Introduction
Lung cancer is a major cause of cancer-related
mortality globally, with >2.4 million new cases and when it
spreads progressively to distant metastatic sites, including the
liver, adrenal glands, bone and brain, it accounts for >1.8
million deaths annually (1–3). Bone metastasis occurs in ~40% of
patients with advanced lung cancer and substantially affects the
quality of life, survival and therapeutic management in patients
(1). Spinal cord compression,
hypercalcemia and bone marrow aplasia are frequently reported in
patients with bone metastasis, who often suffer from severe pain,
movement difficulties and frequent fractures (4). The median survival time following the
diagnosis of bone metastasis is typically <1 year, reflecting
the urgent need for an early and accurate diagnosis to optimize
therapeutic decision-making, symptom management, palliative care
and improve survival outcomes (5,6).
Traditional diagnostic workflow for bone metastasis
in patients with lung cancer involves a combination of laboratory
findings, imaging techniques, and when indicated, pathological
examination of bone biopsies (7).
Hypercalcemia is an important finding for screening and
observational monitoring of bone metastasis, although it is not
specific and numerous false-positive results necessitate combined
diagnostic approaches (8). Imaging
techniques, particularly advanced radiological modalities, account
for the majority of diagnostic approaches, including X-ray
radiography, bone scintigraphy (scan), magnetic resonance imaging,
computed tomography (CT) and positron emission tomography/CT with
18F-fluorodeoxyglucose (9). When
imaging results confirm the existence of bone metastasis, a biopsy
examination of the suspicious bone region is requested by the
clinician to ascertain changes in osteocytes and bone tissue
(10).
Despite technological improvements in medical
imaging, their accurate diagnostic values present a substantial
challenge even for senior radiologists due to multifaceted factors,
such as complexity in metastatic sites, lesion heterogeneity and
overlapping appearance with benign bone diseases, particularly in
the early metastatic phase, as well as the difference in instrument
modalities, which may be different in qualification (11). Along with these confounding factors,
the interpretation of radiographic images is completely dependent
on professionals and interobserver variability, which is usually
dependent on the expertise of the radiologist, in most cases is
inevitable (9). These limitations
have prompted researchers to investigate if they could introduce
new accurate and feature-based computer image processing methods
that can be both quantitative and reproducible for interpreting
imaging results.
Over the past decade, computer-based image
processing techniques, such as artificial intelligence (AI),
machine learning, deep learning, ensemble computational algorithms
and radiomics approaches, have emerged as powerful and efficient
tools to extract features from high-dimensional skeletal imaging
data, facilitating the diagnostic accuracy for bone metastasis.
Previous studies in lung cancer and other malignancies have
introduced the use of computer-based models (e.g., radiomics
signatures, convolutional neural network, AI and neural-network
assisted bone scintigraphy) as a promising method to aid disease
management. Nevertheless, the diagnostic values of such techniques
in clinical decision-making yet remains unestablished owing to the
lack of robust pooled evidence (12–16).
For instance, a deep learning algorithm was developed by Noguchi
et al (11) that could
automatically diagnose if bone metastasis existed across CT scanned
images with a sensitivity of 89.8% (P<0.001) (12). Computer-based image processing
techniques focus on the predefined raw data driven from the
original images (such as texture, intensity and shape), use
classical algorithms aiming to determine thresholds for the primary
features, proceed deeper to develop a primary prediction model and
finally learn, as well as improve, determining features' thresholds
hierarchically to establish a precise prediction model for the
diagnosis of bone metastasis. Previous studies in this field have
introduced some diagnostic models such as machine learning
classifiers (e.g., random forests, support vector machines and
k-nearest neighbors) for pattern recognition from radiographic
results, deep learning models such as convolutional neural network
(CNN) and U-Net approaches for directly learning deterministic
features from the original images, along with artificial
intelligent pattern recognition techniques (13,14,16–19).
However, most of them have been typically performed on heterogenous
primary sites, a single or limited computational modalities or
general studies across different cancer types, which confounding
their reliability and application in patients with lung cancer and
bone metastasis.
To address this gap, despite a growing body of
evidence emphasizing the advantages of using computer-based
techniques in detecting bone metastasis, there is substantial a
need to confirm their accuracy through establishing a robust
standardized meta-analysis uniquely in patients with lung cancer
that can be further extended for their application in clinical
workflows. To the best of our knowledge, no prior standardized
meta-analyses have been specifically performed in this field.
Therefore, the present systematic review and meta-analysis
specifically evaluated the diagnostic accuracy of computer-based
image processing techniques by appraising the confusion
matrix-based metrics across eligible studies for the diagnosis of
bone metastasis in patients with lung cancer. By focusing
exclusively on patients with lung cancer and providing the robust
pooled accuracy metrics, such as sensitivity, specificity and area
under the curve (AUC), the findings of this meta-analysis provide
evidence assisting clinicians in applying advanced computational
image feature extracting tools in diagnostic approaches,
particularly for screening and ruling out non-metastatic cases in a
high-risk population.
Materials and methods
Study design
The methodology for the present systematic review
and meta-analysis followed the Preferred Reporting Items for
Systematic reviews and Meta-Analysis (PRISMA) guidelines to ensure
transparency, reproducibility and methodology robustness (20). In addition, its protocol was
registered in the International Prospective Register of Systematic
Reviews (PROSPERO; http://www.crd.york.ac.uk/prospero/) under submission
ID 1134225.
Considering the aim of the present meta-analysis,
informed by the population, intervention, comparator, outcome
(PICO) framework, the research questions and objectives focused on
cross-sectional studies, diagnostic cohort studies and randomized
controlled trial studies ascertaining the accuracy of different
image processing techniques (intervention) in patients with lung
cancer metastatic to the bone tissues (population) that were
compared with other imaging-based diagnostic assays from
non-metastatic solid tumors (comparator). The diagnostic accuracy
of the confusion matrix was regarded as the primary outcome for the
meta-analysis (outcome), including sensitivity, specificity,
positive predictive value (PPV), negative predictive value (NPV)
and AUC (21).
Search strategy and eligibility
criteria
A comprehensive search of the literature published
between January 2010 and the end of December 2024 was conducted
across different electronic databases, including PubMed (https://pubmed.ncbi.nlm.nih.gov/), Scopus
(https://www.scopus.com/), Embase (https://www.embase.com/), Cochrane library (https://www.cochranelibrary.com/), Web of science
(https://clarivate.com/academia-government/scientific-and-academic-research/research-discovery-and-referencing/web-of-science/),
and clinical trials.gov (https://clinicaltrials.gov/). The search strategy was
to combine Boolean operators with the medical subject heading terms
of specific key words related to cancer bone metastasis, image
modalities and processing techniques, along with confusion matrix
parameters. Key words applied in the search across different
databases are reported in Table
I.
 | Table I.Key words assessed for the study
selection. |
Table I.
Key words assessed for the study
selection.
