Introduction
Cervical cancer (CC) is the fourth most common
cancer among women, with an incidence rate of ~10.9 per 100,000 in
China (1). In recent years, methods
of prevention, diagnosis and treatment of CC have improved notably,
but metastasis and drug resistance remain key causes of CC
treatment failure (2–5). Therefore, clarification of the
mechanisms of development of CC remains a key research goal so that
diagnostic and treatment strategies can be potentially enhanced in
order to improve clinical outcomes for patients with CC.
A potentially key mechanism in this context involves
the concept of Vasculogenic mimicry (VM), which was first proposed
by Maniotis et al (6) in
1999 in a study of uveal melanoma. VM leads to the development of
an endothelium-independent vascular system (7) in which a microcirculatory system is
formed by tumor cells themselves under a hypoxic environment
(8). The main process of formation
of this system involves cancer stem cells (CSC) undergoing
epithelial-endothelial transformation (EET). The resulting
endothelial cells generate a duct structure due to remodeling of
the extracellular matrix (ECM) and they ultimately fuse into
vessels. This process is favored in the hypoxic microenvironment,
where highly differentiated tumor cells gradually transform into
lower-differentiated CSC with high differentiation potential;
notably, these CSC have a decreased sensitivity to chemotherapy
agents and an increased susceptibility to the development of drug
resistance (9–13). The tube formed by CSC with partial
endothelial properties eventually fuses with endothelium-dependent
microvessels to transport oxygen and nutrients to hypoxic regions
of tumors.
The lack of endothelial cell coverage within the VM
channel means that the tumor cells are directly exposed to the
blood. This situation increases the likelihood that shed tumor
cells will metastasize through the bloodstream, leading to early
tumor metastasis (14–18). Therefore, VM is a key factor
associated with increased metastasis and drug resistance and it has
been reported to be associated with the occurrence, development and
poor prognosis of multiple types of cancer, including breast
(19), colorectal (20), prostate (21), hepatocellular (22), lung (23), ovarian (24), gastric (25) and bladder (26) cancer types.
Detailed molecular mechanisms underlying VM,
particularly in CC, require further exploration; however, several
molecular factors have been implicated in this process. For
example, enzymes in the matrix metalloproteinase (MMP) family have
been identified as being involved in VM due to their activities in
the remodeling of the ECM (27,28).
Another potential molecular factor is the
modification of RNA by the formation of 5-methylcytosine
(m5C) by methyltransferases. This change is a dynamic,
reversible and widely distributed epigenetic modification (29) that serves a variety of biological
functions (30,31). Notably in the context of VM, the
levels of m5C have been identified to increase in the
hypoxic tumor microenvironment, where the modification promotes
tumor growth, migration, angiogenesis and drug resistance. In
particular, the NOP2/Sun RNA methyltransferase 2 (NSUN2) has been
reported to be a key cancer-associated methyltransferase of this
modification (32–34). Certain studies have explored roles
for NSUN2 in CC (35,36); however, whether NSUN2 promotes the
malignant progression of CC by enhancing VM in the hypoxic tumor
microenvironment remains to be elucidated.
In the present study, bioinformatics and whole-slide
scanning of immunohistochemical images were combined to investigate
the association of VM with prognosis in CC. The present study also
used bioinformatics and expression assays to demonstrate that MMP-9
may be particularly key in VM formation in CC. In order to study
the mechanisms leading to increased expression levels of MMP-9, the
m5C modification of total RNA in CC was first quantified
under hypoxic conditions using dot blot assay. Subsequently, the
protein expression levels of NSUN2 was examined in hypoxia-induced
VM-competent CC cells via western blot. Finally, a RNA stability
and methylated-RNA immunoprecipitation assays were employed to
study the physical and functional relationship between NSUN2 and
MMP-9 mRNA.
Patients and methods
Clinical tissue specimens
In the present study, 44 CC tissue samples and
paired adjacent normal tissue samples were obtained during
surgeries (May to August 2024) performed at the Jiangsu Province
Geriatric Hospital, which is affiliated with Nanjing Medical
University (Nanjing, China). The cohort consisted entirely of
female patients, with a median age of 52 years (range, 42–68
years). For each patient, one sample of tumor tissue and one sample
of histologically normal adjacent cervical tissue (obtained ≥3 cm
from the tumor margin) were collected intraoperatively. The
diagnosis of CC and the state of paracancerous tissues were
confirmed by pathological evidence. All samples were immediately
snap-frozen in liquid nitrogen and stored at −80°C for at least 72
h prior to analysis. Patient inclusion criteria were as follows: i)
Histopathologically confirmed primary cervical carcinoma; ii)
scheduled for curative-intent surgery; iii) no prior radiotherapy,
chemotherapy, or other anticancer treatments; and iv) provision of
written informed consent for the use of resected tissue for
research. Patient exclusion criteria included: i) A history of any
other malignant tumor; ii) evidence of distant metastasis at
presentation; iii) receipt of neoadjuvant therapy; and iv)
insufficient quantity or poor quality of paired tissue samples as
confirmed by pathological review. The present study was approved by
the Ethics Committee of the Jiangsu Province Geriatric Hospital
[approval no. (2024) Court Ethics Opinion No. 028-1] and written
informed consent was obtained from all participants.
Cell culture and transfection
Normal cervical cells [human skin keratinocytes cell
line (HaCaT; cat. no. TCH-C388; Haixing Biosciences Co., Ltd.;
http://www.hycyte.com/sys-pd/133.html)] and CC cell
lines [human cervical cancer cell line (HeLa; cat. no. TCH-C193;
Haixing Biosciences Co., Ltd.; http://www.hycyte.com/sys-pd/65.html), human cervical
squamous cell line (SiHa; cat. no. TCH-C326; Haixing Biosciences
Co., Ltd.; http://www.hycyte.com/sys-pd/70.html), human CC cell
line with intestinal metastasis (CaSki; cat. no. TCH-C145; Haixing
Biosciences Co., Ltd.; http://www.hycyte.com/sys-pd/119.html), human CC cell
line (C33A; cat. no. TCH-C143; Haixing Biosciences Co., Ltd.;
http://www.hycyte.com/sys-pd/134.html) and human CC
cell line (HT-3; cat. no. CL-0630; Wuhan Punoise Life Technology
Co., Ltd.; http://www.procell.com.cn/p/ht-3-cl-0630-72192) were
cultured in DMEM supplemented with 10% fetal bovine serum (FBS)
purchased from Inner Mongolia Opcel Biotechnology Co., Ltd.
(https://www.nmjyk.com/?tnxq/169.html)
and penicillin/streptomycin (100 U/ml) in an incubator at 37°C with
5% CO2, Furthermore, cellular hypoxia was induced by
exposing the cells to an atmosphere of 5% CO2 and 0.1%
O2 for durations of 24, 48 and 72 h. All cell lines were
free of mycoplasma. The cell lines used were authenticated using
short tandem repeat analysis.
Lipofectamine® 2000 (Invitrogen; Thermo
Fisher Scientific, Inc.) was used for transfection of cells with
short hairpin RNA (shRNA) plasmid (10 µg; 1 µg/µl) and DNA
transfection reagent (Roche Diagnostics) was used for transfection
of cells with overexpression plasmids (4 µg; 1 µg/µl). Cells that
were cultured to 50–70% confluence in 6-well plates were
transfected according to the manufacturer's instructions in a 37°C
incubator. For shRNA plasmid transfection, the medium was changed 6
h after transfection, but the medium was not changed for
overexpression plasmid transfections. RNA was collected 24 h after
transfection and protein was collected 48 h after transfection.
Construction of plasmids and
interference sequences
The NSUN2-targeting shRNA plasmid was
purchased from GeneCopoeia (cat. no. CSHCTR001-CU6; GeneCopoeia,
Inc.) and its sequences were as follows: shNUSN2-1,
5′-CCCGGCCACTTTAAGAATTAC-3′; shNUSN2-2,
5′-CCTCATCATAAGATCTTAGAT-3′; and shNSUN2-3,
5′-GCGATGCCTTAGGATATTACC-3′. The negative control sequence was
5′-GCTTCGCGCCGTAGTCTTA-3′.
The full lengths of the human MMP-9
(NM_004994.3) and NSUN2 (NM_017755) open-reading frames, as
well as the full length open-reading frame of GAPDH, were
cloned into the plasmid cloning DNA3.1(+) vector (cat. no. V80020;
Invitrogen; Thermo Fisher Scientific, Inc.).
DNA and RNA extraction and reverse
transcription-quantitative PCR (RT-qPCR)
Plasmids containing the NSUN2 and
MMP-9 sequences were extracted from E. coli DH5α
using the Endo-free Plasmid Mini Kit II (Omega Bio-Tek Inc.)
according to the manufacturer's instructions. Total RNA was
extracted from CC cell lines or frozen tissue samples using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.) according to the manufacturer's protocol. RNA was reverse
transcribed into cDNA using the PrimeScript™ RT reagent Kit
(Perfect Real Time) Kit (cat. no. RR037A; Takara Bio, Inc.)
according to the manufacturer's protocol and the DNA was stored at
−20°C. Quantitative PCR was performed on an ABI StepOne Plus
Real-Time PCR System (Applied Biosystems; Thermo Fisher Scientific,
Inc.) with three replicates for each sample, using TB
Green® Premix Ex Taq™ (Tli RNaseH Plus) (cat. no.
RR420A; Takara Bio Inc.). The thermal cycling profile was as
follows: Initial denaturation at 95°C for 30 sec, 40 cycles of
denaturation at 95°C for 5 sec, followed by combined
annealing/extension at 60°C for 30–34 sec (fluorescence data
collection was performed at this step). Melt curve analysis was
performed from 60–95°C with a continuous fluorescence measurement
at a ramp rate of 0.3°C per sec. qPCR data were analyzed using the
comparative Cq (2−ΔΔCq) method (37). The primer sequences used were as
follows: NSUN2 F, 5′-GAACTTGCCTGGCACACAAAT-3′ and R,
5′-TGCTAACAGCTTCTTGACGACTA-3′; MMP-9 F,
5′-GGGACGCAGACATCGTCATC-3′ and R, 5′-TCGTCATCGTCGAAATGGGC-3′; and
GAPDH F, 5′-TGGTATGAGAGCTGGGGAATG-3′; and R,
5′-CCTCCCCACCTTGAAAGGAA-3′).
In vitro vasculogenic mimicry
assay
First, 50 µl thawed matrix gel solution (ABW
Matrigel; cat. no. 082704; Shanghai Nova Medical Technology Co.,
Ltd.; http://www.abwbio.com/content/show/593/34) were evenly
spread within each well of a 96-well plate. The plate was kept on
ice for the duration of the experiment (4°C; 30 min). Transfected
CC and control CC cells (3×104) were dropped into the
coagulated matrix glue. The plates were cultured at 37°C, 5%
CO2 for 18 h to allow tuber formation. Images were
captured at ×4 magnification using a light microscope (IX53;
Olympus Corporation). Image Pro (version 6.0; Media Cybernetics,
Inc.) was used to calculate the number of branch points. Each
experiment was performed in triplicate.
Migration and invasion assays
For migration assays, cells (5.0×104)
were suspended in 300 µl serum-free medium and then seeded in the
upper chamber of an 8-µm pore Transwell® insert
(Corning, Inc.). Subsequently, 700 µl fresh medium with 10% FBS
were added to the lower chamber of the Transwell insert and
incubated for 24 h in a 37°C incubator. For invasion assays, a 100
µl aliquot of Matrigel diluted 10-fold in PBS was added to the
upper chamber of a Transwell insert and allowed to solidify for 10
min in a 37°C incubator. cells (1.0×105) were suspended
in 300 µl serum-free medium and then seeded in the upper chamber of
an 8-µm pore Transwell® Matrigel insert (Corning, Inc.).
