Introduction
Pancreatic cancer (PC) is a lethal malignancy with a
5-year survival rate of approximately 10% in the USA (1). It is the third leading cause of cancer
death in the US (2), and fourth in
Japan (3). Radical resection
remains the only way to cure this malignancy, despite advancements
in systemic chemotherapy and molecular targeted therapy. One reason
for the poor prognosis of PC is that most patients present no
symptoms until the advanced stage, losing the chance for curative
surgery at initial diagnosis (1).
Additionally, the specific origin of each pancreatic tumor is still
not fully understood. Recent studies using mouse models have
demonstrated that pancreatic damage prompts acinar cells to acquire
ductal features, eventually contributing to tumorigenesis (4). Epigenetic mechanisms induced by
cellular injury or inflammation in the pancreas are involved in the
acinar-to-ductal metaplasia (ADM) process, transforming acinar
cells into ductal-like cells with plasticity (5), which plays a crucial role in PC
development.
The epigenetic landscape of PC is actively explored
(6). For example, DNA methylation,
one of the epigenetic mechanisms, has been found to be upregulated
in chronic pancreatitis (7), a
well-known major inducer of ADM (8). Consequently, genes whose expression is
perturbed by aberrant DNA methylation may play a critical role in
the ADM process. Tobacco use, alcohol consumption, chronic
pancreatitis, obesity, and diabetes are well-known modifiable risk
factors for PC (9,10) that induce epigenetic abnormalities.
Numerous studies have investigated the relationship between these
risk factors and epigenetic mechanisms (11), including DNA methylation, histone
modifications, chromatin remodeling, and non-coding RNA-mediated
gene regulation. These mechanisms regulate gene expression without
altering the underlying DNA sequence. Considering this knowledge,
developing an understanding of the molecular biology, mechanisms,
and origins of pancreatic tumors will be a primary challenge for
clinical oncologists.
We previously identified 56 genes with decreased
expression due to DNA hypermethylation in esophageal cancer
compared with normal esophageal mucosa tissues (12). Among these gene, we focused on the
NCCRP1 gene because Zuo et al (13) reported that NCCRP1 was expressed in
normal pancreas but decreased in PC. However, the precise
expression pattern of NCCRP1 protein in pancreatic tissue,
specifically in acinar and ductal cells, has not been identified.
Additionally, the mechanism regulating NCCRP1 expression in PC has
not been elucidated. In this study, we examined the precise
expression and DNA methylation status of NCCRP1, which has been
reported as one of the methylation target genes in esophageal
cancers, in paired adjacent normal pancreas and PC tissues. We
found that NCCRP1 expression is frequently lost in PC, probably due
to promoter hypermethylation. We further analyzed the relationship
between DNA methylation status of NCCRP1 and PC prognosis and found
that hypermethylation of the NCCRP1 in PC may predict a
poorer prognosis in PC patients.
Materials and methods
Patients
We enrolled 63 patients diagnosed with PC who
underwent surgery at the National Center for Global Health and
Medicine (NCGM) between January 2011 and March 2022. To analyze the
association with anaplastic carcinoma of the pancreas (ACP), we
additionally enrolled 3 ACP cases resected at NCGM before 2010 and
9 ACP cases resected at Juntendo University between January 2011
and September 2018. In all 75 PC cases, 73 formalin-fixed,
paraffin-embedded sections were used for IHC to evaluate NCCRP1
expression. Consent was retrospectively obtained from these
patients in an opt-out format by posting documents on the website
for a comprehensive study in accordance with the guidelines of the
National Center for Global Health and Medicine Research Ethics
Committee (approval no: 2417). For this study, we accessed the
medical records of the patients from April 2024 to October 2025 and
retrospectively obtained patient data. Clinical and pathological
tumor stages were evaluated using the Union for International
Cancer Control (UICC) TNM Classification of Malignant Tumors, 8th
edition (14). In addition, 52 of
75 PC cases with available frozen samples were examined by
pyrosequencing to assess DNA methylation status of NCCRP1. Written
informed consent was obtained from these 52 patients before sample
collection (approval no: 2464).
Surgical procedure and treatment
We perform standard pancreatectomy for PC by
experienced pancreatic surgeons at our institution. Distal
pancreatectomy is performed if the cancer is located to the body
and tail of the pancreas, with or with no concomitant splenectomy.
Pancreaticoduodenectomy (Whipple procedure) is performed to remove
the head of the pancreas involving the resection of the distal
stomach, common bile duct, duodenum, gallbladder.
Pancreaticoduodenectomy with portal vein (PV) and/or superior
mesenteric vein (SMV) resection is performed for PC when PV/SMV
involvement is determined before or during surgery. Patients who
are in a good physical condition after surgery receive adjuvant
chemotherapy with S-1 (TS-1; Taiho Pharmaceutical) for 6 months,
based on the JASPAC 01 study (15).
In recent years, neoadjuvant chemotherapy (NAC) has emerged as a
preferred choice for patients with resectable, borderline
resectable, and locally advanced PC, compared to upfront surgery,
and is considered for patients with a predicted R0 resection. We
have currently been using two cycles of the GS regimen (gemcitabine
plus S-1) as NAC, based on the results of the Prep-02/JSAP05 study
(16).
Immunohistochemical analysis
Immunohistochemical (IHC) staining was performed in
accordance with the method previously reported by Yamada et
al (17) An anti-NCCRP1
antibody (HPA048141; Sigma-Aldrich, St. Louis, MO) and the
ImmPACTTM DAB Peroxidase Substrate kit (Vector Laboratories,
Burlingame, CA) were used to determine NCCRP1 immunoreactivity. All
slides were reviewed by two observers without access to clinical or
pathological data. Based on the proportion of NCCRP1-positive areas
in the immunohistochemically stained tumor tissue, patients were
classified as those who were NCCRP1-negative (≤30% NCCRP1-positive
tumor cells) and NCCRP1-positive groups (>30% NCCRP1-positive
tumor cells).
Cell lines and culture
Human cell lines derived from PC (Capan-2, AsPC-1,
and MiaPaca-2), those derived from head and neck squamous cell
carcinoma (FaDu and Detroit562), and those derived from lung
cancers (HTB-174 and HCC827) were obtained from the American Type
Culture Collection. A human colorectal cancer cell line HCT116 and
an esophageal squamous cell carcinoma (ESCC) cell line KYSE140 were
obtained from the RIKEN BRC (Tsukuba, Japan) through the National
Bio-Resource Project of the MEXT. Capan-2 and HCT116 cells were
cultured in McCoy's 5A medium supplemented with 10% fetal calf
serum (FCS). AsPC-1, HTB-174, and HCC827 cells were maintained in
RPMI1640 medium supplemented with 10% FCS. MiaPaca-2 cells were
cultured in DMEM medium supplemented with 10% FCS and 1 mM sodium
pyruvate. FaDu and Detroit562 cells were cultured in Minimum
Essential Medium supplemented with 10% FCS and 1-mM sodium
pyruvate. KYSE140 were maintained in Ham's F12/RPMI1640 medium
containing 2% FCS. HCT116 cells with genetic disruption of DNMT1
(DNMT1 KO) or DNMT3b (DNMT3b KO), which were previously established
and molecularly validated (18),
were used in the present study. In some experiments, the cells were
cultured in a 24-well plate at a density of 5×104
cells/well for 18 h, and then treated with 5-aza-2′-deoxycytidine
(5-aza-dC; Sigma-Aldrich) or butyrate (Selleck Chemicals, Houston,
TX, USA) for 4 days. These human cell lines, which were
authenticated by the supplier using short tandem repeat testing,
were passaged in our laboratory for fewer than 6 months after
resuscitation.
