Introduction
Glioblastoma (GBM) is the most aggressive and among
the most common primary malignant brain tumors in adults (1). The incidence of GBM increases with age
from 0.15 per 100,000 in children compared to 15.3 per 100,000 in
aged of 75–84 years. Patients diagnosed with GBM typically die
within a few months if left untreated (2,3).
Despite extensive research over the past decades, GBM continues to
exhibit one of the poorest prognoses among malignancies (4,5). The
standard first-line treatment involves maximal surgical resection
followed by concurrent radiotherapy and temozolomide (TMZ)
chemotherapy. TMZ, the primary chemotherapeutic agent for GBM,
exerts its effect by inducing DNA alkylation, predominantly at the
N7 and O6 positions of guanine and the
N3 position of adenine or guanine. Among TMZ-induced DNA
lesions, N7-methylguanine accounts for 80–85%,
N3-methyladenine or -methylguanine constitutes 8–20%,
and O6-methylguanine (O6-MeG) represents ~8%.
The principal cytotoxic effect of TMZ therapy arises from DNA
mismatches induced by O6-MeG, leading to stalled DNA
replication and the generation of DNA double-strand breaks
(6). This severe DNA damage
activates the ataxia-telangiectasia mutated (ATM)/checkpoint kinase
2 signaling pathway, resulting in p53-mediated G2/M cell
cycle arrest (7) and ultimately
triggering tumor cell apoptosis and necrosis (8).
O6-MeG-DNA methyltransferase (MGMT) is a
DNA repair enzyme that counteracts alkylation-induced damage by
transferring the methyl group from the O6 position of
guanine to a cysteine residue within its active site, thereby
restoring the damaged guanine nucleotide (nt) (9). Elevated mRNA and protein expression
levels of MGMT are strongly associated with tumor resistance to TMZ
chemotherapy (10). MGMT gene
expression is regulated by multiple epigenetic mechanisms,
including promoter methylation and modulation by non-coding RNAs
(ncRNAs). Promoter methylation is a principal mechanism in
suppressing MGMT transcriptional activity (11,12),
thereby reducing protein levels. Clinically, the MGMT+
status typically indicates an unmethylated promoter, active gene
expression and high protein levels. By contrast, the
MGMT− status corresponds to a methylated promoter,
suppressed gene expression and minimal protein production. Patients
with MGMT− status exhibit markedly prolonged overall
survival following TMZ-based chemoradiotherapy (13). By contrast, a subset of patients
with MGMT+ status exhibits poor responsiveness to TMZ
(14,15). Since their tumors presumably retain
functional MGMT protein, this treatment failure cannot be solely
attributed to the canonical MGMT-mediated repair pathway,
indicating that mechanisms beyond MGMT status contribute to TMZ
resistance. Therefore, developing strategies to modulate MGMT
expression is key to overcoming chemoresistance in GBM in the
future.
ncRNAs have been recognized as pivotal regulators of
cellular processes and oncogenesis. These functional RNA molecules,
which do not encode proteins, primarily comprise microRNAs
(miRNAs/miR), long ncRNAs (lncRNAs) and circular RNAs (circRNAs)
(16–18). Among these molecules, miRNAs and
lncRNAs serve as key regulators within the ncRNA transcriptome,
modulating gene expression at transcriptional and
post-transcriptional levels (19),
and thus, exerting key influences on tumorigenesis, cancer
progression and acquired drug resistance (20–23).
In GBM, increasing attention has been directed toward their
regulatory effects on MGMT expression (24–28).
miRNAs can bind to MGMT mRNA, promote its degradation and induce
MGMT silencing (29,30). By contrast, lncRNAs may function as
competing endogenous RNAs (ceRNAs) by sequestering miRNAs, thereby
reducing their regulatory impact on target mRNAs and influencing
MGMT expression (31).
Advancements in next-generation high-throughput
sequencing, gene silencing and gene editing technologies have
provided notable evidence that, in addition to miRNAs and lncRNAs,
circRNAs possess regulatory functions and are associated with
various diseases, including malignancies such as GBM and lung
cancer, as well as cardiovascular conditions like atherosclerosis
(32–35). circRNAs can interact with protein
complexes, RNA molecules and DNA to regulate a broad spectrum of
physiological and pathological processes (36–38).
Based on these developments, the present review systematically
summarizes the molecular mechanisms through which distinct classes
of ncRNAs regulate MGMT gene expression (Table I; Fig.
1). The primary aim of this synthesis is to elucidate the
complex regulatory network governing MGMT expression, thereby
identifying novel ncRNA-based therapeutic targets and strategies to
overcome TMZ resistance in GBM.
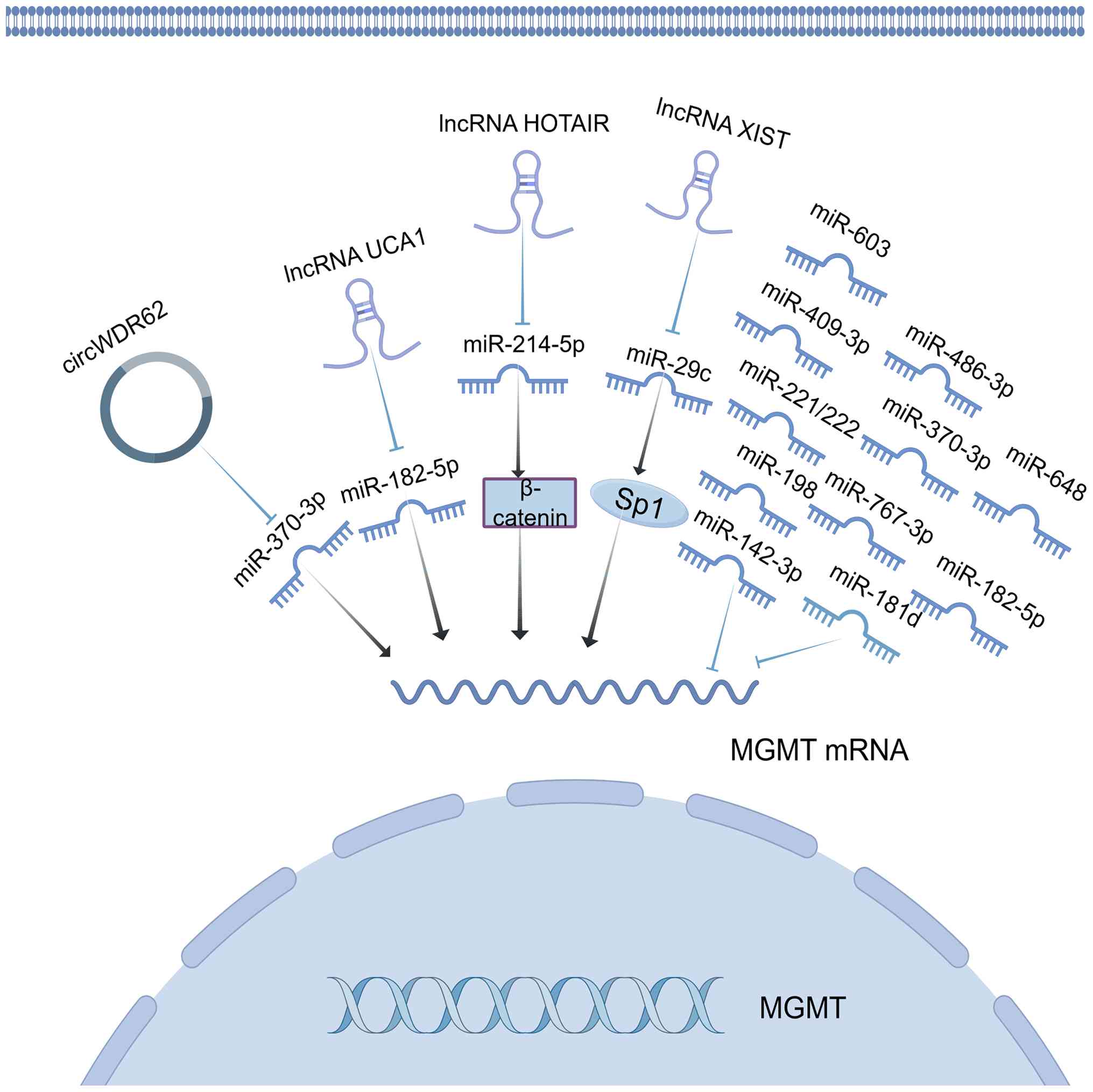 | Figure 1.Core competing endogenous RNA network
of miRNA, lncRNA and circRNA in regulating MGMT expression. Arrows
denote promotion or positive regulation, while blunt-ended arrows
represent inhibition or negative regulation. miRNA/miR, microRNA;
lncRNA, long non-coding RNA; circRNA/circ, circular RNA; MGMT,
O6-methylguanine-DNAmethyltransferase; Sp1, specificity
protein 1; HOTAIR, HOX transcript antisense RNA; UCA1, urothelial
carcinoma-associated 1; WDR62, WD repeat domain 62. |
 | Table I.Mechanisms of non-coding RNAs in
regulating MGMT expression. |
Table I.
Mechanisms of non-coding RNAs in
regulating MGMT expression.
| First author/s,
year | RNA | Mechanism in
regulating MGMT | Effect on TMZ
chemotherapy | (Refs.) |
|---|
| Kirstein et
al, 2020; Kreth et al, 2013 | miRNA | Direct targeting of
the 3′-UTR of MGMT; regulating the expression of MGMT by modulating
relevant pathways | Enhance the
chemotherapeutic sensitivity to TMZ | (29,53) |
| Nadhan et
al, 2024; Shahzad et al, 2021; Stackhouse et al,
2020 | lncRNA | As a specific
‘sponge’ of miRNA, it weakens the regulatory effect of miRNA on
mRNA; regulating the expression of MGMT through epigenetic
mechanisms | Promote
chemo-resistance to TMZ | (94,105,106) |
| Wu et al,
2024; Geng et al, 2022 | circRNA | Functioning as a
miRNA sponge, it modulates downstream MGMT expression via the
constructed circRNA-miRNA-mRNA regulatory network | Promote
chemoresistance to TMZ | (153,154) |
miRNAs
miRNAs are small ncRNA molecules 19–22 nts in length
that regulate gene expression either by sequence-specific
inhibition of mRNA translation or by promoting mRNA degradation
(29,39). As central regulators of cellular
homeostasis, miRNAs govern key cellular processes, including
proliferation, migration, cell cycle progression and apoptosis,
thereby exerting a notable influence on overall cellular function
(40,41). Dysregulated miRNA expression is
closely associated with the pathogenesis of various clinical
disorders, including cancer, neurodegenerative diseases and
cardiovascular conditions (42–45).
In GBM, aberrant miRNA expression serves as a key predictive
biomarker for tumorigenesis and disease progression (46). These miRNAs act as oncogenes or
tumor suppressors and directly modulate key pathways involved in
tumor suppression (29). For
instance, in GBM, dysregulated miRNAs such as miR-21, miR-10b and
miR-7 have been shown to modulate tumor proliferation, invasion,
and apoptosis by targeting key factors including PTEN, Rho and EGFR
(29).
miRNAs in GBM
Extensive studies have established that dysregulated
miRNA expression is closely associated with tumorigenesis and
progression across various cancer types, including lung cancer,
breast cancer, colorectal cancer and gastric cancer, and is closely
associated with tumor initiation and progression (47–50).
In GBM, aberrant miRNA expression constitutes a prominent molecular
feature. Functionally, miRNAs act as oncogenes or tumor suppressors
in GBM, modulating angiogenesis, regulating metabolic pathways and
associated enzymatic activities, and influencing the
differentiation of glioma stem cells (GSCs) (51). Previous studies have indicated that
miRNAs regulate ~3% of glioma-related genes and 30% of
protein-coding genes, with individual miRNAs capable of modulating
the expression levels of up to 100 distinct mRNAs implicated in GBM
(41), highlighting their pivotal
role in GBM pathogenesis (51).
Comprehensive analysis of miRNA expression patterns may facilitate
the identification of numerous potential therapeutic agents,
establishing miRNAs as notable candidates for GBM treatment.
miRNAs regulating MGMT expression
The methylation status of the MGMT promoter is a key
determinant of therapeutic efficacy in GBM (52). Beyond promoter methylation, MGMT
protein expression is also regulated through additional mechanisms,
including post-transcriptional control. Among these, miRNA-mediated
regulation via the mRNA 3′-untranslated region (3′-UTR) serves a
key role (53). Kreth et al
(53) identified two distinct MGMT
transcript variants in GBM with differing 3′-UTR lengths (53). In normal brain tissue, only a
shorter transcript of ~440 bp is expressed, containing a canonical
poly(A) signal and a 105 nt 3′-UTR. By contrast, in GBM a longer
transcript of ~850 bp is specifically expressed, which harbors an
alternative poly(A) signal spanning 522 nt. In the study by Kreth
et al (53), the analysis of
patient samples revealed a notable negative association between
MGMT expression and 3′-UTR length, with normal-length 3′-UTRs
associated with reduced MGMT expression. This differential
expression is closely associated with the sequence architecture of
the 3′-UTR. Bioinformatics analyses indicated that the longer
3′-UTR contained a markedly higher density of potential
miRNA-binding sites, rendering it more susceptible to
miRNA-mediated post-transcriptional degradation. These findings
provided a mechanistic perspective in understanding the
heterogeneity of MGMT expression in GBM. The following section
systematically reviews specific miRNAs that directly or indirectly
target MGMT expression in GBM (Table
II).
 | Table II.miRNAs involved in the regulation of
MGMT in glioblastoma. |
Table II.
miRNAs involved in the regulation of
MGMT in glioblastoma.
