Introduction
Neuroblastic tumors (NTs) histopathologically
include neuroblastoma (NB), ganglioneuroblastoma (GNB), and
ganglioneuroma (GN) (1–4). NB and GNB exhibit various clinical
features, from cases with a good prognosis that may undergo
spontaneous regression without treatment to those with a poor
prognosis, where survival rates remain at 30–40%, even with
intensive multimodal treatment (4,5). GN is
generally considered a benign tumor; however, there have been
reports of malignant transformation from GN to NB (6). Thus, NTs have diverse prognoses and
treatment responses, which necessitate a risk classification system
to guide effective treatment strategies for each tumor type. In
2009, the International Neuroblastoma Risk Group (INRG) criteria
were proposed for NB tumor classification, incorporating seven
potential prognostic factors: tumor stage, histology, MYCN
amplification, age at diagnosis, 11q aberration, and DNA ploidy.
These criteria stratify patients into four risk categories: very
low, low, intermediate, and high risk (7).
Many tumor cells extend their telomeres to maintain
proliferation and evade apoptosis by avoiding telomere shortening
during cell division. Recently, a comprehensive genomic analysis of
NB samples has revealed that the telomere maintenance mechanisms
(TMM) is frequently activated in high-risk (HR) NBs. Telomeric DNA
is maintained through transcriptional activation of the telomerase
reverse transcriptase (TERT) gene via MYCN
overexpression in MYCN-amplified NBs, genomic rearrangement
resulting in enhancer hijacking or promoter mutations of
TERT in MYCN-not-amplified HR NBs (8–10).
An alternative TMM involves telomere elongation via
the alternative lengthening of telomeres (ALT), which is strongly
correlated with ATRX alterations and predominantly observed
in MYCN-not-amplified HR NBs (8,11–13).
Recent reports have indicated that TMM-activated NB have poor
prognoses (14–16), even those with non-HR group
(14). Therefore, TMM is considered
a new prognostic marker for the next version of the INRG
classification. However, only a few reports on TMM in GNB and GN
(17–20), as well as NBs in Asian populations
have been published. Therefore, further studies on TMM in NTs are
required.
In this study, we aimed to examine 321 NTs diagnosed
in Japan for TMM, including 48 GNBs and 18 GNs. TERT qPCR
and C-circle assay demonstrated that TMM activation occurred in a
subpopulation of GNB and GN, in addition to NB. Genomic
characterization of ALT(+) and TERT-high NT tumors was
conducted using array comparative genomic hybridization (CGH) and
mutation analyses.
Materials and methods
Clinical samples
Written informed consent was obtained from the
parents or legal guardians of all patients, and assent was obtained
from the patients themselves when appropriate, at hospitals
participating in the Japan Childhood Cancer Group Neuroblastoma
Committee (JCCG-JNBSG). A total of 321 NT samples were collected
from patients aged between 0 months and 9 years who underwent
surgery or biopsy between November 2014 and April 2018. All tumor
samples analyzed in this study were obtained at the time of initial
diagnosis, prior to any treatment. These tumors were
histopathologically diagnosed as NB, GNB, or GN by a pathological
central review of the JCCG-JNBSG and staged according to the
International Neuroblastoma Staging System (INSS) (7). The MYCN gene copy number and
DNA ploidy were determined as part of the routine diagnostic
procedures by the central molecular diagnosis team at Saitama
Cancer Center (21,22). The study design was approved by the
Ethics Committee of Saitama Cancer Center (approval nos. 1528 and
1529).
C-circle and telomere content
assays
Genomic DNA from tumors and the ALT(+) neuroblastoma
cell line SK-N-FI was extracted using the standard proteinase K
digestion and phenol- chloroform extraction method. The assay was
performed on each sample with and without phai29 polymerase.
C-circle and telomere content (TC) assays using qPCR were performed
in triplicate by QuantStudio 7 Flex Real-Time PCR System (Applied
Biosystems), as previously described (23–25).
Primer sequences for qPCR are listed in Table SI. TC was relative to that of ALT+
cell line SK-N-FI with arbitrary value of 14. C-circle was relative
to that of ALT+ cell line SK-N-FI with arbitrary value of 196.
Telomere elongation and ALT were defined as TC >12 and C-circle
level >7.5, respectively, based on previously established
calculations and cut-off values (24,25).
SK-N-FI cells were grown in Dulbecco's modified Eagle's minimal
essential medium (DMEM, FujiFilm) supplemented with 10% FBS, 100
µg/ml penicillin/streptomycin, and 1% MEM NonEssential Amino Acids
(Fujifilm). Cultures were maintained at 37°C under 5%
CO2 in air. We confirmed SK-N-FI was mycoplasma-free and
had characteristic STR markers.
TERT expression and genomic
rearrangement analysis
Total RNA from human frozen tissue was extracted by
ISOGEN II (NIPPON GENE) and cDNA synthesis was performed by
ReverTra Ace® (TOYOBO) and random primers (Takara Bio)
according to the manufacturer's instructions. RT-qPCR analysis was
conducted in triplicate using QuantStudio 7 Flex Real-Time PCR
System (Applied Biosystems). TERT mRNA expression was
measured using qPCR (Table SI).
The median TERT mRNA expression in NB diagnosed at >18
months of age (arbitrary unit=0.31) was used to define the
TERT-high (>0.31) and TERT-low (<0.31)
expression groups. The genomic status of the TERT locus was
assessed by break-apart fluorescence in situ hybridization
(FISH) using custom SureFISH probes (Agilent Technologies
cat#G110997R-8) following the manufacturer's protocol. Images were
acquired using the BZ-X710 and BZ-X Analyzer (Keyence). Structural
abnormalities of the TERT gene were defined as cases in
which the signals were separated by more than 10% of the nuclear
major axis length. In TERT-high tumors without TERT
structural abnormalities, TERT promoter mutation analysis
was performed by Sanger sequencing (Table SI) (26).
Array comparative genomic
hybridization analysis and genomic subgrouping
Microarray-based CGH analysis was performed on 38
samples using the 8×60 K Human Genome CGH (Agilent Technologies
#G4450A) or the Human Genome customized 8×60 K CGH + SNP Microarray
Kit (Agilent Technologies #G4885A), and CytoGenomics software
(Agilent Technologies) following the manufacturer's protocol. CNVs
were identified using CytoGenomics 5.4.0.11 with the ADM-2
algorithm under default settings, with a minimal absolute average
log ratio of 0.25 as the cut-off. Based on the chromosomal
aberration profiles, including 1p loss, 11q loss, 17q gain, and
MYCN amplification, each tumor was categorized into genomic
subgroups (GGs; partial/segmental, GG-P; whole/numerical, GG-W)
(21,27,28).
ATRX mRNA expression and mutation
analysis
ATRX mRNA expression was analyzed using
TaqMan qPCR (Table SI). ALT(+) NBs
that showed <70% (<0.12) of the lowest ATRX mRNA
expression in ALT(−) NBs were considered to have decreased
ATRX mRNA expression. The copy number alteration of the
ATRX gene was assessed by qPCR and/or customized 8 × 60 K
CGH + SNP analysis (Agilent Technologies #G4885A). The relative
value of ATRX genomic copy number in qPCR assay was
calculated as the ratio of ATRX/SMARCA1: ATRX
copy number ‘loss’ was defined as <0.66 in tumors. Mutation
analysis was performed using Sanger sequencing for ALT(+) NB and
GNB without ATRX deletion (Table SI). RT-PCR products were generated
using the following primer pairs: 1F/1R, 2F/2R, 3F/3R, and 4F/4R.
For sequencing of each RT-PCR product, the following primers were
used: RT-PCR 1F-1R product (RT-PCR1F, RT-PCR1R, Sequencing 1F-1,
1F-2, 1F-3, 1R-1, and 1R-2); RT-PCR 2F-2R product (RT-PCR2F,
RT-PCR2R, Sequencing 2F-1, 2F-2, 2R-1 and 2R-2); RT-PCR 3F-3R
product (RT-PCR3F, RT-PCR3R, Sequencing 3F-1, and 3R-1); RT-PCR
4F-4R product (RT-PCR4F, RT-PCR4R, Sequencing 4F-1, 4F-2, and
4F-3).
Statistical analyses
The patients were grouped according to various
biological and clinical aspects of the disease. The significance of
differences in characteristics between the groups was examined
using the χ2 or Fisher's exact test for categorical
variables. Comparisons between two groups of continuous variables
were performed using the Mann-Whitney U test. Comparisons among
three or more groups were performed using the Kruskal-Wallis test
followed by Dunn's multiple comparisons test. Statistical analyses
were conducted using GraphPad Prism version 6 (Dotmatics).