Subsequently, 700 µl fresh medium with 10% FBS were added to the
lower chamber of the Transwell insert and incubated for 24 h in a
37°C incubator. After incubation for 24 h at 37°C, the cells in the
upper chamber were fixed with 4% paraformaldehyde for 2 h at RT and
stained with 0.1% crystal violet for 1 h at RT. A light microscope
used for observation of migratory/invasive cells in the lower
chamber. The number of migrated cells were counted in 5 fields of
view (original magnification, ×10) using an Olympus IX53 microscope
(Olympus Corporation).
Western blotting
After collecting and lysing the cells, the protein
concentration in the soluble portion was determined using the BCA
protein assay (cat. no. C0050; TargetMol Chemicals Inc.). The
protein extraction buffer contained RIPA (cat. no. P0013B; Shanghai
Beyotime Biotechnology Co., Ltd.), protease inhibitor cocktail
(cat. no. C0001; TargetMol Chemicals Inc.) and PMSF (cat. no.
ST506-2; Shanghai Beyotime Biotechnology Co., Ltd.). Equal amounts
of protein (30 µg/lane) were separated using SDS-PAGE (10%) and
then transferred to methanol-activated 0.45 µm PVDF membranes. The
membranes were blocked with 5% milk at 20±5°C for 1 h. The blocked
membranes were incubated at 4°C overnight with the following
antibodies: A rabbit monoclonal anti-NSUN2 antibody (1:1,000; cat.
no. AB259941; Abcam), a rabbit monoclonal anti-transfer RNA
aspartic acid methyltransferase 1 (TRDMT1) antibody (1:1,000; cat.
no. 19221-1-AP; Proteintech Group, Inc.; Wuhan Sanying
Biotechnology), a rabbit polyclonal anti-MMP-9 antibody (1:1,000;
cat. no. 10375-2-AP; Proteintech Group, Inc.; Wuhan Sanying
Biotechnology), a rabbit polyclonal anti-aldehyde dehydrogenase 1
(ALDH1) antibody (1:1,000; cat. no. 15910-1-AP; Proteintech Group,
Inc.; Wuhan Sanying Biotechnology), a rabbit polyclonal anti-ephrin
type-A receptor 2 (EPHA2) antibody (1:1,000; cat. no. AF5 238;
Affinity Biosciences) and a rabbit polyclonal anti-GAPDH antibody
(1:1,000; TA309157 OriGene Technologies, Inc.). Following extensive
washing, the membranes were incubated at room temperature for 1 h
with an HRP-conjugated sheep anti-rabbit secondary antibody
(1:5,000; cat. no. A0208; Shanghai Beyotime Biotechnology Co.,
Ltd.). Following reaction with ECL reagent (cat. no. P0018S;
Shanghai Beyotime Biotechnology Co., Ltd.), immunoreactive proteins
bands were visualized using a Tanon™ 5200 chemiluminescence gel
imaging system (Tanon Science and Technology Co., Ltd.). The
results of the western blotting analyses were quantified using
ImageJ (version 1.53; National Institutes of Health) and normalized
to either the loading control or GAPDH.
Immunohistochemistry (IHC)
The standard staining protocol in identifying VM is
double staining with periodic acid-Schiff (PAS) and anti-CD31
antibodies (38,39). Tissue samples from patients with CC
or paracancerous tissues and a CC tissue microarray (cat. no.
HUteS140Su01; Shanghai Outdo Biotech Co., Ltd) were fixed and then
cut into 4 µm sections. Paraffin sections were dewaxed to water
using a dewaxing agent (three baths of 10 min each), followed by
gradient alcohol dehydration for 5 min per bath and finally rinsed
thoroughly using distilled water. The blocking process was
performed with 10% goat serum (cat. no. AR1009; Boster Biological
Technology) at room temperature for 30 min. The tissue samples were
treated with PH9.0 EDTA repair solution for antigen retrieval and
then treated with a rabbit polyclonal anti-CD31 antibody (1:2,000;
cat. no. AB76533; Abcam), a rabbit polyclonal anti-NSUN2 antibody
(1:200; cat. no. AB259941; Abcam) or a rabbit polyclonal anti-MMP-9
antibody (1:200; cat. no. 10375-2-AP; Proteintech Group, Inc.;
Wuhan Sanying Biotechnology). The sections were incubated overnight
at 4°C, washed and incubated with a secondary antibody. The
secondary antibody used was goat anti-rabbit HRP (1:2,000; cat. no.
ab205718; Abcam), incubated at 37°C for 50 min. To perform PAS
staining, the sections were first oxidized with periodic acid
solution for 10 min, rinsed thoroughly with distilled water and
blotted dry with absorbent paper. The sections were then stained
with Schiff's reagent for 10–15 min and rinsed under running tap
water for 10 min. The sections were then dehydrated through a
graded series of ethanol (95% and absolute ethanol), cleared with a
clearing agent and subsequently mounted. The stained sections were
imaged using an optical microscope (original magnification, ×20).
The microarrays were scanned using the Hamamatsu NanoZoomer S360
(Hamamatsu Photonics K.K.). H&E staining was performed using
reagents purchased from Hubei BIOSSCI Biotech Co., Ltd (cat. no.
BP0211). The staining procedure was carried out as follows: Tissue
sections were immersed in clearing solution for 10 min twice, with
excess liquid gently shaken off between each step. Sections were
rehydrated through a graded ethanol series: Absolute ethanol for 5
min, 95% ethanol for 5 min, 85% ethanol for 5 min and 75% ethanol
for 5 min, followed by rinsing in distilled water for 1 min.
Sections were stained with Harris hematoxylin solution for 4 min at
room temperature and then rinsed under tap water for 2 min until no
excess dye was released. Differentiation was performed using 0.8%
hydrochloric acid alcohol for 2 sec, followed by rinsing with tap
water. Sections were stained with alcohol-soluble eosin solution
for 20 sec at room temperature without rinsing, then treated with
95% ethanol for 5 min and dehydrated via two changes of absolute
ethanol (2 min each). Finally, sections were cleared in xylene and
mounted with neutral balsam for microscopic examination.
Methylated RNA immunoprecipitation
(IP)
Total RNA was extracted from CC cell lines using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.). The RNA (each sample contained 300 µg of RNA, with a
concentration of 1 µg/µl) was then fragmented at 94°C for 5 min
using fragmentation reagents (30 µl; cat. no. 3735438; Merck KGaA).
After stopping the fragmentation process with stop buffer and
vortexing, the RNA fragments were divided into two tubes, isolated
by centrifugation and dissolved in 100 µl diethylpyrocarbonate
(DEPC)-treated water. The RNA fragments were incubated with 10 µg
antibodies against m5C (cat. no. AB10805; Abcam) or 10
µg NSUN2 (cat. no. AB259941; Abcam) in methylated-RNA IP buffer (5X
IP buffer with RNase inhibitor) for 4 h at 4°C. 30 µl Protein A/G
(cat. no. 3835713; Merck KGaA) magnetic beads were added and the
mixture was incubated for an additional 4 h. The washing reagent
used was IP Buffer (1X), prepared by diluting IP Buffer (5X) (cat.
no. 3841555; Merck KGaA). Each complex was eluted with 100 µl of
Elution Buffer (1X), which was prepared by combining 90 µl of IP
Buffer (5X), 150 µl of m5C (20 mM stock; cat. no. 58366-64-6; Merck
KGaA), 7 µl of RNase inhibitor and 203 µl of molecular
biology-grade, RNase-free water. Subsequently, the complexes were
further eluted with an additional 100 µl of 1X IP Buffer. The
eluates were collected and precipitated by centrifugation at 15,000
× g for 30 min at 4°C. The precipitate RNA was dissolved in an
equal volume of DEPC-treated water and reversed transcribed into
cDNA using the PrimeScript™ RT reagent Kit (Perfect Real Time) Kit
(cat. no. RR037A; Takara Bio, Inc.). Quantitative PCR was used TB
Green® Premix Ex Taq™ (Tli RNaseH Plus) (cat. no.
RR420A; Takara Bio Inc.). The MMP-9 mRNA IP assay primers
used were as follows: 47–153 bp region F,
5′-GTGCTCCTGGTGCTGGGCTG-3′ and R, 5′-GCCAGCTGCCTGTCGGTGAG-3′;
185–279 bp region F, 5′-CGGGTGGCAGAGATGCGTGG-3′ and R,
5′-TCCAGCTCACCGGTCTCGGG-3′; 293–440 bp region F,
5′-AAGGCCATGCGAACCCCACG-3′ and R, 5′-AGGCGTCGTCAATCACCGCC-3′;
493–595 bp region F, 5′-CGTGTACAGCCGGGACGCAG-3′ and R,
5′-AAAGGCGTGTGCCAGGAGCC-3′; 594–676 bp region F,
5′-TTTCCTCCTGGCCCCGGCAT-3′ and R, 5′-TGGACCACGACGCCCTTGC-3′;
730–842 bp region F, 5′-GGGCCGCTCCTACTCTGCCT-3′ and R,
5′-TCTCGCTGGGGCAGAAGCCA-3′; 823–933 bp region F,
5′-TGGCTTCTGCCCCAGCGAGA-3′ and R, 5′-TCCGTGGTGCAGGCGGAGTA-3′;
1070–1170 bp region F, 5′-TTCACTTTCCTGGGTAAG-3′ and R,
5′-TTCTTGTCGCTGTCAAAG-3′; 1,261-1,390 bp region F,
5′-GCCGGAGGCGCTCATGTACC-3′ and R, 5′-GGTGGTGGTTGGAGGCCGTG-3′;
1,565-1,651 bp region F, 5′-TGCAACGTGAACATCTTC-3′ and R,
5′-CTCAGAGAATCGCCAGTA-3′; 1,869-2,004 bp region F,
5′-TCCGGAGTGGCAAGGGGAG-3′ and R, 5′-TGCTGTCCAAAGGCACCCC-3′;
2,230-2,332 bp region F, 5′-TATTCTGTTCTGGAGGAA-3′ and R,
5′-GGTTAGAGAATCCAAGTT-3′. The thermal cycling profile was as
follows: Initial denaturation at 95°C for 30 sec, followed by 55
cycles of at 95°C for 5 sec and 60°C for 30–34 sec (fluorescence
data collection was performed at this step). Melt curve analysis
was performed from 60–95°C with a continuous fluorescence
measurement at a ramp rate of 0.3°C per sec. Analysis of relative
gene expression data using real-time quantitative PCR and the
2−ΔΔCt method and the input RNA as an internal
control.