| First author/s,
year | miRNA | Expression | Regulation | Type | Molecular
function | (Refs.) |
|---|
| Lee et al,
2018 | miR-142-3p | Downregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | MGMT protein
Inhibits expression; enhances TMZ sensitivity | (54) |
| Zhang et al,
2012; Khalil et al, 2016 | miR-181d | Downregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | Reduces MGMT mRNA
and protein expression; enhances TMZ sensitivity | (56,58) |
| Cheng et al,
2022 | miR-182-5p | Downregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | Inhibits MGMT
protein expression; enhances TMZ sensitivity | (61) |
| Nie et al,
2017 | miR-198 | Downregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | Inhibits MGMT
protein expression; enhances TMZ sensitivity | (64) |
| Quintavalle et
al, 2013 | miR-221/222 | Upregulated | Directly targets
the 3′-UTR of MGMT | Oncogenic | Reduces MGMT mRNA
and protein expression; enhances TMZ sensitivity | (65) |
| Zhou et al,
2022 | miR-214-5p | Downregulated | Targets β-catenin
to indirectly regulate MGMT expression | Tumor
suppressor | Inhibits MGMT
protein expression; enhances TMZ sensitivity | (73) |
| Xiao et al,
2016 | miR-29c | Downregulated | Targets Sp1 to
indirectly regulate MGMT expression | Tumor
suppressor | Inhibits MGMT
protein expression; enhances TMZ sensitivity | (76) |
| Gao et al,
2016; Geng et al, 2022 | miR-370-3p | Downregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | Reduces MGMT mRNA
and protein expression; enhances TMZ sensitivity | (81,154) |
| Khalil et
al, 2016 | miR-409-3p | Upregulated | Directly targets
the 3′-UTR of MGMT | Oncogenic | Reduces MGMT mRNA
and protein expression; enhances TMZ sensitivity | (58) |
| Wu et al,
2020 | miR-486-3p | Downregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | Inhibits MGMT
protein expression; enhances TMZ sensitivity | (90) |
| Kushwaha et
al, 2014 | miR-603 | Upregulated | Directly targets
the 3′-UTR of MGMT | Tumor suppressor
and oncogenic | Reduces MGMT mRNA
and protein expression; enhances TMZ sensitivity | (59) |
| Kreth et al,
2013 | miR-648 | Upregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | Inhibits MGMT
protein expression; enhances TMZ sensitivity | (53) |
| Kreth et al,
2013 | miR-767-3p | Upregulated | Directly targets
the 3′-UTR of MGMT | Tumor
suppressor | Degradation of MGMT
mRNA; enhances TMZ sensitivity | (53) |
miR-142-3p
In GBM cell lines, Lee et al (54) demonstrated a marked inverse
association between MGMT expression and miR-142-3p levels. Elevated
MGMT expression was associated with low miR-142-3p levels, whereas
low MGMT expression was associated with high miR-142-3p levels
(54). Overexpression of miR-142-3p
did not alter MGMT mRNA levels but markedly decreased protein
expression, suggesting a direct regulatory interaction with the
MGMT 3′-UTR. This mechanism was subsequently confirmed using a
luciferase reporter assay (54). In
chemotherapy sensitivity assays, cells overexpressing miR-142-3p
exhibited markedly enhanced sensitivity to alkylating agents,
including TMZ and carmustine (BCNU), with the most pronounced
effect observed in the BCNU-treated group (54). These findings underscored the key
role of miR-142-3p in regulating MGMT expression and enhancing
chemosensitivity, positioning it as a potential biomarker and
therapeutic target in GBM.
Additionally, a previous study has demonstrated that
the pro-oncogenic cytokine interleukin-6 suppressed miR-142-3p
expression, further supporting its role as a tumor suppressor
(55). In summary, miR-142-3p
enhances the sensitivity of GBM cells to alkylating chemotherapy by
directly targeting the MGMT 3′-UTR and inhibiting its protein
translation. This mechanism establishes a clear target for the
development of personalized therapeutic strategies in GBM.
miR-181d
Zhang et al (56) established a miRNA-mediated
regulatory mechanism for MGMT and demonstrated that miR-181d binds
directly to the long 3′-UTR of MGMT, thereby suppressing its
expression at the post-transcriptional level (53,56).
In vitro, transfection of miR-181d mimics markedly reduced
both MGMT mRNA and protein levels, and sensitized GBM cells to TMZ
treatment (56). Clinical analyses
revealed a notable association between elevated miR-181d expression
and improved patient survival (56), indicating its potential as a
favorable prognostic indicator. Furthermore, miR-181d expression is
increased following TMZ monotherapy, radiotherapy or combination
therapy, with the most pronounced elevation observed following
combination treatment (57). These
findings suggest that miR-181d may serve as a predictive biomarker
in evaluating the efficacy of both chemotherapy and
radiotherapy.
Subsequent studies have confirmed the direct
regulatory effect of miR-181d on MGMT and elucidated its central
role within a coordinated regulatory network. This regulatory role
was established through systematic genome-wide microarray screening
and analysis of clinical datasets. Notably, miR-181d can cooperate
with other miRNAs, including miR-409-3p (58), miR-603 (59), miR-648 (60) and miR-661 (53), forming a synergistic regulatory
network that modulates MGMT expression. Khalil et al
(58) demonstrated a pronounced
synergistic negative regulatory pattern between miR-181d and
miR-409-3p in MGMT+ GBM. Functional assays revealed that
overexpression of either miRNA mimic in T98G cells reduced MGMT
expression, whereas their combined application produced a marked
synergistic enhancement. Furthermore, analysis of two major
independent datasets, The Cancer Genome Atlas and the Chinese
Glioma Genome Atlas, provided key insights: miR-181d was the only
miRNA to exhibit a notable inverse association with MGMT mRNA
levels; however, combination with miR-409-3p markedly improved the
prediction of MGMT expression. These findings indicated that, in
the complex in vivo environment, MGMT expression is governed
by a precise regulatory network centered on miR-181d with
cooperative contributions from additional miRNAs (58).
In summary, miR-181d functions as a
tumor-suppressive miRNA and directly suppresses MGMT by targeting
its mRNA, thereby reducing transcript and protein levels. Acting as
a central hub within a cooperative regulatory network, miR-181d
integrates signals from other miRNAs to precisely modulate MGMT
expression. This coordinated regulation enhances the
chemosensitivity of GBM cells to TMZ. These findings enhance the
understanding of the regulatory network governing MGMT, and
establish miR-181d and its associated network as promising
biomarkers with therapeutic potential in GBM.
miR-182-5p
Cheng et al (61) demonstrated that miR-182-5p was
downregulated in high-grade GBM tissues and inversely associated
with MGMT protein levels. Mechanistic investigation revealed that
miR-182-5p directly binds to the 3′-UTR of MGMT mRNA, inhibiting
MGMT protein expression via a post-transcriptional mechanism
without altering mRNA levels. The specificity of this interaction
was confirmed using a dual-luciferase reporter assay. Functional
experiments indicated that transfection of miR-182-5p mimics
markedly enhanced the chemosensitivity of GBM cells to TMZ, as
evidenced by reduced cell viability and increased apoptosis. This
effect was closely associated with MGMT protein downregulation and
a concomitant increase in DNA damage markers, such as γ-H2A histone
family member X and ATM (61).
Beyond MGMT, miR-182-5p targets additional genes, such as
neuropilin-1, and through this targeting, it further inhibits
GBM-cell proliferation and migration (62).
In summary, miR-182-5p exhibits multifaceted
tumor-suppressive functions in GBM. The findings highlighted its
potential as a therapeutic target, providing novel avenues for GBM
treatment. Future research should investigate its translational
potential and assess its efficacy in combination with existing
therapies to potentially improve clinical outcomes and prognosis
for patients with GBM in the future.
miR-198
Previous research has indicated that miR-198 is
expressed at low levels in GBM tissue, and inversely associated
with tumor malignancy and progression (63). Analysis of clinical samples further
revealed that GBM tissues with high MGMT expression typically
exhibit low miR-198 levels. Reduced miR-198 expression was strongly
associated with poor patient prognosis. Overexpression of miR-198
markedly enhanced the chemosensitivity of GBM cells to TMZ both
in vitro and in vivo, highlighting its pivotal role
in regulating the chemotherapeutic response. Mechanistic
experiments using a luciferase reporter assay confirmed that
miR-198 directly binds to the 3′-UTR of MGMT mRNA, thereby
inhibiting MGMT protein expression. Rescue experiments demonstrated
that MGMT overexpression effectively reversed the TMZ-sensitizing
effect mediated by miR-198, establishing a complete and functional
‘miR-198/MGMT/TMZ sensitivity’ regulatory axis (64).
In summary, miR-198 acts as a tumor-suppressive
miRNA, enhancing TMZ chemosensitivity by directly targeting MGMT
mRNA and inhibiting its translation. This finding elucidated a key
molecular mechanism underlying GBM chemoresistance and provides a
theoretical foundation for the development of precision therapeutic
strategies targeting the miR-198/MGMT axis.
miR-221/222
In GBM and hepatocellular carcinoma (HCC) (65,66),
miR-221/222 exerts a complex, dual-functional regulatory effect on
MGMT. In GBM, Quintavalle et al (65) demonstrated that miR-221/222 directly
targets the 3′-UTR of MGMT mRNA, suppressing its protein expression
at the post-transcriptional level, a finding confirmed by
luciferase reporter assays. Furthermore, investigations across
multiple GBM and melanoma cell lines revealed that miR-221/222
overexpression markedly downregulated MGMT mRNA and protein levels.
Functionally, miR-221/222 overexpression enhanced GBM cell
sensitivity to TMZ and promoted apoptosis. However, the resulting
impairment of MGMT-mediated DNA repair led to the accumulation of
DNA damage markers, which may foster genomic instability and
contribute to tumor progression (65).
This regulation is neither unidirectional nor devoid
of potential adverse effects. In hepatocellular carcinoma, Chen
et al (66) confirmed that
miR-221-3p, the primary functional isoform of miR-221, directly
targets MGMT. Inhibition of MGMT expression enhanced the
proliferative, migratory, invasive and clonogenic capacities of HCC
cells, while suppressing apoptosis. The study validated the binding
site using bioinformatics prediction and dual-luciferase reporter
assays, and demonstrated, via rescue experiments, that MGMT
overexpression reversed the oncogenic effects of miR-221-3p. These
findings established the pivotal role of the miR-221/MGMT axis in
HCC pathogenesis (66).
Although miR-221/222 can enhance tumor sensitivity
to alkylating agents such as TMZ by downregulating MGMT, it is
highly expressed in various cancer types, including GBM, HCC,
colorectal cancer and cervical cancer, and exhibits
well-established oncogenic properties (65–68).
For instance, miR-221/222 promotes malignant progression by
targeting multiple tumor suppressors, including PTEN, p27 and
tissue inhibitor of metalloproteinase 3 (69,70).
This duality between enhancing chemosensitivity and driving tumor
progression indicates a potential ‘double-edged sword’ effect of
miR-221/222 in clinical applications. While its transient
expression may improve the initial chemotherapy response rate,
sustained high expression could ultimately result in poor prognosis
by impairing DNA repair mechanisms and promoting tumor
invasiveness.
In summary, miR-221/222 is a well-characterized
oncomiR, frequently upregulated across various cancer types and
strongly associated with poor patient prognosis. In GBM,
miR-221/222 enhances tumor cell sensitivity to TMZ chemotherapy by
directly targeting MGMT mRNA and suppressing its protein
expression. While this finding provides a novel approach to
overcoming TMZ resistance, it also underscores the dual role of
miR-221/222. Future research should systematically elucidate the
dynamic roles of miR-221/222 across distinct pathological stages
and molecular contexts, and carefully assess its suitability as a
therapeutic target, including delivery methods and potential risks
in combination strategies.
miR-214-5p
miR-214-5p functions as a tumor-suppressive miRNA
that exerts notable effects in multiple malignancies, including
cervical and prostate cancers, by inhibiting key oncogenic
phenotypes, including cell proliferation, migration and invasion
(71,72). In GBM, a key regulatory mechanism
involving miR-214-5p was revealed: Its overexpression markedly
downregulates β-catenin and MGMT expression, whereas miR-214-5p
inhibition produces the opposite effect. These findings underscore
the pivotal regulatory role of miR-214-5p in GBM (73).
Cyanidin-3-O-glucoside (C3G), a natural flavonoid
widely distributed in the plant kingdom, exhibits notable antitumor
potential. Previous studies have indicated that C3G enhanced the
chemosensitivity of GBM cells to TMZ and markedly upregulated
miR-214-5p expression (73). Zhou
et al (73) further
demonstrated that C3G upregulated miR-214-5p, which directly binds
to the 3′-UTR of catenin β-1, the gene encoding β-catenin, thereby
suppressing β-catenin protein expression. As a key transcriptional
co-activator in the Wnt signaling pathway, β-catenin directly
initiates MGMT transcription. Therefore, miR-214-5p indirectly
attenuates MGMT transcription by regulating β-catenin, thereby
decreasing protein expression. Functional experiments further
revealed that C3G and miR-214-5p enhanced TMZ-induced apoptosis in
GBM cells, an effect reversed by a miR-214-5p inhibitor, indicating
that the chemosensitizing action of C3G was highly dependent on
miR-214-5p (73).
In summary, miR-214-5p functions as a
tumor-suppressive miRNA in GBM, enhancing TMZ chemosensitivity by
targeting the β-catenin/MGMT signaling axis. The natural
small-molecule compound C3G, which activates this pathway, exhibits
potential therapeutic value in reversing TMZ resistance in GBM,
potentially providing a theoretical basis for the development of
adjuvant anti-glioma agents.
miR-29c
miR-29 is frequently downregulated in most solid
tumors, including breast cancer, lung cancer and glioblastoma,
where it exerts notable tumor-suppressive activity and is
associated with a favorable patient prognosis (74–76).
Functional heterogeneity exists within the miR-29 family.
Specifically, miR-29c enhances TMZ chemosensitivity in GBM via dual
mechanisms. miR-29c indirectly suppresses MGMT transcriptional
activity by targeting the transcription factor specificity protein
1 (Sp1) (76,77) and restores the expression of
multiple tumor suppressor genes via demethylation (78), collectively enhancing
chemosensitivity. By contrast, miR-29b exhibits oncogenic
properties in certain contexts and may promote tumorigenesis by
repressing tumor suppressor genes, such as PTEN (79). This functional divergence
underscores the complexity of miRNA regulatory networks, in which
the net biological effect is determined by the relative influence
and interactions among their targets within a particular cellular
context. In GBM models of TMZ resistance, the tumor-suppressive
function of miR-29c, mediated through the Sp1/MGMT axis,
predominates over its potential oncogenic effects observed in other
biological settings (79).
Mechanistically, Xiao et al (76) systematically delineated the
regulatory pathway of miR-29c in GBM. This miRNA does not directly
target MGMT mRNA; instead, it binds specifically to the 3′-UTR of
the transcription factor Sp1, thereby inhibiting Sp1 protein
translation. This suppression reduces Sp1-mediated transcriptional
activation of the MGMT promoter, ultimately leading to decreased
MGMT protein expression (76). Sp1,
a ubiquitously expressed zinc-finger transcription factor present
in virtually all mammalian cells and tissues (80), serves a key role in sustaining MGMT
transcription. Therefore, the identification of the
‘miR-29c/Sp1/MGMT’ axis not only enhances the current understanding
of chemoresistance mechanisms in GBM but also potentially provides
a foundation for the development of novel therapeutic
strategies.