NCCRP1 ectopic expression
To construct the expression vector of the
NCCRP1 gene, a DNA fragment encoding the full-length ORF
(OriGene Technologies, Rockville, MD, USA) was used as a template,
and the NCCRP1 gene was amplified with polymerase chain
reaction (PCR) using Primestar (Takara Bio, Shiga, Japan). The
NCCRP1 fragment was ligated into the pIRES2-EGFP vector
(CLONTECH, Palo Alto, CA, USA) using an In-Fusion HD Cloning Kit
(CLONTECH). The resulting vector or vector alone was transfected
into MiaPaca-2 cells using lipofectamine LTX reagent (Life
Technologies, Carlsbad, CA, USA). Stably transfectants were
isolated using a MoFlo™ XDP Cell Sorter (Beckman
Coulter, Brea, CA, USA).
Cell proliferation assay
MiaPaca-2 transfectants were seeded in 96-well
plates at 5,000 cells/well, and cell proliferation was measured
using an MTT assay kit (Nacalai Tesque, Kyoto, Japan). Cell
proliferation was investigated at 24-h intervals.
Reverse transcription-quantitative
PCR
Total RNA was isolated from cultured cells using the
RNA easy Mini Kit (QIAGEN, Hilden, Germany). After treating RNA
with DNase I, double-stranded complementary DNA (cDNA) was
synthesized using the High-Capacity cDNA Reverse Transcription Kit
(Applied Biosystems, Foster City, CA, USA). Quantitative PCR was
performed using ABI TaqMan. Threshold cycle numbers (Ct) were
determined using the Sequence Detector software and transformed
using the 2−ΔΔCq method (19) as described by the manufacturer, with
glyceraldehyde-3-phosphate dehydrogenase (GAPDH) as the calibrator
gene according to the manufacturer's instructions. The TaqMan Gene
expression assay IDs for the genes used in this study were NCCRP1,
Hs01583969_m1 and GAPDH, Hs00266705_g1 (Applied Biosystems).
Pyrosequencing
Bisulfite modification was performed by using an
EpiTect Bisulfite Kit (QIAGEN). To assess the NCCRP1 methylation
status, pyrosequencing was performed with PyroMark Gold Q24
reagents and a PyroMark Q24 pyrosequencing machine (QIAGEN). The
PCR primers used in this study were 5′-GGATAGAGAAGGAAGTGGTAGAG-3′
and 5′-TTAAATCCCCAAACTTCCTACCC-3′. The primer used for
pyrosequencing was 5′-ATGTTGGATTTTAATGAGG-3′.
Methylation-specific PCR (MSP)
Using a PyroMark PCR Kit (QIAGEN) as previously
reported, MSP was performed to assess the ZNF382 methylation status
(20). The methylation-specific
primers were 5′-GGCGATTAACGGGTCGTTTC-3′ and
5′-AAAATTTCCAAACCCGACTCG-3′. The unmethylated primers were
5′-GTGGTGATTAATGGGTTGTTTT-3′ and 5′-CAAAATTTCCAAACCCAACTCA-3′.
Following agarose gel electrophoresis of the PCR products, each
band was quantified using the NIH ImageJ software, and the ZNF382
methylation status was presented as the methylated-to-unmethylated
ZNF382 ratio.
Statistical analysis
Data were expressed as means ± standard deviation.
One-way ANOVA followed by Tukey's multiple comparisons test was
used to analyze three or more groups. Paired Student's t test was
used for the comparison of DNA methylation frequencies between
paired adjacent normal and PC tissues. Clinicopathological
characteristics were compared between groups using the unpaired
Student's t-test for continuous variables (age), whereas the
Chi-square test or Fisher's exact test, as appropriate, was used
for categorical variables (sex, tumor type, pT status, pN status,
pM status, and disease stage). These analyses were applied to
comparisons between NCCRP1-positive and NCCRP1-negative patients,
as well as between NCCRP1 hypermethylation and non-hypermethylation
groups. Comparisons between NCCRP1-transfected and
mock-transfected cells were performed using the unpaired Welch's
t-test. Overall survival (OS) was defined as the time from the date
of surgery to death from any cause or the last known follow-up
visit. Comparisons between groups were performed using the log-rank
test. Statistical analyses were performed using the Prism 7
statistical program (GraphPad Software, Inc., La Jolla, CA, USA).
In the univariate and multivariate analyses, the Cox proportional
hazards regression model was used to evaluate the risk ratio using
JMP statistical analysis software (version 17; SAS Institute Inc.,
Cary, NC, USA). All tests were two-tailed, and P-values of <0.05
were considered statistically significant.
Results
Participants
We enrolled 66 patients diagnosed with PC who
underwent surgery at the National Center for Global Health and
Medicine (NCGM) and 9 patients diagnosed with ACP who underwent
surgery at Juntendo University. Among 75 PC cases, we performed IHC
on 73 cases to investigate the protein expression and localization
of NCCRP1 in paired adjacent normal and PC tissues, and
bisulfite-pyrosequencing for 52 cases to assess the frequency of
NCCRP1 methylation in PC tissues. There was no significant
difference in mean age between the IHC group (70.1 years) and the
pyrosequencing group (71.0 years). Eleven (15.1%) cases for IHC
analysis and 16 (30.8%) cases for pyrosequencing received NAC. The
IHC group of 73 cases included 46 patients (63%) received
pancreaticoduodenectomy (PD), and 27 patients (37%) received distal
pancreatectomy (DP). The pyrosequencing group of 52 cases included
32 (61.5%) PD and 20 (38.5%) DP patients. The characteristics of
the cases in each assay are summarized in Table I.
 | Table I.Characteristics of the patients with
pancreatic cancer included in each assay. |
Table I.
Characteristics of the patients with
pancreatic cancer included in each assay.
