Introduction
Oral squamous cell carcinoma (OSCC) represents a
predominant malignant neoplasm worldwide, accounting for ~90% of
oral malignancies and markedly impacting both physiological
function and aesthetic appearance. Data from the Global Cancer
Observatory (1) indicates that the
incidence of OSCC is expected to increase by ~40% by 2040, with a
corresponding rise in mortality rates (1). Despite advancements in therapeutic
approaches and diagnostic methods, the 5-year overall survival rate
of OSCC has shown only modest improvements in recent years,
particularly in cases where distant metastasis is detected at
diagnosis (2). Moreover, it has
been reported that less than 1/2 patients survive >5 years,
partly due to the incomplete understanding of the molecular
mechanisms underlying the disease and the notable challenge of drug
resistance (3).
In the field of oncology, RNA chemical
modifications, particularly N6-methyladenosine (m6A)
modification, have emerged as key regulators of gene expression
dynamics and neoplastic evolution (4,5). These
modifications are predominantly identified on messenger RNA (mRNA)
and ribosomal RNA (rRNA), and they have notable effects on RNA
stability, processing, translation and decay (6,7).
Notably, methyltransferase 5, N6-adenosine (METTL5) has been
identified as a principal m6A-modifying enzyme,
responsible for orchestrating m6A modification at the
A1832 locus of 18S rRNA. This specific modification serves a key
role in ribosomal biogenesis and function, thereby exerting a
notable influence on an extensive range of physiological and
pathological processes, including the initiation and progression of
cancer (8–13).
Preliminary investigations into METTL5 across a
spectrum of malignancies have highlighted its complex involvement
through several mechanistic pathways (14–16).
Notably, a previous study has reported that METTL5/transfer RNA
methyltransferase activator subunit 11-2-mediated m6A
modification at the A1832 site of 18S rRNA is upregulated in
nasopharyngeal carcinoma, fostering tumorigenesis both in
vitro and in vivo (17).
Mechanistically, METTL5 contributes to carcinogenesis by promoting
the assembly of 80S ribosomes, which in turn enhances the
translation of mRNA with a 5′-terminal oligopyrimidine structure.
Furthermore, METTL5 enhances the assembly of 80S ribosomes, which
in turn activates the transcription of heat shock factor 4b. This
process hinders the degradation of mutant p53 protein, thereby
promoting nasopharyngeal carcinoma tumorigenesis and contributing
to chemotherapy resistance (17).
In renal cell carcinoma, increased METTL5 expression is associated
with immune dysregulation within the tumor microenvironment. This
may influence tumor progression by modulating immune-related
pathways, such as the differentiation of T-helper (Th)17 and
Th1/Th2 cells, as well as the activity of phosphatidylinositol
3-kinase (18). Furthermore, METTL5
has been identified as a key factor in determining the composition
of the tumor microenvironment and the infiltration of immune cells.
This is particularly evident in lung adenocarcinoma, where
increased METTL5 expression has been associated with unfavorable
patient outcomes. The influence of METTL5 in this context is likely
due to its ability to modulate immune pathways, which can promote
tumor growth and spread by shaping the immune microenvironment
(19). Furthermore, in
hepatocellular carcinoma, the upregulation of METTL5 has been
reported to drive metabolic reprogramming, notably in glucose
metabolism pathways, which in turn promotes cancer cell
proliferation and metastasis. This metabolic reconfiguration is
likely facilitated by the upregulation of key enzymes, including
lactate dehydrogenase A, enolase 1, triosephosphate isomerase 1,
solute carrier family 2 member 1 and pyruvate kinase M2.
Furthermore, METTL5 exerts regulatory control over c-Myc protein
stability, which in turn can further modulate tumor cell
proliferation (20,21). In breast cancer, augmented METTL5
expression is associated with a more aggressive tumor behavior and
reduced survival rates compared with low METTL5 expression.
Mechanistically, METTL5 enhances translational capacity and protein
synthesis within breast cancer cell lines, thus promoting cancer
cell proliferation and survival (22). However, current investigations on
the role and mechanistic details of METTL5 in OSCC is still in its
early stages. Building upon the pivotal role of METTL5 in diverse
types of malignancies, we hypothesize that it may also serve a key
role in the pathogenesis and progression of OSCC.
OSCC is a major head and neck malignancy with
unclear molecular mechanisms underlying its progression. METTL5 is
associated with cancer development, but its role and mechanisms in
OSCC remain unknown. The present study aimed to explore the
clinical significance of METTL5 in patients with OSCC, its
functional role in tumorigenesis and progression, and the
underlying molecular mechanisms. The present study analyzed the
association of METTL5 with clinical outcomes, conducted in
vitro/in vivo experiments to assess its impact on OSCC cell
proliferation, metastasis and tumor growth, and used RNA-seq and
bioinformatics to identify its regulated pathways and targets. The
present study aims to fill the knowledge gap of METTL5 in OSCC and
lay a theoretical foundation for potential therapeutic target
discovery.
Materials and methods
Patient samples
All clinical samples used in the present study were
collected from The First Affiliated Hospital of Sun Yat-sen
University (Guangzhou, China). Prior to sample collection, written
informed consent was obtained from all the patients. All
patient-related studies were reviewed and approved by the
Independent Ethics Committee for Clinical Research and Animal
Trials of The First Affiliated Hospital of Sun Yat-sen University
[approval no. (2022)229], and performed in accordance with the
Declaration of Helsinki. Inclusion criteria for the present study
were as follows: i) Pathologically confirmed primary OSCC; ii) no
neoadjuvant therapy; iii) complete clinical/follow-up data; and iv)
an age of ≥18 years with normal liver/kidney function. Exclusion
criteria were as follows: i) Secondary OSCC/synchronous
malignancies; ii) history of other cancer types (5 years); iii)
severe systemic diseases; and iv) incomplete data. Samples (n=4)
were collected from August to October 2022. Table SI presents the demographic
information and tumor characteristics of patients in the present
study.
Cell culture and lentiviral
transduction/transfection
The HSC3 cell line was purchased from the Japanese
Collection of Research Bioresources Cell Bank (cat. no. JCRB0623).
SCC15 cells were purchased from the American Type Culture
Collection (cat. no. CRL-1623). The Human Oral Keratinocytes (HOK)
cell line was purchased from ScienCell Research Laboratories, Inc.
(cat. no. 2610). Cells were maintained in a humidified incubator
with 5% CO2 at 37°C and cultured in DMEM/F-12 medium
(cat. no. C11330500BT) supplemented with 1% penicillin/streptomycin
(cat. no. 15140-122; Gibco; Thermo Fisher Scientific, Inc.) and 10%
FBS(Gibco; Thermo Fisher Scientific, Inc.).
Both METTL5 knockout and overexpression systems were
constructed for functional assays: Knockout system
(lentivirus-mediated): The fourth-generation CRISPR plasmid system
was used, including packaging plasmids (TAT, RAII, HEMP2 and VSVG)
and target plasmids (LentiCRISPRv2 [negative control, NC group],
lentiCRISPRv2-METTL5-human-sg1, lentiCRISPRv2-METTL5-human-sg2
[target groups]; purchased from Guangzhou IGE Biotechnology Co.,
Ltd.]). The interim cell line for lentiviral packaging was 293T
cells (purchased from the American Type Culture Collection), which
were used to produce viral particles.
Overexpression system (transient transfection): The
overexpression plasmid (pFLAG-CMV2-METTL5/pFLAG-CMV2-CCND3;
overexpression, OE) and empty vector (pFLAG-CMV2; negative control,
NC) were purchased from Guangzhou IGE Biotechnology Co., Ltd.), and
introduced into cells via transient transfection (no lentiviral
packaging required).
Lentiviral transfection (for knockout) was performed
as follows: 1×106 293T cells were seeded in T25 flasks 1
day prior to transfection at 37°C and cultured until reaching 60%
confluence. The transfection mixture was prepared by combining the
packaging plasmids (0.5 µg TAT, 0.5 µg RAII, 0.5 µg HEMP2, 0.5 µg
VSVG) and target plasmids (2 µg per construct) with 200 µl Opti-MEM
medium (Gibco; Thermo Fisher Scientific, Inc.). Separately,
Lipofectamine 2000 (Invitrogen; Thermo Fisher Scientific, Inc.) was
gently mixed with 200 µl Opti-MEM and incubated at room temperature
for 5 min. The plasmid mixture and Lipofectamine 2000 solution were
then combined, gently mixed, and incubated at room temperature for
20 min (plasmid: Lipofectamine 2000 ratio=3.5 µg: 7 µl). The
complete medium of 293T cells was aspirated and replaced with 5 ml
antibiotic-free DMEM containing 10% FBS, followed by the addition
of the entire plasmid-Lipofectamine 2000 complex. Cells were
incubated at 37°C in a 5% CO2 incubator for 12–24 h
before medium replacement.
Lentiviral particles were collected every 24 h for
2–3 consecutive days. The collected medium was centrifuged at ~450
× g (2,000 rpm) for 5 min at 4°C, and the viral supernatant was
aliquoted into sterile 1.5 ml EP tubes and stored at −80°C. For
infection of target OSCC cells (HSC3 and SCC15), cells were seeded
in T25 flasks and cultured until 60% confluence. Lentiviral
particles were added at a multiplicity of infection (MOI) of 10
along with polybrene (final concentration: 8 µg/ml) to enhance
transduction efficiency. Cells were incubated at 37°C in a 5%
CO2 incubator for 24–48 h.