In summary, miR-29c acts as a tumor-suppressive
miRNA in GBM, enhancing TMZ sensitivity by indirectly regulating
MGMT expression via Sp1 targeting. This mechanism provides a novel
perspective on chemoresistance in GBM and establishes a strong
theoretical basis for the development of miRNA-based adjuvant
therapeutic strategies.
miR-370-3p
Gao et al (81) reported that miR-370-3p possesses
potent tumor-suppressive properties in GBM. miR-370-3p is markedly
downregulated in low-grade (WHO grade II) and high-grade (WHO grade
IV) gliomas, as well as in GBM cell lines (82). Overexpression of miR-370-3p markedly
suppresses tumor cell viability, reduces proliferation, and
decreases the proportion of cells in the S and G2/M
phases of the cell cycle (81).
A negative association has been identified between
MGMT and miR-370-3p expression (81). Mechanistically, miR-370-3p directly
targets the 3′-UTR of MGMT mRNA, thereby downregulating its protein
expression. In vitro, miR-370-3p upregulation markedly
reduced MGMT mRNA and protein levels, enhancing GBM cell
sensitivity to TMZ and increasing cell mortality. This direct
regulatory relationship was confirmed using a luciferase reporter
assay, in which mutation of the MGMT 3′-UTR binding site abolished
the effect of miR-370-3p, indicating sequence-specific action.
In vivo, combined administration of TMZ and miR-370-3p
inhibited subcutaneous xenograft tumor growth, and analysis of the
resected tumor tissues revealed notable downregulation of MGMT
expression, further supporting the effectiveness of this regulatory
pathway (83). Nadaradjane et
al (83) proposed monitoring
miR-370-3p levels during standard treatment, noting that a single
measurement of its expression was not associated with overall
survival. Notably, patients exhibiting sustained high miR-370-3p
expression prior to disease recurrence exhibited prolonged survival
(83).
In summary, miR-370-3p functions as a
tumor-suppressive miRNA in GBM by downregulating MGMT expression at
the mRNA and protein levels, thereby enhancing tumor sensitivity to
TMZ chemotherapy. Although miR-370-3p is not a key determinant of
overall patient survival, it represents a potential strategy in
increasing TMZ sensitivity in patients with MGMT+
GBM.
miR-409-3p
Khalil et al (58) reported that miR-409-3p expression
was upregulated 4–5-fold in GBM samples compared with healthy brain
tissues, and was negatively associated with MGMT expression.
Specifically, miR-409-3p levels were elevated in samples with low
MGMT expression but reduced in those with high MGMT expression.
Experimental validation demonstrated that transfection of
miR-409-3p mimics into MGMT-overexpressing GBM cells markedly
downregulated MGMT mRNA and protein levels, enhancing cellular
sensitivity to TMZ. These results indicated that miR-409-3p
suppressed MGMT expression by promoting mRNA degradation and
inhibiting translation (58).
Notably, co-transfection of miR-409-3p mimics with the key
regulator miR-181d produced a synergistic effect, suggesting that
miR-409-3p functions as a co-regulatory factor alongside miR-181d,
jointly modulating MGMT expression and influencing GBM response to
alkylating agent chemotherapy (58).
In summary, miR-409-3p upregulation in human GBM
samples suppresses MGMT transcription and protein expression. This
property indicates the potential of miR-409-3p as a therapeutic
strategy to enhance the responsiveness of patients with
MGMT+ GBM to alkylating chemotherapy agents; however,
the upstream mechanisms driving miR-409-3p upregulation in GBM
remain to be elucidated. Future studies should employ integrated
approaches, including chromatin immunoprecipitation, protein-RNA
interaction assays, and in vitro and in vivo rescue
experiments, to comprehensively elucidate its underlying molecular
mechanisms. These investigations could potentially establish a
robust foundation for subsequent clinical development.
miR-486-3p
miR-486 is a key tumor-suppressive miRNA that is
frequently downregulated in numerous malignant tumors (84,85).
Dong et al (86) reported
that upregulation of its family member miR-486-3p markedly enhanced
the sensitivity of neuronal tumors to chemotherapeutic agents.
Allicin (diallyl trisulfide), the principal bioactive constituent
of garlic, is widely utilized as a dietary supplement and has
garnered increasing research attention for its notable antitumor
activity (87). Evidence indicated
that allicin exerts antitumor effects by inducing tumor cell
apoptosis and inhibiting proliferation, among other mechanisms
(88). Allicin has exhibited
promising therapeutic potential in multiple malignancies, including
GBM (89).
In GBM cells, allicin treatment markedly increases
miR-486-3p expression, as reported by Wu et al (90). Mechanistic analyses indicated that
miR-486-3p directly binds to the 3′-UTR of MGMT mRNA, thereby
inhibiting its translation, and thus, reducing MGMT protein levels.
Functionally, miR-486-3p overexpression markedly augmented
TMZ-induced DNA damage and apoptosis and specifically reversed
chemoresistance in TMZ-resistant cell lines. Notably, this effect
was absent in MGMT− GBM cell lines. Furthermore, rescue
assays confirmed that MGMT overexpression effectively reversed the
TMZ-sensitizing activity mediated by miR-486-3p, thereby validating
the proposed regulatory axis. In vivo, the combined
administration of allicin and TMZ markedly prolonged the survival
of treated mice, with a more notable effect observed in the
miR-486-3p overexpression group (90).
In summary, allicin enhances GBM sensitivity to TMZ
chemotherapy by upregulating miR-486-3p, which subsequently
inhibits MGMT protein translation through targeted repression. This
mechanism provides a molecular rationale for the combined
therapeutic strategy of ‘allicin + TMZ’ and identifies miR-486-3p
as a potential target in improving GBM treatment efficacy.
miR-603
Guo et al (91) reported that miR-603 expression was
upregulated in GBM tissues. The study further demonstrated that
miR-603 promoted malignant tumor progression by enhancing tumor
cell proliferation and accelerating cell cycle progression.
Mechanistic investigations revealed that miR-603 directly targets
the tumor suppressor genes Wnt inhibitory factor 1 and catenin
β-interacting protein 1, thereby activating the Wnt/β-catenin
signaling pathway, which subsequently augments the proliferative
and migratory capacities of tumor cells (91).
However, genome-wide screening and clinical data
analysis conducted by Kushwaha et al (59) revealed a more complex, dual
regulatory role for miR-603 in GBM. In addition to its oncogenic
functions, miR-603 mediates post-transcriptional repression of MGMT
by binding specifically to the 3′-UTR of its mRNA (59). In vitro experiments confirmed
that miR-603 overexpression markedly decreased MGMT mRNA and
protein levels and enhanced GBM cell sensitivity to TMZ
chemotherapy. This effect was further validated in animal models
(59). Notably, the study
demonstrated a notable synergistic interaction between miR-603 and
miR-181d in suppressing MGMT expression, wherein their combined
application produced a markedly stronger inhibitory effect compared
with that of either miRNA alone (59).
Based on these findings, miR-603 represents a
therapeutic target with a dual regulatory mechanism. miR-603
enhances chemosensitivity by suppressing MGMT, while promoting
malignant tumor progression through the Wnt/β-catenin signaling
pathway. Abate et al (92)
developed an innovative self-assembling nanoparticle (SANP)-based
co-delivery system capable of simultaneously loading miR-603 and
miR-221 to achieve efficient MGMT suppression. This delivery system
markedly improved miRNA transport across the blood-brain barrier.
In an orthotopic GBM model, it effectively reversed TMZ resistance,
providing a novel nano-therapeutic strategy to overcome
chemoresistance in GBM (92).
In summary, the SANP platform exhibits therapeutic
potential for brain diseases by enabling efficient miRNA delivery.
Notably, the present review underscores the role of miRNAs in
overcoming chemoresistance in cancer therapy, highlighting the
prospect of developing more effective combination antitumor
therapies.
miR-648 and miR-767-3p
Kreth et al (53) identified a key regulatory mechanism
for MGMT in GBM beyond promoter methylation: Post-transcriptional
regulation mediated by miRNA targeting of an elongated 3′-UTR. The
study emphasized the key roles of miR-648 and miR-767-3p in this
regulation. Although both miRNAs target the long 3′-UTR isoform of
MGMT, their mechanisms of action differ markedly.
Both miR-767-3p and miR-648 are upregulated in GBM
tissue. Co-transfection of these miRNAs enhanced GBM
chemosensitivity to TMZ (53).
Mechanistic investigation revealed that miR-767-3p binds to the
long 3′-UTR isoform of MGMT mRNA, inducing its degradation and
thereby suppressing MGMT expression at the post-transcriptional
level. By contrast, miR-648 predominantly inhibits MGMT protein
expression at the translational stage without markedly reducing
mRNA levels (53). However,
Kushwaha et al (59)
reported that the effect of miR-648 transfection on MGMT expression
was inconsistent, reflecting its highly context-dependent nature,
which may arise from variable ratios of long and short MGMT 3′-UTR
isoforms across different cellular models. The specific isoform
ratio directly influences the accessibility of the target site for
miR-648 (59).
In summary, miR-767-3p suppresses MGMT expression
via mRNA degradation, whereas miR-648 primarily exerts its effect
through translational inhibition. Both miRNAs are upregulated in
GBM and their co-transfection markedly enhances GBM sensitivity to
TMZ. This synergistic mechanism establishes a novel molecular
foundation and therapeutic approach in reversing TMZ resistance.
Nonetheless, its clinical translational potential requires
validation in larger sample cohorts and in vivo models.
lncRNAs
lncRNAs are a class of ncRNA molecules >200 nts
in length. Initially regarded as transcriptional ‘noise’ lacking
notable biological function (93),
lncRNAs are now established as key regulatory factors, a
recognition driven by advances in transcriptomics. In cancer
biology, lncRNAs orchestrate core cellular processes in tumor cells
through complex regulatory networks, including proliferation,
apoptosis and metabolic reprogramming, as well as key pathological
processes such as epithelial-mesenchymal transition, migration,
invasion, metastasis, maintenance of cancer stem cell properties
and chemoresistance (94). These
diverse functions position lncRNAs as integral components of cancer
gene regulatory networks, establishing them as key nodes that
determine malignant phenotypes (95). lncRNAs are dysregulated across
various tumor types, including breast, prostate, lung and
colorectal cancers and glioblastoma. Their aberrant expression can
exert either oncogenic or tumor-suppressive roles, driving tumor
progression through remodeling of the cancer epigenome (94). Several studies have indicated that
dysregulated lncRNAs serve a key regulatory role in the initiation
and progression of GBM (96–99).
Therefore, lncRNAs hold promise both as biomarkers for GBM
diagnosis and prognosis and as novel therapeutic targets.
lncRNAs in GBM
Dysregulated lncRNA expression is closely associated
with the initiation, progression and therapeutic resistance of GBM
(100). Mechanistically, lncRNAs
influence malignant behaviors in GBM through multiple strategies,
including epigenetic modifications, ceRNA network regulation,
signaling pathway intervention and tumor microenvironment
remodeling. Functionally, lncRNAs serve a dual role in GBM, acting
both as oncogenic promoters and tumor suppressors (101). Accordingly, lncRNAs exert
essential regulatory effects on key pathological processes,
including tumor proliferation, invasion, maintenance of stemness,
metabolic reprogramming and immune evasion (102,103). These findings highlight the
potential of lncRNAs as novel biomarkers for early diagnosis,
prognostic evaluation and monitoring of therapeutic response in
GBM. Furthermore, their roles in GBM tumorigenesis, progression and
metastasis are influenced not only by altered expression levels but
also by specific genomic localization (104). lncRNAs participate in complex
regulatory networks of GBM by modulating the expression levels of
proximal or distal coding genes through cis- or trans-regulatory
mechanisms (104).
This mechanistic insight enhances the current
understanding of the functional complexity of lncRNAs in GBM
pathogenesis and establishes a theoretical basis for the
development of novel therapeutic strategies targeting these
molecules. In summary, lncRNAs serve a key role in GBM pathogenesis
and progression.
lncRNAs regulating MGMT
expression
In GBM, lncRNAs regulate MGMT expression through
multiple molecular mechanisms, thereby influencing tumor
sensitivity to TMZ chemotherapy. The principal mechanism involves
lncRNAs functioning as ceRNAs, molecular sponges that sequester
specific miRNAs, thereby relieving miRNA-mediated inhibition of
MGMT and indirectly upregulating its expression. Alternatively,
certain lncRNAs modulate MGMT expression by epigenetically
regulating its promoter. These lncRNAs recruit chromatin-modifying
complexes to the MGMT locus, altering histone modification patterns
or DNA methylation, thereby promoting chromatin remodeling and
directly regulating MGMT transcription (94,105).
Furthermore, lncRNAs can indirectly modulate MGMT by regulating
upstream signaling pathways, such as the c-Met/AKT and NF-κB
signaling pathways. By interfering with these signaling networks,
which serve as key regulatory hubs for MGMT expression, lncRNAs
exert precise control over MGMT expression (106).
Collectively, these mechanisms demonstrate that
lncRNAs influence MGMT expression through multi-layered regulatory
networks, thereby contributing to chemoresistance in GBM. Targeting
these relevant lncRNAs may provide a novel strategy to reverse TMZ
resistance. The following section systematically reviews lncRNAs
currently identified as regulators of MGMT expression (Table III).
 | Table III.lncRNAs involved in the regulation of
MGMT in glioblastoma. |
Table III.
lncRNAs involved in the regulation of
MGMT in glioblastoma.
| First author/s,
year | lncRNA | Expression | Regulation | Type | Molecular
function | (Refs.) |
|---|
| Cheng et al,
2022 | UCA1 | Upregulated | Targeting
miR-182-5p can indirectly regulate MGMT expression | Oncogenic | Regulates MGMT
expression; enhances TMZ resistance | (61) |
| Lan et al,
2024 | HOTAIR | Upregulated | Activates the
miR-214/β-catenin/MGMT pathway | Oncogenic | Regulates MGMT
expression; enhances TMZ resistance | (115) |
| Du et al,
2017 | XIST | Upregulated | Activates the
miR-29c/Sp1/MGMT pathway | Oncogenic | Regulates MGMT
expression; enhances TMZ resistance | (118) |
| Wu et al,
2019 | TALC | Upregulated | Competitively binds
to miR-20b-3p and activates the c-Met/STAT3/p300 signaling axis,
promotes acetylation in the MGMT promoter region | Oncogenic | Regulates MGMT
expression; enhances TMZ resistance | (122) |
| Xin et al,
2023 | PRADX | Upregulated | PRADX and EZH2
binding promotes methylation in the MGMT promoter region | Oncogenic | Regulates MGMT
expression; enhances TMZ resistance | (125) |
lncRNA urothelial carcinoma-associated
1 (UCA1)
lncRNA UCA1 is a primate-specific lncRNA transcribed
in human trophoblasts, where it promotes the proliferation of
trophoblast stem cells. The ectopic expression of lncRNA UCA1
impairs trophoblast syncytialization and is associated with
activation of the interferon signaling pathway (107). As a well-established oncogene,
UCA1 regulates key biological processes in various tumors,
including cell proliferation, metastasis and apoptosis (108–110). In high-grade GBM tissues and
cells, MGMT and lncRNA UCA1 are upregulated. Knockdown of lncRNA
UCA1 or upregulation of miR-182-5p enhances GBM sensitivity to TMZ,
characterized by reduced cell viability, increased apoptosis,
decreased MGMT protein levels and accumulation of DNA damage
markers. Notably, inhibition of miR-182-5p reverses the sensitizing
effect of lncRNA UCA1 knockdown, restores MGMT expression and
diminishes TMZ efficacy (61).