Dot blotting
Total RNA was incubated with oligo dT(25) beads
(cat. no. S1550S; New England Biolabs, Inc.) for 10 min at RT and
then washed with Lysis/Binding buffer (cat. no. B1556AVIAL; New
England Biolabs, Inc.), Wash Buffer I (cat. no. B1558AVIAL; New
England Biolabs, Inc.), Wash Buffer II (cat. no. B1559AVIAL; New
England Biolabs, Inc.)and Low Salt Buffer (cat. no. B1557AVIAL; New
England Biolabs, Inc.). The mRNA was eluted with Elution Buffer
(cat. no. B1561AVIAL; New England Biolabs, Inc.) and samples
containing 300 or 600 ng of mRNA were transferred to nylon
membranes. The RNA was cross-linked under UV light (λ=254 nm) at
0.3 J/cm2 and the membranes were blocked with 5% skim
milk in PBST for 1 h at RT. The membranes were incubated with
anti-m5C antibodies (1:1,000; cat. no. AB10805; Abcam)
overnight at 4°C, washed extensively with PBST (0.1% Tween) and
then incubated with an HRP-conjugated sheep anti-mouse secondary
antibody (1:5,000; cat. no. AF2819; Shanghai Beyotime Biotechnology
Co., Ltd.) for 2 h at RT. The membranes were washed with PBST (0.1%
Tween) and then stained with methylene blue. The nylon membranes
containing the target mRNA were visualized using a Tanon™ 5200
chemiluminescence gel imaging system (Tanon Science and Technology
Co., Ltd.)
RNA stability assay
CC cells were cultured in 6-well plates overnight
and then treated with 5 µg/ml actinomycin D (cat. no. HY-17559;
MedChemExpress) for 3, 6 or 9 h at 37°C. Total RNA was then
extracted using TRIzol® reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) RNA was reverse transcribed into cDNA
using the PrimeScript™ RT reagent Kit (Perfect Real Time) Kit (cat.
no. RR037A; Takara Bio, Inc.) according to the manufacturer's
protocol and the DNA was stored at −20°C. Quantitative PCR was
performed on an ABI StepOne Plus Real-Time PCR System (Applied
Biosystems; Thermo Fisher Scientific, Inc.) with three replicates
for each sample, using TB Green® Premix Ex Taq™ (Tli
RNaseH Plus) (cat. no. RR420A; Takara Bio Inc.). Analysis of
relative gene expression data using RT-qPCR as aforementioned and
data were analyzed using the comparative Cq method. Levels of gene
expression were normalized to that of GAPDH. The mRNA
half-life time was estimated using linear regression analysis.
CC data acquisition and statistical
analysis
RNA-sequencing data and corresponding clinical
information for cervical squamous cell carcinoma and endocervical
adenocarcinoma (CESC) were downloaded from The Cancer Genome Atlas
(TCGA) database via the Genomic Data Commons (GDC) data portal
(https://portal.gdc.cancer.gov/). Gene
expression profiles that were normalized using fragments per
kilobase million (FPKM) were obtained for subsequent analysis,
while clinical data including survival time and survival. A
comprehensive list of VM-promoting factors (Myc, Sox2, POU5F1,
SNAIL family members, ZEB2, TWIST, MMP-2, MMP-9, and MMP-14) and
5-methylcytosine (m5C) RNA methylation-related genes was compiled
through literature review and publicly available databases. The
m5C-related genes primarily included ‘writers’ (methyltransferases
such as NSUN family members and TRDMT1), ‘readers’ (YBX1, ALYREF),
and ‘erasers’ (TET family members). Expression profiles of these
VM-promoting factors and m5C-related genes were extracted from the
FPKM expression matrix for further analysis. Student's t-test was
performed to evaluate the differential expression of VM-promoting
factors and m5C-related genes between tumor and normal tissues,
with a P-value of <0.05 considered statistically significant.
For survival analysis, patients were stratified into high- and
low-expression groups based on the median expression value of each
VM-promoting factor and m5C-related gene. Kaplan-Meier survival
curves were generated to assess the association between gene
expression levels and overall survival (OS), and the log-rank test
was employed to determine statistical significance between survival
curves. Hazard ratios (HR) and 95% confidence intervals (CI) were
calculated using Cox proportional hazards regression models. All
statistical analyses were conducted using R software (version
4.2.1; Posit Software, PBC), with the ‘survival’ (version 3.8.2;
http://github.com/therneau/survival)
and ‘survminer’ (version 0.5.0; http://rpkgs.datanovia.com/survminer/index.html)
packages utilized for survival analysis and visualization. All
tests were two-sided, and P-values <0.05 were considered
statistically significant.
Statistical analysis
Data are presented as mean ± SD from at least three
independent biological replicates. For comparisons between paired
samples (tumor vs. adjacent normal tissue), statistical
significance was assessed using a paired two-tailed Student's
t-test when data passed normality (Shapiro-Wilk test) and
homogeneity of variance assumptions. For comparisons between two
independent groups, statistical significance was determined using
an unpaired two-tailed Student's t-test for data that passed both
normality (assessed by the Shapiro-Wilk test) and homogeneity of
variance tests. For data that did not meet the assumptions for
parametric testing, a two-tailed Mann-Whitney U test was employed
For comparisons among ≥3 groups, one-way ANOVA followed by Tukey's
post hoc test was performed. Linear relationships between variables
were assessed through Pearson's correlation analysis and linear
regression. Survival analysis was carried out using the
Kaplan-Meier method, and differences between groups were compared
with the log-rank test. P<0.05 was considered to indicate a
statistically significant difference. Statistical analyses were
performed using GraphPad Prism (version 7; Dotmatics).
Results
Vasculogenic mimicry is increased in
CC and is associated with poor prognosis
In order to explore the prevalence of VM in CC,
three pairs of CC and paracancerous tissues from human subjects
were selected for IHC staining using the diagnostic standard of
CD31/PAS double staining (Fig. 1A).
Among the three pairs of tissue samples, VM was only revealed to
exist in CC tissues and not in paracancerous tissues and VM was
increased in CC tissues. To study the impact of VM on CC prognosis,
a CC tissue microarray was first analyzed by IHC staining and then
the results were subjected to a double-blind statistical comparison
to the prognoses of the subjects (Fig.
1B). VM-positivity was revealed to be associated with shorter
biochemical progression-free survival after surgery (P=0.004;
Kaplan-Meier analysis) (Fig. 1C;
P<0.01). These results suggest that VM exists at higher levels
in CC tissues and that its presence is associated with a poor
prognosis.
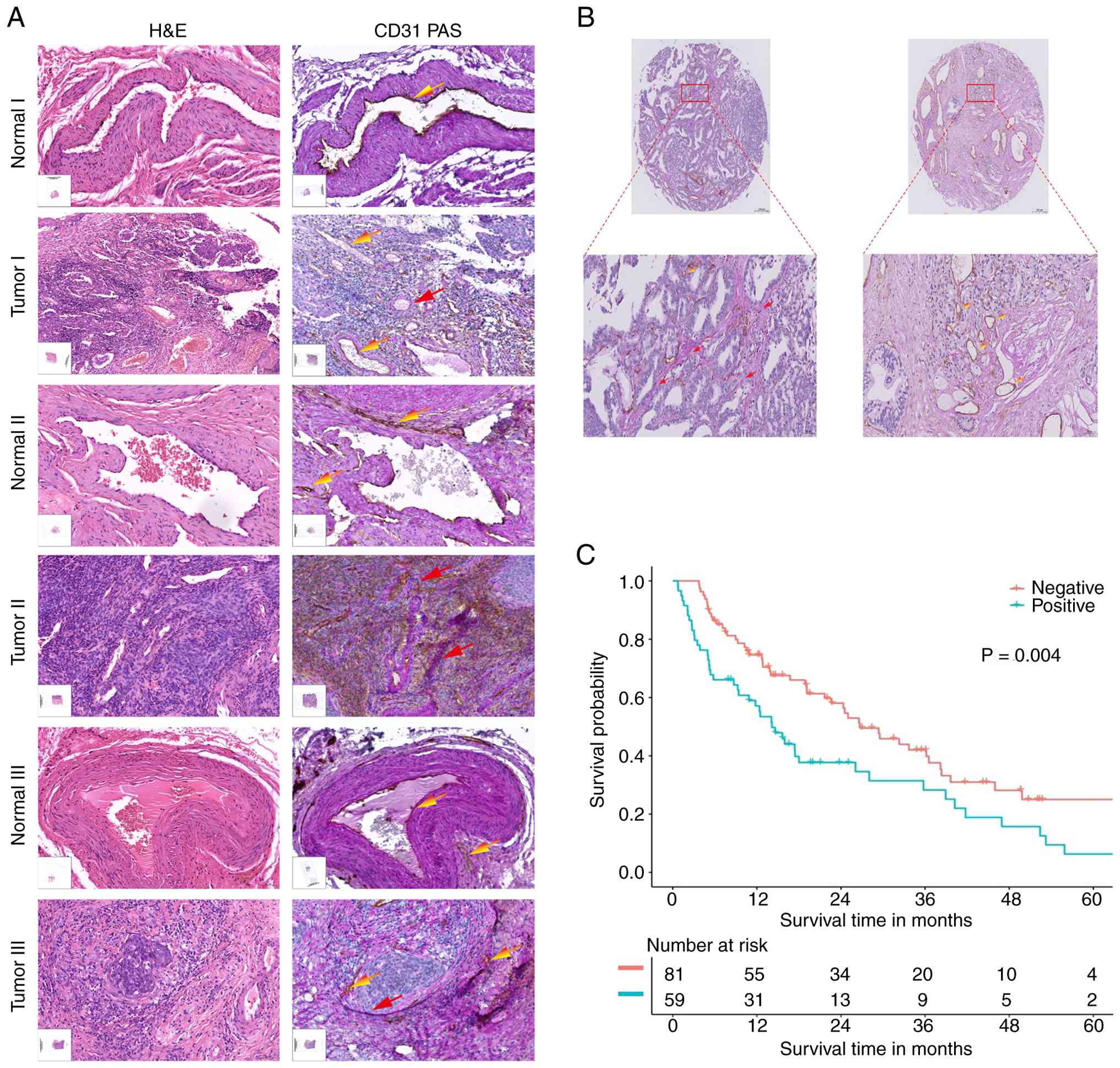 | Figure 1.Vasculogenic mimicry is increased in
cervical cancer and is associated with poor prognosis. (A) Three
pairs of CC and paracancerous tissues were analyzed by H&E and
PAS staining and immunohistochemical detection of CD31, with
CD31−/PAS+ serving as the diagnostic standard
for VM. Red arrows indicate VM and yellow arrows indicate
endothelial angiogenesis. Scale bar, 50 mm. Original magnification,
×20. (B) A CC tissue microarray, consisting of tissues from 140
cases of CC, was subjected to CD31/PAS analysis. Red arrows
indicate VM and gold arrows indicate endothelial angiogenesis.
Original magnification, ×20. (C) Level of VM positivity was
compared with patient outcomes in the form of biochemical
progression-free survival (P=0.004, log-rank test). PAS, periodic
acid-Schiff; VM, vasculogenic mimicry; CC, cervical cancer. |
MMP-9, a key Vasculogenic
mimicry-promoting factor, is highly expressed in CC and is
associated with poor prognosis
In order to further explore the association of VM
with CC, a CC database from TCGA was used to analyze the expression
levels of >12 key factors affecting the three steps that lead to
VM generation: CSC formation, EET and ECM reconstruction (Fig. 2A). The genes encoding SRY-box
transcription factor 2 (Sox2) and MMP-9 were revealed
to be highly expressed in CC (Fig.
2A). Further analysis of the impact of these two factors on the
prognosis of CC demonstrated that patients with CC exhibiting
higher MMP-9 expression levels tend to have a poorer
prognosis, while those with higher Sox2 expression levels
generally experience an improved prognosis (Fig. 2B and C).