| Database | Key words | Results, n |
|---|
| Cochrane
library | (‘bone metastasis’
OR ‘bone neoplasms’ OR metastasis) AND (‘lung cancer’ OR ‘pulmonary
carcinoma’ OR ‘lung neoplasm’ OR ‘Pulmonary Neoplasm’ OR ‘Pulmonary
Cancer’ OR ‘Non-Small-Cell Lung cancer’ OR ‘Small Cell Lung
Carcinoma’ OR ‘Lung Adenocarcinoma’ OR ‘Bronchial Neoplasms’) AND
(‘image processing’ OR radiomics OR ‘deep learning’ OR ‘artificial
intelligence’) AND (‘predictive performance’ OR ‘predictive
accuracy’ OR ‘prediction performance’ OR ‘confusion matrix’ OR
‘performance metrics’ OR ‘diagnostic accuracy’ OR sensitivity OR
recall OR ‘True positive rate’ OR tpr OR ‘True negative rate’ OR
specificity OR accuracy OR ‘area under the roc curve’ OR AUC) | 26 |
| PubMed | ((‘bone
metastasis’[Title/Abstract] OR ‘bone neoplasms’[Title/Abstract] OR
metastasis [Title/Abstract]) AND (‘lung cancer’[Title/Abstract] OR
‘pulmonary carcinoma’[Title/Abstract] OR ‘lung
neoplasm’[Title/Abstract] OR ‘Pulmonary Neoplasm’[Title/Abstract]
OR ‘Pulmonary Cancer’[Title/Abstract] OR ‘Non-Small-Cell Lung
cancer’[Title/Abstract] OR ‘Small Cell Lung
Carcinoma’[Title/Abstract] OR ‘Lung Adenocarcinoma’[Title/Abstract]
OR ‘Bronchial Neoplasms’[Title/Abstract]) AND (‘image
processing’[Title/Abstract] OR radiomics [Title/Abstract] OR ‘deep
learning’[Title/Abstract] OR ‘artificial
intelligence’[Title/Abstract] OR image[Title/Abstract]) AND
(‘confusion matrix’ OR ‘performance metrics’ OR ‘diagnostic
accuracy’ OR sensitivity OR recall OR ‘True positive rate’ OR tpr
OR ‘True negative rate’ OR specificity OR accuracy OR ‘area under
the roc curve’ OR AUC)) AND (‘2010/01/01’ [Date-Publication] :
‘2024/12/31’[Date-Publication]) | 288 |
| Scopus | ( ALL ( ‘predictive
performance’ OR ‘predictive accuracy’ OR ‘prediction performance’
OR ‘confusion matrix’ OR ‘performance metrics’ OR ‘diagnostic
accuracy’ OR sensitivity OR recall OR ‘True positive rate’ OR tpr
OR ‘True negative rate’ OR specificity OR accuracy OR ‘area under
the roc curve’ OR AUC ) AND TITLE-ABS-KEY ( ‘bone metastasis’ OR
‘bone metastases’ OR ‘osseous metastasis’ OR ‘Neoplasm Metastasis’
OR ‘Neoplasm Metastases’ ) AND TITLE-ABS-KEY ( ‘image processing’
OR ‘image analysis’ OR ‘medical imaging’ OR ‘computer vision’ OR
radiomics OR ‘deep learning’ OR ‘artificial intelligence’ ) AND
TITLE-ABS-KEY ( ‘lung cancer’ OR ‘pulmonary carcinoma’ OR ‘lung
neoplasm’ OR ‘Pulmonary Neoplasm’ OR ‘Pulmonary Cancer’ OR
‘Non-Small-Cell Lung cancer’ OR ‘Small Cell Lung Carcinoma’ OR
‘Lung Adenocarcinoma’ OR ‘Bronchial Neoplasms’ ) ) AND PUBYEAR >
2009 AND PUBYEAR < 2025 | 225 |
| Embase | (‘lung cancer’ OR
‘pulmonary carcinoma’ OR ‘lung neoplasm’ OR ‘pulmonary neoplasm’ OR
‘pulmonary cancer’ OR ‘non-small-cell lung cancer’ OR ‘small cell
lung carcinoma’ OR ‘lung adenocarcinoma’ OR ‘bronchial neoplasms’)
AND (‘bone metastasis’ OR ‘bone metastases’ OR ‘osseous metastasis’
OR ‘neoplasm metastasis’ OR ‘neoplasm metastases’) AND (‘predictive
performance’ OR ‘predictive accuracy’ OR ‘prediction performance’
OR ‘confusion matrix’ OR ‘performance metrics’ OR ‘diagnostic
accuracy’ OR sensitivity OR recall OR ‘true positive rate’ OR tpr
OR ‘true negative rate’ OR specificity OR accuracy OR ‘area under
the roc curve’ OR auc) AND (‘image processing’ OR ‘image analysis’
OR ‘medical imaging’ OR ‘computer vision’ OR radiomics OR ‘deep
learning’ OR ‘artificial intelligence’) AND [2010-2024]/py | 167 |
| Web of science | (‘predictive
performance’ OR ‘predictive accuracy’ OR ‘prediction performance’
OR ‘confusion matrix’ OR ‘performance metrics’ OR ‘diagnostic
accuracy’ OR sensitivity OR recall OR ‘True positive rate’ OR tpr
OR ‘True negative rate’ OR specificity OR accuracy OR ‘area under
the roc curve’ OR AUC) AND (‘bone metastasis’ OR ‘bone metastases’
OR ‘osseous metastasis’ OR ‘Neoplasm Metastasis’ OR ‘Neoplasm
Metastases’) AND TS=(‘image processing’ OR ‘image analysis’ OR
‘medical imaging’ OR ‘computer vision’ OR radiomics OR ‘deep
learning’ OR ‘artificial intelligence’) AND (‘lung cancer’ OR
‘pulmonary carcinoma’ OR ‘lung neoplasm’ OR ‘Pulmonary Neoplasm’ OR
‘Pulmonary Cancer’ OR ‘Non-Small-Cell Lung cancer’ OR ‘Small Cell
Lung Carcinoma’ OR ‘Lung Adenocarcinoma’ OR ‘Bronchial Neoplasms’)
AND PY=2010-2024 | 102 |
| Clinical
trials.gov | Lung AND (‘bone
metastasis’ OR ‘bone neoplasms’ OR metastasis) AND (‘image
processing’ OR radiomics OR ‘deep learning’ OR ‘artificial
intelligence’) AND (‘confusion matrix’ OR ‘performance metrics’ OR
‘diagnostic accuracy’ OR sensitivity OR specificity OR accuracy OR
AUC) AND (‘diagnose’ OR ‘diagnosis’ OR ‘predict’ OR
‘predictive’) | 45 |
Inclusion and exclusion criteria for
article selection
Studies were eligible for inclusion if they were
original research articles (cross-sectional, diagnostic cohorts or
randomized controlled studies), written in English, that met the
following criteria: i) Studies on adult patients with lung cancer
and bone metastasis confirmed by the results of histological or
radiological examination; and ii) studies that evaluated the
diagnostic accuracy of at least one computer-based image processing
technique, such as radiomics, machine learning or deep learning,
considering sufficient data, including true-positive,
false-positive, true-negative, false-negative, or at least one
parameter associated with the performance matrix, such as
sensitivity, specificity, accuracy, precision and AUC.
Studies were excluded if they had been written in a
non-English language, reported non-original studies such as review
articles (systematic, narrative or meta-analysis), or were
conference abstracts, editorials, case reports and animal studies.
In addition, considering the aim of the present study, studies that
were performed focusing on primary tumors, those that did not
report any parameters associated with the confusion matrix and
those that did not utilize computer-based image processing
modalities (e.g., traditional radiographic interpretation), or used
them only as an additional technique without further processing,
were also excluded from the study. The other criteria for exclusion
were associated with quality assessment, and the studies were
excluded if three or more high-risk domains were identified.
Study selection process
A comprehensive literature search across the
databases was performed by two independent reviewers, who screened
the titles, abstracts and key words. All screened studies were
imported into EndNote 21 reference manager software (Clarivate Plc)
and duplications were removed. Reviewers were equipped with a
designed checklist in Microsoft Excel (Microsoft Corporation) to
select studies through full-text assessment in the next step
according to the predefined inclusion and exclusion criteria for
eligibility of selected articles. Discrepancies between reviewers
were resolved through consensus or consultation with a third
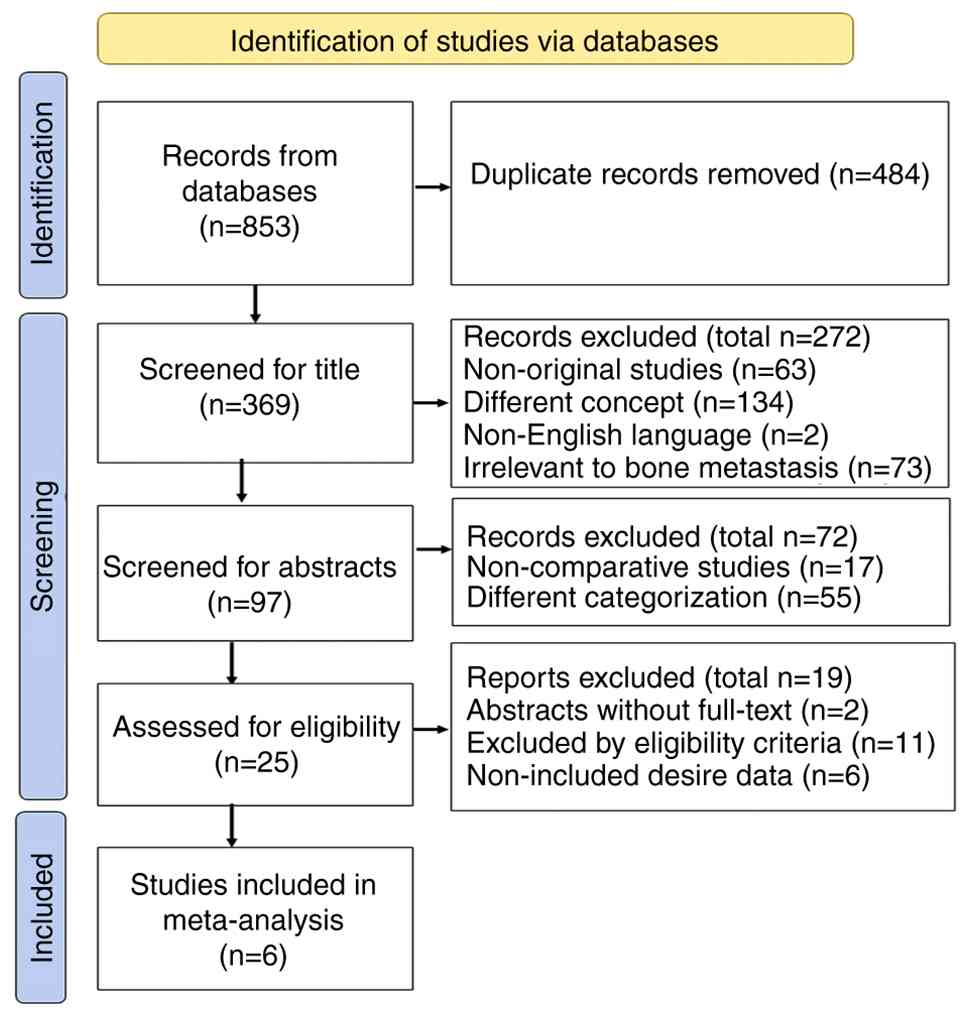
reviewer. The entire study selection process, documented in a
PRISMA flow diagram, is displayed in Fig. 1.
