Introduction
Cholangiocarcinoma (CCA), the second most prevalent
primary hepatic malignancy originating from bile duct epithelial
cells, accounts for ~15% of all primary liver tumors, with a rising
global incidence and mortality (1,2). It is
characterized by pronounced pro-fibroplasia, a complex tumor
microenvironment, and considerable genetic heterogeneity, all
contributing to heightened drug resistance. Non-specific clinical
manifestations often lead to delayed diagnoses, precluding timely
surgical interventions and resulting in an unfavorable prognosis.
Epidemiological data highlight an increasing incidence and
mortality, with CCA accounting for 15% of liver malignancies
(3–5). Consequently, there is an urgent need
to elucidate novel strategies and pharmacological interventions to
counteract CCA tumor metastasis and combat chemotherapy
resistance.
Schisandrin B (Sch B), derived from the fruit of
Schisandra chinensis Baill and used in traditional Chinese
medicine, has demonstrated therapeutic efficacy across various
malignant tumors, including glioma (6,7),
gallbladder cancer (8), breast
cancer (9), prostate cancer
(10) and hepatic cancer (11), by eliciting anti-proliferative and
pro-apoptotic effects. Previous research has indicated
mitochondria-mediated intrinsic apoptotic pathways as one of its
validated biological mechanisms.
Reactive oxygen species (ROS), byproducts of aerobic
metabolism, play a key role in physiological processes and REDOX
balance maintenance (12–14). ROS act as initiators and mediators
of multiple signal transduction pathways, exerting inhibitory
effects on tumor cell proliferation. They are involved in mediating
various anti-tumorigenic signaling pathways inducing DNA damage,
genetic instability and oxidative stress-associated tumor cell
apoptosis (15). Mounting evidence
has established that the ROS-activated mitochondria-mediated
intrinsic apoptosis pathway represents a central mechanism
underlying these effects (16–18).
Mitochondria, the main site of oxygen-free radical production,
undergo alterations in permeability with increased ROS leading to
decreased mitochondrial transmembrane potential (ΔΨm). The
resulting decrease in ΔΨm facilitates the release of cytochrome
c into the cytoplasm, initiating apoptosis via the caspase
pathway (19).
Network pharmacology enables comprehensive
target-based functional analysis and prediction of drug components
and diseases, thus providing a robust framework for elucidating the
complex mechanisms of drug action and disease pathology (20–22).
The present study was designed to investigate the anti-CCA
mechanism of Sch B using an integrated strategy combining network
pharmacology, molecular docking and in vitro experiments,
with a focus on the ROS/p38 MAPK/NF-κB signaling pathway. This
multi-faceted approach may provide a foundation for understanding
the therapeutic potential of Sch B as a natural-derived agent
against CCA.
Materials and methods
Network pharmacology
Potential targets of Sch B were predicted using the
PharmMapper database (http://www.lilab-ecust.cn/pharmmapper/), followed by
standardization via the UniProt database (https://www.uniprot.org/) to obtain relevant target
information. Disease-related targets for CCA were retrieved from
the GeneCards (https://www.genecards.org/) and DisGeNET databases
(https://www.disgenet.org/) using the
keyword ‘Cholangiocarcinoma’. After merging datasets, non-human
genes and duplicate entries were removed to generate a standardized
target list. An EVenn online tool (http://www.ehbio.com/test/venn) was employed to
construct a Venn diagram illustrating overlapping targets between
Sch B and CCA.
The shared targets were imported into the STRING
database (https://string-db.org/) to construct a
protein-protein interaction (PPI) network under the ‘Homo sapiens’
setting. Disconnected nodes were hidden, and a high confidence
interaction score of ≥0.900 was applied. The resulting network was
exported in Tab-Separated Values format and visualized using
Cytoscape 3.8.0 (https://cytoscape.org/). Core targets are identified
based on three topological parameters: Degree centrality (DC),
betweenness centrality (BC) and closeness centrality (CC).
The drug-disease intersection targets were uploaded
to the Metascape platform (http://metascape.org/) for gene ontology (GO)
(https://geneontology.org/) enrichment
analysis, including biological process (BP), cellular component and
molecular function (MF) categories. Analysis parameters included a
minimum overlap of 3, a P-value cut-off of 0.01 and an enrichment
of ≥1.5. The top 10 enriched terms from each GO category were
selected based on ascending log P-values. Kyoto Encyclopedia of
Genes and Genomes (KEGG) pathway analysis was performed using the
ClueGO plugin in Cytoscape, with thresholds set to include pathways
containing ≥16 genes and a κ score >0.6. A total of 11 key
pathways were identified as significantly enriched.
Molecular docking analysis
Molecular docking was performed with the selected
core targets. The structure of Sch B in .mol2 format was obtained
from the TCMSP database (http://tcmsp-e.com/) and prepared using AutoDock
Tools-1.5.6 by removing water molecules, adding hydrogen atoms and
assigning atom types. Rotatable bonds were defined, and the file
was saved in .pdbqt format, which can be opened and used for
molecular docking with AutoDock Vina (version 1.1.2; The Scripps
Research Institute).
Crystal structures of MAPK1, EGFR, ESR1, HSP90AA1,
AKT1, GRB2, SRC, and HRAS in PDB format were retrieved from the PDB
database (https://www.wwpdb.org/). Using PyMOL
3.7.1, non-essential small molecules were removed from the protein
structures. The proteins were then processed in AutoDock
Tools-1.5.6 to remove water molecules, add hydrogens and assign
atom types, and these were saved as .pdbqt files.
The prepared Sch B ligand and the eight core protein
receptors were subjected to molecular docking using AutoDock Vina
with a batch processing script. The binding site and grid
parameters were appropriately defined for each target. The
resulting binding modes were visualized and analyzed using PyMOL
3.7.1.
Cell lines and culture
Human CCA HuccT1 cell line was sourced from The Cell
Bank of Type Culture Collection of The Chinese Academy of Sciences.
