Introduction
Pancreatic ductal adenocarcinoma (PDAC) is one of
the most malignant cancers, with a 5-year overall survival rate of
~10%, and is characterized by extensive cellular heterogeneity and
resistance to treatment (1–6). A number of PDAC cell lines
specifically express epithelial proteins, whereas other cells lines
express mesenchymal proteins (7–9) and
the difference in sphere morphology, proliferative capacity,
migratory capacity and anticancer drug resistance of epithelial and
mesenchymal PDAC cell lines, becomes more apparent when cultured in
three-dimension (3D) (10–14).
PDACs rarely contain squamous cell carcinoma
components that proliferate in sheets and stain positive for
cytokeratin (CK) 5, 6, p63 or p40 upon immunohistochemical
staining. Adenosquamous carcinoma (ASC) is diagnosed when squamous
cell components are found in >30% of pathological tissue
specimen (15). ASC is observed in
<1% of patients with PDAC, but it shows resistance to anticancer
drugs and has a worse prognosis compared with that of PDAC
(16–21). In addition to its aggressive
clinical behavior, pancreatic ASC shares major driver mutations
with conventional PDAC, including frequent tumor protein (TP)-53,
KRAS and CDKN2A alterations (22).
However, these common genetic traits are accompanied by alterations
that affect chromatin regulators and epigenomic control, which
distinguish ASC from typical PDAC. These observations suggest that
ASCs share the same tumor lineage as PDAC but subsequently acquire
additional molecular programs that promote squamous differentiation
and therapeutic resistance.
Squamous cells are not present in the healthy
pancreas, thus the origin of the squamous components in PDAC is
unclear. To date, numerous hypotheses have been proposed regarding
the origin of the squamous epithelium, including differentiation,
squamous metaplasia and collision theories (23–26).
In 3D culture systems, the intercellular interactions and
concentration gradients of oxygen and nutrients enable the
reproduction of an environment closer to the tumor microenvironment
in vivo compared with that achieved through a traditional
two-dimensional (2D) culture (27,28).
However, to the best of our knowledge, the effects of 3D culture on
the expression and spatial distribution of ductal and squamous
epithelial markers in PDAC cells have not been systematically
investigated.
In the present study, the aim was to investigate the
expression profiles of ductal or squamous components in a panel of
epithelial and mesenchymal PDAC cell lines grown in 2D and 3D
cultures using proteomic and immunocytochemical analyses.
Materials and methods
Cell culture
Human PDAC cell lines PK-8 (cat. no. RCB2700),
PK-45P (cat. no. RCB2141), T3M-4 (cat. no. RCB1021) and KP4 (cat.
no. RCB1005) were provided by RIKEN BioResource Research Center
through The National Bio-Resource Project of the Ministry of
Education, Culture, Sports, Science and Technology and The Japan
Agency for Medical Research and Development (29). Further human PDAC cell lines PK-59
(cat. no. TKG0492), PK-1 (cat. no. TKG0239), PANC-1 (cat. no.
TKG0606) and MIA PaCa-2 (cat. no. TKG0227) were obtained from The
Cell Resource Center for Biomedical Research through the Institute
of Development, Aging and Cancer of Tohoku University (Sendai,
Japan). Previous studies describing the establishment of PK-1 and
T3M-4, as well as the other PDAC cell lines used in the present
study, have indicated that they were derived from patients with
histologically confirmed pancreatic adenocarcinoma (30,31).
Cells were cultured in RPMI-1640 medium (Nacalai Tesque, Inc.)
supplemented with 10% FBS (Gibco; Thermo Fisher Scientific, Inc.)
at 37°C in a humidified 5% CO2 atmosphere. Using a
Mycoplasma PCR Detection Kit (cat. no. 25239, iNtRON
Biotechnology) according to the manufacturer's instructions, it was
determined that none of the cells exhibited Mycoplasma
contamination. Genomic DNA was extracted from PDAC cells using the
DNeasy Blood and Tissue Kit (Qiagen), following the manufacturer's
protocol. Short tandem repeats were analyzed using the
GenePrint® 10 System (Promega Corporation), following
the manufacturer's protocol. All PDAC cell lines were correctly
genotyped and showed no evidence of contamination.
Proteomic analysis
For the 2D culture, adherent cells were collected by
trypsinization, centrifuged (170 × g; 5 min; 20°C) and washed with
PBS. This procedure was performed twice. For 3D cultures, PDAC
cells were seeded at 3.0×103 cells/well in
low-attachment 96-well plates (Thermo Fisher Scientific, Inc.) for
7 days. The detailed protocols for 3D sphere formation were
conducted as previously described (12).
Protein extraction
Cell pellets from 2D and 3D cultures were suspended
in 100 µl lysis buffer [5% SDS; 50 mM triethylammonium bicarbonate
(TEAB; Thermo Fisher Scientific, Inc.) buffer; pH 8.5], sonicated
and centrifuged (16,000 × g for 10 min at room temperature). The
supernatants were collected, and protein concentrations were
measured using Pierce 660 nm Protein Assay Reagent (Thermo Fisher
Scientific, Inc.) with Ionic Detergent Compatibility Reagent
(Thermo Fisher Scientific, Inc.).
Trypsin digestion
Supernatants containing 30 µg protein were diluted
with lysis buffer up to 23 µl and digested by trypsin (Promega
Corporation) using a S-Trap™ Micro Spin column (ProtiFi, LLC),
according to the manufacturer's instructions with minor
modifications. Briefly, samples were reduced with 5 mM
tris(2-carboxyethyl)phosphine (Thermo Fisher Scientific, Inc.) at
55°C for 15 min, alkylated with 20 mM iodoacetamide (Thermo Fisher
Scientific, Inc.) at 25°C for 20 min in the dark and digested with
3 µg trypsin (protein/trypsin, 10:1) at 47°C for 2 h. Eluates were
evaporated in vacuo to dryness and dissolved in 100 mM TEAB
buffer (pH 8.5). The sample corresponding to 5 µg protein was
desalted using GL-Tip SDB (GL Sciences Inc.) and dissolved in 5%
acetonitrile (MeCN; FUJIFILM Wako Pure Chemical Corporation)
containing 0.1% formic acid (FA; FUJIFILM Wako Pure Chemical
Corporation) at a concentration of 0.25 µg/µl.
Liquid chromatography-tandem mass
spectrometry analysis
Liquid chromatography-tandem mass spectrometry
analysis of tryptic peptides was performed using the Vanquish Neo
coupled with the Orbitrap Fusion Lumos mass spectrometer (Thermo
Fisher Scientific, Inc.) by injecting 2 µl sample (0.5 µg protein).
The PepMap™ Neo Trap Cartridge (0.3×5.0 mm; 5 µm particle size;
C18; Thermo Fisher Scientific, Inc.) and NANO HPLC Capillary column
(75 µm × 12 cm; 3 µm particle size; C18; Nikkyo Technology, Co.,
Ltd.) were used as the trap and analytical columns, respectively.
Peptide separation was performed using water containing 0.1% FA
(solvent A) and MeCN containing 0.1% FA (solvent B) at a flow rate
of 300 nl/min. The linear gradient for peptide separation was as
follows: 0–1 min, 5% B; 1–91 min, 5–22% B; 91–96 min, 22–40% B;
96–100 min, 40–90% B; and 100–110 min, 90% B. The mass spectrometer
was operated in data-dependent acquisition mode. All mass spectra
(MS1) were acquired with the following settings: Spray voltage, 2.0
kV; ion transfer tube temperature, 275°C; detector type, orbitrap;
resolution, 60,000; mass range, normal; scan range, 350–1,500 m/z;
radiofrequency lens, 30%; automatic gain control (AGC) target,
standard; maximum ion injection time (IT), auto; and polarity,
positive. All MS/MS spectra (MS2) were acquired using the following
settings: Isolation mode, quadrupole; isolation window, 1.6 m/z;
activation type, higher-energy collisional dissociation (HCD);
collision energy mode, fixed; HCD collision energy type,
normalized; HCD collision energy, 30%; detector type, orbitrap;
orbitrap resolution, 15,000; mass range, normal; scan range mode,
auto; AGC target, standard; maximum IT mode, auto; dynamic
exclusion, 20 sec; and charge states, 2–7.