Following transduction, the selection method was
antibiotic selection using puromycin; the selection concentration
was 2.5–3 µg/ml, and the maintenance concentration was 1.0–1.5
µg/ml. Cells were cultured for >48 h until all untransfected
cells died. After selection, cells were maintained in medium
containing the maintenance concentration of puromycin. The
transduced cells were then passaged, and protein extraction was
performed for western blot analysis to verify the gene knockout
efficiency of METTL5. The time interval between transduction and
subsequent experimentation was 48–72 h after successful
transduction and selection.
Transient transfection (for METTL5 overexpression):
OSCC cells (HSC3 and SCC15) were seeded in 6-well plates and
cultured until 60% confluence. The transfection mixture was
prepared by combining 4 µg of pFLAG-CMV2-METTL5/pFLAG-CMV2-CCND3
(or pFLAG-CMV2) with 250 µl Opti-MEM medium, then mixed with 5 µl
Lipofectamine 2000 (incubated at room temperature for 20 min). The
mixture was added to cells in antibiotic-free medium, and cells
were incubated at 37°C for 48 h. The time interval between
transfection and subsequent experimentation was 48 h. Western blot
analysis was performed to verify the METTL5 overexpression
level.
CCND3 overexpression in METTL5-depleted HSC3 cells:
METTL5-depleted HSC3 cells (sgMETTL5 HSC3 cells) were seeded in
6-well plates and cultured until 60% confluence. The CCND3
overexpression plasmid (pFLAG-CMV2-CCND3) and empty vector
(pFLAG-CMV2; negative control) were introduced via transient
transfection: 4 µg of plasmid (or empty vector) was mixed with 250
µl Opti-MEM medium, followed by 5 µl Lipofectamine 2000. The
mixture was incubated at room temperature for 20 min, then added to
cells in antibiotic-free medium. Cells were incubated at 37°C (5%
CO2) for 48 h. The time interval between transfection
and subsequent experimentation was 48 h. Western blot analysis was
performed to verify CCND3 overexpression efficiency before
subsequent functional assays. The sgRNA sequences used in the
present study are listed in Table
SII.
Immunohistochemistry (IHC)
staining
Tissues from the mouse model were first fixed in 4%
paraformaldehyde (PFA) (w/v in phosphate-buffered saline, PBS) at
4°C for 24 h. Fixed tissues were embedded in paraffin resin, and
serial sections were cut into 4 µm-thick slices using a microtome.
For dewaxing and rehydration, sections were processed through a
descending alcohol series: xylene (twice for 10 min each), 100%
ethanol (twice for 5 min each), 95% ethanol (5 min), 80% ethanol (5
min), 70% ethanol (5 min), and finally rinsed with distilled water
for 5 min. Antigen retrieval was performed by immersing sections in
0.01 M citrate buffer (pH 6.0) and heating in a microwave oven at
95–100°C for 15 min. After natural cooling to room temperature,
sections were washed with PBS (3 times for 5 min each).
To block endogenous peroxidase activity, sections
were treated with 3% hydrogen peroxide at room temperature for 10
min, followed by three washes with PBS (5 min each). Non-specific
binding sites were blocked using 5% BSA (w/v in PBS; cat. no.
4240GR100; Saiguo Biotechnology Co., Ltd.) at room temperature for
30 min. Subsequently, sections were incubated overnight with
primary antibodies at 4°C: Anti-Ki-67 and anti-METTL5. After three
washes with PBS (5 min each), sections were incubated with a
horseradish peroxidase (HRP)-conjugated goat anti-rabbit secondary
antibody at room temperature for 60 min. Details of all primary and
secondary antibodies are in Table
SIII.
Immunoreactivity was visualized using a DAB
detection kit (cat. no. GK6007; Beijing Genetech Co., Ltd.) as the
chromogen substrate. Sections were incubated with DAB working
solution at room temperature for 3–5 min (monitoring staining
intensity under a light microscope to avoid over-staining). The
reaction was terminated by rinsing with distilled water. Sections
were then counterstained with hematoxylin at room temperature for 2
min, differentiated with 1% hydrochloric acid in ethanol for 30
sec, and dehydrated through an ascending alcohol series (70, 80, 95
and 100% ethanol for 5 min each) followed by xylene (twice for 10
min each). Finally, sections were mounted with neutral gum and a
coverslip was applied.
Stained sections were imaged using a Leica inverted
light microscope (Leica Microsystems) at magnifications of 400×;
scale bar, 50 µm. IHC scoring was performed based on a
semiquantitative immunoreactivity scoring system (4). Briefly, the proportion of positive
staining cells was scored as follows: 0 (absent), 1 (<10%), 2
(10–50%), and 3 (>50%). Staining intensity was scored as: 0
(negative), 1 (weak), 2 (moderate), and 3 (strong). The final IHC
score was calculated by multiplying the percentage score by the
intensity score, resulting in a range of 0–9. Image analysis and
scoring were performed using ImageJ software (Version 1.53;
National Institutes of Health).
Western blotting
Total protein was extracted from OSCC cells
(HSC3/SCC15), HOK and human tissues using RIPA buffer (cat. no.
C1053; Applygen Technologies, Inc.). Protein concentration was
determined via the BCA protein assay kit (cat. no. P0012; Beyotime
Biotechnology) following the manufacturer's instructions.
Equal amounts of protein (20 µg per lane) were
loaded onto 10% SDS-PAGE gels for separation, then transferred onto
a PVDF membrane (cat. no. IPVH00010; Merck KGaA). The membrane was
blocked with 5% non-fat skimmed milk (dissolved in TBS-Tween; TBST)
for 2 h at room temperature; TBST buffer contained 0.1% Tween-20
(v/v) in Tris-buffered saline.
Subsequently, the membrane was incubated with
primary antibodies (listed in Table
SIII) at 4°C overnight. After three washes with TBST (10 min
each), the membrane was incubated with species-matched secondary
antibodies (HRP-conjugated) at room temperature for 1 h.
Protein bands were detected using a super-sensitive
ECL kit (cat. no. P0018FS; Beyotime Biotechnology). The membrane
was incubated with the ECL working solution (1:1 mixture of reagent
A and B) for 5 min at room temperature, then visualized using the
Tanon 5200 Multi Intelligent Imaging System (Tanon Science and
Technology Co., Ltd.). Band intensities were quantified using
ImageJ software (Version 1.53; National Institutes of Health) for
densitometric analysis.
Cell Counting Kit-8 (CCK-8)
Cell viability was monitored using a CCK-8 kit (cat.
no. CK04; Dojindo Laboratories, Inc.). A total of 1×103
cells (HSC3 or SCC15) per well were plated onto 96-well plates and
cultured in DMEM/F-12 with 1% FBS. CCK-8 working solution was then
applied to the cells for 2 h, according to the manufacturer's
instructions. A microplate reader (cat. no. A51119500C; Thermo
Fisher Scientific, Inc.) was then used to measure the absorbance of
the supernatant at 450 nm.
Cell colony formation assay
HSC3 and SCC15 cells were seeded in 6-well plates at
a density of 1×103 cells/well without additional drug
treatment. Cells were incubated in a humidified incubator with 5%
CO2 at 37°C for 2 weeks to allow colony formation. After
the incubation period, cells were washed twice with PBS (5 min each
wash). Subsequently, cells were fixed with 4% formaldehyde at room
temperature for 15 min. The fixative was discarded, and cells were
stained with 0.1% crystal violet solution (w/v in distilled water)
at room temperature for 20 min. Excess crystal violet was rinsed
off with distilled water, and the plates were air-dried. A colony
was defined as a cluster containing ≥50 cells. Colony
quantification was performed using ImageJ software (Version 1.53;
National Institutes of Health). Briefly, images of stained wells
were captured by camera (cat. no. ILCE-7M4, Sony Corporation), and
the ImageJ software (Version 1.53; National Institutes of Health)
was used to count the number of discrete colonies based on color
thresholding and size gating.
Transwell assay
Transwell migration assays were performed using
Transwell chambers with an 8 µm pore size (cat. no. 353097;
Falcon®; Corning Life Sciences). No Matrigel was used in
the assay, confirming this as a migration experiment. The lower
chamber of the 24-well Transwell system was filled with DMEM
supplemented with 10% FBS. HSC3 and SCC15 cells were resuspended in
serum-free DMEM, and 5×104 cells/well were seeded into
the upper chamber in a total volume of 200 µl. Cells were incubated
in a humidified incubator with 5% CO2 at 37°C for 48 h
to allow migration through the pores.
After incubation, non-migrated cells on the upper
surface of the membrane were gently removed with a cotton swab.
Migrated cells adhering to the lower surface were fixed with 4%
formaldehyde at room temperature for 30 min, then stained with 0.1%
crystal violet solution at room temperature for 30 min. Excess
stain was rinsed off with PBS, and the membranes were air-dried.
Migrated cells were observed and counted using a Leica inverted
light microscope (Leica Microsystems) at a magnification of 100×.
Scale bar, 100 µm were included in the figure legends for
reference. Three random fields were selected per membrane to count
migrated cells, and the average number was calculated for
statistical analysis.
Wound healing assay
A total of 2×105 HSC3 and SCC15 cells
were seeded into each well of a 6-well plate. The cells were
cultured in DMEM/F-12 medium supplemented with 10% FBS (Gibco;
Thermo Fisher Scientific, Inc.) without additional drug treatment,
and no serum starvation was performed before the assay. When cell
confluence reached ~90% of the plate surface, a sterile 200 µl
pipette tip was used to scratch a straight wound across each well.