Data extraction
Data were systematically extracted using a
structured Microsoft spreadsheet form, piloted on a subset of
samples and iteratively improved. The data extraction form was
constructed in three main modules including the study
characteristics (authors, country and institution, study design,
sample size, primary cancer types and imaging modality), image
processing techniques (type of machine learning algorithm, CNN, AI
and radiomics) and diagnostic performance metrics [confusion matrix
values, sensitivity, specificity, accuracy, area under the summary
receiver operating characteristic (AUC SROC) curve, and confidence
intervals (CIs) or standard errors].
Quality and risk of bias
assessment
The methodological quality and risk of bias of the
included studies were evaluated independently by two reviewers
using the Newcastle-Ottawa scale (NOS) and the Quality Assessment
of Diagnostic Accuracy Studies-2 (QUADAS-2) tools (22,23).
In this context, four main domains were considered for QUADAS-2
scoring, including patient selection (possibility of bias owing to
mistakes in including participants), index test (blindness and
predefined image processing protocol), reference standard, and flow
and timing, were scored as low, high or unclear risk of bias.
Studies with three or more high-risk domains were excluded from the
meta-analysis. On the other hand, NOS scoring was considered
according to three main domain including selectivity,
comparability, and exposure/outcome with a maximum value of nine
score. Disagreements in bias assessments were resolved through
consensus.
Statistical analysis and
meta-analysis
A bivariant random-effect meta-analysis applying the
Reitsma model was performed to calculate pooled sensitivity and
specificity across finally included studies (24,25).
The extracted data from the included studies were imported into a
Microsoft Excel spreadsheet and the statistical analyses were
conducted in R (version 4.4.1), (https://cran.r-project.org/bin/windows/base/old/4.4.1/)
using the ‘mada’ (version 0.5.12), (https://cran.r-project.org/web/packages/mada/index.html),
and ‘binom’ (version 1.1–1.1), (https://cran.r-project.org/web/packages/binom/index.html)
packages. The Reitsma model was selected to justify the association
between sensitivity and specificity, as well as to incorporate
between-study variability (24).
The restricted maximum likelihood (REML) method was utilized to
estimate parameters, including pooled sensitivity, specificity,
negative likelihood ratio (LR), positive LR and diagnostic odds
ratio with 95% CI (26).
The confusion matrix components for each study,
including true-positive, true-negative, false-positive and
false-negative results, were extracted from the studies or
alternatively calculated based on the reported sensitivity,
specificity and the sample sizes. The Wilson method, implemented
through ‘binom’ package was employed to compute the 95% CI for the
sensitivity and specificity results (27).
In order to determine the pooled sensitivity and
specificity and their corresponding 95% CIs for meta-analysis,
χ2 equality assessments, and forest plots were generated
using the ‘ggplot2’ R package (https://cran.r-project.org/package=ggplot2), labeled
by the name of the first authors and publication year. To evaluate
the trade-off between sensitivity and specificity, a SROC curve was
plotted using the ‘mada’ package (https://cran.r-project.org/web/packages/mada/index.html).
The overall diagnostic accuracy was assessed by calculating the AUC
and partial AUC (restricted to observed false-positive rate,
normalized).
The robustness of pooled estimates was examined
through one-in/one-out sensitivity analysis. In brief, pooled
sensitivity was estimated iteratively by systematically excluding
one study in each step and recalculation of the bivariate model.
Publication bias was appraised through Deeks' funnel plot asymmetry
test, which is strongly recommended for meta-analyses on the
diagnostic accuracy tests.
The model fit was assessed using log-likelihood,
Akaike information criterion (AIC) and Bayesian information
criterion (BIC). PPVs and NPVs were calculated for a normal
distribution with a mean prevalence of 10% (range, 5–15%).
Continuity correction (0.5) was utilized, where it was necessary to
handle null results in the confusion matrix.
Heterogeneity and inconsistency
assessment
Between-study heterogeneity was assessed both
statistically and clinically. The statistical heterogeneity between
studies was assessed using variance components in the key
performance metrics, including sensitivity, specificity, AUC,
segmentation metrics, such as Dice similarity coefficient (Dice),
intersection over union (IoU), model performance metrics and
I2 estimates as the portion of variability attributed to
heterogeneity. I2 values >50% indicated substantial
heterogeneity. Sources of possible clinical heterogeneity were
ascertained considering patient demographics, differences in
imaging modalities, image processing methods and the study design
characteristics (28,29).
Exploring the source of
heterogeneity
Following the heterogeneity assessment, post-hoc
subgroup analyses and univariate meta-regression were conducted to
examine the potential source of heterogeneity between studies. The
included studies were categorized according to two methodological
characteristics, namely, imaging modalities and computational
interpretating methods. Based on the imaging modalities, studies
were categorized as ‘CT-based’ [including single-source dual-energy
CT (ssDECT), deep learning-based CT and CT based radiomics] versus
‘scintigraphy/single photon emission CT (SPECT)-based’ (including
SPECT bone scintigraphy, bone scintigraphy and AI-based bone
scintigraphy). Based on computational interpretation, the
classifier algorithms were categorized as ‘AI/Deep learning’
(including deep neural networks and CNN-based approaches) versus
‘Other/Radiomics’ (including radiomics feature extraction method
and material decomposition analysis assays). Subsequently
meta-regressions using the REML method were performed to explore if
there were any associations between mediators and diagnostic
performance measures. It should be noted that, owing to the limited
number of eligible studies (n=6), the heterogeneity trends across
studies were also visualized by annotating subgroup characteristics
on forest plots (Fig. 2).
Results
Literature overview, heterogeneity and
inconsistency results
A total of 6 original research studies were included
in the present study (Table II)
(15,30–34).
The studies explored the diagnostic accuracy of image processing
techniques to distinguish metastatic from non-metastatic bone
tissue. The studies employed diverse imaging modalities, including
bone scintigraphy, dual-energy and conventional CT, by applying
different classes of machine learning classifiers, such as CNN,
neural network, deep learning and radiomics (Table SI).
 | Table II.Study characteristics and diagnostic
performance of included research. |
Table II.
Study characteristics and diagnostic
performance of included research.
| First author,
year | Imaging
modality | Processing
technique | Cases, n | Controls, n | Sensitivity % (95%
CI) | Specificity % (95%
CI) | DOR (95% CI) | (Refs.) |
|---|
| Dong et al,
2015 | ssDECT | Spectral curve | 79 | 43 | 93.0 | 93.3 | 162.2 | (30) |
|
|
| analysis |
|
| (0.84–0.96) | (0.81–0.98) | (38.5–683) |
|
| Zhao et al,
2020 | Bone | AI-based | 265 | 337 | 93.5 | 93.5 | 208.9 | (31) |
|
| scintigraphy |
|
|
| (0.90–0.96) | (0.90–0.96) | (108–401) |
|
| Liu et al,
2021 | Bone | CNN | 567 | 686 | 76.2 | 82.5 | 15.1 | (32) |
|
| scintigraphy |
|
|
| (0.72–0.79) | (0.79–0.85) | (11.4–19.9) |
|
| Huo et al,
2023 | CT | Deep learning | 57 | 69 | 89.4 | 85.7 | 50.1 | (33) |
|
|
|
|
|
| (0.79–0.95) | (0.75–0.92) | (17–147) |
|
| Su et al,
2024 | CT | Radiomics | 33 | 117 | 72.7 | 89.7 | 23.3 | (15) |
|
|
|
|
|
| (0.56–0.85) | (0.83–0.94) | (8.8–61.6) |
|
| Wang et al,
2024 | SPECT bone | CNN | 267 | 260 | 80.4 | 80.4 | 16.9 | (34) |
|
| scintigraphy |
|
|
| (0.75–0.85) | (0.75–0.85) | (11–26) |
|
All studies reported diagnostic performance metrics
according to the confusion matrix components (Table SII). Dong et al (30) (2015) evaluated spectral CT imaging
to differentiate osteoblastic metastasis from normal bone lesions.
Statistical heterogeneity was assessed by the results of CT values,
spectral curve slopes and material densities. Although the formal
inconsistency results were not explicitly reported in the study,
however, a high AUC indicated a low inconsistency.
Zhao et al (31) utilized a neural network model for
bone scintigraphy images (31). The
statistical heterogeneity in this study was checked using the range
of the AUC values across cancer subtypes. Low inconsistency was
observed with a consistently high AUC among cancer subtypes.
Liu et al (32) proposed a CNN-based method from bone
scintigraphy to identify bone metastasis. Statistical heterogeneity
was stratified based on lesion burden and included Dice scores
(0.85) and an IoU value of 0.789. Moderate inconsistencies were
detected with a high AUC.
Huo et al (33) proposed a CNN algorithm from CT
imaging in patients with lung cancer and bone metastasis.
Statistical heterogeneity was assessed using Dice coefficient and
IoU values (0.856 and 0.789, respectively). Low inconsistency was
observed, along with a stable AUC (0.879), among the different
cohorts.
