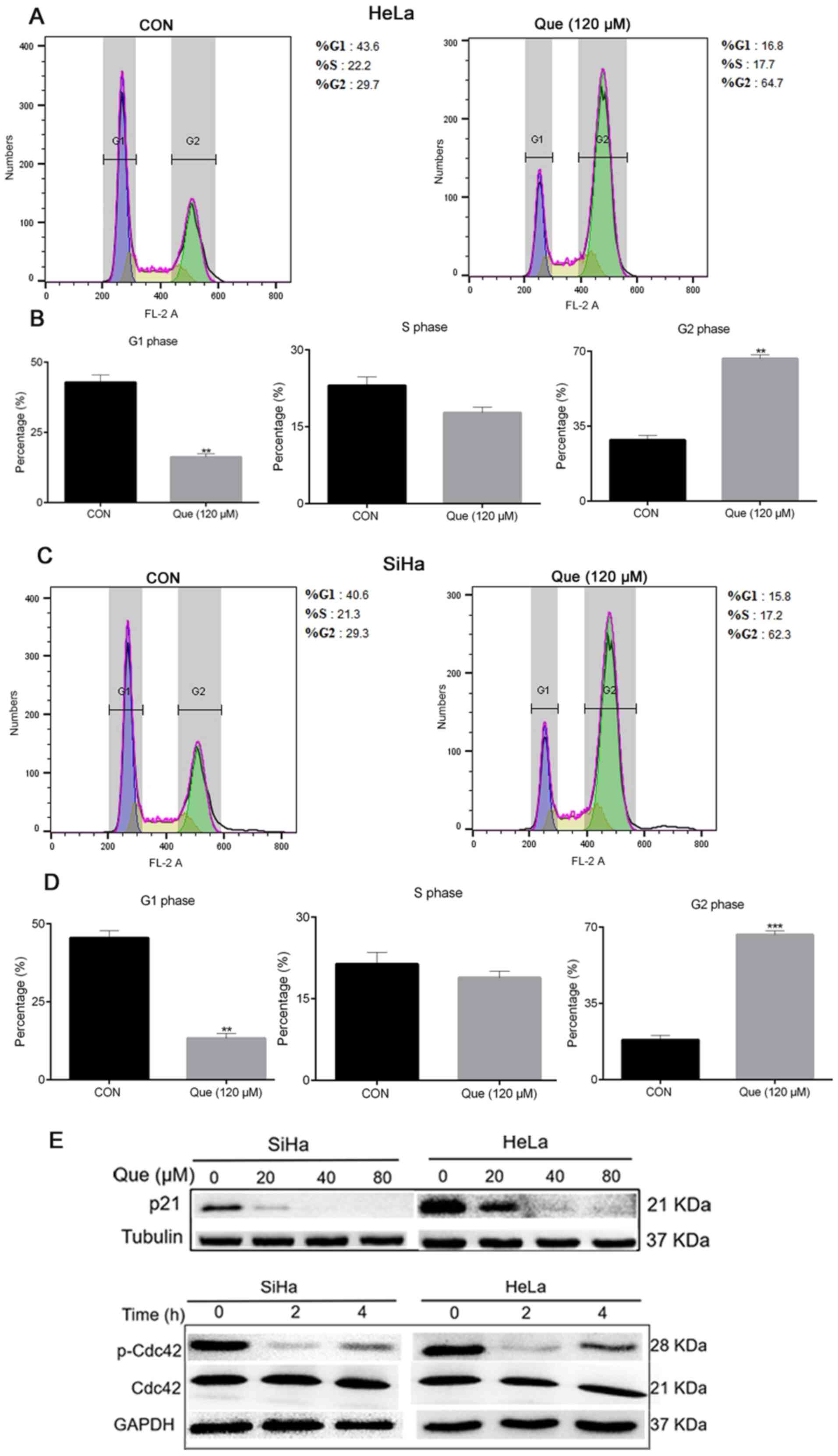
Next, it was investigated whether Que was associated
with the cell cycle of the cervical cancer cells. The HeLa and SiHa
cell lines were treated with Que for 8 h, following which flow
cytometry was performed. The results demonstrated that Que
significantly increased the percentage of SiHa and HeLa cells in
the G2/M phase. It was found that the percentages of
G2/M cells were 29.3 and 62.3% for the SiHa cells and
29.7 and 54.7% for the HeLa cells, at 0 and 120 µM, respectively
(Fig. 3A-D), indicating that Que
induced G2/M arrest in the cervical cancer cell lines.
Western blot analysis revealed that Que caused a decline in p-Cdc42
and p21 protein expression level in a time and
concentration-dependent manner, respectively, which served a key
role in the regulation of cell cycle (Fig. 3E). Collectively, it was suggested
that Que notably blocked the cervical cancer cell cycle in the
G2/M phase.