The scratched cells were gently washed away with PBS (twice) to
remove cell debris, and fresh DMEM/F-12 medium containing 10% FBS
was added to each well. Wound width was measured at 0 h and the
endpoint time (8, 12 or 24 h) under the bright field of a Leica
inverted light microscope (Leica Microsystems). The percentage of
wound healing was calculated using the formula: Wound healing
(%)=1-(widtht/width0) ×100, where
width0 is the initial wound width at 0 h and
widtht is the wound width at the endpoint time.
Cell cycle analysis
Cell cycle progression of HSC3 and SCC15 cells was
assessed using a Cell Cycle Kit (cat. no. CCS012; MultiSciences
(Lianke) Biotech Co., Ltd.) (no cytokines/chemokines used).
2×105−1×106 harvested cells were washed with
PBS, then mixed with 1 ml DNA staining solution + 10 µl
permeabilization solution (kit components), vortexed 5–10 sec, and
incubated at room temperature in the dark for 30 min. Stained cells
were analyzed on a BD FACSCanto™ II flow cytometer (low injection
speed), and data (G0/G1, S, G2/M phases) were processed via FlowJo
v10.8.1 (BD Biosciences). No antibodies were used.
Xenograft mouse model
A total of 12 6-week-old female BALB/c nude mice
(18–22 g) were purchased from the Animal Experiment Center of The
First Affiliated Hospital, Sun Yat-Sen University. Mice were housed
under specific pathogen-free conditions (20–25°C; relative
humidity, 40–70%; 12 h light/12 h dark cycle and ad libitum access
to sterile food and water).
1×106 tumor cells (METTL5 knockout and
control SCC15 cells, in 50 µl sterile PBS) were subcutaneously
injected into the right dorsal flank of nude mice. Palpable tumors
were monitored once weekly. Humane endpoints: Excess tumor size
(maximum permitted size was 20 mm), >20% weight loss, poor
mobility or distress.
Experiment ended 18 days post-injection. Mice were
euthanized by cervical dislocation, and death confirmed by 5-min
absence of breathing/heartbeat. Tumors were excised, weighed,
measured and imaged. IHC samples were fixed with 4%
paraformaldehyde at 4°C for 24 h.
All experiments were approved by the Ethical
Committee of The First Affiliated Hospital, Sun Yat-Sen University
[approval no. (2024)051] and conducted per institutional
guidelines.
Ribosome nascent-chain (RNC)
complex-bound mRNA sequencing (mRNA-seq)
To examine the alteration of mRNA translation after
METTL5 depletion, mRNA-seq was performed on total mRNAs and
RNC-mRNAs from METTL5-knocked-out or control HSC3 cells. RNC-mRNAs
were extracted as previously described. Cells were quickly washed
with ice-cold PBS containing 100 mg/ml cycloheximide and then
collected and lysed in 1.2 ml lysis buffer containing 1% Triton
X-100 in ribosome buffer [which contains 20 mM HEPESKOH (pH 7.4),
15 mM MgCl2, 200 mM KCl, 100 mg/ml cycloheximide and 2 mM
dithiothreitol] for 30 min on ice. Cell lysates were centrifuged at
16,200 × g for 10 min at 4°C to remove the cell debris. The
supernatants were transferred on the surface of 12 ml of ribosome
buffer containing 30% sucrose. RNC-mRNAs were pelleted after
ultra-centrifugation at 185,000 × g for 5 h at 4°C. RNAs in total
cell lysates and RNC-mRNAs were extracted with TRIzol®
reagent (cat. no. 15596018; Thermo Fisher Scientific, Inc.) and
subjected to RNA-sequencing (RNA-seq).
For RNA-seq experiments, the following details were
followed: i) RNA sample preparation kit: TRIzol® reagent
(cat. no. 15596018; Thermo Fisher Scientific, Inc.); ii) sample
quality/integrity verification: RNA quality was assessed by Q20
(>90%), Q30 (>85%) values, and RNA integrity number (RIN)
≥7.0 using a Microplate Reader (Molecular Devices, Gemini XPS) and
Qubit® 4 Fluorometer (cat. no. Q33216, Thermo Fisher
Scientific, Inc.); iii) sequencing type: Paired-end sequencing
(PE150) on the DNBSEQ platform; iv) sequencing kit: DNBSEQ-T7RS
High-throughput Sequencing Kit (cat. no. 940-000268-00; Shenzhen
BGI Genomics Co., Ltd.); v) Final library loading concentration: 2
nM, measured by reverse transcription-quantitative PCR (qPCR) using
the Qubit® ssDNA Assay Kit (cat. no. Q10212, Invitrogen;
Thermo Fisher Scientific, Inc.) for molar concentration
quantification; vi) Data analysis software: SOAPnuke (version
1.5.6; http://github.com/BGI-flexlab/SOAPnuke) for raw data
filtering, HISAT2 (version 2.1.0; http://daehwankimlab.github.io/hisat2/) for genome
alignment, Bowtie2 (version 2.3.4.3; http://bowtie-bio.sourceforge.net/bowtie2/index.shtml)
for gene alignment, RSEM (version 1.2.28; http://deweylab.github.io/RSEM/) for gene expression
quantification, DESeq2 (version 1.40.2; http://bioconductor.org/packages/3.16/bioc/html/DESeq2.html)
for differential expression analysis.
The TE was calculated using the following formula:
TE=transcripts per million (TPM) in RNC-seq/TPM in input RNA-seq.
Genes with a TE ratio of sgMETTL5 (sgM5)/NC <0.667 and >1.5
were classified as decreased TE (TE down) and enhanced TE (TE up),
respectively.
Gene Ontology (GO) and pathway
analyses
GO and pathway analyses of TE-down mRNAs identified
in RNC-seq data were performed using the Bioinformatics Webtool
(https://www.bioinformatics.com.cn/),
and significantly enriched pathways were identified.
Datasets
Gene Expression Profiling Interactive Analysis
(GEPIA) database (http://gepia.cancer-pku.cn/): The dataset ‘TCGA-HNSC’
(The Cancer Genome Atlas Head and Neck Squamous Cell Carcinoma) was
queried with the search term ‘METTL5’ to analyze METTL5 mRNA
expression levels in tumor vs. normal tissues. Survival analysis
was performed via GEPIA using the ‘TCGA-HNSC’ dataset with METTL5
expression stratified into ‘low’ and ‘high’ groups (median cutoff).
University of Alabama at Birmingham Cancer data analysis Portal
(UALCAN; http://ualcan.path.uab.edu/index.html): The dataset
‘TCGA-HNSC’ was queried with the search term ‘METTL5’ to analyze
the association between METTL5 expression and clinical parameters
(lymph node metastasis, cancer stage, tumor grade).
Statistical analysis
Quantitative data are presented as mean ± standard
deviation. Unpaired two-tailed Student's t-test was used to assess
statistically significant differences between two experimental
groups, while one-way ANOVA followed by Tukey's honest significant
difference post hoc test was applied for multiple groups. For
non-parametric data, the Kruskal-Wallis test was used, with Dunn's
post hoc test for pairwise comparisons following a significant
result. Pearson's correlation analysis was performed for the data
presented in Fig. S4C (from
GEPIA2; http://gepia2.cancer-pku.cn/). All
experiments were performed with three independent biological
repeats. Statistical significance was defined as P<0.05; for
differential gene analysis (where applicable), the significance
cut-off levels were set as adjusted P<0.05 and |log2 fold
change| >1. Statistical analyses were performed using GraphPad
Prism (version 8.0; Dotmatics). *P<0.05, **P<0.01;
***P<0.001; ****P<0.0001.
Results
METTL5 expression is elevated in head
and neck squamous cell carcinoma (HNSCC) and is associated with
poor prognosis of HNSCC
To ascertain whether METTL5 contributes to cancer
progression, the present study performed an analysis of METTL5 mRNA
levels in HNSCC using the Gene Expression Profiling Interactive
Analysis database (http://gepia.cancer-pku.cn/, dataset are provided in
Methods section: Datasets). The findings indicated that METTL5
expression was significantly higher in tumor tissues compared with
that in normal tissues (Fig. 1A).
In addition, data from University of Alabama at Birmingham Cancer
data analysis Portal (https://ualcan.path.uab.edu/index.html, dataset are
provided in Methods section: Datasets) revealed that elevated
METTL5 expression was significantly associated with increased lymph
node metastasis (vs. no lymph node metastasis), advanced cancer
stages (vs. early stages) and higher tumor grades (vs. lower
grades) in HNSCC (Fig. 1C-E).
Furthermore, patients with higher METTL5 expression exhibited
significantly worse overall survival than those with low METTL5
expression (Fig. 1B). Collectively,
these findings suggest that METTL5 is key to regulating the
progression of HNSCC.