Cells were cultured in RPMI-1640 medium (Gibco; Thermo Fisher
Scientific, Inc.) supplemented with 10% FBS (Gibco; Thermo Fisher
Scientific, Inc.), 100 µg/ml streptomycin and 100 U/ml penicillin
(HyClone™; Cytiva) and maintained in a CO2 incubator at
37°C. Upon reaching 80–90% confluency as observed under a
microscope, cells were digested with 1 ml of trypsin containing
0.25% EDTA at 37°C for 1–3 min. Digestion was monitored
microscopically and terminated immediately when cells became
rounded and detached from the culture surface.
Drugs and antibodies
Sch B was purchased from Merck KGaA and dissolved in
DMSO (Merk KGaA) to create a 100 mmol/l stock solution. This stock
solution was further diluted with culture media to obtain the
desired concentrations. The control groups received treatment with
equivalent volumes of DMSO. N-acetyl-L-cysteine (NAC) was acquired
from BD Biosciences, 2′,7′-dichlorofluorescein diacetate (DCFH-DA)
was purchased from Merck KGaA and Bay 11–7082 was purchased from
Merck KGaA. Detailed information for all primary and secondary
antibodies used in this study, including antibody names, catalog
numbers, dilutions and suppliers, is provided in Table I.
 | Table I.Details of antibodies. |
Table I.
Details of antibodies.
| Antibody name | Cat. no. | Dilution | Supplier |
|---|
| Primary
antibodies |
|
|
|
|
BIP | 3183 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
CHOP | 2895 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
XBP1s | 12782 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
Bax | 5023 | 1:1,000 | Cell Signaling
Technology, Inc. |
| p-p38
MAPK (Thr180/Tyr182) | 4511 | 1:1,000 | Cell Signaling
Technology, Inc. |
| p38
MAPK | 8690 | 1:1,000 | Cell Signaling
Technology, Inc. |
| p-p65
NF-κB (Ser536) | 3033 | 1:1,000 | Cell Signaling
Technology, Inc. |
| p65
NF-κB | 8242 | 1:1,000 | Cell Signaling
Technology, Inc. |
| p-IκBα
(Ser32) | 2859 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
IκBα | 4814 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
IL-6 | 12153 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
IL-8 | 94407 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
TNF-α | 6945 | 1:1,000 | Cell Signaling
Technology, Inc. |
|
GAPDH | 5174 | 1:5,000 | Cell Signaling
Technology, Inc. |
| Secondary
antibodies |
|
|
|
|
Anti-rabbit IgG,
HRP-linked | 7074 | 1:3,000 | Cell Signaling
Technology, Inc. |
|
Anti-mouse IgG,
HRP-linked | 7076 | 1:3,000 | Cell Signaling
Technology, Inc. |
|
AP-labeled Anti-rabbit
IgG | WB0120 | 1:5,00 | Vigorous
Biotechnology Beijing Co., Ltd. |
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from treated HuccT1 cells
using TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.). Subsequently, 1 µg of RNA from each sample was
used for complementary DNA synthesis using the PrimeScript™ RT
reagent Kit with gDNA Eraser (Takara Bio, Inc.), according to the
manufacturer's protocol. For RT-qPCR, TB Green® Premix
Ex Taq™ (Takara Bio, Inc.) was used on a CFX Connect Real-Time PCR
Detection System (Bio-Rad Laboratories, Inc.). The thermocycling
conditions were as follows: Initial denaturation at 95°C for 30
sec, followed by 45 cycles of denaturation at 95°C for 5 sec and
combined annealing/extension at 58°C for 30 sec. All reactions were
performed in triplicate. The relative expression levels of target
genes were normalized to GAPDH and calculated using the
2−ΔΔCq method (23). The
primer sequences are listed in Table
II.
 | Table II.Primer sequences. |
Table II.
Primer sequences.
| Target gene | Primer sequence (5′
to 3′) |
|---|
| GAPDH-FP |
GAAGGTGAAGGTCGGAGTC |
| GAPDH-RP |
GAAGATGGTGATGGGATTTC |
| XBP1s-FP |
AGGAGTTAAGACAGCGCTTGGGG |
| XBP1s-RP |
AATACCTGCACCTGCTGCGGACTCAGCAGA |
| BIP-FP |
TGACATTGAAGACTTCAAAGCT |
| BIP-RP |
CTGCTGTATCCTCTTCACCAGT |
| CHOP-FP |
CAACTGCAGAGATGGCAGCTGA |
| CHOP-RP | CTGATGCTCCCAATT
GTTCAT |
| IL6-FP |
TCAGGGATGCAATGCCACTT |
| IL6-RP |
TGCAGAAGAGAGCCAACCAA |
| IL8-FP |
ACAAGCTTCTAGGACAAGAGCC |
| IL8-RP |
ACTTCTCCACAACCCTCTGC |
| TNFα-FP |
CGAGTGACAAGCCTGTAGC |
| TNFα-RP |
CCTTCTCCAGCTGGAAGAC |
Western blot analysis
HuccT1 cells were treated with various
concentrations of Sch B (0, 40, 80 and 160 µM) for 48 h at 37°C.
After treatment, cells were harvested and lysed on ice using RIPA
buffer (Beyotime Biotechnology) supplemented with a protease
inhibitor cocktail (Roche Applied Science) for 15 min. The lysates
were centrifuged at 13,500 × g for 30 min at 4°C to collect the
supernatant. Protein concentration was determined using the BCA
assay. Subsequently, 60 µg of total protein per sample was
separated by SDS-PAGE on 10% polyacrylamide gels and
electrophoretically transferred onto PVDF membranes
(MilliporeSigma). Following transfer, the membranes were blocked
with 5% skim milk prepared in Tris-buffered saline with Tween-20
(TBST) for 2 h at room temperature. The membranes were then
incubated overnight at 4°C with the indicated primary antibodies
(Table I). After incubation, the
membranes were washed three times for 10 min each with TBST. A
total of two distinct detection systems were employed: Horseradish
Peroxidase (HRP)-chemiluminescence detection: For HRP-based
detection, the membranes were incubated with appropriate
HRP-conjugated secondary antibodies for 1 h at room temperature.