Que inhibits cell migration and
invasion
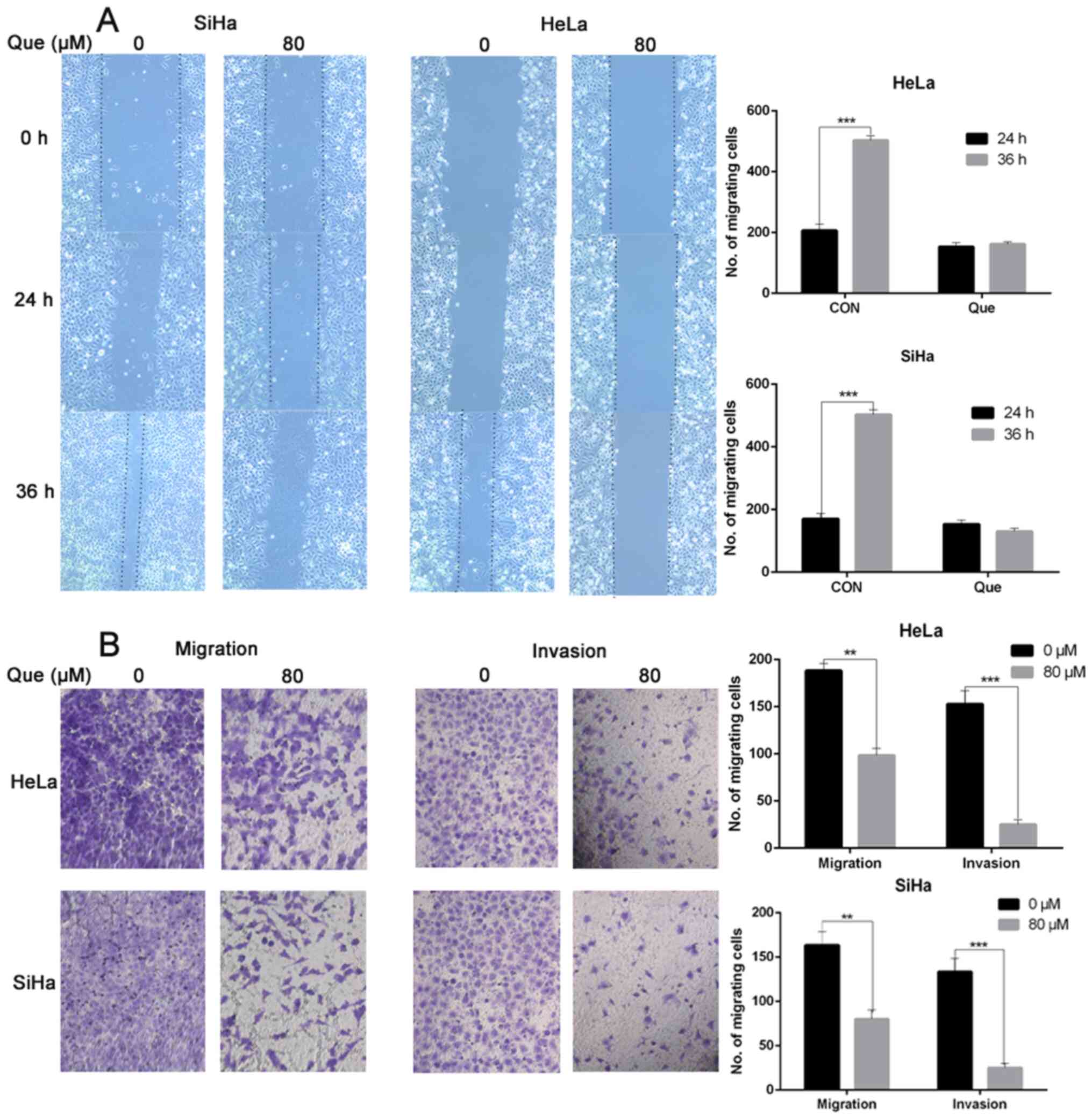
A wound healing assay was performed to investigate
whether Que inhibited cervical cancer cell migration. It was
identified that Que significantly suppressed the wound healing rate
of the HeLa and SiHa cells (Fig.
4A). Furthermore, a Transwell assay was conducted to determine
whether Que inhibits cervical cancer cell migration and invasion.
The number of cells passing through the membrane and Matrigel to
the lower chamber was decreased with Que was gradually decreased
with increasing Que concentrations (Fig.
4B), suggesting that Que suppressed the migration and invasion
of the SiHa and HeLa cells.
Que promotes cervical cancer cell
apoptosis
The cells treated with Que for 24 h presented with
the morphological features of apoptosis, including shrinking in
size, becoming round, losing connections and breaking away from the
surrounding cells (data not shown). Subsequently, Que-induced
apoptosis was further evaluated using Annexin V-PE/7-AAD staining.
As presented in Fig. 5A, the
proportion of cells undergoing apoptosis was gradually enhanced
with the increasing Que concentrations compared with that in the
control group.
Caspase-3 serves an important role in cell apoptosis
and this process can be blocked by Bcl-2 (14). The results demonstrated that Que
significantly promoted the activation of caspase-3. Furthermore,
Que increased the expression levels of the pro-apoptotic protein,
Bax and decreased the expression level of the anti-apoptotic
protein, Bcl-2 (Fig. 5C).
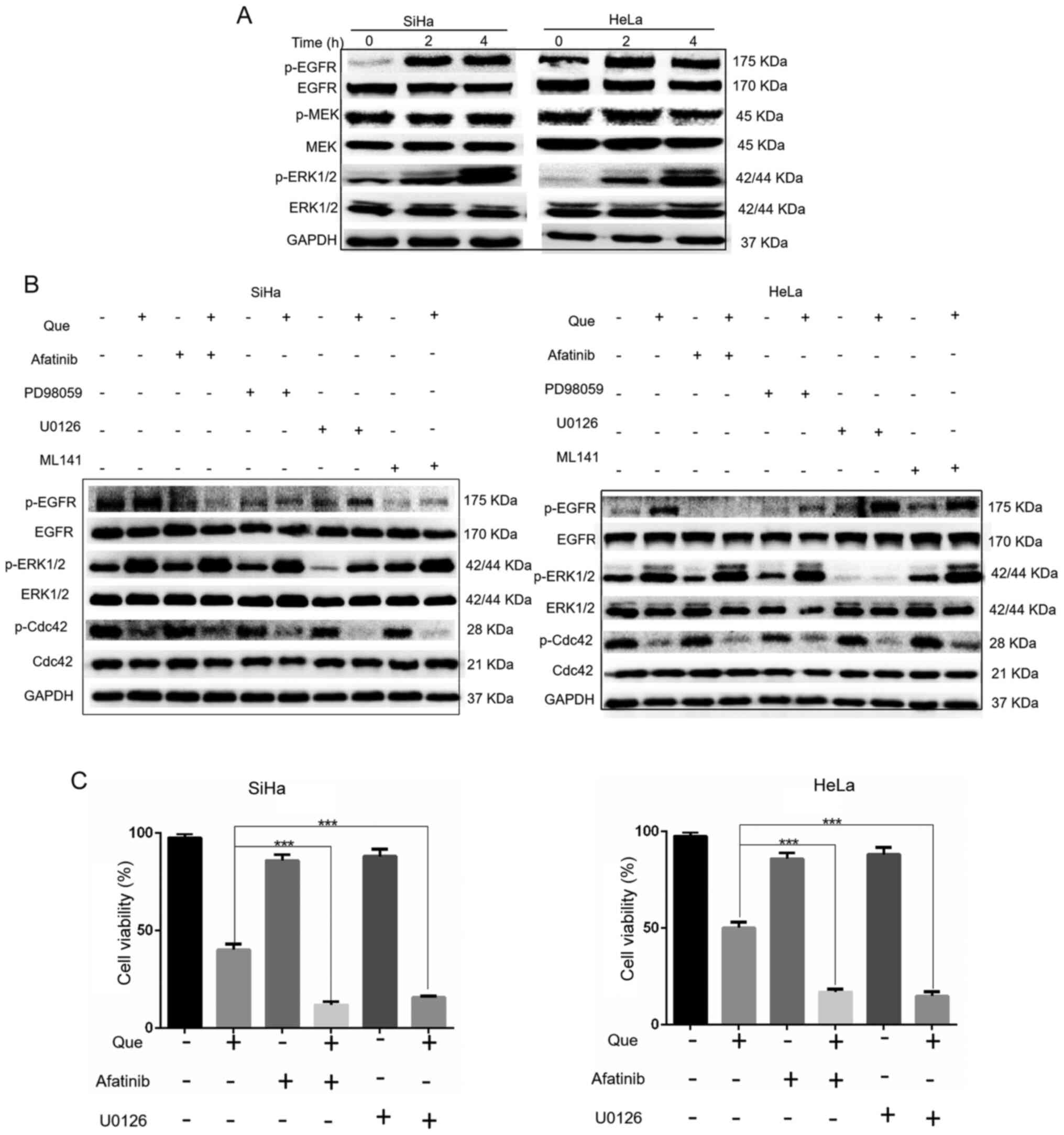
EGFR and ERK activation resists
Que-induced cytotoxicity in cervical cancer cells
According to the results of network pharmacology,
EGFR may be associated with the Que-induced anticancer effects in
cervical cancer. The results indicated that Que markedly increased