Two-sided P<0.05 was considered to indicate a statistically
significant difference.
Results
Analysis of ALT, TERT mRNA expression,
and TC in NB
The clinical information of NB (n=255), GNB (n=48),
and GN (n=18) is presented in Table
SII, and the TMM status, including ALT, TERT mRNA
expression, and TC, for each tumor, is shown in Fig. 1. ALT(+) and TERT-high were
mutually exclusive in all cases except for two NB samples and one
GNB sample. ALT(+) or TERT-high cases were classified as
TMM, excluding ‘ever-shorter telomeres’ characterized by long
telomeres without an abundance of C-circle formation associated
with TMM (29). The correlations
between ALT and TERT mRNA expression levels and age, INSS,
MYCN amplification status, risk classification, and DNA
ploidy status were analyzed. Of the 255 NBs, 38 (14.9%) were
ALT(+). ALT(+) cases were significantly more prevalent among
patients aged ≥18 months (P<0.0001), those with INSS stage 4
(P=0.0054), those in the HR group (P<0.0001), and those with
diploidy (P=0.0037) (Table I).
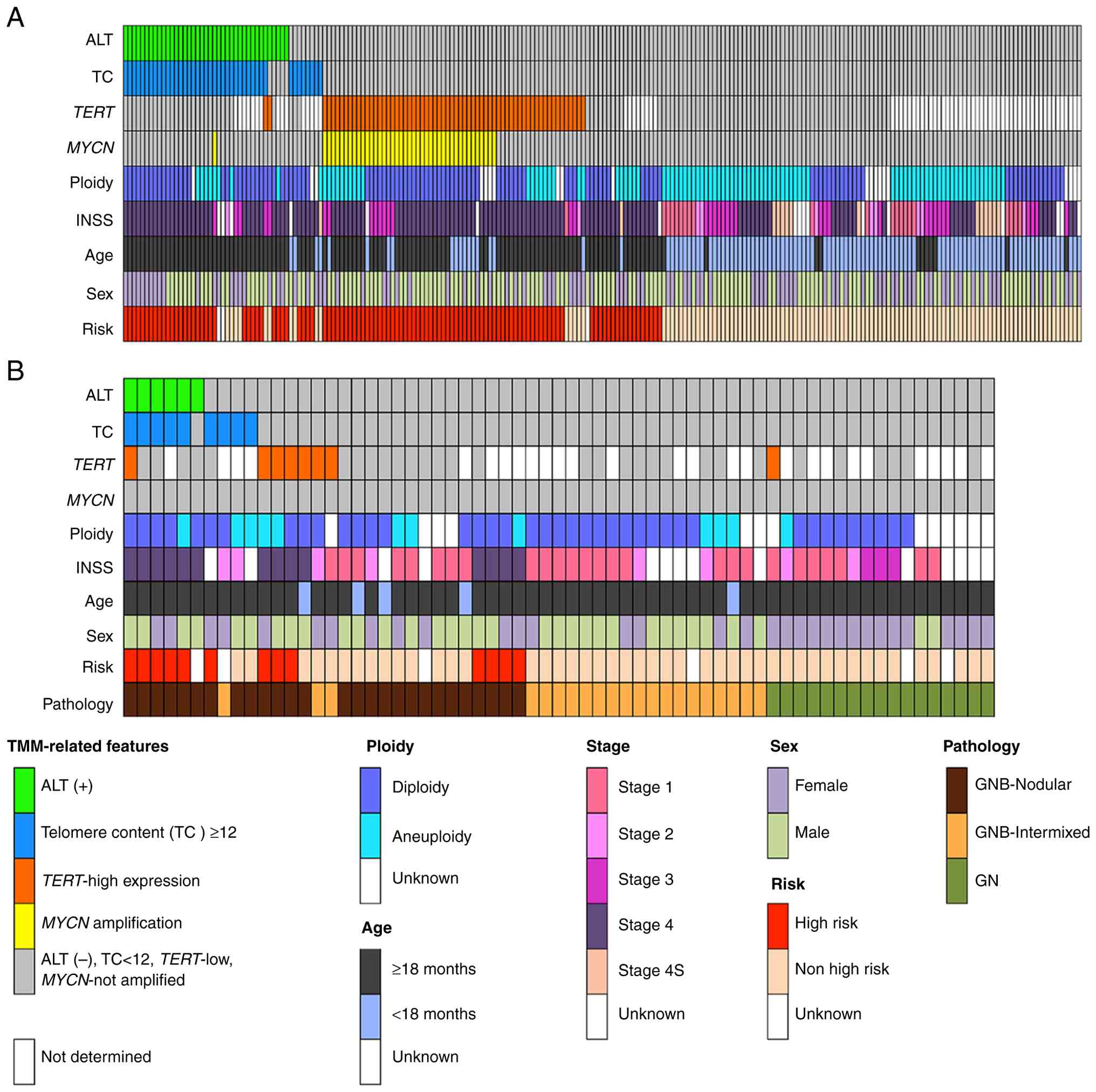 | Figure 1.Landscape of TMM and clinical and
pathological features of 255 NBs, 48 GNBs and 17 GNs. (A) NBs
(n=255), (B) GNBs (n=48), and GNs (n=17). MYCN status,
TERT mRNA expression, presence of C-circle (alternative
lengthening of telomeres), telomere content, and clinical and
pathological variables are shown. ALT, alternative lengthening of
telomeres; TC, telomere content; INSS, International Neuroblastoma
Staging System; TMM, telomere maintenance mechanisms; NB,
neuroblastoma; GNB, ganglioneuroblastoma; GN, ganglioneuroma. |
 | Table I.Patient characteristics of
alternative lengthening of telomeres and TERT expression status in
neuroblastoma cases. |
Table I.
Patient characteristics of
alternative lengthening of telomeres and TERT expression status in
neuroblastoma cases.
|
| ALT (n=255) | TERT
(n=169) |
|---|
|
|
|
|
|---|
| Characteristic | ALT(+) (n=38) | ALT(−) (n=217) | ALT(+), % | P-value | TERT-high
(n=64) | TERT-low
(n=105) | TERT-high,
% | P-value |
|---|
| Age |
|
|
|
|
|
|
|
|
| <18
months | 0 | 115 | 0.0 | <0.0001 | 14 | 55 | 20.3 | <0.0001 |
| ≥18
months | 38 | 102 | 27.1 |
| 50 | 50 | 50.0 |
|
| Sex |
|
|
|
|
|
|
|
|
|
Male | 19 | 129 | 12.8 | n.s. | 42 | 58 | 42.0 | n.s. |
|
Female | 19 | 88 | 17.8 |
| 22 | 47 | 31.9 |
|
| INSS |
|
|
|
|
|
|
|
|
| 1, 2 or
3 | 6 | 79 | 7.0 | 0.0054 | 16 | 37 | 30.2 | 0.013 |
| 4 | 30 | 117 | 20.4 |
| 48 | 55 | 46.6 |
|
| 4S | 0 | 15 | 0.0 |
| 0 | 7 | 0.0 |
|
|
Unknown | 2 | 6 | 25.0 |
| 0 | 6 | 0.0 |
|
| MYCN
statu |
|
|
|
|
|
|
|
|
| Not
amplified | 37 | 146 | 20.2 | <0.0001 | 23 | 93 | 19.8 | <0.0001 |
|
Amplified | 1 | 71 | 1.4 |
| 41 | 12 | 77.4 |
|
| Risk (INRG) |
|
|
|
|
|
|
|
|
|
Non-high risk | 5 | 108 | 4.4 | <0.0001 | 7 | 58 | 10.8 | <0.0001 |
| High
risk | 31 | 108 | 22.3 |
| 57 | 44 | 56.4 |
|
|
Unknown | 2 | 1 | 66.7 |
| 0 | 3 | 0.0 |
|
| Ploidy |
|
|
|
|
|
|
|
|
|
Aneuploidy | 8 | 92 | 8.0 | 0.037 | 19 | 46 | 29.2 | n.s. |
|
Diploidy | 29 | 102 | 22.1 |
| 39 | 50 | 43.8 |
|
|
Unknown | 1 | 23 | 4.2 |
| 6 | 9 | 40.0 |
|
The number of patients with TERT-high NB was
64 of 169 (37.9%). Patients with TERT-high were
significantly more prevalent among those aged >18 months
(P<0.0001), those with INSS stage 4 (P=0.013), and those in the
HR group (P<0.0001) (Table
I).