After washing with TBST, protein bands were visualized using an
enhanced chemiluminescence substrate and imaged with a Gel Doc 2000
system (Bio-Rad Laboratories, Inc.). Alkaline Phosphatase
(AP)-colorimetric detection: For AP-based colorimetric detection,
an AP western detection kit (Vigorous Biotechnology Beijing Co.,
Ltd.) was used. After primary antibody incubation and washing, the
membranes were probed with AP-conjugated secondary antibodies for
30 to 120 min at room temperature. The membranes were then
thoroughly washed 3–5 times with TBST to remove any phosphate
residues. A color development solution was freshly prepared
immediately before use by adding 0.1 ml of
5-bromo-4-chloro-3-indolyl phosphate (BCIP) and 0.1 ml of nitroblue
tetrazolium (NBT) per 10 ml of reaction buffer (Vigorous
Biotechnology, Beijing Co., Ltd.). After removing the wash buffer,
sufficient development solution was added to completely cover the
membrane, followed by incubation at room temperature for 5 to 15
min in the dark, with close monitoring of band intensity. The
reaction was stopped by rinsing the membrane with distilled water
once the target bands were clearly visible with minimal background.
The membrane was air-dried and imaged directly.
Measurement of ROS production
HuccT1 cells were prepared as single cell
suspensions and then seeded in 6-well plates at a density of
1×106 cells per well in 100 µl of culture medium.
Subsequently, the cells were treated with various concentrations of
Sch B (0, 40 and 160 µM) for 24 h at 37°C in a CO2
incubator. DCFH-DA was diluted with serum-free medium at a ratio of
1:1,000 to reach a final concentration of 10 µmol/l. The CCA cells
were incubated with DCFH-DA at 37°C for 20 min. Following three
washes with PBS, the cells were stimulated using a positive control
for ROS. Finally, the excitation and emission wavelengths for the
fluorescence enzyme were adjusted to 488 and 525 nm,
respectively.
Lactate dehydrogenase (LDH) release
assay
An LDH release assay was performed according to the
manufacturer's protocol (Beyotime Biotechnology). The amount of
color formed is proportional to the number of lysed cells. Cells
treated with DMSO or 160 µmol/l Sch B + different concentrations of
NAC (0, 0.5, 1, 3, 6 µmol/l) were seeded in a 96-well plate. After
24 h, LDH levels were determined by analyzing the amount of LDH
released into the cell culture supernatant. Absorbance signals at
490 nm were obtained using a microplate reader. To determine the
percentage of LDH release, the experimental LDH release quantities
were calculated relative to the control LDH release quantities, as
stated in the provided instructions.
Cell viability assay by calcein
AM-PI
HuccT1 cells were treated with Sch B (0 and 160 µM)
in the presence or absence of various concentrations of NAC (0,
0.5, 1, 3 and 6 µM) for 24 h at 37°C. After treatment, cells were
incubated with 2 µM calcein-AM (CAS 148504-34-1; MedChemExpress)
for 20 min at 37°C, followed by a 5 min co-incubation with 4 µM
propidium iodide (PI; CAS 25535-16-4; MedChemExpress) at 37°C.
Fluorescence images were captured using a Cytation 3 Cell Imaging
Multi-Mode Reader (Agilent Technologies, Inc.) with excitation at
488 nm and emission recorded at 525 nm for calcein-AM.
Dual luciferase reporter gene
experiment
HuccT1 cells were seeded in 24-well plates and
cultured for 24 h at 37°C in a 5% CO2 atmosphere prior
to transfection. Cells were co-transfected with 100 ng of reporter
plasmid (pGL4.2–3×AP-1-Luc or pGL4.2-NF-κB-Luc, both from Promega
Corporation) and 1 ng of internal control plasmid (pRL-TK, Promega
Corporation), corresponding to a mass ratio of 100:1
(reporter:internal control), using Lipofectamine® 2000
(Invitrogen; Thermo Fisher Scientific, Inc.). Transfection was
performed at 37°C, and the medium was replaced with fresh culture
medium 4 h post-transfection. At 24 h post-transfection, cells were
treated with various concentrations of Sch B (0, 10, 20, 40, 80 and
160 µM) for an additional 24 h at 37°C. After treatment, the
culture medium was removed, and cells were lysed with Reporter Gene
Cell Lysis Buffer (Beyotime Biotechnology). A 20 µl aliquot of cell
lysate was used to measure firefly and Renilla luciferase
activities sequentially using the Dual Luciferase Reporter Gene
Assay Kit II (cat. no. RG029; Beyotime Biotechnology) on a
luminometer. Relative luciferase activity was calculated as the
ratio of firefly luminescence to Renilla luminescence.
Colony formation assay
After cell counting, the HuccT1 cells were diluted
to 1×105 cells/ml, then 1×104 cells/ml was
inoculated on a 6-well plate (100 µl/well). After inoculation,
complete medium was added (2 ml/well). After 1 week of culture at
37°C in 5% CO2, the medium was removed and cells were
rinsed twice with PBS or normal saline for 10 sec each, fixed with
4% paraformaldehyde for 15 min at room temperature, and washed
twice with PBS twice for 10 sec each. Subsequently, 200 µl of
crystal violet staining solution was added to cover the bottom of
each well and incubation for 20 min at room temperature. The 6-well
plate was then rinsed under running water for 10 sec, air-dried,
and the number of colonies (containing >50 cells) was counted
manually.