EGFR activities, but suppressed the activity of Cdc42 (Figs. 6A and 3E, respectively). It has been
well-documented that EGFR activation leads to downstream MEK/ERK
pathway activation (15). As shown
in Fig. 6A, Que notably induced ERK
activation, but only marginally affected the activities of
MEK1.
To further examine the role of EGFR and ERK in
Que-induced anticancer effects, afatinib and U0126 were used to
block the activities of EGFR and ERK, respectively. It was found
that both afatinib and U0126 effectively inhibited EGFR and ERK
activation induced by Que. EGFR is the upstream signal of ERK;
therefore, if Que acts on EGFR, then the EGFR inhibitor afatinib
will reverse the activation of ERK under the action of quercetin.
Unexpectedly, afatinib did not reverse Que-induced ERK activation,
indicating that Que may increase ERK activity in an
EGFR-independent manner. In addition, PD98059 and ML141 were used
to detect the Ras/Raf/MEK/ERK/MAPK signaling pathway, ML141 further
promoted the reduction of Cdc42 protein expression in combination
with Que (Fig. 6B).
Next, it was investigated whether EGFR and ERK were
associated with Que-induced anticancer activities. It was found
that the suppression of EGFR and ERK, using the afatinib and U0126
inhibitors, enhanced Que-induced cell viability inhibition
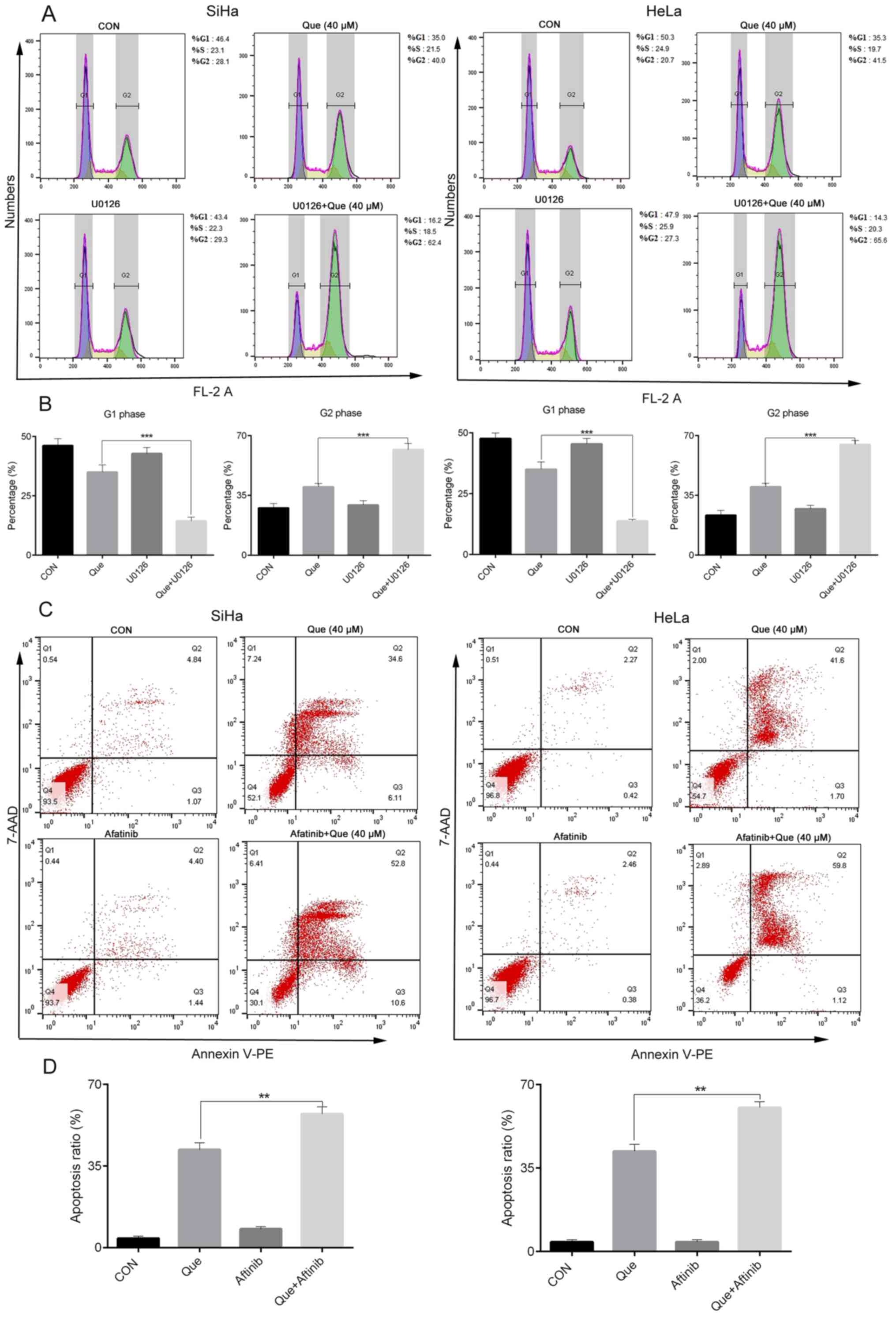
(Fig. 6C), cell cycle arrest and
apoptosis (Fig. 7A-D). For example,
the apoptotic rate of the HeLa and SiHa cells with Que alone was
43.3 and 40.71%, respectively, but when afatinib and Que or U0126
and Que were added together, the apoptotic rate increased to 60.92
and 63.4%. The G2/M phase ratio in the HeLa and SiHa
cells with Que alone was 41.5 and 40.0%, but when U0126 and Que
were combined, the ratio increased to 65.6 and 62.4%, respectively
suggesting that EGFR and ERK activation had a cytoprotective role
in the Que-induced anticancer effects in cervical cancer cell
lines.