Mechanistic studies have indicated that miR-182-5p directly targets
both lncRNA UCA1 and the 3′-UTR of MGMT mRNA, thereby negatively
regulating MGMT expression (61).
Furthermore, miR-182-5p regulates C-X-C motif chemokine ligand 14
secretion, glycolysis and tumor invasion within glioma-associated
stromal cells (111), and
contributes to GBM proliferation and migration (112). Collectively, these findings
establish the central role of the lncRNA UCA1/miR-182-5p/MGMT
regulatory axis in mediating chemoresistance in GBM.
In summary, miR-182-5p expression is negatively
associated with lncRNA UCA1 and MGMT levels, whereas lncRNA UCA1
and MGMT exhibit a notable positive association at the tissue
level. Mechanistic studies have indicated that lncRNA UCA1
sequesters miR-182-5p, thereby alleviating its post-transcriptional
suppression of MGMT (61). This
process results in upregulation of MGMT protein, enhanced DNA
repair capacity and promotion of TMZ resistance (61). These findings underscore the complex
regulatory role of lncRNAs in tumor drug resistance and establish a
theoretical foundation for the development of combination therapies
targeting lncRNA UCA1 or miR-182-5p.
lncRNA HOX transcript antisense RNA
(HOTAIR)
HOTAIR is a well-characterized oncogenic lncRNA.
lncRNA HOTAIR is minimally expressed in normal brain tissues but is
upregulated in human GBM tissues and TMZ-resistant GBM cells
(113). Exosome-derived lncRNA
HOTAIR from serum has been reported to promote TMZ resistance in
GBM cells, whereas knockdown of lncRNA HOTAIR enhances TMZ
sensitivity both in vivo and in vitro (114). Clinical data further indicate that
elevated lncRNA HOTAIR expression is associated with poor response
to TMZ therapy in patients with GBM (115). Functional experiments demonstrated
that modulating lncRNA HOTAIR expression directly impacted TMZ
sensitivity in GBM cells: Knockdown enhanced cytotoxicity, while
overexpression induced a chemoresistant phenotype. lncRNA HOTAIR is
among the most notably upregulated lncRNAs in GBM cell lines
compared with parental human GSCs, underscoring its key role in GBM
malignancy and potential clinical relevance (116).
Mechanistic studies have demonstrated that lncRNA
HOTAIR functions as a ceRNA by sequestering miR-214-3p, thereby
relieving its post-transcriptional inhibition of β-catenin and
activating the Wnt/β-catenin signaling pathway (115). This pathway is markedly enriched
in patients with high lncRNA HOTAIR expression and is closely
associated with poor prognosis and TMZ resistance in GBM. Upon
activation, β-catenin translocates into the nucleus and acts as a
transcriptional co-activator, directly binding to the lymphoid
enhancer factor/T cell factor sites within the MGMT promoter to
promote MGMT transcription (115).
This β-catenin-mediated transcriptional activation results in
elevated MGMT protein levels and confers resistance to TMZ.
Experimental validation confirmed this regulatory axis: Knockdown
of lncRNA HOTAIR decreased β-catenin and MGMT levels, whereas
treatment with the β-catenin agonist SKL2001 reversed the
TMZ-sensitizing effect of lncRNA HOTAIR silencing (115,117). Furthermore, lncRNA HOTAIR
expression is notably negatively associated with MGMT promoter
methylation, with patients exhibiting promoter methylation
typically displaying lower lncRNA HOTAIR levels (115). These findings suggest that lncRNA
HOTAIR may participate in the MGMT regulatory network by modulating
the epigenetic landscape, although the precise underlying mechanism
remains to be elucidated (115).
In summary, upregulation of lncRNA HOTAIR
constitutes a notable risk factor for TMZ resistance in GBM. The
regulation of the miR-214/β-catenin/MGMT pathway via ceRNA activity
by lncRNA HOTAIR represents a central mechanism mediating this
chemoresistance. However, the regulation of MGMT by lncRNA HOTAIR
extends beyond a single pathway. Key areas for further
investigation include its crosstalk with epigenetic regulatory
networks, context-dependent functions across distinct GBM molecular
subtypes and its translational potential as a therapeutic
target.
lncRNA X-inactive specific transcript
(XIST)
The lncRNA XIST is upregulated in multiple
malignancies, including GBM (118), ovarian cancer (119) and non-small cell lung cancer
(120), where it functions as an
oncogene. In GBM, elevated lncRNA XIST expression is closely
associated with larger tumor volume, advanced World Health
Organization grade and reduced overall survival, underscoring its
potential as a biomarker of poor prognosis (121). Functional experiments have
indicated that lncRNA XIST knockdown effectively inhibits GBM cell
migration, invasion and proliferation, while promoting apoptosis.
Furthermore, lncRNA XIST silencing enhances the chemosensitivity of
GBM cells to TMZ (121). These
findings highlight the pivotal role of lncRNA XIST in regulating
malignant progression and mediating drug resistance in GBM
(121).
At the mechanism level, lncRNA XIST directly binds
and sequesters miR-29c, reducing its biological activity and
alleviating its inhibitory effect on downstream target genes. In
TMZ-resistant GBM models, the antitumor and chemosensitizing
effects of lncRNA XIST knockdown were partially reversed by a
miR-29c inhibitor, confirming the central role of the XIST/miR-29c
axis in regulating chemoresistance (118). Further experiments demonstrated
that this regulatory axis enhanced DNA damage repair capacity and
promoted TMZ resistance by modulating the expression levels of the
transcription factor SP1 and MGMT. Specifically, SP1, a
transcriptional regulator of the mismatch repair key protein MutS
homolog 6 (MSH6), is negatively regulated by miR-29c, whereas MGMT
directly reverses O6-MeG lesions induced by TMZ.
Knockdown of XIST markedly reduced the protein levels of MSH6, SP1
and MGMT, an effect reversed by miR-29c inhibition. These findings
indicated that the lncRNA XIST/miR-29c axis coordinately regulates
SP1 and MGMT expression in TMZ-resistant GBM cells (118).
In summary, upregulation of lncRNA XIST constitutes
a notable risk factor for TMZ resistance in GBM, primarily by
regulating the miR-29c/Sp1/MGMT signaling pathway, thereby
modulating cellular chemosensitivity.
lncRNA TMZ-associated lncRNA in GBM
recurrence (TALC)
A previously uncharacterized lncRNA in GBM,
designated lncRNA TALC, is highly expressed in TMZ-resistant
tissues. lncRNA TALC is located at the AL358975 locus, spans two
exons and comprises 418 nts. Mechanistically, lncRNA TALC
expression is regulated by the phosphorylated-AKT/FOXO3 signaling
axis. Specifically, phosphorylated-AKT inhibits FOXO3 nuclear
translocation, relieving the FOXO3-mediated repression effect of
the lncRNA TALC promoter and promoting its transcription (122). Highly expressed lncRNA TALC
competitively binds to miR-20b-3p, alleviating its
post-transcriptional inhibition of c-Met mRNA and activating the
c-Met/STAT3 signaling axis, which enhances TMZ resistance in GBM
(122). Subsequent investigation
demonstrated that lncRNA TALC activated the c-Met/STAT3/p300
pathway, facilitating the formation of a STAT3-histone
acetyltransferase p300 complex that localizes at the MGMT promoter
region. This process increases histone acetylation levels at
histone H3 lysine (H3K) 9, H3K27 and H3K36 sites, remodels
chromatin accessibility, and augments MGMT transcription (122). Functional experiments revealed
that miR-20b-3p inhibition increased MGMT expression, whereas its
overexpression suppressed it. These findings confirmed the central
role of the lncRNA TALC/miR-20b-3p/c-Met axis in the epigenetic
regulation of MGMT (Fig. 2)
(122).
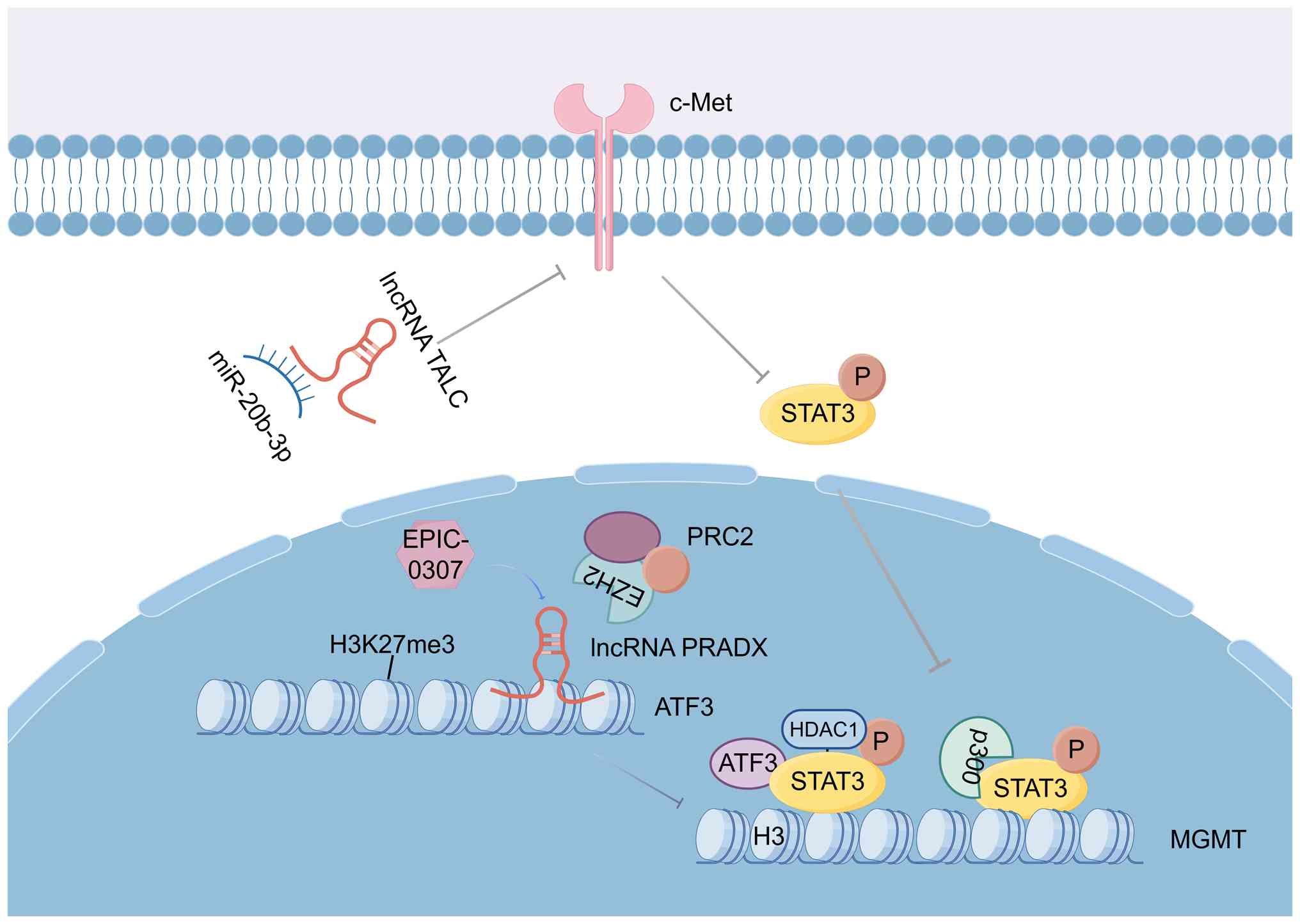 | Figure 2.Schematic diagram illustrating the
mechanisms by which lncRNAs TALC and PRADX regulate MGMT
expression. Arrows denote promotion or positive regulation, while
blunt-ended arrows represent inhibition or negative regulation.
lncRNA, long non-coding RNA; MGMT, O6-methylguanine-DNA
methyltransferase; PRADX, PRC2 and DDX5-associated lncRNA; TALC,
temozolomide-associated lncRNA in glioblastoma recurrence; ATF3,
activating transcription factor 3; P, phosphorylation; H3K27me3,
trimethylation of histone H3 lysine 27; HDACI, histone deacetylase
inhibitor; miR, microRNA. |
In summary, within GBM, the AKT signaling pathway
upregulates lncRNA TALC expression. Functioning as a ceRNA, lncRNA
TALC sequesters miR-20b-3p, thereby activating downstream c-Met
signaling. This cascade ultimately remodels histone acetylation at
the MGMT promoter, resulting in increased MGMT expression and the
induction of TMZ resistance. Therefore, targeting lncRNA TALC
offers a promising strategy to overcome TMZ resistance and provides
a novel strategic avenue to enhance therapeutic outcomes in
patients with GBM.
PRC2 and DDX5-associated lncRNA
(PRADX)
Li et al (123) identified an lncRNA designated
PRADX (transcript ID, ENST00000449248.1). This lncRNA is
upregulated in GBM and colon adenocarcinoma (COAD) and is
predominantly localized within the nucleus (103). A subsequent study demonstrated
that lncRNA PRADX is upregulated in mesenchymal GBM, where it
promotes tumor invasion and metastasis by modulating the tumor
microenvironment (124).