Data analysis
Protein identification and label-free quantification
(LFQ) were performed using the Proteome Discoverer software
(version 3.0; Thermo Fisher Scientific, Inc.). The parameters for
protein identification were as follows: Search engine, CHIMERYS
(version 1.7; MSAID GmbH); fragment mass tolerance, 0.02 Da;
enzyme, trypsin (full); maximum missed cleavage sites, 2; static
modification, carbamidomethyl (Cys, +57.021 Da); dynamic
modification, oxidation (Met, +15.995 Da); and protein database,
Homo sapiens (20,426 proteins downloaded on 09-20-2023 from
UniProtKB; http://www.uniprot.org/uniprotkb). The parameters for
LFQ were as follows: Precursor quantification, area; normalization
mode, total peptide amount. The results of the proteomic analysis
of ductal and squamous cell markers are summarized in Table SI.
Immunocytochemical analysis
Immunocytochemical analysis was performed as
previously described (12,32). Adherent PDAC cells were collected
after trypsin treatment, centrifuged (170 × g, 5 min, 20°C) and
fixed with 10% neutral-buffered formalin for 3 h at 22°C. For
sphere formation in 3D culture, PDAC cells were seeded in a growth
medium consisting of RPMI-1640 medium (Nacalai Tesque) supplemented
with 10% FBS at 3.0×103 cells/well in 96-well
low-attachment plates (cat. no. 174925; Thermo Fisher Scientific,
Inc.). After 7 days, the spheres were aspirated using micropipettes
and fixed in formalin for 3 h at 22°C. Formalin was removed using a
micropipette and both PDAC cells and spheres were dehydrated in
graded ethanol and embedded in paraffin. Subsequently, serial
sections of the cell blocks (3 µm thick) were stained using the
BOND Polymer Refine Detection kit (Leica Biosystems) according to
the manufacturer's instructions.
Mouse monoclonal anti-CK7 (cat. no. 713481;
ready-to-use) and anti-p40 (cat. no. 418171; ready-to-use)
antibodies were purchased from Nichirei Biosciences, Inc. Mouse
monoclonal anti-CK5/6 (cat. no. M7237; 1:1,000), anti-p63 (cat. no.
M7317; 1:50) and anti-CK20 (cat. no. M7019; 1:5,000) antibodies
were purchased from Dako; Agilent Technologies, Inc. Furthermore,
rabbit monoclonal anti-ATP binding cassette subfamily C (ABCC)-1
(cat. no. D5C1X; 1:100), anti-MMP-2 (cat. no. D4M2N; 1:100),
anti-MMP-14 (cat. no. E3S5S; 1:100) and anti-ATP-binding cassette
sub-family G (ABCG)-2 (cat. no. D5V2K; 1:200) antibodies were
purchased from Cell Signaling Technology, Inc. and the rabbit
monoclonal anti-ABCC2 (anti-MRP2; cat. no. EPR10998; 1:1,000)
antibody was purchased from Abcam. For CK7 and CK20, antigen
retrieval was performed using the Bond Enzyme Pretreatment Kit
(cat. no. AR9551; Leica Biosystems) according to the manufacturer's
instructions. For CK5/6, p63, p40, ABCC1, ABCC2, ABCG2, MMP-, and
MMP-14, antigen retrieval was performed using the BOND Epitope
Retrieval Solution 2 (cat. no. AR964; Leica Biosystems) according
to the manufacturer's instructions. Tissue sections were incubated
with each primary antibody for 15 min at 22°C. Antigen detection
was performed using DAB, followed by counterstaining with
hematoxylin for 2 min at 22°C. Negative controls were generated by
omitting the primary antibodies. Images were captured using the
PhenoImager Mantra™ 2 multispectral microscope (Akoya Biosciences,
Inc.).
Pathway analysis
Pathway analysis was performed using KeyMolnet
software (version 6.2; KM Data Inc.) (33,34).
To identify the molecular pathways associated with squamous
metaplasia-related phenotypes, PK-45P was selected as the reference
epithelial PDAC cell line, as immunocytochemical analysis
demonstrated the absence of p40 expression under 2D and 3D culture
conditions. Using PK-45P as the denominator, pathway analyses were
conducted for PK-1 and T3M-4 cells, which exhibited squamous
metaplasia-associated features. Proteomic datasets obtained through
label-free quantitative proteomic analysis were used as input for
pathway analysis. The list of identified proteins with abundance
ratios and P-values was imported into KeyMolnet software and
pathway analysis was performed using the ‘interrelation search’
algorithm to generate molecular interaction networks within a
single path from the input molecules, including direct
activation/inactivation and transcriptional activation/repression.
Pathways associated with the input protein sets were extracted from
the curated KeyMolnet knowledge base.
Results
Proteomic and immunocytochemical
analyses of CK7 in PDAC cells
Proteomic and immunocytochemical analyses were
performed using five PDAC cell lines exhibiting epithelial
properties with high E-cadherin and low vimentin expression (PK-8,
PK-45P, PK-59, PK-1 and T3M-4) and three PDAC cell lines (PANC-1,
KP4 and MIA PaCa-2) exhibiting the opposite expression pattern
(12). PK-8, PK-45P, PK-59, PK-1,
T3M-4, PANC-1, KP4 and MIA PaCa-2 were listed in descending order
of E-cadherin mRNA levels, as previously assessed using
quantitative real-time PCR analysis in our previous study (12) (Fig.
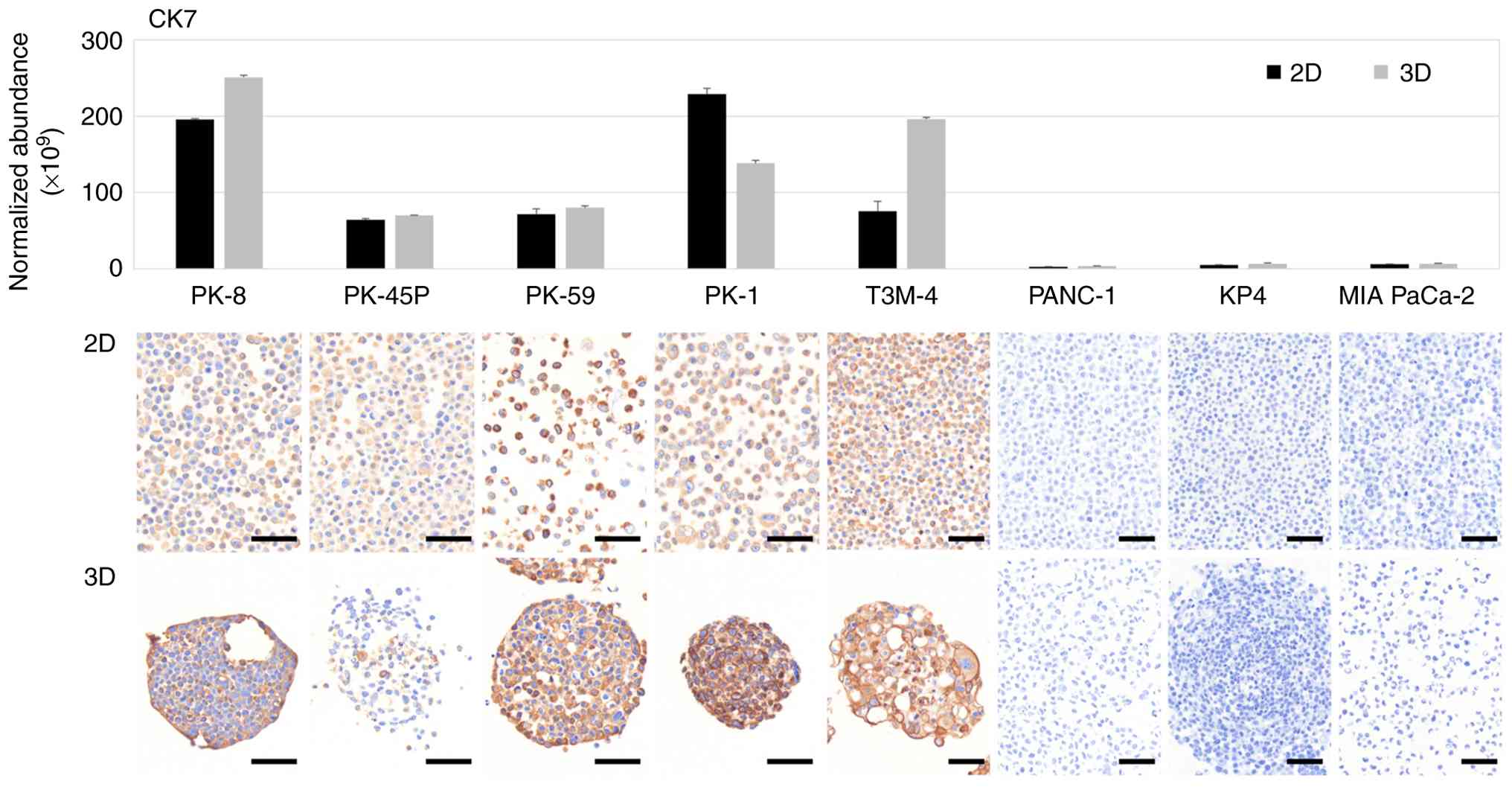
1). Proteomic analysis of PDAC cells cultured in 2D and 3D
showed that CK7, an intermediate filament found in ductal cells and
highly expressed in pancreatic cancer, was detected in five
epithelial PDAC cell lines but was expressed at lower levels in
three mesenchymal PDAC cell lines (Fig.