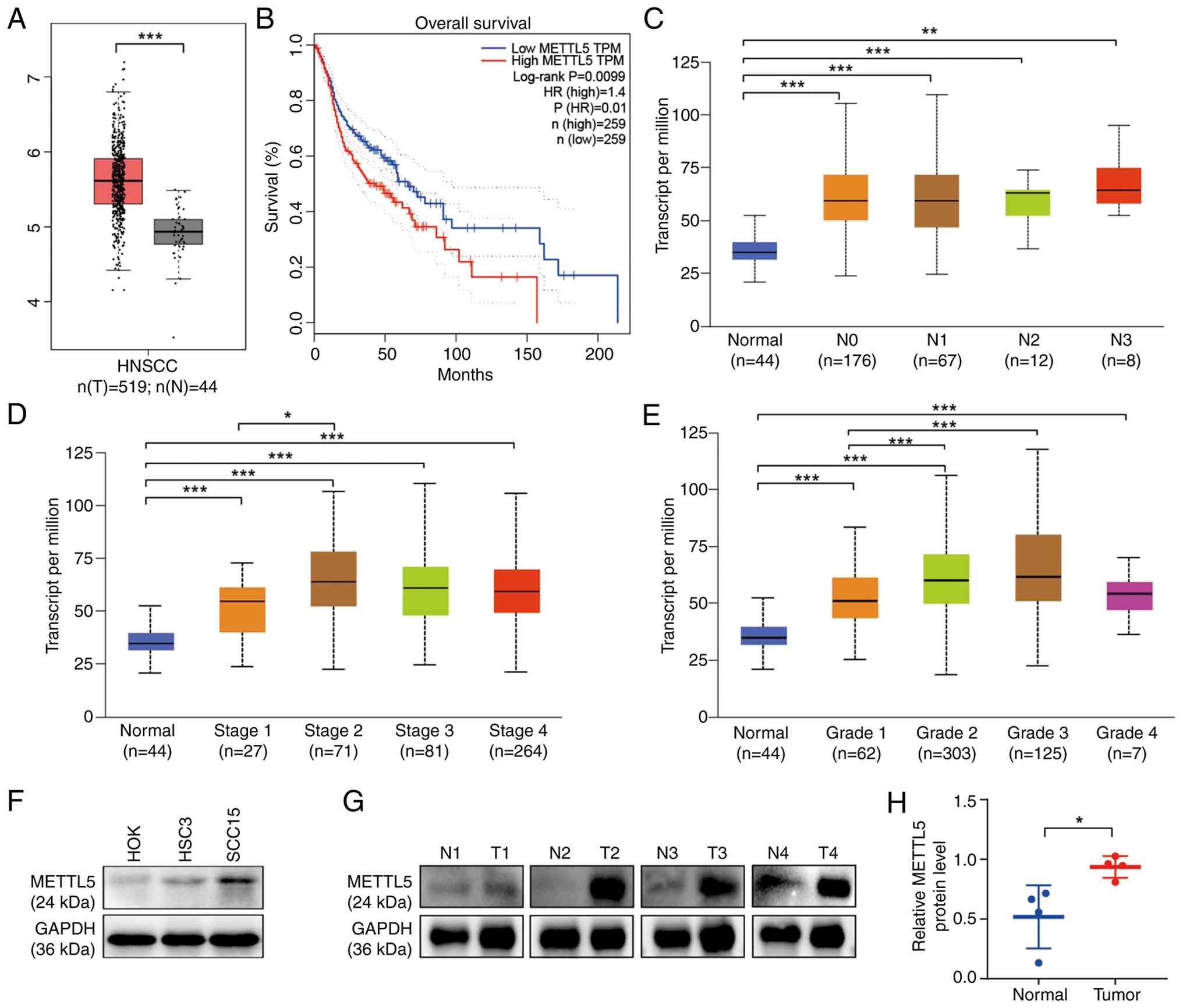 | Figure 1.METTL5 is elevated in HNSCC and is
associated with poor prognosis of HNSCC. (A) Comparison of mRNA
levels of METTL5 in HNSCC with normal tissues in TCGA-HNSCC
dataset. (B) Overall survival of patients between METTL5-high
(n=259) and METTL5-low (n=259) groups. (C) METTL5 expression level
in normal and HNSCC tissues with different stages of lymph node
metastasis. (D) METTL5 expression level in normal tissues and
different stages of HNSCC tissues. (E) METTL5 expression level in
normal HNSCC tissues and different grades of HNSCC tissues. (F)
METTL5 protein level in OSCC(HSC3, SCC15) and HOK cell lines. (G)
Representative bands of METTL5 protein expression levels in human
OSCC and paired normal tissues measured by western blotting. (H)
Semi-quantification of METTL5 protein expression levels in human
OSCC and paired normal tissues. *P<0.05, **P<0.01;
***P<0.001. METTL5, methyltransferase 5, N6-adenosine; HNSCC,
head and neck squamous cell carcinoma; TCGA, The Cancer Genome
Atlas; OSCC, oral squamous cell carcinoma; HOK, human normal oral
epithelial keratinocyte; HR, hazard ratio; TPM, transcripts per
million; N, normal; T, tumor. |
Furthermore, western blotting analysis was used to
assess METTL5 expression in two OSCC cell lines and a human normal
oral epithelial keratinocyte (HOK) cell line. The results revealed
that OSCC cells exhibited markedly higher METTL5 expression
compared with that in HOK cells (Fig.
1F). Furthermore, western blotting analysis demonstrated
significantly elevated METTL5 protein levels in human OSCC tissues
compared with that in normal tissues (Fig. 1G and H).
Knockout of METTL5 inhibits the
progression of OSCC in vitro
To elucidate the function of METTL5 in OSCC, the
present study established stable knockout cell lines using SCC15
and HSC3 cells, targeting METTL5 with independent sgMETTL5
(Figs. 2A and S1A). Notably, the METTL5 knockout cells
demonstrated a significant decrease in proliferative capacity
(Figs. 2B and S1B) and a colony formation (Figs. 2C and S1C) compared with that in the control
group. Furthermore, the Transwell migration and wound healing
assays indicated that METTL5 knockout significantly impaired the
migration of OSCC cells in vitro compared with that in
control cells (Figs. 2D, E,
S1D and E).
