|
1
|
Cao W, Chen HD, Yu YW, Li N and Chen WQ:
Changing profiles of cancer burden worldwide and in China: A
secondary analysis of the global cancer statistics 2020. Chin Med J
(Engl). 134:783–791. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Anwanwan D, Singh SK, Singh S, Saikam V
and Singh R: Challenges in liver cancer and possible treatment
approaches. Biochim Biophys Acta Rev Cancer. 1873:1883142020.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kudo M: Systemic therapy for
hepatocellular carcinoma: 2017 update. Oncology. 93 (Suppl
1):135–146. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Shao YY, Hsu CH and Cheng AL: Predictive
biomarkers of sorafenib efficacy in advanced hepatocellular
carcinoma: Are we getting there? World J Gastroenterol.
21:10336–10347. 2015. View Article : Google Scholar
|
|
5
|
Mossenta M, Busato D, Dal Bo M and Toffoli
G: Glucose metabolism and oxidative stress in hepatocellular
carcinoma: Role and possible implications in novel therapeutic
strategies. Cancers (Basel). 12:16682020. View Article : Google Scholar
|
|
6
|
Lai Y, Huang H, Abudoureyimu M, Lin X,
Tian C, Wang T, Chu X and Wang R: Non-coding RNAs: Emerging
regulators of glucose metabolism in hepatocellular carcinoma. Am J
Cancer Res. 10:4066–4084. 2020.PubMed/NCBI
|
|
7
|
Li Z and Zhang H: Reprogramming of
glucose, fatty acid and amino acid metabolism for cancer
progression. Cell Mol Life Sci. 73:377–392. 2016. View Article : Google Scholar
|
|
8
|
Hay N: Reprogramming glucose metabolism in
cancer: Can it be exploited for cancer therapy? Nat Rev Cancer.
16:635–649. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar
|
|
10
|
Lunt SY and Vander Heiden MG: Aerobic
glycolysis: Meeting the metabolic requirements of cell
proliferation. Annu Rev Cell Dev Biol. 27:441–464. 2011. View Article : Google Scholar
|
|
11
|
Warburg O: On the origin of cancer cells.
Science. 123:309–314. 1956. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Praly JP and Vidal S: Inhibition of
glycogen phosphorylase in the context of type 2 diabetes, with
focus on recent inhibitors bound at the active site. Mini Rev Med
Chem. 10:1102–1126. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Agius L: Role of glycogen phosphorylase in
liver glycogen metabolism. Mol Aspects Med. 46:34–45. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Favaro E, Bensaad K, Chong MG, Tennant DA,
Ferguson DJ, Snell C, Steers G, Turley H, Li JL, Günther UL, et al:
Glucose utilization via glycogen phosphorylase sustains
proliferation and prevents premature senescence in cancer cells.
Cell Metab. 16:751–764. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Philips KB, Kurtoglu M, Leung HJ, Liu H,
Gao N, Lehrman MA, Murray TG and Lampidis TJ: Increased sensitivity
to glucose starvation correlates with downregulation of glycogen
phosphorylase isoform PYGB in tumor cell lines resistant to
2-deoxy-D-glucose. Cancer Chemother Pharmacol. 73:349–361. 2014.
View Article : Google Scholar
|
|
16
|
Zhou Y, Jin Z and Wang C: Glycogen
phosphorylase B promotes ovarian cancer progression via
Wnt/β-catenin signaling and is regulated by miR-133a-3p. Biomed
Pharmacother. 120:1094492019. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
de Luna N, Brull A, Guiu JM, Lucia A,
Martin MA, Arenas J, Martí R, Andreu AL and Pinós T: Sodium
valproate increases the brain isoform of glycogen phosphorylase:
Looking for a compensation mechanism in McArdle disease using a
mouse primary skeletal-muscle culture in vitro. Dis Model Mech.
8:467–472. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res.
45(W1):W98–W102. 2017. View Article : Google Scholar
|
|
19
|
Chandrashekar DS, Bashel B, Balasubramanya
SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BVSK and
Varambally S: UALCAN: A portal for facilitating tumor subgroup gene
expression and survival analyses. Neoplasia. 19:649–658. 2017.
View Article : Google Scholar
|
|
20
|
Cancer Genome Atlas Research Network, .
Comprehensive genomic characterization defines human glioblastoma
genes and core pathways. Nature. 455:1061–1068. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Rath VL, Ammirati M, LeMotte PK, Fennell
KF, Mansour MN, Danley DE, Hynes TR, Schulte GK, Wasilko DJ and
Pandit J: Activation of human liver glycogen phosphorylase by
alteration of the secondary structure and packing of the catalytic
core. Mol Cell. 6:139–148. 2000. View Article : Google Scholar
|
|
22
|
Lukacs CM, Oikonomakos NG, Crowther RL,
Hong LN, Kammlott RU, Levin W, Li S, Liu CM, Lucas-McGady D,
Pietranico S and Reik L: The crystal structure of human muscle
glycogen phosphorylase a with bound glucose and AMP: An
intermediate conformation with T-state and R-state features.
