Introduction
Diabetes mellitus (DM) is a chronic metabolic
disorder characterized by elevated blood glucose (hyperglycemia),
insulin (hyperinsulinemia) and lipids (hyperlipidemia), and it
represents a growing global health challenge (1). According to the International Diabetes
Federation, in 2024, >589 million individuals aged 20–79 years
were diagnosed with DM, and this number is projected to increase to
853 million by 2050, predominantly affecting populations in low-
and middle-income countries (2,3). DM is
classified into different types, including type 1 DM (T1DM), which
is primarily driven by genetic defects leading to autoimmune
destructions of pancreatic insulin-producing cells (4). Type 2 DM (T2DM) arises from
dysregulated protein, carbohydrate and lipid metabolism due to
insulin resistance, impaired insulin secretion or a combination of
both (5). Mainly influenced by
environmental factors such as diet, lack of exercise and aging,
T2DM is notably more prevalent when compared with gestational
diabetes or T1DM, accounting for 90–95% of all cases of diabetes
(3). The economic and clinical
burden of DM is substantial, as patients frequently experience
complications, including vascular malformations, cardiovascular
diseases, neurodegenerative disorders and various types of cancer,
despite treatment with antidiabetic agents such as metformin, a
glucagon-like peptide-1 receptor agonist (5–9). These
observations underscore the urgent need for early screening and the
development of therapeutic strategies for patients with DM who have
an increased risk of developing cancer, often diagnosed at an
advanced and clinically challenging stage due to drug resistance
(10–12).
Numerous epidemiological studies indicate that T2DM
is associated with an increased risk for several types of cancer,
particularly breast cancer (BC), including early-stage
non-metastatic BC and a 20–30% higher risk of invasive BC compared
with non-diabetic individuals (13–16).
BC is the most frequently diagnosed cancer in women worldwide, with
>2.3 million new cases and 665,000 mortalities reported in 2022
(17). The impact of T2DM on BC,
particularly invasive BC, has been estimated to increase both
incidence and mortality by ~20%, primarily due to poor glycemic
control, delayed diagnosis, advanced disease stage, poor prognosis
and limited efficacy of systemic anticancer treatments such as
chemotherapy and immunotherapy (18–20).
Furthermore, specific cancer therapies induce insulin resistance in
patients with BC, contributing to T2DM development and underscoring
shared and overlapping pathophysiological mechanisms between the
two conditions (21,22). Advanced glycation end products
(AGEs) abundantly generated through non-enzymatic glycation under
hyperglycemic conditions, and insulin-like growth factors (IGFs),
overproduced during hyperinsulinemia, contribute to tumorigenesis
through multiple mechanisms (23,24).
In addition to inducing oxidative stress and inflammation, both
AGEs and IGFs activate oncogenic signaling pathways, including
RAS/MAPK, PI3K/Akt/mTOR (PAM), NF-κB and JAK/STAT pathways, through
their respective receptors, receptor for AGEs (RAGE) and IGF
receptors, thereby stimulating tumor development and progression
via metastasis (25,26).
BC stem-like cells (BCSCs) have been identified in
invasive ductal carcinoma (IDC), which originates in milk ducts and
accounts for 80% of invasive cases of BC (27,28).
Generally, cancer stem-like cells (CSCs) are characterized by their
capacity for self-renewal, differentiation, motility, metastasis
and resistance to anticancer therapies, traits that contribute to
tumor aggressiveness and progression (29,30).
In various BC subtypes, these small cell populations, representing
~2% of tumor tissue, express a range of stemness markers, such as
CD44+/CD24−, high aldehyde dehydrogenase
(ALDH) activity, octamer-binding transcription factor (OCT)3/4 and
vimentin expression (31,32).
In the present study, a naturally immortalized cell
line, KAIMRC1, derived from IDC tissue obtained from a Saudi female
patient established by Ali et al (33) was used. KAIMRC1 was characterized as
a stem-like based on the expression of canonical stem cell markers,
including CD44+/CD24−, and its tumorigenicity
was confirmed through spheroid formation in soft agar (33). Hyperglycemia promotes CSC generation
by creating a tumor environment that supports its expansion,
tumorigenicity and aggressiveness (34,35).
Clinical observations have reported elevated glycated albumin (GA)
levels in blood collected from individuals with DM (36,37).
The pro-oncogenic effects of GA on triple-negative BC (TNBC) and
estrogen receptor-positive BC cell lines have been previously
reported (38,39); however, the biological impact of GA
on the stemness characteristics of IDC-derived CSCs remains poorly
understood. Therefore, in the present study, the tumorigenic
response of the IDC-derived BCSC line KAIMRC1 to GA at the cellular
and molecular levels was examined, including cell surface and
signaling proteins, to identify potential biomarkers for the
precise detection of IDC-derived stem-like cells and the
development of tailored therapeutic strategies. Bioinformatics
analyses were conducted to determine the expression levels of
stemness-associated proteins of interest in patients with invasive
BC and DM and to evaluate their impact on overall survival.
Materials and methods
Reagents
Mouse primary monoclonal antibodies directed against
extracellular signal-regulated kinase (ERK)1 (clone G-8; cat. no.
sc-271269), phosphorylated-ERK1/2 (p-ERK1/2; clone E-4; Tyr204 of
ERK1; cat. no. sc-7383), OCT3/4 (clone A-9; cat. no. sc-365509) and
RAGE (clone E-1; cat. no. sc-74473) were obtained from Santa Cruz
Biotechnology, Inc. Mouse monoclonal antibodies targeting vimentin
(V9; cat. no. 790-2917) and glyceraldehyde 3-phosphate
dehydrogenase (GAPDH; clone 6C5; cat. no. ab8245) were sourced from
Roche Diagnostics GmbH and Abcam, respectively. Mouse monoclonal
anti-RAGE antibody (clone 02; cat. no. MA5-29007) and isotype
control immunoglobulin (IgG; cat. no. 31235) were purchased from
Invitrogen, Thermo Fisher Scientific, Inc. The secondary
antibodies, IRDye 680RD (red)-conjugated goat anti-rabbit (cat. no.
926-68071) and IRDye 800RD (green)-conjugated goat anti-mouse (cat.
no. 926-32210) were provided by LI-COR Biosciences. All other
reagents were obtained from Thermo Fisher Scientific, Inc. unless
otherwise specified.
Preparation of GA and unglycated
albumin
GA was generated as previously described (38–40).
Briefly, bovine serum albumin (BSA; fraction V; 10 mg/ml) was
incubated with 0.1 M methylglyoxal (MG) in 0.1 M sodium phosphate
buffer (pH 7.4) containing 3 mM sodium azide for 72 h at 37°C to
induce glycation. Unglycated albumin was processed using the same
procedure but without MG. Free MG (unbound sugar) was removed by
dialysis against distilled water, using a Slide-A-lyzer®
mini dialysis device with a molecular weight cutoff of 3.5 kDa
(cat. no. 69550). Endotoxin levels were negligible after
purification with Detoxi-gel endotoxin-removing gel columns (cat.
no. 20344) and assessment with the E-TOXATE kit (cat. no. ET0200;
MilliporeSigma), according to the manufacturers' instructions. As
shown in Fig. S1, GA formation was
characterized and confirmed by the increase in fluorescence
intensity measured using a SpectraMax M5 fluorescence
spectrophotometer (Molecular Devices, LLC). Protein concentrations
were determined using the Qubit™ protein assay (cat. no. Q33211) on
the Invitrogen Qubit 3 fluorometer. To ensure the stability of the
GA and unglycated BSA solutions, aliquots were prepared and stored
at −20°C until use.
Culture and treatment of IDC-derived
KAIMRC1 stem-like cells and TNBC cell line MDA-MB-231
The IDC-derived KAIMRC1 stem-like cell line
(Resource Identification Initiative: CVCL_RW19), classified as
ER+/HER2− subtype exhibiting stemness
features, was cultured in a complete medium consisting of advanced
Dulbecco's modified eagle (DMEM) supplemented with 10%
heat-inactivated fetal bovine serum (FBS), 2 mM glutamine and
antibiotics (100 IU/ml penicillin; 100 µg/ml streptomycin)
(33). Cultures were maintained at
37°C in a humidified incubator containing 5% CO2. The
TNBC cell line MDA-MB-231 (cat. no. HTB-26, American Type Culture
Collection) was maintained in complete DMEM supplemented with the
aforementioned components. At confluence, MDA-MB-231 cells were
passaged every 2–3 days using enzymatic digestion with TrypLE™
Express Enzyme solution, whereas KAIMRC1 stem-like cells were
easily detached from the culture flasks. Both cell lines were
subcultured at a ratio of 1:2 or 1:3. For treatment, cells were
exposed to GA and unglycated BSA at concentrations of 25, 50, 100
and 200 µg/ml in low-serum medium (LSM), consisting of low glucose
(1 g/l) DMEM supplemented with 2.5% FBS, for designated incubation
periods. Untreated cells served as controls.
ALDH activity assay
The nicotinamide adenine dinucleotide
(NAD)-dependent ALDH activity colorimetric assay (cat. no.
ab155893; Abcam) was performed according to the manufacturer's
instructions. Briefly, a reduced NAD (NADH) standard curve was
generated by preparing 0, 2, 4, 6, 8 and 10 nmol/well NADH
standards corresponding to 0, 2, 4, 6, 8 and 10 µl NADH solutions
dispensed in duplicate into a 96-well plate. The final volume for
each standard was adjusted to 50 µl/well using an ALDH assay
buffer. Untreated cells (1×106) were homogenized in 200
µl ice-cold assay buffer for 10 min and centrifuged at 12,000 × g
for 5 min at 4°C and 50 µl of the resulting supernatant was
transferred to a 96-well plate and mixed with 50 µl assay buffer.
Subsequently, 50 µl reaction mix was added to each well, and the
plate was incubated at room temperature (RT) for 5 min. Optical
density was recorded at 450 nm for the samples and their
corresponding backgrounds controls at the initial time points
(A1 and A1B), and again after 60 min
(A2 and A2B). ALDH activity (mU/ml) was
calculated using the following equation:
ALDH
activity=B/{[(A2-A2B)-(A1-A1B)]
× V × dilution factor} where B, represents the amount of NADH
generated, and V represents the sample volume.
Monitoring real-time cell
proliferation, directional motility and invasion using the
xCELLigence system
The tumorigenic effects of GA on real-time cell
proliferation, directional motility and invasion were assessed
using the xCELLigence Real-time Cell Analyzer Dual Purpose
(RTCA-DP) system (Agilent Technologies, Inc.) following the
manufacturer's instructions. This label-free, impedance-based
platform continuously monitors cellular adhesion, growth and
motility by measuring changes in electrical impedance across plates
incorporating microelectrodes, thus providing a dynamic and
quantitative assessment of cell behavior. For cell proliferation,
16-well E-plates were used (cat. no. 0-060-0890; Agilent
Technologies, Inc.) and for cell invasion and motility, 16-well
cellular invasion/motility (CIM)-plates (cat. no. 05-665-817-001;
Agilent Technologies, Inc.) were used for cell proliferation and
motility/invasion, respectively. For the cell proliferation assay,
7,500 cells were seeded per well in the E-plate, which was then
inserted into RTCA-DP station. A total of 6 h after seeding the
TNBC MDA-MB-231 cells and 24 h after seeding the IDC-derived
KAIMRC1 stem-like cells, 100 µl complete medium was replaced with
100 ml LSM containing different concentrations (25, 50, 100 and 200
µg/ml) GA and unglycated BSA. The E-plate was then reinserted into
the RTCA-DP station. For the cell motility assay, the CIM-16 plate,
consisting of an upper and lower chamber, was prepared by hydrating
the upper membrane with serum-free medium and filling the lower
chamber membrane with LSM containing effective concentrations 50
and 100 µg/ml GA, which served as a chemoattractant. After the
prescribed incubation period and before cell seeding, the CIM-16
plates were placed on the RTCA-DP station, and a baseline impedance
measurement was obtained using a cell-free medium. Cells (15,000)
were then seeded into each well of the upper chamber. The CIM-16
plates were subsequently returned to the RTCA-DP station, and
directional cell motility was quantified based on the electrical
impedance changes detected as cells migrated through the membrane.
The same experimental approach was applied to assess cell invasion,
defined as a motility across a reconstituted basement membrane,
using the xCELLigence system and CIM-16 plate, except that the
upper chamber was first coated with Geltrex™ LDEV-free,
reduced-growth factor basement membrane matrix (cat. no. A1413202).
Data acquisition and cell index calculations were performed using
RTCA software 1.2.1 (cat. no. 30-060-0890; Agilent Technologies,
Inc.), and cell index curves were monitored every 15 min for 72–80
h for cell proliferation and 24 h to assess cell motility and
invasion.
Colony formation assays
Soft agar colony formation assay for IDC-derived
KAIMRC1 stem-like cells was performed in 24-well plates, each well
containing two layers of agarose (cat. no. 121853; Merck KGaA) in
complete medium, as previously described (33). The bottom layer contained 0.6%
agarose, while the top layer contained 0.35% agarose. To assess
colony formation, IDC-derived KAIMRC1 stem-like cells (20,000/well)
were seeded into the top agarose layer. After 2–3 days of
incubation at 37°C, the complete medium was replaced with LSM
containing GA at concentrations 50 and 100 µg/ml for 72 h at 37°C.