Regarding MYCN status, ALT(+) was enriched in
MYCN-not amplified NBs compared with MYCN-amplified
cases (P<0.0001). In an ALT(+) NB with MYCN
amplification, we observed the coexistence of MYCN-amplified
cells (approximately 10% of the population) and non-amplified tumor
cells on a FISH slide. This tumor was thought to be composed of a
mixture of ALT(+) and MYCN-amplified clones because previous
reports have described MYCN amplification and ALT(+) as
mutually exclusive (8,12,30).
TERT-high expression was more abundant in
MYCN-amplified cases than in MYCN-not amplified cases
(77.4% vs. 19.8%, P<0.0001), which is consistent with the fact
that the TERT gene is a transcriptional target of MYCN.
To assess the difference in telomere length between
ALT(+) and TERT-high NBs without MYCN amplification,
we compared the distribution of TC in four NB subgroups, ALT(+)
(n=35), TERT-high (n=21), TMM(−) HR group (n=10), and TMM(−)
non-HR group (n=56), excluding two cases with both ALT(+) and
TERT-high NB. The TC values in the ALT(+) NB group were
significantly higher than those in the other groups [ALT(+) vs.
TERT-high, TMM(−) HR, or TMM(−) non-HR, P<0.0001] and
were significantly lower in the TERT-high NB group compared
with the TMM(−) non-HR group (P=0.0015) (Fig. 2A).
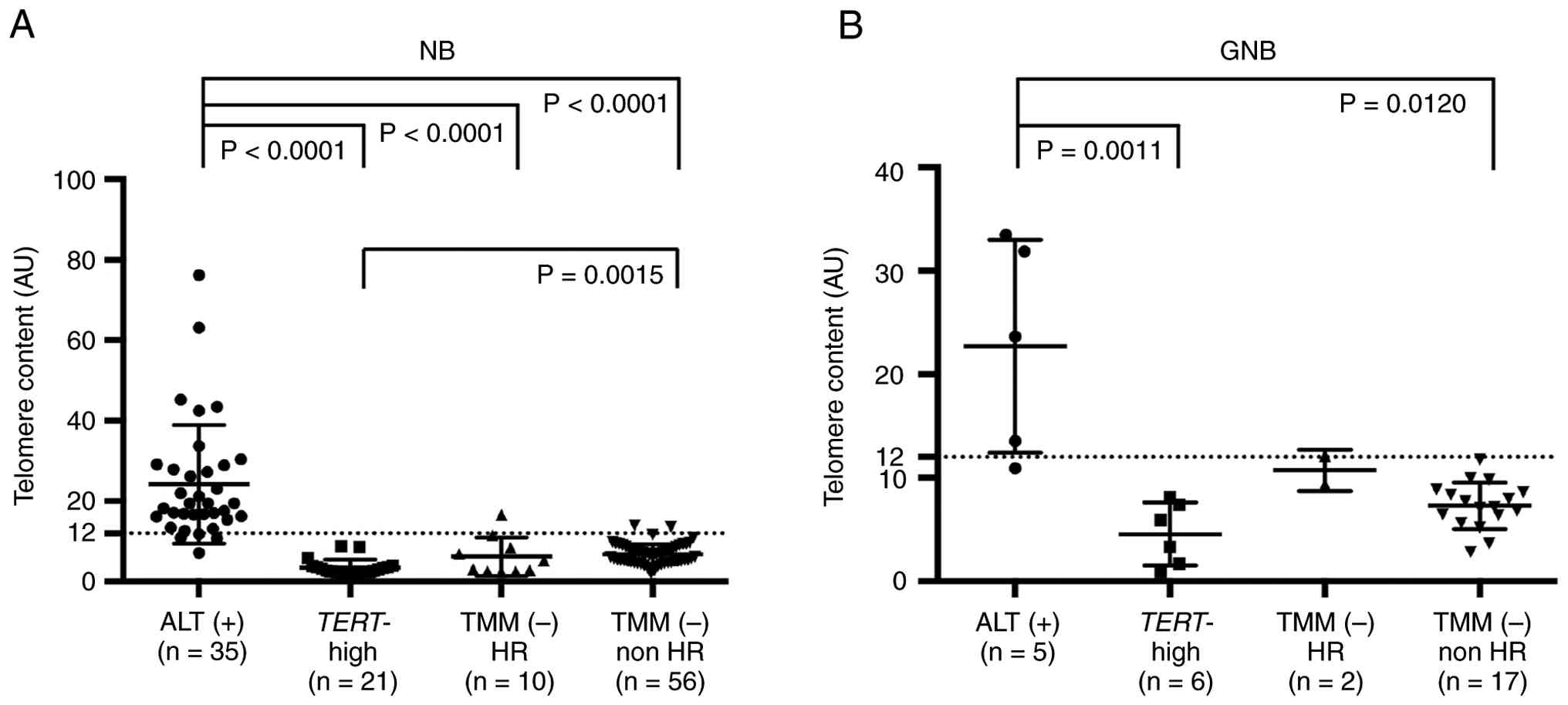 | Figure 2.TCs in MYCN-not-amplified
neuroblastic tumors. (A) Relative TC measured by qPCR in
neuroblastomas with ALT(+) (n=35), TERT-high (n=21), TMM(−)
HR (n=10), and TMM(−) non-HR (n=56). (B) Relative TC measured by
qPCR in ganglioneuroblastomas in ALT(+) (n=5), TERT-high
(n=6), TMM(−) HR (n=2), and TMM(−) non-HR (n=17). The horizontal
dot line demarcates TC=12. AU, arbitrary unit; TC, telomere
content; qPCR, quantitative PCR; ALT, alternative lengthening of
telomeres; TMM, telomere maintenance mechanisms; HR, high risk; NB,
neuroblastoma; GNB, ganglioneuroblastoma. |
Analysis of ALT, TERT mRNA expression,
and TC in GNB and GN
The C-circle assay (n=48) and TERT mRNA qPCR
(n=31) were performed in GNB, and the relationship between these
measurements and age, INSS, risk classification, histology, and DNA
ploidy status was analyzed. MYCN amplification was not
observed in any of the GNBs. We identified 6 ALT(+) cases (12.5%)
and 7 TERT-high cases (22.6%) among patients with GNB.
Patients with ALT(+) GNB comprised a significantly higher
proportion of those with INSS stage 4 (P=0.0018) and those in the
HR group (P=0.0285) (Table II),
whereas patients with TERT-high GNB made up a significantly
higher proportion of those with INSS stage 4 (P=0.0302). Owing to
the small sample size, no significant differences in age or ploidy
status were observed in either C-circle or TERT mRNA
measurements. Nodular tumor histology was significantly correlated
with ALT(+) GNB (P=0.0285).
 | Table II.Patient characteristics of
alternative lengthening of telomeres and TERT expression status in
ganglioneuroblastoma cases. |
Table II.
Patient characteristics of
alternative lengthening of telomeres and TERT expression status in
ganglioneuroblastoma cases.
|
| ALT (n = 48) | TERT (n =
31) |
|---|
|
|
|
|
|---|
| Characteristic | ALT (+) (n=6) | ALT (−) (n=42) | ALT (+), % | P-value | TERT-high
(n=7) | TERT-low
(n=24) | TERT-high,
% | P-value |
|---|
| Age |
|
|
|
|
|
|
|
|
| <18
months | 0 | 5 | 0 | n.s. | 1 | 2 | 33 | n.s. |
| ≥18
months | 6 | 37 | 14 |
| 6 | 22 | 21 |
|
| Sex |
|
|
|
|
|
|
|
|
|
Male | 4 | 27 | 13 | n.s. | 4 | 15 | 21 | n.s. |
|
Female | 2 | 15 | 12 |
| 3 | 9 | 25 |
|
| INSS |
|
|
|
|
|
|
|
|
| 1, 2 or
3 | 0 | 30 | 0 | 0.0018 | 2 | 17 | 11 | 0.0302 |
| 4 | 5 | 9 | 36 |
| 5 | 5 | 50 |
|
|
Unknown | 1 | 3 | 25 |
| 0 | 2 | 0 |
|
| MYCN
status |
|
|
|
|
|
|
|
|
| Not
amplified | 6 | 42 | 13 | - | 7 | 24 | 23 | - |
|
Amplified | 0 | 0 | - |
| 0 | 0 | - |
|
| Risk (INRG) |
|
|
|
|
|
|
|
|
|
Non-high risk | 0 | 31 | 0 | 0.0012 | 3 | 17 | 15 | n.s. |
| High
risk | 5 | 8 | 38 |
| 4 | 5 | 44 |
|
|
Unknown | 1 | 3 | 25 |
| 0 | 2 | 0 |
|
| Ploidy |
|
|
|
|
|
|
|
|
|
Aneuploidy | 1 | 9 | 10 | n.s. | 2 | 5 | 29 | n.s. |
|
Diploidy | 5 | 28 | 15 |
| 4 | 16 | 20 |
|
|
Unknown | 0 | 5 | 0 |
| 1 | 3 | 25 |
|
| Histology |
|
|
|
|
|
|
|
|
|
Nodular | 6 | 21 | 22 | 0.0285 | 5 | 15 | 25 | n.s. |
|
Intermixed | 0 | 21 | 0 |
| 2 | 9 | 18 |
|
Similar to NB, four subgroups based on the status of
ALT and TERT expression were established for analysis:
ALT(+) (n=5), TERT-high (n=6), TMM(−) HR (n=2), and TMM(−)
non-HR (n=17), excluding one case with both ALT(+) and
TERT-high NB. Owing to the small sample size, the TC values
in the TMM(−) HR group did not significantly differ from those in
the ALT(+) group; however, TC values were significantly higher in
the ALT(+) group compared with the TERT-high and TMM(−)
non-HR groups [ALT(+) vs. TERT-high or TMM(−) non-HR,
P=0.0011 or P=0.0120, respectively] (Fig. 2B).
