Introduction
We reported previously that aldehyde dehydrogenase
(ALDH)1A1, a potential marker of cancer stem cells (CSCs) (1,2), was
related to tumorigenic potential in endometrioid carcinoma (EC)
(3). CSCs refer to tumor cells
that have the ability to self-renew and generate the diverse cells
that comprise the tumor (4,5).
Recent years, CSCs are reported to emerge from non-CSCs, called
tumor cell plasticity (6,7), however, the mechanism has not been
fully elucidated. In this study, we investigated the tumor cell
plasticity from the viewpoint of ALDH1A1 activity.
In EC cell line HEC-1B, ALDH1A1-low population
spontaneously yielded ALDH1A1-high population, mimicking tumor cell
plasticity, and we found that the mixture of ALDH1A1-high
population sometimes accelerated the transition from ALDH1A1-low to
ALDH1A1-high population. We established two distinct HEC-1B
sublines, in which ALDH1A1-high population accelerated such
transition and ALDH1A1-high population did not show such
acceleration. By comparing two sublines, we focused neuronal
membrane glycoprotein M6-b (GPM6B) as the candidate mediating tumor
cell plasticity.
GPM6B is a transmembrane protein that belongs to the
proteolipid protein family. GPM6B is expressed in the central
nervous system (8). GPM6B is
related to the process of neuronal myelination, stabilizes the
axonal membranes, and promotes neuronal differentiation (9–11).
In the functional analysis of GPM6B in tumors, there is only a
report that GPM6B has a cancer-suppressing effect in prostate
cancer (12), and the function of
GPM6B in EC is unknown.
In this study, we investigated the tumor cell
plasticity from the viewpoint of ALDH1A1 activity, and found that
the tumor cell plasticity sometimes accelerated by direct contact
between cancer cells. We focused on GPM6B as the candidate
mediating tumor cell plasticity and started functional analysis of
GPM6B. The knocked-out of GPM6B decreased and its overexpression
increased the expression of ALDH1A1. Thus, it might be suggested
that GPM6B mediated the induction of ALDH1A1 and the plasticity of
CSCs.
Materials and methods
Patients
The study was approved by the Ethical Review Board
of the Graduate School of Medicine, Osaka University (approval no.
15234), and was performed in accordance with the committee
guidelines and regulations. We examined 47 patients undergoing
surgery for EC at Osaka University Hospital from 2011 to 2014. All
patients provided written informed consent. The clinicopathological
features of the enrolled patients were shown in Table I. Resected specimens were fixed in
10% formalin and processed for paraffin embedding. Specimens were
stored at room temperature in a dark room, sectioned at 4 µm
thickness, and subjected to immunohistochemical analysis.
 | Table I.Clinicopathological features of
enrolled cases. |
Table I.
Clinicopathological features of
enrolled cases.
| Characteristic | Number of cases |
|---|
| Histological
grade |
|
| Grade
1 | 24 |
| Grade
2 | 16 |
| Grade
3 | 7 |
|
Total | 47 |
| Clinical stage |
|
| IA | 26 |
| IB | 2 |
| II | 7 |
| III | 12 |
|
Total | 47 |
Cell lines and cell culture
The human EC cell lines HEC-1B and HEC108 were
obtained from the Health Science Research Resources Bank of Osaka,
Japan. Cell lines were cultured in Dulbecco's modified Eagle's
medium (DMEM)-High glucose supplemented with 10% FBS, penicillin
(100 IU/ml), and streptomycin (100 µg/ml) and maintained at 37°C in
5% CO2.
Antibodies and reagents
The antibodies against ALDH1A1 and β-actin were used
as previously reported (13). The
antibody against GPM6B (HPA002913, Sigma-Aldrich) was used for
immunohistochemistry (dilution at 1:200). The antibody against GFP
(#2956, Cell Signaling Technology) was used for immunoblotting
(dilution at 1:500).
Plasmid
The plasmids Empty-EGFP (pRP-EGFP-CMV) and GPM6B
[pRP-EGFP-CAG-FLAG/3×GGGGS/hGPM6B (NM_001001995.3)] were obtained
from Vector Builder, Inc (Chicago, IL, USA).
Flow cytometry
The ALDEFLUOR kit (STEM CELL Technologies) was used
according to the manufacturer's instructions. Cells were analyzed
by FACS CantoII and AriaII flow cytometer (BD Biosciences, Franklin
Lakes, NJ, USA). Data analysis was perfomed using Cell Quest
software (BD Biosciences).
Immunofluorescence
The ALDEFLUOR kit and PKH26 Red Fluorescent Cell
Linker Kit (SIGMA-ALDRICH) were used according to the
manufacturer's protocols. Fluorescence signals were visualized
using fluorescence microscope (BZ-8000, KEYENCE, Osaka, Japan).
Generation of subline cells using
plasma-activated medium
For generation of HEC-1B subline cells, original
cells were treated with threefold diluted plasma-activated medium
(PAM) which was culture medium irradiated by non-thermal plasma
device that containing of gas, electrons, ions, radicals, and
ultraviolet light as previously described (14). PAM induced apoptosis in a large
number of cells due to production of reactive oxygen species (ROS)
such as H2O2. Therefore, we selected the single cell survived from
ROS using microscope. After the selected cell was cultured in
96-well plate, cells proliferated in 6-well plate. The same
procedure was repeated once more to prepare HEC-1B subline
cells.