1, upper panel). To further determine the results of the
proteomic analysis, cell blocks of 2D and 3D cultured PDAC cells
were prepared and stained with a specific antibody against CK7.
Immunocytochemical staining showed that CK7 was positive in all
epithelial PDAC cell lines cultured in 2D and 3D but negative in
mesenchymal PDAC cell lines (Fig.
1, lower panels). By contrast, CK20 was not detected above the
proteomic analysis threshold in any PDAC cell line cultured in 2D
or 3D conditions and was consistently shown to be negative using
immunocytochemistry (Fig. S1).
Proteomic and immunocytochemical
analyses of CK5/6 in PDAC cells
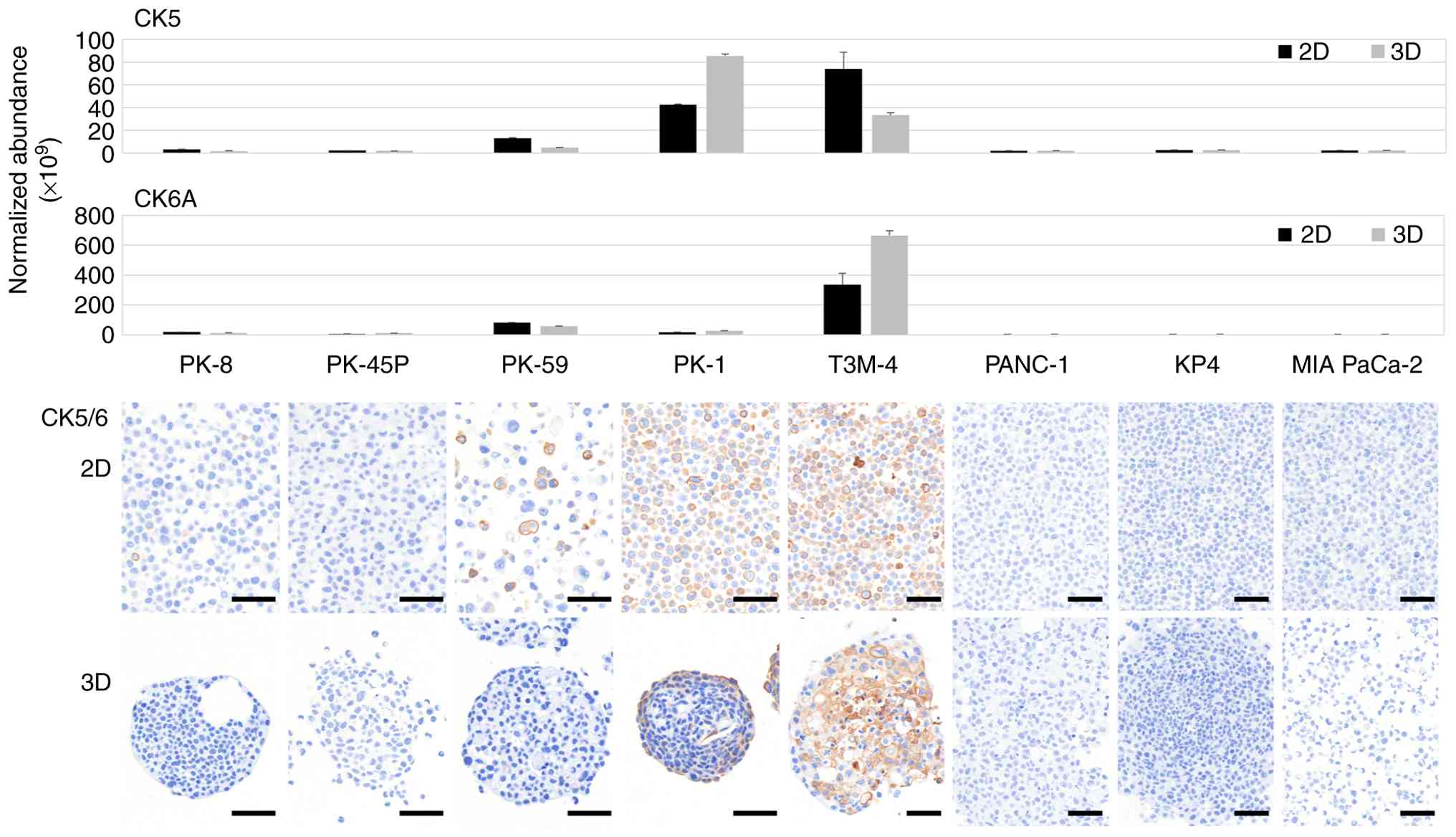
CK5 is a product of the keratin (KRT)-5 gene,
whereas CK6A, CK6B and CK6C are products of KRT6A, KRT6B and KRT6C,
respectively. CK5/6 is not expressed in PDAC cells but is expressed
in squamous carcinomas (35,36).
Proteomic analysis revealed that CK5 was highly expressed in PK-1
and T3M-4 cells, whereas CK6A was highly expressed in T3M-4 cells
only (Fig. 2, upper panels). Low
levels of CK6B and CK6C were detected, with a slight increase
observed in T3M-4 cells (Fig. S2).
No increase in the expression of these CKs was observed in the
mesenchymal PDAC cell lines. Immunocytochemical staining of the
cell blocks showed diffuse CK5/6 localization in the majority of
PDAC cells in 2D cultured PK-1 and T3M-4 cell lines. By contrast,
in 3D cultures, CK5/6 was localized to the periphery of the sphere
of PK-1 cells and the center of the sphere of T3M-4 cells (Fig. 2, lower panels).