All 18 GNs were ALT(−), but TERT-high was
observed in one case diagnosed at 10 years and 7 months [1 of 8
cases (12.5%)].
Analysis of the mechanism of high TERT
gene expression
Increased TERT expression is regulated by
MYCN amplification, mutations in TERT promoter
region, or TERT rearrangements, which result in
super-enhancer hijacking. To further investigate the mechanism in
each TERT-high tumor, 23 NBs, 7 GNBs, and 1 GN exhibiting
TERT-high without MYCN amplification were analyzed by
FISH to detect TERT rearrangements. Structural abnormalities
of the TERT gene were identified in 14 NBs (60.9%, 13
rearrangements, and 1 amplification) and 1 GNB (16.7%,
rearrangement), but not in GN. In one case of NB without
TERT rearrangement, a mutation (C228T) in the promoter
region of the TERT gene was observed using Sanger sequencing
(Case #144; Table III; Fig. S1). No mutations in the TERT
promoter region were observed in GNB and GN. These results are
consistent with previous reports, suggesting that mutations or
structural abnormalities in the TERT promoter region are the
main mechanisms of TERT-high MYCN-not amplified NBs,
whereas they are less frequent in TERT-high GNBs. This
suggests that GNB utilizes other mechanisms to achieve
TERT-high phenotype. Based on this notion, we further
performed CGH analysis to explore whether common chromosomal
aberrations occurred in 12 NTs (5 NB cases, 6 GNB cases, and 1 GN
case) with TERT-high but no promoter mutations using
sequencing or TERT rearrangement using FISH (Fig. S2). Among the 12 cases, 3 GNBs and 1
GN showed no chromosomal abnormalities, and no additional copy
number changes in the TERT genomic region on chromosome 5
were detected in the remaining 8 tumors. A partial (segmental) gain
of 17q was observed in 7 of the 12 cases, and a partial loss of 11q
was observed in 6 of the 12 cases. Additionally, a gain of 7q was
observed in 5 of the 12 cases (whole q-arm gain in 4 cases and
partial gain in 1 case), and a partial gain of 11q was observed in
5 of the 12 cases. The genome group (GG) based on the previously
proposed CGH signature showed that 7 of the 12 cases were
classified as segmental chromosomal alterations (P3s and P1s),
which are correlated with poor prognosis in patients with NB
(21,27,28,31).
 | Table III.TERT abnormalities in
MYCN-not-amplified, TERT-high neuroblastoma and
ganglioneuroblastoma. |
Table III.
TERT abnormalities in
MYCN-not-amplified, TERT-high neuroblastoma and
ganglioneuroblastoma.
| A, NB |
|---|
|
|---|
| Case no. | INSS stage | Age, months | TERT
expression, TERT/ACTB | TERT genomic
abnormality |
|---|
| 3 | 4 | 18 | 0.37 | Rearrangement |
| 14 | 2 | 27 | 1.65 | Rearrangement |
| 61 | 4 | 66 | 0.51 | Rearrangement |
| 92 | 4 | 34 | 1.57 | Rearrangement |
| 93 | 4 | 47 | 1.35 | Rearrangement |
| 142 | 3 | 42 | 0.7 | Rearrangement |
| 156 | 4 | 48 | 0.62 | Rearrangement |
| 234 | 4 | 48 | 7.74 | Rearrangement |
| 237 | 4 | 76 | 1.45 | Rearrangement |
| 251 | 4 | 36 | 2.81 | Rearrangement |
| 285 | 4 | 40 | 0.48 | Rearrangement |
| 317 | 4 | 36 | 0.61 | Rearrangement |
| 320 | 4 | 69 | 7.35 | Rearrangement |
| 180 | 4 | 42 | 1.33 | Amplification |
| 144 | 4 | 42 | 0.43 | Promoter
mutation |
| 31 | 4 | 22 | 0.55 | - |
| 178 | 4 | 17 | 2.36 | - |
| 246 | 4 | 37 | 1.7 | - |
| 292 | 3 | 29 | 0.43 | - |
| 295 | 3 | 56 | 1.59 | - |
| 196 | 4 | 62 | 0.64 | ND |
| 211 | 3 | 39 | 0.52 | ND |
| 240 | 1 | 19 | 0.44 | ND |
|
| B, GNB |
|
| Case
no. | INSS
stage | Age,
months | TERT
expression, TERT/ACTB | TERT
genomic abnormality |
|
| 192 | 4 | 36 | 6.55 | Rearrangement |
| 9 | 4 | 57 | 0.71 | - |
| 151 | 4 | 14 | 0.49 | - |
| 205 | 1 | 44 | 0.41 | - |
| 230 | 4 | 39 | 0.76 | - |
| 267 | 4 | 66 | 0.58 | - |
| 277 | 2 | 32 | 1.24 | - |
Analysis of ATRX genetic alterations
in ALT (+) NB and GNB
ALT(+) NBs frequently exhibit ATRX gene
alterations (15,16). To assess ATRX status, we
performed genomic qPCR targeting ATRX exon 9, array CGH
using a custom CGH + SNP array, and TaqMan qPCR for ATRX
mRNA expression. Copy number loss of ATRX was detected in at
least 25 of 38 ALT(+) NBs (65.8%, Table IV). Additionally, 16 of the 32
ALT(+) NBs (50%, Table IV) showed
decreased ATRX mRNA expression (relative expression value
<0.12). Among the 25 cases with ATRX deletion, 13 (13/19,
68.4%) exhibited decreased ATRX mRNA expression, whereas 5
were not analyzed owing to the unavailability of frozen samples.
Three NBs showed decreased ATRX mRNA (exon 9) expression
despite retaining the ATRX (exon 9) region. Sanger
sequencing of the ATRX region revealed at least two deletion
mutations, c.5162del and c.4356_4361del, in both cases (Fig. S3A and B). Collectively, 29 of 36
ALT(+) NBs (80.5%) exhibited ATRX genetic alterations in the
sample set.
 | Table IV.ATRX status in alternative
lengthening of telomeres (+) neuroblastoma and
ganglioneuroblastoma. |
Table IV.
ATRX status in alternative
lengthening of telomeres (+) neuroblastoma and
ganglioneuroblastoma.