Discussion
At present, the morbidity and mortality rates of
cervical cancer are at the forefront of malignant tumors, which
seriously threatens the health of women (16). In recent years, plant-derived
compounds, with the potential to inhibit the development and the
proliferation of tumors, have attracted increased attention from
scientific researchers (17–19). In the present study, based on network
pharmacology, Que is one of the effective ingredients of SH, then
the hub gene, EGFR was identified, which could be associated with
Que. Que is a natural flavonoid compound, widely present in
numerous plants, such as onions, shallots, asparagus, cabbage,
mustard greens and green peppers and has a preventive and
therapeutic effect on a variety of cancer types, including stomach,
breast, liver and cervical cancers (20). Most in vivo and in
vitro studies have shown that Que was not carcinogenic
(21–23). In 1999, the International Agency for
Research on Cancer classified Que as a non-carcinogenic substance
to human (24). Phase I and Phase II
clinical trials in England have confirmed that Que inhibited the
progression of a variety of tumors, and it has been circulated as a
commodity in the United States and some European countries
(25,26). Nutrigenomics aims to understand how
the active ingredients of food directly or indirectly affect the
changes in the human genome structure, and to investigate the
effects of dietary factors and nutrients on the human genome. In
addition, to investigate which chronic or hereditary diseases, such
as irritable bowel syndrome, musculoskeletal pain, chronic pelvic
pain and dry eye are susceptible to dietary factors, and explore
the differences in the sensitivity of healthy humans and diseased
patients to different dietary factors based on differences in human
genetic polymorphisms (27).
The occurrence of cervical cancer may be associated
with the activation of certain proto-oncogenes, such as EGFR, which
is widely distributed in mammalian epithelial cells and binds to
EGF or TNF (28,29). EGFR is a glycoprotein that belongs to
the tyrosine kinase type receptor family and its different
phosphorylation sites regulate various downstream signal pathways,
such as PI3K/AKT and Ras/Raf/MEK/ERK (30). The current study focused on signaling
regulation after activation at Tyr1068. Tyrosine kinase receptor,
which is activated by growth factors, binds to growth factor
receptor-bound protein 2 (GRB2) directly or indirectly, following
which GRB2 recruits the guanylate exchange factor, SOS (Ras/Rac
guanine nucleotide exchange factor 1), which is located on the cell
membrane adjacent to Ras and a series of cascade amplification
reactions occur after Ras is activated (31,32). The
transcriptome refers to the collection of all transcripts in a
specific type of cell. In other words, it is a snapshot of the
expression profile of a cell at a specific time. In the
transcriptome, it includes both the familiar messengers that encode
proteins (mRNA), non-coding RNA, such as microRNA, which does not
code for protein and long non-coding (lnc)RNA, which are newly
discovered (33,34). These RNA transcripts combine to
regulate a series of important physiological processes, such as
cell growth, development and apoptosis (35). After autophosphorylation, EGFR binds
directly or indirectly to the GRB2 complex to activate the Ras
protein. The activated Ras activates the downstream
serine/threonine protein kinase, Raf to phosphorylate ERK-1 and 2.
The signal is transmitted to the nucleus, leading to the
phosphorylation of transcription factors in the nucleus, starting
the transcription of target genes, and ultimately leading to
biological effects, such as cell proliferation and proliferation
(36). In total, 4 major MAPKs have
been identified: ERK1/2, JNK/stress activated protein kinases, p38
MAPK and ERK5 (37).
ERK activation can exert an anti-apoptotic or
proapoptotic effects, depending on the stimulation and cell types.
ERKs are mainly, but not only, activated by growth factors and
participate in mitosis regulation (38). It also transfers to the nucleus and
activates transcription factors to produce corresponding biological
effects directly after being activated (39). The activation of Raf is not
completely dependent on Ras and ERK can also be activated by other
factors, such as SOS and ILs, thus forming a complex network
regulatory structure (40). The
present results suggested that Que activated ERK independent of
EGFR, and published studies have confirmed that receptor tyrosine
kinases, G protein-coupled receptors and some cytokine receptors,
such as Ras, Raf and MEK can all activate the ERK signal
transduction pathway (41,42). Deng et al (43) reported that Que induced human breast
cell apoptosis by inhibiting ERK, high-affinity potassium
transporter and survivin. In addition, Kim et al (44) observed that Que promoted cycle arrest
and apoptosis of melanoma cells by increasing the ratio of
Bax/Bcl-2, and inhibited the PI3K/AKT and ERK/JNK pathways involved
in cell proliferation and signal transduction, while Balakrishnan
et al (45) revealed that Que
induced MCF-7 breast cancer cell apoptosis by activating
AMP-activated protein kinase, thereby inhibiting the expression of
cyclooxygenase-2.