Proteomic and immunocytochemical
analyses of p63 in PDAC cells
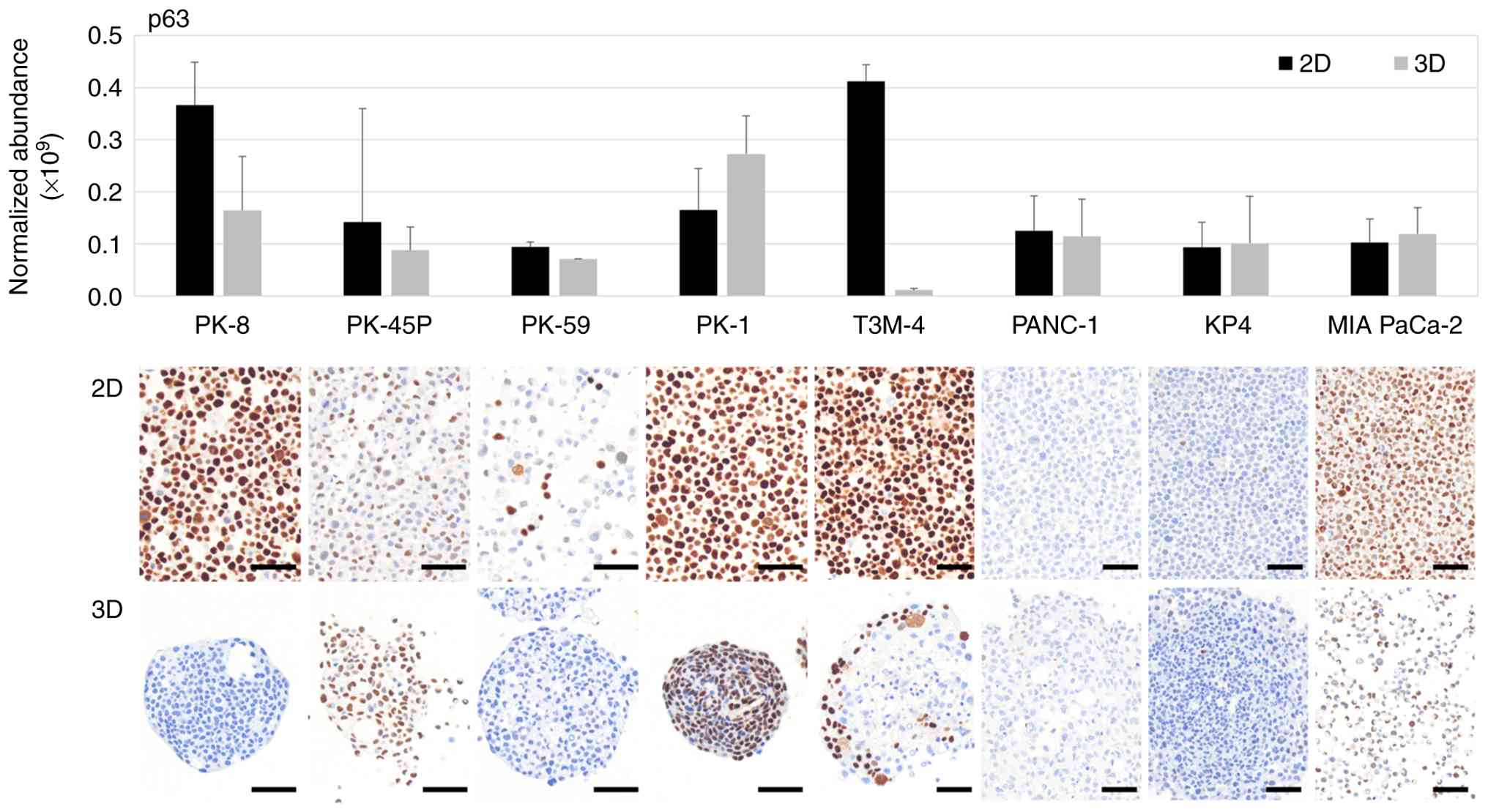
p63 is transcribed from the TP63 gene and p40 is one
of its isoforms generated through alternative promoter usage
(37). Proteomic analysis revealed
that p63 was highly expressed in PK-8 and T3M-4 cells under 2D
culture conditions, whereas relatively higher p63 expression was
observed in PK-1 cells under 3D culture conditions (Fig. 3, upper panel). Notably, due to the
extensive shared peptide sequences between p63 isoforms, liquid
chromatography mass spectrometry-based proteomic analysis does not
discriminate between the transactivating p63 (Tap63) and
dominant-negative (ΔNp63; p40) isoforms. Immunocytochemical
staining showed that p63 was positive in the majority of PK-8 cells
cultured in 2D but was negative when cultured in 3D (Fig. 3, lower panels). Furthermore, p63 was
present in the majority of PK-1 cells in 2D and 3D cultures. In
T3M-4 cells cultured in 3D, only the cells at the periphery of the
spheres were found to be p63-positive. In addition, a small number
of p63-positive cells were found in PK-45P and MIA PaCa-2 cells in
2D and 3D cultures and in PK-59 cells in 2D culture. Although p63
immunoreactivity was occasionally observed in MIA PaCa-2 cells,
these cells were consistently negative for p40 expression and
demonstrated no elevation in p63 levels by proteomic analysis,
indicating the absence of bona fide squamous differentiation.
Immunocytochemical analysis of p40 in
PDAC cells
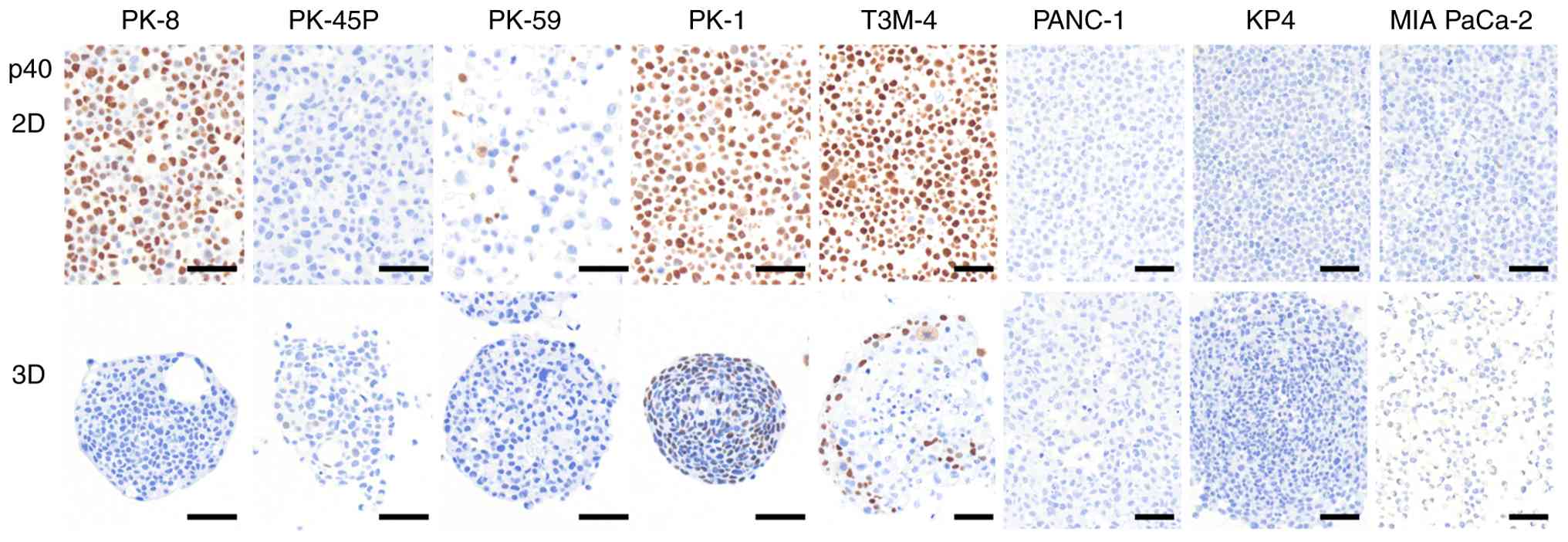
p40 is considered a more specific marker than p63 in
squamous cell carcinoma, however it cannot be detected by proteomic
analysis due to structural overlaps (38). p40 was detected by
immunocytochemical staining in PK-8, PK-1 and T3M-4 cells cultured
in 2D (Fig. 4, upper panel). In 3D
culture, PK-8 cells were p40-negative, whereas PK-1 and T3M-4 cells
located at the periphery of the spheres were p40-positive (Fig. 4, lower panel). Proteomic analysis
detected p63-related signals in PK-8 cells cultured under 3D
conditions; however, p40 expression was not observed by
immunocytochemical analysis under the same conditions. This
discrepancy indicates that proteomic detection of p63 does not
necessarily reflect p40 (ΔNp63) expression in 3D-cultured PK-8
cells. Immunocytochemical analyses of MMP-14 and ABCC1 revealed
positive immunoreactivity predominantly in cells at the periphery
of spheres exhibiting squamous metaplasia, as indicated by p40
positivity (Fig. S3). By contrast,
ABCC2, ABCG2 and MMP-2 showed no detectable immunoreactivity in
either the sphere core or peripheral regions (Fig. S3).