Following treatment, the cells were incubated for 2–3 weeks, and
fresh LSM containing GA (50 and 100 µg/ml) was added twice weekly.
Colonies were fixed with 4% paraformaldehyde for 5 min at RT and
visualized under an inverted ZEISS microscope, and colonies
containing >50 cells were counted manually.
For the TNBC cell line MDA-MB-231, cells were
harvested and resuspended in serum-free cell culture medium to a
final concentration of 1×106 cells/ml. An appropriate
number of cells (3,000–5,000 per well) were seeded into 24-well
plates. After 24 h, the culture medium was replaced with LSM
containing GA at concentrations of 50 and 100 µg/ml. Following
treatment, the plates were incubated at 37°C in a humidified 5%
CO2 incubator for 1 week, until untreated control cells
formed colonies of substantial size (>50 cells per colony).
Cells were fixed with 4% paraformaldehyde for 5 min at RT. After
removing the fixative, 1 ml of 0.5% crystal violet solution was
added to each well and incubated for 2 h at RT. Excess stain was
carefully removed by rinsing the plates in distilled water, and the
plates were air-dried. Colonies were imaged using a ZEISS
microscope, colonies containing >50 cells were counted (41).
Preparation of cell lysates and
western blot analysis
IDC-derived KAIMRC1 stem-like cells and TNBC
MDA-MB-231 cells (3×105 cells) were seeded in complete
medium in 12-well plates and incubated for 24 h at 37°C. The medium
was subsequently replaced with LSM and incubated for an additional
24 h at 37°C. Cells were treated with GA and unglycated BSA at
concentrations of 25, 50, 100 and 200 µg/ml for the indicated
incubation periods (10 min and 48 h). After washing in ice-cold
phosphate-buffered saline, proteins were extracted by lysing the
cells with NP40 lysis buffer. Cell debris were removed by
centrifugation at 20,000 × g for 30 min at 4°C. Protein
concentration for each sample was determined using the Qubit
Protein Assay, and total protein amount was adjusted to 100 or 200
µg for equal protein loading. Protein samples were mixed with an
equal volume of 4X Laemmli sample buffer in 1.5 ml
Eppendorf® tubes, denatured by boiling at 95°C for 15
min and then briefly centrifuged at 1,000 × g for 7 sec at RT.
Samples were resolved on 11% SDS-PAGE gels alongside prestained
molecular weight markers. Proteins were then electroblotted onto
PVDF membranes, which were blocked for 1 h at RT in Tris-buffered
saline (TBS)-0.1% Tween (pH 7.4) containing 1% BSA. PVDF membranes
were incubated overnight at 4°C on a rotating shaker with mouse or
rabbit primary antibodies diluted in the blocking buffer, targeting
p-ERK1 (1:500 dilution), total ERK1 (1:500 dilution), OCT3/4 (1:500
dilution), RAGE (1:1,000 dilution), vimentin (1:1,000 dilution) and
the housekeeping protein GAPDH (1:5,000 dilution). Following five
washes for 10 min each in TBS-0.1% Tween at RT, membranes were
incubated for 1 h at RT with infrared fluorescent secondary
antibodies diluted (1:5,000 dilution) in LI-COR Odyssey®
blocking buffer: IRDye 680RD (red)-conjugated goat anti-rabbit or
IRDye 800RD (green)-conjugated goat anti-mouse (LI-COR Biosciences)
with continuous mixing. After washing, protein bands were
visualized using the LI-COR Odyssey CLx Scanner, and protein
expression levels were quantified using ImageJ software version
1.53e (https://imagej.net/ij/index.html) as previously
described (39,42).
Bioinformatics
The MammOnc-DB database (http://resource.path.uab.edu/MammOnc-Home.html;
accessed, 19 September 2025) was used to analyze proteins of
interest (43). Gene expression
profiles of the genes encoding RAGE (AGER), vimentin
(VIM) and OCT4 (POU5F1), were stratified by biopsy
type and TNBC status, as well as their associations with 5-year
survival in patients with BC, retrieved from the Swedish Cancerome
Analysis Network-Breast (SCAN-B) within MammOnc-DB database
(43,44). Additional gene expression profiles
for these genes, categorized according to three-gene classifier
subtypes (HER2+, ER+HER2− and
ER−HER2−), were obtained from the Molecular
Taxonomy of Breast Cancer International Consortium (METABRIC)
dataset available through MammOnc-DB (43,45).
Further analyses were conducted using the cBioPortal database
(https://www.cbioportal.org/; accessed,
19 September 2025) (46–48). The dataset ‘Breast Cancer (METABRIC,
Nature 2012 & Nat Commun 2016)’ within the ‘breast’ section was
applied to assess gene expression profiles of the analyzed genes
based on prediction analysis of microarray 50 (PAM50) and claudin
subtype classifications (45,49,50).
Patients were classified into low- and high-expression groups using
the median mRNA expression values (Illumina HT-12 v3 microarray;
Illumina, Inc.) for each gene. Corresponding clinical data were
downloaded from the database.
Data from Panigrahi et al (51) was retrieved from the Gene Expression
Omnibus data repository (GSE202922; accessed on 23 June 2025).
Processed RNA sequencing count data were normalized using counts
per million (CPM) after trimmed mean of M-values (TMM)
normalization (CPM-TMM) method (51).
RAGE neutralization
RAGE was neutralized in the cells by treating them
with an anti-RAGE antibody to determine whether GA exerts its
effects through this receptor. IDC-derived KAIMRC1 stem-like and
TNBC MDA-MB-231 cells (3×105 cells) were seeded in
complete media as previously described51 a40). After medium
replacement with LSM, 20 µg/ml mouse monoclonal anti-RAGE antibody
(clone 02; cat. no. MA5-29007) or 20 µg/ml isotype control IgG
(cat. no. 31235) was added. Following a 2 h incubation at 37°C, the
cells were stimulated with 100 µg/ml GA for 48 h, which represented
the most effective concentration used in earlier experiments of the
present study. Protein lysates were then collected for western blot
analysis.
Statistical analysis
All data are presented as the mean ± SD from three
independent experiments. Statistical comparisons between the two
groups were performed using an unpaired two-tailed student's
t-test. For analysis involving more than two groups, one-way ANOVA
statistical test followed by Tukey's post hoc test was used to
assess statistical differences among multiple comparisons.
Statistical outputs for bioinformatics analyses were derived from
the SCAN-B and METABRIC datasets within the MammOnc-DB database, as
well as the ‘Breast Cancer (METABRIC, Nature 2012 & Nat Commun
2016)’ dataset accessed through the cBioPortal database. Pairwise
comparisons for datasets generated by Panigrahi et al
(51) were evaluated using the
Mann-Whitney U non-parametric test. P<0.05 was considered to
indicate a statistically significant difference.
Results
IDC-derived KAIMRC1 stem-like cells
exhibit higher ALDH activity when compared with TNBC MDA-MB-231
cells
High ALDH activity is widely recognized as a
valuable biomarker for CSCs, particularly BCSCs, because it
supports their stemness and self-renewal properties (52). Therefore, ALDH activity was first
assessed in IDC-derived KAIMRC1 and TNBC MDA-MB-231 cells to
identify the cell line with greater stem-like properties. The
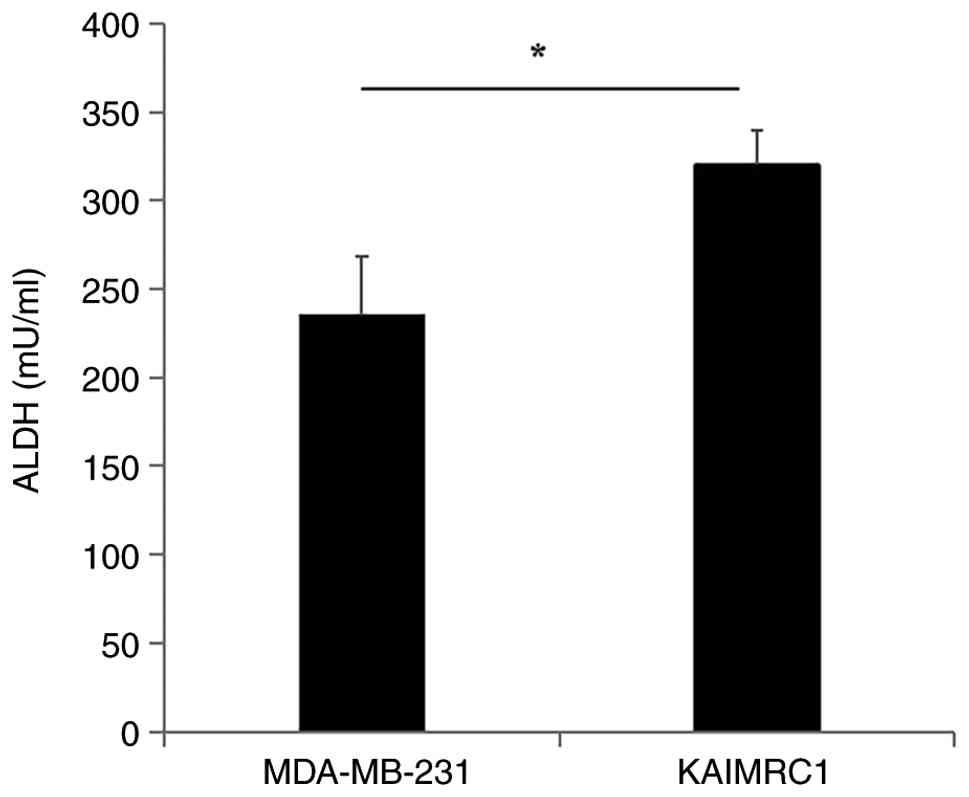
KAIMRC1 cell line showed significantly higher ALDH enzyme activity
(1.36-fold; P=0.019) compared with the MDA-MB-231 cell line
(Fig. 1).
GA weakly increases the cell
proliferation rate of IDC-derived KAIMRC1 stem-like cells, unlike
its effect on TNBC MDA-MB-231 cells
Cell proliferation was monitored using the
xCELLigence RTCA-DP system to assess the mitogenic response of
stem-like cells. Before examining KAIMRC stem-like cells, the
effects of various GA concentrations (25–200 µg/ml) and the
expected lack of effect of unglycated BSA on the proliferation of
TNBC MDA-MB-231 cells (Fig. S2)
were verified and confirmed, consistent with findings using the
trypan blue exclusion method (38).
Thus, using the real-time system on MDA-MB-231 cells, unglycated
BSA did not alter cell proliferation at any concentrations tested,
except for a significant reduction at 200 µg/ml (0.83-fold; P=0.02;
Fig. S2A). Conversely, GA
stimulated the cell proliferation rate in a dose-dependent manner
that followed a bell-shaped curve characterized by increased
proliferation at intermediate doses (1.28-fold at 50 µg/ml;
P=0.00009; 1.59-fold at 100 µg/ml; P=0.00024; Fig. S2B), followed by a decreased
proliferation at the highest GA dose (0.79-fold at 200 µg/ml;
P=0.019; Fig. S2B). This pattern
is consistent with previously described activation kinetics of
dimeric receptors such as RAGE (38). After validating the biological
effect of GA, KAIMRC1 stem-like cells were treated with
intermediate doses of 50 and 100 µg/ml of GA and unglycated BSA.
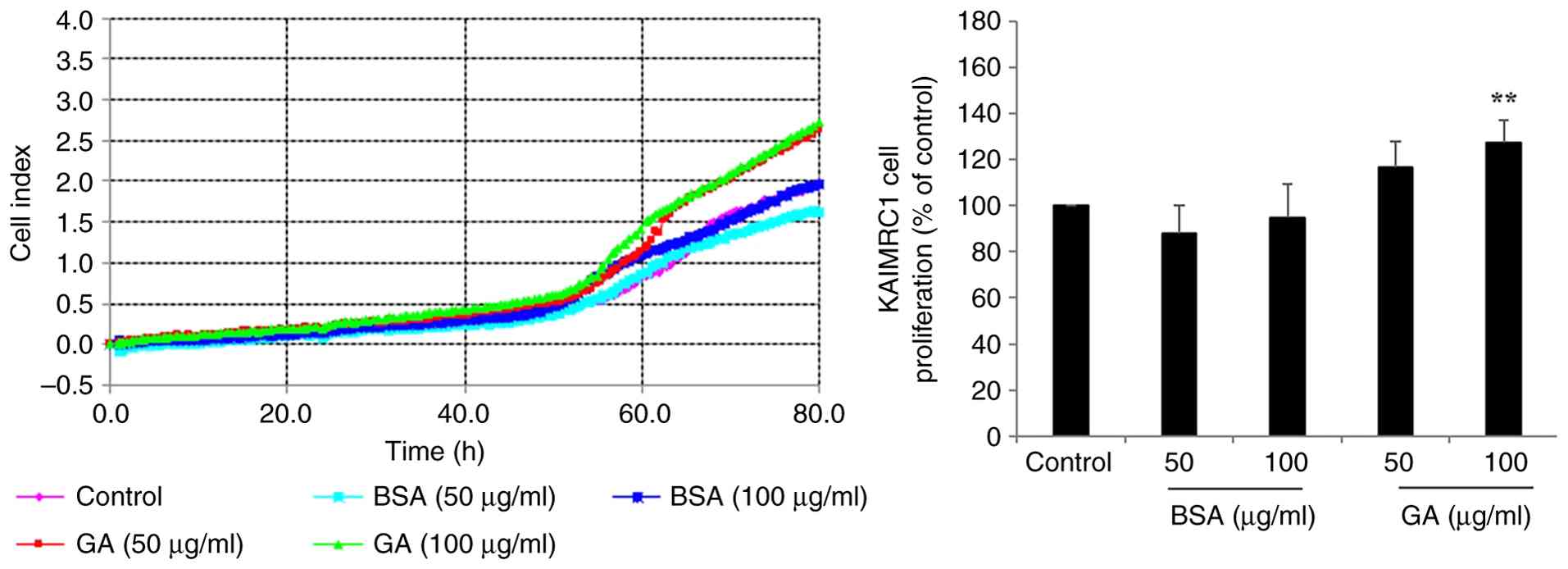
Similar to the findings in MDA-MB-231 cells, 100 mg/ml GA
significantly increased cell proliferation of KAIMRC1 stem-like
cells (1.27-fold; P=0.0076; Fig.