Mechanistic investigations have revealed that
lncRNA PRADX interacts with enhancer of zeste homolog 2 (EZH2), the
catalytic subunit of the polycomb repressive complex 2 (PRC2). This
interaction facilitates the recruitment of PRC2 to specific gene
promoter regions, where it catalyzes trimethylation of H3K27
(H3K27me3) deposition. Therefore, it induces the epigenetic
silencing of multiple tumor suppressor genes, including p21 and p53
upregulated modulator of apoptosis. Furthermore, the PRADX-EZH2
interaction indirectly influences MGMT expression through a
sophisticated multi-layered regulatory network. PRADX suppresses
activating transcription factor 3 (ATF3) transcription by
maintaining H3K27me3 at its promoter. ATF3, functioning as a
pivotal transcription factor, forms a complex with
phosphorylated-STAT3 and histone deacetylase 1 (HDAC1). This
complex is recruited to the MGMT promoter, where it represses MGMT
transcription by reducing H3K27 acetylation levels. Although this
indirect regulatory mechanism introduces complexity to the
therapeutic targeting, it provides valuable insights for developing
combination treatment strategies. The small-molecule inhibitor
EPIC-0307 disrupts the PRADX-EZH2 interaction, thereby relieving
the transcriptional repression of ATF3 (125). Through the ATF3/pSTAT3/HDAC1 axis,
EPIC-0307 downregulates MGMT expression (Fig. 2) and markedly enhances the cytotoxic
efficacy of TMZ (125).
In summary, PRADX is an oncogenic lncRNA that is
highly expressed in GBM and COAD. PRADX regulates MGMT expression
by directly binding to EZH2 and mediating epigenetic silencing. The
small-molecule compound EPIC-0307 targets the PRADX-EZH2
interaction and exhibits synergistic antitumor activity when
combined with TMZ. This regimen markedly enhances therapeutic
efficacy against GBM and offers a potential approach to reversing
TMZ resistance.
circRNAs
circRNAs constitute a class of single-stranded
ncRNAs composed of 100–1,000s of nts. circRNAs form through
back-splicing of precursor mRNA or linear RNA, producing a
covalently closed circular structure with joined 3′- and 5′-ends.
This conformation confers resistance to ribonuclease R degradation
(126,127), thereby enhancing the stability of
circRNAs compared with their linear mRNA counterparts. Studies have
demonstrated that circRNAs can arise from diverse genomic region
and can be generated through multiple circularization patterns.
These characteristics confer notable heterogeneity in circRNA
sequence composition and length, which contributes to their broad
functional diversity (127–129).
circRNAs participate in key biological processes in numerous human
cancer types, such as cervical cancer, colorectal cancer,
pancreatic cancer and glioblastoma (130–135). Their functions include acting as
miRNA sponges, interacting with proteins, regulating gene splicing
or transcription, translating into proteins or peptides, and
contributing to epigenetic regulation (128). Notably, circRNA expression is
dysregulated in neurological disorders, cardiovascular diseases and
multiple cancer types, including breast cancer, colorectal cancer,
hepatocellular carcinoma and glioblastoma (128,135–139). In oncogenesis, specific circRNAs
possess oncogenic or tumor-suppressive roles (140,141). These properties underscore their
potential as diagnostic biomarkers and therapeutic targets
(142,143).
circRNAs in GBM
circRNAs are abundantly expressed in brain tissue
and perform key biological functions. circRNAs serve a key role in
cancer development and progression (139). Previous studies have demonstrated
that circRNAs contribute to the pathological processes of GBM
(144,145). Numerous circRNAs are dysregulated
in GBM tissues and this dysregulation is closely associated with
tumorigenic mechanisms. These circRNAs regulate proliferation,
invasion, migration and cell cycle progression of GBM cells,
underscoring their potential in diagnosis, prognostic evaluation
and therapeutic strategies for GBM (146). circRNAs are either upregulated or
downregulated in GBM cells and tissues, functioning through diverse
molecular mechanisms. In therapeutic contexts, circRNAs have
emerged as a promising class of targets. Oncogenic circRNAs can be
precisely silenced using gene knockdown technologies. By contrast,
tumor-suppressive circRNAs can be restored or enhanced through
vector-based delivery or synthetic biology, thereby modulating key
tumor-associated signaling pathways (145,147). Furthermore, exosome-based circRNA
delivery systems offer a novel method to modulate the tumor
microenvironment and reverse treatment resistance (147). Collectively, these strategies are
advancing GBM diagnosis and therapy toward greater precision,
dynamic monitoring and individualized treatment.
circRNAs regulating MGMT
expression
Previous research has indicated that circRNAs
regulate gene expression at transcriptional and
post-transcriptional levels. Their primary mechanisms include
functioning as miRNA sponges to sequester miRNA activity (148), acting as RNA-binding protein (RBP)
sponges or protein scaffolds to participate in diverse biological
processes (149,150), and directly regulating translation
to mediate polypeptide synthesis (151,152). As central components of the ceRNA
mechanism, circRNAs modulate downstream target gene expression by
establishing a circRNA/miRNA/mRNA regulatory axis, which
contributes to malignant tumor progression (153). Notably, in GBM, specific circRNAs
act as molecular sponges by competitively binding to miRNAs that
target MGMT. This interaction relieves post-transcriptional
inhibition of MGMT mRNA, leading to increased MGMT protein
expression and subsequent TMZ resistance (154). The following section details
circRNAs that regulate MGMT expression in GBM (154).
circWDR62
The WD repeat domain 62 (WDR62) gene comprises 32
exons and encodes a protein consisting of 1,523 amino acid residues
that localizes to the minus ends of mitotic spindle microtubules.
During neurogenesis, WDR62 is highly expressed in the forebrain,
particularly in the ventricular and subventricular zones, where it
serves a key role in neuronal proliferation and migration (155). The host gene WDR62 is markedly
upregulated in various tumor tissues, such as bladder cancer,
breast cancer and renal cancer, and is closely associated with poor
prognosis (156,157). Studies have demonstrated that
circWDR62 expression is elevated in TMZ-resistant GBM tissues and
their derived exosomes, with high expression being positively
associated with poor prognosis in patients with GBM (154). Knockdown of circWDR62 effectively
suppressed TMZ resistance and malignant progression in GBM. At the
molecular level, circWDR62 was predominantly localized in the
cytoplasm and functioned as a ceRNA by sequestering miR-370-3p,
thereby alleviating the inhibitory effect of this miRNA on
downstream target genes. Dual-luciferase reporter assays confirmed
direct binding between circWDR62 and miR-370-3p, as well as between
miR-370-3p and the 3′-UTR of MGMT mRNA. Furthermore, rescue
experiments demonstrated that MGMT overexpression reversed the TMZ
sensitization and suppression of malignant phenotypes induced by
circWDR62 silencing, functionally validating the
circWDR62/miR-370-3p/MGMT regulatory axis (29,154).
Notably, circWDR62 can be packaged into exosomes and transferred
from TMZ-resistant to sensitive cells, inducing similar resistance
and malignant progression in recipient cells. This mechanism
facilitates the ‘horizontal transmission’ of chemoresistance within
the tumor microenvironment, revealing a novel biological role for
exosomal circRNA in the dissemination of chemoresistance in GBM
(154).
However, this study had certain limitations
(154). Although the ceRNA
mechanism was experimentally validated, whether circWDR62 regulates
MGMT expression via RBPs or other epigenetic mechanisms, such as
transcriptional regulation, remains to be elucidated. Furthermore,
the potential roles of circWDR62 in remodeling the tumor immune
microenvironment and metabolic reprogramming warrant further
research.
In summary, exosome-encapsulated circWDR62
functions as a miR-370-3p sponge to regulate MGMT, thereby
promoting TMZ resistance and the malignant progression of GBM in
vitro and in vivo. These findings provided novel
insights into the mechanisms underlying TMZ resistance in GBM and
suggest potential avenues for developing therapeutic strategies
against TMZ-resistant GBM.
Conclusion
Despite advances in surgical techniques and
radiochemotherapy, GBM survival rates have not improved, with
chemoresistance remaining the primary cause of treatment failure
(5). Elucidating the mechanisms of
TMZ resistance in GBM and developing strategies to overcome it are
key to improve therapeutic regimens and enhance glioma prognosis.
Increasing evidence indicates that ncRNAs serve a key role in tumor
chemoresistance (101,129,158,159). The present review systematically
examined how miRNAs, lncRNAs and circRNAs modulate GBM sensitivity
to TMZ through direct or indirect regulation of MGMT expression.
Multiple miRNAs, including miR-142-3p, miR-181d and miR-182-5p,
directly target the 3′-UTR of the MGMT mRNA, promoting its
degradation or inhibiting translation, thereby reducing MGMT
protein levels and enhancing TMZ efficacy. Furthermore, certain
miRNAs, such as miR-29c, indirectly regulate MGMT expression by
targeting transcription factors, including Sp1. The expression
levels of these miRNAs are closely associated with GBM prognosis,
indicating their potential as biomarkers or therapeutic targets.
Notably, synergistic interactions between miRNAs, such as the
combination of miR-181d and miR-409-3p, can more effectively
suppress MGMT expression, offering novel insights for combination
therapy strategies.
lncRNAs regulate MGMT expression either by
competitively binding to miRNAs via the ceRNA mechanism or by
recruiting epigenetic modification complexes, such as PRC2. For
instance, lncRNA UCA1 alleviates MGMT inhibition by sequestering
miR-182-5p, whereas HOTAIR promotes TMZ resistance by activating
the miR-214-3p/β-catenin/MGMT pathway. Aberrant expression levels
of these lncRNAs are closely associated with malignant progression
and chemoresistance in GBM, indicating that targeting of lncRNAs
may represent a promising strategy in overcoming drug resistance in
the future.
circRNAs function as miRNA sponges, regulating
downstream gene expression through ceRNA networks. For example,
circWDR62 upregulates MGMT expression by sequestering miR-370-3p,
thereby promoting TMZ resistance. Their high stability and
tissue-specific expression render circRNAs promising targets for
GBM diagnosis and therapeutic intervention.
Although existing studies have preliminarily
elucidated the key role of ncRNAs in MGMT regulation and
highlighted their translational potential, the field faces notable
challenges. The high complexity of ncRNA regulatory networks, and
their context-dependent functions and limitations, such as in
vivo delivery efficiency, remain major obstacles to clinical
translation. Therefore, future research should further define the
precise roles of ncRNAs within specific cell populations and
spatial contexts, and develop efficient, targeted delivery
strategies. This dual focus is key to translating mechanistic
insights into clinical practice.
Challenges and future perspectives
The present review systematically examines the
mechanisms by which ncRNAs regulate MGMT expression to influence
TMZ resistance in GBM, providing a theoretical foundation in
targeting ncRNAs to overcome chemoresistance. Nevertheless,
translating these findings into clinical practice remains
challenging. To advance this field and enable clinical translation,
future research should employ spatial and single-cell multi-omics
technologies to resolve heterogeneity of ncRNA regulation. Analyses
based on bulk tissue samples capture averaged signals from diverse
cell populations and tumor regions, thereby obscuring the inherent
complexity of ncRNA regulatory networks. GBM exhibits pronounced
intratumoral heterogeneity, encompassing distinct tumor cell
subpopulations and varied components of the tumor microenvironment
(160–162). This heterogeneity contributes to
the apparent functional discrepancies observed for the same ncRNA
across different studies or within distinct regions of a single
tumor. Integrating spatial transcriptomics with single-cell RNA
sequencing holds considerable potential in mapping ncRNA expression
at single-cell resolution. This approach enables the identification
of specific ncRNA expression patterns across cellular
subpopulations and allows precise assignment of their functions to
particular cell types. Simultaneously, it provides insights into
the spatial distribution of ncRNA expression within tumor regions,
such as the core and invasive front, facilitating investigation of
key questions, such as whether molecules such as exosomal circWDR62
undergo directional ‘horizontal transmission’. Furthermore,
integration of multi-omics data supports the construction of cell
type-specific ceRNA networks, enhancing the precise elucidation of
regulatory mechanisms by which ncRNAs modulate key genes, such as
MGMT.
Future research should address the challenges
associated with the technical and clinical translation of ncRNAs as
liquid biopsy biomarkers. Although ncRNAs derived from blood or
cerebrospinal fluid, particularly exosomal circRNAs, demonstrate
potential due to their stability (139,146,147), the absence of standardized
protocols in sample processing and detection remains a major
obstacle. Variability in experimental procedures across studies
limits reproducibility and comparability of results. Furthermore,
improving the specificity of detection methods to distinguish
tumor-derived signals from background noise and enhancing
sensitivity to identify early-stage or minimal residual disease are
primary objectives for future technological advancements.
Simultaneously, the development of advanced bioinformatics
pipelines is key to accurately identify and functionally annotate
low-abundance ncRNAs amid large-scale sequencing data.
Future research should also address the barriers to
clinical translation of ncRNA-targeted therapies. ncRNA-based
therapeutics, including miRNA mimics and inhibitors, face
challenges such as low delivery efficiency, off-target effects and
potential drug resistance. The blood-brain barrier and blood-tumor
barrier constitute major physical obstacles to drug delivery,
necessitating the development of smart carriers that can
specifically target GBM cells and efficiently penetrate these
barriers. Since a single miRNA can regulate hundreds of targets,
artificial intervention may induce unforeseen off-target effects
and toxicity. These off-target effects and potential toxicities can
be mitigated through tumor-specific promoters, chemically modified
antisense oligonucleotides to enhance specificity or intraoperative
local administration to reduce systemic toxicity. Furthermore,
tumor cells may develop resistance to ncRNA therapies through
mechanisms such as target mutation or pathway activation (105,163,164). Therefore, clinical trial designs
should consider combination therapies and dynamic resistance
monitoring. Emerging technologies, including long-read sequencing,
interactomics (such as cross-linking and immunoprecipitation
sequencing) and clustered regularly interspaced short palindromic
repeat-based high-throughput functional screening, are expanding
the current understanding of ncRNA regulatory dimensions (127). These approaches extend beyond the
classical ceRNA mechanism, elucidating transcriptional regulatory
roles and enabling the identification of novel key regulatory
nodes.
Therefore, these advancements are anticipated to
converge in the realization of ncRNA-guided personalized precision
medicine. By integrating patient-derived multi-omics profiles with
dynamic liquid biopsy monitoring, a digital predictive model can be
developed to assess the risk of both intrinsic and acquired TMZ
resistance in a patient, identify optimal ncRNA targets or
combination strategies and guide dynamic adjustment of treatment
regimens, thereby progressing toward truly individualized precision
medicine. In summary, ncRNAs serve a key role in regulating MGMT
and mediating TMZ resistance in GBM. Research in this domain is
transitioning from a discovery phase to a key stage of mechanistic
elucidation and clinical translation. The full potential of ncRNAs
to improve the prognosis of patients with GBM can be realized only
by utilizing insights from emerging technologies and fostering
multidisciplinary collaboration to address technical and biological
challenges in biomarker development and targeted therapy.