RNA sequencing analysis
Total RNA was extracted using the miRNeasy Mini Kit
(Qiagen, Hilden, Germany) according to the manufacturer's
instructions. cDNA libraries were constructed using the TruSeq
Stranded mRNA Sample Prep Kit (Illumina, San Diego, CA, USA)
according to the manufacturer's protocol. Sequencing was undertaken
on the Illumina HiSeq 2500 platform in 75-base single-end mode.
Casava version 1.8.2 software (Illumina) was used for base calling.
The sequenced reads were mapped to a human reference genome
sequence (hg19) using TopHat version 2.0.13 (http://ccb.jhu.edu/software/tophat/index.shtml),
Bowtie2 version 2.2.3 (http://bowtie–bio.sourceforge.net/bowtie2/index.shtml),
and SAMtools version 0.1.19 (http://samtools.sourceforge.net/). The fragments per
kilobase of exon per million mapped fragments values were
calculated using Cuffnorm version. 2.2.1 (http://cole–trapnell–lab.github.io/cufflinks/) to
identify upregulated (2.0-fold, P<0.05) and downregulated
(−0.5-fold, P<0.05) genes.
Generation of GPM6B-KO cell lines
GPM6B in HEC-1B was disrupted using the TrueGuide™
CRISPR/Cas9 system (Invitrogen, Carlsbad, CA, USA), in accordance
with the manufacturer's instructions. The crRNA (A35509,
CRISPR714997_CR, sequence: GUGUUGCUCAAGAAUCGCCA, target exon:
exon2, Invitrogen) was annealed with TrueCut™ tracrRNA
(Invitrogen). HEC-1B cells was co-transfected with the gRNA
(crRNA:tracrRNA duplex) and TrueCut™ Cas9 Protein v2 (Invitrogen)
using Lipofectamine™ CRISPRMAX™ Cas9 Transfection Reagent
(Invitrogen). Single cell clones were then isolated by using
limiting dilution cloning in 96-well plates. The positive clones
were confirmed the absence of GPM6B by PCR of genomic sequence.
Untransfected HEC-1B parent cells (WT) was used as a negative
control.
Generation of HEC-1B and HEC108 cells
expressing GPM6B-GFP
The plasmid Empty-GFP was transfected into HEC-1B
and HEC108 cells using Lipofectamine 3000 reagent (Thermo Fisher
Scientific). The plasmid GPM6B-GFP was transfected into HEC-1B
(GPM6B-KO) and HEC108 cells. Green population was sorted with Cell
Sorter SH800ZDP (SONY, Tokyo, Japan). When colonies formed after
passage, we picked up different colonies and named OE1 and OE2.
Untransfected HEC108 parent cells (WT) was used as a negative
control. HEC1B and HEC-108 transfected with empty vector (EV) was
used as another negative control.
Reverse transcription-quantitative PCR
(RT-qPCR)
The RT-qPCR was performed with StepOnePlus™
Real-Time PCR instrument (Applied Biosystems, Foster City, CA)
using Taqman probe/primer sets specific for human GPM6B
(Hs01041077_m1). GAPDH was used as a reference for gene
amplification (Applied Biosystems).
PCR
Cells were rinsed three times with PBS and lysed in
500 µl of lysis buffer [1×SSC (418 µl), 1M Tris-HCl (pH 7.5) (5
µl), 0.5M EDTA (pH 8.0) (1 µl), 10% SDS (50 µl), 20 mg/ml
Proteinase K (FUJIFILM) (25 µl), 10 mg/ml RNase A (Invitrogen) (1
µl)]. The samples were mixed by vortexing and centrifugation at
15,310 g for 10 min at room temperature. After centrifugation at
high speed, the upper phase was carefully removed and transferred
to a new tube. A mixture of Phenol: Chloroform: Isoamyl Alcohol
(Nacalai) was added in equal volumes to samples and the samples
were mixed gently and the aqueous layer was transferred into a new
tube. After ethanol precipitation, precipitated DNA was dissolved
in 30 µl of sterile 1×TE buffer (pH 8.0). Extracted genomic DNA was
amplified with KOD FX Neo (TOYOBO). The following primers which
include protospacer adjacent motif were used to amplify DNA:
(Forward) 5′-CCGTGGCGATTCTTGAGCAAC-3′
(Reverse) 5′-ATGCCCTGGGATCTGCTCTTC-3′
The following primers were used to check the
disrupted alleles:
Human GPM6B:
Exon 2:
(Forward) 5′-ACTGCTCTGCCATTCACTACCCTTCCAG-3′
(Reverse) 5′-ACGCACCACCACGCCCAGCTAAATTTTT-3′.
PCR was done on the T100 Thermal Cycler (Bio-Rad,
USA). The PCR amplification consisted of an initial denaturation
for 2 min at 94°C, followed by 5 cycles of denaturation (10 sec,
98°C) and extension (30 sec, 74°C), 5 cycles of denaturation (10
sec, 98°C) and extension (30 sec, 72°C), 5 cycles of denaturation
(10 sec, 98°C) and extension (30 sec, 70°C), 30 cycles of
denaturation (10 sec, 98°C) and extension (30 sec, 68°C). The final
extension step was carried out at 68°C for 7 min. Its analyzing was
performed using 1.5% agarose gel electrophoresis and visualized
using gel documentation (AE-9000 E-Graph, ATTO, Japan).