Su et al (15) combined CT-based radiomics and
clinical features for developing a predictive model for bone
metastasis in patients with lung adenocarcinoma. Low heterogeneity
and moderate inconsistency were observed, with an AUC of 0.866 for
the proposed combined radiomics predictive model.
Wang et al (34) developed a CNN classifier algorithm
for the diagnosis of bone metastasis based on SPECT images through
continuous preprocessing steps, such as bladder removal and image
fusion. Low statistical heterogeneity was confirmed by an accuracy
of 0.803 and an AUC of 0.848 in the preprocessed bladder-removed
images. Low inconsistency with a stable performance was
observed.
All included studies utilized different CT-based
imaging modalities with different processing techniques, including
ssDECT, neural network algorithms and radiomics techniques, that
were compared with traditional image interpretation in the
diagnosis of bone metastasis. Overall high sensitivities, ranging
from 72.7% in the study by Su et al (15) to 93.5% in the study by Zhao et
al (31), with a median of
86.0%, were reported (95% CI, 78.2–91.3). The results of the
χ2 equality assessment for sensitivity and specificity
confirmed significant differences between studies for sensitivity
(χ2=47.8; df=5; P<0.001) and specificity
(χ2=31.52; df=5; P<0.001).
Quality assessment
The quality of the included studies was ascertained
using NOS and QUADAS-2 tools. Although most of the studies were
retrospectively designed, the results generally indicated
high-quality with robustness in terms of methodology. A total of 4
studies scored eight or more out of a total score of nine on NOS
criteria, with strength in cohort selection, exposure ascertainment
and outcome assessment. The quality assessments in this
meta-analysis were basically assessed using QUADAS2, as the present
study was aimed to determine the diagnostic performance accuracy.
However, it also checked NOS criteria to ensure robustness of the
study selection criteria. In the present study, NOS scoring was
considered according to three main domain including selectivity,
comparability and exposure/outcome and specific scores for each
domain, with a maximum value of nine scores (Table SIII). The QUADAS-2 risk of bias
assessment demonstrated low to moderate risk across the patient
selection, index test interpretation, reference standard assessment
and flow/timing domains. The exclusion criteria for studies were ≥3
high-risk domains. A total of two independent reviewers reported.
One of them reported moderate to high risk of bias and the other
reported moderate risk of bias. In consensus, risk of bias was
reported high, but in only two domains and therefore, the study was
not excluded. The results of the quality assessment are outlined in
Table SIV.
Assessment of the robustness and
publication bias
The results of leave one-in/one-out sensitivity
analysis indicated the robustness of findings in the present study
(Table III). When systematically
excluding process was fallowed, minimal fluctuations were observed
in recalculated pooled estimates: Sensitivity ranged from 0.832 to
0.878, specificity ranged from 0.854 to 0.891, and AUC from 0.908
to 0.942. These minimal fluctuations indicate that the findings are
robust and are not disproportionately influenced by any single
study.
 | Table III.Leave-one-out sensitivity analysis of
pooled diagnostic estimates. |
Table III.
Leave-one-out sensitivity analysis of
pooled diagnostic estimates.
| Study removed | Sensitivity (95%
CI) | Specificity (95%
CI) | AUC | τ_sens | τ_fpr | (Refs.) |
|---|
| Dong et al,
2015 | 0.845
(0.752–0.907) | 0.868
(0.785–0.923) | 0.922 | 0.597 | 0.461 | (30) |
| Zhao et al,
2020 | 0.832
(0.734–0.899) | 0.854
(0.767–0.913) | 0.908 | 0.465 | 0.294 | (31) |
| Liu et al,
2021 | 0.878
(0.792–0.933) | 0.888
(0.812–0.937) | 0.942 | 0.565 | 0.500 | (32) |
| Huo et al,
2023 | 0.857
(0.763–0.918) | 0.881
(0.802–0.932) | 0.932 | 0.668 | 0.545 | (33) |
| Su et al,
2024 | 0.870
(0.779–0.928) | 0.880
(0.800–0.931) | 0.936 | 0.671 | 0.509 | (15) |
| Wang et al,
2024 | 0.870
(0.779–0.928) | 0.891
(0.816–0.939) | 0.940 | 0.647 | 0.456 | (34) |
| Original model | 0.860
(0.782–0.913) | 0.878
(0.822–0.926) | 0.931 | 0.609 | 0.471 |
|
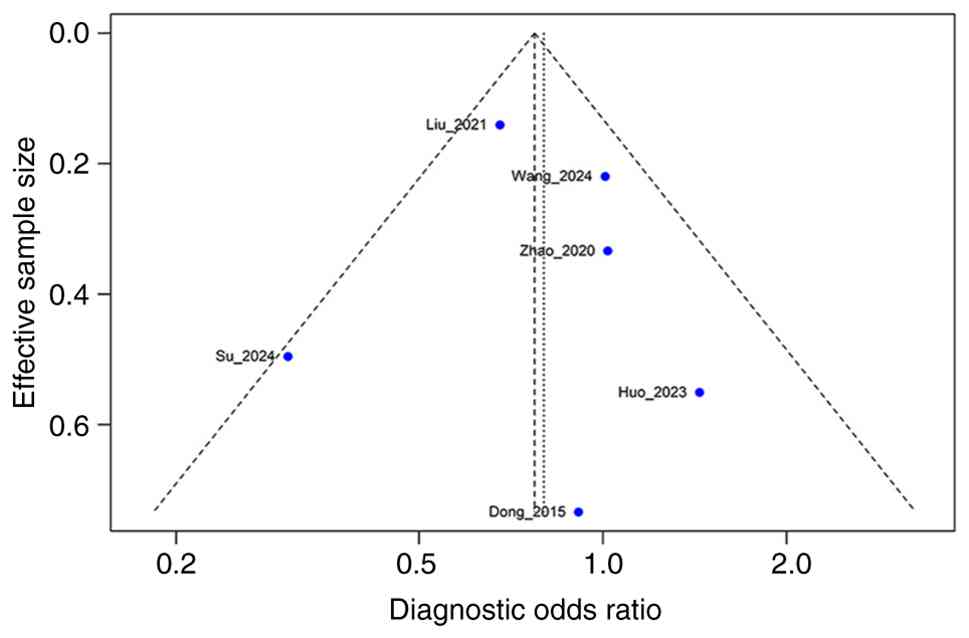
In addition, although the statistical power was
limited by inclusion of only 6 studies, no significant publication
bias was observed when the Deeks' funnel plot asymmetry test was
applied (t=0.18; df=0.4; P=0.867), with a bias estimation of 1.27
(standard error=7.14) (Fig. 2).
Although the statistical power is limited, this non-significant
result refutes any substantial publication bias in the present
meta-analysis.
Exploration of the sources of
heterogeneity
The results of meta-regression following the
subgroup analysis demonstrated clinically important patterns. When
categorization by imaging modalities was considered (Fig. 3, blue lines vs. orange lines),
scintigraphy/SPECT-based studies (n=4) demonstrated a higher mean
sensitivity (88.0±8.1%) compared with CT-based studies (n=2)
(76.5±5.4%); however, the specificity was comparable between groups
(88.8 vs. 85.0%, respectively). The absolute difference in
sensitivities of 12% across distinct categorization suggests that
scintigraphy-based techniques were superior in diagnostic
efficiency for bone metastasis. Considering computational technique
categorization, although a greater variability in sensitivity
metrics was observed in AI/deep learning studies, similarity in
mean sensitivity was observed between AI/deep learning (n=3) and
other/radiomics approaches (n=3) (83.4 vs. 85%, respectively).
Despite limitations in statistical significance due
to the limited number of included studies, the results of
meta-regression of coefficients indicated meaningful effect size as
follows: Imaging modality revealed a coefficient of 0.852 (95% CI,
−0.316 to 2.021; P=0.153) for sensitivity, and the algorithm type
presented a coefficient of 0.165 (95% CI, −1.144 to 1.474;
P=0.805).