2), although the onset of the KAIMRC1 cell response was
observed after 60 h of incubation (Fig.
2) when compared with MDA-MB-231 cell response observed after
10 h of incubation (Fig. S2B).
Unglycated BSA did not affect the cell proliferation of KAIMRC1
stem-like cells compared with untreated controls (Fig. 2).
GA has an increased stimulatory effect
on the invasiveness of IDC-derived KAIMRC1 stem-like cells when
compared with TNBC MDA-MB-231 cells
Cell motility, which drives migration, invasion and
metastasis, is a key process in cancer progression (7). To evaluate directional motility and
basement membrane invasion, the automated xCELLigence RTCA-DP
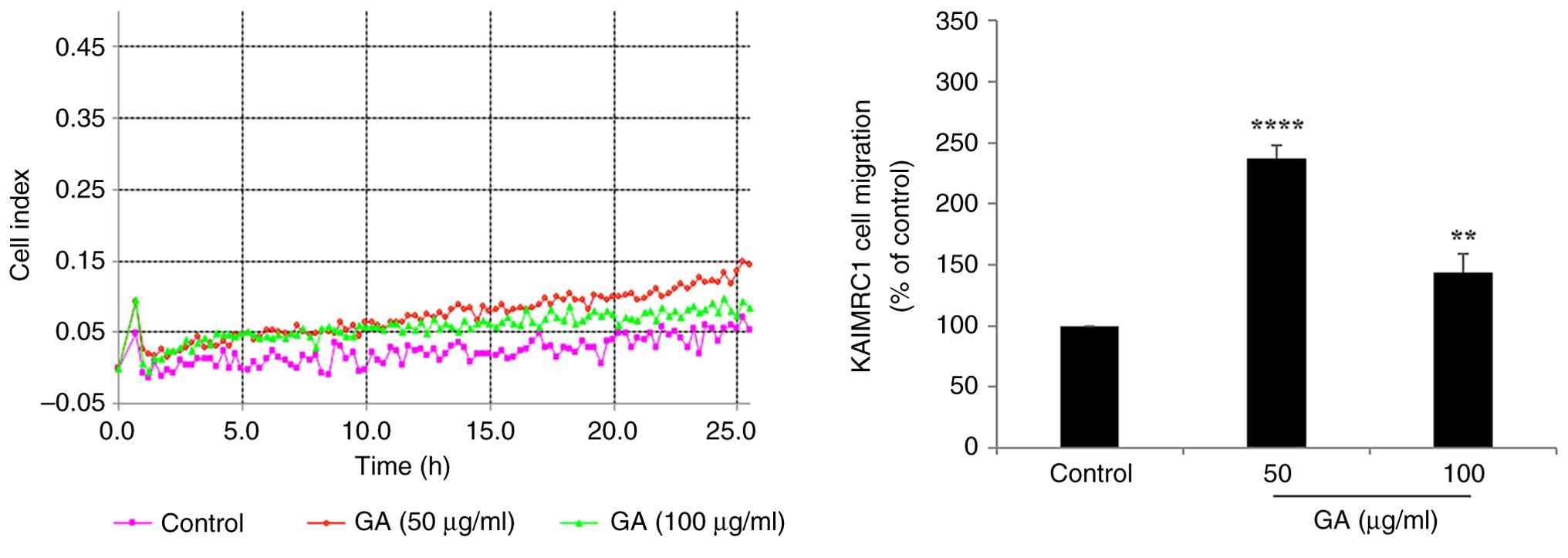
system was employed. In IDC-derived KAIMRC1 stem-like cells,
treatment with effective GA concentrations, at 50 and 100 µg/ml,
significantly enhanced motility capacity, with a 2.37-fold
(P=0.00003) and 1.44-fold (P=0.0096) increase, respectively,
compared with untreated controls (Fig.
3). TNBC MDA-MB-231 cells exhibited a similar response, showing
a 2.52-fold (P=0.0096) and 1.6-fold, (P=0.03) increase at the same
GA concentrations, respectively (Fig.
S3). When motility was assessed through a protein-based matrix
to determine invasive capacity, GA significantly enhanced invasion
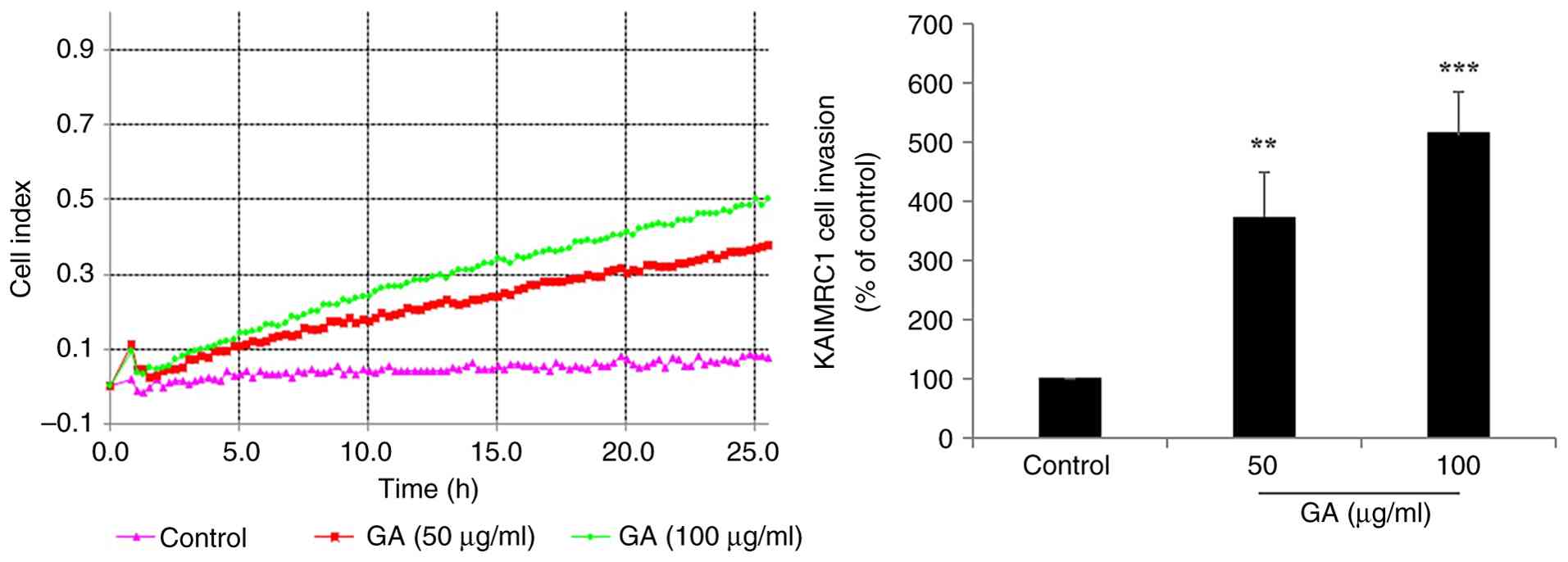
in both cell lines, though to differing extents. IDC-derived
KAIMRC1 stem-like cells exhibited a marked increase in invasiveness
upon GA treatment, with 3.71-fold (P=0.004) and 5.14-fold
(P=0.0005) enhancements at 50 and 100 µg/ml GA, respectively
(Fig. 4). Conversely, TNBC
MDA-MB-231 cells showed a significant increase in invasion, with
1.42-fold (P=0.009) and 1.47-fold (P=0.04) enhancements at the
effective (50–100 µg/ml) GA concentrations compared with the
untreated controls, respectively (Fig.
S4). These findings indicate that IDC-derived KAIMRC1 stem-like
cells are markedly more responsive to GA-induced invasiveness when
compared with MDA-MB-231 cells.
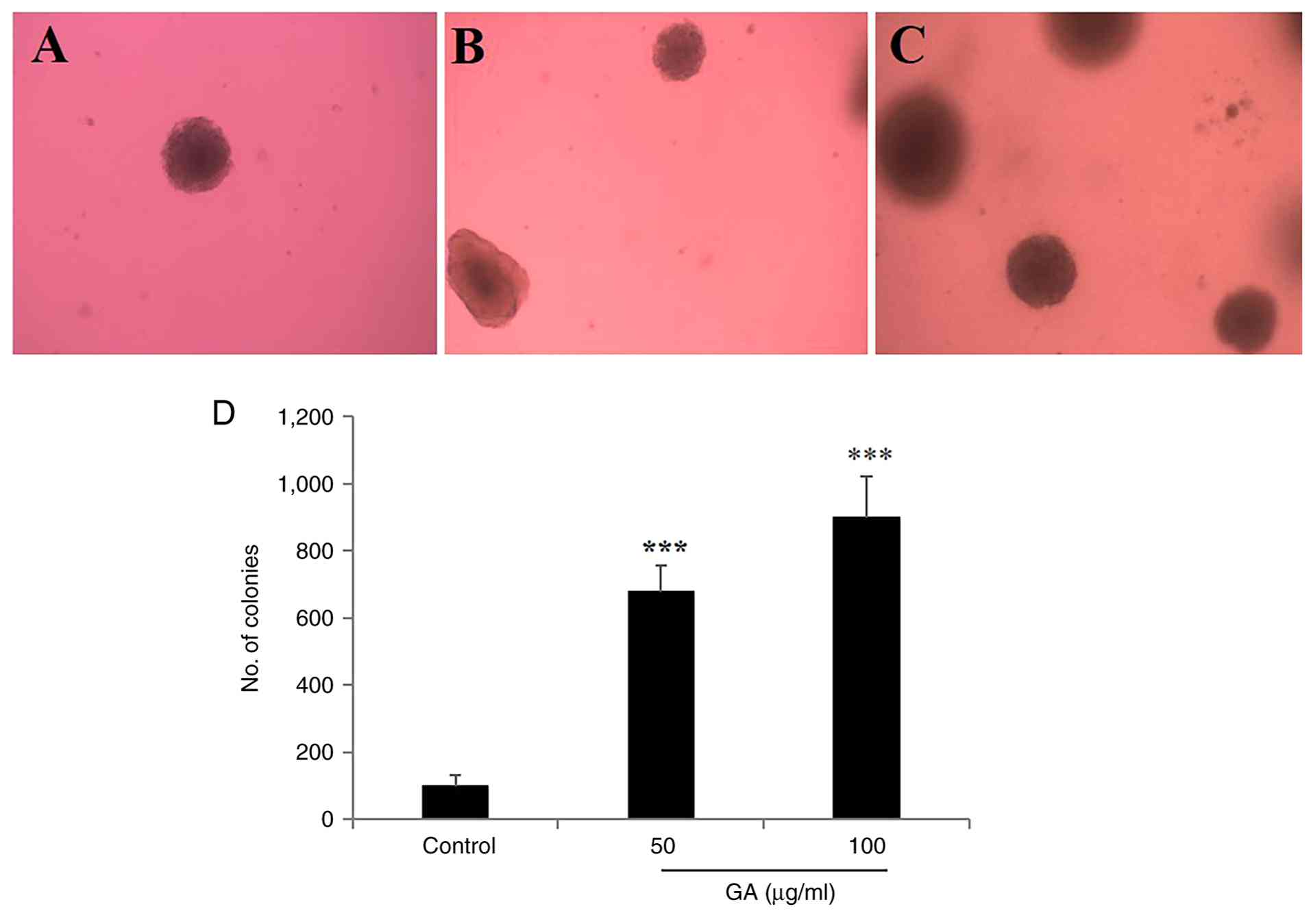
GA promotes colony formation in
IDC-derived KAIMRC1 stem-like cells and TNBC MDA-MB-231 cells
Colony formation assays were conducted to evaluate
the capacity of individual cells to survive, proliferate and form
colonies over an extended period in response to GA treatment. In
IDC-derived KAIMRC1 stem-like cells, GA treatment markedly enhanced
colony formation in a dose-dependent manner (Fig. 5A-D). Specifically, treatment with 50
µg/ml GA increased colony numbers by 6.77-fold (P=0.0003), whereas
100 µg/ml GA resulted in a 9.01-fold enhancement (P=0.0004)
compared with untreated controls (Fig.