Proteins. 63:1123–1126. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Mathieu C, Li de la Sierra-Gallay I, Duval
R, Xu X, Cocaign A, Léger T, Woffendin G, Camadro JM, Etchebest C,
Haouz A, et al: Insights into Brain glycogen metabolism: The
structure of human brain glycogen phosphorylase. J Biol Chem.
291:18072–18083. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar
|
|
25
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
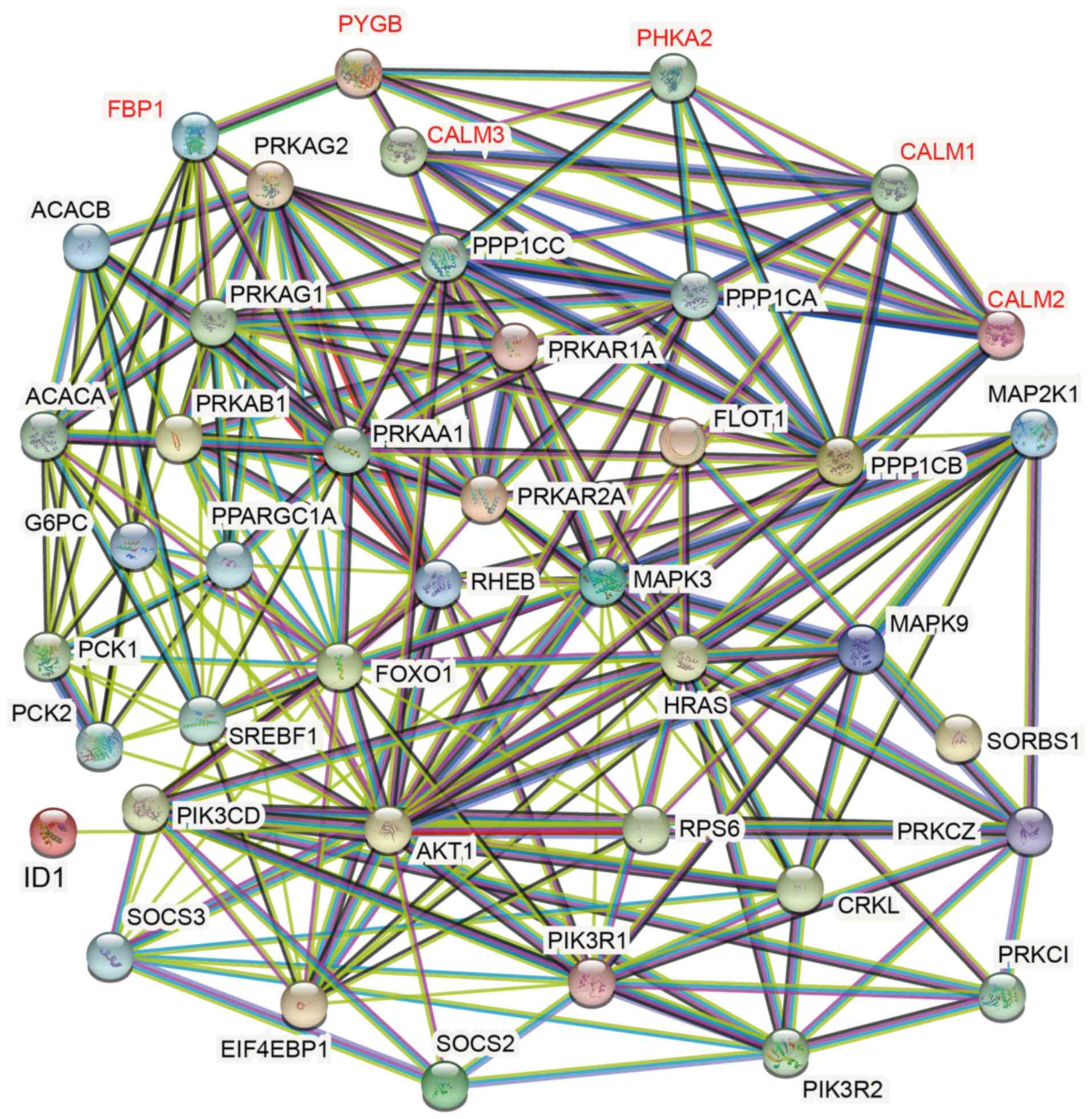
Jensen LJ, Kuhn M, Stark M, Chaffron S,
Creevey C, Muller J, Doerks T, Julien P, Roth A, Simonovic M, et
al: STRING 8-a global view on proteins and their functional
interactions in 630 organisms. Nucleic Acids Res. 37:(Database
issue). D412–D416. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Castven D, Fischer M, Becker D, Heinrich
S, Andersen JB, Strand D, Sprinzl MF, Strand S, Czauderna C,
Heilmann-Heimbach S, et al: Adverse genomic alterations and
stemness features are induced by field cancerization in the
microenvironment of hepatocellular carcinomas. Oncotarget.