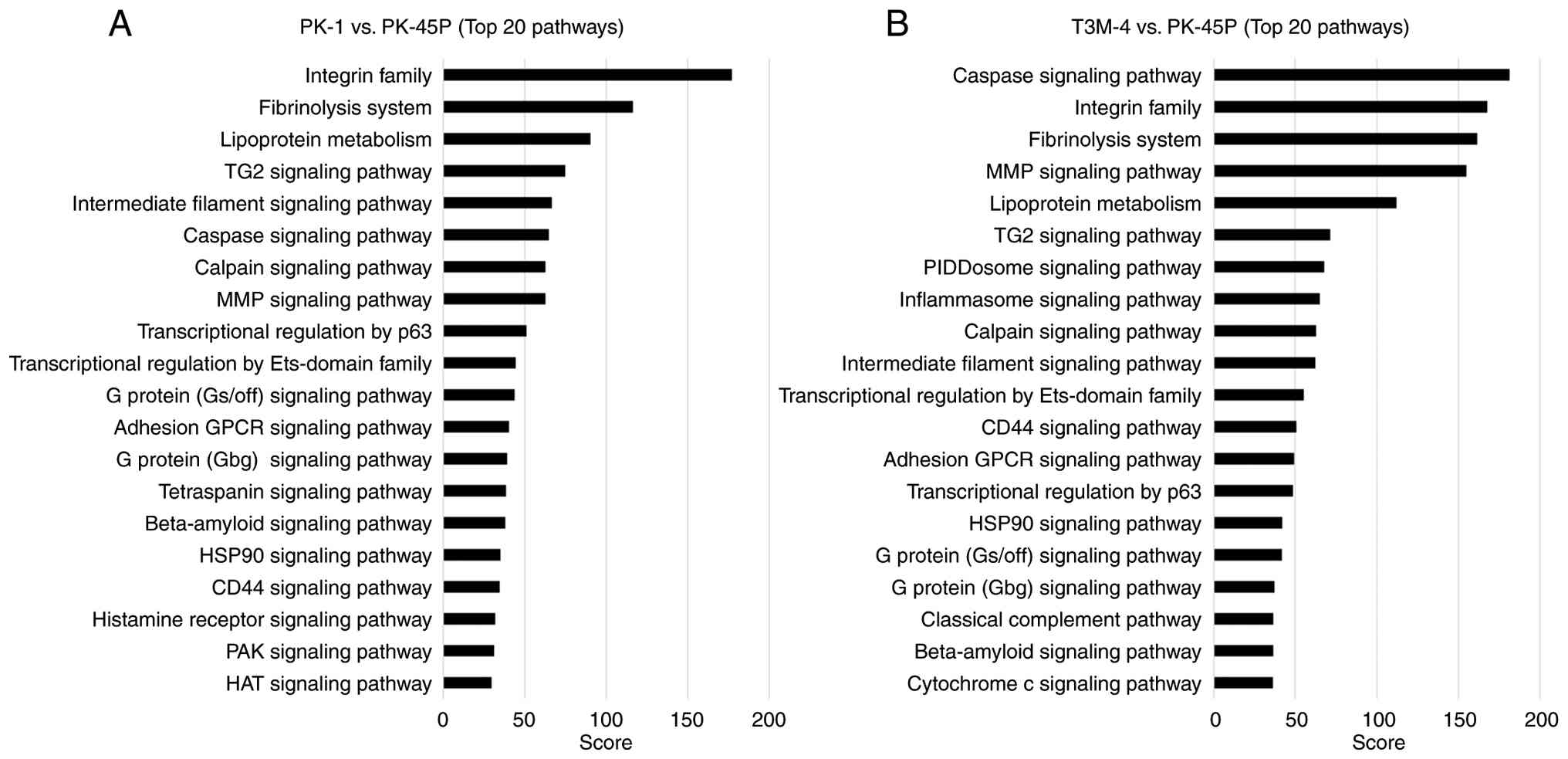
Pathway analysis of squamous
metaplasia-associated proteomic profiles
To explore molecular pathways associated with
squamous metaplasia-associated phenotypes in PDAC cell lines,
pathway analysis was performed using KeyMolnet software based on
proteomic comparisons of PK-1 or T3M-4 cells with the epithelial
reference cell line PK-45P. The top 20 enriched pathways identified
in each comparison are shown in Fig.
5A (PK-1/PK-45P) and Fig. 5B
(T3M-4/PK-45P). In the PK-1/PK-45P comparison (Fig. 5A) pathways related to cell adhesion
and extracellular matrix interactions, including the ‘integrin
family’, ‘fibrinolysis system’ and the ‘MMP signaling pathway’,
showed high enrichment scores. Pathways associated with the
‘intermediate filament signaling pathway’, the transglutaminase 2
‘(TG2) signaling pathway’ and ‘transcriptional regulation by p63’
were also prominently enriched. In the T3M-4/PK-45P comparison
(Fig. 5B), enrichment of pathways
associated with cell adhesion and extracellular matrix remodeling,
such as ‘integrin family’, fibrinolysis system’, ‘MMP signaling
pathway’, ‘TG2 signaling pathway’ and ‘intermediate filament
signaling pathway’, was observed. By contrast with PK-1 cells,
T3M-4 cells exhibited marked enrichment of pathways associated with
‘caspase signaling pathway’, ‘PIDDosome signaling pathway’,
‘inflammasome signaling pathway’, ‘classical complement pathway’
and ‘cytochrome c signaling pathway’. Numerous pathways, including
the ‘integrin family’, ‘fibrinolysis system’, ‘MMP signaling
pathway’, ‘TG2 signaling pathway, ‘intermediate filament signaling
pathway’ and ‘transcriptional regulation by p63’, were commonly
enriched in PK-1 and T3M-4 cells. Collectively, these results
indicate that PK-1 and T3M-4 cells share a core set of molecular
pathways associated with squamous differentiation, while each cell
line exhibits a distinct pathway enrichment profile.
Discussion
Previously, it was reported that CK7, an indicator
of pancreatic ductal differentiation, was expressed in PK-1 cells
(epithelial PDAC cells) but not in PANC-1 cells (mesenchymal PDAC
cells) in both 2D and 3D cultures (11). In the present study, four epithelial
PDAC cell lines, in addition to PK-1 cells, expressed CK7, whereas
two mesenchymal PDAC cell lines and PANC-1 cells did not express
CK7. The majority of PDAC cells are characterized by being
CK7-positive and CK20-negative and are expressed in the epithelial
cells of the digestive tract, such as the large intestine (39). Epithelial PDAC cell lines exhibited
characteristics of typical PDAC cells
(CK7+/CK20−), whereas mesenchymal PDAC cells
were CK7−/CK20−. This highlights the
diversity of PDAC and emphasizes the need for caution in the
pathological diagnosis of PDAC with mesenchymal features.
CK7 expression was restricted only to
epithelial-type PDAC cell lines and was absent in mesenchymal-type
cell lines. By contrast, squamous cell markers were detected only
in a subset of epithelial-type PDAC cell lines, namely PK-1 and
T3M-4, and were absent in mesenchymal-type PDAC cell lines. These
findings indicated that squamous differentiation preferentially
occurred in PDAC cells with a potential for pancreatic ductal
epithelial differentiation, whereas mesenchymal-type PDAC cell
lines lack both ductal and squamous differentiation features,
representing a distinct differentiation state.