5D). In TNBC MDA-MB-231 cells, GA also significantly promoted
colony formation, although to a lesser extent (Fig. S5). Colony numbers increased by
1.46-fold (P=0.0097) and 1.47-fold (P=0.00099) at 50 and 100 µg/ml,
respectively, compared with untreated controls (Fig. S5).
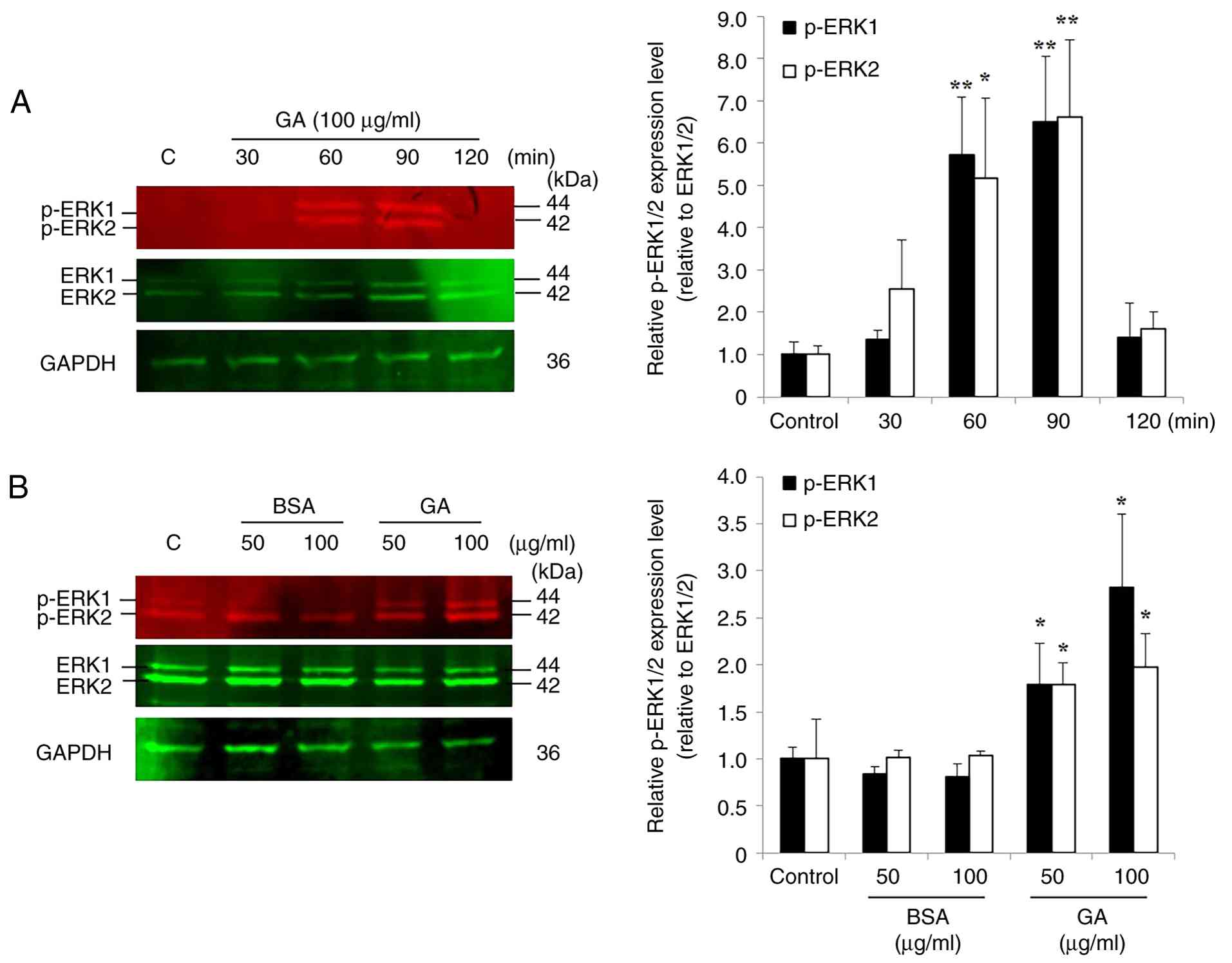
GA induces delayed ERK1/2
phosphorylation in IDC-derived KAIMRC1 stem-like cells
Previous studies using TNBC MDA-MB-231 cells
reported that GA induced optimal phosphorylation of the key
oncogenic signaling protein ERK1/2 after 10 min exposure to 100
µg/ml GA (38,40), a finding that was corroborated under
the present experimental conditions (Fig. S6). In KAIMRC1 stem-like cells, the
optimal duration of ERK1/2 phosphorylation was first determined
using the same effective GA concentration (100 µg/ml) across
multiple incubation periods (30, 60, 90 and 120 min). Upon GA
treatment, ERK1/2 phosphorylation level significantly increased at
60 min (5.70-fold for p-ERK1; P=0.0043 and 5.16-fold for p-ERK2;
P=0.019) and 90 min (6.50-fold for p-ERK1; P=0.0037 and 6.62-fold
for p-ERK2; P=0.0059) compared with untreated control cells,
whereas no significant change was observed at 30 and 120 min
(Fig. 6A). Based on these results,
90 min was selected as the optimal incubation period for maximal
ERK1/2 activation in KAIMRC1 stem-like cells. Subsequently, cells
were exposed to increasing concentrations of GA and unglycated BSA
(50 and 100 µg/ml) for 90 min. GA induced a significant,
concentration-dependent increase in ERK1/2 phosphorylation levels,
with a 1.79-fold (P=0.04) increase for p-ERK1 and a 1.78-fold
(P=0.047) increase for p-ERK2 at 50 µg/ml, and a 2.81-fold
(P=0.016) increase for p-ERK1 and a 1.97-fold (P=0.037) increase
for p-ERK2 at 100 µg/ml, compared with untreated control cells
(Fig. 6B). Conversely, both
concentrations (50 and 100 µg/ml) of unglycated BSA did not alter
ERK1/2 phosphorylation levels compared with untreated controls
(Fig. 6B).
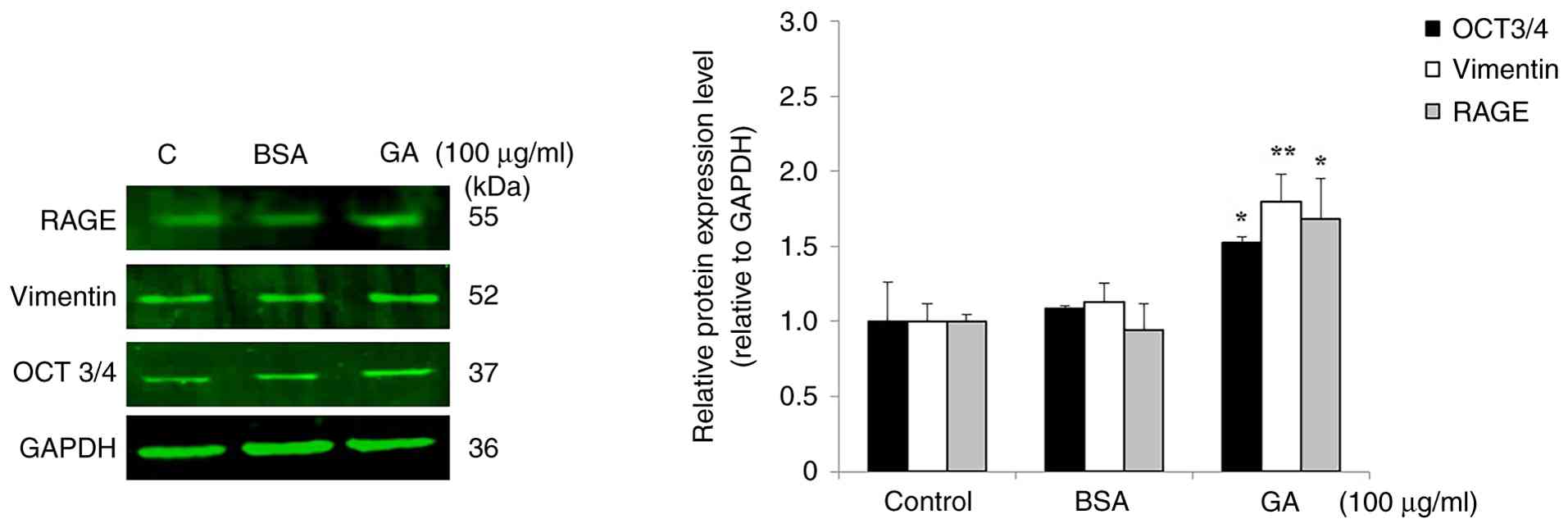
GA upregulates RAGE, vimentin and
OCT3/4 expression in IDC-derived KAIMRC1 stem-like cells and TNBC
MDA-MB-231 cells
To determine whether GA stimulates RAGE signaling,
induces epithelial-mesenchymal transition (EMT) and promotes
stemness features in BC cells, protein expression levels of RAGE,
vimentin and OCT3/4 were examined using western blot analysis.
Treatment with 100 µg/ml GA markedly increased the expression of
all three proteins in both IDC-derived KAIMRC1 stem-like and TNBC
MDA-MB-231 cell lines, compared with their basal levels in
untreated control cells. In KAIMRC1 stem-like cells, GA treatment
significantly elevated the expression of OCT3/4 (1.52-fold;
P=0.02), vimentin (1.79-fold; P=0.005) and RAGE (1.68-fold;
P=0.012) compared with the basal protein expression levels in
untreated control cells (Fig. 7).
Similarly, in MDA-MB-231 cells, GA treatment significantly
increased the expression of OCT3/4 (1.67-fold; P=0.026), vimentin
(2.07-fold; P=0.022) and RAGE (1.56-fold; P=0.044), compared with
control cells (Fig. S7). Treatment
with unglycated BSA did not alter the expression levels of any of
the proteins of interest in either cell line (Figs. 7 and S7).
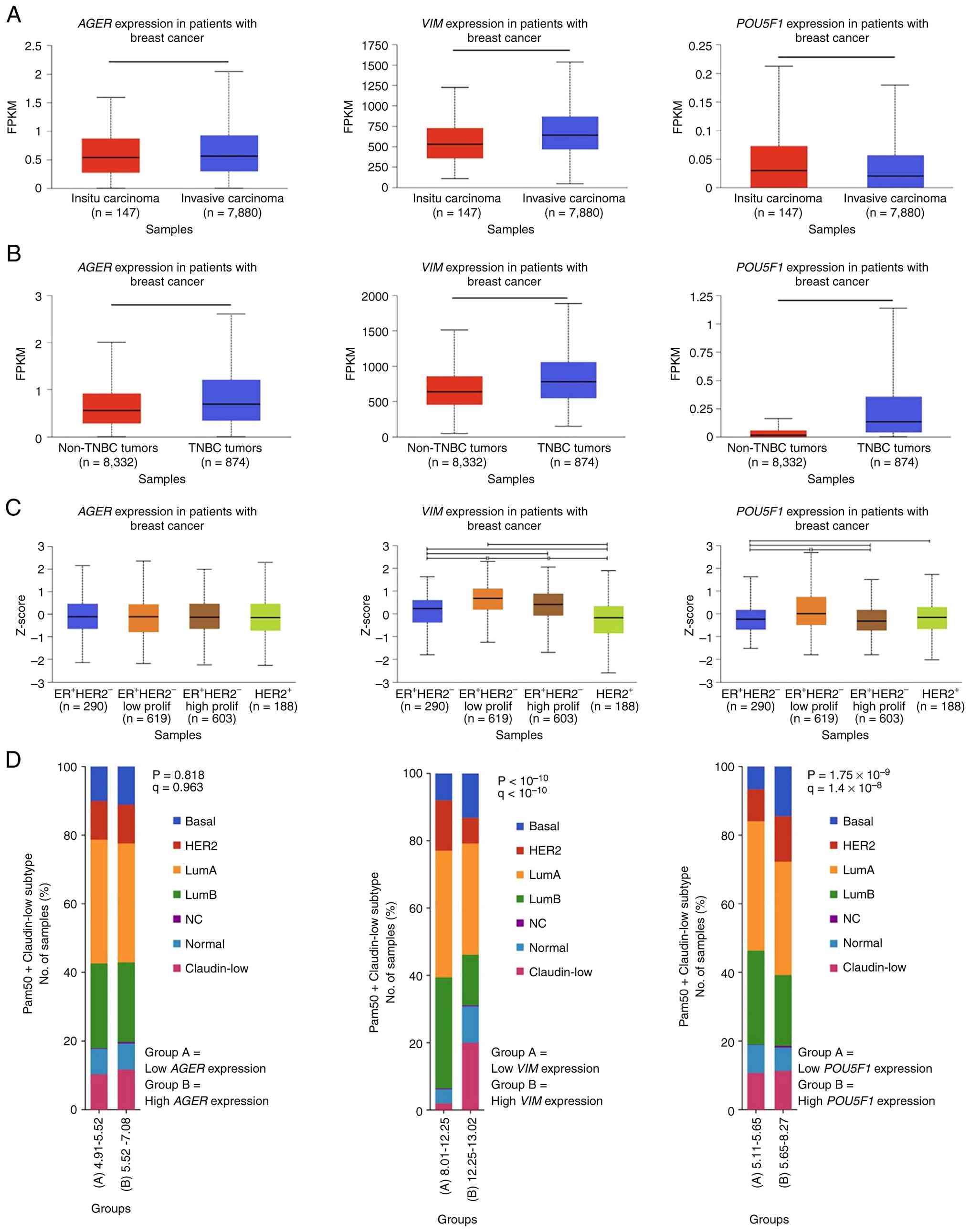
Bioinformatics analysis reveals
elevated VIM gene expression in invasive breast carcinoma, TNBC and
ER+/HER2− tumors, associated with reduced
5-year survival
Gene expression levels of RAGE (AGER),
vimentin (VIM) and OCT4 (POU5F1) were evaluated
utilizing data from the SCAN-B dataset within the MammOnc-DB
database (43,44). Analysis by biopsy type (Fig. 8A) showed that AGER and
VIM were significantly higher in invasive carcinoma
(n=7,880) compared with carcinoma in situ (n=147;
P=2.2×10−2 and P=3.35×10−3; respectively).