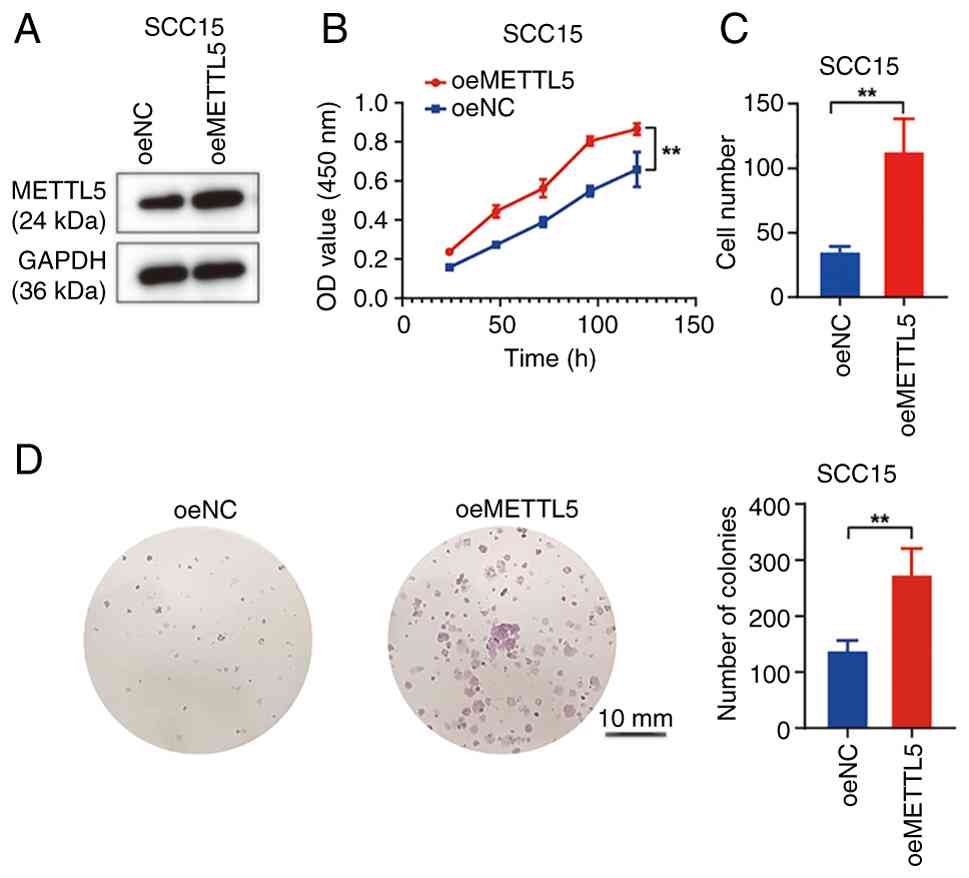
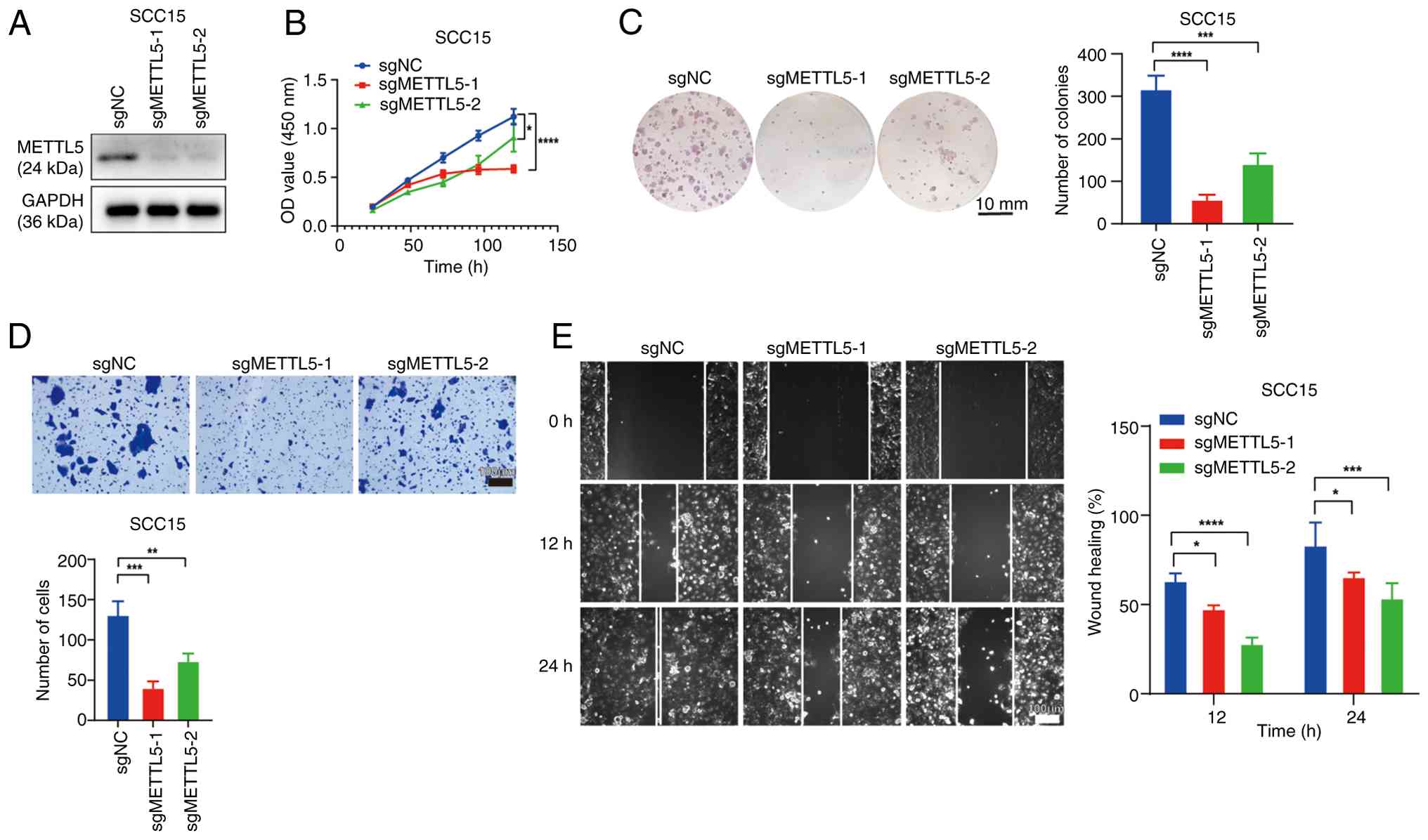 | Figure 2.Knockout of METTL5 inhibits the
progression of OSCC in vitro. (A) Validation of METTL5
stable knockout in SCC15 cells by western blotting. Results of (B)
Cell Counting Kit-8 assay, (C) colony formation assay, (D)
Transwell assay and (E) wound healing assay after knockout of
METTL5 in SCC15 cells. *P<0.05, **P<0.01; ***P<0.001;
****P<0.0001. METTL5, methyltransferase 5, N6-adenosine; NC,
negative control; OSCC, oral squamous cell carcinoma; sg, small
guide RNA; NC, negative control; OD, optical density. |
Overexpression of METTL5 promotes the
progression of OSCC
A METTL5 overexpression system was also constructed
using lentivirus to investigate the functional impact of METTL5 in
OSCC (Figs. 3A and S2A). The findings revealed that METTL5
overexpression led to a significant increase in cellular
proliferation (Figs. 3B and
S2B) and colony formation
(Fig. 3D) in OSCC cells.
Furthermore, the overexpression of METTL5 enhanced the migration of
OSCC cells (Figs. 3C and S2C), indicating a role for METTL5 in
promoting the malignancy of OSCC.
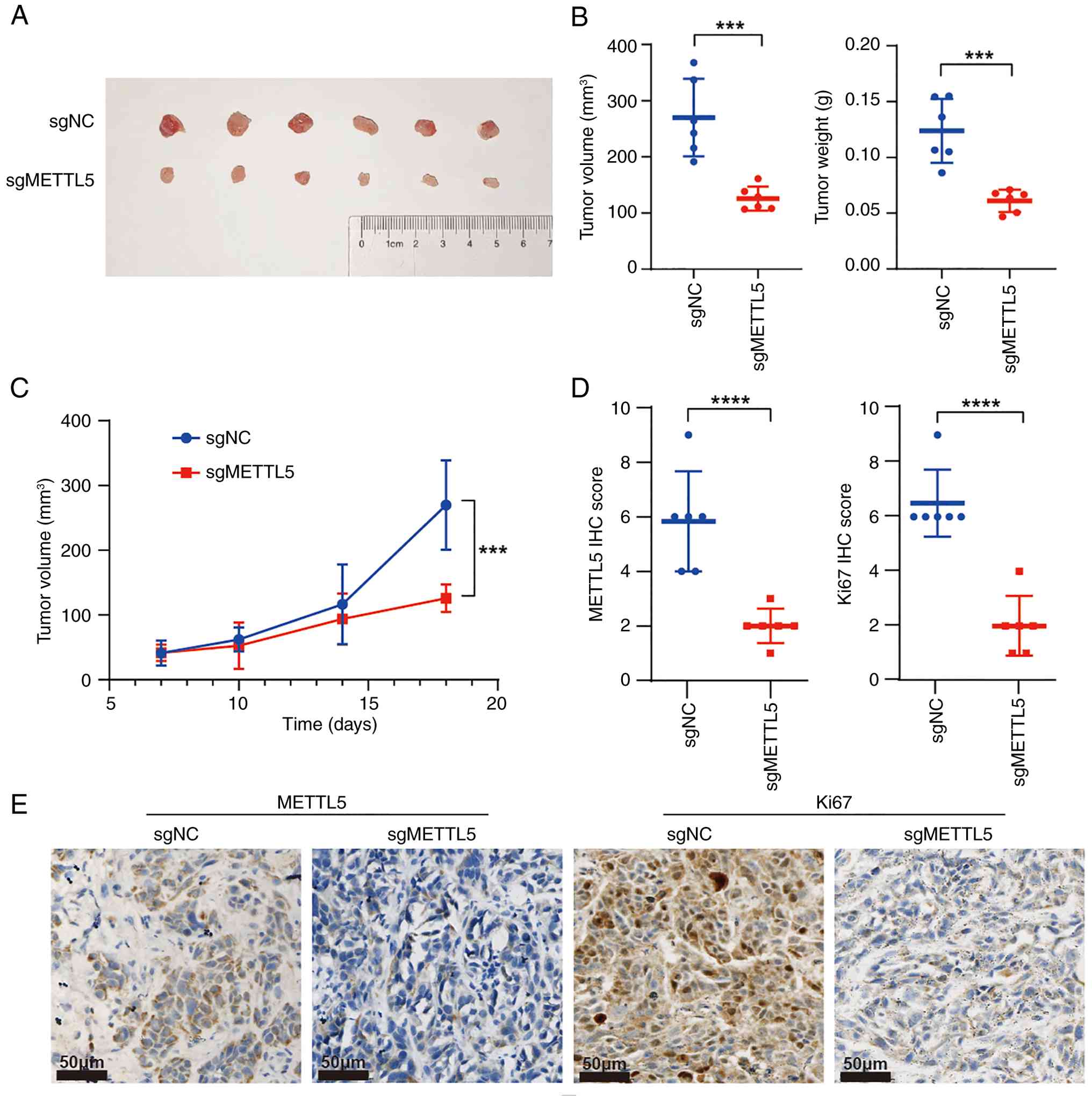
Inhibition of METTL5 suppresses tumor
growth in vivo
Xenograft tumor-formation assays were subsequently
performed to assess the role of METTL5 in the regulation of OSCC
tumorigenesis in vivo. METTL5 knockout and control SCC15
cells were inoculated subcutaneously into the right dorsal flank of
nude mice, with tumor volumes monitored throughout the experiment.
The results indicated that the subcutaneous tumors were notably
larger in the control group compared with that in the sgMETTL5
group (Fig. 4A). Compared with that
in the control group, the growth curve of the xenograft tumor and
the final tumor weights further demonstrated that the knockout of
METTL5 significantly impeded tumor growth in vivo (Fig. 4B and C). Moreover, IHC analysis
revealed that the expression level of METTL5 was significantly
diminished in xenograft tumors derived from the sgMETTL5 group
compared with that in the control group (Fig. 4D and E). In addition, xenograft
tumors from the sgMETTL5 groups exhibited significantly lower Ki-67
expression compared with that in the control group, indicating that
METTL5-knockout xenograft tumors were potentially less malignant
(Fig. 4D and E).