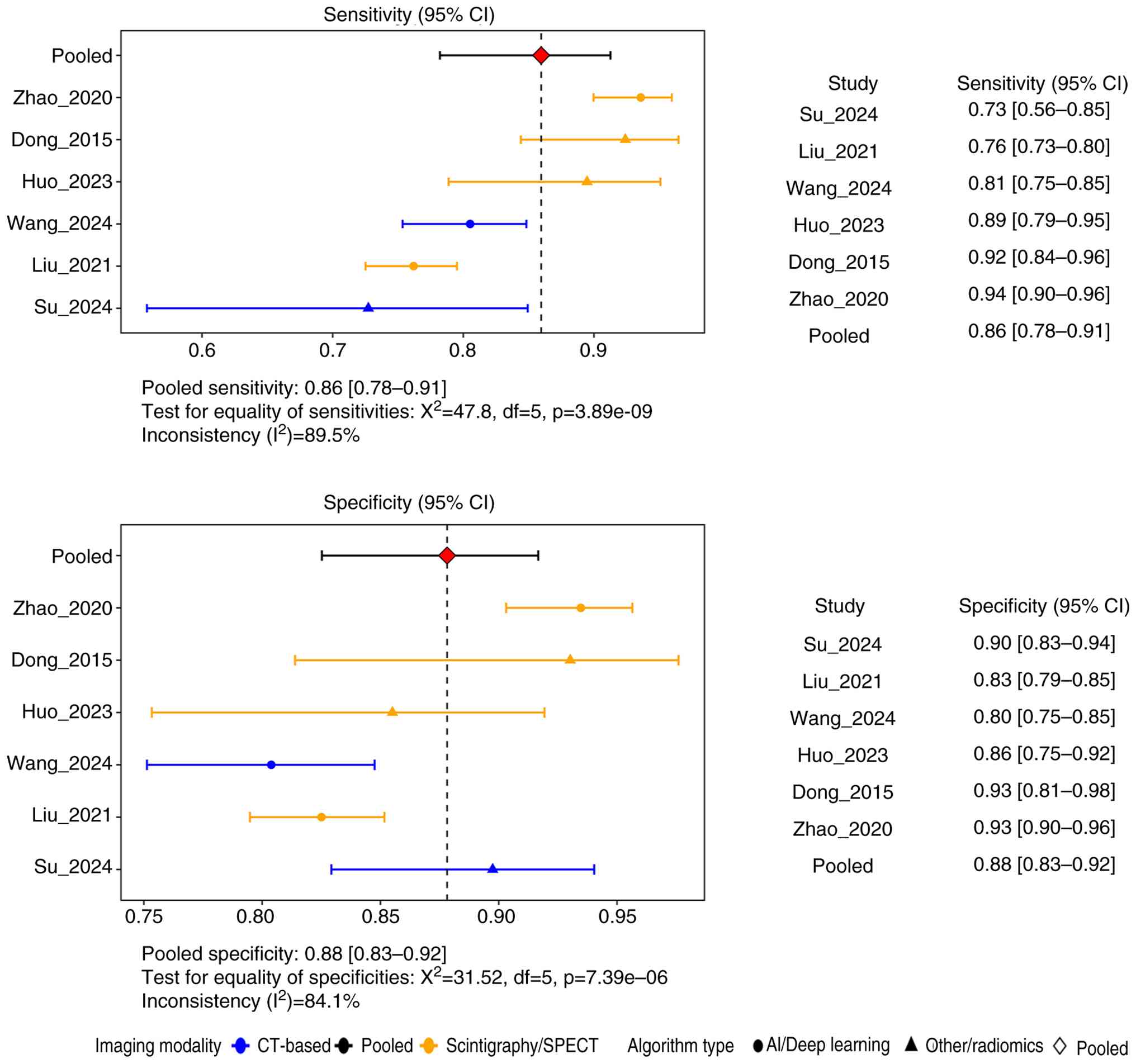
Forest plot visualization and
bivariate Reitsma meta-analysis
In order to calculate the pooled estimate metrics, a
bivariate random-effects Reitsma model was employed for analyzing
the 2,780 subjects from the 6 included studies. Enhanced forest
plots (Fig. 3) exhibit the point
estimates and 95% CIs for diagnostic sensitivities and
specificities, which were subgroup annotated for imaging modality
and algorithm type. As presented in Fig. 3, the pointed sensitivity and 95% CI
had a high variability ranging from 0.73 (95% CI, 0.56–0.85) in the
study by Su et al (15) to
0.94 (95% CI, 0.90–0.96) in the study by Zhao et al
(31), with the specificity ranging
from 0.80 (95% CI, 0.75–0.85) in the study by Wang et al
(34) to 0.93 (95% CI, 0.90–0.96)
in the study by Zhao et al (31). The forest plots represent a wider
range of sensitivity than specificity, with clear clustering of
scintigraphy/SPECT-based studies within higher sensitivity ranges.
Studies using advanced techniques (e.g., scintigraphy/SPECT-based
methods) consistently presented with narrower CIs, indicating
higher precision.
The pooled sensitivity and specificity, with 95%
CIs, are demonstrated in Fig. 2 as
red rectangles with a horizontal boundary of 95% CIs. In this
meta-analysis, the pooled sensitivity and specificity were
estimated as 0.86 (95% CI, 0.78–0.91) and 0.88 (95% CI, 0.83–0.92),
respectively. Log-transformed intersects were 1.81 (95% CI,
1.28–2.35) for sensitivity and 1.98 (95% CI, −2.4 to −1.55) for
false-positive rate (Table IV).
High between-study variability was observed with a standard
deviation of 0.61 for sensitivity and 0.471 for false-positive
rates. In addition, the presence of a significant negative
correlation (the correlation was derived directly from the
variance-covariance matrix of the bivariate Reitsma model) between
the false-positive rate and sensitivity (correlation coefficient,
−0.968; P<0.001) indicating that fewer false-positive results
were achieved in the studies with higher sensitivity.
 | Table IV.Parameter estimates and
goodness-of-fit indices from the bivariate Reitsma meta-analysis
model. |
Table IV.
Parameter estimates and
goodness-of-fit indices from the bivariate Reitsma meta-analysis
model.
|
Parameter/Index | Estimate | 95% CI |
Interpretation/Notes |
|---|
| Fixed effects
(logit scale) |
|
|
|
|
Intercept (sensitivity) | 1.81 | 1.28–2.35 |
|
|
Intercept (false-positive
rate) | −1.98 | −2.40 - −1.55 |
|
| Between-study
variability |
|
|
|
| τ
(sensitivity) | 0.61 | - | High
heterogeneity |
| τ
(false-positive rate) | 0.47 | - | High
heterogeneity |
| Correlation |
|
|
|
|
Sensitivity ~FPR (ρ) | −0.97 | - | P<0.001 |
| Model fit
indices |
|
|
|
|
Log-likelihood | 17.59 | - |
|
|
AIC | −25.19 | - | Lower is
preferred |
|
BIC | −22.76 | - | Lower is
preferred |
| Transformed pooled
estimates |
|
|
|
|
Sensitivity | 0.86 | 0.78–0.91 |
|
|
Specificity | 0.88 | 0.82–0.93 |
|
| AUC (SROC) | 0.93 | - | Excellent
accuracy |
The status of model fit was evaluated by
log-likelihood, AIC and BIC indices. The results supported the
model fit, with a log-likelihood of 17.593, AIC of −25.186 and BIC
of −22.762. These results suggest that the model fit was adequate.
The AUC of the developed model was estimated to be 0.931. This high
AUC indicated an excellent overall diagnostic accuracy of
computational image processing techniques over traditional image
interpretation.
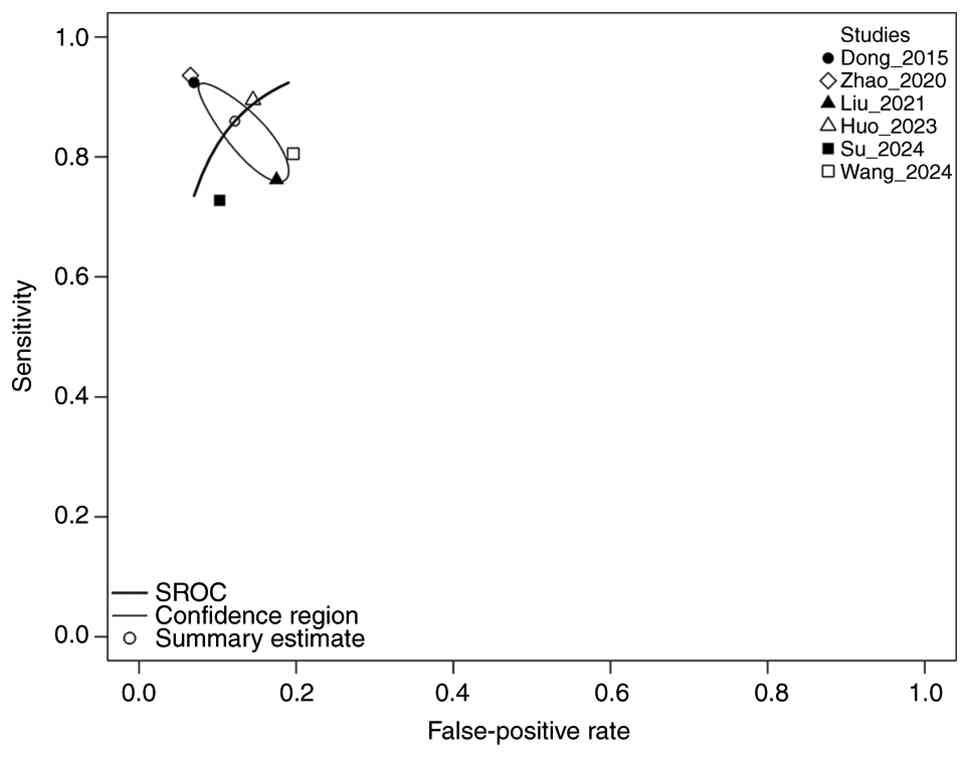
Diagnostic performance
visualization
The SROC curve, a suitable visualization tool for
the trade-off between sensitivity and specificity, was plotted
using the ‘mada’ R package (Fig.