Further analysis (Fig. 8B)
indicated that patients with TNBC (n=874) exhibited significantly
higher mRNA levels of AGER, VIM and POU5F1 than those
without TNBC (n=8,332; P<1×10−12). Gene expression
profiles stratified by three-gene classifier subtypes from the
METABRIC dataset were also examined (43,45).
Patients with BC were categorized as ER−HER2−
(n=290), ER+HER2− low proliferative (n=619),
ER+HER2− high proliferative (n=603) and
HER2+ (n=188). Fig. 8C
indicates that patients with ER+HER2−
low-proliferative tumors had significantly higher VIM and
POU5F1 mRNA levels than patients with
ER−HER2− subtype (P=4.99×10−7 and
P=1.63×10−10, respectively). Comparative analysis of
gene expression based on PAM50 and claudin subtypes using the
‘Breast Cancer (METABRIC, Nature 2012 & Nat Commun 2016)’
dataset from cBioPortal (45–50)
revealed that P with high VIM expression were enriched in
basal and claudin-low subtypes (P<1×10−10), whereas
elevated POU5F1 expression was primarily associated with the
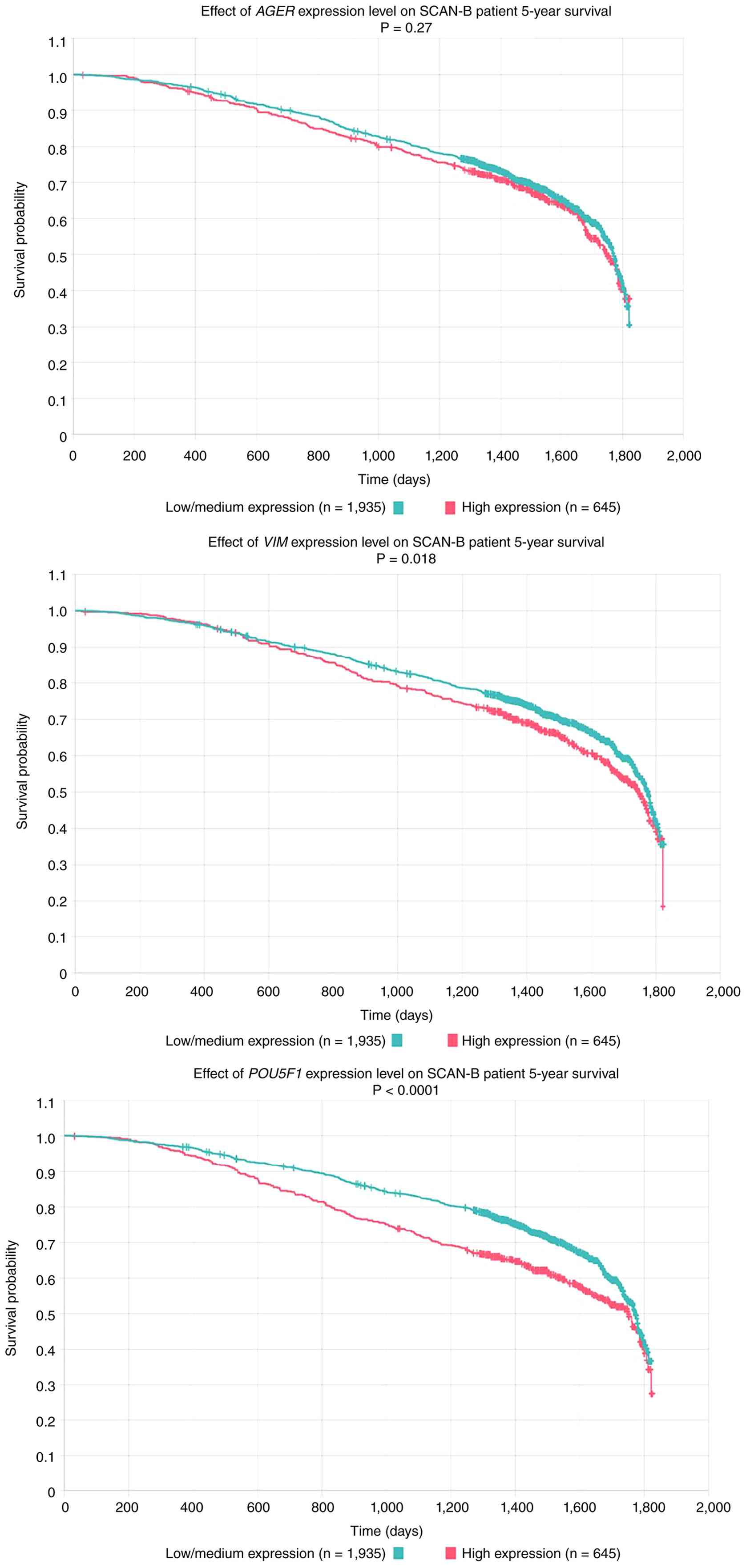
basal subtype (P=1.75×10−9; Fig. 8D). Survival analysis using SCAN-B
dataset indicated that patients with BC with low-to-moderate
VIM (n=1,935) or POU5F1 (n=1,935) expression had
significantly favourable 5-year survival (P=0.018 and P<0.0001,
respectively), whereas AGER expression did not notably
affect the survival of patients with BC (Fig. 9).
Bioinformatics analysis indicates
elevated mRNA expression levels of VIM, AGER and POU5F1 in patients
with BC and T2DM
Panigrahi et al (51) examined 73 patients with BC,
including 36 patients with DM and 37 without DM. The diabetic group
was further stratified into T1DM (n=6) and T2DM (n=30). VIM
mRNA expression was significantly higher in patients with DM when
compared with those without DM, with the most pronounced increase
in patients with T2DM (Mann-Whitney U test; P=0.003; Fig. 10A). Although pairwise comparisons
did not reach statistical significance, a small increase in
POU5F1 (Fig. 10B) and
AGER (Fig. 10C) mRNA
expressions were observed in the diabetic group, particularly in
the T2DM counterparts. A correlation heatmap (Fig. S8) illustrating expression profiles
of VIM, POU5F1 and AGER across all patients showed
marked differences in VIM expression between patients with
BC with and without DM. Conversely, no discernible differences were
observed for POU5F1 and AGER expression (Fig. S8).
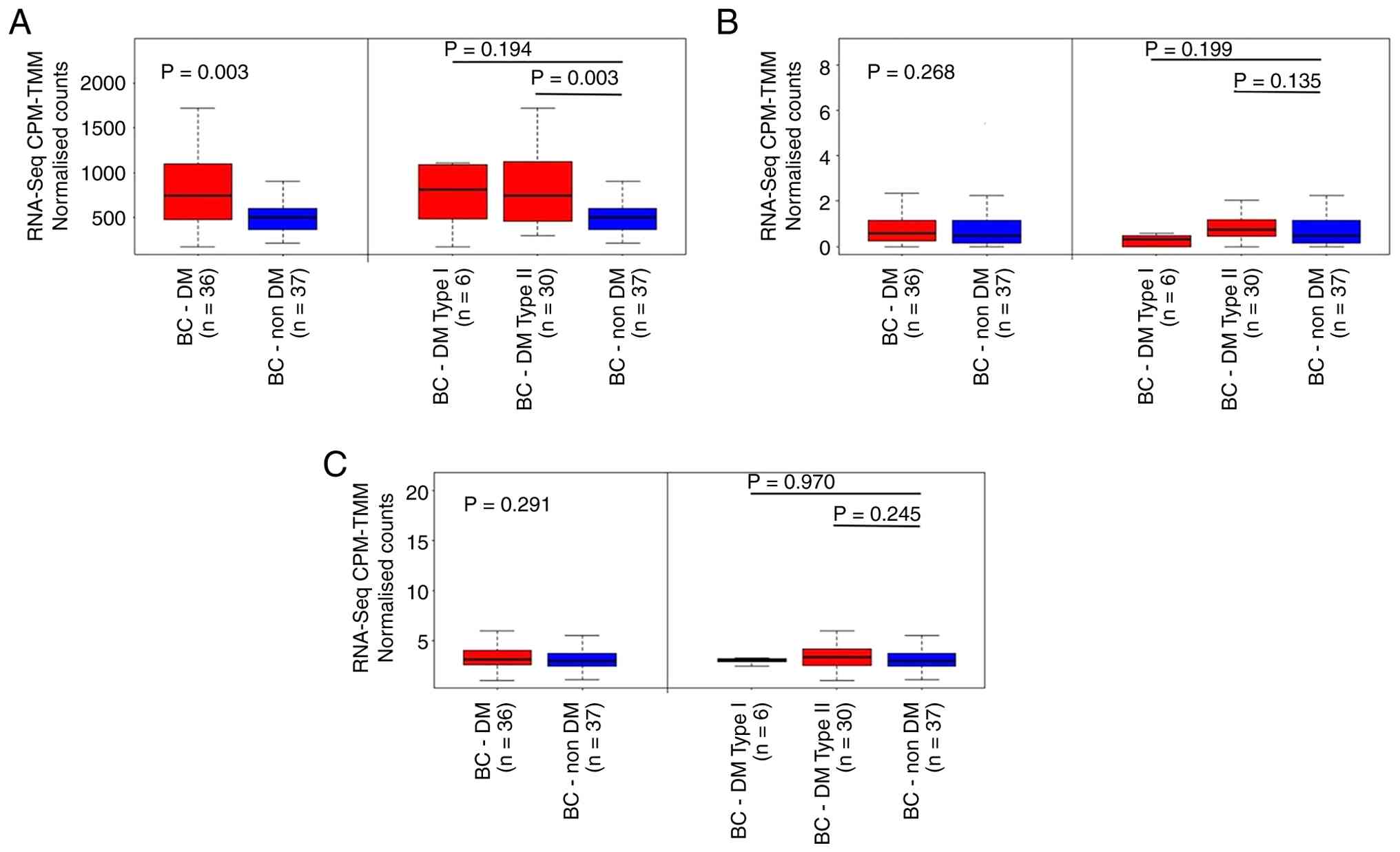 | Figure 10.Upregulation of (A) VIM, (B)
POU5F1 and (C) AGER mRNA levels in patients with BC
with DM. Higher expression was particularly observed in those with
T2DM compared with those without DM. Pairwise comparisons were
conducted using the Mann-Whitney U test. The data were obtained
from the study of Panigrahi et al (51). BC, breast cancer; DM, diabetes
mellitus; T1DM, type 1 diabetes mellitus; T2DM, type 2 diabetes
mellitus, RNA-seq, RNA sequencing; CPM-TMM, counts per million
after trimmed mean of M-values, FPKM, fragments per kilobase of
transcript per million mapped reads. |
GA upregulates vimentin protein
expression in IDC-derived KAIMRC1 stem-like and TNBC MDA-MB-231
cells through RAGE
To determine whether GA activates KAIMRC1 stem-like
and MDA-MB-231 cells through the RAGE signaling pathway, cells were
pretreated with a neutralizing anti-RAGE antibody to inhibit RAGE
signaling or with an IgG isotype control antibody as a negative
control. Cells were exposed to 100 µg/ml GA for 48 h of incubation.