| A, NB |
|---|
|
|---|
| Case no. | INSS stage | Age, months | Sex | ATRX mRNA
expression, ATRX/ACTB | Genomic DNA qPCR
for ATRX exon 9 | ATRX genomic
abnormality by CGH | ATRX
sequencing | ATRX
aberration status |
|---|
| 128 | 4 | 45 | M | 0.02 | Loss | Loss | Normal | Mutation |
| 75 | 4 | 68 | M | 0.02 | Loss | Loss | ND | Mutation |
| 125 | 4 | 50 | M | 0.02 | Loss | Loss | Normal | Mutation |
| 63 | 4 | 45 | M | 0.04 | Loss | Loss | ND | Mutation |
| 49 | 4 | 58 | M | 0.05 | Loss | Loss | Normal | Mutation |
| 157 | 4 | 86 | M | 0.07 | Retain | Loss | Normal | Mutation |
| 263 | 4 | 43 | M | 0.08 | Loss | Loss | Normal | Mutation |
| 47 | 4 | 41 | M | 0.13 | Loss | ND | Normal | Mutation |
| 64 | 4 | 41 | M | ND | Loss | Retain | Normal | Mutation |
| 105 | 4 | 30 | M | ND | Loss | Loss | ND | Mutation |
| 271 | 4 | 56 | M | ND | Loss | Loss | ND | Mutation |
| 80 | 4 | 51 | M | 0.1 | Retain | Chr. X loss | Normal | Mutation |
| 113 | 4 | 88 | M | 0.09 | Retain | Retain | Normal | Low expression |
| 13 | 4 | 121 | M | 0.1 | Retain | Retain | Normal | Low expression |
| 282 | - | 45 | M | 0.37 | Retain | Retain | Normal | Normal |
| 89 | 4 | 28 | M | 0.38 | Retain | Retain | Normal | Normal |
| 268 | 4 | 120 | M | 0.51 | Retain | Retain | Normal | Normal |
| 212 | 2 | 76 | M | 0.23 | Retain | ND | Normal | ND |
| 142 | 3 | 42 | M | 0.28 | Retain | Retain | ND | ND |
| 18 | 4 | 59 | F | 0.02 | Loss | Chr. X
loss/Loss | Normal | Mutation |
| 217 | 4 | 71 | F | 0.06 | Loss | Chr. X
loss/Loss | Normal | Mutation |
| 51 | 4 | 56 | F | 0.1 | Loss | Chr. X
loss/Loss | Normal | Mutation |
| 295 | 3 | 56 | F | ND | Retain | Chr. X
loss/Loss | ND | Mutation |
| 259 | 4 | 56 | F | ND | Retain | Chr. X
loss/Loss | ND | Mutation |
| 79 | 4 | 91 | F | 0.02 | Loss | Loss/Retain | ND | Mutation |
| 171 | 3 | 73 | F | 0.14 | Retain | Loss/Retain | Normal | Mutation |
| 243 | 4 | 49 | F | 0.17 | Retain | Loss/Retain | Normal | Mutation |
| 36 | 4 | 45 | F | 0.49 | Retain | Loss/Retain | Normal | Mutation |
| 226 | - | 95 | F | 0.17 | Retain | Chr. X
loss/Retain | c.5162del | Mutation |
| 311 | 3 | 41 | F | ND | Retain | Chr. X
loss/Retain | Normal | Mutation |
| 257 | 3 | 49 | F | 0.08 | Retain | Chr. X
loss/Retain | ND | Mutation |
| 207 | 4 | 39 | F | 0.3 | Retain | Chr. X
loss/Retain | Normal | Mutation |
| 294 | 4 | 37 | F | 0.27 | Retain | Retain | c.4356_4361del | Mutation |
| 96 | 4 | 48 | F | 0.03 | Retain | Retain | ND | Low expression |
| 273 | 4 | 42 | F | 0.42 | Retain | Retain | Normal | Normal |
| 45 | 4 | 71 | F | 0.44 | Retain | Retain | Normal | Normal |
| 6 | 4 | 75 | F | 3.04 | Retain | Retain | Normal | Normal |
| 140 | 4 | 20 | F | 0.27 | Retain | Chr.X gain | Normal | Normal |
|
| B, GNB |
|
| Case
no. | INSS
stage | Age,
months | Sex | ATRX mRNA
expression, ATRX/ACTB | Genomic DNA qPCR
for ATRX exon 9 | ATRX
genomic abnormality by CGH | ATRX
sequencing | ATRX
aberration status |
|
| 214 | 4 | 39 | M | 0.02 | Loss | Loss | Normal | Mutation |
| 258 | - | 40 | M | 0.05 | Loss | Loss | Normal | Mutation |
| 77 | 4 | 25 | M | 0.26 | Loss | Loss | Normal | Mutation |
| 230 | 4 | 39 | M | 0.79 | Retain | Retain | c.2518dup | Mutation |
| 86 | 4 | 53 | F | 0.09 | Loss | Loss/Retain | c.2518dup | Mutation |
| 118 | 4 | 43 | F | 0.11 | Loss | Chr. X
loss/Loss | ND | Mutation |
To date, no ATRX gene abnormalities have been
reported in GNBs; therefore, we performed a detailed genome
analysis of GNBs. We searched for ATRX mutations in all six
ALT(+) GNBs using genomic qPCR, CGH + SNP array, and Sanger
sequencing. Because ATRX is located on the X chromosome,
boys generally have only one copy of the ATRX gene. In three
of the four boys, copy number loss of ATRX was identified by
both qPCR and CGH +SNP array, and expression analysis confirmed a
marked decrease in ATRX mRNA expression in two cases (boys
#214 and #258). In the case without ATRX deletion (boy
#230), c.2518dup was identified (Fig.
S3C). In one girl with GNB, a deletion of one X chromosome and
an internal deletion of ATRX (exon 8–12 deletion) in the
remaining allele were detected using qPCR and CGH+SNP array
analysis (girl #118). In another girl with GNB, ATRX
deletion in one allele (c.178_2220del2023) was confirmed by qPCR
and CGH+SNP array analysis, and c.2518dup was detected by Sanger
sequencing (girl #86) (Fig. S3D).
Thus, ATRX abnormalities were identified in all six ALT (+)
GNBs (Table IV).
Chromosomal aberrations in ALT(+)
NTs
Copy number analysis was subsequently performed on
36 ALT(+) NB cases and 6 ALT (+) GNB cases to identify the
characteristic chromosomal aberrations in ALT(+) NTs. A whole
and/or partial chromosomal gain of 17q was observed in 41 (97.6%)
cases, and a partial loss of 11q was observed in 37 (88.1%) cases
(Fig. S3). Other frequently
observed chromosomal aberrations included partial and overall gains
of 7q (n=35, 83.3%), 7p (n=21, 50.0%), 11p (n=19, 45.2%), and 18q
(n=23, 54.8%) and a partial gain of 2p (n=23, 54.8%). Based on the
CGH-based genome group classifications (21,26,27,30),
41 (97.6%) tumors were classified as segmental/partial types (P3s,
P2s, and P4s), and 1 was classified as numerical/whole type (W5s)
(Fig. S4). Compared with
TERT-high ALT(−) tumors, ALT(+) NTs appeared to exhibit a
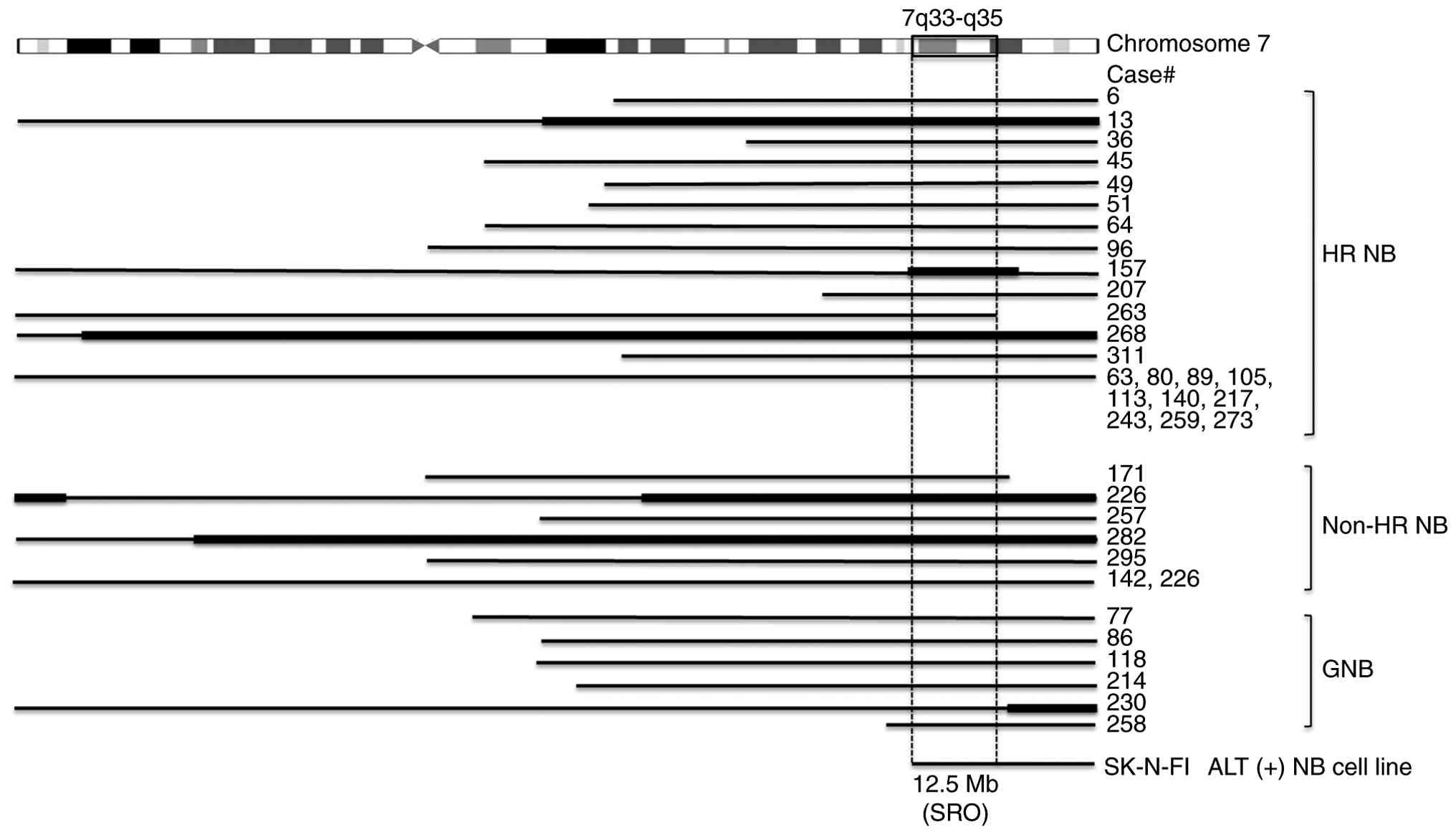
higher frequency of copy number alterations (Figs. S2 and S4). Furthermore, we analyzed the 7q gain
region in ALT(+) NTs, which was the third most frequent alteration
after 17q gain and 11q loss. We also observed a 7q gain in
TERT-high ALT(−) NTs; however, most of these cases exhibited
a whole-arm gain of 7q, rather than the partial (segmental) gains
observed in ALT(+) NTs. We identified a common 12.5 Mb region of
overlap (hg19: chr7:131,876,751-144,419,786) involving at least 115
gene IDs from the University of California Santa Cruz database
(Fig. 3).