Immunohistochemistry
Immunohistochemical staining was conducted by the
Dako Autostainer Link 48 + (Dako/Agilent Technologies, Inc.)
according to the manufacturer's instructions. Primary antibodies
are incubated for 30 min at room temperature.
Immunoblotting
Studies were performed as previously reported
(13). LAS-4000 Image Analyzer (GE
Healthcare, Chicago, IL, USA) or ChemiDoc Touch (Bio-Rad) were used
for the detection of antibody reaction. The expression of β-Actin
was used as a loading control.
Statistical analysis
Statistical analyses were performed using JMP Pro 14
software (SAS Institute). In vitro experiments were
performed at least two times. The data are presented as means ±
standard error of the mean of independent experiments. The
significance of the differences was determined using Mann Whitney U
test. The log-rank test was used for survival analysis.
Kaplan-Meier survival plots were made by using GraphPad Prism 9.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Confirmation of plasticity of cancer
cells
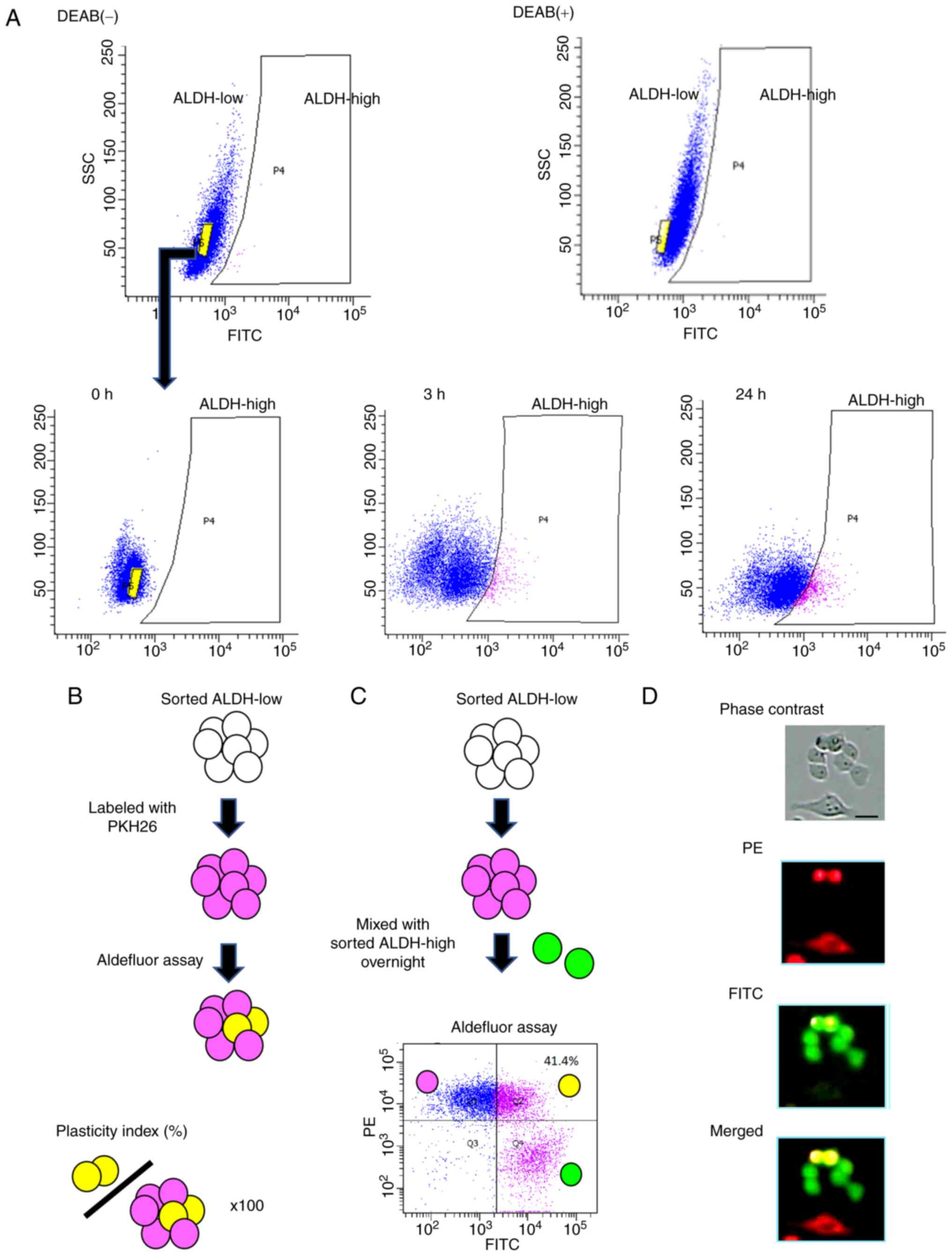
To validate the tumor cell plasticity, we conducted
Aldefluor assay using EC cell line HEC-1B. After culturing
ALDH1A1-low cells, the cell distribution was analyzed by the assay.
ALDH1A1-low population spontaneously yielded ALDH1A1-high
population (Fig. 1A). Next, we
stained ALDH1A1-low cells red using the PKH26 Red Fluorescent Cell
Linker Kit and conducted Aldefluor assay. The percentage of cells
that turned yellow in the total cell number was defined as
Plasticity index (Fig. 1B).