This approach allowed assessment of whether RAGE is required for
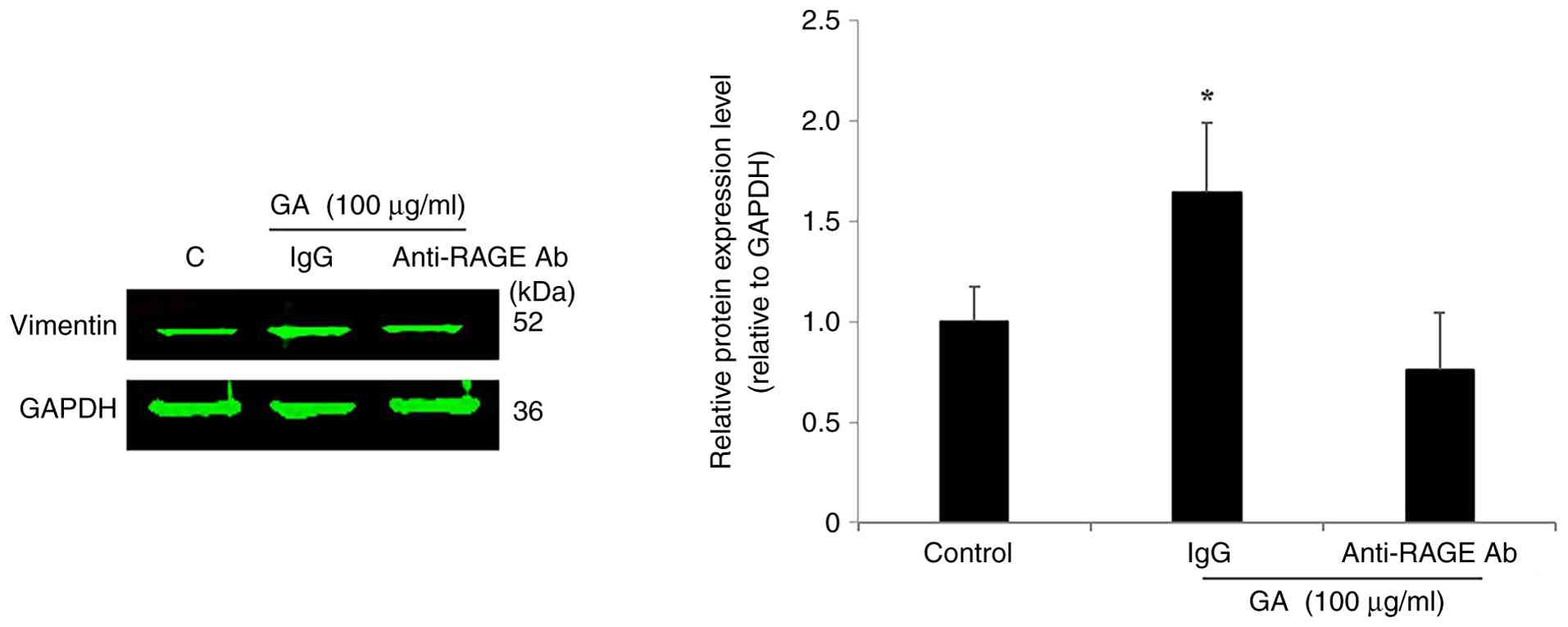
the cellular response to GA stimulation. As shown in Fig. 11, treatment with GA significantly
increased vimentin expression in KAIMRC1 stem-like cells
(1.64-fold; P=0.045; Fig. 11) and
MDA-MB-231 cells (1.39-fold; P=0.012; Fig. S9) pretreated with the IgG control
antibody, compared with untreated controls. Conversely, GA-induced
vimentin upregulation was not observed in either cell line when
RAGE signaling was blocked by pretreatment with an anti-RAGE
antibody (Figs. 11 and S9).
Discussion
Management of patients with T2DM who develop IDC
remains suboptimal, partly because these patients often present at
advanced stages and frequently develop aggressive TNBC (11,53,54).
Challenges include late detection and the limited effectiveness of
currently available systemic treatments, including chemotherapy and
immunotherapy (55). CSCs,
identified in both TNBC and IDC tissues, have garnered considerable
interest in clinical research because they represent a small
population of undifferentiated tumor cells associated with tumor
onset, metastasis, therapy resistance and recurrence (28). KAIMRC1, a spontaneously immortalized
BC cell line derived from the surgical biopsy of a Saudi female
patient diagnosed with IDC was previously established and
characterized (33). Notably, these
cells exhibit CSC-associated phenotypes and CSC markers, including
CD24−/CD44+, CD47 and ALDH1 (33). However, molecular evidence
associating GA, the most abundant globular circulating glycated
protein used as a clinical biomarker for DM (56), to the best of our knowledge,
CSC-associated tumorigenesis in BC remains limited. MG was used for
the generation of GA however, due to its low expression in TNBC
compared with other BC subtypes (57), the biological effects of MG were not
investigated in the present study. Thus, the pro-tumorigenic
effects of GA on IDC-derived KAIMRC1 stem-like cells compared with
MDA-MB-231 cells, a mature and heterogeneous TNBC cell line known
to contain a CSC subpopulation were explored (58). A detailed examination of the
functional and molecular characteristics of BCSCs under
hyperglycemic conditions may help identify new biomarkers and
molecular targets, ultimately supporting the early detection and
targeted therapeutic strategies for patients with T2DM and invasive
BC.
Stemness properties in IDC-derived KAIMRC1 cells
were initially assessed by comparing ALDH activity with that of the
poorly differentiated TNBC cell line MDA-MB-231. ALDH is a highly
active enzyme in both normal stem cells and CSCs, where it
contributes to key stem cell properties, including self-renewal,
differentiation and drug resistance, and is considered a potential
therapeutic and prognostic marker (52,59,60).
In the present study, IDC-derived KAIMRC1 cells exhibited
significantly higher ALDH activity when compared with MDA-MB-231
cells. However, a previous study reported higher cell surface
expression of the CSC-associated protein ALDH1 on MDA-MB-231 cells
when compared with that on KAIMRC1 cells (33). KAIMRC1 cells were previously shown
to possess CSC-like properties, including robust spheroid formation
in soft agar (33). In the present
study, the elevated ALDH activity observed in KAIMRC1 cells further
emphasizes their relevance as a model for studying BC stemness
under hyperglycemic conditions, compared with MDA-MB-231 cells, a
TNBC model frequently associated with T2DM-related BC (16).
At the functional and tumorigenic levels, GA
significantly enhanced cell proliferation, motility, invasion,
colony formation and ERK1/2 phosphorylation in both IDC-derived
KAIMRC1 stem-like and TNBC MDA-MB-231 cells, although the magnitude
and timing of these responses varied between the two models. In
MDA-MB-231 cells, GA exhibited a bell-shaped, dose-dependent
proliferation response, with an optimal effect at 100 µg/ml,
indicating the involvement of a dimeric receptor such as RAGE.
These findings are consistent with a previous study reporting that
GA at 100 µg/ml enhances TNBC MDA-MB-231 cell proliferation,
migration and invasion, accompanied by increased MMP-9 gelatinase
activity mediated through RAGE, using other in vitro
cell-based assays such as trypan blue exclusion, scratch wound and
Boyden chamber assays (38). In the
present study, GA also significantly increased the proliferation of
IDC-derived KAIMRC1 stem-like cells, although to a lesser extent
than that in TNBC MDA-MB-231 cells, indicating a comparatively
slower proliferative rate. This observation is consistent with the
well-recognized tendency of adult CSCs to remain in a quiescent
phase to preserve long-term self-renewal potential and minimize
replication-associated errors (61).
The biological effects of GA on cell motility and,
notably, on the ability to cross a reconstituted basement membrane
revealed that IDC-derived KAIMRC1 stem-like cells were more
responsive than TNBC MDA-MB-231 cells. Although TNBC cells are
highly invasive, CSCs are well known for their metastatic
potential, and previous mechanobiology studies have shown that they
exhibit markedly elevated invasive potential (62,63).
These findings suggest that GA may play an important role in
increasing CSC invasiveness, a hallmark of metastatic progression
and an indicator of tumor aggressiveness. Additionally, a BC
invasion scoring model was recently developed to predict tumor
aggressiveness, therapeutic response and prognosis by analyzing the
PAM pathway (64). Phosphorylation
of p70S6K1, the downstream effector of the PAM pathway, was
detected in IDC tissues from patients with T2DM, and
pharmacological inhibition of p70S6K1 using PF-4708671 suppressed
GA-induced invasion in TNBC (14,40).
These findings indicate that additional modeling of PAM/p70S6K1
pathway in patients with T2DM diagnosed with IDC or TNBC may help
refine predictions of clinical outcomes, including recurrence and
therapeutic responses.
From a tumorigenic standpoint, the present findings
reveal that GA significantly enhanced colony-forming ability in
both KAIMRC1 stem-like cells and MDA-MB-231 cells. The clonogenic
assay is a quantitative in vitro method used to evaluate the
capacity of a single cell to undergo clonal expansion and form a
colony, thereby reflecting tumorigenic potential (65,66).
In the present study, treatment with GA produced more colonies in
KAIMRC1 stem-like cells when compared with that in the more
differentiated MDA-MB-231 cells, indicating a potent tumorigenic
effect in CSCs-enriched cells. Consistent with these findings, a
commercial AGE-BSA at 40–80 µg/ml promoted colony formation in
human primary TNBC cells (67).
Additionally, several AGEs-BSA preparations, including MG-derived
AGEs, have been shown to exert pro-tumorigenic effects in prostate
cancer cells and xenografts mouse models, accompanied by RAGE
overexpression and increased levels of EMT markers, including
N-cadherin and vimentin (68).
Verification of the tumorigenic potential of GA on IDC-derived
KAIMRC1 stem-like cells could be strengthened using in vivo
models, such as xenografts mouse models.
Focusing on the main signaling oncoprotein ERK1/2,
GA was shown to transiently activate the ERK1/2 signaling pathway
in TNBC MDA-MB-231 cells and IDC-derived KAIMRC1 stem-like cells.
Previous studies reported that, in MDA-MB-231 cells, ERK1/2
phosphorylation peaks rapidly, ~10 min posttreatment with an
effective concentration of GA, and exhibits a characteristic
bell-shaped response, indicating the involvement of the dimeric
receptor RAGE (38,40). Additional in vitro studies
have confirmed that ERK signaling is a key component of the
AGEs-RAGE axis promoting carcinogenesis (67,69,70).
In contrast to the rapid response observed in MDA-MB-231 cells,
KAIMRC1 stem-like cells exhibited delayed ERK1/2 activation, with
phosphorylation significantly increasing only after 60–90 min of
treatment with GA. Notably, stem-like cells, including those
exhibiting mesenchymal features, often show altered sensitivity to
external stimuli due to their unique epigenetic and signaling
profiles (71,72). Therefore, the delayed
phosphorylation response in KAIMRC1 stem-like cells may reflect
differences from mature cancer cells in membrane receptor
expression, intracellular signaling dynamics or metabolic
activity.
The regulatory role of GA in key stemness markers,
such as OCT3/4 and vimentin, was also examined. The present results
revealed that treatment with GA induced a significant upregulation
of mesenchymal markers OCT3/4 and vimentin in both IDC-derived
KAIMRC1 stem-like cells and TNBC MDA-MB-231 cells compared with
untreated controls. These findings are consistent with clinical
reports indicating that T2DM promotes differentiation of
mesenchymal stem cells (MSCs), particularly adipose tissue-derived
MSCs, generating MSC-derived extracellular vesicles in the blood
circulation, that contribute to BC metastasis (73,74).
In vivo experimental evidence from diabetic mouse models has
similarly shown that hyperglycemia induces a mesenchymal phenotype
in BC cells, thereby increasing their motility and invasive
potential (75). OCT3/4 is a
transcription factor that plays a key role in expressing and
maintaining pluripotency and self-renewal (76). Reactivation of OCT3/4 expression has
been observed in several human BC cell lines but not in
non-malignant cells (77). Notably,
previous studies have shown that OCT4 expression in CSC promotes
colony formation, metastasis and cancer progression, as shown in
human cervical and liver cancer models (78,79).
Consistent with these findings, the present data suggest that
GA-induced elevation of OCT3/4 may mediate greater enhancement of
colony formation in KAIMRC1 stem-like cells when compared with that
in MDA-MB-231 cells. Vimentin, a pivotal cytoskeletal protein
typically associated with mesenchymal cells, is a key marker of EMT
(80). EMT mechanisms are essential
for improving cancer cell invasion and migration, which are key
factors influencing metastatic potential (81). In the present study, GA-induced
vimentin overexpression coincided with augmented invasiveness and
motility in both cell lines. Furthermore, elevated vimentin
expression was observed in patients with TNBC and in
ER+/HER2− subtypes, including the cellular
model KAIMRC1, associating with poor-prognosis in patients with
T2DM diagnosed with BC compared with those without DM. These
findings corroborate previous clinical studies reporting that
vimentin is a poor prognostic biomarker and therapeutic target in
patients with BC and TNBC, primarily due to its involvement in
tumor progression and metastasis (82,83).
Furthermore, using the invasive BC cell line
SK-BR3, metformin was shown to suppress vimentin expression
(84). It would therefore be
beneficial to monitor vimentin expression in patients with T2DM
diagnosed with invasive BC and determine overall survival.
Importantly, RAGE neutralization significantly inhibited GA-induced
enhancement of vimentin expression, indicating that RAGE is an
important mediator of GA-driven EMT and potentially
stemness-related phenotypes. This observation aligns with prior
evidence suggesting that AGEs, particularly GA, modulate EMT
processes through RAGE-dependent mechanisms (26). Other research tools such as small
interfering RNA technology could be applied to verify and
strengthen the key role of RAGE signaling in GA-induced vimentin
expression. Additionally, an in silico analysis of the
publicly available gene expression data from Panigrahi et al
(51) was conducted on 36 patients
with DM and 37 without DM. The small sample size may constrain the
ability to draw statistically significant or clinically meaningful
conclusions, particularly when considering potential confounding
factors such as disease duration, severity, HbA1c levels and
treatment regimens. To the best of our knowledge, no other publicly
available diabetic BC datasets of sufficient sample size exist to
allow further in silico validation. Despite this limitation,
the analyzed dataset offers valuable preliminary insights that
warrant further exploration. These findings underscore the
importance of monitoring the gene expression levels of VIM,
POU5F1 and AGER in local Saudi cohorts of patients with BC and
with DM.