Next, the expression of CK5/6, p63 and p40 was
examined, which are commonly used markers of squamous epithelial
components in the pathological diagnosis of PDAC cell lines. CK5/6
expression has been reported in the basal cell types of pancreatic
cancer, pancreatic cancer with squamous cell differentiation and
ASC (40,41). p63 is a transcription factor
belonging to the p53 family and two key (TAp63 and ΔNp63) are
produced from the TP63 gene by an alternative promoter (37). p63 is expressed in squamous
epithelial cells, squamous carcinoma cells, basal cells and cells
with stem cell-like properties (42). ΔNp63, also known as p40 has been
suggested to be more specific for squamous epithelial cells
compared with p63 (43). In the
present study, p63 and p40 were extensively upregulated in the
majority of the epithelial PDAC cell lines, PK-1 and T3M-4, in 2D
culture. By contrast, in 3D culture, p40 expression was observed
only at the periphery of the spheres of these PDAC cells. In the
PDAC cell lines PK-1 and T3M-4, three squamous cell markers were
detected by both proteomics and immunohistochemistry, suggesting
that these cell lines have the ability to produce squamous
components.
Furthermore, no consistent changes in squamous
marker expression were observed between 2D and 3D culture
conditions. Some epithelial PDAC cell lines showed relatively
uniform and high squamous marker expression throughout the cell
population in 2D culture; however, under 3D culture conditions,
squamous marker expression was either overall reduced or became
spatially restricted to cells located at the periphery of the
spheres, rather than being uniformly increased across all cells.
Notably, 3D culture did not result in a global upregulation of
squamous markers compared with 2D culture, but instead altered
their spatial distribution in a cell line-dependent manner. These
observations do not support the notion that 3D culture uniformly
promotes squamous differentiation, but rather indicate that 3D
culture modulates the localization of squamous marker expression
without consistently enhancing squamous differentiation. Therefore,
it cannot be concluded that 3D culture intrinsically induces
squamous differentiation.
In addition, the technical and biological
distinction between p63 and its ΔNp63 (p40) isoform should be
considered. Although the p63 protein was detected using proteomic
analysis, liquid chromatography-mass spectrometry does not
discriminate between individual p63 isoforms due to extensive
shared peptide sequences. By contrast, immunocytochemical analysis
with isoform-specific antibodies revealed a dissociation between
total p63 and ΔNp63 (p40) expression, as exemplified by PK-8 cells
cultured under 3D conditions, which showed p63 detection by
proteomics but lacked p40 by immunocytochemistry. This indicates
that p63 positivity alone does not necessarily reflect activation
of ΔNp63-driven squamous differentiation programs. On the surface
of PK-1 cell spheres, scanning and transmission electron microscopy
has revealed a flattened layer of cells (11,12).
These cells may indicate the morphological differentiation of
squamous epithelial cells. As the expression of squamous components
differs between 2D and 3D cultures, these spatially distinct
expression patterns suggest a role for intercellular interactions
in the regulation of squamous differentiation.
Notably, the localization patterns of CK5/6 under 3D
culture conditions differed between PK-1 and T3M-4 cells, although
the precise mechanisms underlying this difference remain unclear.
However, additional immunocytochemical analyses of proteins in
relation to anticancer drug resistance and extracellular matrix
degradation, revealed distinct spatial associations with CK5/6
localization. These findings suggest that the differential
localization of CK5/6 may reflect distinct biological contexts or
stages of squamous differentiation, potentially corresponding with
structurally stabilized squamous metaplasia in PK-1 cells and a
more stress-associated or dynamic differentiation process in T3M-4
cells. Furthermore, the spatial coincidence of CK5/6 expression
with selected drug resistance- and extracellular matrix-related
proteins indicates that squamous marker-expressing cells may
possess enhanced adaptive properties, particularly at the periphery
of tumor spheres. Consistent with these observations, the
immunoreactivity of MMP-14 and ABCC1 was preferentially localized
to the periphery of p40-positive spheres, whereas other invasion-
and drug resistance-related markers, including ABCC2, ABCG2 and
MMP-2, were not detected. These findings indicate that the
peripheral expression of MMP-14 and ABCC1 does not reflect a
generalized stress response, but rather a selective phenotypic
feature associated with squamous metaplasia and spatial
heterogeneity in 3D PDAC spheres.
Pathway analysis implicated numerous signaling
pathways in epithelial plasticity and squamous differentiation,
which may contribute to this process. Notably, pathway analysis of
proteomic data identified enrichment of transcriptional
regulation-related and histone acetylation-associated pathways,
suggesting the involvement of epigenomic regulation in squamous
differentiation in PDAC. These findings provide a rationale for
future studies to focus on epigenetic regulators, such as
chromatin-modifying enzymes, as potential drivers of lineage
plasticity and squamous transdifferentiation. Pathways associated
with the ‘integrin family’, extracellular matrix remodeling and the
‘intermediate filament signaling pathway’ were commonly enriched,
supporting the notion that cell-cell and cell-matrix interactions
serve a central role in initiating squamous differentiation under
3D culture conditions. In addition, transcriptional regulation by
p63, a marked regulator of squamous epithelial lineage commitment,
was consistently enriched in PDAC cell lines capable of expressing
squamous markers. This supports the notion that squamous
differentiation in PDAC cells is regulated at the transcriptional
level rather than exhibiting a non-specific phenotypic alteration.
The spatial confinement of p40 expression to the sphere periphery
further suggests that local microenvironmental cues, such as
mechanical stress, cell density or paracrine signaling, may
modulate p63-driven transcriptional programs (44). Therefore, no subsequent direct
inhibition experiments were conducted, however a number of the
present results provide mechanistic support for proposing such
experiments. This finding was further supported by the spatially
limited expression of p40 at the sphere periphery under 3D culture
conditions, suggesting localized activation of p63-driven
transcriptional programs. These results indicate that squamous
differentiation in PDAC cells is regulated at the transcriptional
level rather than representing a non-specific phenotypic change and
provide a biologically plausible rationale for future inhibition
studies targeting p63-mediated transcriptional regulation. Future
studies should therefore aim to investigate the inhibition of
candidate proteins involved in integrin signaling, cytoskeletal
remodeling or p63-mediated transcription to potentially regulate
squamous differentiation in PDAC.
Numerous signaling pathways previously implicated in
squamous differentiation, including NOTCH, TGF-β and EGFR
signaling, are modulated by cell-cell interactions and 3D tissue
organization (45–48). Although direct inhibition or
functional pathway analyses were not performed in the present
study, the enrichment of pathways associated with intercellular
interaction, transcriptional regulation and cytoskeletal remodeling
is consistent with the involvement of these regulatory axes in
squamous differentiation under 3D culture conditions. In addition,
enrichment of histone acetylation-related pathways suggests that
epigenetic regulation may contribute to the observed phenotypic
plasticity, as chromatin remodeling is implicated in lineage
switching and squamous transdifferentiation in numerous carcinomas
(49,50).
Notably, the differential expression and spatial
localization of squamous markers under 2D and 3D culture conditions
could have direct clinical implications for patients with PDAC.
Specifically, the acquisition or regional enrichment of squamous
markers, such as CK5/6 and p40, in 3D cultures, may reflect the
phenotypic plasticity associated with tumor aggressiveness,
therapeutic resistance and poor prognosis, which are characteristic
features of ASC (51). These
findings suggest that conventional 2D cultures may underestimate
squamous differentiation, whereas 3D culture systems may exhibit
improved mimicking of the in vivo tumor microenvironment
relevant to diagnosis and drug response. The expression of squamous
markers by specific PDAC cell lines in 3D culture demonstrates the
utility of this in vitro model for investigating the
biological behavior of ASC and testing therapeutic strategies
targeting squamous-like PDAC phenotypes. The present study suggests
that a number of the squamous components observed in PDAC tissues
and ASCs were formed by the differentiation of PDAC cells into
cancer cells with squamous components. The regulatory mechanisms of
squamous cell component expression in PDAC cell lines, particularly
with regard to epigenetic regulation, require further
investigation. Immunocytochemical staining of p63 alone showed
positive results in both 2D- and 3D-cultured MIA PaCa-2 cells,
however p40 staining was negative and no increase was demonstrated
by proteomics. Therefore, it is unlikely that squamous epithelial
components are expressed in MIA PaCa-2 cells.