4). In Fig. 4, each individual
study is represented by a unique symbol, while the pooled SROC is
demonstrated by a line with the confidence and prediction region to
reflect uncertainty and variability. The two studies clustered in
the top-left corner of Fig. 4 [Dong
et al (30) and Zhao et
al (31)] exhibited the highest
performance, with high sensitivity and specificity values. The
studies by Liu et al (32),
Su et al (15) and Wang
et al (34) demonstrated a
wider prediction region that indicated lower sensitivity and
moderate heterogeneity, especially in the study by Su 2024, which
can be attributed to the variability in image processing
techniques. However, despite heterogeneity, the AUC of 0.931 and
partial AUC of 0.848 confirmed the acceptable diagnostic accuracy
of the model.
Diagnostic metrics and predictive
values
The ‘SummaryPts’ function was employed to calculate
the summary of diagnostic metrics for bone metastasis. The
diagnostic metrics and predictive values are presented in Table V. The positive LR in the studies by
Zhao et al (31) and Dong
et al (30) had the highest
values (14.34 and 13.25, respectively), with a mean positive LR of
7.22 (95% CI, 4.53–10.9), indicating the moderate to strong ability
of models in the detection of bone metastasis. In the same way, the
studies by Zhao et al (31)
and Dong et al (30)
reported the lowest negative LRs (0.069 and 0.082, respectively),
with a mean of 0.165 (95% CI, 0.096–0.262), which also confirms the
high ability of models to rule out bone metastasis in patients with
lung cancer. The mean inverse negative LR was estimated as 6.48
(95% CI, 3.82–10.40), which also reinforced the reliability of the
negative test results. The mean diagnostic odds ratio (DOR) in the
present study was calculated as 49.30 (95% CI, 17.50–111.00), which
confirmed the strong diagnostic ability of the model for the
diagnosis of bone metastasis.
 | Table V.Diagnostic metrics and predictive
values of constructed model. |
Table V.
Diagnostic metrics and predictive
values of constructed model.
| Metric | Mean | 95% CI |
|---|
| Sensitivity | 0.860 | 0.782–0.913 |
| Specificity | 0.878 | 0.822–0.926 |
| Positive likelihood
ratio | 7.22 | 4.53–10.90 |
| Negative likelihood
ratio | 0.165 | 0.096–0.262 |
| Inverse negative
likelihood ratio | 6.48 | 3.82–10.40 |
| Diagnostic odds
ratio | 49.30 | 17.50–111.00 |
| Negative predictive
value (5–15%) | - | 0.972–0.991 |
| Positive predictive
value (5–15%) | - | 0.273–0.554 |
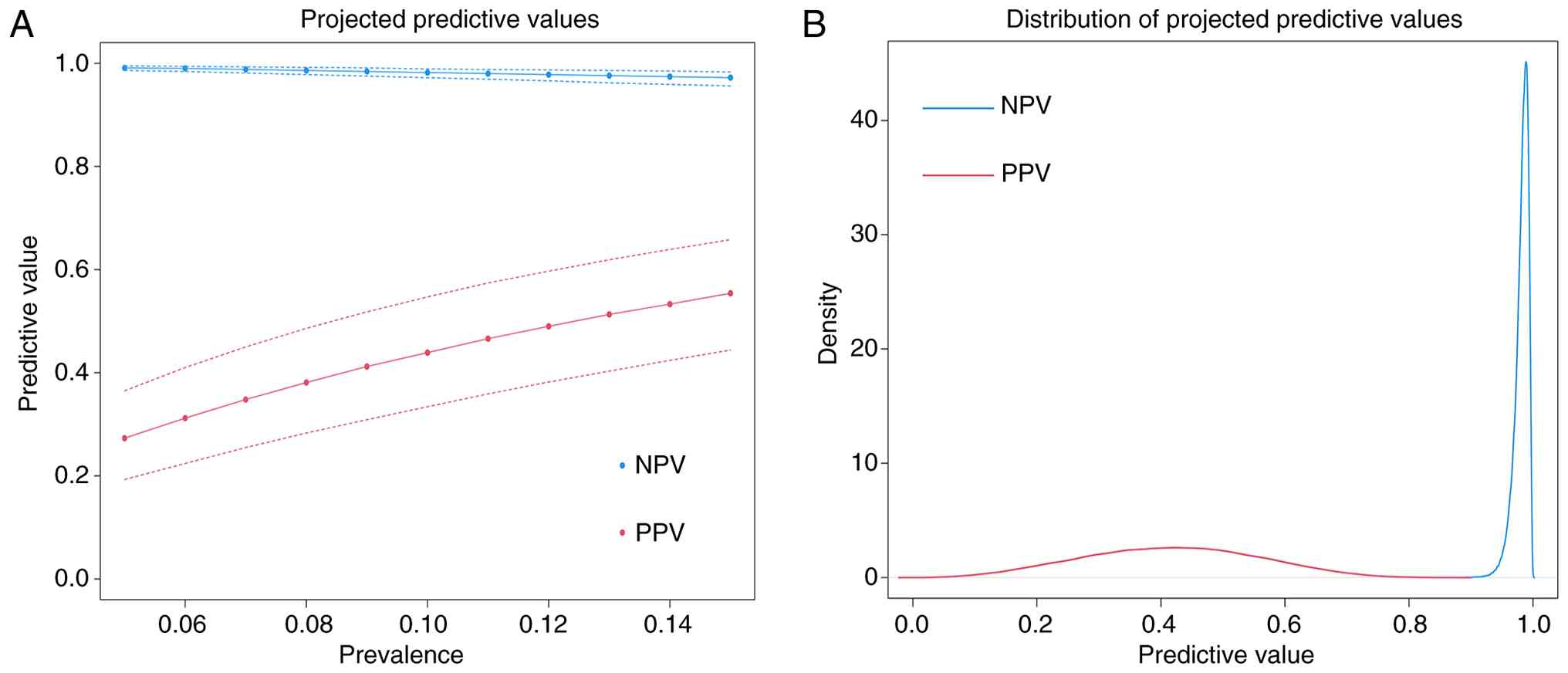
Predictive values for a mean distribution of 10%
(ranging from 5 to 15%) were calculated as follows: An NPV of
0.972–0.991, indicating a very high probability that a negative
test result is truly negative, consistent across studies due to
high specificity, and a PPV of 0.273–0.554, indicating that the
probability of a positive test being truly positive increases with
prevalence. The highest PPVs were observed in the studies by Zhao
et al (31) and Dong et
al (30). The results of the
variation in predictive values across the range of 5–15% prevalence
are presented in Fig. 5. As
observed, the results highlight the stability of NPV in ruling out
bone metastasis.
Discussion
The present systematic review and meta-analysis
evaluated the diagnostic accuracy and predictive performance of
computer-based image-processing techniques for detecting bone
metastases in patients with lung cancer. The bivariate Reitsma
model yielded a pooled sensitivity of 86% and a pooled specificity
of 87.8%, with an AUC of 0.931, supporting the robustness and high
diagnostic accuracy of these models for identifying metastatic
lesions. These findings highlight the importance of integrating
advanced imaging techniques and computer-based processing methods
into the clinical diagnostic workflow, especially where early
decision-making is critical to optimize treatment strategies and
supportive care.
Data synthesis was executed from 6 primary studies
that employed different imaging modalities and machine learning
classifier algorithms, including convolutional neural networks,
radiomics and AI-based methodologies, for identifying bone
metastasis. The pooled diagnostic metrics were high and suggest
that computer-based image processing techniques have advantages
over traditional radiographic interpretation approaches in
improving diagnostic potential for bone metastasis in clinical
decision-making.
The present study also demonstrated some variability
in the performance metric values across different studies, with the
sensitivity ranging from 72.7 to 93.5%. These variabilities can be
attributed to several factors, such as different imaging
modalities, the efficacy of the employed machine learning
algorithms and the basic characteristics of the original studies.
For instance, advanced techniques such as ssDECT and AI-based bone
scintigraphy seem to perform better, possibly owing to the enhanced
quantitative information methodological advances and pipeline for
the study design.
Computer-based processing algorithms are useful for
assisting radiologists in extracting sufficient information from
imaging results and precisely reducing time wastage for diagnostic
approaches (12,35). Beyond the diagnostic accuracy
metrics, the practical aspects of integrating AI-based image
processing in clinical settings is evident, considering several
practical factors. First, the interpretation of the radiological
images is crucial for clinical adaptation, as radiologists and
oncologists need to carefully assess every change in radiological
images, which is a time-consuming process. AI-based algorithms with
the ability to process high-dimensional data can integrate all
imaging features by applying a role-based diagnostic model, boosted
with the exact, feature-extracting algorithms. Thereby, these
techniques can effectively minimize personal mistakes, mitigate
overlapping features and effectively assist radiologists in the
accurate diagnosis and multidisciplinary decision-making process.
Furthermore, cost effectiveness is an essential factor, especially
when advanced modalities are limited. In this context,
computer-based approaches are promising as they have the potential
of an accurate diagnosis, minimize misinterpreting and reducing
unnecessary downstream clinical examinations and management
(36–38).