Immunofluorescent staining
HuccT1 cells were cultured on circular coverslips in
6-well plates, fixed with 4% paraformaldehyde for 30 min at room
temperature, and then permeabilized with 0.3% Triton X-100 (Merck
KGaA) for 15 min at room temperature. Following permeabilization,
cells were incubated in a solution containing 5% bovine serum
albumin (Merck KGaA) for 40–60 min at room temperature. After
washing three times with PBS, the cells were incubated overnight at
4°C with the corresponding primary antibodies (dilution 1:100;
Table I). The next day, cells were
washed three times with PBS and incubated with fluorescence-labeled
secondary antibodies (dilution 1:200; Table I) for 1 h at room temperature in the
dark. Subsequently, cells were stained with Hoechst 33342 (cat. no.
HY-D0983; MedChemExpress) for 15 min at room temperature in the
dark. The coverslips were then mounted onto glass slides, and
fluorescence images were captured using a Nikon fluorescence
microscope (Nikon Corporation).
Statistical analysis
The data are presented as mean ± SD from at least
three independent experiments. Statistical analyses were performed
using SPSS 23.0 (IBM Corp.). All datasets were confirmed to follow
a normal distribution by the Shapiro-Wilk test. For comparisons
among multiple experimental groups with a single control group,
homogeneity of variances was assessed using Brown-Forsythe test.
When the assumption of homogeneity of variances was met, ANOVA
followed by Dunnett's post hoc test was applied to compare each
experimental group with the control group. In cases where variances
were heterogeneous, Welch's ANOVA followed by Dunnett T3 post hoc
test was employed. The specific statistical test used for each
dataset is indicated in the corresponding figure legend. A value of
*P<0.05, **P<0.01 and ***P<0.001 was considered to
indicate a statistically significant difference.
Results
Identification of core targets and
validation by molecular docking
Potential targets of Sch B and CCA were retrieved
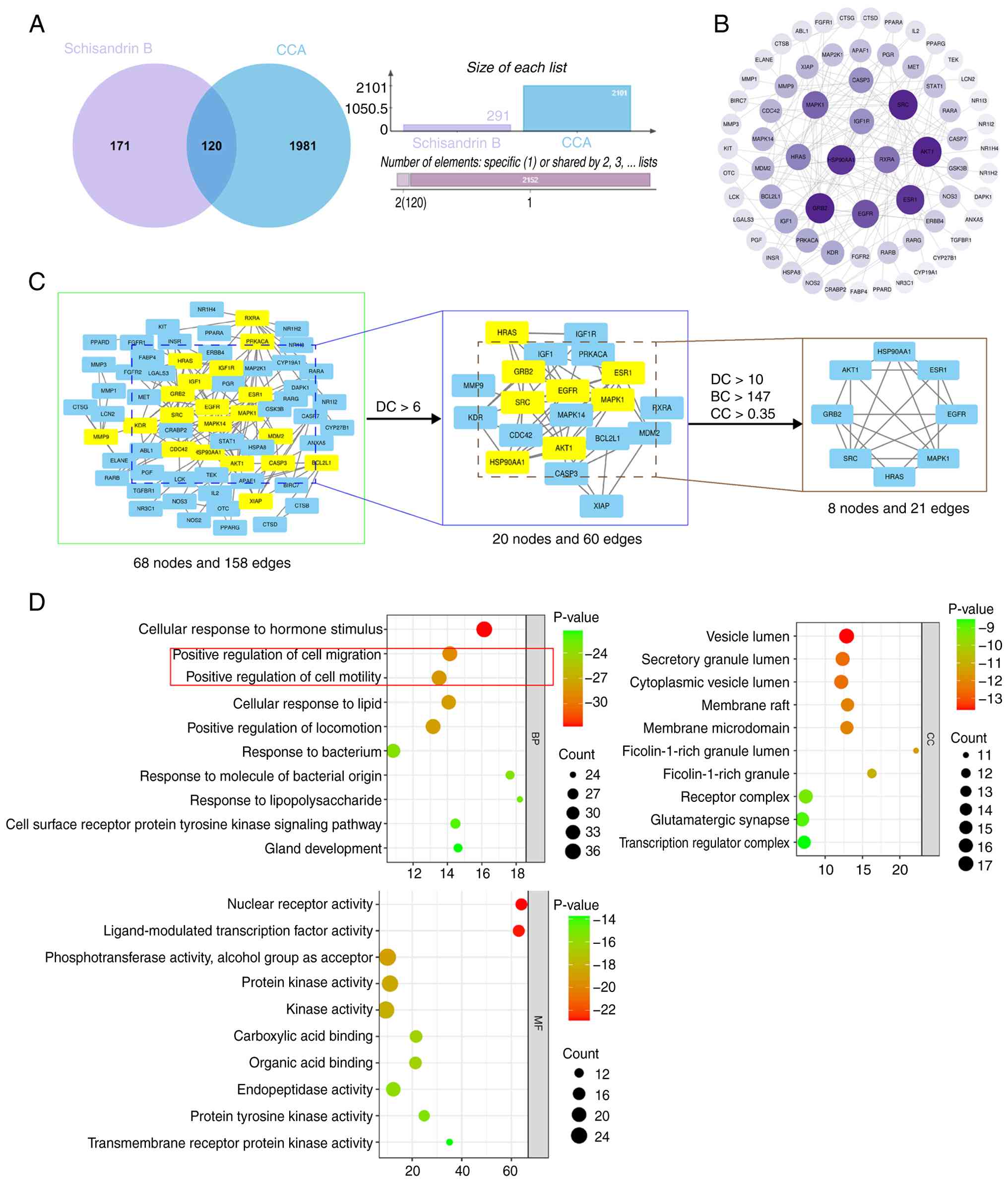
from relevant databases. A Venn diagram revealed 120 overlapping
targets between Sch B and CCA (Fig.