Furthermore, we found that the mixture of ALDH1A1-high population
sometimes accelerated the plasticity index and observed changes in
tumor cells with low ALDH1A1 activity under a fluorescence
microscope (Fig. 1C and D). Many
color changes were observed in the adhesive areas between tumor
cells (Fig. 1D).
Direct contact with cancer cells
promotes tumor plasticity
We speculated that physical contact between cells
contributed to the activation of ALDH1A1. First, we generate some
sublines of HEC-1B cells using PAM and co-cultured ALDH1A1-high
cells and ALDH1A1-low cells by a method of culturing two types of
cells in the same dish and a method of culturing the two types of
cells so that they do not contact with each other via transwell
(Permeable Supports 3.0 µm Polycarbonate membrane, Corning)
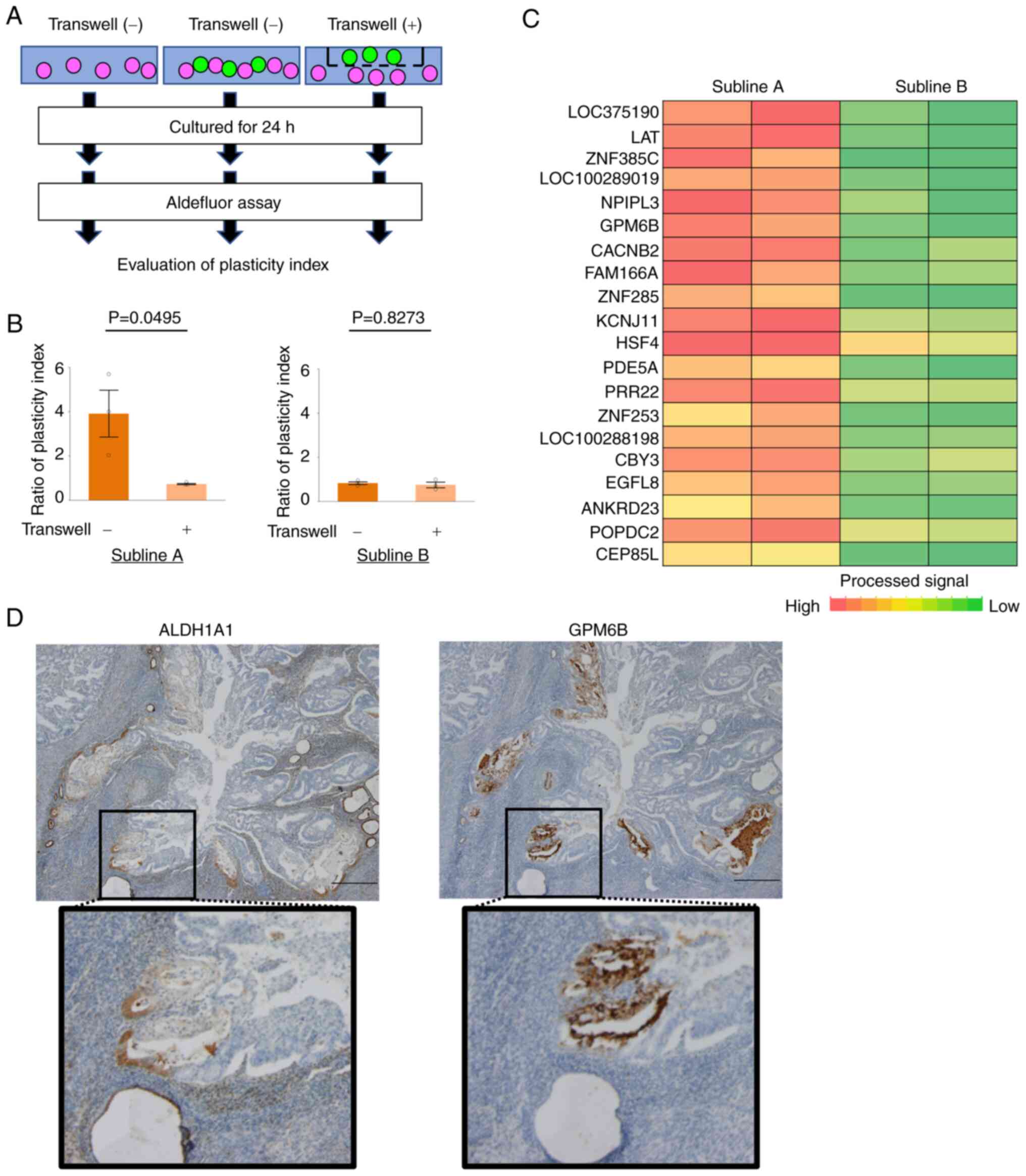
(Fig. 2A). In subline A, the
former method had a higher plasticity index than the latter method,
however, there was no difference in subline B (Fig. 2B). And then, we extracted RNAs of
subline A and B. We compared the RNA expression of both cells and
focused on GPM6B as the candidate mediating tumor cell plasticity
(Fig. 2C; Tables II and SI). Next, we performed
immunohistochemistry analysis of ALDH1A1 and GPM6B in clinical
samples of EC tissues and found that GPM6B tended to express in the
border of ALDH1A1 expressing tumor cells and non-expressing tumor
cells (Fig. 2D).
 | Table II.Top 20 genes upregulated in subline
A. |
Table II.