EdU is a thymidine analogue and the alkyne group
attached to this factor is rare in natural compounds. This group
can replace thymine into a synthetic DNA molecule during DNA
replication (46). Based on the
specific reaction between Apollo fluorescent dye and EdU, DNA
replication activity can be directly and accurately detected
(47). The present MTT and EdU assay
results demonstrated that Que had a notable inhibitory activity on
the viability of cervical cancer cells, while flow cytometry
detection identified that Que significantly promoted
G2/M phase arrest and apoptosis. Furthermore, the wound
healing and Transwell assays suggested that Que inhibited cellular
migration and invasion.
Apoptosis and cell cycle processes are strictly
governed by multiple genes, such as Bcl-2, caspase and the CDK
family (48). Caspase-3 is one of
the most important enzymes, which is a terminal splicing factor in
the process of cell apoptosis and plays an irreplaceable role in
this process. The main substrate of caspase-3 is poly (ADP-ribose)
polymerase 1, and caspase-3 can be cleaved by this factor, leading
to an inhibition of DNA repair and initiation of degradation
(49,50). Cdc42 is a member of the ρ family, and
serves an important role in the establishment and maintenance of
epithelial cell polarity by regulating the formation of
microtubules and microfilament skeletons and intercellular
connection (51). It was found that
the protein expression level of p-Cdc42 was markedly decreased
following the addition of Que; however, its expression after 4 h of
Que treatment was slightly increased compared with that after 2 h.
This may be caused by the negative feedback regulation mechanism,
which requires further investigation. p21 was first discovered as a
downstream gene of p53, and subsequently, scientists have conducted
numerous studies examining p21 and have shown that it is an
important cell cycle government target (52,53). p21
is involved in the process of cell proliferation, differentiation,
senescence and death (54). At the
same time, it is associated with the occurrence of tumors and
serves a role in the physiological and pathological processes.
Previous studies have confirmed that p21 can form a complex with
cyclin D/CDK to block the cell cycle in G1 phase and it can also
interact with proliferating cell nuclear antigen (PCNA) via the
C-terminus to block the activity of PCNA-activated DNA polymerase,
thereby inhibiting DNA synthesis and arresting the cell cycle
(55,56).
MAPK can transduce extracellular signals into the
nucleus via a 3-level kinase cascade (MAPKKK>MAPKK>MAPK),
thereby regulating cell proliferation, differentiation, apoptosis,
inflammatory response and other biological behaviors, such as
antitumor effects (57). The
RAS/RAF/MEK/ERK axis has been the mostly widely investigated of the
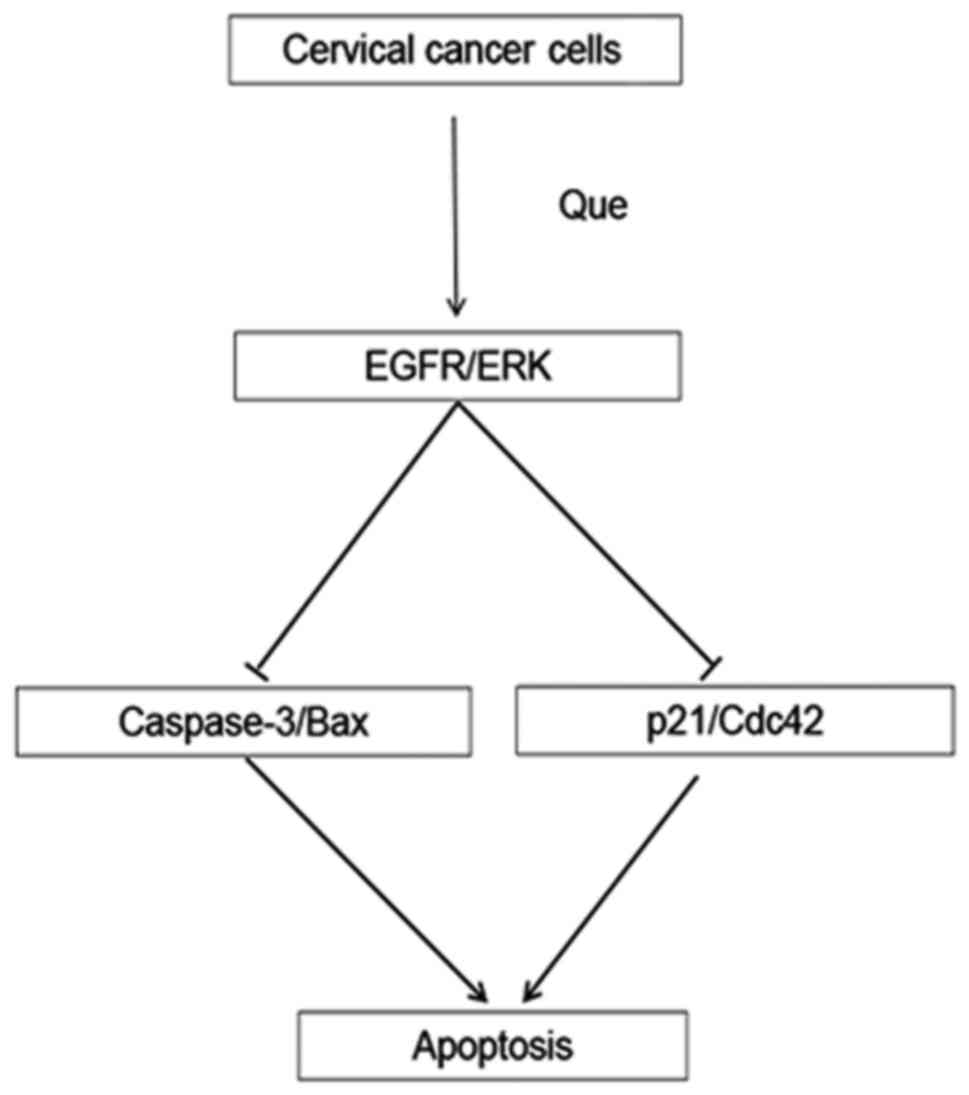
aforementioned cascades. The present study analyzed the negative
regulatory effect of the activation of EGFR and ERK when the
cervical cancer cell lines were treated with Que, as both EGFR and
ERK inhibitors enhanced the Que-induced anticancer effects. Que
activates EGFR and ERK and inhibits the activity of related
kinases, indicating that the activation of EGFR and ERK resists the
effect of Que-mediated apoptosis in cervical cancer cells (Fig. 8).