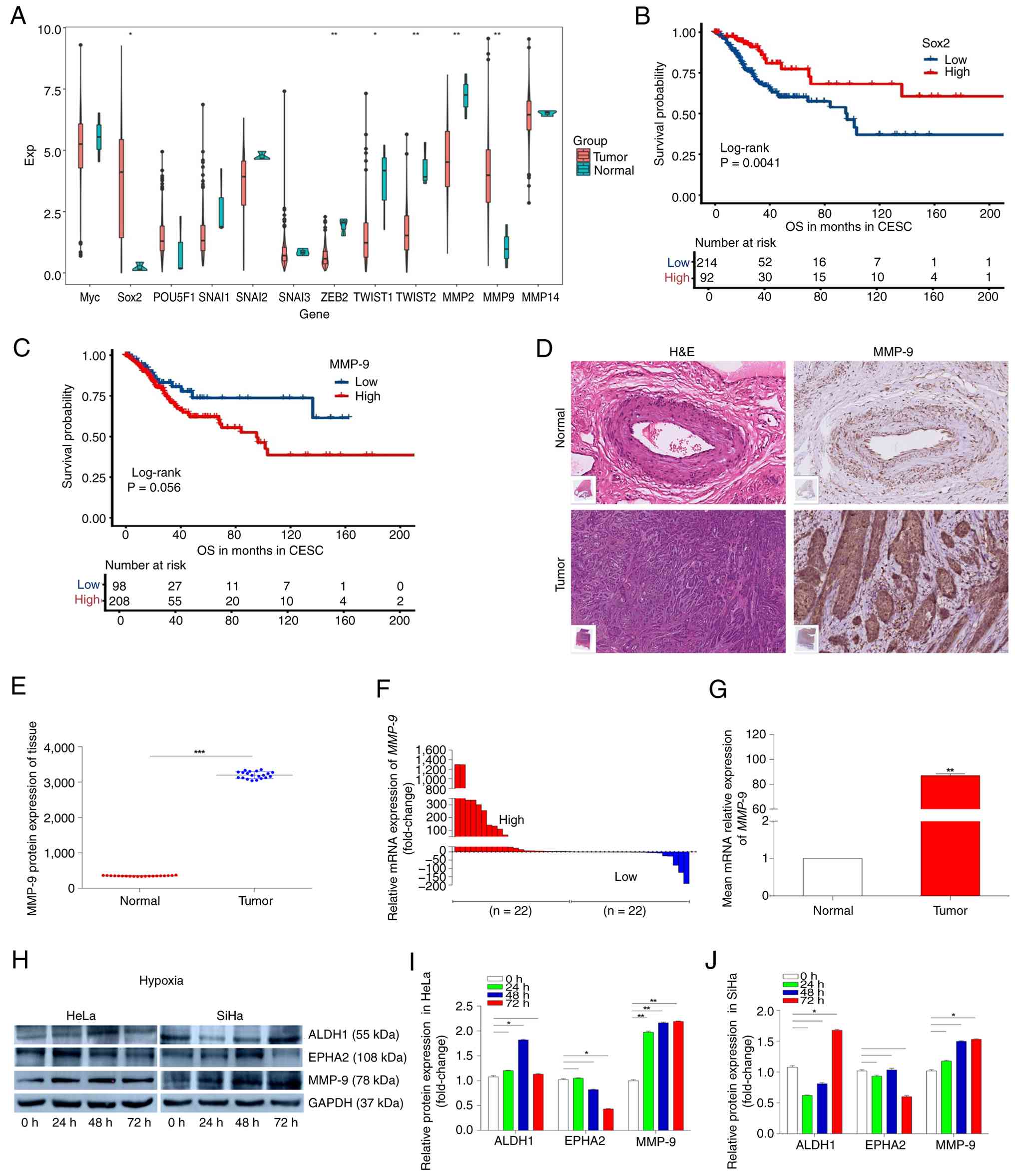 | Figure 2.MMP-9, a factor that promotes
Vasculogenic mimicry, is highly expressed in CC and is associated
with poor prognosis. (A) CC database of TCGA was used to analyze
key factors associated with VM. (B) Association of Sox2
expression with overall survival in CC (log-rank test). (C)
Association of MMP-9 expression with overall survival in CC
(log-rank test). (D) Panoramic scans after immunohistochemical
detection of MMP-9 and H&E staining in samples from cancerous
and paracancerous tissues from subjects with CC. Scale bar, 50 µm.
Original magnification, ×20. (E) Protein levels of MMP-9 in 20
paired samples, with the MMP-9 level in CC tissue expressed
compared with that in the paired normal tissue. (F) Expression
levels of MMP-9 mRNA in 44 paired CC and paracancerous
tissues, with MMP-9 expression in CC tissue expressed
compared with that in the paired normal tissue. (G) Comparison of
the average expression levels of MMP-9 mRNA in CC tissues
compared with paracancerous tissues. (H) HeLa and SiHa cells were
incubated under hypoxia (0.1% O2) and proteins collected
at 24, 48 and 72 h for western blotting of ALDH1, EPHA2, MMP-9 and
GAPDH. ImageJ was used to semi-quantify western blotting signals
from HeLa (I) and SiHa (J) cells. GAPDH served as an internal
reference. *P<0.05, **P<0.01 and ***P<0.001. MMP-9, matrix
metalloproteinase 9; VM, vasculogenic mimicry; ALDH1, aldehyde
dehydrogenase 1; EPHA2, ephrin type-A receptor 2; TCGA, The Cancer
Genome Atlas; Sox2, SRY-box transcription factor 2; CC, cervical
cancer; CESC, cervical squamous cell carcinoma. |
Similarly, IHC staining of 20 pairs of CC and
paracancerous tissue samples demonstrated that the protein levels
of MMP-9 were significantly higher in cancerous tissues compared
with paired normal tissues (P<0.001) (Fig. 2D and E). The expression levels of
MMP-9 mRNA was further analyzed using RT-qPCR of 44 pairs of
samples. The present study identified that MMP-9 mRNA was
significantly highly expressed in cancerous tissues compared with
paired normal tissues (P<0.01) (Fig.
2F and G).
To further study the potential role of MMP-9 in
promoting VM in CC, VM-competent CC cell lines were subjected to
hypoxic stimulation. HeLa and SiHa cells were induced in a 0.1%
O2 incubator (40–42)
and proteins collected at 24, 48 and 72 h were analyzed using
western blotting (Fig. 2H). The
level of ALDH1, a key factor affecting VM through the CSC pathway,
was revealed to be slightly increased under continuous hypoxia,
although the increase did not rise to the level of statistical
significance (P<0.05) (43,44).
The level of EPHA2, a key factor affecting VM through the EET
pathway (45,46), was revealed to be decreased. By
contrast, the level of MMP-9, a key factor affecting VM through the
ECM pathway, increased over time and the level of MMP-9 at 72 h was
significantly higher compared with in non-hypoxic cells (P<0.01)
(Fig. 2I and J). These results
suggest that compared with the CSC pathway, the effect of VM
generation on CC mainly depends on the ECM remodeling pathway and
the increased expression levels of MMP-9 may serve a key role.
m5C methyltransferase NSUN2
is upregulated in CC and is associated with poor prognosis
Next, the mechanism by which MMP-9 levels are
increased in VM in the context of CC were investigated. A dot blot
test demonstrated that the m5C methylation modification
was significantly increased upon hypoxic induction of VM-competent
HeLa (P<0.001) and SiHa (P<0.01) cells (Fig. 3A-C). Therefore, TCGA CC database was
then used to analyze the expression levels of all enzymes that
generate the m5C modification. The present study
identified a high expression level of NSUN2 and low
expression level of TRDMT1 (which encodes transfer RNA
aspartic acid methyltransferase 1) among the methyltransferases
(Fig. 3D).
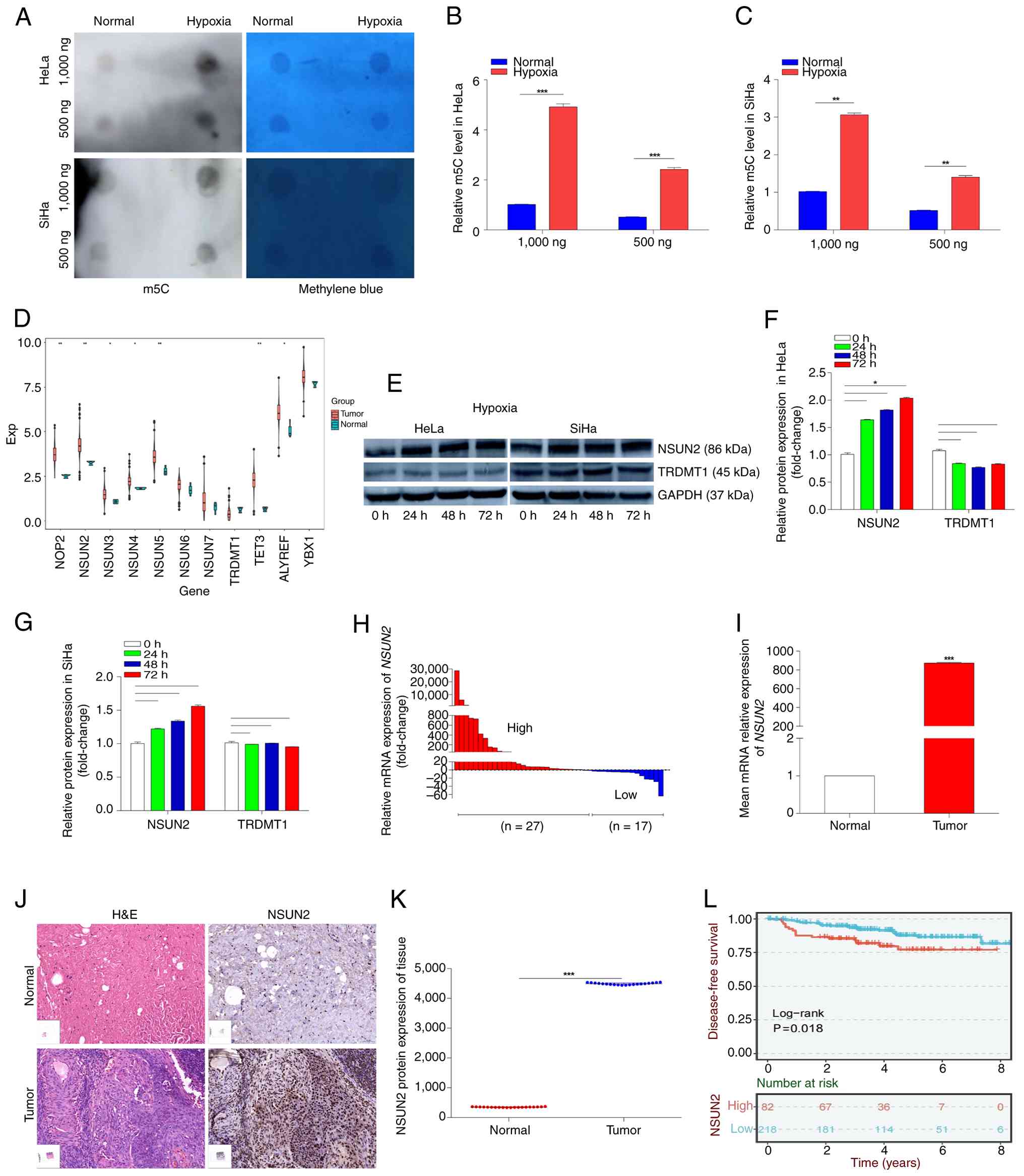 | Figure 3.m5C RNA methyltransferase
NUSN2 is significantly upregulated in CC and its gene expression is
associated with poor prognosis. (A) The level of m5C
modification of RNA in CC cells was analyzed using a dot blot
assay. Methylene blue staining served as an internal reference. (B)
Semi-quantitative analysis of dot blot results in HeLa cells. (C)
Semi-quantitative analysis of dot blot results in SiHa cells. (D)
Expression levels of genes encoding m5C
methyltransferases was analyzed within a CC database of TCGA. (E)
HeLa and SiHa cells were incubated under hypoxia (0.1%
O2) and proteins collected at 24, 48 and 72 h were
analyzed for NSUN2, TRDMT1 and GAPDH by western blotting. ImageJ
was used to quantify western blotting signals from (F) HeLa and (G)
SiHa cells. GAPDH served as an internal reference. (H) mRNA
expression level of NSUN2 in 44 pairs of CC and
paracancerous tissues were quantified using RT-qPCR. Gene
expression in cancer tissues is expressed compared with expression
in normal tissues. (I) Comparison of the average expression level
of NSUN2 mRNA in CC tissues compared with paracancerous
tissues. (J) Panoramic scans after IHC detection of NSUN2 and
H&E staining in CC and paracancerous tissues. Scale bar, 50 µm.