The present study has several limitations. Although
eight PDAC cell lines, including five epithelial-type and three
mesenchymal-type lines, were analyzed, this panel may not fully
represent the heterogeneity of squamous metaplasia observed in
PDAC. In addition, the analyses were largely descriptive and no
formal statistical comparisons were performed, warranting cautious
interpretation of the observed differences. Furthermore,
immunocytochemical staining was evaluated qualitatively without
quantitative assessment of staining intensity or positive cell
ratios, which limits direct comparison across cell lines and
culture conditions. Future studies incorporating a larger panel of
cell lines, statistical analyses, and quantitative image-based
evaluation will be required to strengthen the robustness and
generalizability of the findings.
In conclusion, the present study demonstrated that
3D culture conditions markedly affected both the expression levels
and spatial distribution of squamous cell markers in PDAC cell
lines. This suggests that squamous differentiation in PDAC is
regulated by extracellular signals from outside the cells. Yet,
additional studies are required to further elucidate the regulation
of the expression of squamous cell components in PDAC cells.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
The authors would like to thank Mr. Mamoru Nomura
(Kiko Tech Co., Ltd.) for their technical support with the imaging
of the immunocytochemical staining.
Funding
The present study was supported by The Japan Society for the
Promotion of Science KAKENHI program (grant nos. 24K11881,
22K08835, 25K12018 and 22K08882; Grant-in-Aid for Scientific
Research C).
Availability of data and materials
The data generated in the present study may be found
in the jPOST repository under accession number JPST004363 or at the
following URL: https://repository.jpostdb.org/entry/JPST004363.
Authors' contributions
YS, HT and TI designed the present study and drafted
the manuscript. YS and MF conducted the cell culture experiments.
HT and YM performed the proteomic analysis and contributed to data
analysis and interpretation. YH performed the immunocytochemical
analysis and contributed to the interpretation of the
immunocytochemical data. KN, SS, HR, KT and TA contributed to data
acquisition, analysis and interpretation, and provided critical
methodological input and intellectual contributions to the
experimental design and data interpretation. All authors reviewed
and edited the manuscript. All authors read and approved the final
version of the manuscript. YS and HT confirm the authenticity of
all the raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Kratzer TB, Wagle NS, Sung H
and Jemal A: Cancer statistics, 2026. CA Cancer J Clin.
76:e700432026.PubMed/NCBI
|
|
2
|
Ushio J, Kanno A, Ikeda E, Ando K, Nagai
H, Miwata T, Kawasaki Y, Tada Y, Yokoyama K, Numao N, et al:
Pancreatic ductal adenocarcinoma: Epidemiology and risk factors.
Diagnostics (Basel). 11:5622021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Blackford AL, Canto MI, Dbouk M, Hruban
RH, Katona BW, Chak A, Brand RE, Syngal S, Farrell J, Kastrinos F,
et al: Pancreatic cancer surveillance and survival of High-Risk
individuals. JAMA Oncol. 10:1087–1096. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yasinzai AQK, Tareen B, Tracy K, Jamil N,
Khan M, Ullah H, Raza M, Khan AU, Arif D, Waheed A, et al:
Pancreatic ductal adenocarcinoma: Exploring clinicopathological
trends and racial disparities in a comprehensive U.S.
population-based study. Clin Transl Oncol. 26:2618–2628. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Latenstein AEJ, van der Geest LGM, Bonsing
BA, Groot Koerkamp B, Haj Mohammad N, de Hingh IHJT, de Meijer VE,
Molenaar IQ, van Santvoort HC, van Tienhoven G, et al: Nationwide
trends in incidence, treatment and survival of pancreatic ductal
adenocarcinoma. Eur J Cancer. 125:83–93. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lei S, Mao Y, Yang Q, Yan H and Wang J:
Trends in pancreatic cancer incidence, prevalence, and survival
outcomes by histological subtypes: A retrospective cohort study.
Gastroenterol Rep (Oxf). 13:goaf0302025. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Collisson EA, Sadanandam A, Olson P, Gibb
WJ, Truitt M, Gu S, Cooc J, Weinkle J, Kim GE, Jakkula L, et al:
Subtypes of pancreatic ductal adenocarcinoma and their differing
responses to therapy. Nat Med. 17:500–503. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kong R, Qian X and Ying W: Pancreatic
cancer cells spectral library by DIA-MS and the phenotype analysis
of gemcitabine sensitivity. Sci Data. 9:2832022. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Schmidtlein PM, Volz C, Braun R, Thürling
I, Lapshyna O, Wellner UF, Konukiewitz B, Lehnert H, Marquardt JU
and Ungefroren H: A comparative endocrine trans-differentiation
approach to pancreatic ductal adenocarcinoma cells with different
EMT phenotypes identifies quasi-mesenchymal tumor cells as those
with highest plasticity. Cancers (Basel). 13:46632021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Shichi Y, Gomi F, Sasaki N, Nonaka K, Arai
T and Ishiwata T: Epithelial and mesenchymal features of pancreatic
ductal adenocarcinoma cell lines in Two- and Three-dimensional
cultures. J Pers Med. 12:7462022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Shichi Y, Sasaki N, Michishita M, Hasegawa
F, Matsuda Y, Arai T, Gomi F, Aida J, Takubo K, Toyoda M, et al:
Enhanced morphological and functional differences of pancreatic
cancer with epithelial or mesenchymal characteristics in 3D
culture. Sci Rep. 9:108712019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Minami F, Sasaki N, Shichi Y, Gomi F,
Michishita M, Ohkusu-Tsukada K, Toyoda M, Takahashi K and Ishiwata
T: Morphofunctional analysis of human pancreatic cancer cell lines
in 2- and 3-dimensional cultures. Sci Rep. 11:67752021. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Shichi Y, Fujiwara M, Gomi F, Nonaka K,
Hasegawa F, Shinji S, Rokutan H, Arai T, Takahashi K and Ishiwata
T: Transmission electron microscopic analysis of pancreatic ductal
adenocarcinoma cell spheres formed in 3D cultures. Med Mol Morphol.
58:298–306. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Shichi Y, Gomi F, Hasegawa Y, Nonaka K,
Shinji S, Takahashi K and Ishiwata T: Artificial intelligence-based
analysis of time-lapse images of sphere formation and process of
plate adhesion and spread of pancreatic cancer cells. Front Cell
Dev Biol. 11:12907532023. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Gill AJ, Lam AK and Washington MK: Tumours
of the Pancreas. Digestive System Tumours WHO classification of
tumours series. 5th edition. Arends MJ FM and Klimstra DS: Lyon:
pp. 296–372. 2019
|
|
16
|
Boyd CA, Benarroch-Gampel J, Sheffield KM,
Cooksley CD and Riall TS: 415 patients with adenosquamous carcinoma
of the pancreas: A population-based analysis of prognosis and
survival. J Surg Res. 174:12–19. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kaiser J, Hinz U, Mayer P, Hank T, Niesen
W, Hackert T, Gaida MM, Büchler MW and Strobel O: Clinical
presentation and prognosis of adenosquamous carcinoma of the
pancreas-Matched-pair analysis with pancreatic ductal
adenocarcinoma. Eur J Surg Oncol. 47:1734–1741. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Moslim MA, Lefton MD, Ross EA, Mackrides N
and Reddy SS: Clinical and histological basis of adenosquamous
carcinoma of the pancreas: A 30-year experience. J Surg Res.
259:350–356. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wild AT, Dholakia AS, Fan KY, Kumar R,
Moningi S, Rosati LM, Laheru DA, Zheng L, De Jesus-Acosta A,
Ellsworth SG, et al: Efficacy of platinum chemotherapy agents in
the adjuvant setting for adenosquamous carcinoma of the pancreas. J
Gastrointest Oncol. 6:115–125. 2015.PubMed/NCBI
|
|
20
|
Regi P, Butturini G, Malleo G, Pedica F,
D'Onofrio M and Bassi C: Clinicopathological features of
adenosquamous pancreatic cancer. Langenbecks Arch Surg.