The idea for integrating imaging-based AI with
clinical biomarkers has been proposed in the literature. For
instance, a study on colorectal cancer showed that the integration
of composite score with inflammation-related (for example,
c-Reactive protein or systemic immune inflammation score) and
nutrition-related (for example, albumin) biomarkers can
significantly enhance prognostic and clinical values in disease
stratification and precise risk categorization (38,39).
In addition, the incorporation of these biomarkers with imaging or
AI information could potentially yield a more powerful diagnostic
model for metastatic outcomes (40). In addition, the recent advances in
multi-omics and metabolic reprogramming studies have established a
framework to improve investigation of the mechanisms of the disease
states (37). These findings
indicate a prospective direction wherein radiomics and deep
learning models for bone metastasis extend beyond imaging data
alone, incorporating multi-omics signatures and metabolic
reprogramming patterns to develop imaging-genomic and
imaging-metabolomic models that more accurately reflect tumor
biology, microenvironmental context and risk of progression.
Over the last decade, the clinical utility of
computational image processing techniques has been studied
extensively. Cao et al (35)
retrospectively explored clinical and imaging results of 273
patients with lung adenocarcinoma to identify predictive risk
factors for bone metastasis. Su et al investigated the
integration of imaging results with the clinical information of
patients (15). Based on the
results, a logistic regression model was developed by integrating
genomic mutations, laboratory results, and imaging data, with an
AUC of 0.91, to validate bone metastasis. Noguchi et al
(12), emphasizing the importance
of early diagnosis of metastatic status, especially in bones,
proposed a deep learning algorithm that could automatically process
the CT images and detect if bone metastasis had occurred with a
sensitivity of 89.8%. This result was strongly consistent with the
sensitivity of 86% in the present study, which was achieved using a
predictive bivariate Reitsma model.
Recent studies have developed various methodological
innovations underscoring the clinical application of multi-panel or
computational aided methodologies for bone metastasis in diverse
diseases and malignancies (36,41–43).
For instance, a deep learning architecture was applied by Crasta
et al (44) to improve the
accuracy of detecting bone lesions. The authors introduced the
integration of the imaging results in a multi-modality approach,
emphasizing the importance of applying combination techniques, such
as AI for diagnostic goals, as highlighted in the present pooled
metrics and meta-analysis. Liu et al (18) proposed the incorporation of radiomic
features combined with clinical parameters to build robust
predictive models, highlighting the methodological advances
discussed in the present study regarding feature-based and
learning-based approaches. Lastly, So et al (14) introduced a novel machine learning
model with improved sensitivity and specificity to predict the risk
of bone metastasis in patients with lung cancer by employing
clinical and radiological variables including T staging,
consumption of EGFR inhibitors, American Joint Committee on Cancer
(AJCC) staging, and presence of lymphovascular invasion. This
evidence supports the results in the present meta-analysis towards
enhanced performance metrics using advanced computer-based image
processing techniques.
The results of the present methodology comparison
among the included studies implied that the studies with advanced
imaging modalities, such as ssDECT in the study by Dong et
al (25) and bone scintigraphy
in the study by Zhao et al (26), were performed more successfully
owing to the advancement instrumental and, possibly, computational
analysis (30,31). A strong negative correlation between
sensitivity and false-positive rate was observed in the present
meta-analysis, suggesting a favorable trade-off for the
high-performing included studies. The other finding in the present
study was the higher value of the diagnostic model, with an AUC of
0.93, and the stable high NPV in all included studies, which is
useful for ruling out non-metastatic cases. Collectively, despite
some inconsistencies driven by the included studies, such as
difference in imaging modalities, study design and population,
these findings highlight the importance of applying advanced image
processing techniques over the traditional approach for the
interpretation of radiographic results.
One of the most relevant findings of the present
study with regard to the clinical perspective is the high NPV
(0.972–9.991) across a realistic prevalence range of 5–15%. Such a
high NPV strongly indicates its reliable application as a screening
tool for ruling out non-metastatic cases. It means that some lung
cancer patients complain of bone pain or other non-specific
symptoms. However, if they are considered ‘low risk’ because of a
negative imaging result, they should be assessed for other bone
problems before using expensive or invasive methods.
The present meta-analysis was written with adherence
to the PRISMA guidelines and utilized the Reitsma model to explore
the realistic diagnostic accuracy of image processing techniques
over traditional image interpretation for bone metastasis. However,
there were some inevitable limitations, which are addressed as
follows: First, each meta-analysis is completely dependent on the
presence of original research articles according to predefined
eligibility criteria. In the present comprehensive search, only 6
studies were included, which may potentially lower the statistical
power of the findings. The heterogeneity in imaging modalities and
also computational processing methodologies were other limitations
of the study. In addition, most of the included studies were
retrospectively designed, and the lack of external validation was
evident in the included studies. However, most of the mentioned
limitations were associated with the primary research, and
considering the importance of the objectives, they were disregarded
in the present study. External validation was overlooked in the
included studies and only a limited number of studies reported
detailed information on providing a precise standard diagnostic
protocol for calibration and decision-making processes in
computer-based processing algorithms. Future research should aim to
perform multi-center, prospective studies to systematically
validate and standardize computer-based image processing techniques
for bone metastasis in patients with lung cancer. Such studies
should establish a standard prospective protocol and evaluation
framework in order to facilitate clinical translation, optimizing
the patients' management.
In conclusion, the present systematic review and
meta-analysis demonstrated the advantages of computational-based
image processing techniques, such as AI, neural networks and
machine learning classifier algorithms, particularly CNN, with high
diagnostic accuracy for bone metastasis in patients with lung
cancer. Despite the limitations and challenges in standardizing
protocols, the findings of the present study support the
integration of the aforementioned advanced techniques into clinical
practice, indicating the potential for better decision-making and
improving the patient outcome.
Supplementary Material
Supporting Data
Supporting Data
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
This study was supported by the Primary Health Development
Research Center of Sichuan Province Program (grant no.
SWFZ24-Z-11), the Chengdu Medical Research Project (grant no.
2025246) and the Key R&D Project of Chengdu Science and
Technology Bureau (grant nos. 2024-YF05-00119-SN and
2024-YF05-00947-SN).
Availability of data and materials
The data presented in this study are available on
request from the corresponding author.
Authors' contributions
YH contributed to the study conceptualization and
design, protocol registration, data acquisition, data analysis and
data interpretation. YH, WL and YE contributed to the study design,
literature search, data collection, formal analysis and data
interpretation. Technical assistance and project administration was
provided by WL and YE. QT was involved in all the study processes,
including conceptualization and study design, funding acquisition,
supervision, and data analysis and interpretation. YH and QT
confirm the authenticity of all the raw data. All authors have read
and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
AIC
|
Akaike information criterion
|
|
AUC SROC
|
area under the summary receiver
operating characteristic curve
|
|
BIC
|
Bayesian information criterion
|
|
CI
|
confidence interval
|
|
CT
|
computed tomography
|
|
CNN
|
convolutional neural network
|
|
Dice
|
Dice similarity coefficient
|
|
DOR
|
diagnostic odds ratio
|
|
IoU
|
intersection over union
|
|
LR
|
likelihood ratio
|
|
NOS
|
Newcastle-Ottawa scale
|
|
NPV
|
negative predictive value
|
|
PPV
|
positive predictive value
|
|
PRISMA
|
Preferred Reporting Items for
Systematic reviews and Meta-Analysis
|
|
QUADAS-2
|
Quality Assessment of Diagnostic
Accuracy Studies-2
|
|
REML
|
restricted maximum likelihood
|
|
ssDECT
|
single-source dual energy CT
|
References
|
1
|
Riihimäki M, Hemminki A, Fallah M, Thomsen
H, Sundquist K, Sundquist J and Hemminki K: Metastatic sites and
survival in lung cancer. Lung Cancer. 86:78–84. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Milovanovic IS, Stjepanovic M and Mitrovic
D: Distribution patterns of the metastases of the lung carcinoma in
relation to histological type of the primary tumor: An autopsy
study. Ann Thorac Med. 12:191–198. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Zhou J, Xu Y, Liu J, Feng L, Yu J and Chen
D: Global burden of lung cancer in 2022 and projections to 2050:
Incidence and mortality estimates from GLOBOCAN. Cancer Epidemiol.
93:1026932024. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Macedo F, Ladeira K, Pinho F, Saraiva N,
Bonito N, Pinto L and Gonçalves F: Bone metastases: An overview.
Oncol Rev. 11:3212017.PubMed/NCBI
|
|
5
|
Jagadeesan S: Predictors of survival in
patients with bone metastasis of lung cancer. Ann Oncol.
28:II532017. View Article : Google Scholar
|
|
6
|
Gong L, Xu L, Yuan Z, Wang Z, Zhao L and
Wang P: Clinical outcome for small cell lung cancer patients with
bone metastases at the time of diagnosis. J Bone Oncol.
19:1002652019. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Duan J, Fang W, Xu H, Wang J, Chen Y, Ding
Y, Dong X, Fan Y, Gao B, Hu J, et al: Chinese expert consensus on
the diagnosis and treatment of bone metastasis in lung cancer (2022
edition). J Natl Cancer Cent. 3:256–265. 2023.PubMed/NCBI
|
|
8
|
Yetiskul E, Salak J, Arafa F, Agarwal A,
Matra A, Niazi M and Odaimi M: Hypercalcemia and bone metastasis in
a case of large cell neuroendocrine carcinoma with unknown primary.