Acknowledgements
Not applicable.
Funding
The present review was funded by The National Natural Science
Foundation of China (grant no. 82360609), The Program of Science
and Technology Department of Guizhou Province [grant no. Qian Ke He
Ji Chu-ZK (2022) Yiban 619], The Key Construction Discipline of
Immunology and Pathogen Biology in Zhuhai Campus of Zunyi Medical
University (grant no. ZHGF2024-1) and The Program for High Level
Innovative Talents in the Guizhou Province [grant no. QKHRC-CXTD
(2025)046].
Availability of data and materials
Not applicable.
Authors' contributions
YG reviewed the literature and wrote the first
draft. JG produced the figures. WX collected part of the data. YL
supervised and revised the manuscript. YG, JG, WX and YL were
involved in writing the paper. Data authentication is not
applicable. All authors read and approved the final version of the
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Horbinski C, Solomon DA, Lukas RV, Packer
RJ, Brastianos P, Wen PY, Snuderl M, Berger MS, Chang S, Fouladi M,
et al: Molecular testing for the world health organization
classification of central nervous system tumors: A review. JAMA
Oncol. 11:317–328. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Berger TR, Wen PY, Lang-Orsini M and
Chukwueke UN: World Health Organization 2021 Classification of
Central Nervous System Tumors and Implications for Therapy for
Adult-Type Gliomas: A Review. JAMA Oncol. 8:1493–1501. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yousefi Y, Nejati R, Eslahi A, Alizadeh F,
Farrokhi S, Asoodeh A and Mojarrad M: Enhancing temozolomide (TMZ)
chemosensitivity using CRISPR-dCas9-mediated downregulation of
O6-methylguanine DNA methyltransferase (MGMT). J
Neurooncol. 169:129–135. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Han X, Abdallah MOE, Breuer P, Stahl F,
Bakhit Y, Potthoff AL, Pregler BEF, Schneider M, Waha A, Wüllner U
and Evert BO: Downregulation of MGMT expression by targeted editing
of DNA methylation enhances temozolomide sensitivity in
glioblastoma. Neoplasia. 44:1009292023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Pouyan A, Ghorbanlo M, Eslami M,
Jahanshahi M, Ziaei E, Salami A, Mokhtari K, Shahpasand K, Farahani
N, Meybodi TE, et al: Glioblastoma multiforme: Insights into
pathogenesis, key signaling pathways, and therapeutic strategies.
Mol Cancer. 24:582025. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Duan C, He B, Wang Y, Liu W, Bao W, Yu L,
Xin J, Gui H, Lei J, Yang Z, et al: Stanniocalcin-1 promotes
temozolomide resistance of glioblastoma through regulation of MGMT.
Sci Rep. 14:201992024. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jackson CB, Noorbakhsh SI, Sundaram RK,
Kalathil AN, Ganesa S, Jia L, Breslin H, Burgenske DM, Gilad O,
Sarkaria JN and Bindra RS: Temozolomide sensitizes MGMT-deficient
tumor cells to ATR inhibitors. Cancer Res. 79:4331–4338. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Szames D and Kelley SO:
Mitochondria-targeted temozolomide probe for overcoming
MGMT-Mediated resistance in glioblastoma. Chembiochem.
26:e2024009352025. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Jiang J, Xu J, Ji S, Yu X and Chen J:
Unraveling the mysteries of MGMT: Implications for neuroendocrine
tumors. Biochim Biophys Acta Rev Cancer. 1879:1891842024.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sarkaria JN, Ballman KV, Kizilbash SH,
Sulman EP, Giannini C, Friday BB, Butowski NA, Mohile NA, Piccioni
DE, Battiste JD, et al: Efficacy of adding veliparib to
temozolomide for patients with MGMT-Methylated glioblastoma: A
randomized clinical trial. JAMA Oncol. 10:1637–1644. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Umeda K, Yamada Y, Ito H, Tasaka K, Isobe
K, Akazawa R, Kawabata N, Saida S, Kato I, Hiramatsu H, et al: MGMT
protein expression is a reliable predictive biomarker for
temozolomide-containing chemotherapy in osteosarcoma. Cancer Sci.
115:3394–3402. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wu Z, Dai J, Li J, Zhang Z and Shen X:
Exploiting the role of
O6-methylguanine-DNA-methyltransferase (MGMT) in
gastrointestinal cancers. Naunyn Schmiedebergs Arch Pharmacol.
398:319–327. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Deng M, Ren B, Zhao J, Guo X, Yang Y, Shi
H, Bian X, Wu M, Xu C, Tao M, et al: Temozolomide-promoted MGMT
transcription contributes to chemoresistance by activating the ERK
signalling pathway in malignant melanoma. J Cell Mol Med.
29:e703802025. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Preusser M, Charles Janzer R, Felsberg J,
Reifenberger G, Hamou MF, Diserens AC, Stupp R, Gorlia T, Marosi C,
Heinzl H, et al: Anti-O6-methylguanine-methyltransferase
(MGMT) immunohistochemistry in glioblastoma multiforme: Observer
variability and lack of association with patient survival impede
its use as clinical biomarker. Brain Pathol. 18:520–532. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ramakrishnan V, Kushwaha D, Koay DC, Reddy
H, Mao Y, Zhou L, Ng K, Zinn P, Carter B and Chen CC:
Post-transcriptional regulation of O(6)-methylguanine-DNA
methyltransferase MGMT in glioblastomas. Cancer Biomark.
10:185–193. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chen LL and Kim VN: Small and long
non-coding RNAs: Past, present, and future. Cell. 187:6451–6485.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Vafadar A, Shirazi-Tehrani E, Vosough P,
Khalili Alashti S, Kargar Jahromi H, Bagheri Lankarani K,
Savardashtaki A and Ehtiati S: Non-coding RNAs in celiac disease.
Clin Chim Acta. 576:1204172025. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tonon F, Grassi C, Tierno D, Biasin A,
Grassi M, Grassi G and Dapas B: Non-coding RNAs as potential
diagnostic/prognostic markers for hepatocellular carcinoma. Int J
Mol Sci. 25:122352024. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shi Z, Jin S, Liu X, Jiang M, Fang Y,
Khadaroo PA, Lin H and Fan X: The epigenetic regulatory network of
long noncoding RNAs in hepatocellular carcinoma. Genes Dis.
12:1015342025. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li J, Wang X and Wang H: RNA modifications
in long non-coding RNAs and their implications in cancer biology.
Bioorg Med Chem. 113:1179222024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ma S, Wang W, Zhang D, Zhao G and Lu Z:
Long non-coding RNA colon cancer-associated transcript 2: Role and
function in human cancers. Chin Med J (Engl). 135:2785–2797. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Lin S and Gregory RI: MicroRNA biogenesis
pathways in cancer. Nat Rev Cancer. 15:321–333. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yue J and Tigyi G: MicroRNA trafficking
and human cancer. Cancer Biol Ther. 5:573–578. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Dang Y, Wei X, Xue L, Wen F, Gu J and
Zheng H: Long Non-Coding RNA in Glioma: Target miRNA and signaling
pathways. Clin Lab. 64:887–894. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li J, Liang R, Song C, Xiang Y and Liu Y:
Prognostic and clinicopathological significance of long non-coding
RNA in glioma. Neurosurg Rev. 43:1–8. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Palumbo S, Miracco C, Pirtoli L and
Comincini S: Emerging roles of microRNA in modulating cell-death
processes in malignant glioma. J Cell Physiol. 229:277–286. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tumilson CA, Lea RW, Alder JE and Shaw L:
Circulating microRNA biomarkers for glioma and predicting response
to therapy. Mol Neurobiol. 50:545–558. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wong JW: MicroRNA-induced silencing of
glioma progression. J Neurosci. 30:3868–3869. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kirstein A, Schmid TE and Combs SE: The
role of miRNA for the treatment of MGMT unmethylated glioblastoma
multiforme. Cancers. 12:10992020. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Cheng W, Ren X, Cai J, Zhang C, Li M, Wang
K, Liu Y, Han S and Wu A: A five-miRNA signature with prognostic
and predictive value for MGMT promoter-methylated glioblastoma
patients. Oncotarget. 6:292852015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chen LL: The expanding regulatory
mechanisms and cellular functions of circular RNAs. Nat Rev Mol
Cell Biol. 21:475–490. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Cao YZ, Sun JY, Chen YX, Wen CC and Wei L:
The roles of circRNAs in cancers: Perspectives from molecular
functions. Gene. 767:1451822021. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Yu Z, Huang Q, Zhang Q, Wu H and Zhong Z:
CircRNAs open a new era in the study of cardiovascular disease
(Review). Int J Mol Med. 47:49–64. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zaiou M: circRNAs signature as potential
diagnostic and prognostic biomarker for diabetes mellitus and
related cardiovascular complications. Cells. 9:6592020. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li L, Sun D, Li X, Yang B and Zhang W:
Identification of key circRNAs in Non-small cell lung cancer. Am J
Med Sci. 361:98–105. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Koch L: CRISPR-Cas13 targets circRNAs. Nat
Rev Genet. 22:682021. View Article : Google Scholar
|
|
37
|
Li Y, Feng W, Kong M, Liu R, Wu A, Shen L,
Tang Z and Wang F: Exosomal circRNAs: A new star in cancer. Life
Sci. 269:1190392021. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Luo Q, Li X, Fu B, Zhang L, Fang L, Qing
C, Guo Y, Huang Z and Li J: Expression profile and diagnostic value
of circRNAs in peripheral blood from patients with systemic lupus
erythematosus. Mol Med Rep. 23:12021. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Gebert LFR and MacRae IJ: Regulation of
microRNA function in animals. Nat Rev Mol Cell Biol. 20:21–37.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Henriksen M, Johnsen KB, Andersen HH,
Pilgaard L and Duroux M: MicroRNA expression signatures determine
prognosis and survival in glioblastoma multiforme-a systematic
overview. Mol Neurobiol. 50:896–913. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Balandeh E, Mohammadshafie K, Mahmoudi Y,
Hossein Pourhanifeh M, Rajabi A, Bahabadi ZR, Mohammadi AH,
Rahimian N, Hamblin MR and Mirzaei H: Roles of Non-coding RNAs and
angiogenesis in glioblastoma. Front Cell Dev Biol. 9:7164622021.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Bautista-Sánchez D, Arriaga-Canon C,
Pedroza-Torres A, De La Rosa-Velázquez IA, González-Barrios R,
Contreras-Espinosa L, Montiel-Manríquez R, Castro-Hernández C,
Fragoso-Ontiveros V, Álvarez-Gómez RM and Herrera LA: The promising
role of miR-21 as a cancer biomarker and its importance in
RNA-Based therapeutics. Mol Ther Nucleic Acids. 20:409–420. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Ruppert M, Korkmaz-Icöz S, Benczik B, Ágg
B, Nagy D, Bálint T, Sayour AA, Oláh A, Barta BA, Benke K, et al:
Pressure overload-induced systolic heart failure is associated with
characteristic myocardial microRNA expression signature and
post-transcriptional gene regulation in male rats. Sci Rep.
13:161222023. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Keuls RA, Ochsner SA, O'Neill MB, O'Day
DR, Miyauchi A, Campbell KM, Lanners N, Goldstein JA, Yee C,
McKenna NJ, et al: Post-transcriptional regulation in cranial
neural crest cells expands developmental potential. Proc Natl Acad
Sci USA. 120:e22125781202023. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Alkhazaali-Ali Z, Sahab-Negah S, Boroumand
AR and Tavakol-Afshari J: MicroRNA (miRNA) as a biomarker for
diagnosis, prognosis, and therapeutics molecules in
neurodegenerative disease. Biomed Pharmacother. 177:1168992024.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Pottoo FH, Javed MN, Rahman JU, Abu-Izneid
T and Khan FA: Targeted delivery of miRNA based therapeuticals in
the clinical management of Glioblastoma Multiforme. Semin Cancer
Biol. 69:391–398. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Menon A, Abd-Aziz N, Khalid K, Poh CL and
Naidu R: miRNA: A Promising Therapeutic Target in Cancer. Int J Mol
Sci. 23:115022022. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zhu Y, Huang C, Zhang C, Zhou Y, Zhao E,
Zhang Y, Pan X, Huang H, Liao W and Wang X: LncRNA MIR200CHG
inhibits EMT in gastric cancer by stabilizing miR-200c from
target-directed miRNA degradation. Nat Commun. 14:81412023.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Maltseva D, Zhiyanov A, Lange T and
Tonevitsky A: CD44 knockdown alters miRNA expression and their
target genes in colon cancer. Front Immunol. 16:15526652025.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Han L, Chen S, Luan Z, Fan M, Wang Y, Sun
G and Dai G: Immune function of colon cancer associated miRNA and
target genes. Front Immunol. 14:12030702023. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Uddin MS, Mamun AA, Alghamdi BS, Tewari D,
Jeandet P, Sarwar MS and Ashraf GM: Epigenetics of glioblastoma
multiforme: From molecular mechanisms to therapeutic approaches.
Semin Cancer Biol. 83:100–120. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Hegi ME, Oppong FB, Perry JR, Wick W,
Henriksson R, Laperriere NJ, Gorlia T, Malmström A and Weller M: No
benefit from TMZ treatment in glioblastoma with truly unmethylated
MGMT promoter: Reanalysis of the CE.6 and the pooled Nordic/NOA-08
trials in elderly glioblastoma patients. Neuro Oncol. 26:1867–1875.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Kreth S, Limbeck E, Hinske LC, Schütz SV,
Thon N, Hoefig K, Egensperger R and Kreth FW: In human
glioblastomas transcript elongation by alternative polyadenylation
and miRNA targeting is a potent mechanism of MGMT silencing. Acta
Neuropathol. 125:671–681. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Lee YY, Yarmishyn AA, Wang ML, Chen HY,
Chiou SH, Yang YP, Lin CF, Huang PI, Chen YW, Ma HI, et al:
MicroRNA-142-3p is involved in regulation of MGMT expression in
glioblastoma cells. Cancer Manag Res. 10:775–785. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Chiou GY, Chien CS, Wang ML, Chen MT, Yang
YP, Yu YL, Chien Y, Chang YC, Shen CC, Chio CC, et al: Epigenetic
regulation of the miR142-3p/interleukin-6 circuit in glioblastoma.