396:217–222. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Katz MH, Taylor TH, Al-Refaie WB, Hanna
MH, Imagawa DK, Anton-Culver H and Zell JA: Adenosquamous versus
adenocarcinoma of the pancreas: A population-based outcomes
analysis. J Gastrointest Surg. 15:165–174. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Lenkiewicz E, Malasi S, Hogenson TL,
Flores LF, Barham W, Phillips WJ, Roesler AS, Chambers KR,
Rajbhandari N, Hayashi A, et al: Genomic and epigenomic landscaping
defines new therapeutic targets for adenosquamous carcinoma of the
pancreas. Cancer Res. 80:4324–4334. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Borazanci E, Millis SZ, Korn R, Han H,
Whatcott CJ, Gatalica Z, Barrett MT, Cridebring D and Von Hoff DD:
Adenosquamous carcinoma of the pancreas: Molecular characterization
of 23 patients along with a literature review. World J Gastrointest
Oncol. 7:132–140. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kardon DE, Thompson LD, Przygodzki RM and
Veffess CS: Adenosquamous carcinoma of the pancreas: A
clinicopathologic series of 25 cases. Mod Pathol. 14:443–451. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Madura JA, Jarman BT, Doherty MG, Yum MN
and Howard TJ: Adenosquamous carcinoma of the pancreas. Arch Surg.
134:599–603. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Motojima K, Tomioka T, Kohara N, Tsunoda T
and Kanematsu T: Immunohistochemical characteristics of
adenosquamous carcinoma of the pancreas. J Surg Oncol. 49:58–62.
1992. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Han SJ, Kwon S and Kim KS: Challenges of
applying multicellular tumor spheroids in preclinical phase. Cancer
Cell Int. 21:1522021. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Costa EC, Moreira AF, de Melo-Diogo D,
Gaspar VM, Carvalho MP and Correia IJ: 3D tumor spheroids: An
overview on the tools and techniques used for their analysis.
Biotechnol Adv. 34:1427–1441. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kurata N, Satoh H, Kitano H, Nagato Y,
Endo T, Sato K, Akashi R, Ezura H, Kusaba M, Kobayashi M, et al:
NBRP, National Bioresource Project of Japan and plant bioresource
management. Breeding Sci. 60:461–468. 2010. View Article : Google Scholar
|
|
30
|
Kobari M, Matsuno S, Sato T, Kan M and
Tachibana T: Establishment of a human pancreatic cancer cell line
and detection of pancreatic cancer associated antigen. Tohoku J Exp
Med. 143:33–46. 1984. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Okabe T, Yamaguchi N and Ohsawa N:
Establishment and characterization of a carcinoembryonic antigen
(CEA)-producing cell line from a human carcinoma of the exocrine
pancreas. Cancer. 51:662–668. 1983. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Shichi Y, Gomi F, Ueda Y, Nonaka K,
Hasegawa F, Hasegawa Y, Hinata N, Yoshimura H, Yamamoto M,
Takahashi K, et al: Multiple cystic sphere formation from PK-8
cells in three-dimensional culture. Biochem Biophys Rep.
32:1013392022.PubMed/NCBI
|
|
33
|
Miura Y, Hayakawa A, Kikuchi S, Tsumoto H,
Umezawa K, Chiba Y, Soejima Y, Sawabe M, Fukui K, Akimoto Y, et al:
Fumarate accumulation involved in renal diabetic fibrosis in
Goto-Kakizaki rats. Arch Biochem Biophys. 678:1081672019.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Sato M, Tsumoto H, Toba A, Soejima Y, Arai
T, Harada K, Miura Y and Sawabe M: Proteome analysis demonstrates
involvement of endoplasmic reticulum stress response in human
myocardium with subclinical left ventricular diastolic dysfunction.
Geriatr Gerontol Int. 21:577–583. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Boecker W, Tiemann K, Boecker J, Toma M,
Muders MH, Löning T, Buchwalow I, Oldhafer KJ, Neumann U,
Feyerabend B, et al: Cellular organization and histogenesis of
adenosquamous carcinoma of the pancreas: Evidence supporting the
squamous metaplasia concept. Histochem Cell Biol. 154:97–105. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Volkel C, De Wispelaere N, Weidemann S,
Gorbokon N, Lennartz M, Luebke AM, Hube-Magg C, Kluth M, Fraune C,
Möller K, et al: Cytokeratin 5 and cytokeratin 6 expressions are
unconnected in normal and cancerous tissues and have separate
diagnostic implications. Virchows Arch. 480:433–447. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Amelio I, Grespi F,
Annicchiarico-Petruzzelli M and Melino G: p63 the guardian of human
reproduction. Cell Cycle. 11:4545–4551. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Alomari AK, Glusac EJ and McNiff JM: p40
is a more specific marker than p63 for cutaneous poorly
differentiated squamous cell carcinoma. J Cutan Pathol. 41:839–845.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Hamidov Z, Altendorf-Hofmann A, Chen Y,
Settmacher U, Petersen I and Knosel T: Reduced expression of
desmocollin 2 is an independent prognostic biomarker for shorter
patients survival in pancreatic ductal adenocarcinoma. J Clin
Pathol. 64:990–994. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Raper SE: One more piece of the puzzle?
Adenosquamous carcinoma in a pancreatic mass. Gastroenterol Hepatol
(NY). 7:630–632. 2011.PubMed/NCBI
|
|
41
|
Shibayama T, Hayashi A, Toki M, Kitahama
K, Ho YJ, Kato K, Yamada T, Kawamoto S, Kambayashi K, Ochiai K, et
al: Combination immunohistochemistry for CK5/6, p63, GATA6, and
HNF4a predicts clinical outcome in treatment-naive pancreatic
ductal adenocarcinoma. Sci Rep. 14:155982024. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Steurer S, Riemann C, Buscheck F, Luebke
AM, Kluth M, Hube-Magg C, Hinsch A, Höflmayer D, Weidemann S,
Fraune C, et al: p63 expression in human tumors and normal tissues:
A tissue microarray study on 10,200 tumors. Biomark Res. 9:72021.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Bishop JA, Teruya-Feldstein J, Westra WH,
Pelosi G, Travis WD and Rekhtman N: p40 (DeltaNp63) is superior to
p63 for the diagnosis of pulmonary squamous cell carcinoma. Mod
Pathol. 25:405–415. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Barton CE, Johnson KN, Mays DM, Boehnke K,
Shyr Y, Boukamp P and Pietenpol JA: Novel p63 target genes involved
in paracrine signaling and keratinocyte differentiation. Cell Death
Dis. 1:e742010. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Shi Q, Xue C, Zeng Y, Yuan X, Chu Q, Jiang
S, Wang J, Zhang Y, Zhu D and Li L: Notch signaling pathway in
cancer: From mechanistic insights to targeted therapies. Signal
Transduct Target Ther. 9:1282024. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Pan L, Lemieux ME, Thomas T, Rogers JM,
Lipper CH, Lee W, Johnson C, Sholl LM, South AP, Marto JA, et al:
IER5, a DNA damage response gene, is required for Notch-mediated
induction of squamous cell differentiation. Elife. 9:e580812020.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Oshimori N, Oristian D and Fuchs E: TGF-β
promotes heterogeneity and drug resistance in squamous cell
carcinoma. Cell. 160:963–976. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zheng C, Wang J, Wang J, Zhang Q and Liang
T: Cell of origin of pancreatic cancer: Novel findings and current
understanding. Pancreas. 53:e288–e297. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Dawson MA and Kouzarides T: Cancer
epigenetics: From mechanism to therapy. Cell. 150:12–27. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Boumahdi S and de Sauvage FJ: The great
escape: Tumour cell plasticity in resistance to targeted therapy.
Nat Rev Drug Discov. 19:39–56. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Simone CG, Zuluaga Toro T, Chan E, Feely
MM, Trevino JG and George TJ Jr: Characteristics and outcomes of
adenosquamous carcinoma of the pancreas. Gastrointest Cancer Res.
6:75–79. 2013.PubMed/NCBI
|