Case Rep Oncol Med. 2024:87922912024.PubMed/NCBI
|
|
9
|
Isaac A, Dalili D, Dalili D and Weber MA:
State-of-the-art imaging for diagnosis of metastatic bone disease.
Radiologe. 60:1–16. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Łukaszewski B, Nazar J, Goch M,
Łukaszewska M, Stępiński A and Jurczyk MU: Diagnostic methods for
detection of bone metastases. Contemp Oncol (Pozn). 21:98–103.
2017.PubMed/NCBI
|
|
11
|
Wu S, Pan Y, Mao Y, Chen Y and He Y:
Current progress and mechanisms of bone metastasis in lung cancer:
A narrative review. Transl Lung Cancer Res. 10:439–451. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Noguchi S, Nishio M, Sakamoto R, Yakami M,
Fujimoto K, Emoto Y, Kubo T, Iizuka Y, Nakagomi K, Miyasa K, et al:
Deep learning-based algorithm improved radiologists' performance in
bone metastases detection on CT. Eur Radiol. 32:7976–7987. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Papalia GF, Brigato P, Sisca L, Maltese G,
Faiella E, Santucci D, Pantano F, Vincenzi B, Tonini G, Papalia R
and Denaro V: Artificial intelligence in detection, management, and
prognosis of bone metastasis: A systematic review. Cancers (Basel).
16:27002024. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
So KWL, Leung EMC, Ng T, Tsui R, Cheung
JPY and Choi SW: Machine learning models to predict bone metastasis
risk in patients with lung cancer. Cancer Med. 13:e703832024.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Su Q, Wang B, Guo J, Nie P and Xu W:
CT-based radiomics and clinical characteristics for predicting bone
metastasis in lung adenocarcinoma patients. Transl Lung Cancer Res.
13:721–732. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li T, Lin Q, Guo Y, Zhao S, Zeng X, Man Z,
Cao Y and Hu Y: Automated detection of skeletal metastasis of lung
cancer with bone scans using convolutional nuclear network. Phys
Med Biol. 67:10.1088/1361–6560/ac4565. 2022. View Article : Google Scholar
|
|
17
|
Fan X, Zhang X, Zhang Z and Jiang Y: Deep
learning on MRI images for diagnosis of lung cancer spinal bone
metastasis. Contrast Media Mol Imaging. 2021:52943792021.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liu Z, Yin R, Ma W, Li Z, Guo Y, Wu H, Lin
Y, Chekhonin VP, Peltzer K, Li H, et al: Bone metastasis prediction
in non-small-cell lung cancer: Primary CT-based radiomics signature
and clinical feature. BMC Med Imaging. 24:2032024. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kim DH, Seo J, Lee JH, Jeon ET, Jeong D,
Chae HD, Lee E, Kang JH, Choi YH, Kim HJ, et al: Automated
detection and segmentation of bone metastases on spine MRI using
U-Net: A multicenter study. Korean J Radiol. 25:363–373. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Parums DV: Editorial: Review articles,
systematic reviews, meta-analysis, and the updated preferred
reporting items for systematic reviews and meta-analyses (PRISMA)
2020 guidelines. Med Sci Monit. 27:e9344752021. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Eriksen MB and Frandsen TF: The impact of
patient, intervention, comparison, outcome (PICO) as a search
strategy tool on literature search quality: A systematic review. J
Med Libr Assoc. 106:420–431. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Carra MC, Romandini P and Romandini M:
Risk of bias evaluation of cross-sectional studies: Adaptation of
the newcastle-ottawa scale. J Periodontal Res. 28:1–10. 2025.
|
|
23
|
Lee J, Mulder F, Leeflang M, Wolff R,
Whiting P and Bossuyt PM: QUAPAS: An adaptation of the QUADAS-2
tool to assess prognostic accuracy studies. Ann Intern Med.
175:1010–1018. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Shim SR, Kim SJ and Lee J: Diagnostic test
accuracy: Application and practice using R software. Epidemiol
Health. 41:e20190072019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Vilca-Alosilla JJ, Candia-Puma MA,
Coronel-Monje K, Goyzueta-Mamani LD, Galdino AS, Machado-de-Ávila
RA, Giunchetti RC, Coelho EA and Chávez-Fumagalli MA: A systematic
review and meta-analysis comparing the diagnostic accuracy tests of
COVID-19. Diagnostics. 13:10.3390/diagnostics13091549. PubMed/NCBI
|
|
26
|
Bellio R and Brazzale AR: Restricted
likelihood inference for generalized linear mixed models.
Statistics and Computing. 21:173–183. 2011. View Article : Google Scholar
|
|
27
|
Wilson C, Harley C and Steels S:
Systematic review and meta-analysis of pre-hospital diagnostic
accuracy studies. Emerg Med J. 35:757–764. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sedgwick P: Meta-analyses: What is
heterogeneity? BMJ. 350:h14352015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Balduzzi S, Rücker G and Schwarzer G: How
to perform a meta-analysis with R: A practical tutorial. Evid Based
Ment Health. 22:153–160. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Dong Y, Zheng S, Machida H, Wang B, Liu A,
Liu Y and Zhang X: Differential diagnosis of osteoblastic
metastases from bone islands in patients with lung cancer by
single-source dual-energy CT: Advantages of spectral CT imaging.
Eur J Radiol. 84:901–907. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhao Z, Pi Y, Jiang L, Xiang Y, Wei J,
Yang P, Zhang W, Zhong X, Zhou K, Li Y, et al: Deep neural network
based artificial intelligence assisted diagnosis of bone
scintigraphy for cancer bone metastasis. Sci Rep. 10:170462020.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Liu Y, Yang P, Pi Y, Jiang L, Zhong X,
Cheng J, Xiang Y, We J, Li L, Yi Z, et al: Automatic identification
of suspicious bone metastatic lesions in bone scintigraphy using
convolutional neural network. BMC Med Imaging. 21:1312021.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Huo T, Xie Y, Fang Y, Wang Z, Liu P, Duan
Y, Zhang J, Wang H, Xue M, Liu S and Ye Z: Deep learning-based
algorithm improves radiologists' performance in lung cancer bone
metastases detection on computed tomography. Front Oncol.
13:11256372023. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang Y, Lin Q, Zhao S, Zeng X, Zheng B,
Cao Y and Man Z: Automated diagnosis of bone metastasis by
classifying bone scintigrams using a self-defined deep learning
model. Curr Med Imaging. 20:e157340562815782024. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Cao Z, Zheng R, Li J, Wang X, Ding C,
Zhang F, Geng J, Wei Z and Fan R: Risk factors of bone metastasis
in lung adenocarcinoma. BMC Pulm Med. 25:2992025. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Ouyang W, Deng Z, Li Y, Chi W, Huang Z,
Zhan C, Li M, Wang D, Li F, Liu Y, et al: Traditional Chinese
medicine in cerebral infarction: Integrative strategies and future
directions. Phytomedicine. 143:1568412025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Chen Y, Bai M, Liu M, Zhang Z, Jiang C, Li
K, Chen Y, Xu Y and Wu L: Metabolic reprogramming in lung cancer:
Hallmarks, mechanisms, and targeted strategies to overcome immune
resistance. Cancer Med. 14:e713172025. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Wang K, Li K, Zhang Z, Zeng X, Wu Z, Zhang
B, Pan Y, Lau LY, Zhao Z and Chen Y: Combined preoperative
platelet-albumin ratio and cancer inflammation prognostic index
predicts prognosis in colorectal cancer: A retrospective study. Sci
Rep. 15:295002025. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Li S, Han H, Yang K, Li X, Ma L, Yang Z
and Zhao YX: Exosome-mediated metabolic reprogramming: Effects on
thyroid cancer progression and tumor microenvironment remodeling.
Mol Cancer. 24:2472025. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Teng X, Han K, Jin W, Ma L, Wei L, Min D,
Chen L and Du Y: Development and validation of an early diagnosis
model for bone metastasis in non-small cell lung cancer based on
serological characteristics of the bone metastasis mechanism.
EClinicalMedicine. 72:2024. View Article : Google Scholar
|
|
41
|
Ouyang W, Lai Z, Huang H and Ling L:
Machine learning-based identification of cuproptosis-related lncRNA
biomarkers in diffuse large B-cell lymphoma. Cell Biol Toxicol.
41:722025. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ouyang W, Zhu C, Li Y, Huang H, Li F and
Ling L: Assessing the neurotoxic risks of triethyl citrate in daily
environmental exposure using network toxicology and molecular
docking. Ecotoxicol Environ Saf. 297:1182252025. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Ouyang W, Huang Z, Wan K, Nie T, Chen H
and Yao H: RNA ac4C modification in cancer: Unraveling
multifaceted roles and promising therapeutic horizons. Cancer Lett.
601:2171592024. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Crasta LJ, Neema R and Pais AR: A novel
deep learning architecture for lung cancer detection and diagnosis
from computed tomography image analysis. Healthcare Analytics.
5:1003162024. View Article : Google Scholar
|