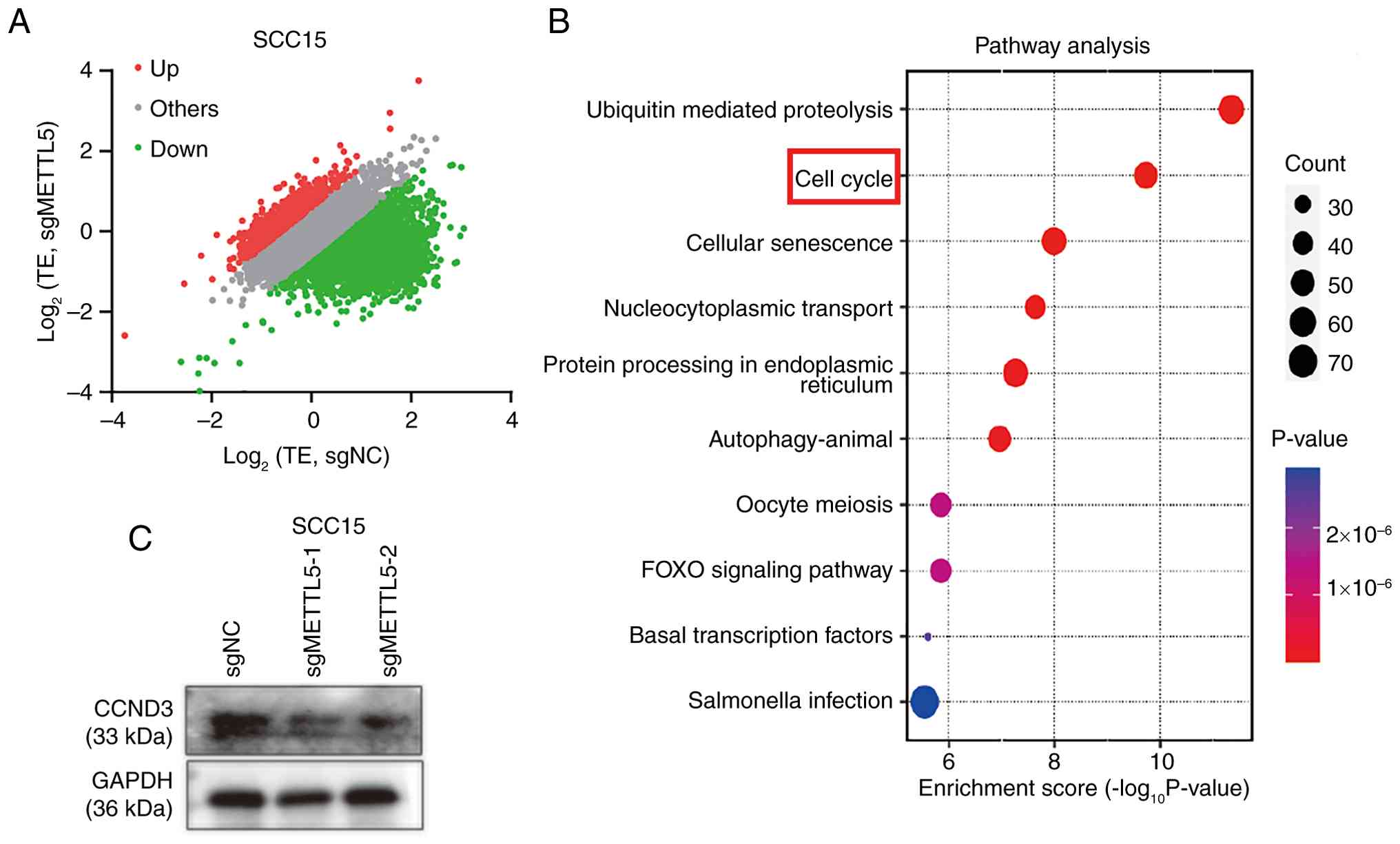
Knockout of METTL5 selectively
inhibits the translation of oncogenic mRNAs
The present study further utilized RNC-mRNA-seq to
assess mRNA translation profiles in METTL5-depleted and control
cells. The RNC-mRNA-seq analysis revealed 3,391 genes with TE-down
and 1,020 genes with TE-up in METTL5-depleted OSCC cells (Fig. 5A). This suggests that
METTL5-knockout predominantly leads to a decrease in TE rather than
an increase. Furthermore, GO analysis of genes with TE-down
revealed that ‘Cell cycle’-related processes were significantly
enriched (Fig. 5B). Among the cell
cycle-related TE-down genes identified, the present study
prioritized cyclin D3 (CCND3) as it is a well-characterized cyclin
driving cell cycle phase transition, and its dysregulation is
directly associated with tumor growth (20). Subsequently, western blotting
demonstrated that the levels of the cell-cycle regulator CCND3 were
markedly decreased in sgMETTL5 cells compared with that in control
cells (Fig. 5C). Consistent with
this, cell cycle assays revealed that knockout of METTL5 induced
the proportion of OSCC cells in the G0/G1
phase and decreased the proportion in the G2/M phase
(Fig. 6D and S3). In summary, the results indicate that
the depletion of METTL5 in OSCC cells significantly alters the
translational profile. This alteration not only results in a
general decrease in TE but also specifically impacts the
translation of certain mRNAs, particularly those associated
involved in cell cycle regulation.
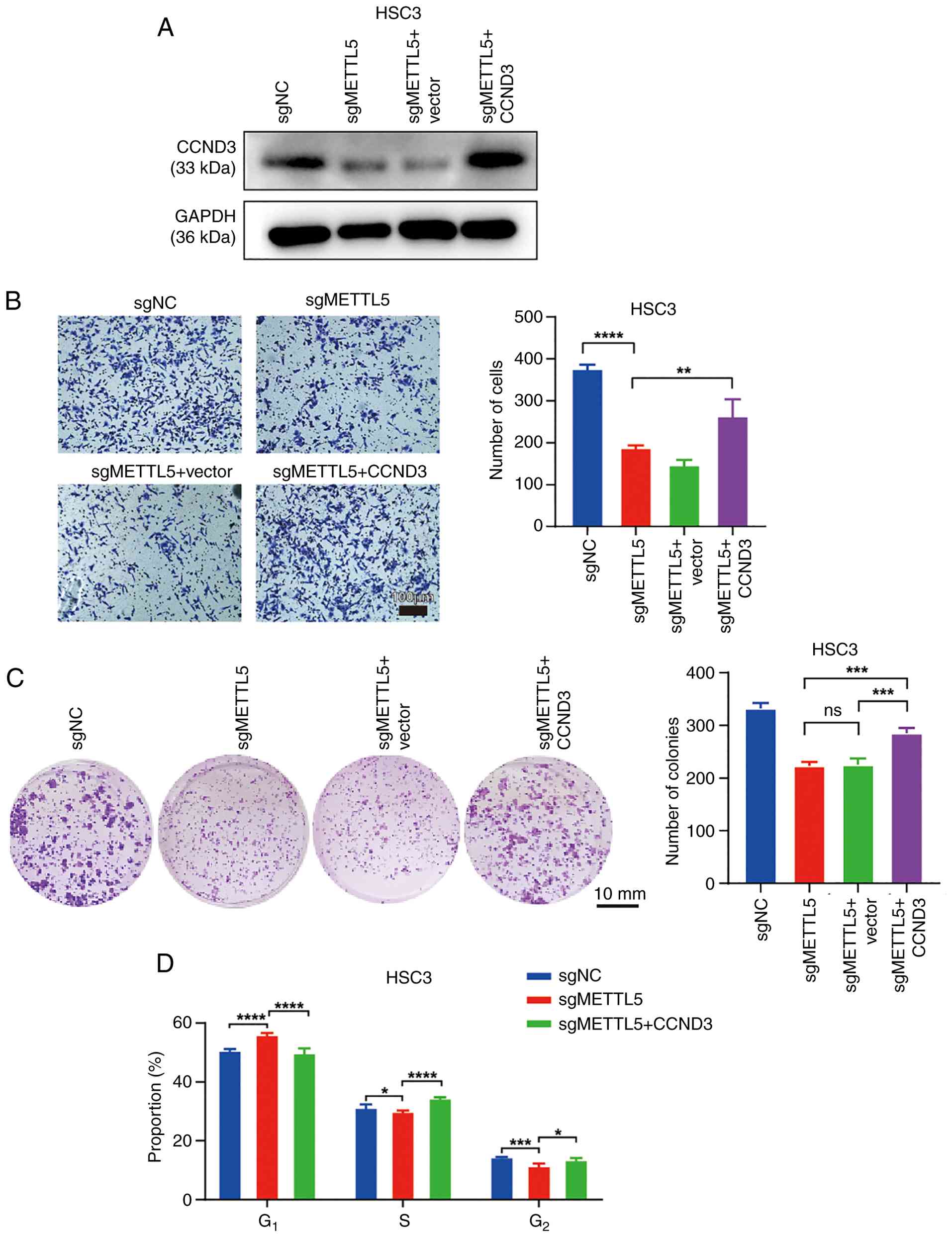
CCND3 overexpression partially rescues
cancer cell phenotypes in METTL5-depleted OSCC cells
The aforementioned results indicate that the
downregulation of METTL5 expression disrupts the translation
process, particularly for mRNAs involved in the ‘Cell cycle’
pathway. Consequently, rescue experiments (Fig. 6) were performed using HSC3 cells due
to their notably increased TE and higher baseline invasion
capacity, which are key to robustly evaluate the functional
recovery of migration ability. Although SCC15 cells demonstrated
higher basal METTL5 expression (Fig.
1F), the characteristics of HSC3 made it a more suitable model
for validating the rescue effect of CCND3 overexpression.
Therefore, the present study overexpressed CCND3 in METTL5-depleted
HSC3 cells (Fig. 6A). The results
revealed that overexpression of CCND3 partially rescued the
migration and colony formation capacity of METTL5-depleted OSCC
cells (Fig. 6B and C). Moreover,
for cell cycle analysis, METTL5 knockout induced a cell cycle
arrest in the G0/G1 phase compared with that
in control cells, and CCND3 overexpression reversed this (Figs. 6D and S3). Collectively, the results suggest
that METTL5-mediated m6A modification contributes to
translational dysregulation, potentially influencing OSCC
progression. To explore the potential broader relevance of the
findings of the present study, the expression patterns of METTL5
and CCND3 were analyzed across several cancer types using TCGA
data. The results demonstrated that METTL5 (Fig. S4A) and CCND3 (Fig. S4B) exhibited differential
expression in multiple cancer types compared with that in adjacent
normal tissues. Moreover, although there is no existing literature
reporting a direct association between METTL5 and CCND3 in other
cancer types, to the best of our knowledge, the weak but
statistically significant mRNA-level correlation demonstrated in
the generated scatter plot indicates that METTL5 transcription
minimally drives CCND3 mRNA expression (Fig. S4C).