|
Characteristics | Immunohistology
(n=73) | Pyrosequencing
(n=52) |
|---|
| Mean age ± SD,
years | 70.1±10.8 | 71.0±9.5 |
| Sex, n (%) |
|
|
|
Male | 47 (64.4) | 33 (63.5) |
|
Female | 26 (35.6) | 19 (36.5) |
| pT classification,
n (%) |
|
|
|
T1/2 | 11 (15.1) | 11 (21.2) |
|
T3/4 | 62 (84.9) | 41 (78.8) |
| pN classification,
n (%) |
|
|
| N0 | 26 (35.6) | 19 (36.5) |
| N1 | 47 (64.4) | 33 (63.5) |
| pM classification,
n (%) |
|
|
| M0 | 70 (95.9) | 48 (92.3) |
| M1 | 3 (4.1) | 4 (7.7) |
| Cancer
stagea, n (%) |
|
|
|
IA/IB/IIA | 26 (35.6) | 19 (36.5) |
|
IIB/III/IV | 47 (64.4) | 33 (63.5) |
| Tumor site, n
(%) |
|
|
|
Head | 46 (63.0) | 32 (61.5) |
|
Tail | 27 (37.0) | 20 (38.5) |
| Neoadjuvant
chemotherapy, n (%) | 11 (15.1) | 16 (30.8) |
NCCRP1 expression in adjacent normal
and cancerous tissues of pancreas
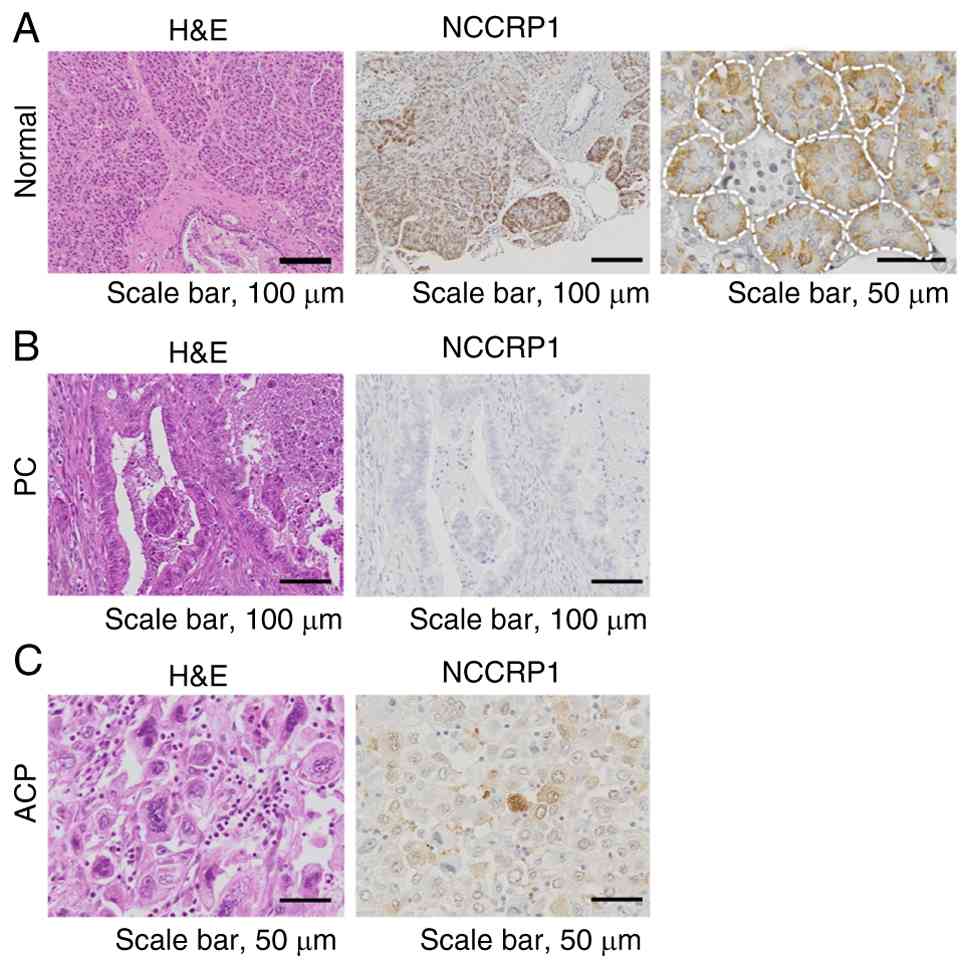
In adjacent normal pancreatic tissues obtained from
patients with PC, NCCRP1 protein was expressed in acinar cells but
not in ductal and islet cells (Fig.
1A). The protein expression of NCCRP1 was confined to the
cytoplasm, with no detection in the nuclei of acinar cells. In
contrast, we found the loss of NCCRP1 expression in the majority of
pancreatic carcinomas (64/73, 86.3%) (Fig. 1B). To investigate the
clinicopathological features of NCCRP1-expressing PC, we classified
PC cases into the NCCRP1-negative and NCCRP1-positive groups. No
significant differences in clinical features including sex, pT
stage, pN stage, pM stage, or cancer stage, were observed between
these groups. Of the nine NCCRP1-positive cases, eight were
pathologically classified as anaplastic carcinoma (ACP), which is
characterized by pleomorphic giant and spindle-shaped cells lacking
glandular differentiation (Fig.
1C). The ratio of ACP in NCCRP1-positive group was
significantly higher than that of pancreatic ductal adenocarcinoma
(PDAC), which exhibits typical ductal structures with desmoplastic
stroma (P<0.0001) (Table
II).
 | Table II.Cancer types of NCCRP1-negative and
-positive patients with pancreatic cancer. |
Table II.
Cancer types of NCCRP1-negative and
-positive patients with pancreatic cancer.
|
Characteristics | Total |
NCCRP1-negative |
NCCRP1-positive | P-value |
|---|
| No. of patients
(%) | 73 | 64 (87.7) | 9 (12.3) |
|
| Cancer subtype,
n |
|
|
|
|
| Ductal
adenocarcinoma | 52 | 51 | 1 |
|
|
Well
differentiated | 16 | 16 | 0 |
|
|
Moderately
differentiated | 23 | 23 | 0 |
|
|
Poorly
differentiated | 13 | 12 | 1 |
|
| Adenosquamous
carcinoma | 2 | 2 | 0 |
|
| Mucinous
carcinoma | 4 | 4 | 0 |
|
| Anaplastic
carcinoma | 13 | 5 | 8 |
<0.0001a |
| Acinar cell
carcinoma | 2 | 2 | 0 |
|
Aberrant DNA hypermethylation
contributes the loss of NCCRP1 in PC
To clarify whether DNA methylation in the NCCRP1
promoter contributes to the reduced NCCRP1 expression in PCs, we
compared its mRNA expression levels in three PC cell lines to those
in cells treated with 5-aza-2′-deoxycytidine (5-aza-dC), a DNA
methyltransferase inhibitor. In all cell lines, treatment with
5-aza-dC resulted in increased expression of NCCRP1 mRNA (Fig. 2A). Next, we quantified NCCRP1
methylation in PC tissues by analyzing CpG methylation levels in 52
paired adjacent normal and PC tissue samples. Significantly
elevated NCCRP1 methylation was observed in cancer tissues compared
to paired adjacent normal mucosa (P<0.0001, Fig. 2B), suggesting that NCCRP1 expression
was silenced by DNA hypermethylation in PCs. To identify which DNA
methyltransferase contributes to NCCRP1 promoter methylation, we
examined NCCRP1 expression in human colorectal cancer HCT116 cells
with knockout (KO) of DNMT1 or DNMT3b. NCCRP1 expression was higher
in DNMT1 KO cells than in parental HCT116 cells, whereas DNMT3b KO
cells showed no change (Fig. 2C),
suggesting that DNMT1 is primarily responsible for NCCRP1 promoter
methylation. Furthermore, we evaluated the effect of
5-aza-2′-deoxycytidine in other cancer cell types, including head
and neck squamous cell carcinoma, esophageal cancer, and lung
cancer. Treatment with 5-aza-2′-deoxycytidine induced NCCRP1
expression in all cancer cell types examined (Fig. 2D), indicating that DNA
methylation-mediated NCCRP1 silencing occurs across multiple cancer
types.