1A). A PPI network was constructed, comprising 68 nodes and 158
edges, with node size proportional to the degree value (Fig. 1B). Using a DC threshold >6, a
subnetwork with 20 nodes and 60 edges was obtained. Further
applying criteria of DC >10, BC >147 and CC >0.35, a core
network of 8 nodes and 21 edges was identified. A total of eight
core targets were ultimately selected: MAPK1, EGFR, ESR1, HSP90AA1,
AKT1, GRB2, SRC and HRAS (Fig.
1C).
GO enrichment analysis of the 120 overlapping
targets was performed, covering BP, cellular component and MF
categories. The top 10 enriched terms from each category are
displayed in a bubble chart (Fig.
1D), suggesting that Sch B may regulate the migration and
motility of CCA cells.
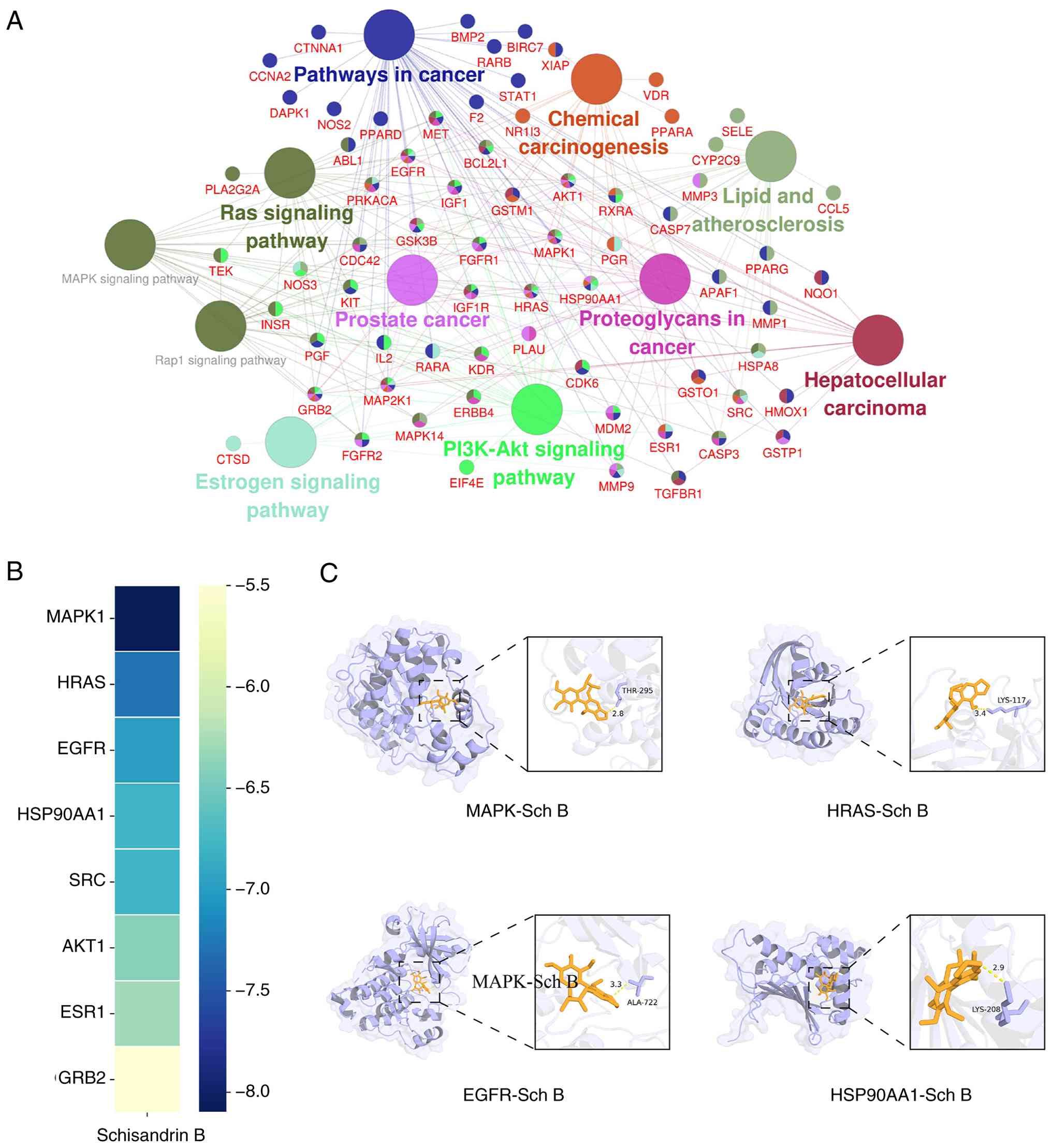
KEGG pathway analysis was conducted using the ClueGO
plugin in Cytoscape 3.8.0, with thresholds set to include pathways
containing ≥16 genes and a κ score >0.6. A total of 11
significantly enriched pathways were identified, primarily
associated with ‘Pathways in cancer’ and the ‘MAPK signaling
pathway’ (Fig. 2A). Molecular
docking was performed between Sch B and the top 8 core target
proteins (MAPK1, EGFR, ESR1, HSP90AA1, AKT1, GRB2, SRC and HRAS)
identified from the PPI network. The lowest binding energies for
all ligand-receptor complexes were below-5 kcal/mol, indicating
spontaneous binding and stable conformations (Table III). Among them, MAPK1 exhibited
the strongest binding affinity with Sch B. The binding modes of Sch
B with MAPK1, HRAS, EGFR and HSP90AA1 were visualized using PyMOL
3.7.1 (Fig. 2B and C).
 | Table III.Molecular docking binding affinity of
Sch B with core targets. |
Table III.