In conclusion, the present study conducted network
pharmacology analysis, and the results demonstrated that Que may be
one of the active components of SH. Furthermore, Que notably
inhibited the viability of cervical cancer cells, and induced
G2/M arrest and apoptosis. EGFR and ERK activation was
associated with the resistance to Que-induced anticancer
activities, as both EGFR and ERK inhibitors enhanced the
Que-induced anticancer effects. Thus, it would be important to
examine the combination of Que and EGFR inhibitors in the treatment
of cervical cancer. Collectively, the present study suggested that
EGFR and ERK activation served a protective role in Que-induced
anticancer activities in cervical cancer in vitro. If Que is
used for clinical application, the combination of afatinib may
enhance the antitumor effect of Que one day, but this topic is
based on the conclusions obtained from in vitro cell
experiments and in vivo animal experiments have not been
conducted, therefore further investigation is required to confirm
the conclusions.
Acknowledgements
Not applicable.
Funding
This study was supported in part by the National
Natural Science Foundation of China (grant nos. 81560429, 81602545
and 81760519) and the Yunnan Natural Science Foundation of China
[grant nos. 2017FF116 (−023) and 2019FF002 (−011)]
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
XC completed the experiments. PX and LG made
substantial contributions to conception and design. HZ, XS, LG, XZ,
JW and PH analyzed the data, drafted the manuscript and revised it
critically for important intellectual content. RS and QZ approved
the final version of the manuscript, confirmed the authenticity of
the raw data generated during the study, and guided and planned the
study. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Woo S, Atun R, Ward ZJ, Scott AM, Hricak H
and Vargas HA: Diagnostic performance of conventional and advanced
imaging modalities for assessing newly diagnosed cervical cancer:
Systematic review and meta-analysis. Eur Radiol. 30:5560–5577.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Tran KN, Park Y, Kim BW, Oh JK and Ki M:
Incidence and mortality of cervical cancer in Vietnam and Korea
(1999–2017). Epidemiol Health. 42:e20200752020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Benitez-Restrepo CC, Arias-Ortiz NE and
Arboleda-Ruiz WA: Cervical cancer incidence and patient survival in
Manizales, Colombia, 2008–2012. Rev Peru Med Exp Salud Publica.
37:438–445. 2020.(In Spanish, English). View Article : Google Scholar : PubMed/NCBI
|
|
4
|
D'Alton P, Craddock F, Bergin N and Lynch
J: Exploring the lived experience of partners of women impacted by
cervical cancer and the CervicalCheck screening failure in Ireland.
Psychooncology. Jun 7–2021.(Epub ahead of print). View Article : Google Scholar
|
|
5
|
Malikova H, Burghardtova M, Fejfarova K,
Nadova K and Weichet J: Advanced cervical cancer in young women:
Imaging study of late and very late radiation-related side effects
after successful treatment by combined radiotherapy. Quant Imaging
Med Surg. 11:21–31. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhang FY, Li JJ, Zhou Y and Xu XY: Review
for sedative and hypnotic mechanism of sedative traditional Chinese
medicine and relative active components on neurotransmitters.
Zhongguo Zhongyao Zazhi. 41:4320–4327. 2016.(In Chinese).
PubMed/NCBI
|
|
7
|
Qian C, Kuang M and Wang Y: Effect of
qianghuo erhuang decoction on T regulatory and T Helper 17 cells in
treatment of adjuvant-induced arthritis in rats. Sci Rep.
7:171982017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Chen H, Feng X, Gao L, Mickymaray S,
Paramasivam A, Abdulaziz Alfaiz F, Almasmoum HA, Ghaith MM,
Almaimani RA and Aziz Ibrahim IA: Inhibiting the PI3K/AKT/mTOR
signalling pathway with copper oxide nanoparticles from
Houttuynia cordata plant: Attenuating the proliferation of
cervical cancer cells. Artif Cells Nanomed Biotechnol. 49:240–249.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zeng Y, Liu J, Zhang Q, Qin X, Li Z, Sun G
and Jin S: The traditional uses, phytochemistry and pharmacology of
Sarcandra glabra (Thunb.) Nakai, a Chinese herb with
potential for development: Review. Front Pharmacol.
12:6529262021.Review. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhang Y, Yuan T, Li Y, Wu N and Dai X:
Network pharmacology analysis of the mechanisms of compound Herba
Sarcandrae (Fufang Zhongjiefeng) aerosol in chronic pharyngitis
treatment. Drug Des Devel Ther. 15:2783–2803. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Guo X, Shen L, Tong Y, Zhang J, Wu G, He
Q, Yu S, Ye X, Zou L, Zhang Z, et al: Antitumor activity of caffeic
acid 3,4-dihydroxyphenethyl ester and its pharmacokinetic and
metabolic properties. Phytomedicine. 20:904–912. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Trusov NV, Apryatin SA, Shipelin VA and
Gmoshinski IV: Full transcriptome analysis of gene expression in
liver of mice in a comparative study of quercetin efficiency on two
obesity models. Probl Endokrinol (Mosk). 66:31–47. 2020.(In
Russian). View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Flater D: A system of quantities from
software metrology. Measurement (Lond). 168:2021.doi:
10.1016/j.measurement.2020.108435. PubMed/NCBI
|
|
14
|
Mokhtari Sangdehi SR, Hajizadeh Moghaddam
A and Ranjbar M: Anti-apoptotic effect of silymarin-loaded chitosan
nanoparticles on hippocampal caspase-3 and Bcl-2 expression
following cerebral ischemia/reperfusion injury. Int J Neurosci. Jan
21–2021.(Epub ahead of print). View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhang X, Huang R, Gopalakrishnan S,
Cao-Milán R and Rotello VM: Bioorthogonal nanozymes: Progress
towards therapeutic applications. Trends Chem. 1:90–98. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Luckett R and Feldman S: Will HPV
vaccination affect cervical cancer morbidity and mortality
world-wide? Hum Vaccin Immunother. 12:1373–1374. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Dehelean CA, Marcovici I, Soica C, Mioc M,
Coricovac D, Iurciuc S, Cretu OM and Pinzaru I: Plant-derived
anticancer compounds as new perspectives in drug discovery and
alternative therapy. Molecules. 26:262021. View Article : Google Scholar
|
|
18
|
Mohsenpour H, Pesce M, Patruno A, Bahrami
A, Pour PM and Farzaei MH: A review of plant extracts and
plant-derived natural compounds in the prevention/treatment of
neonatal hypoxic-ischemic brain injury. Int J Mol Sci. 22:222021.