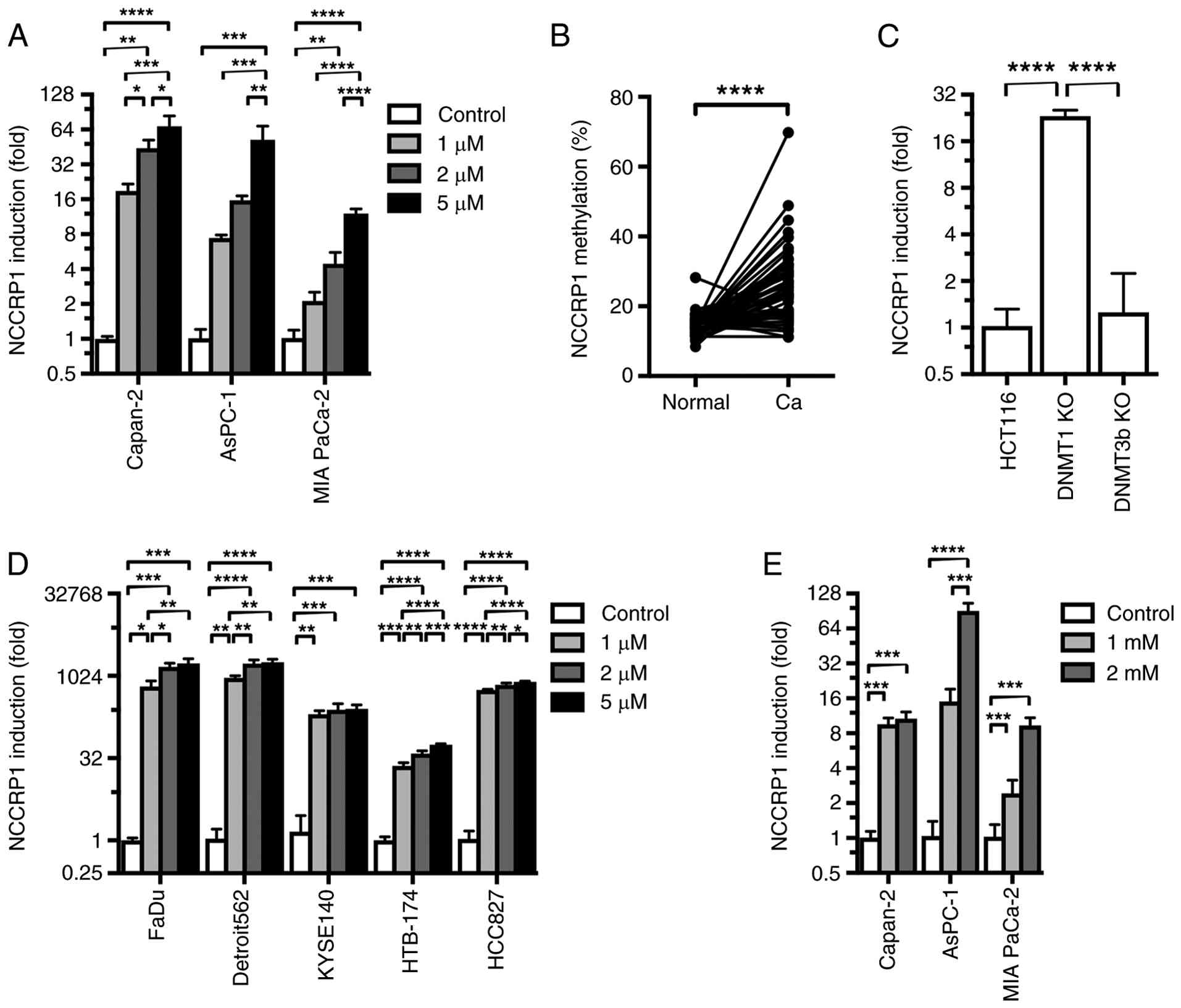 | Figure 2.NCCRP1 expression is silenced by DNA
hypermethylation in PC. (A) NCCRP1 mRNA induction, which is defined
as the fold increase in NCCRP1 mRNA expression levels in PC cell
lines treated with 1, 2 or 5 µM 5-aza-2′-deoxycytidine relative to
untreated control cells. Data from one representative experiment
out of three independent experiments are presented as the mean ± SD
of assays conducted in triplicate, because the absolute values
varied substantially between experiments; however, similar trends
were observed across all independent experiments. (B) DNA
methylation of NCCRP1 was determined via pyrosequencing of paired
samples from 40 patients with PC. The data indicated
hypermethylation in cancerous tissues compared with the mean levels
in paired adjacent normal tissues. (C) Induction of NCCRP1 mRNA in
DNMT1 KO and DNMT3b KO cells compared with parental HCT116 cells.
Data from one representative experiment out of three are presented
as the mean ± SD of assays conducted in triplicate, because
substantial inter-experiment variability in absolute values
precluded pooling of data across independent experiments; however,
consistent trends were reproducibly observed in all experiments.
(D) Induction of NCCRP1 mRNA in head and neck squamous cell
carcinoma cell lines (FaDu and Detroit562), an esophageal cancer
cell line (KYSE140), and lung cancer cell lines (HTB-174 and
HCC827) following treatment with 1, 2 or 5 µM
5-aza-2′-deoxycytidine presented as fold increases in induction
relative to untreated cells. Data from one representative
experiment out of three are presented as the mean ± SD of assays
conducted in triplicate, because substantial inter-experiment
variability in absolute values precluded pooling of data across
independent experiments; however, consistent trends were
reproducibly observed in all experiments. (E) Induction of NCCRP1
mRNA in PC cell lines following treatment with 1 or 2 mM butyrate
presented as fold increases in induction relative to untreated
controls. Data from one representative experiment out of three are
presented as the mean ± SD of triplicate assays, because
substantial inter-experiment variability in absolute values
precluded pooling of data across independent experiments; however,
consistent trends were reproducibly observed in all experiments. (A
and C-E) Statistical analysis was performed using one-way ANOVA
followed by Tukey's multiple comparisons test. (B) Statistical
analysis was performed using a paired Student's t-test. *P<0.05,
**P<0.01, ***P<0.001, ****P<0.0001. Ca, cancerous tissue;
DNMT, DNA methyltransferase; KO, knockout; NCCRP1, nonspecific
cytotoxic cell receptor protein 1; PC, pancreatic cancer. |
To explore alternative regulatory mechanisms, we
treated PC cell lines with butyrate, a histone deacetylase
inhibitor. Butyrate increased NCCRP1 mRNA expression in all three
cell lines, suggesting that histone modifications, in addition to
DNA methylation, may also contribute to NCCRP1 downregulation
(Fig. 2E).