Magnification, ×20. (K) Protein levels of NSUN2 in 20 paired CC and
paracancerous tissues, with the NSUN2 level in CC tissue expressed
compared with that in the paired normal tissue. (L) Level of
NSUN2 expression in CC was associated with a poor prognosis
in the form of biochemical progression-free survival (P=0.018;
log-rank test). *P<0.05, **P<0.01 and ***P<0.001. MMP9,
matrix metalloproteinase 9; VM, vasculogenic mimicry; CC, cervical
cancer; IHC, immunohistochemistry; NSUN2, NOP2/Sun RNA
methyltransferase 2; m5C, 5-methylcytidine; TRDMT1,
transfer RNA aspartic acid methyltransferase 1; TCGA, The Cancer
Genome Atlas; RT-qPCR, reverse transcription-quantitative PCR; IHC,
immunohistochemistry. |
To investigate the impact of hypoxia on the levels
of these proteins, VM-competent HeLa and SiHa cells were induced in
a 0.1% O2 incubator and proteins collected at 24, 48 and
72 h were analyzed using western blotting. The expression levels of
NSUN2 (P<0.05) were consistently high under continuous hypoxia,
while the expression levels of TRDMT1 were not significantly
changed and was even slightly downregulated. Therefore, the present
study concluded that NSUN2 is a key methyltransferase in the VM
process under these conditions (Fig.
3E-G).
Regarding human tissue samples, NSUN2 mRNA
expression was revealed to be higher in CC tissue samples compared
with in normal control samples (Fig.
3H). Similarly, a quantitative analysis of 44 pairs of CC and
paracancerous tissue samples demonstrated that NSUN2 mRNA
levels were significantly higher in CC tissues compared with
paracancerous tissues (P<0.001) (Fig. 3I) and IHC analyses of 20 pairs of
samples revealed that expression was also significantly higher at
the protein level (P<0.001) (Fig. 3J
and K).
Notably, the level of NSUN2 expression in CC
was associated with a poor prognosis in the form of biochemical
progression-free survival (P=0.018; log-rank test; Fig. 3L). These results suggest that
m5C methylation of RNA increases in VM-competent CC
cells under hypoxic conditions and that the methyltransferase NSUN2
serves a key role in this process.
NUSN2 promotes vasculogenic mimicry,
invasion and migration of CC cells
As NSUN2 was a key candidate for the direct mediator
of increased m5C methylation in hypoxic CC cells, the
potential impact of this methyltransferase on the VM process.
First, in vitro experiments were performed in order to
explore whether NSUN2 can promote VM in CC cells. Preliminary
experiments involved the normal cervical cell line HaCaT and the CC
cell lines HeLa, SiHa, CaSki, C33A and HT-3. According to RT-qPCR
analyses, NSUN2 was revealed to be most significantly highly
expressed in HeLa (P<0.001) and SiHa (P<0.01) cells (Fig. 4A); therefore, these two CC cell
lines were selected for further analyses.
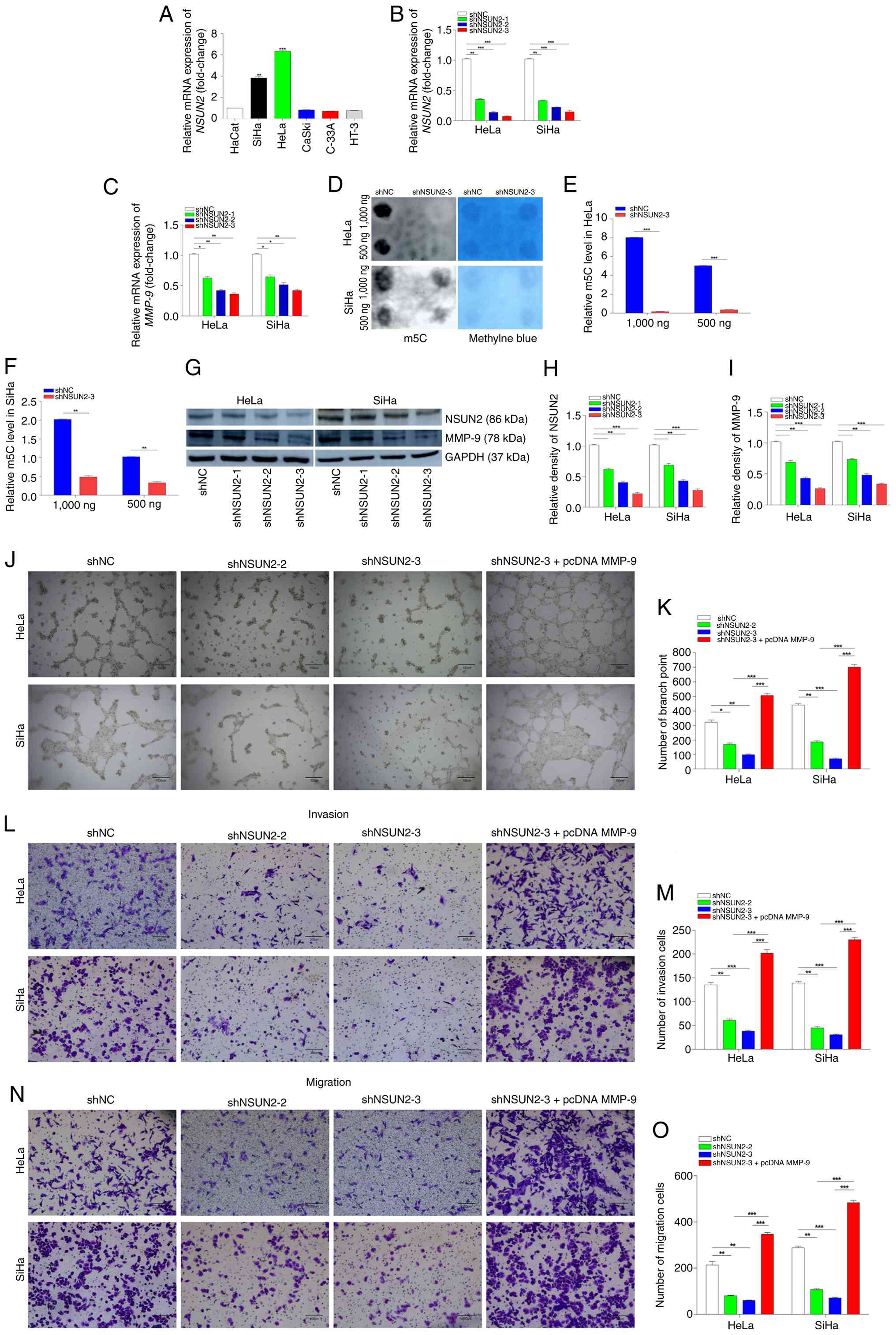 | Figure 4.NSUN2 promotes Vasculogenic mimicry,
invasion and migration of CC cells under hypoxic conditions. (A)
Expression levels of NSUN2 in CC cells lines (HeLa, SiHa,
CaSki, C33A and HT-3) and in a normal cervical cell line (HaCaT).
(B) RT-qPCR was used to determine relative expression levels of
NSUN2 mRNA in HeLa and SiHa cells after transfection of
shRNAs and incubation under hypoxia for 24 h. (C) Relative
expression levels of MMP-9 mRNA in HeLa and SiHa cells
transfected with the NSUN2-interfering plasmid and incubated
under hypoxia for 24 h. (D) Dot blot assay analysis of
m5C expression levels in HeLa and SiHa cells after 48 h
of hypoxia culture following transfection with NSUN2
knockdown plasmids. (E) Semi-quantitative analysis of dot blot
results in HeLa cells. (F) Semi-quantitative analysis of dot blot
results in SiHa cells. (G) Western blotting was used to investigate
NSUN2 and MMP-9 protein levels in HeLa and SiHa cells transfected
with the NSUN2-interfering plasmid and incubated under
hypoxia for 48 h. ImageJ was used to semi-quantify western blotting
bands for (H) NSUN2 and (I) MMP-9 protein levels. (J) 2D
tube-forming assays of HeLa and SiHa cells transfected with an
NSUN2-interfering plasmid and incubated under hypoxia. Scale
bar, 100 µm. Original magnification, ×4. (K) Assay displayed in
panel (J) was quantified using Image Pro. (L) Invasion assay of
HeLa and SiHa cells transfected with a control plasmid, shNSUN2-2
or shNSUN2-3 or with shNSUN2 and pcDNA MMP-9. Scale bar, 200 µm.
Original magnification, ×10. (M) Assay displayed in panel was
quantified using Image Pro. (N) Migration assay of HeLa and SiHa
cells transfected with a control plasmid, shNSUN2-2 or shNSUN2-3 or
with shNSUN2 and pcDNA MMP-9. Scale bar, 200 µm. Original
magnification, ×10. (O) Assay displayed in panel (N) was quantified
using Image Pro. *P<0.05, **P<0.01 and ***P<0.001. VM,
Vasculogenic mimicry; CC, cervical cancer; IHC,
immunohistochemistry; NSUN2, NOP2/Sun RNA methyltransferase 2;
m5C, 5-methylcytidine; RT-qPCR, reverse
transcription-quantitative PCR; IHC, immunohistochemistry; CaSki,
human cervical cancer cell line with intestinal metastasis; C33A,
human cervical cancer cell line; HaCaT, human skin keratinocytes
cell line; HeLa, human cervical cancer cell line; HT-3, human
cervical cancer cell line; MMP9, matrix metalloproteinase 9; SiHa,
human cervical squamous cell line; pcDNA, plasmid cloning DNA;
shRNA, short hairpin RNA; NC, negative control. |
Plasmids were designed to produce shRNA to interfere
with expression levels of NSUN2. Following transfection of
these plasmids and a control plasmid into HeLa and SiHa cells, the
cells were subjected to hypoxic conditions for 24 h and RNA was
collected for RT-qPCR analyses. Compared with the control group,
the level of NSUN2 mRNA was significantly decreased in cells
transfected with NSUN2-interfering plasmids, particularly with
plasmid shNUSN2-3 (P<0.001) (Fig.
4B). Notably, the level of MMP-9 mRNA was also
significantly decreased in cells transfected with shNUSN2-3
(P<0.01) (Fig. 4C).
To further investigate whether NSUN2 serves a key
role in the methylation modification and thus the level of
m5C, cells transfected with the interfering plasmid were
subjected to hypoxia and the mRNA was subjected to dot blotting.
The present study identified that the level of m5C was
significantly reduced in HeLa (P<0.001)and SiHa (P<0.01)
cells and this effect was particularly strong in HeLa cells
(Fig. 4D-F). The results suggest
that NSUN2 serves a key role in m5C methylation
modification in CC.