Discussion
Ribosomes are intricate macromolecular machines that
serve as the cellular catalysts for protein synthesis. They are key
to translating all cellular proteins and eukaryotic cytoplasmic
ribosomes, and are intricately assembled from four types of rRNA
and ~80 distinct ribosomal proteins. Universally, they are
organized into two subunits, each consisting of a complex array of
proteins and several rRNAs (23).
rRNAs are the architects of the ribosomal subunits, shaping both
their form and function, and they provide the enzymatic activity
key to protein synthesis. Emerging evidence indicates that rRNAs
may influence multiple aspects of protein synthesis, including the
recruitment of mRNA, modulation of TE for specific mRNAs and the
facilitation of ribosomal shunting (24,25).
rRNAs are distinguished as one of the most extensively modified
class of RNAs, with ~2% of their nucleotides subject to
post-transcriptional modifications. These modifications constitute
a notable source of ribosome diversity and are increasingly
acknowledged for their role in ribosome specialization. There is
extensive research that implicates abnormalities in the enzymes
responsible for rRNA modification, as well as variations in the
extent of these modifications, in the etiology of developmental
disorders, genetic conditions and cancer (24).
rRNA constitutes >80% of the total RNA in the
cell (26), yet the physiological
functions of rRNA modifications remain to be elucidated.
m6A modification refers to the reversible process of
methylation on the 6th nitrogen atom of RNA adenine catalyzed by
methyltransferase (27). The
m6A methylation mark is also present within the decoding
center of 18S rRNA (28); however,
the specific biological function of m6A on 18S rRNA
remains to be elucidated. METTL5, an enzyme responsible for the
m6A modification of rRNA, has been associated with
several types of cancer, including breast cancer, liver cancer and
nasopharyngeal carcinoma. Moreover, the role of METTL5 in
catalyzing m6A modifications appears to differ across
distinct cancer types, such as breast cancer (promoting cell
growth) and nasopharyngeal carcinoma (enhancing chemoresistance)
(17,22,29).
Research suggests that METTL5-mediated m6A modifications
are associated with intracellular TE (30), tumor cell proliferation, cell
pluripotency (10) and stress
response (31). However, the
specific role and mechanism of METTL5 in OSCC remain to be fully
elucidated.
The present study identified that the m6A
methyltransferase METTL5 is significantly overexpressed in OSCC and
is associated with a poor patient prognosis. The results also
demonstrated that downregulation of METTL5 significantly reduced
the proliferative and migratory capacities of human OSCC cells,
whilst its overexpression promoted OSCC progression in
vitro. Furthermore, the in vivo OSCC models in the
present study utilizing METTL5-deficient SCC15 cells, corroborated
the role of METTL5 in the development and progression of OSCC
within a living organism. At the molecular level, the present study
identified the CCND3 signaling pathway as a key
m6A-modified target that mediates the effects of METTL5
on OSCC progression. Collectively, the results of the present
study, including both in vitro and in vivo
experiments, provide evidence highlighting the notable role of the
m6A modification in regulating OSCC tumorigenesis and
its progression. Although METTL5 depletion phenocopies
translational disruption, meaning that METTL5 knockout leads to
phenotypes (such as reduced translation efficiency) similar to
those caused by direct disruption of translational machinery,
direct causality requires further validation to confirm
m6A-dependent mechanisms.
Emerging evidence has revealed the oncogenic
functions of CCND3 in regulating the progression of several
diseases, including cancer. In diffuse large B-cell lymphoma, it
has been reported that pathways associated with cell cycle and DNA
replication are markedly upregulated in patients harboring
mutations in the CCND3 gene (32).
MicroRNA-4779 has been reported to suppress cancer cell
proliferation by inducing cell cycle arrest and apoptosis through
the direct targeting of p21 (RAC1) activated kinase 2 and CCND3.
Furthermore, the downregulation of CCND3 has been shown to induce
cell cycle arrest and apoptosis (33). In addition, polypyrimidine
tract-binding protein 1 has been identified to enhance the
translation of CCND3 by binding to the 5′-untranslated region of
the CCND3 mRNA, thereby facilitating cell cycle progression and
tumor growth (23). Therefore,
targeting the CCND3 represents a promising therapeutic strategy for
cancer treatment. Moreover, METTL5-mediated rRNA m6A
modification is known to modulate overall translation activity and
the selective translation of certain mRNAs (31). The results of the present study
demonstrated that CCND3 mRNA was affected by METTL5 expression, and
that knockout of METTL5 diminished CCND3 protein levels. This
indicates that METTL5-mediated m6A modification
regulates the TE of CCND3 mRNA. Furthermore, the present study
revealed that the forced activation of CCND3 restored the migration
capacity of METTL5 knockout cells. This finding suggests that
METTL5-mediated m6A deposition is a predominant,
although not exclusive, regulator of CCND3 translation.
Notably, the HSC3 and SCC15 cell lines used in the
present study are derived from tongue squamous cell carcinoma, with
HSC3 from lymph node metastasis (34). Due to the heterogeneity of OSCC
across oral subsites, the findings of the present study may not
fully represent OSCC from other sites, which warrants validation in
more diverse cell lines and samples. Furthermore, he limitations in
evaluating the role of METTL5 in OSCC invasion and metastasis are
acknowledged. The in vivo experiments in the present study
were restricted to subcutaneous xenografts, which do not fully
recapitulate the oral mucosal microenvironment, including
epithelial-mesenchymal interactions and immune cell infiltration.
This lack of microenvironment-relevant models limits the clinical
translatability of the findings, as the effects of METTL5 cannot be
definitively associated with the context-specific mechanisms
driving invasion and metastasis in native oral tissues. Moreover,
the potential involvement of METTL5 in OSCC metastasis is supported
by in vitro migration data and the results indicating that
high METTL5 expression is associated with lymph node metastasis and
poor prognosis rather than direct in vivo evidence. Future
studies utilizing orthotopic models or patient-derived xenografts
with intact microenvironments will be key to validate these
associations and clarify the role of METTL5 in clinically relevant
invasion and metastasis processes.
Furthermore, although the present study demonstrated
the promotional role of METTL5 in OSCC progression through a series
of in vitro and in vivo experiments, and revealed its
potential function in regulating the TE of CCND3, certain
limitations should be acknowledged. Notably, the present study did
not directly verify whether the m6A modification level
at the A1832 site of 18S rRNA was indeed reduced after METTL5
knockout. Due to the current unavailability of commercial
site-specific m6A antibodies targeting the A1832 site of
18S rRNA, and the technical challenges in conventional
m6A-quantitive (q) PCR and associated techniques to
resolve adjacent modification sites in rRNA, the present study
could not obtained direct experimental evidence regarding the
changes in modification levels at this specific site. Therefore,
the mechanism by which METTL5 selectively regulates the TE of
specific mRNAs (particularly CCND3) by catalyzing m6A
modification at the A1832 site of 18S rRNA currently remains a
hypothesis based on existing research findings and known mechanisms
in associated fields. In future studies, with the development of
specific detection technologies, further verification of the
dynamic changes of this modification site under METTL5 regulation
using methods such as m6A-qPCR and
m6A-cross-linking and immunoprecipitation/RNA
immunoprecipitation-qPCR targeting the A1832 site of 18S rRNA
should be performed to further clarify this molecular mechanism.
However, due to the conserved functions of METTL5 in regulating
translation and the extensive involvement of CCND3 in cell cycle
control across malignancies, future studies should explore whether
such an axis exists in other cancer types, which could further
expand the translational significance of this regulatory
relationship.
Additionally, the functional validation of CCND3 as
a downstream mediator of METTL5 in the present study was limited to
in vitro assays, and the absence of in vivo rescue
experiments (for example, overexpressing CCND3 in METTL5-knockout
xenografts) prevents the definitive establishment of the extent to
which CCND3 mediates the pro-tumorigenic effects of METTL5 in
vivo. This suggests that other translationally downregulated
genes may also contribute markedly to OSCC progression downstream
of METTL5.
In conclusion, an elevated expression level of
METTL5 in patients with OSCC is associated with an unfavorable
prognosis. Moreover, the downregulation of METTL5 expression
through is associated with a decrease in m6A rRNA
methylation and a selective decrease in the TE of oncogene CCND3 in
OSCC. These findings indicate the potential inhibitory effect of
METTL5 knockout on OSCC tumorigenesis and progression in both in
vitro and in vivo settings. The results of the present
study contribute to furthering the understanding of the epigenetic
network that governs OSCC progression. However, although the
results demonstrate the prognostic value of METTL5, its potential
as a therapeutic target requires further validation in future
studies. METTL5 is anticipated to offer potential novel biomarkers
and therapeutic targets for the diagnosis and treatment of OSCC in
the future.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was funded by the National Natural Science
Foundation of China (grant no. 82002871), Natural Science
Foundation of Guangdong Province (grant no. 2023A1515220258) and
Foundation and Application Foundation Project in Guangzhou (grant
no. SL2023A04J02127).
Availability of data and materials
The data generated in the present study may be found
in the China National Center for Bioinformation (CNCB) under
accession number OMIX012047 or at the following URL: https://share.cncb.ac.cn/AZg3Lt9OjY/OMIX012047/. The
remaining data generated in the present study may be requested from
the corresponding author.