Correlation between survival and
NCCRP1 expression or methylation status in PC
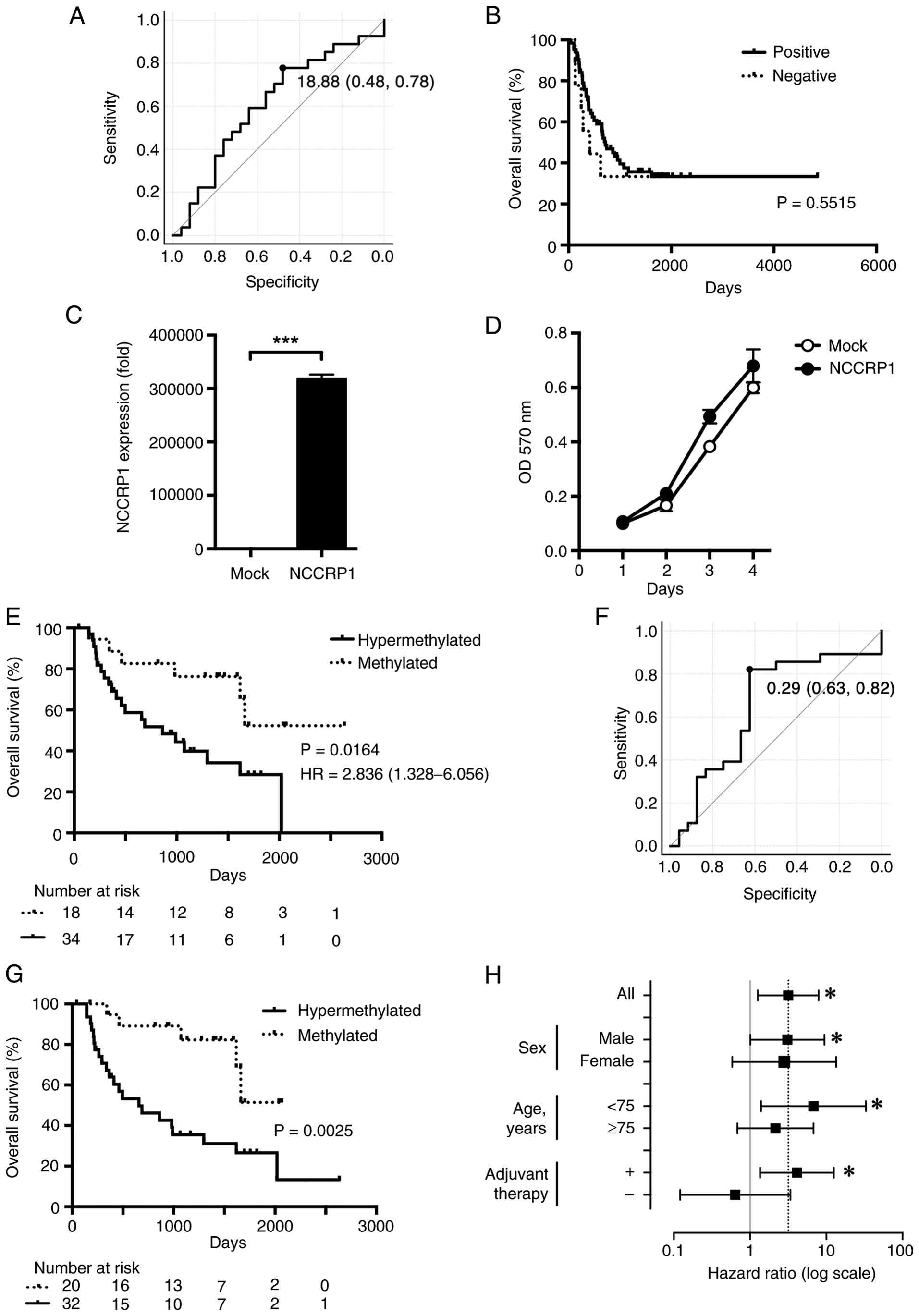
To analyze the relation between the methylation
status of the NCCRP1 gene and clinicopathological features
of PC, we classified PC cases into low and high methylation groups
based on a cut-off value (18.88%) determined using receiver
operating characteristic (ROC) curve analysis (Fig. 3A). Out of the 52 cases examined, 34
(65.4%) demonstrated hypermethylation of NCCRP1. No statistically
significant associations were observed in patients' characteristics
or tumor stages between these groups (Table III). Kaplan-Meier estimates were
then evaluated within both NCCRP1 expression and NCCRP1 methylation
groups. In the prognostic analyses of patients assessed by IHC,
there was no difference between NCCRP1-positive and NCCRP1-negative
groups when considering the entire cohort (Fig. 3B). To clarify the function of the
NCCRP1, we prepared NCCRP1-transfected and
mock-transfected MIAPaCa cells and investigated the effect of
NCCRP1 expression on their growth. Stably
NCCRP1-transfected MIAPaCa cells displayed >30,000-fold
higher NCCRP1 expression than mock-transfected counterparts
(Fig. 3C); however,
NCCRP1-transfected and mock-transfected cells demonstrated
no significant difference in growth (Fig. 3D). In contrast, among patients
analyzed by pyrosequencing, those with hypermethylated NCCRP1
exhibited significantly poorer survival compared to those with
low-methylated NCCRP1 (Fig. 3E,
P=0.0164). In the same cohort, we investigated the methylation
status of ZNF382, one of methylation makers already examined in PC
(20), and classified PC cases into
low and high methylation groups on the basis of the cut-off value
(0.289) determined by ROC curve analysis (Fig. 3F). Patients with ZNF382
hypermethylation also exhibited significantly poorer survival
(Fig. 3G, P=0.0025). Univariate
analysis of OS revealed that NCCRP1 hypermethylation (P=0.014),
advanced pT classification (P=0.022), advanced pN classification
(P=0.012), advanced pStage (P=0.013), and lack of adjuvant
chemotherapy (P=0.003) were significant risk factors (Table IV). No significant differences were
observed with respect to sex or age; however, poorer survival was
noted in the hypermethylated NCCRP1 group compared with the
low-methylated group among male and younger patients (<75 years;
Fig. 3H). In addition, poorer
survival in the hypermethylated group was observed only among
patients who received adjuvant chemotherapy (Fig. 3H). Multivariate analysis identified
NCCRP1 hypermethylation [hazard ratio (HR), 2.602; 95% confidence
interval (CI), 1.012–6.689; P=0.0471], advanced stage (HR, 2.977;
95% CI, 1.215–7.294; P=0.0017), and lack of adjuvant chemotherapy
(HR, 3.771; 95% CI, 1.487–9.563; P=0.005) as independent prognostic
factors (Table V).
 | Table III.Nonspecific cytotoxic cell receptor
protein 1 methylation status and clinical features of pancreatic
cancer. |
Table III.
Nonspecific cytotoxic cell receptor
protein 1 methylation status and clinical features of pancreatic
cancer.
|
Characteristics |
Hypermethylation | Methylation | P-value |
|---|
| No. of patients
(%) | 34 (65.4) | 18 (34.6) |
|
| Mean age ± SD,
years | 70.9±9.9 | 71.3±9.2 | 0.886 |
| Sex, n (%) |
|
| 0.727 |
|
Male | 21 (61.8) | 12 (66.7) |
|
|
Female | 13 (38.2) | 6 (33.3) |
|
| pT classification,
n (%) |
|
| 0.482 |
|
T1/2 | 6 (17.7) | 5 (27.8) |
|
|
T3/4 | 28 (82.4) | 13 (72.2) |
|
| pN classification,
n (%) |
|
| 0.389 |
| N0 | 11 (32.4) | 8 (44.4) |
|
| N1 | 23 (67.7) | 10 (55.6) |
|
| pM classification,
n (%) |
|
| 0.999 |
| M0 | 31 (91.2) | 17 (94.4) |
|
| M1 | 3 (8.8) | 1 (5.6) |
|
| Cancer
stagea, n (%) |
|
| 0.389 |
|
IA/IB/IIA | 11 (32.4) | 8 (44.4) |
|
|
IIB/III/IV | 23 (67.6) | 10 (55.6) |
|
| Tumor site, n
(%) |
|
| 0.963 |
|
Head | 21 (61.8) | 11 (61.1) |
|
|
Tail | 13 (38.2) | 7 (38.9) |
|
| Neoadjuvant
chemotherapy, n (%) | 9 (26.5) | 7 (38.9) | 0.356 |
| Adjuvant
chemotherapy, n (%) | 20 (58.8) | 15 (83.3) | 0.073 |
 | Table IV.Univariate analysis of the risk
factors for overall survival in 52 patients with pancreatic
cancer. |
Table IV.