The levels of NSUN2 and MMP-9 proteins were also
investigated under these conditions. After transfection of the
NSUN2-interfering plasmid into HeLa and SiHa cells and
subjection to hypoxic conditions for 48 h, proteins were collected
and analyzed using western blotting. Compared with the control,
transfection of NSUN2-interfering plasmids led to
significant decreases in the levels of NSUN2 protein, particularly
with the shNUSN2-3 plasmid (P<0.001) (Fig. 4G and H). The level of MMP-9 protein
was also significantly decreased upon interference with
NUSN2 expression (P<0.001) (Fig. 4I).
The effects of NSUN2 knockdown on the
two-dimensional tube formation that characterizes VM were
investigated under hypoxic conditions. After NSUN2
knockdown, the tube formation ability of HeLa and SiHa cells was
significantly decreased (P<0.01); however, overexpression of
MMP-9 significantly reversed this effect of NSUN2
knockdown on cell tube formation (P<0.001) (Fig. 4J and K). Similarly, both invasion
and migration abilities of HeLa and SiHa cells under hypoxic
conditions were significantly diminished after NSUN2
knockdown (P<0.01); however, overexpression of MMP-9
significantly reversed these effects (P<0.001) (Fig. 4L-O). These results suggest that
NSUN2 is required for generation of VM of CC cells and they
demonstrate that NSUN2 promotes the invasion and migration of CC
cells under hypoxic conditions. It was also identified that MMP-9
is a potential downstream mediator of these effects of NSUN2.
NSUN2 maintains the stability of MMP-9
mRNA
Because MMP-9 was revealed to be a potential
downstream mediator of the effects of NSUN2, the present study
further investigated the relationships between these two factors.
First, a Pearson's correlation analysis of the levels of
NSUN2 and MMP-9 mRNA in 44 pairs of CC tissue samples
was performed. The levels of NSUN2 mRNA was revealed to be
significantly positively correlated with the level of MMP-9
mRNA (P<0.0001; Fig. 5A). In
addition, methylated-RNA immunoprecipitation assays demonstrated a
marked enrichment of m5C in MMP-9 mRNA in HeLa
and SiHa cells in two regions: Between nucleotides 1,070 and 1,170
and between nucleotides 1,565 and 1,651 (Fig. 5B and C). Similarly, RNA
immunoprecipitation assays indicated that NSUN2 was highly enriched
on MMP-9 mRNA within the 1,565-1,651 region in both HeLa and
SiHa cells (Fig. 5D and E).
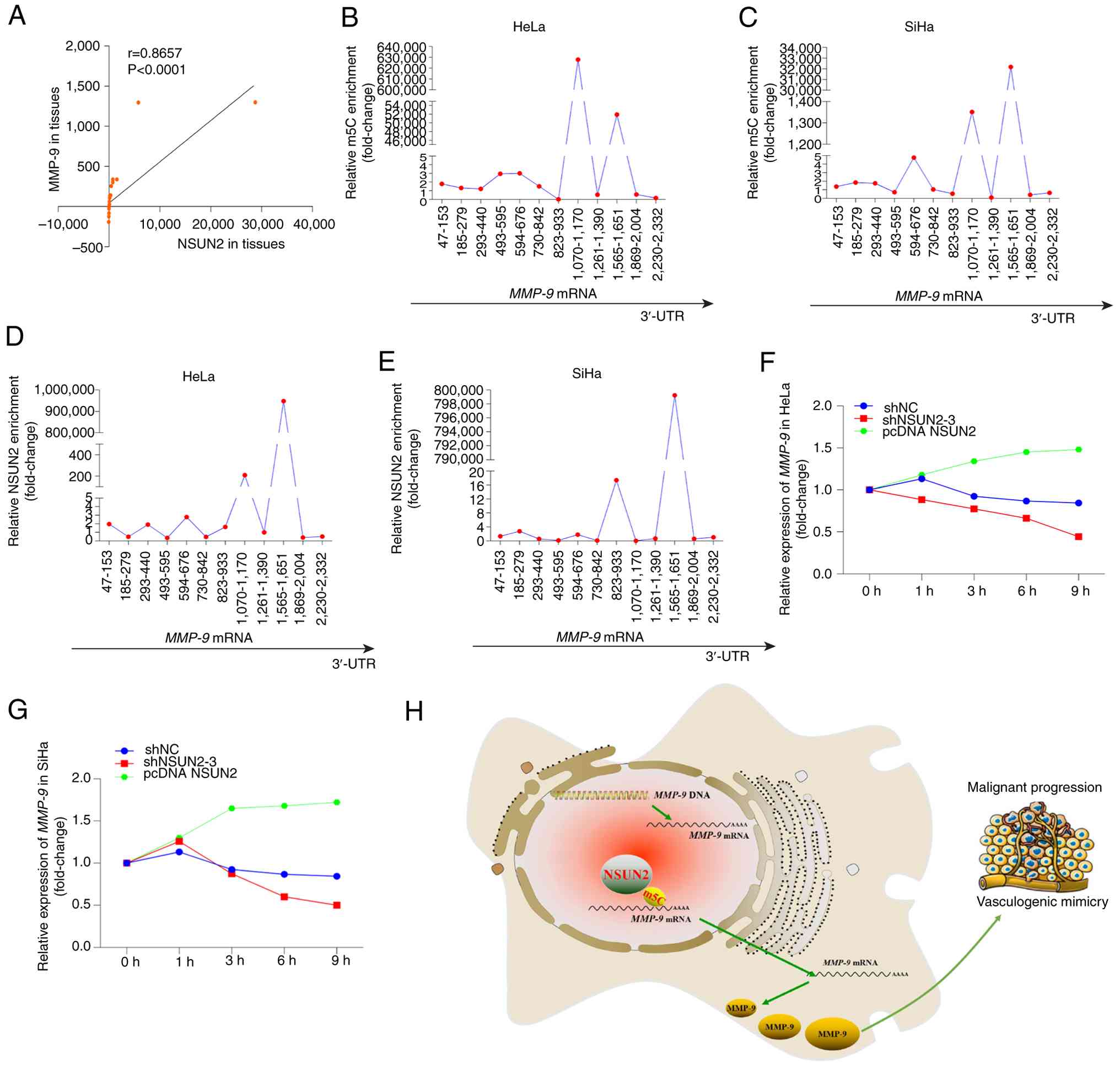 | Figure 5.NSUN2 increases the stability of
MMP-9 mRNA. (A) A positive correlation was observed between
NSUN2 and MMP-9 mRNA expression levels in 44 pairs of
samples from subjects with CC. Enrichment of the m5C
modification of MMP-9 mRNA in HeLa (B) and SiHa (C) Cells
were measured with anti-m5C methylated-RNA IP assays.
Interaction of NSUN2 with MMP-9 mRNA in (D) HeLa and (E)
SiHa cells was measured with anti-NSUN2 RNA IP assays.
Stability of MMP-9 mRNA after interference with and
overexpression of NSUN2 was measured in (F) HeLa and (G) SiHa
cells. (H) A model illustrating the proposed mechanism by which
NSUN2-mediated stabilization of MMP-9 mRNA promotes
Vasculogenic mimicry in CC. NSUN2, NOP2/Sun RNA methyltransferase
2; m5C, 5-methylcytidine; pcDNA, plasmid cloning DNA;
shRNA, short hairpin RNA; HeLa, human cervical cancer cell line;
MMP9, matrix metalloproteinase 9; SiHa, human cervical squamous
cell line; IP, immunoprecipitation; CC, cervical cancer; NC,
negative control. |
The present study also identified that in HeLa and
SiHa cells, the stability of MMP-9 mRNA markedly decreased
upon transfection of an NSUN2-interference plasmid and the
stability markedly increased upon overexpression of NSUN2
(Fig. 5F and G). These results were
consistent with a model in which NUSN2 modifies MMP-9 mRNA
in the region of 1,565 to 1,651 bp and thus, that MMP-9 is a
downstream mediator of the effects of NUSN2 on the hypoxic VM
response in CC cells. In this model, NSUN2 increases the expression
levels of MMP-9 by maintaining the stability of the MMP-9
mRNA, leading to the promotion of VM generation by CC cells and
enhanced malignant progression (Fig.
5H).
Discussion
Globally, CC accounted for about 7.7% of all female
cancer deaths globally, making it one of the deadliest cancers for
women, with ~348,000 deaths annually. (1). While radiation therapy and radical
surgery can improve the 5-year survival rate for patients with
early-stage CC (47,48), there is an urgent need to explore
novel strategies for the treatment of distant metastases and drug
resistance for patients with advanced CC. Neovascularization is a
potential vulnerability in advanced cancer, as when the diameter of
a tumor >2 mm, the inner cells become starved for oxygen and
nutrients and new blood vessels are needed. Therefore, tumor cells
secrete a large number of vascular growth factors, such as VEGF and
FGF-2, to promote the formation of blood vessels around the
tumor.
Certain cancer cells mimic endothelial cells and
align along a deposited basement membrane to form lumenized,
interconnecting vessel-like structures that perfuse blood and fuse
to the endothelium, in a process called VM (49). Tumor cells that shed from VM ducts
are more likely to undergo early tumor metastasis to distal organs
through the VM-dependent vasculature and endothelium-dependent
vessels (38,39). In the hypoxic tumor
microenvironment, tumor cells that have the ability to engage in VM
typically revert to a poorly differentiated state, thus forming
CSCs, which tend to be resistant to chemotherapy agents (4,5).
Therefore, it is key to study VM to develop novel strategies for
the treatment of metastasis and drug resistance in advanced CC and
exploring the molecular mechanisms underlying VM in CC may provide
a theoretical basis for novel tools to diagnose and treat CC in the
future.
Notably, the present study provides further evidence
that VM influences the invasiveness and metastatic potential of
malignant tumors and that VM status might serve as an independent
tool to evaluate prognoses in CC and other cancer types.
In the hypoxic tumor microenvironment, the VM
process involves CSC, EET or ECM, but the specific mechanism
leading to VM in CC remained unclear. In the present study, the
statuses of key factors affecting CSC, EET and ECM in CC were
analyzed using a CC database in TGCA and the results suggested that
Sox2, which affects CSC and MMP-9, and thus the ECM,
were highly expressed in CC. However, the present analysis also
demonstrated that only MMP-9 expression was associated with
poor prognosis in CC; therefore, the present study performed
further analysis of MMP-9. Experimentally, the present study
identified that expression levels of MMP-9 were increased at
both the mRNA and protein levels in CC tissues compared with
paracancerous normal tissues. VM-competent CC cell lines (HeLa and
SiHa) were selected to explore the possible mechanisms that
influence the formation of VM under hypoxic conditions. Notably,
hypoxic induction led to an increase in the levels of MMP-9, a
protein that influences the ECM pathway in the process of VM. By
contrast, the levels of ALDH1, which affects the CSC pathway were
revealed to increase only slightly under continuous hypoxia and
EPHA2, which affects the EET pathway, was revealed to be decreased.
These results provided further evidence that increased VM in CC is
mainly dependent on the ECM pathway and that MMP-9 serves a key
role. Notably, MMP-9 expression was elevated in CC and
hypoxia-induced conditions, with MMP-mediated degradation serving a
pivotal role in CC metastatic progression.