Top 20 genes upregulated in subline
A.
| Fold change | P-value | Gene symbol | Description |
|---|
| 4.003 | 0.049 | LOC375190 | N/A |
| 3.585 | 0.009 | LAT | Linker for activation
of T cells |
| 3.399 | 0.023 | ZNF385C | Zinc finger protein
385C |
| 3.322 | 0.050 | LOC100289019 | Uncharacterized
LOC100289019 |
| 3.268 | 0.039 | NPIPL3 | Nuclear pore complex
interacting protein-like 3 |
| 3.091 | 0.019 | GPM6B | Neuronal membrane
glycoprotein M6B |
| 2.856 | 0.012 | CACNB2 | Calcium channel,
voltage-dependent, beta 2 subunit |
| 2.665 | 0.040 | FAM166A | Family with sequence
similarity 166, member A |
| 2.593 | 0.005 | ZNF285 | Zinc finger protein
285 |
| 2.589 | 0.008 | KCNJ11 | Potassium
inwardly-rectifying channel, subfamily J, member 11 |
| 2.407 | 0.024 | HSF4 | Heat shock
transcription factor 4 |
| 2.362 | 0.021 | PDE5A | Phosphodiesterase
5A, cgmp-specific |
| 2.285 | 0.005 | PRR22 | Proline rich
22 |
| 2.278 | 0.030 | ZNF253 | Zinc finger protein
253 |
| 2.240 | 0.004 | LOC100288198 | Uncharacterized
LOC100288198 |
| 2.179 | 0.008 | CBY3 | Chibby homolog 3
(Drosophila) |
| 2.171 | 0.015 | EGFL8 | EGF-like-domain,
multiple 8 |
| 2.073 | 0.027 | ANKRD23 | Ankyrin repeat
domain 23 |
| 2.067 | 0.013 | POPDC2 | Popeye domain
containing 2 |
| 2.028 | 0.008 | CEP85L | Centrosomal protein
85kda-like |
GPM6B promotes ALDH1A1 expression in
EC cell lines
To examine the significance of GPM6B in EC, we
disrupted the GPM6B gene in HEC-1B cells, using the CRISPR/Cas9
system and successfully established GPM6B-knockout (KO) HEC-1B
cells (Fig. 3A and B).
Furthermore, we showed the transfection efficiency (Fig. S1A), and we constructed HEC-1B
cells (GPM6B-KO) stably expressing GPM6B (Figs. 3C and S1B). GPM6B knockout in HEC-1B cells
resulted in a decreased level of ALDH1A1 expression and GPM6B
expressing HEC-1B cells (GPM6B-KO) resulted in an increased level
of ALDH1A1 (Fig. 3D and Table SII). Similarly, we constructed
GPM6B-expressing HEC108 cells (OE1 and OE2) (Figs. 3E and S1B). GPM6B expressing HEC108 cells
resulted in an increased level of ALDH1A1 (Fig. 3F).
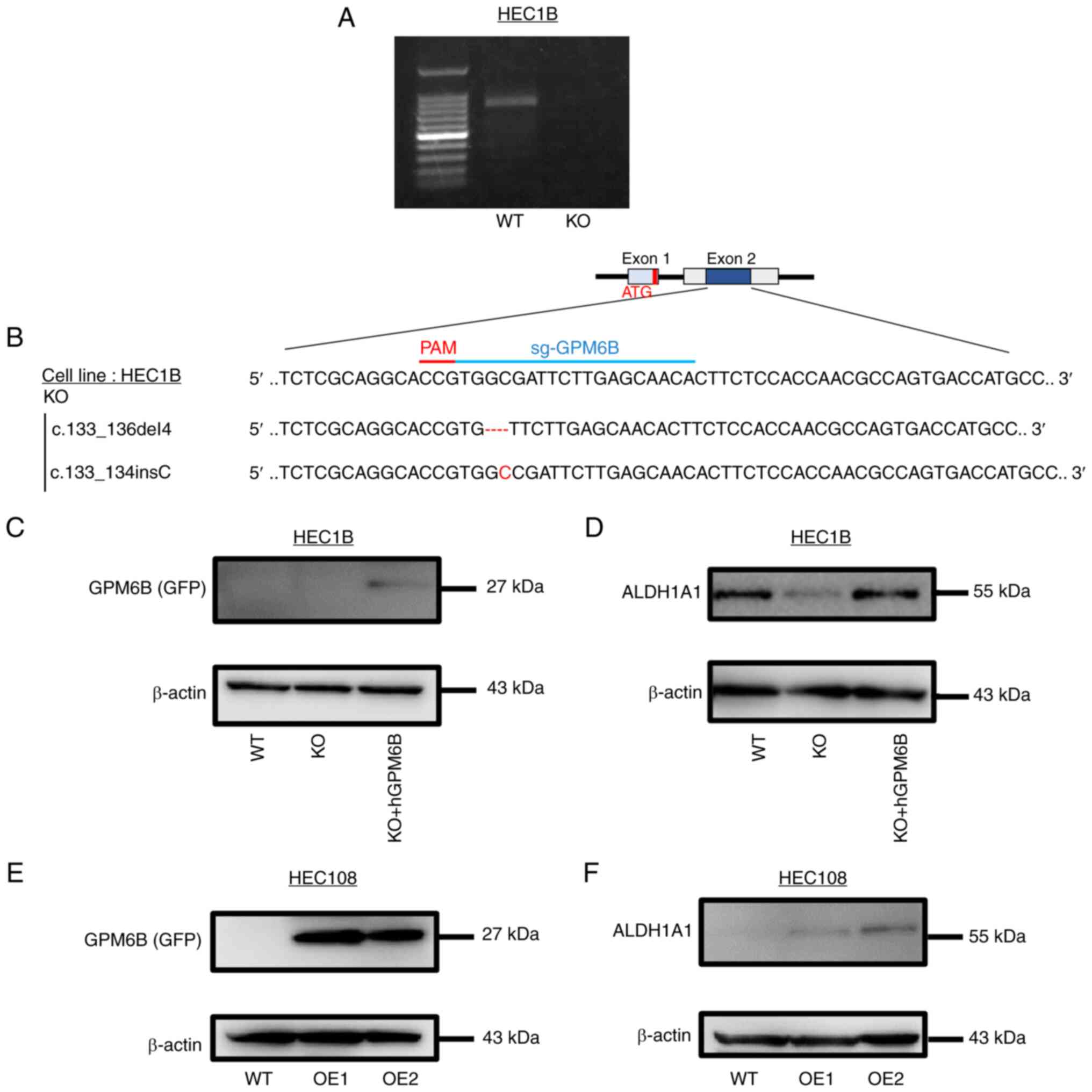 | Figure 3.GPM6B promotes ALDH1A1 expression in
EC cell lines. (A) PCR of GPM6B in parent cells (WT) and GPM6B-KO
HEC-1B cells. (B) A schematic of the single-guide (sg)
RNA-targeting sites in the human GPM6B gene. The guide sequence of
the human GPM6B gene was targeting exon 2 and it was