Mol Cell. 52:693–706. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Zhang W, Zhang J, Hoadley K, Kushwaha D,
Ramakrishnan V, Li S, Kang C, You Y, Jiang C, Song SW, et al:
miR-181d: A predictive glioblastoma biomarker that downregulates
MGMT expression. Neuro Oncol. 14:712–719. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Lizarte Neto FS, Rodrigues AR, Trevisan
FA, de Assis Cirino ML, Matias CCMS, Pereira-da-Silva G, Peria FM,
Tirapelli DPDC and Carlotti CG Jr: microRNA-181d associated with
the methylation status of the MGMT gene in Glioblastoma multiforme
cancer stem cells submitted to treatments with ionizing radiation
and temozolomide. Brain Res. 1720:1463022019. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Khalil S, Fabbri E, Santangelo A, Bezzerri
V, Cantù C, Di Gennaro G, Finotti A, Ghimenton C, Eccher A,
Dechecchi M, et al: miRNA array screening reveals cooperative
MGMT-regulation between miR-181d-5p and miR-409-3p in glioblastoma.
Oncotarget. 7:28195–28206. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Kushwaha D, Ramakrishnan V, Ng K, Steed T,
Nguyen T, Futalan D, Akers JC, Sarkaria J, Jiang T, Chowdhury D, et
al: A genome-wide miRNA screen revealed miR-603 as a
MGMT-regulating miRNA in glioblastomas. Oncotarget. 5:4026–4039.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Jesionek-Kupnicka D, Braun M,
Trąbska-Kluch B, Czech J, Szybka M, Szymańska B, Kulczycka-Wojdala
D, Bieńkowski M, Kordek R and Zawlik I: MiR-21, miR-34a, miR-125b,
miR-181d and miR-648 levels inversely correlate with MGMT and TP53
expression in primary glioblastoma patients. Arch Med Sci.
15:504–512. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Cheng M, Wang Q, Chen L, Zhao D, Tang J,
Xu J and He Z: LncRNA UCA1/miR-182-5p/MGMT axis modulates glioma
cell sensitivity to temozolomide through MGMT-related DNA damage
pathways. Hum Pathol. 123:59–73. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Feng YA, Liu TE and Wu Y: microRNA-182
inhibits the proliferation and migration of glioma cells through
the induction of neuritin expression. Oncol Lett. 10:1197–1203.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Deng Y, Zhu H, Xiao L, Liu C and Meng X:
Circ_0005198 enhances temozolomide resistance of glioma cells
through miR-198/TRIM14 axis. Aging (Albany NY). 13:2198–2211. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Nie E, Jin X, Wu W, Yu T, Zhou X, Shi Z,
Zhang J, Liu N and You Y: MiR-198 enhances temozolomide sensitivity
in glioblastoma by targeting MGMT. J Neurooncol. 133:59–68. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Quintavalle C, Mangani D, Roscigno G,
Romano G, Diaz-Lagares A, Iaboni M, Donnarumma E, Fiore D, De
Marinis P, Soini Y, et al: MiR-221/222 target the DNA
methyltransferase MGMT in glioma cells. PLoS One. 8:e744662013.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Chen Z, Xiang B, Qi L, Zhu S and Li L:
miR-221-3p promotes hepatocellular carcinogenesis by downregulating
O6-methylguanine-DNA methyltransferase. Cancer Biol
Ther. 21:915–926. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Tian F, Wang P, Lin D, Dai J, Liu Q, Guan
Y, Zhan Y, Yang Y, Wang W, Wang J, et al: Exosome-delivered
miR-221/222 exacerbates tumor liver metastasis by targeting SPINT1
in colorectal cancer. Cancer Sci. 112:3744–3755. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Yang J, Yan Z, Wang Y, Xu J, Li R, Li C,
Liu S, Shi L and Yao Y: Association study of relationships of
polymorphisms in the miR-21, miR-26b, miR-221/222 and miR-126 genes
with cervical intraepithelial neoplasia and cervical cancer. BMC
Cancer. 21:9972021. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Song Q, An Q, Niu B, Lu X, Zhang N and Cao
X: Role of miR-221/222 in tumor development and the underlying
mechanism. J Oncol. 2019:72520132019. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Garofalo M, Di Leva G, Romano G, Nuovo G,
Suh SS, Ngankeu A, Taccioli C, Pichiorri F, Alder H, Secchiero P,
et al: Retraction notice to: miR-221&222 regulate TRAIL
resistance and enhance tumorigenicity through PTEN and TIMP3
downregulation. Cancer Cell. 40:14402022. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Guo M, Lin B, Li G, Lin J and Jiang X:
LncRNA TDRG1 promotes the proliferation, migration, and invasion of
cervical cancer cells by sponging miR-214-5p to target SOX4. J
Recept Signal Transduct Res. 40:281–293. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Terada N, Karim MR, Izawa T, Kuwamura M
and Yamate J: Immunolocalization of β-catenin, E-cadherin and
N-cadherin in neonate and adult rat kidney. J Vet Med Sci.
79:1785–1790. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Zhou Y, Chen L, Ding D, Li Z, Cheng L, You
Q and Zhang S: Cyanidin-3-O-glucoside inhibits the β-catenin/MGMT
pathway by upregulating miR-214-5p to reverse chemotherapy
resistance in glioma cells. Sci Rep. 12:77732022. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Amirian M, Jafari-Nozad AM, Darroudi M,
Farkhondeh T and Samarghandian S: Overview of the miR-29 family
members' function in breast cancer. Int J Biol Macromol.
230:1232802023. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Kim S, Lee KW, Yoo Y, Park SH, Lee JW,
Jeon S, Illia S, Joshi P, Park HW, Lo HE, et al: MiR-29 and MiR-140
regulate TRAIL-induced drug tolerance in lung cancer. Anim Cells
Syst (Seoul). 28:184–197. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Xiao S, Yang Z, Qiu X, Lv R, Liu J, Wu M,
Liao Y and Liu Q: miR-29c contribute to glioma cells temozolomide
sensitivity by targeting O6-methylguanine-DNA
methyltransferases indirectely. Oncotarget. 7:50229–50238. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Shaw R, Karmakar S, Basu M and Ghosh MK:
DDX5 (p68) orchestrates β-catenin, RelA and SP1 mediated MGMT gene
expression in human colon cancer cells: Implication in TMZ
chemoresistance. Biochim Biophys Acta Gene Regul Mech.
1866:1949912023. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Hui P, Wang Y, Chen B, Wang Z and Qin S:
Mir-29c expression in glioma and its effects on tumor cell
proliferation and apoptosis. Iran J Public Health. 49:304–311.
2020.PubMed/NCBI
|
|
79
|
Li G, Zhao J, Peng X, Liang J, Deng X and
Chen Y: The mechanism involved in the loss of PTEN expression in
NSCLC tumor cells. Biochem Biophys Res Commun. 418:547–552. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Ding J, Fayyaz AI, Ding Y, Liang D and Luo
M: Role of specificity protein 1 (SP1) in cardiovascular diseases:
Pathological mechanisms and therapeutic potentials. Biomolecules.
14:8072024. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Gao YT, Chen XB and Liu HL: Up-regulation
of miR-370-3p restores glioblastoma multiforme sensitivity to
temozolomide by influencing MGMT expression. Sci Rep. 6:329722016.
View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Louis DN, Perry A, Wesseling P, Brat DJ,
Cree IA, Figarella-Branger D, Hawkins C, Ng HK, Pfister SM,
Reifenberger G, et al: The 2021 WHO classification of tumors of the
central nervous system: A summary. Neuro Oncol. 23:1231–1251. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Nadaradjane A, Briand J, Bougras-Cartron
G, Disdero V, Vallette FM, Frenel JS and Cartron PF: miR-370-3p is
a therapeutic tool in Anti-glioblastoma therapy but is not an
intratumoral or Cell-free circulating biomarker. Mol Ther Nucleic
Acids. 13:642–650. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Chou ST, Peng HY, Mo KC, Hsu YM, Wu GH,
Hsiao JR, Lin SF, Wang HD and Shiah SG: MicroRNA-486-3p functions
as a tumor suppressor in oral cancer by targeting DDR1. J Exp Clin
Cancer Res. 38:2812019. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Yang S, Sui J, Liu T, Wu W, Xu S, Yin L,
Pu Y, Zhang X, Zhang Y, Shen B and Liang G: Expression of
miR-486-5p and its significance in lung squamous cell carcinoma. J
Cell Biochem. 120:13912–13923. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Dong N, Qi W, Wu L, Li J, Zhang X, Wu H,
Zhang W, Jiang J, Zhang S, Fu W, et al: LINC00606 promotes
glioblastoma progression through sponge miR-486-3p and interaction
with ATP11B. J Exp Clin Cancer Res. 43:1392024. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Ravindra J, Ug Y, Pandyanda Nanjappa D,
Kalladka K, Dhakal R, Chakraborty A and Chakraborty G: Allicin
extracted from Allium sativum shows potent anti-cancer and
antioxidant properties in zebrafish. Biomed Pharmacother.
169:1158542023. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Rauf A, Abu-Izneid T, Thiruvengadam M,
Imran M, Olatunde A, Shariati MA, Bawazeer S, Naz S, Shirooie S,
Sanches-Silva A, et al: Garlic (Allium sativum L.): Its
chemistry, nutritional composition, toxicity, and anticancer
properties. Curr Top Med Chem. 22:957–972. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Yang Z, Du J, Zhu J, Rong Y, Chen S, Yu L,
Deng X, Zhang X, Sheng H, Yang L, et al: Allicin inhibits
proliferation by decreasing IL-6 and IFN-β in HCMV–Infected glioma
cells. Cancer Manag Res. 12:7305–7317. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Wu H, Li X, Zhang T, Zhang G, Chen J, Chen
L, He M, Hao B and Wang C: Overexpression miR-486-3p promoted by
allicin enhances temozolomide sensitivity in glioblastoma via
targeting MGMT. Neuromolecular Med. 22:359–369. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Guo M, Zhang X, Wang G, Sun J, Jiang Z,
Khadarian K, Yu S, Zhao Y, Xie C, Zhang K, et al: miR-603 promotes
glioma cell growth via Wnt/β-catenin pathway by inhibiting WIF1 and
CTNNBIP1. Cancer Lett. 360:76–86. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Abate M, Porru M, Campani V, Leonetti C,
Nele V, Di Paola R, De Martino M, Russo M, Tathode M, Cossu AM, et
al: Self-assembling nanoparticles for delivery of miR-603 and
miR-221 in glioblastoma as a new strategy to overcome resistance to
temozolomide. J Control Release. 377:458–469. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Yuan Y, Tang Y, Fang Z, Wen J, Wicha MS
and Luo M: Long Non-coding RNAs: Key regulators of tumor
Epithelial/Mesenchymal plasticity and cancer stemness. Cells.
14:2272025. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Nadhan R, Isidoro C, Song YS and
Dhanasekaran DN: LncRNAs and the cancer epigenome: Mechanisms and
therapeutic potential. Cancer Lett. 605:2172972024. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Chi SW, Zang JB, Mele A and Darnell RB:
Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature.
460:479–486. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Pokorná M, Černá M, Boussios S, Ovsepian
SV and O'Leary VB: lncRNA biomarkers of glioblastoma multiforme.
Biomedicines. 12:9322024. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Yang L, Tang L, Min Q, Tian H, Li L, Zhao
Y, Wu X, Li M, Du F, Chen Y, et al: Emerging role of RNA
modification and long noncoding RNA interaction in cancer. Cancer
Gene Ther. 31:816–830. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Tai F and Zhao X: Research progress on
function and mechanism of long non-coding RNA in glioma. Cell Mol
Biol (Noisy-le-Grand). 70:233–237. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Goenka A, Song X, Tiek D, Iglesia RP, Lu
M, Zeng C, Horbinski C, Zhang W, Hu B and Cheng SY: Oncogenic long
noncoding RNA LINC02283 enhances PDGF receptor A-mediated signaling
and drives glioblastoma tumorigenesis. Neuro Oncol. 25:1592–1604.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Sun P, Bao A, Hua X, Cao J and Ding Y:
RP5-1148A21.3 (lncRP5) exerts oncogenic function in human ovarian
carcinoma. Acta Biochim Biophys Sin (Shanghai). 54:209–219. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Ran R, Gong CY, Wang ZQ, Zhou WM, Zhang
SB, Shi YQ, Ma CW and Zhang HH: Long non-coding RNA PART1: Dual
role in cancer. Hum Cell. 35:1364–1374. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Shi D, Zhong W, Liu D, Sun X, Hao S, Yang
Y, Ao L, Zhou J, Xia Y, Zhou Y, et al: Computational identification
of immune-related lncRNA signature for predicting the prognosis and
immune landscape of human glioblastoma multiforme. Front Immunol.
13:9329382022. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Zhang B, Cheng Y, Li R, Lian M, Guo S and
Liang C: Development of a novel angiogenesis-related lncRNA
signature to predict the prognosis and immunotherapy of
glioblastoma multiforme. Transl Cancer Res. 12:13–30. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Li Q, Jia H, Li H, Dong C, Wang Y and Zou
Z: LncRNA and mRNA expression profiles of glioblastoma multiforme
(GBM) reveal the potential roles of lncRNAs in GBM pathogenesis.
Tumour Biol. 37:14537–14552. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Shahzad U, Krumholtz S, Rutka JT and Das
S: Noncoding RNAs in Glioblastoma: Emerging biological concepts and
potential therapeutic implications. Cancers (Basel). 13:15552021.
View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Stackhouse CT, Gillespie GY and Willey CD:
Exploring the roles of lncRNAs in GBM pathophysiology and their
therapeutic potential. Cells. 9:23692020. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Kong X, Li R, Chen M, Zheng R, Wang J, Sun
C and Qu Y: Endogenous retrovirus HERVH-derived lncRNA UCA1
controls human trophoblast development. Proc Natl Acad Sci USA.
121:e23181761212024. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Duan DM, Zhang L and Hua F: LncRNA UCA1
inhibits proliferation and promotes apoptosis of cervical cancer
cells by regulating β-catenin/TCF-4. Eur Rev Med Pharmacol Sci.