Molecular docking binding affinity of
Sch B with core targets.
| Compound | Lowest binding
energy (kcal/mol) |
|---|
| MAPK1 | −8.1 |
| EGFR | −7 |
| HRAS | −7.3 |
| HSP90AA1 | −6.8 |
| AKT1 | −6.4 |
| GRB2 | −5.5 |
| SRC | −6.8 |
| ESR1 | −6.3 |
Sch B induced a dose-dependent
increase in ROS levels in CCA cells
Based on the KEGG pathway enrichment analysis, which
indicated strong associations between the predicted MAPK signaling
pathway and key BPs such as chemical carcinogenesis, and supported
by established literature evidence highlighting the pivotal role of
oxidative stress as a key mediator in these pathways, the present
study was guided to investigate the involvement of ROS in the
mechanism of Sch B. Guided by these predictions, the biological
effects were subsequently validated through a series of in
vitro experiments. CCA cells were exposed to various
concentrations of Sch B, ranging from 0 to 160 µmol/l, to assess
the mRNA expression levels of BIP, CHOP and XBP1s. The results
showed an increasing trend, indicative of a dose-dependent effect
(Fig. 3A-C). Western blot analysis
further confirmed the elevated expression levels of BIP, CHOP and
XBP1s, showing an upward trend (Fig.
3D).
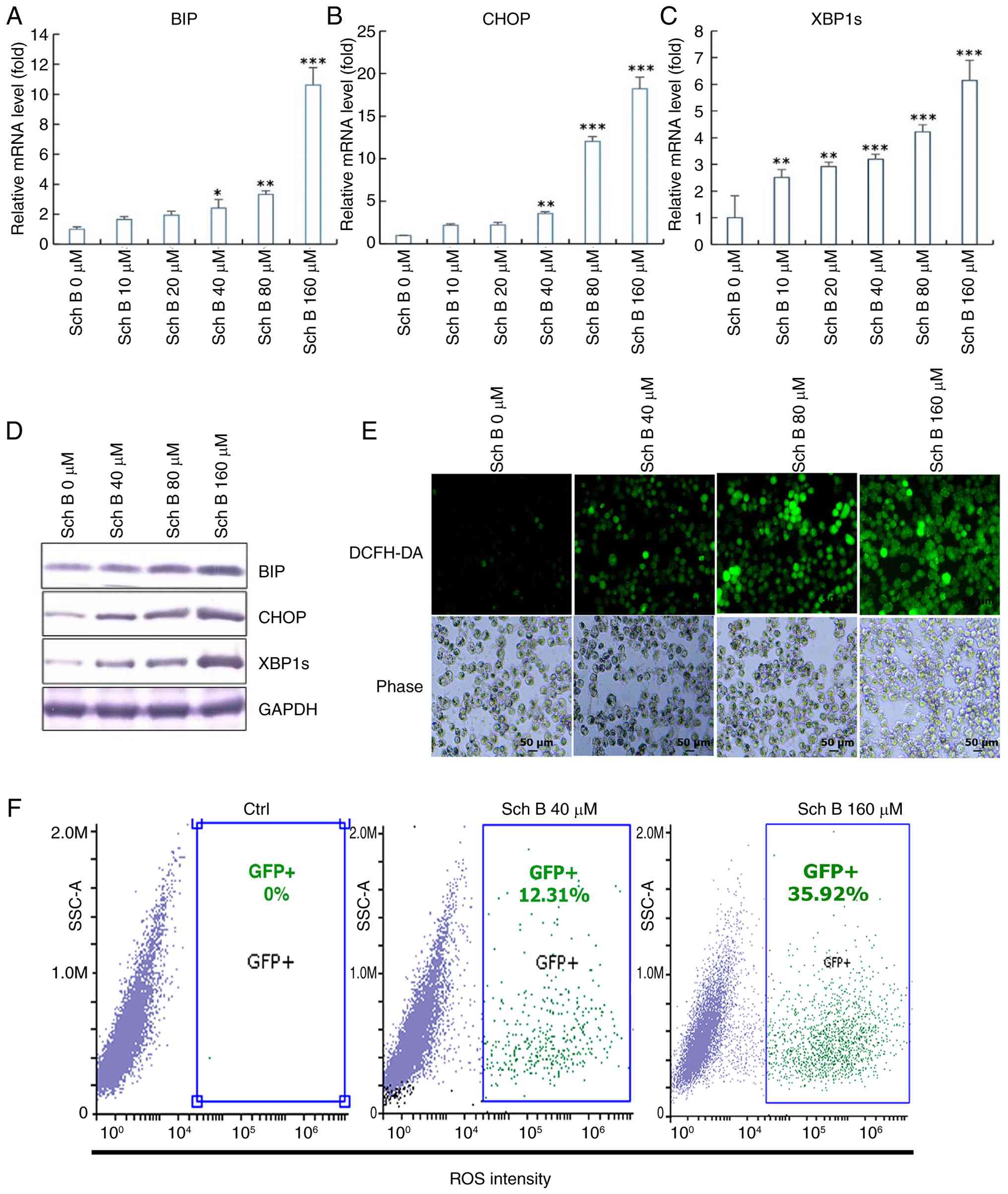 | Figure 3.Sch B induced a dose-dependent
increase in ROS levels in CCA cells. (A) BIP, (B) CHOP and (C)
XBP1s expressions were determined by reverse-transcription
quantitative PCR in CCA cells treated with different concentrations
of Sch B (0, 10, 20, 40, 80, 160 µmol/l), (D) BIP, CHOP and XBP1s
expression levels were determined by western blotting in CCA cells
treated with different concentrations of Sch B (0, 40, 80, 160
µmol/l). (E) ROS levels in CCA cells treated with different
concentrations of Sch B (0, 40, 80, 160 µmol/l) were measured using
the ROS probe DCFH-DA. (F) Flow cytometry analysis of ROS levels in
CCA cells following treatment with various Sch B concentrations (0,
40, 160 µmol/l). Statistical analysis was performed using ANOVA and
Dunnett's post hoc test. *P<0.05, **P<0.01, ***P<0.001.