Authors' contributions
LL performed the experiments and drafted the
manuscript. LL and XZ confirm the authenticity of all the raw data.
SS and XZ performed the experiments and acquired the data. SH
participated in the analysis of the data. LW, DS, LT and QH
designed the present study. LT and QH supervised the present study
and revised the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
Human data and samples were obtained with written
informed consent and the use was approved by the Independent Ethics
Committee for Clinical Research and Animal Trials of The First
Affiliated Hospital of Sun Yat-Sen University [approval no.
(2022)229]. The nude mouse experiments performed were approved by
the Independent Ethics Committee for Clinical Research and Animal
Trials of The First Affiliated Hospital of Sun Yat-Sen University
[approval no. (2024)051].
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Tan Y, Wang Z, Xu M, Li B, Huang Z, Qin S,
Nice EC, Tang J and Huang C: Oral squamous cell carcinomas: State
of the field and emerging directions. Int J Oral Sci. 15:442023.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hedberg ML, Goh G, Chiosea SI, Bauman JE,
Freilino ML, Zeng Y, Wang L, Diergaarde BB, Gooding WE, Lui VW, et
al: Genetic landscape of metastatic and recurrent head and neck
squamous cell carcinoma. J Clin Invest. 126:169–180. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lo Nigro C, Denaro N, Merlotti A and
Merlano M: Head and neck cancer: Improving outcomes with a
multidisciplinary approach. Cancer Manag Res. 9:363–371. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Han H, Yang C, Zhang S, Cheng M, Guo S,
Zhu Y, Ma J, Liang Y, Wang L, Zheng S, et al: METTL3-mediated
m6A mRNA modification promotes esophageal cancer
initiation and progression via Notch signaling pathway. Mol Ther
Nucleic Acids. 26:333–346. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chen Z, Chen J, Xu X, Li Q, Zhang C, Li S,
Liu L, Cao C, Chen D and He Q: METTL3-mediated ALDH m6A
methylation regulates the malignant behavior of BMI1+
HNSCC stem cells. Oral Dis. 30:1061–1071. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Pan L, She H, Wang K, Xia W, Tang H, Fan Y
and Ye J: Characterization of the m6A regulator-mediated
methylation modification patterns in oral squamous cell carcinoma.
Sci Rep. 13:66172023. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Qin Z, Bai J, He H and Li B: Expression
and significance of m6A-RNA-methylation in oral cancer
and precancerous lesion. Front Oncol. 13:10130542023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Li H, Wu H, Wang Q, Ning S, Xu S and Pang
D: Dual effects of N6-methyladenosine on cancer
progression and immunotherapy. Mol Ther Nucleic Acids. 24:25–39.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Qi YN, Liu Z, Hong LL, Li P and Ling ZQ:
Methyltransferase-like proteins in cancer biology and potential
therapeutic targeting. J Hematol Oncol. 16:892023. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Xing M, Liu Q, Mao C, Zeng H, Zhang X,
Zhao S, Chen L, Liu M, Shen B, Guo X, et al: The 18S rRNA
m6 A methyltransferase METTL5 promotes mouse embryonic
stem cell differentiation. EMBO Rep. 21:e498632020. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ignatova VV, Stolz P, Kaiser S, Gustafsson
TH, Lastres PR, Sanz-Moreno A, Cho YL, Amarie OV, Aguilar-Pimentel
A, Klein-Rodewald T, et al: The rRNA m6A
methyltransferase METTL5 is involved in pluripotency and
developmental programs. Genes Dev. 34:715–729. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang L, Liang Y, Lin R, Xiong Q, Yu P, Ma
J, Cheng M, Han H, Wang X, Wang G, et al: Mettl5 mediated 18S rRNA
N6-methyladenosine (m6A) modification controls stem cell
fate determination and neural function. Genes Dis. 9:268–274. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Liu X, Ma H, Ma L, Li K and Kang Y: The
potential role of methyltransferase-like 5 in deficient mismatch
repair of uterine corpus endometrial carcinoma. Bioengineered.
13:5525–5536. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang Z, Liu J, Yang Y, Xing C, Jing J and
Yuan Y: Expression and prognostic potential of ribosome 18S RNA
m6A methyltransferase METTL5 in gastric cancer. Cancer
Cell Int. 21:5692021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Shinde H and Kadam US: Growing prospects
of RNA therapeutics: A case of METTL5 and 18S rRNA m6A
modification. Mol Ther. 32:11–12. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
van Tran N, Ernst FGM, Hawley BR, Zorbas
C, Ulryck N, Hackert P, Bohnsack KE, Bohnsack MT, Jaffrey SR,
Graille M and Lafontaine DLJ: The human 18S rRNA m6A
methyltransferase METTL5 is stabilized by TRMT112. Nucleic Acids
Res. 47:7719–7733. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chen B, Huang Y, He S, Yu P, Wu L and Peng
H: N6-methyladenosine modification in 18S rRNA promotes
tumorigenesis and chemoresistance via HSF4b/HSP90B1/mutant p53
axis. Cell Chem Biol. 30:144–158.e10. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wei Z, Chen Y, Zeng Z, Peng Y, Li L, Hu N,
Gao X, Cai W, Yin L, Xu Y, et al: The novel m6A writer METTL5 as
prognostic biomarker probably associating with the regulation of
immune microenvironment in kidney cancer. Heliyon. 8:e120782022.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sun S, Fei K, Zhang G, Wang J, Yang Y, Guo
W, Yang Z, Wang J, Xue Q, Gao Y and He J: Construction and
comprehensive analyses of a METTL5-associated prognostic signature
with immune implication in lung adenocarcinomas. Front Genet.
11:6171742020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Xia P, Zhang H, Lu H, Xu K, Jiang X, Jiang
Y, Gongye X, Chen Z, Liu J, Chen X, et al: METTL5 stabilizes c-Myc
by facilitating USP5 translation to reprogram glucose metabolism
and promote hepatocellular carcinoma progression. Cancer Commun
(Lond). 43:338–364. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang L and Peng JL: METTL5 serves as a
diagnostic and prognostic biomarker in hepatocellular carcinoma by
influencing the immune microenvironment. Sci Rep. 13:107552023.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Rong B, Zhang Q, Wan J, Xing S, Dai R, Li
Y, Cai J, Xie J, Song Y, Chen J, et al: Ribosome 18S m6A
Methyltransferase METTL5 promotes translation initiation and breast
cancer cell growth. Cell Rep. 33:1085442020. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kang H, Heo S, Shin JJ, Ji E, Tak H, Ahn
S, Lee KJ, Lee EK and Kim W: A miR-194/PTBP1/CCND3 axis regulates
tumor growth in human hepatocellular carcinoma. J Pathol.
249:395–408. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Sloan KE, Warda AS, Sharma S, Entian KD,
Lafontaine DLJ and Bohnsack MT: Tuning the ribosome: The influence
of rRNA modification on eukaryotic ribosome biogenesis and
function. RNA Biol. 14:1138–1152. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Burman LG and Mauro VP: Analysis of rRNA
processing and translation in mammalian cells using a synthetic 18S
rRNA expression system. Nucleic Acids Res. 40:8085–8098. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Noller HF: Ribosomal RNA and translation.
Annu Rev Biochem. 60:191–227. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tian S, Lai J, Yu T, Li Q and Chen Q:
Regulation of gene expression associated with the
N6-Methyladenosine (m6A) enzyme system and its significance in
cancer. Front Oncol. 10:6236342021. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sepich-Poore C, Zheng Z, Schmitt E, Wen K,
Zhang ZS, Cui XL, Dai Q, Zhu AC, Zhang L, Sanchez Castillo A, et
al: The METTL5-TRMT112 N6-methyladenosine
methyltransferase complex regulates mRNA translation via 18S rRNA
methylation. J Biol Chem. 298:1015902022. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Peng H, Chen B, Wei W, Guo S, Han H, Yang
C, Ma J, Wang L, Peng S, Kuang M and Lin S:
N6-methyladenosine (m6A) in 18S rRNA promotes
fatty acid metabolism and oncogenic transformation. Nat Metab.
4:1041–1054. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Dai Z, Zhu W, Hou Y, Zhang X, Ren X, Lei
K, Liao J, Liu H, Chen Z, Peng S, et al: METTL5-mediated 18S rRNA
m6A modification promotes oncogenic mRNA translation and
intrahepatic cholangiocarcinoma progression. Mol Ther.
31:3225–3242. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Lei K, Lin S and Yuan Q:
N6-methyladenosine (m6A) modification of
ribosomal RNAs (rRNAs): Critical roles in mRNA translation and
diseases. Genes Dis. 10:126–134. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hua W, Li Y, Yin H, Du KX, Zhang XY, Wu
JZ, Liang JH, Shen HR, Gao R, Li JY, et al: Analysis of CCND3
mutations in diffuse large B-cell lymphoma. Ann Hematol.
103:5729–5739. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Koo KH and Kwon H: MicroRNA miR-4779
suppresses tumor growth by inducing apoptosis and cell cycle arrest
through direct targeting of PAK2 and CCND3. Cell Death Dis.
9:772018. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ribeiro IP, Rodrigues JM, Mascarenhas A,
Kosyakova N, Caramelo F, Liehr T, Melo JB and Carreira IM:
Cytogenetic, genomic, and epigenetic characterization of the HSC-3
tongue cell line with lymph node metastasis. J Oral Sci. 60:70–81.
2018. View Article : Google Scholar : PubMed/NCBI
|