Next, the mechanisms by which the levels of MMP-9
increase in this context were considered. Various tumor cells,
including CC (50), pancreatic
cancer (51), glioma (52), breast cancer (53) and liver cancer (54), have been revealed to undergo changes
to RNA methylation under hypoxic conditions. The present study
specifically focused on the RNA modification m5C, which
has gained notable attention at the frontiers of epigenomics
research. Roles for this modification in promoting proliferation,
migration, angiogenesis and drug resistance have been reported in
multiple tumor types (such as NSCLC, bladder cancer and glioma)
(32–34). In the present study, the results of
a dot blot assay revealed that the levels of the m5C
modification increased significantly in the RNA of VM-competent
HeLa and SiHa cells under hypoxic conditions. TCGA CC database
further indicated that the methyltransferase NSUN2 is
significantly upregulated in CC. In order to determine if NSUN2
might be responsible for the changes to the m5C
modification, the proteins in VM-competent HeLa and SiHa cells
under continuous hypoxia were analyzed using western blotting. The
results suggested that compared with methyltransferase TRDMT1, the
protein level of NSUN2 continued to increase with the time of
hypoxia induction, in accordance with the increasing m5C
levels under these same conditions. By contrast, when the levels of
NSUN2 were decreased using shRNA, the levels of
m5C were significantly decreased. Therefore, the present
study concluded that NSUN2 serves a key role in establishing the
m5C methylation modification in CC. Subsequently, the
present study identified that NSUN2 levels are significantly higher
in CC and that high expression level of the NSUN2 gene is
associated with a poor prognosis in CC. These results suggest that
NSUN2 might contribute to the malignant progression of CC by
influencing the VM process.
Subsequently, the present study performed further
analysis of the model in which NUSN2 contributes to VM generation
and thus to invasion and migration of CC. Interfering with the
expression of NUSN2 using a shRNA-expressing plasmid system
inhibited the VM process as well as invasion and migration of HeLa
and SiHa cells. Notably, the present study also observed that
decreasing the levels of NSUN2 was associated with decreased
expression of MMP-9 at the mRNA and protein levels;
accordingly, overexpression of MMP-9 significantly reversed
the inhibitory effects of NSUN2 knockdown on VM generation,
invasion and migration. Therefore, these results indicated that
NSUN2 may act as an oncogene, as its increased expression
promotes VM and malignant progression of CC by increasing the
expression levels of MMP-9.
The m5C modulation alters gene expression
at the post-transcriptional level by affecting RNA stability.
Therefore, we hypothesized that NSUN2 might interact with and
modify MMP-9 mRNA. This potential relationship between NSUN2
and MMP-9 mRNA was investigated by focusing on the
m5C modifications within this mRNA. The present study
identified that MMP-9 mRNA is markedly enriched in
m5C in two regions of the mRNA: Between nucleotides
1,070 and 1,170 and between nucleotides 1,565 and 1,651. Further
investigation in the present study demonstrated that NSUN2 formed a
stable interaction with MMP-9 mRNA within the region from
nucleotides 1,565-1,651, suggesting that NSUN2 might modify and
thus stabilize MMP-9 mRNA. Subsequently, the stability of
MMP-9 mRNA was revealed to decrease upon shRNA-mediated
interference of NSUN2 expression and increase upon
overexpression of NSUN2. Therefore, the effect of NSUN2 on
VM generation in CC is likely mediated at least in part by
increasing levels of MMP-9. Thus, the present study enhances
current understanding of the VM process during the malignant
progression of CC and provides insights into the regulation of VM
by the m5C RNA modification. It also suggests that VM,
NSUN2 and MMP-9 are potentially valuable targets in the treatment
of solid tumors in the future.
Future research should address the limitations of
the present study to further validate the findings. While th
present work establishes a novel link between NSUN2, m5C
modification of MMP9 mRNA and VM in CC, the enzymatic activity of
the upregulated MMP-9 protein was not assessed. Therefore,
subsequent investigation will be to determine if increased
expression translates to heightened proteolytic function using
activity-based assays. This would provide a more robust foundation
for evaluating repurposed drugs such as the MMP inhibitor
doxycycline (which shows anti-VM efficacy in other types of cancer)
(55,56), within the present experimental
model. Additionally, the specific molecular interplay, including
whether m5C readers such as YBX1 (32,57–59)
are involved in conveying the NSUN2-mediated effect, remains an
open and mechanistically important avenue for future research.
The present study demonstrated that NUSN2
influences the cellular level of MMP-9 by stabilizing its mRNA. If
NUSN2 expression increases, its effect on MMP-9 promotes malignant
progression of CC by enhancing VM. These results indicate that both
NSUN2 and MMP-9 may serve as biomarkers for early diagnosis and
prognosis in CC and these two enzymes may serve as targets for
novel anti-vascular therapies in the future.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
JL and HS conceptualized and designed the present
study. MY, KS and GL provided study materials, recruited patients,
were involved in the interpretation of the clinical data and
contributed to the critical revision of the manuscript for
important intellectual conten. XX, LQ and MQ collected and
assembled the data. JH, XX and MQ conducted data analysis and
interpreted the data. JL wrote the manuscript. JL and HS confirmed
the authenticity of all the raw data. All authors have read and
approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Nanjing Medical University [approval no. (2024) Court
Ethics Opinion No. 028-1; Nanjing, China] and written informed
consent was obtained from all participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
CC
|
cervical cancer
|
|
CSC
|
cancer stem cells
|
|
CaSki
|
human cervical cancer cell line with
intestinal metastasis
|
|
C33A
|
human cervical cancer cell line
|
|
DEPC
|
diethylpyrocarbonate
|
|
TRDMT1
|
transfer RNA aspartic acid
methyltransferase 1
|
|
EET
|
epithelial-endothelial
transformation
|
|
ECM
|
extracellular matrix
|
|
FBS
|
fetal bovine serum
|
|
HaCaT
|
human skin keratinocytes cell
line
|
|
HeLa
|
human cervical cancer cell line
|
|
HT-3
|
human cervical cancer cell line
|
|
m5C
|
5-methylcytidine
|
|
MMP-9
|
matrix metalloproteinase 9
|
|
NSUN2
|
NOP2/Sun RNA methyltransferase 2
|
|
TCGA
|
The Cancer Genome Atlas
|
|
VM
|
vasculogenic mimicry
|
|
Sox2
|
SRY-box transcription factor 2
|
|
SiHa
|
human cervical squamous cell line
|
References
|
1
|
Bray F, Laversanne M, Sung HY, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI
|
|
2
|
Zhou M, Gao Y, Zhang Y, He L, Gao B, Zhang
Y, Claret FX, Calin GA and Wang D: CircZFR/YTHDF3 axis drives lymph
node metastasis in cervical cancer via FASN translation. Mol
Cancer. 24:2182025. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Zhang C, Yuan L, Wen W, Shao C, Liao Y,
Jia Y, Zhao X, Liao Y, Xu D, Chen L, et al: LNMAC promotes cervical
squamous cell carcinoma lymphatic metastasis via epigenetic
regulation of FGF2-induced Lymphangiogenesis. Adv Sci (Weinh).
11:e24046452024. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hyeon DY, Nam D, Shin HJ, Jeong J, Jung E,
Cho SY, Shin DH, Ku JL, Baek HJ, Yoo CW, et al: Proteogenomic
characterization of molecular and cellular targets for
treatment-resistant subtypes in locally advanced cervical cancers.
Mol Cancer. 24:772025. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Cao G, Wang Y, Zeng H, Zhi Y, Guo Y, Xu M,
Ruan Y, Wang Y, Xiao Y, Lu J, et al: Oligoclonal tumor-specific CD8
T-cell revival and IRE1α/XBP1-GDF15-mediated immunosuppressive
niches determine neoadjuvant chemoimmunotherapy efficacy in
cervical cancer. J Immunother Cancer. 13:e0126302025. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Maniotis AJ, Folberg R, Hess A, Seftor EA,
Gardner LM, Pe'er J, Trent JM, Meltzer PS and Hendrix MJ: Vascular
channel formation by human melanoma cells in vivo and in vitro:
Vasculogenic mimicry. Am J Pathol. 155:739–752. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Harris AL: Hypoxia-a key regulatory factor
in tumour growth. Nat Rev Cancer. 2:38–47. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sun B, Zhang D, Zhao N and Zhao X:
Epithelial-to-endothelial transition and cancer stem cells: Two
cornerstones of vasculogenic mimicry in malignant tumors.
Oncotarget. 8:30502–30510. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhao B, Wu M, Hu Z, Ma Y, Wang Q, Zhang Y,
Li Y, Yu M, Wang H and Mo W: Thrombin is a therapeutic target for
non-small-cell lung cancer to inhibit vasculogenic mimicry
formation. Signal Transduct Target Ther. 5:1172020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Huang M, Lin Y, Wang C, Deng L, Chen M,
Assaraf YG, Chen ZS, Ye W and Zhang D: New insights into
antiangiogenic therapy resistance in cancer: Mechanisms and
therapeutic aspects. Drug Resist Updat. 64:1008492022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yeh YH, Hsiao HF, Yeh YC, Chen TW and Li
TK: Inflammatory interferon activates HIF-1α-mediated
epithelial-to-mesenchymal transition via PI3K/AKT/mTOR pathway. J
Exp Clin Cancer Res. 37:702018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
He M, Yang H, Shi H, Hu Y, Chang C, Liu S
and Yeh S: Sunitinib increases the cancer stem cells and
vasculogenic mimicry formation via modulating the
lncRNA-ECVSR/ERβ/Hif2-α signaling. Cancer Lett. 524:15–28. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lacal PM, Atzori MG, Ruffini F, Scimeca M,
Bonanno E, Cicconi R, Mattei M, Bernardini R, D'Atri S, Tentori L
and Graziani G: Targeting the vascular endothelial growth factor
receptor-1 by the monoclonal antibody D16F7 to increase the
activity of immune checkpoint inhibitors against cutaneous
melanoma. Pharmacol Res. 159:1049572020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yu P, Han Y, Meng L, Tian Y, Jin Z, Luo J,
Han C, Xu W, Kong L and Zhang C: Exosomes derived from pulmonary
metastatic sites enhance osteosarcoma lung metastasis by
transferring the miR-194/215 cluster targeting MARCKS. Acta Pharm
Sin B. 14:2039–2056. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang Q, Huang Y, Jiang M, Tang Y, Wang Q,
Bai L, Yu C, Yang X, Ding K, Wang W, et al: The demethylase ALKBH5
mediates ZKSCAN3 expression through the m6A modification to
activate VEGFA transcription and thus participates in MNNG-induced
gastric cancer progression. J Hazard Mater. 473:1346902024.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhang R, Zhang D, Han F, Song X, Zhang Y,
Zhang J, Zhu Q and Qin Y: The deubiquitinase USP7 and E3 ligase
TRIM21 regulate vasculogenic mimicry and malignant progression of
RMS by balancing SNAI2 homeostasis. J Exp Clin Cancer Res.
43:1352024. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Huang XY, Huang ZL, Huang J, Xu B, Huang
XY, Xu YH, Zhou J and Tang ZY: Exosomal circRNA-100338 promotes
hepatocellular carcinoma metastasis via enhancing invasiveness and
angiogenesis. J Exp Clin Cancer Res. 39:202020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liu Y, Tang R, Cao Y, Wu N, Qin Q, Chen Y,
Wei X, Ren J, Sun Y, Zhou H, et al: LIFU/MMP-2 dual-responsive
release of repurposed drug disulfiram from nanodroplets for
inhibiting vasculogenic mimicry and lung metastasis in
triple-negative breast cancer. J Nanobiotechnology. 22:2092024.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shirakawa K, Kobayashi H, Heike Y,
Kawamoto S, Brechbiel MW, Kasumi F, Iwanaga T, Konishi F, Terada M
and Wakasugi H: Hemodynam-ics in vasculogenic mimicry and
angiogenesis of infammatory breast cancer xenograft. Cancer Res.