5′-TGGCGATTCTTGAGCAACAC-3. Targeting site and protospacer adjacent
motif (PAM) are indicated as red-colored bar. Sequence alignments
of the wild-type GPM6B gene and the disrupted alleles from GPM6B-KO
clone are shown. Deleted regions are indicated with red-colored
dashes and insertion parts are shown in red. (C) Immunoblotting of
GPM6B(GFP) in WT, GPM6B-KO and GPM6B-KO + hGPM6B HEC-1B cells. (D)
Immunoblotting of ALDH1A1 in HEC-1B cells (WT, KO, KO + hGPM6B) (E)
Immunoblotting of GPM6B in WT, GPM6B-OE1 and GPM6B-OE2 HEC108
cells. (F) Immunoblotting of ALDH1A1 in HEC108 cells (WT,
GPM6B-OE1, GPM6B-OE2). ALDH1A1, aldehyde dehydrogenase 1 family
member A1; GPM6B, neuronal membrane glycoprotein M6-b; KO,
knockout; OE, overexpressing; WT, wild type. |
High GPM6B is related with poor
prognosis
Due to a limited number of enrolled cases in this
study, we examined the effect of high GPM6B on prognosis with three
kinds of publicly available datasets; gene expression profiling
interactive analysis (GEPIA)2, human protein atlas, and the cancer
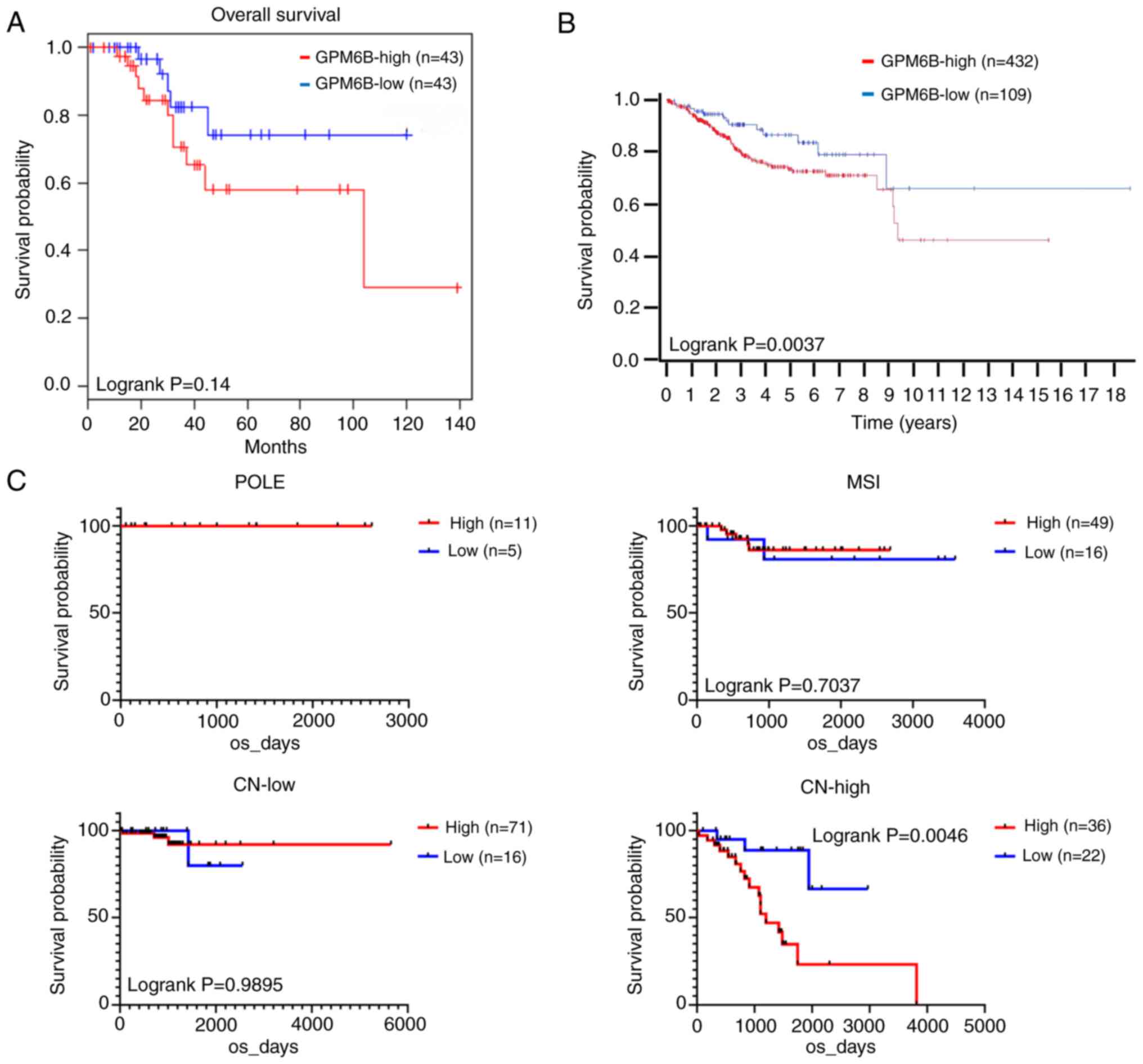
genome atlas (TCGA). In GEPIA2 of uterine corpus endometrial
carcinoma, high GPM6B expression was marginally correlated with
poor overall survival of the patients (Fig. 4A), in which group cutoff was set as
quartile (http://gepia2.cancer-pku.cn/#survival). The data of
human protein atlas revealed high GPM6B expression was correlated
with poor overall survival of endometrial carcinoma (https://www.proteinatlas.org/ENSG00000046653–GPM6B/pathology/endometrial+cancer),
in which 432 cases of GPM6B-high and 109 cases of GPM6B-low were
enrolled and the best expression cut-off score 1.94 was applied
(Fig. 4B). The median follow-up
time of human protein atlas was 2.5 years and the p-value was
0.037. Lastly, we examined TCGA database, in which the enrolled
endometrial carcinoma cases were divided into recently published
classification (15); POLE type
(ultramutated) (POLE), microsatellite instability (MSI), copy
number low (CN-low), and copy number high (CN-high). Sixteen cases
of POLE, 65 cases of MSI, 87 cases of CN-low and 58 cases of
CN-high were examined in TCGA database. When cutoff was set as
quartile, patients with high GPM6B expression had significantly
shorter OS compared with those with low GPM6B expression in CN-high
group but not in other groups (Fig.
4C).
Discussion
We identified a new function of GPM6B in EC cells.