View Article : Google Scholar
|
|
19
|
Yoo S, Yang HC, Lee S, Shin J, Min S, Lee
E, Song M and Lee D: A deep learning-based approach for identifying
the medicinal uses of plant-derived natural compounds. Front
Pharmacol. 11:5848752020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Vu DL, Žabčíková S, Červenka L, Ertek B
and Dilgin Y: Sensitive voltammetric determination of natural
flavonoid quercetin on a disposable graphite lead. Food Technol
Biotechnol. 53:379–384. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Harwood M, Danielewska-Nikiel B,
Borzelleca JF, Flamm GW, Williams GM and Lines TC: A critical
review of the data related to the safety of quercetin and lack of
evidence of in vivo toxicity, including lack of
genotoxic/carcinogenic properties. Food Chem Toxicol. 45:2179–2205.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Elmowafy M, Musa A, Alnusaire TS, et al:
Olive Oil/Pluronic Oleogels for Skin Delivery of Quercetin: In
Vitro Characterization and Ex Vivo Skin Permeability. Polymers
(Basel). 13:2021. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kim JM, Seo SW, Han DG, Yun H and Yoon IS:
Assessment of metabolic interaction between repaglinide and
quercetin via mixed inhibition in the liver: In vitro and in vivo.
Pharmaceutics. May 23–2021.(Epub ahead of print).
|
|
24
|
Boulton DW, Walle UK and Walle T: Fate of
the flavonoid quercetin in human cell lines: Chemical instability
and metabolism. J Pharm Pharmacol. 51:353–359. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ferry DR, Smith A, Malkhandi J, Fyfe DW,
deTakats PG, Anderson D, Baker J and Kerr DJ: Phase I clinical
trial of the flavonoid quercetin: Pharmacokinetics and evidence for
in vivo tyrosine kinase inhibition. Clin Cancer Res. 2:659–668.
1996.PubMed/NCBI
|
|
26
|
Chalet C, Hollebrands B, Duchateau GS and
Augustijns P: Intestinal phase-II metabolism of quercetin in HT29
cells, 3D human intestinal tissues and in healthy volunteers: A
qualitative comparison using LC-IMS-MS and LC-HRMS. Xenobiotica.
49:945–952. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Rashedi J, Ghorbanihaghjo A, Asgharzadeh M
and Baradaran B: Chitosan and quercetin: Potential hand in hand
encountering tumors in oral delivery system. Curr Pharm Des.
25:3074–3086. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Peng Z, Wang Q, Zhang Y, He J and Zheng J:
EBP50 interacts with EGFR and regulates EGFR signaling to affect
the prognosis of cervical cancer patients. Int J Oncol.
49:1737–1745. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhang W, Gao Y, Jiang Y, Ping L, Cheng H
and Zhang J: EGFR promoter methylation detection in cervical cancer
by a hybridization-fluorescence polarization assay. Diagn Mol
Pathol. 22:102–106. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hartman Z, Geldenhuys WJ and Agazie YM: A
specific amino acid context in EGFR and HER2 phosphorylation sites
enables selective binding to the active site of Src homology
phosphatase 2 (SHP2). J Biol Chem. 295:3563–3575. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Guerrero C, Pesce L, Lecuona E, Ridge KM
and Sznajder JI: Dopamine activates ERKs in alveolar epithelial
cells via Ras-PKC-dependent and Grb2/Sos-independent mechanisms. Am
J Physiol Lung Cell Mol Physiol. 282:L1099–L1107. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Liu Y, Shi QF, Ye YC, Tashiro S, Onodera S
and Ikejima T: Activated O2(•-) and
H2O2 mediated cell survival in
SU11274-treated non-small-cell lung cancer A549 cells via
c-Met-PI3K-Akt and c-Met-Grb2/SOS-Ras-p38 pathways. J Pharmacol
Sci. 119:150–159. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Viñas JL, Spence M, Porter CJ, Douvris A,
Gutsol A, Zimpelmann JA, Campbell PA and Burns KD: micro-RNA-486-5p
protects against kidney ischemic injury and modifies the apoptotic
transcriptome in proximal tubules. Kidney Int. Jun 26–2021.(Epub
ahead of print). View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Xu Z, Shi X, Bao M, Song X, Zhang Y, Wang
H, Xie H, Mao F, Wang S, Jin H, et al: Transcriptome-wide analysis
of RNA m6A methylation and gene expression changes among two
arabidopsis ecotypes and their reciprocal hybrids. Front Plant Sci.
12:6851892021. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Toki MI, Carvajal-Hausdorf DE, Altan M,
McLaughlin J, Henick B, Schalper KA, Syrigos KN and Rimm DL:
EGFR-GRB2 protein colocalization is a prognostic factor unrelated
to overall EGFR expression or EGFR mutation in lung adenocarcinoma.
J Thorac Oncol. 11:1901–1911. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Ouyang L, Chen Y, Wang XY, Lu RF, Zhang
SY, Tian M, Xie T, Liu B and He G: Polygonatum odoratum lectin
induces apoptosis and autophagy via targeting EGFR-mediated
Ras-Raf-MEK-ERK pathway in human MCF-7 breast cancer cells.