62:560–566. 2002.PubMed/NCBI
|
|
20
|
Baeten CIM, Hillen F, Pauwels P, de Bruine
AP and Baeten CGMI: Prognostic role of vasculogenic mimicry in
colorectal cancer. Dis Colon Rectum. 52:2028–2035. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Liu R, Yang K, Meng C, Zhang Z and Xu Y:
Vasculogenic mimicry is a marker of poor prognosis in prostate
cancer. Cancer Biol Ther. 13:527–533. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Liu WB, Xu GL, Jia WD, Li JS, Ma JL, Chen
K, Wang ZH, Ge YS, Ren WH, Yu JH, et al: Prognostic signifcance and
mechanisms of patterned matrix vasculogenic mimicry in
hepatocellular carcinoma. Med Oncol. 28 (Suppl 1):S228–S238. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Ding J, Jia X, Zuo B, He J, Yang J and He
Y: A novel monoclonal antibody targeting a novel epitope of
VE-cadherin inhibits vasculogenic mimicry of lung cancer cells.
Oncol Rep. 39:2837–2844. 2018.PubMed/NCBI
|
|
24
|
Tang HS, Feng YJ and Yao LQ: Angiogenesis,
vasculogenesis, and vasculogenic mimicry in ovarian cancer. Int J
Gynecol Cancer. 19:605–610. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Guo QJ, Yuan Y, Jin ZC, Xu T, Gao YB, Wei
HM, Li CH, Hou W and Hua BJ: Association between tumor vasculogenic
mimicry and the poor prognosis of gastric cancer in China: An
updated systematic review and meta-analysis. Biomed Res Int.
2016:24086452016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Fujimoto A, Onodera H, Mori A, Nagayama S,
Yonenaga Y and Tachibana T: Tumour plasticity and extravascular
circulation in ECV304 human bladder carcinoma cells. Anticancer
Res. 26:59–69. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sun B, Zhang D, Zhang S, Zhang W, Guo H
and Zhao X: Hypoxia influences vasculogenic mimicry channel
formation and tumor invasion-related protein expression in
melanoma. Cancer Lett. 249:188–197. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Qie S, Sun BC, Zhao XL, Zhang SW, Sun T,
Gao SY and Wang XH: Correlation between expressions of matrix
metalloproteinase-2& 9 and vasculogenic mimicry in
gastrointestinal stromal tumors. ZhonghuaYi Xue Za Zhi.
89:1106–1109. 2009.(In Chinese).
|
|
29
|
Boccaletto P, Machnicka MA, Purta E,
Piatkowski P, Baginski B, Wirecki TK, de Crécy-Lagard V, Ross R,
Limbach PA, Kotter A, et al: MODOMICS: A database of RNA
modification pathways. 2017 update. Nucleic Acids Res.
46:D303–D307. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
García-Vílchez R, Sevilla A and Blanco S:
Post-transcriptional regulation by cytosine-5 methylation of RNA.
Biochim Biophys Acta Gene Regul Mech. 1862:240–252. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chen YS, Yang WL, Zhao YL and Yang YG:
Dynamic transcriptomic m5 C and its regulatory role in RNA
processing. Wiley Interdiscip Rev RNA. 12:e16392021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wang YQ, Wei JY, Feng LY, Li OW, Huang L,
Zhou SX, Xu YJ, An K, Zhang Y, Chen RY, et al: Aberrant m5C
hypermethylation mediates intrinsic resistance to gefitinib through
the NSUN2/YBX1/QSOX1 axis in EGFR-mutant non-small-cell lung
cancer. Mol Cancer. 22:812023. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang N, Chen RX, Deng MH, Wei WS, Zhou ZH,
Ning K, Li YH, Li XD, Ye YL, Wen JH, et al: m5C-dependent
cross-regulation between nuclear reader ALYREF and writer NSUN2
promotes urothelial bladder cancer malignancy through facilitating
RABL6/TK1 mRNAs splicing and stabilization. Cell Death Dis.
14:1392023. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Pan A, Xue Y, Ruan X, Dong W, Wang D, Liu
Y, Liu L, Lin Y, Tiange E, Lin H, et al: m5C modification of
LINC00324 promotes angiogenesis in glioma through the CBX3/VEGFR2
pathway. Int J Biol Macromol. 257:1284092024. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wang L, Zhang J, Su Y, Maimaitiyiming Y,
Yang S, Shen Z, Lin S, Shen S, Zhan G, Wang F, et al: Distinct
roles of m5C RNA methyltransferase NSUN2 in major gynecologic
cancers. Front Oncol. 25:7862662022. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Chen YJ, Zuo XZ, Wei QL, Xu J, Liu XY, Liu
SL, Wang HC, Luo QY, Wang YY, Yang Y, et al: Upregulation of LRRC
m5C modification-mediated mRNA stability suppresses apoptosis and
facilitates tumorigenesis in cervical cancer. Int J Biol Sci.
19:691–704. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Pearsall SM, Williamson SC, Humphrey S,
Hughes E, Morgan D, Marqués FJ, Awanis G, Carroll R, Burks L, Shue
YT, et al: Lineage plasticity in SCLC generates non-neuroendocrine
cells primed for vasculogenic mimicry. J Thorac Oncol.
18:1362–1385. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Treps L, Faire S and Clere N: Vasculogenic
mimicry, a complex and devious process favoring
tumorigenesis-Interest in making it a therapeutic target. Pharmacol
Ther. 223:1078052021. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Tomita H, Tanaka K, Tanaka T and Hara A:
Aldehyde dehydrogenase 1A1 in stem cells and cancer. Oncotarget.
7:11018–11032. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Bhattacharya S, Calar K and de la Puente
P: Mimicking tumor hypoxia and tumor-immune interactions employing
three-dimensional in vitro models. J Exp Clin Cancer Res.
39:752020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Samuel T, Rapic S, O'Brien C, Edson M,
Zhong Y and DaCosta RS: Quantitative intravital imaging for
real-time monitoring of pancreatic tumor cell hypoxia and stroma in
an orthotopic mouse model. Sci Adv. 9:eade86722023. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Brown BA, Myers PJ, Adair SJ, Pitarresi
JR, Sah-Teli SK, Campbell LA, Hart WS, Barbeau MC, Leong K, Seyler
N, et al: A histone methylation-MAPK signaling axis drives durable
epithelial-mesenchymal transition in hypoxic pancreatic cancer.
Cancer Res. 84:1764–1780. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Sun HZ, Yao N, Cheng SQ, Li LQ, Liu SQ,
Yang Z, Shang GJ, Zhang DF and Yao Z: Cancer stem-like cells
directly participate in vasculogenic mimicry channels in
triple-negative breast cancer. Cancer Biol Med. 16:299–311. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Liu X, He H, Zhang F, Hu X, Bi F, Li K, Yu
H, Zhao Y, Teng X, Li J, et al: m6A methylated EphA2 and VEGFA
through IGF2BP2/3 regulation promotes vasculogenic mimicry in
colorectal cancer via PI3K/AKT and ERK1/2 signaling. Cell Death
Dis. 13:4832022. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Ju RJ, Li XT, Shi JF, Li XY, Sun MG, Zeng
F, Zhou J, Liu L, Zhang CX, Zhao WY and Lu WL: Liposomes, modified
with PTD(HIV-1) peptide, containing epirubicin and celecoxib, to
target vasculogenic mimicry channels in invasive breast cancer.
Biomaterials. 35:7610–7621. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Landoni F, Maneo A, Colombo A, Placa F,
Milani R, Perego P, Favini G, Ferri L and Mangioni C: Randomized
study of radical surgery versus radiotherapy for stage Ib-IIa
cervical cancer. Lancet. 350:535–540. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Quinn MA, Benedet JL, Odicino F,
Maisonneuve P, Beller U, Creasman WT, Heintz APM, Ngan HYS and
Pecorelli S: Carcinoma of the cervix uteri. FIGO 26th annual report
on the results of treatment in gynecological cancer. Int J Gynaecol
Obstet. 95:S43–S103. 2006.
|
|
49
|
Tan LY, Cockshell MP, Moore E, Min KKM,
Ortiz M, Johan MZ, Ebert B, Ruszkiewicz A, Brown MP, Ebert LM and
Bonder CS: Vasculogenic mimicry structures in melanoma support the
recruitment of monocytes. Oncoimmunology. 11:20436732022.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Liang L, Zhu Y, Li J, Zeng J and Wu L:
ALKBH5-mediated m6A modification of circCCDC134 facilitates
cervical cancer metastasis by enhancing HIF1A transcription. J Exp
Clin Cancer Res. 41:2612022. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Wirth M and Schneider G: A
hypoxia-epigenetics axis drives EMT in pancreatic cancer. Cancer
Res. 84:1739–1741. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Guo XF, Qiu W, Li B, Qi YH, Wang SB, Zhao
RR, Cheng B, Han X, Du H, Pan ZW, et al: Hypoxia-induced neuronal
activity in glioma patients polarizes microglia by potentiating RNA
m6A demethylation. Clin Cancer Res. 30:1160–1174. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Xiong JJ, Zhou ZR, Jiang YL, Li QF, Geng
ZH, Guo JH, Yan CJ and Zhang J: Hypoxic stabilization of RIPOR3
mRNA via METTL3-mediated m6A methylation drives breast cancer
progression and metastasis. Oncogene. 43:3426–3441. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Zhang Q, Wei T, Yan LS, Zhu SQ, Jin W, Bai
Y, Zeng YD, Zhang XF, Yin ZX, Yang JL, et al: Hypoxia-Responsive
lncRNA AC115619 encodes a micropeptide that suppresses m6A
modifications and hepatocellular carcinoma progression. Cancer Res.
83:2496–2512. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Meng J, Sun B, Zhao XL, Zhang D, Zhao XM,
Gu Q, Dong XY, Zhao N, Liu PM and Liu YR: Doxycycline as an
inhibitor of the epithelial-to-mesenchymal transition and
vasculogenic mimicry in hepatocellular carcinoma. Mol Cancer Ther.
13:3107–3122. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Sun B, Zhang S, Zhang D, Yin X, Wang S, Gu
Y and Wang Y: Doxycycline influences microcirculation patterns in
B16 melanoma. Exp Biol Med (Maywood). 232:1300–1307. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Liu XY, Wei QL, Yang CY, Zhao HY, Xu J,
Mobet Y, Luo QY, Yang D, Zuo XZ, Chen NX, et al: RNA m5C
modification upregulates E2F1 expression in a manner dependent on
YBX1 phase separation and promotes tumor progression in ovarian
cancer. Exp Mol Med. 56:600–615. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Zhang YY, Li CT, Zhou YJ, Li H, Li J,
Xiong QP, Zhou W, Huang W, Zhang QFC, Xiang YF, et al: A cohort of
mRNAs undergo high-stoichiometry NSUN6-mediated site-specific m5C
modification. Nat Commun. 16:61192025. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Shaw A, Ajit K, Chataignier M and
Gullerova M: Non-coding Y RNA fragments in a complex with YBX1
modulate PARP1 residency at DNA double strand breaks. Nucleic Acids
Res. 53:gkaf5172025. View Article : Google Scholar : PubMed/NCBI
|



