8:48688–48700. 2017. View Article : Google Scholar
|
|
28
|
Chin D and Means AR: Calmodulin: A
prototypical calcium sensor. Trends Cell Biol. 10:322–328. 2000.
View Article : Google Scholar
|
|
29
|
Fu J, Wang T and Xiao X: A novel PHKA2
mutation in a Chinese child with glycogen storage disease type IXa:
A case report and literature review. BMC Med Genet. 20:562019.
View Article : Google Scholar
|
|
30
|
Tsang WY, Spektor A, Luciano DJ, Indjeian
VB, Chen Z, Salisbury JL, Sánchez I and Dynlacht BD: CP110
cooperates with two calcium-binding proteins to regulate
cytokinesis and genome stability. Mol Biol Cell. 17:3423–3434.
2006. View Article : Google Scholar
|
|
31
|
Kebede M, Favaloro J, Gunton JE, Laybutt
DR, Shaw M, Wong N, Fam BC, Aston-Mourney K, Rantzau C, Zulli A, et
al: Fructose-1,6-bisphosphatase overexpression in pancreatic
beta-cells results in reduced insulin secretion: A new mechanism
for fat-induced impairment of beta-cell function. Diabetes.
57:1887–1895. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Rocha S, Lucas M, Araujo AN, Corvo ML,
Fernandes E and Freitas M: Optimization and validation of an in
vitro standardized glycogen phosphorylase activity assay.
Molecules. 26:46352021. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Caiola E, Frapolli R, Tomanelli M, Valerio
R, Iezzi A, Garassino MC, Broggini M and Marabese M: Wee1 inhibitor
MK1775 sensitizes KRAS mutated NSCLC cells to sorafenib. Sci Rep.
8:9482018. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Qu Z, Wu J, Wu J, Luo D, Jiang C and Ding
Y: Exosomes derived from HCC cells induce sorafenib resistance in
hepatocellular carcinoma both in vivo and in vitro. J Exp Clin
Cancer Res. 35:1592016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li S, Yang F and Ren X: Immunotherapy for
hepatocellular carcinoma. Drug Discov Ther. 9:363–371. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Lurje I, Czigany Z, Bednarsch J, Roderburg
C, Isfort P, Neumann UP and Lurje G: Treatment strategies for
hepatocellular carcinoma− a multidisciplinary approach.
Int J Mol Sci. 20:14652019. View Article : Google Scholar
|
|
37
|
Luengo A, Gui DY and Vander Heiden MG:
Targeting metabolism for cancer therapy. Cell Chem Biol.
24:1161–1180. 2017. View Article : Google Scholar
|
|
38
|
Bose S, Zhang C and Le A: Glucose
metabolism in cancer: The Warburg effect and beyond. Adv Exp Med
Biol. 1311:3–15. 2021. View Article : Google Scholar
|
|
39
|
Abdel-Wahab AF, Mahmoud W and Al-Harizy
RM: Targeting glucose metabolism to suppress cancer progression:
Prospective of anti-glycolytic cancer therapy. Pharmacol Res.
150:1045112019. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Gomes AS, Ramos H, Soares J and Saraiva L:
p53 and glucose metabolism: An orchestra to be directed in cancer
therapy. Pharmacol Res. 131:75–86. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Ancey PB, Contat C and Meylan E: Glucose
transporters in cancer-from tumor cells to the tumor
microenvironment. FEBS J. 285:2926–2943. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zois CE and Harris AL: Glycogen metabolism
has a key role in the cancer microenvironment and provides new
targets for cancer therapy. J Mol Med (Berl). 94:137–154. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Ritterson Lew C, Guin S and Theodorescu D:
Targeting glycogen metabolism in bladder cancer. Nat Rev Urol.
12:383–391. 2015. View Article : Google Scholar
|
|
44
|
Zois CE, Favaro E and Harris AL: Glycogen
metabolism in cancer. Biochem Pharmacol. 92:3–11. 2014. View Article : Google Scholar
|
|
45
|
Favaro E and Harris AL: Targeting glycogen
metabolism: A novel strategy to inhibit cancer cell growth?
Oncotarget. 4:3–4. 2013. View Article : Google Scholar
|
|
46
|
Jin Y and Yang Y: Bioinformatics-based
discovery of PYGM and TNNC2 as potential biomarkers of head and
neck squamous cell carcinoma. Biosci Rep. 39:BSR201916122019.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Altemus MA, Goo LE, Little AC, Yates JA,
Cheriyan HG, Wu ZF and Merajver SD: Breast cancers utilize hypoxic
glycogen stores via PYGB, the brain isoform of glycogen
phosphorylase, to promote metastatic phenotypes. PLoS One.
14:e02209732019. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Xia B, Zhang K and Liu C: PYGB promoted
tumor progression by regulating Wnt/β-catenin pathway in gastric
cancer. Technol Cancer Res Treat. 19:15330338209265922020.
View Article : Google Scholar
|