In conclusion, the present study showed that GA
stimulates key tumorigenic and stemness-associated properties in
IDC-derived KAIMRC stem-like cells and TNBC MDA-MB-231 cells,
representing in vitro models of BC subtypes prevalent in
patients with T2DM. GA primarily promotes proliferation in the
poorly differentiated MDA-MB-231 cell line, while concurrently
enhancing stemness-associated features, motility, invasiveness and
EMT in the KAIMRC1 stem-like cell line. These effects are mediated
through the RAGE signaling pathway, as evidenced by the reduction
in GA-induced vimentin upregulation following RAGE neutralization.
Bioinformatics analyses further indicate that low vimentin
expression is associated with improved overall survival in patients
with T2DM and BC. Although additional in vitro studies in
primary IDC-derived BC cells are warranted, these findings
highlight the AGEs-RAGE axis as a key mechanism linking
hyperglycemia to enhanced stemness in IDC-derived BCSCs and TNBC.
Notably, vimentin emerges as a potential prognostic biomarker, and
targeting the AGEs-RAGE pathway may represent a viable therapeutic
strategy. Further clinical research on the impact of VIM
transcript levels and their associations with overall survival in
patients with T2DM with invasive BC is necessary to validate its
prognostic utility.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
This work was supported by King Abdullah International Medical
Research Center (grant no. SP24R/018/01).
Authors' contributions
SMN conceived and conducted the study. SMN and RaA
supervised the study, analysed and interpreted the data, and
reviewed the manuscript. MA, NA, SH, ReA and RiA performed the
experiments, generated the data, collected the data and reviewed
the manuscript. MKA and AA carried out bioinformatics analyses. MA,
MKA, AAA and SMN interpreted the data, wrote and reviewed the
manuscript. SMN and MKA confirm the authenticity of all the raw
data. All authors read and approved the final manuscript.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
AGEs
|
advanced glycation end products
|
|
ALDH
|
aldehyde dehydrogenase
|
|
BC
|
breast cancer
|
|
BSA
|
bovine serum albumin
|
|
CIM
|
cellular invasion/migration
|
|
CPM
|
counts per million
|
|
CSC
|
cancer stem-like cells
|
|
DM
|
diabetes mellitus
|
|
DMEM
|
Dulbecco's modified Eagle's
medium
|
|
EMT
|
epithelial-mesenchymal transition
|
|
ERK
|
extracellular signal-regulated
kinase
|
|
GA
|
glycated albumin
|
|
GAPDH
|
glyceraldehyde 3-phosphate
dehydrogenase
|
|
IDC
|
invasive ductal carcinoma
|
|
IgG
|
immunoglobulin G
|
|
IGF
|
insulin-like growth factor
|
|
LSM
|
low-serum medium
|
|
METABRIC
|
molecular taxonomy of breast cancer
international consortium
|
|
MG
|
methylglyoxal
|
|
NAD
|
nicotinamide adenine dinucleotide
|
|
OCT
|
octamer-binding transcription
factor
|
|
p
|
phosphorylated
|
|
PAM
|
PI3K/Akt/mTOR
|
|
PAM50
|
prediction analysis of microarray
50
|
|
RAGE
|
receptor for advanced glycation end
products
|
|
RT
|
room temperature
|
|
RTCA-DP
|
Real-time Cell Analyzer Dual
Purpose
|
|
SCAN-B
|
Swedish Cancerome Analysis
Network-Breast
|
|
T1DM
|
type 1 diabetes mellitus
|
|
T2DM
|
type 2 diabetes mellitus
|
|
TBS
|
Tris-buffered saline
|
|
TMM
|
Trimmed Mean of M-values
|
|
TNBC
|
triple-negative breast cancer
|
References
|
1
|
Lu X, Xie Q, Pan X, Zhang R, Zhang X, Peng
G, Zhang Y, Shen S and Tong N: Type 2 diabetes mellitus in adults:
Pathogenesis, prevention and therapy. Signal Transduct Target Ther.
9:2622024. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
IDF Diabetes Atlas 2025 (IDF), . IDF,
Brussels. 2025.https://diabetesatlas.org/resources/idf–diabetes–-atlas–2025/
|
|
3
|
GBD 2021 Diabetes Collaborators, . Global,
regional, and national burden of diabetes from 1990 to 2021, with
projections of prevalence to 2050: A systematic analysis for the
global burden of disease study 2021. Lancet. 402:203–234. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Robertson CC, Elgamal RM, Henry-Kanarek
BA, Arvan P, Chen S, Dhawan S, Eizirik DL, Kaddis JS, Vahedi G,
Parker SCJ, et al: Untangling the genetics of beta cell dysfunction
and death in type 1 diabetes. Mol Metab. 86:1019732024. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Młynarska E, Czarnik W, Dzieża N,
Jędraszak W, Majchrowicz G, Prusinowski F, Stabrawa M, Rysz J and
Franczyk B: Type 2 diabetes mellitus: New pathogenetic mechanisms,
treatment and the most important complications. Int J Mol Sci.
26:10942025. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Petrović N, Milosavljević M, Gojak R,
Ðešević M, Lakić D, Stević I and Janković S: Costs of treating type
2 diabetes mellitus and its complications. Biotechnol Biotchnol
Equip. 38:23999412024. View Article : Google Scholar
|
|
7
|
Zhang YY, Li YJ, Xue CD, Li S, Gao ZN and
Qin KR: Effects of T2DM on cancer progression: Pivotal
precipitating factors and underlying mechanisms. Front Endocrinol
(Lausanne). 15:13960222024. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Alshammari AM, Elnaem MH and Ong SC:
Evaluating the economic impact of diabetes mellitus: A
hospital-centric cost analysis in Hail, Saudi Arabia. Clinicoecon
Outcomes Res. 17:473–484. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wen S, Yuan Y, Li Y, Xu C, Chen L, Ren Y,
Wang C, He Y, Li X, Gong M, et al: The effects of non-insulin
anti-diabetic medications on the diabetic microvascular
complications: a systematic review and meta-analysis of randomized
clinical trials. BMC Endocr Disord. 25:1792025. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hao Q, Huang Z, Li Q, Liu D, Wang P, Wang
K, Li J, Cao W, Deng W, Wu K, et al: A novel metabolic
reprogramming strategy for the treatment of diabetes-associated
breast cancer. Adv Sci (Weinh). 9:e21023032022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lu Y, Hajjar A, Cryns VL, Trentham-Dietz
A, Gangnon RE, Heckman-Stoddard BM and Alagoz O: Breast cancer risk
for women with diabetes and the impact of metformin: A
meta-analysis. Cancer Med. 12:11703–11718. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kounatidis D, Vallianou NG, Karampela I,
Rebelos E, Kouveletsou M, Dalopoulos V, Koufopoulos P,
Diakoumopoulou E, Tentolouris N and Dalamaga M: Anti-diabetic
therapies and cancer: From bench to bedside. Biomolecules.
14:14792024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Vulichi SR, Runthala A, Begari N, Rupak K,
Chunduri VR, Kapur S, Chippada AR and Sistla DSM: Type-2 diabetes
mellitus-associated cancer risk: In pursuit of understanding the
possible link. Diabetes Metab Syndr. 16:1025912022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Matou-Nasri S, Aldawood M, Alanazi F and
Khan AL: Updates on triple-negative breast cancer in type 2
diabetes mellitus patients: From risk factors to diagnosis,
biomarkers and therapy. Diagnostics (Basel). 13:23902023.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Durrani IA, John P, Bhatti A and Khan JS:
Network medicine based approach for identifying the type 2
diabetes, osteoarthritis and triple negative breast cancer
interactome: Finding the hub of hub genes. Heliyon. 10:e366502024.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Liu L, Gao Y, Liu P, Hui R and Zhang J:
Association of type 2 diabetes mellitus with histopathological
features of non-metastatic breast cancer in Chinese women: A
retrospective Cross-sectional study. Sci Rep. 15:286452025.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 countries in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI
|
|
18
|
Lawrenson R, Lao C, Stanley J, Teng A,
Kuper-Hommel M, Campbell I, Krebs J, Sika-Paotonu D, Koea J,
Meredith I and Gurney J: Does diabetes affect breast cancer
survival? Cancer Rep (Hoboken). 7:e20402024.PubMed/NCBI
|
|
19
|
Natalicchio A, Marrano N, Montagnani M,
Gallo M, Faggiano A, Zatelli MC, Argentiero A, Del Re M, D'Oronzo
S, Fogli S, et al: Glycemic control and cancer outcomes in
oncologic patients with diabetes: An Italian Association of Medical
Oncology (AIOM), Italian Association of Medical Diabetologists
(AMD), Italian Society of Diabetology (SID), Italian Society of
Endocrinology (SIE), Italian Society of Pharmacology (SIF)
multidisciplinary critical view. J Endocrinol Invest. 47:2915–2928.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Sears AJ, Wild SH, Mesa-Eguiagaray I, Hall
PS and Figueroa JD: Breast cancer survival and mortality among
women with type 2 diabetes: A retrospective cohort study. Sci Rep.
15:261442025. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Jordt N, Kjærgaard KA, Thomsen RW,
Borgquist S and Cronin-Fenton D: Breast cancer and incidence of
type 2 diabetes mellitus: A systematic review and meta-analysis.
Breast Cancer Res Treat. 202:11–22. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ahmed SBM, Radwan N, Amer S, Sharif-Askari
NS, Mahdami A, Samara KA, Halwani R and Jelinek HF: Assessing the
link between diabetic metabolic dysregulation and breast cancer
progression. Int J Mol Sci. 24:118162023. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang AMY, Xia YH, Lin JSH, Chu KH, Wang
WCK, Ruiter TJJ, Yang JCC, Chen N, Chhuor J, Patil S, et al:
Hyperinsulinemia acts via acinar insulin receptors to initiate
pancreatic cancer by increasing digestive enzyme production and
inflammation. Cell Metab. 35:2119–2135.e5. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang W, Hapach LA, Taufalele PV, Bates ME,
Wu Y and Reinhart-King C: Diabetic hyperglycemia-induced glycation
regulates tumor vasculature integrity via NF-kB-mediated GM-CSF
secretion by tumor cells. Cell Biomater. 1:1001862025. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Vella V, Lappano R, Bonavita E, Maggiolini
M, Clarke RB, Belfiore A and De Francesco EM: Insulin/IGF axis and
the receptor for advanced glycation end products: Role in
meta-inflammation and potential in cancer therapy. Endocr Rev.
44:693–723. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Coser M, Neamtu BM, Pop B, Cipaian CR and
Crisan M: RAGE and its ligands in breast cancer progression and
metastasis. Oncol Rev. 18:15079422025. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Oliveira RV, Souza VB, Souza PC, Soares
FA, Vassallo J, Rocha RM and Schenka AA: Detection of putative
stem-cell markers in invasive ductal carcinoma of the breast by
immunohistochemistry: Does it improve prognostic/predictive
assessments? Appl Immunohistochem Mol Morphol. 26:760–768. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Korfias D, Contis J, Frangou-Plemenou M,
Gennatas K, Kondis A and Vlachodimitropoulos D: Stem cells in
ductal breast cancer: Immunohistochemical expression of CD44, CD24,
CD133, and ALDH-1 markers in 104 cases. Eur J Gynaecol Oncol.
41:36–41. 2020. View Article : Google Scholar
|
|
29
|
Bamodu OA, Chung CC, Pisanic TR II and Wu
ATH: The intricate interplay between cancer stem cells and
cell-of-origin of cancer: Implications for therapeutic strategies.
Front Oncol. 14:14046282024. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Mengistu BA, Tsegaw T, Demessie Y, Getnet
K, Bitew AB, Kinde MZ, Beirhun AM, Mebratu AS, Mekasha YT, Feleke
MG and Fenta MD: Comprehensive review of drug resistance in
mammalian cancer stem cells: Implications for cancer therapy.
Cancer Cell Int. 24:4062024. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Park SY, Choi JH and Nam JS: Targeting
cancer stem cells in triple-negative breast cancer. Cancers
(Basel). 11:9652019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Murillo LL, Sugden CJ, Ozsvari B,
Moftakhar Z, Hassan GS, Sotgia F and Lisanti MP:
ALDHHigh breast cancer stem cells exhibit a
mesenchymal-senescent hybrid phenotype, with elevated metabolic and
migratory activities. Cells. 13:20592024. View Article : Google Scholar
|
|
33
|
Ali R, Samman N, Al Zahrani H, Nehdi A,
Rahman S, Khan AL, Al Balwi M, Alriyees LA, Alzaid M, Al Askar A
and Boudjelal M: Isolation and characterization of a new naturally
immortalized human breast carcinoma cell line, KAIMRC1. BMC Cancer.
17:8032017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ambrosio MR, Mosca G, Migliaccio T,
Liguoro D, Nele G, Schonauer F, D'Andrea F, Liotti F, Prevete N,
Melillo RM, et al: Glucose enhances pro-tumorigenic functions of
mammary adipose-derived mesenchymal stromal/stem cells on breast
cancer cell lines. Cancers (Basel). 14:54212022. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ayodeji SA, Bao B, Teslow EA, Polin LA,
Dyson G, Bollig-Fischer A and Fehl C: Hyperglycemia and O-GlcNAc
transferase activity drive a cancer stem cell pathway in
triple-negative breast cancer. Cancer Cell Int. 23:1022023.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Júnior JPL, Brescansin CP, Santos-Weiss
ICR, Welter M, Souza EM, Rego FGM, Picheth G and Alberton D: Serum
fluorescent advanced glycation End (F-AGE) products in gestational
diabetes patients. Arch Endocrinol Metab. 61:233–237. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Kato S, Matsumura T, Sugawa H and Nagai R:
Correlation between serum advanced glycation end-products and
vascular complications in patient with type 2 diabetes. Sci Rep.