Discussion
In the present study, high expression of TERT
mRNA and ALT as TMM in NB was observed more frequently in the older
age group (>18 months), INSS stage 4, and the HR group, with
statistical significance, as in previous reports (15,16).
There have been no reports analyzing TMMs in multiple GNB and GN
samples.
Multiple approaches have been proposed for detecting
ALT, including the C-circle assay, telomere length heterogeneity
(TLH), and ALT-associated PML bodies (APBs), and ongoing debate
remains as to which individual marker, or combination of markers,
most accurately reflects ALT activation and its clinical relevance.
Although the C-circle assay captures only extrachromosomal C-circle
structures and does not encompass other ALT-associated features
such as TLH or APBs, it provides high sensitivity and specificity
and is widely used across tumor types to quantitatively assess ALT
activity. Likewise, although TERT mRNA levels do not
directly measure telomerase enzymatic activity as the TRAP assay
does, TRAP analysis requires fresh tumor tissue, limiting the
number of evaluable cases. For this reason, TERT mRNA
expression has been widely adopted as a practical surrogate in TMM
studies.
In this study, 6 of the 48 (12.5%) GNB cases were
ALT(+) and 7 of the 31 (22.6%) cases were TERT-high. In
total, at least 25% (12/48) of the GNBs were TMM(+), and 10 (83.3%)
were nodular, a subgroup with aggressive histological features. As
additional reference, a recently published meta-analysis that
aggregated GNB cases extracted from multiple reports (total n=12,
with heterogeneous ALT assessment methods) reported that 75% (9 of
12) of nodular GNBs were ALT-positive (32). Our study represents the first
systematic analysis of ALT in a sizable GNB cohort using a single
standardized assay (the C-circle method). GNB and GN are extremely
rare tumors, and the numbers of GNB and GN cases analyzed in this
study were relatively limited. This limitation may reduce the
statistical power, particularly in subgroup analyses. Therefore,
further studies with larger sample sizes will be required to more
robustly determine the clinical and biological significance of TMM
in GNB and GN.
This study also showed a high frequency of
ATRX gene alterations in ALT(+) GNB with a nodular subtype,
although ATRX gene alteration is not the only mechanism of
ALT. Analysis of 16 cases of GN revealed no ALT(+) cases and only
one case exhibited TERT-high expression.
Given that TMM activation has been strongly
associated with poor prognosis in both HR and non-HR-NB (8,11,12,14–16),
it will be important to investigate whether TMM-positive GNB and GN
cases also exhibit adverse clinical outcomes. Because the tumors
analyzed in this study were primary specimens obtained at initial
diagnosis, a solid evaluation of the clinical significance of TMM
status in GN and GNB will require several additional years of
long-term follow-up. Further analyses with larger sample sizes and
more complete prognostic data, including GNB and GN cases, will be
essential to determine the clinical implications of TMM positivity
in these tumors.
In NB, TERT-high is triggered by MYCN
amplification and TERT gene rearrangement (8). In this study, genomic abnormalities in
the TERT gene were observed in 15 of 23 NBs with
TERT-high, indicating that this is a responsible mechanism.
In contrast, among GNB cases, six out of seven with
TERT-high showed no genomic abnormality of the TERT
gene, suggesting that TERT-high is induced by an alternative
mechanism distinct from that in NB. The expression of TERT
is regulated by multiple transcriptional and epigenetic mechanisms
beyond genomic alterations (33,34).
Among these, c-MYC has been reported to directly bind to the
TERT promoter and activate its transcription (35). In addition, NF-κB signaling can
enhance TERT transcription through promoter binding and
cooperation with c-MYC under stress or inflammatory conditions
(36). BRD4, a
bromodomain-containing chromatin reader, also contributes to TERT
activation by maintaining an open chromatin structure at
super-enhancer regions (37). These
findings raise the possibility that, in NT, TERT
upregulation is driven predominantly by epigenetic activation and
transcriptional deregulation rather than by structural alterations
of the TERT locus. Further integrative analyses
incorporating chromatin accessibility and histone modification
profiling will be necessary to elucidate these alternative
TERT regulatory mechanisms.
We performed CGH analysis based on the idea that
unknown genomic abnormalities in the region encoding regulators of
TERT gene expression may occur in NTs that exhibit
TERT-high expression in the absence of TERT gene
abnormalities and MYCN amplification. Despite the high
frequency of 11q deletions and 17q gains, we were unable to
identify appropriate candidates because of similar findings in
ALT(+) tumors exhibiting TERT-low expression. Comprehensive
analyses involving larger sample sizes are necessary to elucidate
the mechanisms underlying TERT-high NBs/GNBs independent of
TERT gene abnormalities and MYCN amplification.
CGH analysis of ALT(+) NTs showed a gain of 17q and
a partial loss of 11q in many cases. These alterations were also
frequently detected in ALT(−) NTs, suggesting that ALT occurs after
17q gains and 11q partial deletions. Although 1q42.2–1 qter
deletions have been previously reported in some ALT(+) tumors
(20), they were rare in our series
(2 of 35 tumors). We specifically focused on 7q gains, which were
the third most frequent alteration after 17q gain and 11q loss in
ALT(+) NTs. It has been observed that 7q gain, but not entire
chromosome 7 gain, contributes to either tumorigenesis or
progression in NBs (38). Moreover,
recurrent gain at 7q21.2-tel was observed in MYCN-not
amplified NB metastases compared with primary tumors (39). These reports may explain the poor
prognostic nature of ALT(+) tumors, which are susceptible to
segmental gain in the 7q region. The 12.5 Mb region of the common
7q gains may serve as a useful prognostic marker in ALT(+) NTs.
Regarding the clinical implications of TMMs,
activation of the TMM by TERT-high has been reported in
various cancers, leading to the development of molecularly targeted
drugs for telomerase inhibition. Recent studies have demonstrated
that 6-thio-2´deoxyguanosine, BET bromodomain inhibitors (such as
JQ1, AZD5153, and OTX015), dinaciclib, and carfilzomib inhibit the
growth of NB cells and/or NB xenografts exhibiting TERT
overexpression or high telomerase activity. These findings suggest
the need for the initiation of clinical trials targeting NTs with
TERT overexpression or high telomerase activity (15,40–42).
In conclusion, the development of molecularly
targeted drugs for ALT(+) tumors is important. In ALT(+) NB, the
synergistic effect of AZD0156, an ATM inhibitor, used in
combination with temozolomide and irinotecan, was reported both
in vitro and in vivo (43). Previous studies have demonstrated
the sensitivity of tumors with ALT(+) or ATRX mutations to
PARP inhibitors (44,45).
From a translational perspective, assay selection
must also take into account the availability of clinical specimens
and methodological reproducibility across laboratories. While
further comparative studies are needed to establish the most
comprehensive and reliable TMM evaluation strategies, the combined
use of the C-circle assay and TERT mRNA analysis currently
represents a robust and widely utilized framework for molecular TMM
assessment in NTs. Importantly, by applying this standardized
approach to a large cohort, including in GNB and GN, our study
provides new biological insights and contributes foundational data
that may inform future refinements of INRG risk stratification and
the clinical integration of TMM markers.