CCA, cholangiocarcinoma; Sch B, Schisandrin B; Ctrl, control; ROS,
reactive oxygen species. |
To support these findings, a ROS probe, DCFH-DA and
flow cytometry were used to measure ROS levels in CCA cells. The
results demonstrated a dose-dependent increase in ROS levels
(Fig. 3E and F). Collectively,
these results suggest that Sch B induces a dose-dependent increase
in ROS levels within CCA cells, offering valuable insights into its
molecular impact on cellular responses.
NAC counteracted the upregulatory
effect of Sch B on ROS expression
Treatment with 160 µmol/l Sch B led to a notable
increase in the expression levels of BIP, CHOP and XBP1s in CCA
cells. The addition of NAC showed a dose-dependent reduction in the
expression levels of BIP, CHOP and XBP1s in the experimental group
(Fig. 4A and B).
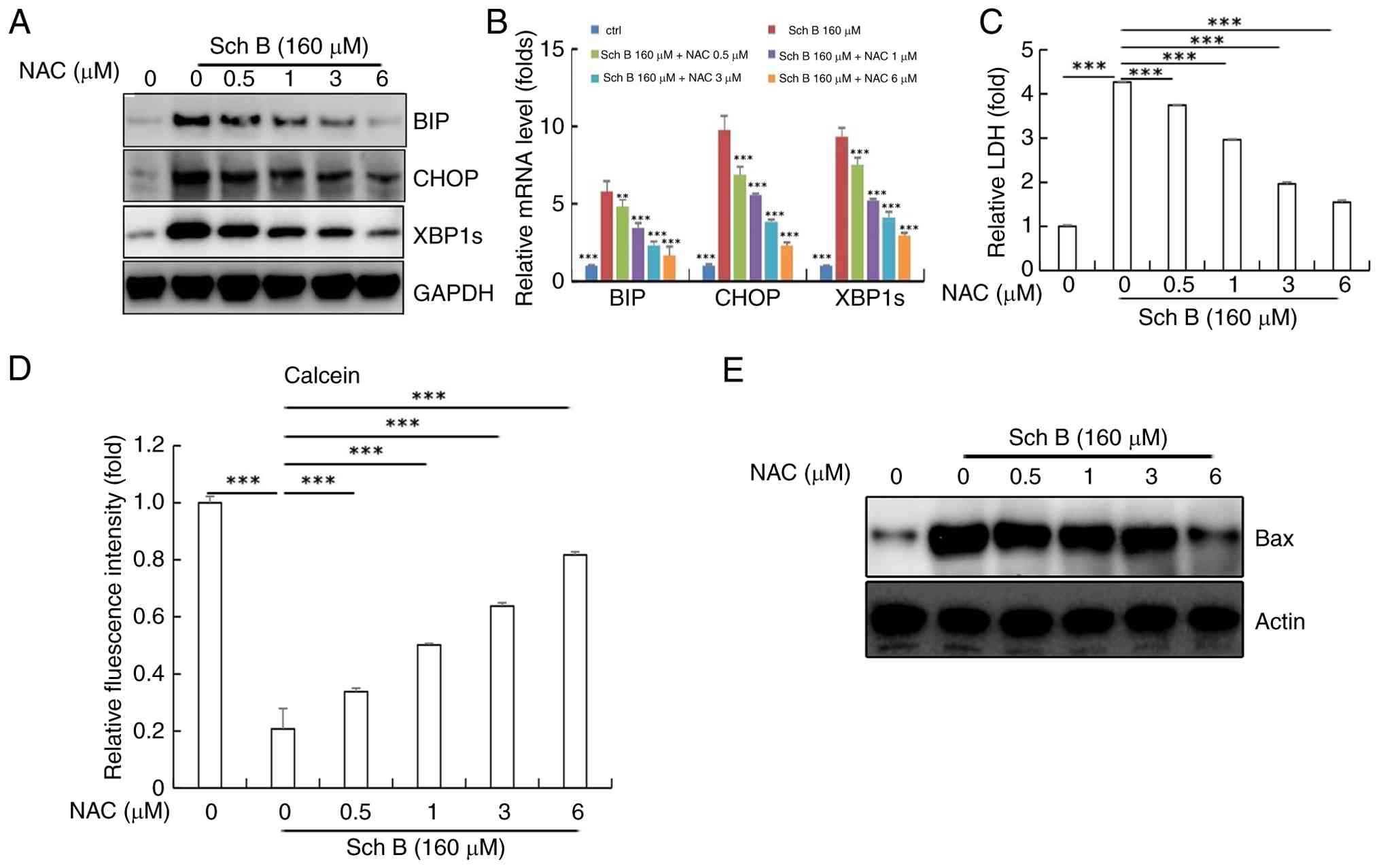 | Figure 4.NAC counteracted the upregulatory
effect of Sch B on ROS expression. After treatment with 160 µmol/l
Sch B + different concentrations of NAC (0, 0.5, 1, 3, 6 µmol/l).
BIP, CHOP and XBP1s expression determined by (A) western blotting
and (B) reverse transcription-quantitative PCR in CCA cells. (C)
LDH activity in cell culture medium. (D) Cell activity levels
detected by the Calcein AM-PI live cell staining. (E) Western blot
analysis of the expression levels of Bax in CCA cells. Statistical
analysis was performed using ANOVA and Dunnett's post hoc test.
**P<0.01, ***P<0.001. ROS, reactive oxygen species; CCA,
cholangiocarcinoma; NAC, N-acetyl-L-cysteine; ctrl, control; Sch B,
Schisandrin B; LDH, lactate dehydrogenase. |
It is well known that NAC, as an antioxidant, can
reduce ROS levels and protect cell membrane integrity (24,25),
thereby lowering LDH release from cells. With increasing
concentrations of NAC, the activity of LDH in the culture medium
decreases (Fig. 4C), A reduction in
Bax expression was observed only at the highest concentration of
NAC (6 µM). Western blotting results in Fig. 4E are representative images and were
not subjected to densitometric quantification. In parallel,
Calcein-AM/PI staining showed that Sch B-induced cytotoxicity was
alleviated by NAC co-treatment, as evidenced by increased green
fluorescence intensity (Fig. 4D).