Phytomedicine. 21:1658–1665. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Correia I, Wilson D, Hube B and Pla J:
Characterization of a Candida albicans mutant defective in
all MAPKs highlights the major role of Hog1 in the MAPK signaling
network. J Fungi (Basel). 6:62020.
|
|
38
|
Dalton GD and Howlett AC: Cannabinoid CB1
receptors transactivate multiple receptor tyrosine kinases and
regulate serine/threonine kinases to activate ERK in neuronal
cells. Br J Pharmacol. 165:2497–2511. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Tsarouhas V, Yao L and Samakovlis C: Src
kinases and ERK activate distinct responses to Stitcher receptor
tyrosine kinase signaling during wound healing in Drosophila. J
Cell Sci. 127:1829–1839. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Qi ZH, Xu HX, Zhang SR, Xu JZ, Li S, Gao
HL, Jin W, Wang WQ, Wu CT, Ni QX, et al: RIPK4/PEBP1 axis promotes
pancreatic cancer cell migration and invasion by activating
RAF1/MEK/ERK signaling. Int J Oncol. 52:1105–1116. 2018.PubMed/NCBI
|
|
41
|
Yu X, Stallone JN, Heaps CL and Han G: The
activation of G protein-coupled estrogen receptor induces
relaxation via cAMP as well as potentiates contraction via EGFR
transactivation in porcine coronary arteries. PLoS One.
13:e01914182018. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Mulner-Lorillon O, Chassé H, Morales J,
Bellé R and Cormier P: MAPK/ERK activity is required for the
successful progression of mitosis in sea urchin embryos. Dev Biol.
421:194–203. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Deng XH, Song HY, Zhou YF, Yuan GY and
Zheng FJ: Effects of quercetin on the proliferation of breast
cancer cells and expression of survivin in vitro. Exp Ther Med.
6:1155–1158. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Kim SH, Yoo ES, Woo JS, Han SH, Lee JH,
Jung SH, Kim HJ and Jung JY: Antitumor and apoptotic effects of
quercetin on human melanoma cells involving JNK/P38 MAPK signaling
activation. Eur J Pharmacol. 860:1725682019. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Balakrishnan S, Mukherjee S, Das S, Bhat
FA, Raja Singh P, Patra CR and Arunakaran J: Gold
nanoparticles-conjugated quercetin induces apoptosis via inhibition
of EGFR/PI3K/Akt-mediated pathway in breast cancer cell lines
(MCF-7 and MDA-MB-231). Cell Biochem Funct. 35:217–231. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Mina U, Smiti K and Yadav P:
Thermotolerant wheat cultivar (Triticum aestivum L. var.
WR544) response to ozone, EDU, and particulate matter interactive
exposure. Environ Monit Assess. 193:3182021. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Troth EV and Kyle DE: EdU incorporation to
assess cell proliferation and drug susceptibility in Naegleria
fowleri. Antimicrob Agents Chemother. 65:e00017212021. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Wu LW, Chiang YM, Chuang HC, Lo CP, Yang
KY, Wang SY and Shyur LF: A novel polyacetylene significantly
inhibits angiogenesis and promotes apoptosis in human endothelial
cells through activation of the CDK inhibitors and caspase-7.
Planta Med. 73:655–661. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Xu P, Cai X, Zhang W, Li Y, Qiu P, Lu D
and He X: Flavonoids of Rosa roxburghii Tratt exhibit
radioprotection and anti-apoptosis properties via the
Bcl-2(Ca(2+))/Caspase-3/PARP-1 pathway. Apoptosis. 21:1125–1143.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Napso T and Fares F: Zebularine induces
prolonged apoptosis effects via the caspase-3/PARP pathway in head
and neck cancer cells. Int J Oncol. 44:1971–1979. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Ikehata M, Yamada A, Fujita K, Yoshida Y,
Kato T, Sakashita A, Ogata H, Iijima T, Kuroda M, Chikazu D, et al:
Cooperation of Rho family proteins Rac1 and Cdc42 in cartilage
development and calcified tissue formation. Biochem Biophys Res
Commun. 500:525–529. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Sha JY, Li JH, Zhou YD, Yang JY, Liu W,
Jiang S, Wang YP, Zhang R, Di P and Li W: The p53/p21/p16 and
PI3K/Akt signaling pathways are involved in the ameliorative
effects of maltol on D-galactose-induced liver and kidney aging and
injury. Phytother Res ptr. 71422021.PubMed/NCBI
|
|
53
|
Costa BP, Nassr MT, Diz FM, Fernandes KHA,
Antunes GL, Grun LK, Barbé-Tuana FM, Nunes FB, Branchini G and de
Oliveira JR: Methoxyeugenol regulates the p53/p21 pathway and
suppresses human endometrial cancer cell proliferation. J
Ethnopharmacol. 267:1136452021. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Zeng R: Expression of p53, p21, PCNA and
COX-2 and its relationship with recurrence in the early-stage
laryngeal cancer with negative surgical margin. Lin Chung Er Bi Yan
Hou Tou Jing Wai Ke Za Zhi. 30:349–352. 3562016.(In Chinese).
PubMed/NCBI
|
|
55
|
Chiappara G, Gjomarkaj M, Sciarrino S,
Vitulo P, Pipitone L and Pace E: Altered expression of p21,
activated caspase-3, and PCNA in bronchiolar epithelium of smokers
with and without chronic obstructive pulmonary disease. Exp Lung
Res. 40:343–353. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Li P, Liu X, Hao Z, Jia Y, Zhao X, Xie D,
Dong J and Zeng F: Dual Repressive Function by Cip1, a Budding
Yeast Analog of p21, in Cell-Cycle START Regulation. Front
Microbiol. 11:16232020. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Avkiran M and Marber MS: Feeling the
stress: MAPKKK-MAPKK-MAPK signaling cascades in heart failure. J
Mol Cell Cardiol. 48:283–285. 2010. View Article : Google Scholar : PubMed/NCBI
|