14:187222024. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Sharaf H, Matou-Nasri S, Wang Q, Rabhan Z,
Al-Eidi H, Al Abdulrahman A and Ahmed N: Advanced glycation
endproducts increase proliferation, migration and invasion of the
breast cancer cell line MDA-MB-231. Biochim Biophys Acta.
1852:429–441. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Matou-Nasri S, Sharaf H, Wang Q, Almobadel
N, Rabhan Z, Al-Eidi H, Yahya WB, Trivilegio T, Ali R, Al-Shanti N
and Ahmed N: Biological impact of advanced glycation endproducts on
estrogen receptor breast cancer cell line MDA-MB-231. Biochim
Biophys Acta Mol Basis Dis. 1863:2808–2820. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Alanazi F, Alsaleh AA, Alamoudi MK,
Alasiri A, Haymond A and Matou-Nasri S: Targeting p70S6K1 inhibits
glycated albumin-induced triple-negative breast cancer cell
invasion and overexpression of galectin-3, a potential prognostic
marker in diabetic patients with invasive breast cancer.
Biomedicines. 13:6122025. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Brix N, Samaga D, Hennel R, Gehr K,
Zitzelsberger H and Lauber K: The clonogenic assay: Robustness of
plating efficiency-based analysis is strongly compromised by
cellular cooperation. Radiat Oncol. 15:2482020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Maashi Y, Almutairi S, Aldawood M, Al-Eidi
H, AlRoshody R, Alghamdi HA, Bahattab S, Alsaleh AA, Alkadi H,
Alghamdi S, et al: In vitro oncogenic effects of glycated albumin
in human colorectal cancer cell lines HT29 and SW620 revealing
EpCAM and Galectin-3 upregulation in Type 2 diabetic colorectal
cancer tissues as potential biomarkers. J Oncol Res Ther.
9:102272024.
|
|
43
|
Karthikeyan SK, Chandrashekar DS, Sahai S,
Shrestha S, Aneja R, Singh R, Kleer CG, Kumar S, Qin ZS, Nakshatri
H, et al: MammOnc-DB, an integrative breast cancer data analysis
platform for target discovery. NPJ Breast Cancer. 11:352025.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Staaf J, Häkkinen J, Hegardt C, Saal LH,
Kimbung S, Hedenfalk I, Lien T, Sørlie T, Naume B, Russnes H, et
al: RNA sequencing-based single sample predictors of molecular
subtype and risk of recurrence for clinical assessment of
early-stage breast cancer. NPJ Breast Cancer. 8:942022. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Pereira B, Chin SF, Rueda OM, Vollan HK,
Provenzano E, Bardwell HA, Pugh M, Jones L, Russell R, Sammut SJ,
et al: The somatic mutation profiles of 2,433 breast cancers
refines their genomic and transcriptomic landscapes. Nat Commun.
7:114792016. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Dicov. 2:401–404.
2012. View Article : Google Scholar
|
|
47
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 2:pl12013.PubMed/NCBI
|
|
48
|
De Bruijn I, Kundra R, Mastrogiacomo B,
Tran TN, Sikina L, Mazor T, Li X, Ochoa A, Zhao G, Lai B, et al:
Analysis and visualization of longitudinal genomic and clinical
data from the AACR project GENIE Biopharma collaborative in
cBioPortal. Cancer Res. 83:3861–3867. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Rueda OM, Sammut SJ, Seoane JA, Chin SF,
Caswell-Jin JL, Callari M, Batra R, Pereira B, Bruna A, Ali HR, et
al: Dynamics of breast-cancer relapse reveal late-recurring
ER-positive genomic subgroups. Nature. 567:399–404. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Curtis C, Shah SP, Chin SF, Turashvili G,
Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, et
al: The genomic and transcriptomic architecture of 2,000 breast
tumours reveals novel subgroup. Nature. 486:346–352. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Panigrahi G, Candia J, Dorsey TH, Tang W,
Ohara Y, Byun JS, Minas TZ, Zhang A, Ajao A, Cellini A, et al:
Diabetes-associated breast cancer is molecularly distinct and shows
a DNA damage repair deficiency. JCI Insight. 8:e1701052023.
View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Wang S, Ma L, Wang Z, He H, Chen H, Duan
Z, Li Y, Si Q, Chuang TH, Chen C, et al: Lactate dehydrogenase-A
(LDH-A) preserves cancer stemness and recruitment of
tumor-associated macrophages to promote breast cancer progression.
Front Oncol. 11:6544522021. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Park YMM, Bookwalter DB, O'Brien KM,
Jackson CL, Weinberg CR and Sandler DP: A prospective study of type
2 diabetes, metformin use, and risk of breast cancer. Ann Oncol.
32:351–359. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Zhang F, de Haan-Du J, Sidorenkov G,
Landman GWD, Jalving M, Zhang Q and de Bock GH: Type 2 diabetes
mellitus and clinicopathological tumor characteristics in women
diagnosed with breast cancer: A systematic review and
meta-analysis. Cancers (Basel). 13:49922021. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Abdelaty WR: Outcome of management of
invasive lobular carcinoma in comparison to invasive ductal
carcinoma. Al-Azhar Int Med J. 5:262024. View Article : Google Scholar
|
|
56
|
Xiong JY, Wang JM, Zhao XL, Yang C, Jiang
XS, Chen YM, Chen CQ and Li ZY: Glycated albumin as a biomarker for
diagnosis of diabetes mellitus: A systematic review and
meta-analysis. World J Clin Cases. 9:9520–9534. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Chiavarina B, Nokin MJ, Durieux F, Bianchi
E, Turtoi A, Peulen O, Peixoto P, Irigaray P, Uchida K, Belpomme D,
et al: Triple negative tumors accumulate significantly less
methylglyoxal specific adducts than other human breast cancer
subtypes. Oncotarget. 5:5472–5482. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Altundag-Erdogan Ö, Çetinkaya H, Öteyaka
MÖ and Çelebi-Slatik B: Targeting MDA-MB-231 cancer stem cells with
temsirolimus in 3D collagen/PGA/NA2SiO3-based bone model. Macromol
Mater Engineer. 310:24003602025. View Article : Google Scholar
|
|
59
|
Gomez-Salazar MA, Wang Y, Thottappillil N,
Hardy RW, Alexandre M, Höller F, Martin N, Gonzalez-Galofre ZN,
Stefancova D, Medici D, et al: Aldehyde dehydrogenase, a marker of
normal and malignant stem cells, typifies mesenchymal progenitors
in perivascular niches. Stem Cells Transl Med. 12:474–484. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Li B, Tian J, Zhang F, Wu C, Li Z, Wang D,
Zhuang J, Chen S, Song W, Tang Y, et al: Self-assembled aldehyde
dehydrogenase-activatable nano-prodrug for cancer stem
cell-enriched tumor detection and treatment. Nat Commun.
16:94172024. View Article : Google Scholar
|
|
61
|
Altshuler A, Wickstrom SA and
Shalom-Feuerstein R: Spotlighting adult stem cells: Advances,
pitfall, and challenges. Trends Cell Biol. 33:477–494. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Alvarez-Elizondo MB and Weihs D: Breast
cancer stem cells: Mechanobiology reveals highly invasive cancer
cell subpopulations. Cell Mol Life Sci. 79:1342022. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Ouyang M, Gui Y, Li N and Zhao L:
Prognostic model based on tumor stemness genes for triple-negative
breast cancer. Sci Rep. 14:308552024. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Li X, Zhang Y, Gong J, Liu W, Zhao H, Xue
H, Ren Z, Bao J and Lin Z: Development of a breast cancer invasion
score to predict tumor aggressiveness and prognosis via
PI3K/AKT/mTOR pathway analysis. Cell Death Disc. 11:1572025.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Nayak A, Warrier NM and Kumar P: Cancer
stem cells and the tumor microenvironment: Targeting the critical
crosstalk through nanocarrier system. Stem Cell Rev Rep.
18:2209–2233. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Chu X, Tian W, Ning J, Xiao G, Zhou Y,
Wang Z, Zhai Z, Tanzhu G, Yang J and Zhou R: Cancer stem cells:
Advances in knowledge and implications for cancer therapy. Signal
Transduct Target Ther. 9:1702024. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Lee KJ, Yoo JW, Kim YK, Choi JH, Ha TY and
Gil M: Advanced glycation end products promote triple negative
breast cancer cells via ERK and NF-kB pathway. Biochem Biophys Res
Commun. 495:2195–2201. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Khoo SH, Wu PR, Yeh KT, Hsu SL and Wu CH:
Biological and clinical significance of the AGE-RAGE axis in the
aggressiveness and prognosis of prostate cancer. J Food Drug Anal.
31:664–682. 2023.PubMed/NCBI
|
|
69
|
Azizian-Farsani F, Abedpoor N, Sheikhha
MH, Gure AO, Nasr-Esfahani MH and Ghaedi K: Receptor for advanced
glycation end products acts as a fuel to colorectal cancer
development. Front Oncol. 10:5522832020. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Nam HK, Jeong SR, Pyo MC, Ha SK, Nam MH
and Lee KW: Methylglyoxal-derived advanced glycation end products
(AGE4) promote cell proliferation and survival in renal cell
carcinoma cells through the RAGE/Akt/ERK signaling pathways. Biol
Pharm Bull. 44:1697–1706. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Galassi C, Manic G, Estellar M, Galluzzi L
and Vitale I: Epigenetic regulation of cancer stemness. Sign
Transduct Target Ther. 10:2432025. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Lee H, Kim B, Park J, Park S, Yoo G, Yum
S, Kang W, Lee JM, Youn H and Youn B: Cancer stem cells: Landscape,
challenges and emerging therapeutic innovations. Sign Transduct
Target Ther. 10:2482025. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Khanh VC, Fukushige M, Moriguchi K,
Yamashita T, Osaka M, Hiramatsu Y and Ohneda O: Type 2 diabetes
mellitus induced paracrine effects on breast cancer metastasis
through extracellular vesicles from human mesenchymal stem cells.
Stem Cells Dev. 29:1382–1394. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Chang YH, Ngo NH, Vuong CK, Yamashita T,
Osaka M, Hiramatsu Y and Ohneda O: Type 2 diabetes mellitus
promotes the differentiation of adipose-derived mesenchymal stem
cells into cancer-associated fibroblasts, induced by breast cancer
cells. Stem Cells Dev. 31:659–671. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Viedma-rodríguez R, Martínez-Hernández MG,
Martínez-Torres DI and Baiza-Gutman A: Epithelial mesenchymal
transition and progression of breast cancer promoted by diabetes
mellitus in mice are associated with increased expression of
glycolytic and proteolytic enzymes. Horm Cancer. 11:170–181. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Baek KH, Choi J and Pei CZ: Cellular
functions of OCT-3/4 regulated by ubiquitination in proliferating
cells. Cancers (Basel). 12:6632020. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Zhao FQ, Misra Y, Li DB, Wadsworth MP,
Krag D, Weaver D, Tessitore J, Li DW, Zhang G, Tian Q and Buss K:
Differential expression of Oct3/4 in human breast cancer and normal
tissues. Int J Oncol. 52:2069–2078. 2018.PubMed/NCBI
|
|
78
|
Shu S, Li Z, Liu L, Ying X, Zhang Y, Wang
T, Zhou X, Jiang P and Lv W: HPV16 E6-activated OCT4 promotes
cervical cancer progression by suppressing p53 expression via
co-repressor NCOR1. Front Oncol. 12:9008562022. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Sun L, Liu T, Zhang S, Guo K and Liu Y:
Oct4 induces EMT through LEF1/b-catenin dependent WNT signaling
pathway in hepatocellular carcinoma. Oncol Lett. 13:2599–2606.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Berr AL, Wiese K, Dos Santos G, Koch CM,
Anekalla KR, Kidd M, Davis JM, Cheng Y, Hu YS and Ridge KM:
Vimentin is required for tumor progression and metastasis in a
mouse model of non-small cell lung cancer. Oncogene. 42:2074–2087.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Pearson GW: Control of invasion by
epithelial-to-mesenchymal transition programs during metastasis. J
Clin Med. 8:6462019. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Yamashita N, Tokunaga E, Kitao Y, Taketani
K, Akiyoshi S, Okada S, Aishima S, Morita M and Maehara Y: Vimentin
as a prognostic factor for triple-negative breast cancer. J Cancer
Res Clin Oncol. 139:739–746. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Chen Z, Fang Z and Ma J: Regulatory
mechanisms and clinical significance of vimentin in breast cancer.
Biomed Pharmacother. 133:1110682021. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Ahmadpour F, Igder S, Moghadam ARE,
Moradipoodeh B, Sepahdar A, Mokarram P, Fallahi J and Mohammadzadeh
G: Metformin as a potential therapeutic agent in breast cancer:
Targeting miR-125a methylation and epigenetic regulation. Int J Mol
Cell Med. 13:272–285. 2024.PubMed/NCBI
|