GPM6B contributed to the tumor cell plasticity in EC. We reported
previously that a potential marker of CSCs; ALDH1A1 was related to
tumorigenic potential in EC (3).
The ALDH1A1-high cells are thought to be target for treatment.
However, the tumor cell plasticity is reported to occur (6,7) and
we should target the ALDH1A1-low cells which have a potential to
become ALDH1A1-high cell.
To find the mechanism of the tumor cell plasticity,
we focused on the direct contact between tumor cells. We collected
some subclones from HEC-1B cells and tested the plasticity index
according to Figs. 1C and 2A. We revealed that the biological
ramification of direct cell-cell contact between tumor cells was
one of triggers for the tumor cell plasticity. We extracted RNAs of
cells with high/low plasticity index and compared the RNA
expression of both cells. Among membrane proteins GPM6B was only
high expression level in cells with high plasticity index.
GPM6B is less understood for cancer research and
recently reported to work as a tumor suppressor in prostate cancer
(12). To examine the significance
of GPM6B in EC, we generated GPM6B-KO and GPM6B-expressing cells.
We found that ALDH1A1 expression was regulated by GPM6B. Thus, we
revealed another aspect of GPM6B for cancer. Notably, GPM6B was
expressed in the border of ALDH1A1 expressing tumor cells and
non-expressing tumor cells in clinical samples of EC. That is,
GPM6B is not expressed in ALDH1A1 high expression area. GPM6B might
be necessary for only increasing ALDH1A1. Further investigation is
needed to uncover the mechanism.
In prognostic analysis, GPM6B was a prognostic
factor. Though ALDH1A1 was a prognostic factor, the gene regulating
tumor cell plasticity was also prognostic factor. It might be
important to target not only CSCs but also non-CSCs which have a
potential to become CSCs for cure EC. Moreover, high GPM6B
expression may be more effective as prognostic factor when the EC
cases are classified to CN-high. CN-high is a worse prognostic
classification as compared to POLE, MSI and CN-low. Factors
enhancing plasticity of non-CSCs to CSCs might play important roles
in prognosis.
In summary, direct cell-cell contact between tumor
cells influenced on the tumor cell plasticity. GPM6B regulated
ALDH1A1 expression. Furthermore, GPM6B was also a prognostic
factor. These results suggest that GPM6B mediated the induction of
ALDH1A1 and we have to consider tumor cell plasticity to cure
EC.