Univariate analysis of the risk
factors for overall survival in 52 patients with pancreatic
cancer.
| Variables | Hazard ratio | 95% CI | P-value |
|---|
| NCCRP1 |
|
|
|
|
Hypermethylated | 3.171 | 1.266–7.944 | 0.014a |
|
Methylated | 1.000 | - |
|
| Age, years |
|
|
|
|
≥75 | 1.385 | 0.643–2.984 | 0.406 |
|
<75 | 1.000 | - |
|
| Sex |
|
|
|
|
Male | 1.000 | - |
|
|
Female | 1.038 | 0.478–2.255 | 0.925 |
| pT
classification |
|
|
|
|
T3/4 | 4.230 | 1.237–14.461 | 0.022a |
|
T1/2 | 1.000 | - |
|
| pN
classification |
|
|
|
|
≥N1 | 3.038 | 1.276–7.234 | 0.012a |
| N0 | 1.000 | - |
|
| pM
classification |
|
|
|
| M1 | 2.387 | 0.552–10.320 | 0.244 |
| M0 | 1.000 | - |
|
| Cancer
stageb |
|
|
|
| IIB,
III, IV | 3.005 | 1.263–7.147 | 0.013a |
| IA, IB,
IIA | 1.000 | - |
|
| Tumor site |
|
|
|
|
Head | 1.303 | 0.593–2.866 | 0.510 |
|
Tail | 1.000 | - |
|
| Neoadjuvant
chemotherapy |
|
|
|
|
Yes | 1.323 | 0.601–2.913 | 0.487 |
| No | 1.000 | - |
|
| Adjuvant
chemotherapy |
|
|
|
|
Yes | 1.000 | - |
|
| No | 3.562 | 1.537–8.258 | 0.003a |
 | Table V.Multivariate analysis of the risk
factors for overall survival in 52 patients with pancreatic
cancer. |
Table V.
Multivariate analysis of the risk
factors for overall survival in 52 patients with pancreatic
cancer.
| Variables | Hazard ratio | 95% CI | P-value |
|---|
| NCCRP1 |
|
|
|
|
Hypermethylated | 2.602 | 1.012–6.689 | 0.047a |
|
Methylated | 1.000 | - |
|
| Age, years |
|
|
|
|
≥75 | 1.315 | 0.560–3.092 | 0.530 |
|
<75 | 1.000 | - |
|
| Sex |
|
|
|
|
Male | 1.000 | - |
|
|
Female | 1.378 | 0.573–3.312 | 0.474 |
| Cancer
stageb |
|
|
|
| IIB,
III, IV | 2.977 | 1.215–7.294 | 0.017a |
| IA, IB,
IIA | 1.000 | - |
|
| Adjuvant
chemotherapy |
|
|
|
|
Yes | 1.000 |
|
|
| No | 3.771 | 1.487–9.563 | 0.005a |
Discussion
In the present study, we first demonstrated that
NCCRP1 is silenced by aberrant DNA hypermethylation in PC.
Furthermore, we clearly indicated significance of the DNA
methylation status of the acinar cell-specific gene, NCCRP1,
as a prognostic factor. Our findings also indicate that NCCRP1
methylation status is a better indicator for selecting patients
with poor prognosis than its protein expression detected by
antibody.
We found that the patients with PC in which the
NCCRP1 gene was DNA hypermethylated have poorer prognosis as
compared with the patients with lower methylation levels of the
NCCRP1 gene. To our knowledge, there are no reports of
NCCRP1 being methylated in cancer, including PC, with
prognostic value. Our immunostaining results showed that almost all
cases had no NCCRP1 expression, suggesting that evaluation of
NCCRP1 expression by IHC is not suitable to predict prognosis.
Instead, DNA methylation status of the NCCRP1 is a better
indicator for selecting patients with poor prognosis. There are
several reports identifying DNA methylation correlated with
survival prognosis in PDAC (21–23).
Among these abnormally methylated genes in PDAC, ZNF154 and ZNF382
function as tumor suppressors, which may contribute to lower
survival rates in PC (22,24). Therefore, loss of NCCRP1 by aberrant
DNA methylation may has a significant role in PC progression.
Moreover, NCCRP1 and ZNF382 had comparable proportions of high
methylation cases (65.4% vs. 61.5%), with concordance in most cases
(both high methylation in 55.8%; both low in 28.8%). A minority
(9.6%) demonstrated high NCCRP1 but low ZNF382 methylation,
suggesting that NCCRP1 can detect additional high-risk cases missed
by existing markers. Ongoing clinical trials of epigenetic drugs in
solid tumors (25) may encourage
combination strategies involving demethylating agents (e.g., 5-aza
analogs) with chemotherapy for PC. Although adjuvant chemotherapy
is a standard treatment after resection of PC, around half of
resected patients can complete adjuvant chemotherapy and over 75%
of patients experience a recurrence within the first two years
post-resection (26–28). Methylation status information will
be valuable in selecting patients who are suitable for personalized
treatments. Wu et al (7)
reported that circulating cell-free DNA (cfDNA) methylation
signatures can distinguish PC from chronic pancreatitis.
Additionally, liquid biopsies containing tumor-derived cfDNA could
be obtained before surgical resection. Although further studies are
needed to develop cfDNA-based assays for clinical application and
to assess the feasibility of detecting NCCRP1 methylation in cfDNA
as a noninvasive prognostic biomarker, preoperative assessment of
NCCRP1 methylation using liquid biopsies may provide additional
clinical value in decision-making for neoadjuvant therapy.
NCCRP1 was initially identified in fish with a
cytolytic function (29).
Therefore, the role of NCCRP1 in humans is mostly unknown. At the
molecular level, NCCRP1 functions as a paralog within the F-box
superfamily of proteins, which are key components of E3 ubiquitin
ligase complexes known for their regulatory roles in the cell cycle
(30). Zuo et al (13) reported a relationship between
ubiquitination-related genes, including NCCRP1, and PC prognosis.
Although further studies using mass spectrometry to identify
proteins that coprecipitated with NCCRP1 are warranted, it may play
a similar regulatory role in PC. To examine the functional
significance of NCCRP1 loss in PC, we generated NCCRP1-transfected
and mock-transfected MIAPaCa cells. However, no significant
difference in growth was observed between the two groups. In this
study, we also demonstrated that NCCRP1 protein expression was
undetectable in almost all PC tissue samples, irrespective of
methylation status. These findings collectively suggest that NCCRP1
molecular functions may not have a major influence on PC prognosis.