24:5963–5969. 2020.PubMed/NCBI
|
|
109
|
Wang CJ, Zhu CC, Xu J, Wang M, Zhao WY,
Liu Q, Zhao G and Zhang ZZ: The lncRNA UCA1 promotes proliferation,
migration, immune escape and inhibits apoptosis in gastric cancer
by sponging anti-tumor miRNAs. Mol Cancer. 18:1152019. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Xue M, Chen W and Li X: Urothelial cancer
associated 1: A long noncoding RNA with a crucial role in cancer. J
Cancer Res Clin Oncol. 142:1407–1419. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
He Z, You C and Zhao D: Long non-coding
RNA UCA1/miR-182/PFKFB2 axis modulates Glioblastoma-associated
stromal cells-mediated glycolysis and invasion of glioma cells.
Biochem Biophys Res Commun. 500:569–576. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
He Z, Wang Y, Huang G, Wang Q, Zhao D and
Chen L: The lncRNA UCA1 interacts with miR-182 to modulate glioma
proliferation and migration by targeting iASPP. Arch Biochem
Biophys. 623–624. 1–8. 2017.
|
|
113
|
Angelopoulou E, Paudel YN and Piperi C:
Critical role of HOX transcript antisense intergenic RNA (HOTAIR)
in gliomas. J Mol Med (Berl). 98:1525–1546. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Zhang J, Chen G, Gao Y and Liang H:
HOTAIR/miR-125 axis-mediated Hexokinase 2 expression promotes
chemoresistance in human glioblastoma. J Cell Mol Med.
24:5707–5717. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Lan T, Quan W, Yu DH, Chen X, Wang ZF and
Li ZQ: High expression of LncRNA HOTAIR is a risk factor for
temozolomide resistance in glioblastoma via activation of the
miR-214/β-catenin/MGMT pathway. Sci Rep. 14:262242024. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Balci T, Yilmaz Susluer S, Kayabasi C,
Ozmen Yelken B, Biray Avci C and Gunduz C: Analysis of dysregulated
long non-coding RNA expressions in glioblastoma cells. Gene.
590:120–122. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Anastas JN and Moon RT: WNT signalling
pathways as therapeutic targets in cancer. Nat Rev Cancer.
13:11–26. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Du P, Zhao H, Peng R, Liu Q, Yuan J, Peng
G and Liao Y: LncRNA-XIST interacts with miR-29c to modulate the
chemoresistance of glioma cell to TMZ through DNA mismatch repair
pathway. Biosci Rep. 37:BSR201706962017. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Ren C, Li X, Wang T, Wang G, Zhao C, Liang
T, Zhu Y, Li M, Yang C, Zhao Y and Zhang GM: Functions and
mechanisms of long noncoding RNAs in ovarian cancer. Int J Gynecol
Cancer. 25:566–569. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Fang J, Sun CC and Gong C: Long noncoding
RNA XIST acts as an oncogene in non-small cell lung cancer by
epigenetically repressing KLF2 expression. Biochem Biophys Res
Commun. 478:811–817. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Yao Y, Ma J, Xue Y, Wang P, Li Z, Liu J,
Chen L, Xi Z, Teng H, Wang Z, et al: Knockdown of long non-coding
RNA XIST exerts tumor-suppressive functions in human glioblastoma
stem cells by up-regulating miR-152. Cancer Lett. 359:75–86. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Wu P, Cai J, Chen Q, Han B, Meng X, Li Y,
Li Z, Wang R, Lin L, Duan C, et al: Lnc-TALC promotes
O6-methylguanine-DNA methyltransferase expression via
regulating the c-Met pathway by competitively binding with
miR-20b-3p. Nat Commun. 10:20452019. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Li Y, Liu X, Cui X, Tan Y, Wang Q, Wang Y,
Xu C, Fang C and Kang C: LncRNA PRADX-mediated recruitment of
PRC2/DDX5 complex suppresses UBXN1 expression and activates NF-κB
activity, promoting tumorigenesis. Theranostics. 11:4516–4530.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Xu C, Zhao J, Song J, Xiao M, Cui X, Xin
L, Xu J, Zhang Y, Yi K, Hong B, et al: lncRNA PRADX is a
mesenchymal glioblastoma biomarker for cellular metabolism targeted
therapy. Front Oncol. 12:8889222022. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Xin L, Tan Y, Zhu Y, Cui X, Wang Q, Zhao
J, Tian S, Xu C, Xiao M, Hong B, et al: EPIC-0307-mediated
selective disruption of PRADX-EZH2 interaction and enhancement of
temozolomide sensitivity to glioblastoma via inhibiting DNA repair
and MGMT. Neuro Oncol. 25:1976–1988. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
126
|
He AT, Liu J, Li F and Yang BB: Targeting
circular RNAs as a therapeutic approach: Current strategies and
challenges. Signal Transduct Target Ther. 6:1852021. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Kristensen LS, Andersen MS, Stagsted LVW,
Ebbesen KK, Hansen TB and Kjems J: The biogenesis, biology and
characterization of circular RNAs. Nat Rev Genet. 20:675–691. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Conn VM, Chinnaiyan AM and Conn SJ:
Circular RNA in cancer. Nat Rev Cancer. 24:597–613. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Ma S, Kong S, Wang F and Ju S: CircRNAs:
Biogenesis, functions, and role in drug-resistant Tumours. Mol
Cancer. 19:1192020. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Wang Y, Zhao J, Wu J, Liu J, Wang Y, Xu T,
Zhang M, Zhuang M, Zou L, Sun W, et al: Genome-wide perturbations
of A-to-I RNA editing dysregulated circular RNAs promoting the
development of cervical cancer. Comput Biol Med. 166:1075462023.
View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Heidari-Ezzati S, Moeinian P,
Ahmadian-Nejad B, Maghbbouli F, Abbasi S, Zahedi M, Afkhami H,
Shadab A and Sajedi N: The role of long non-coding RNAs and
circular RNAs in cervical cancer: Modulating miRNA function. Front
Cell Dev Biol. 12:13087302024. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Fu X, Chen P and Wang H: The role of
circRNA-miRNA-mRNA regulatory network and its potential biomarker
function in colorectal cancer. Protein Pept Lett. 30:1020–1029.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Chen H, Wang X, Liu S, Tang Z, Xie F, Yin
J, Sun P and Wang H: Circular RNA in pancreatic cancer: Biogenesis,
mechanism, function and clinical application. Int J Med Sci.
22:1612–1629. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Ghadami E, Jafari M, Razipour M, Maghsudlu
M and Ghadami M: Circular RNAs in glioblastoma. Clin Chim Acta.
565:1200032025. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Hashemi M, Khosroshahi EM, Daneii P,
Hassanpoor A, Eslami M, Koohpar ZK, Asadi S, Zabihi A, Jamali B,
Ghorbani A, et al: Emerging roles of CircRNA-miRNA networks in
cancer development and therapeutic response. Noncoding RNA Res.
10:98–115. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Ebbesen KK, Kjems J and Hansen TB:
Circular RNAs: Identification, biogenesis and function. Biochim
Biophys Acta. 1859:163–168. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Wang Y and Wang Z: Efficient backsplicing
produces translatable circular mRNAs. RNA. 21:172–179. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Wilusz JE: Circular RNAs: Unexpected
outputs of many protein-coding genes. RNA Biol. 14:1007–1017. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
139
|
Li J, Sun D, Pu W, Wang J and Peng Y:
Circular RNAs in cancer: Biogenesis, function, and clinical
significance. Trends Cancer. 6:319–336. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
140
|
Yang X, Ye T, Liu H, Lv P, Duan C, Wu X,
Jiang K, Lu H, Xia D, Peng E, et al: Expression profiles,
biological functions and clinical significance of circRNAs in
bladder cancer. Mol Cancer. 20:42021. View Article : Google Scholar : PubMed/NCBI
|
|
141
|
Geng X, Zhang Y, Li Q, Xi W, Yu W, Shi L,
Lin X, Sun S and Wang H: Screening and functional prediction of
differentially expressed circular RNAs in human glioma of different
grades. Aging (Albany NY). 13:1989–2014. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
142
|
Wang S, Chao F, Zhang C, Han D, Xu G and
Chen G: Circular RNA circPFKP promotes cell proliferation by
activating IMPDH2 in prostate cancer. Cancer Lett. 524:109–120.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Li B, Zhu L, Lu C, Wang C, Wang H, Jin H,
Ma X, Cheng Z, Yu C, Wang S, et al: circNDUFB2 inhibits non-small
cell lung cancer progression via destabilizing IGF2BPs and
activating anti-tumor immunity. Nat Commun. 12:2952021. View Article : Google Scholar : PubMed/NCBI
|
|
144
|
Wang Y, Yu Y, Yu J, Wang C, Wang Y, Fu R
and Zhang C: The role of the dysregulation of circRNAs expression
in glioblastoma multiforme. J Mol Neurosci. 75:92025. View Article : Google Scholar : PubMed/NCBI
|
|
145
|
Wang H, Bian C, Zhang Y, Zhang L and Wang
F: Circular RNAs in glioma progression: Fundamental mechanisms and
therapeutic potential: A review. Int J Biol Macromol.
313:1443602025. View Article : Google Scholar : PubMed/NCBI
|
|
146
|
Sun W, Zhou H, Han X, Hou L and Xue X:
Circular RNA: A novel type of biomarker for glioma (Review). Mol
Med Rep. 24:6022021. View Article : Google Scholar : PubMed/NCBI
|
|
147
|
Guo X and Piao H: Research progress of
circRNAs in glioblastoma. Front Cell Dev Biol. 9:7918922021.
View Article : Google Scholar : PubMed/NCBI
|
|
148
|
Liu G, Sun J, Yang ZF, Zhou C, Zhou PY,
Guan RY, Sun BY, Wang ZT, Zhou J, Fan J, et al: Cancer-associated
fibroblast-derived CXCL11 modulates hepatocellular carcinoma cell
migration and tumor metastasis through the
circUBAP2/miR-4756/IFIT1/3 axis. Cell Death Dis. 12:2602021.
View Article : Google Scholar : PubMed/NCBI
|
|
149
|
Huang Q, Guo H, Wang S, Ma Y, Chen H, Li
H, Li J, Li X, Yang F, Qiu M, et al: A novel circular RNA,
circXPO1, promotes lung adenocarcinoma progression by interacting
with IGF2BP1. Cell Death Dis. 11:10312020. View Article : Google Scholar : PubMed/NCBI
|
|
150
|
Liu Z, Gu S, Wu K, Li L, Dong C, Wang W
and Zhou Y: CircRNA-DOPEY2 enhances the chemosensitivity of
esophageal cancer cells by inhibiting CPEB4-mediated Mcl-1
translation. J Exp Clin Cancer Res. 40:3612021. View Article : Google Scholar : PubMed/NCBI
|
|
151
|
Wu N, Yuan Z, Du KY, Fang L, Lyu J, Zhang
C, He A, Eshaghi E, Zeng K, Ma J, et al: Translation of
yes-associated protein (YAP) was antagonized by its circular RNA
via suppressing the assembly of the translation initiation
machinery. Cell Death Differ. 26:2758–2773. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
152
|
Li Y, Wang Z, Su P, Liang Y, Li Z, Zhang
H, Song X, Han D, Wang X, Liu Y, et al: circ-EIF6 encodes
EIF6-224aa to promote TNBC progression via stabilizing MYH9 and
activating the Wnt/beta-catenin pathway. Mol Ther. 30:415–430.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
153
|
Wu M, Yuan H, Zou W, Xu S, Liu S, Gao Q,
Guo Q, Han Y and An X: Circular RNAs: Characteristics, functions,
mechanisms, and potential applications in thyroid cancer. Clin
Transl Oncol. 26:808–824. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
154
|
Geng X, Zhang Y, Lin X, Zeng Z, Hu J, Hao
L, Xu J, Wang X, Wang H and Li Q: Exosomal circWDR62 promotes
temozolomide resistance and malignant progression through
regulation of the miR-370-3p/MGMT axis in glioma. Cell Death Dis.
13:5962022. View Article : Google Scholar : PubMed/NCBI
|
|
155
|
Chen H, Zheng Y, Wu H, Cai N, Xu G, Lin Y
and Li JJ: Autosomal recessive primary microcephaly type 2
associated with a novel WDR62 splicing variant that disrupts the
expression of the functional transcript. Front Neurol.
15:13418642024. View Article : Google Scholar : PubMed/NCBI
|
|
156
|
Das R, Sjöström M, Shrestha R, Yogodzinski
C, Egusa EA, Chesner LN, Chen WS, Chou J, Dang DK, Swinderman JT,
et al: An integrated functional and clinical genomics approach
reveals genes driving aggressive metastatic prostate cancer. Nat
Commun. 12:46012021. View Article : Google Scholar : PubMed/NCBI
|
|
157
|
Bu Y, Zhang L, Ma X, Wang R, Zhang X and
Li J: Systematic analysis of the oncogenic role of WDR62 in human
tumors. Dis Markers. 2021:99402742021. View Article : Google Scholar : PubMed/NCBI
|
|
158
|
Kousar K, Ahmad T, Abduh MS, Kanwal B,
Shah SS, Naseer F and Anjum S: miRNAs in regulation of tumor
microenvironment, chemotherapy resistance, immunotherapy modulation
and miRNA therapeutics in cancer. Int J Mol Sci. 23:138222022.
View Article : Google Scholar : PubMed/NCBI
|
|
159
|
Singh D, Assaraf YG and Gacche RN: Long
non-coding RNA mediated drug resistance in breast cancer. Drug
Resist Updat. 63:1008512022. View Article : Google Scholar : PubMed/NCBI
|
|
160
|
Altaf S, Cummins MJ, Ittner LM and Mattick
JS: The emerging roles of long non-coding RNAs in the nervous
system. Nat Rev Neurosci. 26:661–676. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
161
|
Goenka A, Tiek DM, Song X, Iglesia RP, Lu
M, Hu B and Cheng SY: The role of non-coding RNAs in glioma.
Biomedicines. 10:20312022. View Article : Google Scholar : PubMed/NCBI
|
|
162
|
Slack FJ and Chinnaiyan AM: The role of
Non-coding RNAs in oncology. Cell. 179:1033–1055. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
163
|
Masoomabadi N, Gorji A, Ghadiri T and
Ebrahimi S: Regulatory role of circular RNAs in the development of
therapeutic resistance in the glioma: A double-edged sword. Iran J
Basic Med Sci. 28:3–15. 2025.PubMed/NCBI
|
|
164
|
Dai L, Liang W, Shi Z, Li X, Zhou S, Hu W,
Yang Z and Wang X: Systematic characterization and biological
functions of non-coding RNAs in glioblastoma. Cell Prolif.
56:e133752023. View Article : Google Scholar : PubMed/NCBI
|
