Supplementary Material
Supporting Data
Supporting Data
Supporting Data
Acknowledgements
The authors would like to thank Mr. Masaharu Kohara,
Ms. Megumi Nihei, Ms. Etsuko Maeno Fujinami and Ms. Takako Sawamura
from Department of Pathology, Osaka University Graduate School of
Medicine for their technical assistance.
Funding
This work was supported by JSPS KAKENHI (grant nos. A19H034520,
A22J207990, 16K08649 and 21K06881) and by AMED (grant nos.
JP21ae0121049).
Availability of data and materials
The datasets used and/or analyzed during this study
are available acquired with the permission of the corresponding
author. The sequencing datasets generated and/or analyzed during
the current study are available in the Gene Expression Omnibus
repository under accession number GSE212889 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE212889).
Authors' contributions
SK, JII and EM were involved with conception and
design. SK, JII, DO and EM analyzed and interpretated the data. SK,
JII, EMF, MK, ST, TM, SN and EM performed experiments and analyzed
data. SK, JII and EM wrote the manuscript. SK and EM confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript, and are accountable for all aspects of the
work.
Ethics approval and consent to
participate
The study was approved by the Ethical Review Board
of the Graduate School of Medicine, Osaka University (grant no.
15234), and was performed in accordance with the committee
guidelines and regulations. All patients provided written informed
consent to participate in the study.
Patient consent for publication
Patient consent for publication was covered by the
informed consent document.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
EC
|
endometrioid carcinoma
|
|
ALDH1A1
|
aldehyde dehydrogenase 1 family member
A1
|
|
GPM6B
|
neuronal membrane glycoprotein
M6-b
|
References
|
1
|
Ginestier C, Hur MH, Charafe-Jauffret E,
Monville F, Dutcher J, Brown M, Jacquemier J, Viens P, Kleer CG,
Liu S, et al: ALDH1 is a marker of normal and malignant human
mammary stem cells and a predictor of poor clinical outcome. Cell
Stem Cell. 1:555–567. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Jiang F, Qiu Q, Khanna A, Todd NW, Deepak
J, Xing L, Wang H, Liu Z, Su Y, Stass SA and Katz RL: Aldehyde
dehydrogenase 1 is a tumor stem cell-associated marker in lung
cancer. Mol Cancer Res. 7:330–338. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Rahadiani N, Ikeda J, Mamat S, Matsuzaki
S, Ueda Y, Umehara R, Tian T, Wang Y, Enomoto T, Kimura T, et al:
Expression of aldehyde dehydrogenase 1 (ALDH1) in endometrioid
adenocarcinoma and its clinical implications. Cancer Sci.
102:903–908. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Visvader JE and Lindeman GJ: Cancer stem
cells in solid tumours: Accumulating evidence and unresolved
questions. Nat Rev Cancer. 8:755–768. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Merrell AJ and Stanger BZ: Adult cell
plasticity in vivo: De-differentiation and transdifferentiation are
back in style. Nat Rev Mol Cell Biol. 17:413–425. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yuan S, Norgard RJ and Stanger BZ:
Cellular plasticity in cancer. Cancer Discov. 9:837–851. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Yan Y, Narayanan V and Lagenaur C:
Expression of members of the proteolipid protein gene family in the
developing murine central nervous system. J Comp Neurol.
370:465–478. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bang ML, Vainshtein A, Yang HJ,
Eshed-Eisenbach Y, Devaux J, Werner HB and Peles E: Glial M6B
stabilizes the axonal membrane at peripheral nodes of Ranvier.
Glia. 66:801–812. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Mita S, de Monasterio-Schrader P,
Fünfschilling U, Kawasaki T, Mizuno H, Iwasato T, Nave KA, Werner
HB and Hirata T: Transcallosal projections require glycoprotein
M6-dependent neurite growth and guidance. Cereb Cortex.
25:4111–4125. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Werner HB, Krämer-Albers EM, Strenzke N,
Saher G, Tenzer S, Ohno-Iwashita Y, De Monasterio-Schrader P,
Möbius W, Moser T, Griffiths IR and Nave KA: A critical role for
the cholesterol-associated proteolipids PLP and M6B in myelination
of the central nervous system. Glia. 61:567–586. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
He S, Huang Z, Li X, Ding Y, Sheng H, Liu
B and Jia Z: GPM6B inhibit PCa proliferation by blocking prostate
cancer cell serotonin absorptive capacity. Dis Markers.
2020:88107562020. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kusumoto S, Kurashige M, Ohshima K, Tahara
S, Matsui T, Nojima S, Hattori S and Morii E: An immature
inhibin-α-expressing subpopulation of ovarian clear cell carcinoma
cells is related to an unfavorable prognosis. Cancer Med.
10:1485–1500. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ikeda JI, Tanaka H, Ishikawa K, Sakakita
H, Ikehara Y and Hori M: Plasma-activated medium (PAM) kills human
cancer-initiating cells. Pathol Int. 68:23–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Kandoth C, Schultz N, Cherniack AD, Akbani
R, Liu Y, Shen H, Robertson AG, Pashtan I, Shen R, Benz CC, et al:
Integrated genomic characterization of endometrial carcinoma.
Nature. 497:67–73. 2013. View Article : Google Scholar : PubMed/NCBI
|