Existing research, for instance, Zhou et al (31) demonstrated that elevated NCCRP1
expression correlates with poor prognosis in triple-negative breast
cancer, a subtype known for its high malignancy and poorer
outcomes. Similarly, in PC, Zuo et al (13) reported that despite the
down-regulation of NCCRP1 expression in tumor tissue, patients with
high NCCRP1 expression have a poorer prognosis, which aligns with
our study results in which ACP with poor prognosis showed
significantly higher expression of NCCRP1. On the other hand, Miwa
et al (32) observed in ESCC
that low NCCRP1 expression was associated with poor prognosis,
emphasizing that NCCRP1 function is context-dependent.
In adjacent normal pancreas obtained from patients
with PC, NCCRP1 expression is restricted to acinar cells. Although
further studies using lineage-tracing mouse models are warranted,
the observation of NCCRP1 hypermethylation in PC may indicate that
the origin of PC could be acinar cells, where NCCRP1 is silenced by
aberrant DNA methylation induced by risk factors such as
pancreatitis, smoking, alcohol consumption, or diabetes. The
understanding of the cellular origins of pancreatic tumors remains
elusive. Previous studies have shown that both acinar and ductal
cells possess the capability to form PDAC (33). Recently, Del Poggetto et al
(4) reported that acinar cells are
the origin of cancer in mouse models. Our findings indicate that PC
may have originated from distinct cell types: those with NCCRP1
hypermethylation (likely derived from acinar cells) and those with
NCCRP1 low-methylation (likely derived from ductal cells). This
indicates that cases where NCCRP1 was low-methylated may have
developed into cancer through alternative mechanisms. It is also
possible that both acinar cells and ductal cells are originated
from common stem cells (34). Beer
et al (35) summarized the
existence of progenitor cells exhibiting squamous epithelial
characteristics among pancreatic ductal, islet, and acinar cells.
In zebrafish and murine models, these progenitor cells were
demonstrated to possess the ability to differentiate into both
ductal and acinar cells, categorized as centroacinar cells. Since
there are no reports indicating that prognosis differs depending on
cell origin, further studies are needed.
ACP has significantly higher expression of NCCRP1.
According to the World Health Organization (WHO) classification of
pancreatic tumors, ACP is defined as undifferentiated carcinoma in
which a significant component of the neoplasm has no specific signs
of differentiation (36). Although
prognosis of ACP is thought to be poor compared to other types of
PC, the clinical features and treatment of ACP remain unknown
because of its rarity. A stratified analysis of ACP cases showed a
trend toward poorer survival among NCCRP1-positive patients
(HR=3.37), although this was not statistically significant
(P=0.1651, data not shown). Interpretations regarding NCCRP1
expression patterns in ACP should be made cautiously due to the
limited sample size. Nonetheless, ACP may undergo carcinogenesis
through a different molecular pathway, including DNA methylation
status.
A limitation of the present study is the use of a
small cohort for pyrosequencing. In addition, several limitations
are considered, including a small sample size and its retrospective
nature, potentially leading to biases. This study also lacked
analysis of patients with late-stage PC because these patients are
often not candidates for surgical resection. The omission of the
lineage-tracing mouse models is also a limitation of the present
study. Although the limited number of samples in the
methylation-survival analysis reduces statistical power, this study
is the first to demonstrate that NCCRP1 hypermethylation in PC
correlates with poor survival prognosis. The poorer survival in
patients with hypermethylated NCCRP1 suggests the importance of
epigenetic landscapes in determining patient prognosis and proposes
a novel therapeutic target.
Acknowledgements
The authors would like to thank Dr Miwa
Tamura-Nakano and Dr Chinatsu Oyama (Electron Microscopy Support
Unit, National Institute for Global Health and Medicine, Shinjuku,
Tokyo, Japan) for their technical support for histological
analysis.
Funding
The present study was supported by JSPS KAKENHI (grant nos.
JP15K10124 and JP19K08457), and by grants from the National Center
for Global Health and Medicine (grant nos. 26-117, 29-1019,
20A1017, 20A3002 and 23A-3001).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
YIK conceived and designed the study. MN, TH and YIK
acquired data. MN and YIK analyzed and interpreted data. MN, KY,
TI, YF, AS, NT and NK contributed to the collection of clinical
specimens and analyzed clinicopathological data. MN and YIK drafted
the manuscript. MN, KY, TH and YIK obtained funding. MN and YIK
confirm the authenticity of all the raw data. All authors read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
The present study was approved by the National
Center for Global Health and Medicine Research Ethics Committee
(approval no. 2464; Shinjuku, Tokyo, Japan), and written informed
consent was obtained from 52 patients before sample collection and
subsequent pyrosequencing. For the remaining patients in whom
archived pathological specimens were used only in the
immunohistochemical analysis, the requirement for written informed
consent was waived because the study was retrospective and
non-interventional. Instead, an opt-out document was posted on the
hospital website before the study commenced and consent was
retrospectively obtained in accordance with the guidelines of the
National Center for Global Health and Medicine Research Ethics
Committee (approval no. 2417).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Mizrahi JD, Surana R, Valle JW and Shroff
RT: Pancreatic cancer. Lancet. 395:2008–2020. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD, Fuchs HE and Jemal
A: Cancer statistics, 2022. CA Cancer J Clin. 72:7–33.
2022.PubMed/NCBI
|
|
3
|
Cancer Statistics in Japan 2024, .
National Cancer Center Institute for Cancer Control, Division of
Cancer Information Service. November 11–2024https://ganjoho.jp/public/qa_links/report/statistics/2024_en.html
|
|
4
|
Del Poggetto E, Ho IL, Balestrieri C, Yen
EY, Zhang S, Citron F, Shah R, Corti D, Diaferia GR, Li CY, et al:
Epithelial memory of inflammation limits tissue damage while
promoting pancreatic tumorigenesis. Science. 373:eabj04862021.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wang L, Xie D and Wei D: Pancreatic
Acinar-to-ductal metaplasia and pancreatic cancer. Methods Mol
Biol. 1882:299–308. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lomberk G, Blum Y, Nicolle R, Nair A,
Gaonkar KS, Marisa L, Mathison A, Sun Z, Yan H, Elarouci N, et al:
Distinct epigenetic landscapes underlie the pathobiology of
pancreatic cancer subtypes. Nat Commun. 9:19782018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wu Y, Seufert I, Al-Shaheri FN, Kurilov R,
Bauer AS, Manoochehri M, Moskalev EA, Brors B, Tjaden C, Giese NA,
et al: DNA-methylation signature accurately differentiates
pancreatic cancer from chronic pancreatitis in tissue and plasma.
Gut. 72:2344–2353. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Halbrook CJ, Wen HJ, Ruggeri JM, Takeuchi
KK, Zhang Y, di Magliano MP and Crawford HC: Mitogen-activated
protein kinase kinase activity maintains acinar-to-ductal
metaplasia and is required for organ regeneration in pancreatitis.
Cell Mol Gastroenterol Hepatol. 3:99–118. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Huang J, Lok V, Ngai CH, Zhang L, Yuan J,
Lao XQ, Ng K, Chong C, Zheng ZJ and Wong MCS: Worldwide burden of,
risk factors for, and trends in pancreatic cancer.