In conclusion, by expanding TMM analysis, whose
role as a high-risk biomarker in NB is now supported by
increasingly strong evidence, to GNB and GN, our findings may
support a more precise update of the INRG risk classification
across the full spectrum of NTs, including non-high-risk types. In
addition, given that several therapeutic approaches have been
proposed for TMM-positive tumors, incorporating TMM status into the
molecular diagnostic framework for NTs, including GNB and GN, may
help guide more tailored and effective treatment strategies in
future clinical practice.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
The authors would like to thank Ms. Tomoko
Yanagisawa, Ms. Akiyo Yamashita, Ms. Hiroko Odagawa and Ms. Fumiyo
Fukui (Saitama Cancer Center Research Institute, Saitama, Japan)
for their technical assistance, and Dr Matthias Fischer (Department
of Experimental Pediatric Oncology, University Children's Hospital
of Cologne, Germany) for providing SK-N-FI cells.
Funding
The Saitama Cancer Center provided the financial support for
this study. This study was partly supported by the Practical
Research for Innovative Cancer Control (No. 19ck0106468s0301, No.
19ck0106332s0203), Tailor-made Medical Treatment Program (No.
19cm0106603s0103) of the Japan Agency for Medical Research and
Development, Japan, and Japan Society for the Promotion of Science
KAKENHI Grant-in-Aid for Scientific Research (B), Japan
(JP19H03625), and Grant-in-Aid from the National Cancer Center
Research and Development Fund.
Availability of data and materials
Sanger sequencing data generated in the present
study may be found in the DNA Data Bank of Japan and ClinVar under
accession numbers LC911903-LC911906 and SCV007329794-SCV007329796,
respectively, or at the following URLs: LC911903, https://getentry.ddbj.nig.ac.jp/
getentry/na/LC911903/?format=flatfile&filetype=html&trace=
true&show_suppressed=false&limit=10;
LC911904, https://getentry.ddbj.nig.ac.jp/getentry/na/LC911904/?format=
flatfile&filetype=html&trace=true&show_suppressed=false&
limit=10;
LC911905, https://getentry.ddbj.nig.ac.jp/getentry/
na/LC911905/?format=flatfile&filetype=html&trace=true&show_suppressed=false&limit=10;
LC911906, https://getentry.
ddbj.nig.ac.jp/getentry/na/LC911906/?format=flatfile&filetype=
html&trace=true&show_suppressed=false&limit=10;
SCV007329794, http://www.ncbi.nlm.nih.gov/clinvar/
variation/4685457/?oq=SCV007329794&m=NM_000489.6
(ATRX):c.2518dup%20(p.Arg840fs);
SCV007329795, http://www.ncbi.nlm.nih.gov/clinvar/variation/4685458/?oq=SCV
007329795&m=NM_000489.6(ATRX):c.5162del%20
(p.Gly1721fs);
SCV007329796, http://www.ncbi.nlm.nih.
gov/clinvar/variation/4685459/?oq=SCV007329796&m=NM_
000489.6(ATRX):c.4353GGA%5B1%5D%20(p.Glu1463_
Glu1464del).
Other data generated in the present study may be requested from the
corresponding author.
Authors' contributions
TKa conceptualized and designed the study. MS, MH,
MO, JA, RS, RO, KA, AN and HO performed the laboratory experiments
and collected and analyzed the data. TT coordinated the collection
and provided clinical data through the data center. TI, TKo and SU
contributed to the provision of clinical samples and clinical
information as members of the clinical research group. TKa provided
resources. MS and MH drafted the original manuscript, with input
from TKa. MH, MO, TI, TKo, SU and TKa critically reviewed and
edited the manuscript for important intellectual content. MH and
TKa confirm the authenticity of all the raw data. All authors read
and approved the final manuscript, and agree to be accountable for
all aspects of the work.
Ethics approval and consent to
participate
Written informed consent was obtained from patients
and/or their guardians at hospitals participating in the Japan
Childhood Cancer Group Neuroblastoma Committee (JCCG-JNBSG). The
study design was approved by the Ethics Committee of Saitama Cancer
Center (approval nos. 1528 and 1529).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
ALT
|
alternative lengthening of
telomeres
|
|
CGH
|
comparative genomic hybridization
|
|
FISH
|
fluorescence in situ
hybridization
|
|
GN
|
ganglioneuroma
|
|
GNB
|
ganglioneuroblastoma
|
|
HR
|
high-risk
|
|
INRG
|
International Neuroblastoma Risk
Group
|
|
INSS
|
International Neuroblastoma Staging
System
|
|
JCCG
|
Japan Childhood Cancer Group
Neuroblastoma Committee
|
|
NB
|
neuroblastoma
|
|
NTs
|
neuroblastic tumors
|
|
TC
|
telomere content
|
|
TERT
|
telomerase reverse transcriptase
|
|
TMM
|
telomere maintenance mechanisms
|
References
|
1
|
Joshi VV and Silverman JF: Pathology of
neuroblastic tumors. Semin Diagn Pathol. 11:107–117.
1994.PubMed/NCBI
|
|
2
|
Shimada H, Ambros IM, Dehner LP, Hata JI,
Joshi VV and Roald B: Terminology and morphologic criteria of
neuroblastic tumors: Recommendations by the International
Neuroblastoma Pathology Committee. Cancer. 86:349–363. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Shimada H, Umehara S, Monobe Y, Hachitanda
Y, Nakagawa A, Goto S, Gerbing RB, Stram DO, Lukens JN and Matthay
KK: International neuroblastoma pathology classification for
prognostic evaluation of patients with peripheral neuroblastic
tumors: A report from the Children's Cancer Group. Cancer.
92:2451–2461. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Peuchmaur M, d'Amore ES, Joshi VV, Hata
JI, Roald B, Dehner LP, Gerbing RB, Stram DO, Lukens JN, Matthay KK
and Shimada H: Revision of the international neuroblastoma
pathology classification: Confirmation of favorable and unfavorable
prognostic subsets in ganglioneuroblastoma, nodular. Cancer.
98:2274–2281. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Brodeur GM and Bagatell R: Mechanisms of
neuroblastoma regression. Nat Rev Clin Oncol. 11:704–713. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Moschovi M, Arvanitis D, Hadjigeorgi C,
Mikraki V and Tzortzatou-Stathopoulou F: Late malignant
transformation of dormant ganglioneuroma? Med Pediatr Oncol.
28:377–381. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cohn SL, Pearson AD, London WB, Monclair
T, Ambros PF, Brodeur GM, Faldum A, Hero B, Iehara T, Machin D, et
al: INRG task force, The international neuroblastoma risk group
(INRG) classification system: An INRG task force report. J Clin
Oncol. 27:289–297. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Peifer M, Hertwig F, Roels F, Dreidax D,
Gartlgruber M, Menon R, Krämer A, Roncaioli JL, Sand F, Heuckmann
JM, et al: Telomerase activation by genomic rearrangements in
high-risk neuroblastoma. Nature. 526:700–704. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hiyama E, Hiyama K, Yokoyama T, Matsuura
Y, Piatyszek MA and Shay JW: Correlating telomerase activity levels
with human neuroblastoma outcomes. Nat Med. 1:249–255. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Killela PJ, Reitman ZJ, Jiao Y, Bettegowda
C, Agrawal N, Diaz LA, Friedman AH, Friedman H, Gallia GL,
Giovanella BC, et al: TERT promoter mutations occur frequently in
gliomas and a subset of tumors derived from cells with low rates of
self-renewal. Proc Natl Acad Sci USA. 110:6021–6026. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Cheung NK, Zhang J, Lu C, Parker M,
Bahrami A, Tickoo SK, Heguy A, Pappo AS, Federico S, Dalton J, et
al: Association of age at diagnosis and genetic mutations in
patients with neuroblastoma. JAMA. 307:1062–1071. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Valentijn LJ, Koster J, Zwijnenburg DA,
Hasselt NE, van Sluis P, Volckmann R, van Noesel MM, George RE,
Tytgat GAM, Molenaar JJ and Versteeg R: TERT rearrangements are
frequent in neuroblastoma and identify aggressive tumors, Nat.
Genet. 47:1411–1414. 2015.PubMed/NCBI
|
|
13
|
Akter J and Kamijo T: How do telomere
abnormalities regulate the biology of neuroblastoma? Biomolecules.
11:11122021. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ackermann S, Cartolano M, Hero B, Welte A,
Kahlert Y, Roderwieser A, Bartenhagen C, Walter E, Gecht J,
Kerschke L, et al: A mechanistic classification of clinical
phenotypes in neuroblastoma. Science. 362:1165–1170. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Roderwieser A, Sand F, Walter E, Fischer
J, Gecht J, Bartenhagen C, Ackermann S, Otte F, Welte A, Kahlert Y,
et al: Telomerase is a prognostic marker of poor outcome and a
therapeutic target in neuroblastoma. JCO Precis Oncol. 3:1–20.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Koneru B, Lopez G, Farooqi A, Conkrite KL,
Nguyen TH, Macha SJ, Modi A, Rokita JL, Urias E, Hindle A, et al:
Telomere maintenance mechanisms define clinical outcome in
high-risk neuroblastoma. Cancer Res. 80:2663–2675. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Maitra A, Yashima K, Rathi A, Timmons CF,
Rogers BB, Shay JW and Gazdar AF: The RNA component of telomerase
as a marker of biologic potential and clinical outcome in childhood
neuroblastic tumors. Cancer. 85:741–749. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Isobe K, Yashiro T, Omura S, Kaneko M,
Kaneko S, Kamma H, Tatsuno I, Takekoshi K, Kawakami Y and Nakai T:
Expression of the human telomerase reverse transcriptase in
pheochromocytoma and neuroblastoma tissues. Endocr J. 51:47–52.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Heaphy CM, Subhawong AP, Hong SM, Goggins
MG, Montgomery EA, Gabrielson E, Netto GJ, Epstein JI, Lotan TL,
Westra WH, et al: Prevalence of the alternative lengthening of
telomeres telomere maintenance mechanism in human cancer subtypes.
Am J Pathol. 179:1608–1615. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Hartlieb SA, Sieverling L, Nadler-Holly M,
Ziehm M, Toprak UH, Herrmann C, Ishaque N, Okonechnikov K,
Gartlgruber M, Park YG, et al: Alternative lengthening of telomeres
in childhood neuroblastoma from genome to proteome. Nat Commun.
12:12692021. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Tomioka N, Oba S, Ohira M, Misra A,
Fridlyand J, Ishii S, Nakamura Y, Isogai E, Hirata T, Yoshida Y, et
al: Novel risk stratification of patients with neuroblastoma by
genomic signature, which is independent of molecular signature.
Oncogene. 27:441–449. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ambros PF, Ambros IM, Brodeur GM, Haber M,
Khan J, Nakagawara A, Schleiermacher G, Speleman F, Spitz R, London
WB, et al: International consensus for neuroblastoma molecular
diagnostics: Report from the International Neuroblastoma Risk Group
(INRG) Biology Committee. Br J Cancer. 100:1471–1482. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Henson JD, Cao Y, Huschtscha LI, Chang AC,
Au AYM, Pickett HA and Reddel RR: DNA C-circles are specific and
quantifiable markers of alternative- lengthening-of-telomeres
activity. Nat Biotechnol. 27:1181–1185. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Farooqi AS, Dagg RA, Choi LMR, Shay JW,
Reynolds CP and Lau LMS: Alternative lengthening of telomeres in
neuroblastoma cell lines is associated with a lack of MYCN genomic
amplification and with p53 pathway aberrations. J Neurooncol.
119:17–26. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Akter J, Katai Y, Sultana P, Takenobu H,
Haruta M, Sugino RP, Mukae K, Satoh S, Wada T, Ohira M, et al: Loss
of p53 suppresses replication stress-induced DNA damage in
ATRX-deficient neuroblastoma. Oncogenesis. 10:732021. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Heidenreich B, Nagore E, Rachakonda PS,
Garcia-Casado Z, Requena C, Traves V, Becker J, Soufir N, Hemminki
K and Kumar R: Telomerase reverse transcriptase promoter mutations
in primary cutaneous melanoma. Nat Commun. 5:34012014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ohira M and Nakagawara A: Global genomic
and RNA profiles for novel risk stratification of neuroblastoma.
Cancer Sci. 101:2295–2301. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ohira M, Nakamura Y, Takimoto T, Nakazawa
A, Hishiki T, Matsumoto K, Shichino H, Iehara T, Nagase H,
Fukushima T, et al: Retrospective analysis of INRG clinical and
genomic factors for 605 neuroblastomas in Japan: A report from the
Japan Children's Cancer Group Neuroblastoma Committee (JCCG-JNBSG).
Biomolecules. 12:182021. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Dagg RA, Pickett HA, Neumann AA, Napier
CE, Henson JD, Teber ET, Arthur JW, Reynolds CP, Murray J, Haber M,
et al: Extensive proliferation of human cancer cells with
ever-shorter telomeres. Cell Rep. 19:2544–2556. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zeineldin M, Federico S, Chen X, Fan Y, Xu
B, Stewart E, Zhou X, Jeon J, Griffiths L, Nguyen R, et al: MYCN
amplification and ATRX mutations are incompatible in neuroblastoma.
Nat Commun. 11:9132020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Schleiermacher G, Mosseri V, London WB,
Maris JM, Brodeur GM, Attiyeh E, Haber M, Khan J, Nakagawara A,
Speleman F, et al: Segmental chromosomal alterations have
prognostic impact in neuroblastoma: A report from the INRG project.
Br J Cancer. 107:1418–1422. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Avinent-Pérez M, Westermann F, Navarro S,
López-Carrasco A and Noguera R: Tackling ALT-positive
neuroblastoma: Is it time to redefine risk classification systems?
A systematic review with IPD meta-analysis. Neoplasia.
60:1011062025. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ramlee MK, Wang J, Toh WX and Li S:
Transcription regulation of the human telomerase reverse
transcriptase (hTERT) gene. Genes (Basel). 7:502016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Sharma S and Chowdhury S: Emerging
mechanisms of telomerase reactivation in cancer. Trends in Cancer.
8:632–641. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhao Y, Cheng D, Wang S and Zhu J: Dual
roles of c-Myc in the regulation of hTERT gene. Nucleic Acids Res.
42:10385–10398. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Yin L, Hubbard AK and Giardina C: NF kappa
B regulates transcription of the mouse telomerase catalytic
subunit. J Biol Chem. 275:36671–366758. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Donati B, Lorenzini E and Ciarrocchi A:
BRD4 and cancer: going beyond transcriptional regulation. Mol
Cancer. 17:1642018. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Stallings RL, Howard J, Dunlop A,
Mullarkey M, McDermott M, Breatnach F and O'Meara A: Are gains of
chromosomal regions 7q and 11p important abnormalities in
neuroblastoma? Cancer Genet Cytogenet. 140:133–137. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Cobrinik D, Ostrovnaya I, Hassimi M,
Tickoo SK, Cheung IY and Cheung NK: Recurrent pre-existing and
acquired DNA copy number alterations, including focal TERT gains,
in neuroblastoma central nervous system metastases. Genet Chromosom
Cancer. 52:1150–1166. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Huang M, Zeki J, Sumarsono N, Coles GL,
Taylor JS, Danzer E, Bruzoni M, Hazard FK, Lacayo NJ, Sakamoto KM,
et al: Epigenetic targeting of TERT-Associated gene expression
signature in human neuroblastoma with TERT overexpression. Cancer
Res. 80:1024–1035. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Chen J, Nelson C, Wong M, Tee AE, Liu PY,
La T, Fletcher JI, Kamili A, Mayoh C, Bartenhagen C, et al:
Targeted therapy of TERT-rearranged neuroblastoma with BET
bromodomain inhibitor and proteasome inhibitor combination therapy.
Clin Cancer Res. 27:1438–1451. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Wood L, Huang M, Zeki J, Gong M, Taylor J,
Shimada H and Chiu B: Combining inhibitors of Brd4 and
cyclin-dependent kinase can decrease tumor growth in neuroblastoma
with MYCN amplification. J Pediatr Surg. 56:1199–1202. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Koneru B, Farooqi A, Nguyen TH, Chen WH,
Hindle A, Eslinger C, Makena MR, Burrow TA, Wilson J, Smith A, et
al: Reynolds, ALT neuroblastoma chemoresistance due to telomere
dysfunction-induced ATM activation is reversible with ATM inhibitor
AZD0156. Sci Transl Med. 13:57502021. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
George SL, Lorenzi F, King D, Hartlieb S,
Campbell J, Pemberton H, Toprak UH, Barker K, Tall J, da Costa BM,
et al: Therapeutic vulnerabilities in the DNA damage response for
the treatment of ATRX mutant neuroblastoma. EBiomedicine.
59:1029712020. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zimmermann M, Bernier C, Kaiser B,
Fournier S, Li L, Desjardins J, Skeldon A, Rimkunas V, Veloso A,
Young JTF, et al: Guiding ATR and PARP inhibitor combinations with
chemogenomic screens. Cell Rep. 40:1110812022. View Article : Google Scholar : PubMed/NCBI
|

