Gastroenterology. 160:744–754. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Quoc Lam B, Shrivastava SK, Shrivastava A,
Shankar S and Srivastava RK: The Impact of obesity and diabetes
mellitus on pancreatic cancer: Molecular mechanisms and clinical
perspectives. J Cell Mol Med. 24:7706–7716. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Mahmoud AM: An overview of epigenetics in
obesity: The role of lifestyle and therapeutic interventions. Int J
Mol Sci. 23:13412022. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Otsubo T, Yamada K, Hagiwara T, Oshima K,
Iida K, Nishikata K, Toyoda T, Igari T, Nohara K, Yamashita S, et
al: DNA hypermethyation and silencing of PITX1 correlated with
advanced stage and poor postoperative prognosis of esophageal
squamous cell carcinoma. Oncotarget. 8:844342017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zuo H, Chen L, Li N and Song Q:
Identification of a ubiquitination-related gene risk model for
predicting survival in patients with pancreatic cancer. Front
Genet. 11:6121962020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Brierley JD, Gospodarowicz MK and
Wittekind C: UICC TNM Classification of Malignant Tumours. 8th
edition. Wiley Blackwell; New York, NY, USA: pp. 101–104. 2017
|
|
15
|
Uesaka K, Boku N, Fukutomi A, Okamura Y,
Konishi M, Matsumoto I, Kaneoka Y, Shimizu Y, Nakamori S, Sakamoto
H, et al: Adjuvant chemotherapy of S-1 versus gemcitabine for
resected pancreatic cancer: A phase 3, open-label, randomised,
non-inferiority trial (JASPAC 01). Lancet. 388:248–257. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Motoi F, Kosuge T, Ueno H, Yamaue H, Satoi
S, Sho M, Honda G, Matsumoto I, Wada K, Furuse J, et al: Randomized
phase II/III trial of neoadjuvant chemotherapy with gemcitabine and
S-1 versus upfront surgery for resectable pancreatic cancer
(Prep-02/JSAP05). Jpn J Clin Oncol. 49:190–194. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yamada K, Hagiwara T, Inazuka F, Sezaki T,
Igari T, Yokoi C, Nohara K, Yamashita S, Dohi T and Kawamura YI:
Expression of the desmosome-related molecule periplakin is
associated with advanced stage and poor prognosis of esophageal
squamous cell carcinoma. Transl Cancer Res. 72018.doi:
10.21037/tcr.2018.01.03.
|
|
18
|
Rhee I, Bachman KE, Park BH, Jair KW, Yen
RW, Schuebel KE, Cui H, Feinberg AP, Lengauer C, Kinzler KW, et al:
DNMT1 and DNMT3b cooperate to silence genes in human cancer cells.
Nature. 416:552–556. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhuang KR, Chen CF, Chan HY, Wang SE, Lee
DH, Chen SC, Shyr BU, Chou YJ, Chen CC, Yuan SH, et al:
Andrographolide suppresses the malignancy of pancreatic cancer via
alleviating DNMT3B-dependent repression of tumor suppressor gene
ZNF382. Phytomedicine. 132:1558602024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Thompson MJ, Rubbi L, Dawson DW, Donahue
TR and Pellegrini M: Pancreatic cancer patient survival correlates
with DNA methylation of pancreas development genes. PLoS One.
10:e01288142015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Mishra NK, Southekal S and Guda C:
Survival analysis of multi-omics data identifies potential
prognostic markers of pancreatic ductal adenocarcinoma. Front
Genet. 10:6242019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wang SS, Hall ML, Lee E, Kim SC, Ramesh N,
Lee SH, Jang JY, Bold RJ, Ku JL and Hwang CI: Whole-genome
bisulfite sequencing identifies stage-and subtype-specific DNA
methylation signatures in pancreatic cancer. iScience.
27:1094142024. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
He W, Lin S, Guo Y, Wu Y, Zhang LL, Deng
Q, Du ZM, Wei M, Zhu W, Chen WJ, et al: Targeted demethylation at
ZNF154 promotor upregulates ZNF154 expression and inhibits the
proliferation and migration of esophageal squamous carcinoma cells.
Oncogene. 41:4537–4546. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Rossi A, Zacchi F, Reni A, Rota M,
Palmerio S, Menis J, Zivi A, Milleri S and Milella M: Progresses
and pitfalls of epigenetics in solid tumors clinical trials. Int J
Mol Sci. 25:117402024. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chikhladze S, Lederer AK, Kousoulas L,
Reinmuth M, Sick O, Fichtner-Feigl S and Wittel UA: Adjuvant
chemotherapy after surgery for pancreatic ductal adenocarcinoma:
Retrospective real-life data. World J Surg Oncol. 17:1852019.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Klaiber U, Hackert T and Neoptolemos JP:
Adjuvant treatment for pancreatic cancer. Transl Gastroenterol
Hepatol. 4:272019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
van Roessel S, van Veldhuisen E,
Klompmaker S, Janssen QP, Abu Hilal M, Alseidi A, Balduzzi A,
Balzano G, Bassi C, Berrevoet F, et al: Evaluation of adjuvant
chemotherapy in patients with resected pancreatic cancer after
neoadjuvant FOLFIRINOX treatment. JAMA Oncol. 6:1733–1740. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Reimers K, Abu Qarn M, Allmeling C, Bucan
V and Vogt PM: Identification of the non-specific cytotoxic cell
receptor protein 1 (NCCRP1) in regenerating axolotl limbs. J Comp
Physiol B. 176:599–605. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kallio H, Tolvanen M, Jänis J, Pan PW,
Laurila E, Kallioniemi A, Kilpinen S, Tuominen VJ, Isola J,
Valjakka J, et al: Characterization of non-specific cytotoxic cell
receptor protein 1: A new member of the lectin-type subfamily of
F-box proteins. PLoS One. 6:e271522011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhou J, Qian W, Huang C, Mai C, Lai Y, Lin
Z and Lai G: Combined targeting of KRT23 and NCCRP1 as a potential
novel therapeutic approach for the treatment of triple-negative
breast cancer. Gland Surg. 11:16732022. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Miwa T, Kanda M, Koike M, Iwata N, Tanaka
H, Umeda S, Tanaka C, Kobayashi D, Hayashi M and Yamada S:
Identification of NCCRP1 as an epigenetically regulated tumor
suppressor and biomarker for malignant phenotypes of squamous cell
carcinoma of the esophagus. Oncol Lett. 14:4822–4828. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ferreira RM, Sancho R, Messal HA, Nye E,
Spencer-Dene B, Stone RK, Stamp G, Rosewell I, Quaglia A and
Behrens A: Duct-and acinar-derived pancreatic ductal
adenocarcinomas show distinct tumor progression and marker
expression. Cell Rep. 21:966–978. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Huang L, Desai R, Conrad DN, Leite NC,
Akshinthala D, Lim CM, Gonzalez R, Muthuswamy LB, Gartner Z and
Muthuswamy SK: Commitment and oncogene-induced plasticity of human
stem cell-derived pancreatic acinar and ductal organoids. Cell Stem
Cell. 28:1090–1104. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Beer RL, Parsons MJ and Rovira M:
Centroacinar cells: At the center of pancreas regeneration. Dev
Biol. 413:8–15. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
World Health Organization (WHO), . WHO
Classification of Tumours Editorial Board: Digestive System
Tumours. 5th edition. WHO; Geneva: 2019
|