Consequently, these results show that NAC may counter the
inhibitory effects of Sch B on CCA cells proliferation and
facilitate apoptosis by mitigating ROS upregulation.
MAPK signaling pathway inhibition by
Sch B
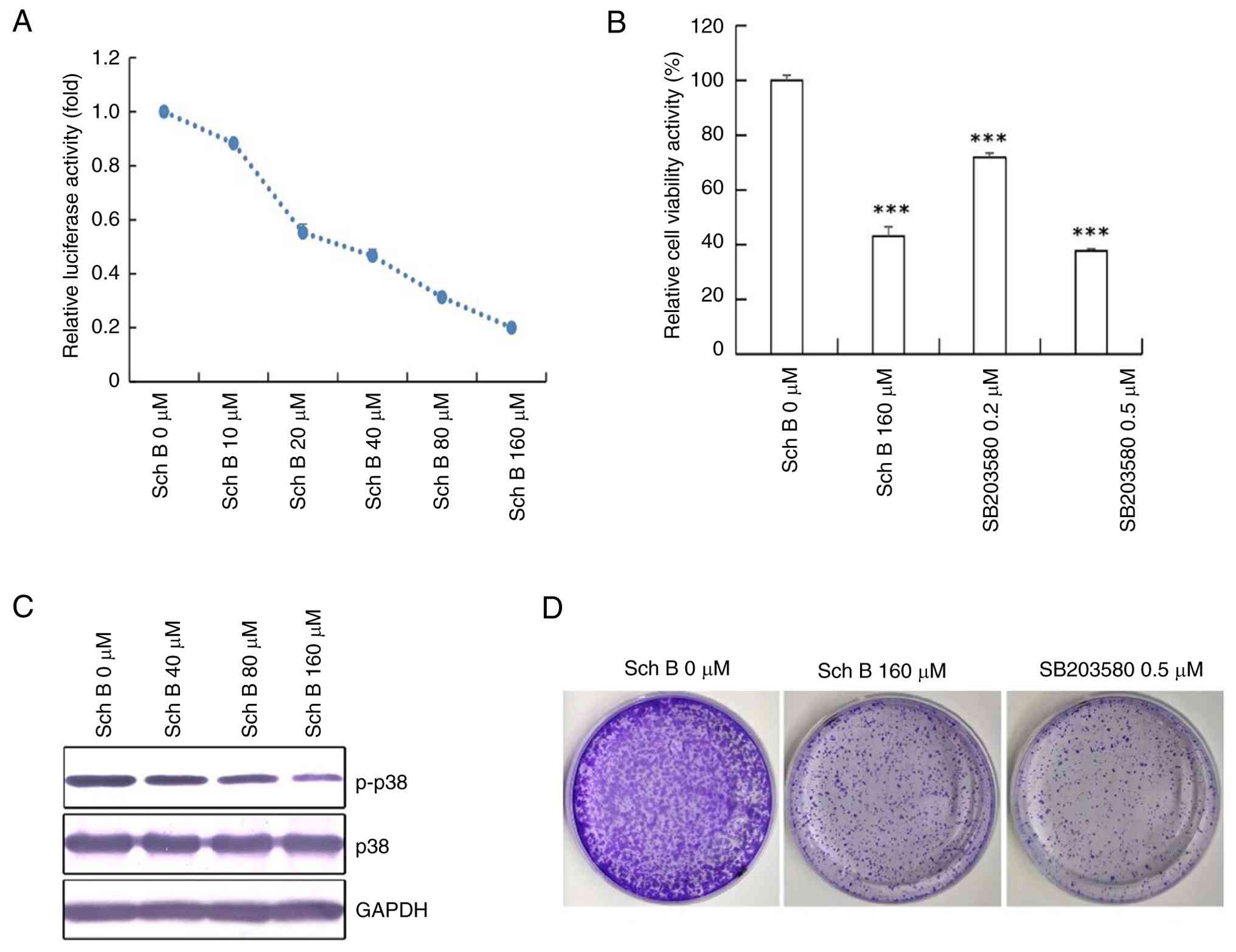
CCA cells treated with different concentrations of
Sch B ranging from 0 to 160 µmol/l showed a dose-dependent decrease
in AP-1 expression, a key regulatory component of the MAPK
signaling pathway, as measured by a double luciferase reporter
assay (Fig. 5A). Additionally,
western blot analysis indicated a decreasing trend in p-p38
expression with higher Sch B concentrations, further demonstrating
inhibition of the MAPK signaling pathway (Fig. 5C).
Treatment with 160 µmol/l Sch B led to significant
inhibition of CCA cell activity (Fig.
5B). The subsequent addition of SB203580, a p38 MAPK inhibitor,
at 0.2 and 0.5 µmol/l concentrations resulted in a notable decrease
in CCA cell activity compared with the control group (Fig. 5B), reflecting the effects seen with
Sch B.
Furthermore, the anti-proliferative effect of Sch B
on CCA cells was evidenced by a trend of reduced colony formation
at 160 µM, an effect qualitatively similar to that observed with
0.5 µM SB203580 (representative images shown in Fig. 5D). These findings together highlight
the effectiveness of Sch B in impeding the MAPK signaling pathway
and reducing CCA cell proliferation.
Sch B inhibits NF-κB signaling pathway
expression
Sch B exhibited dose-dependent suppression of the
NF-κB signaling pathway in CCA cell lines. Treatment with Sch B
(0,40, 80, 160 µmol/l) led to a corresponding decrease in the
expression levels of IL-6, IL-8 and TNF-α. Although the 40 µmol/l
treatment group generally did not show statistically significant
differences compared to the control group, a trend of reduction was
observed with increasing concentrations (Fig. 6A-C). The suppressive effect on NF-κB
expression was shown by a dose-dependent reduction in NF-κB levels,
as shown by a double luciferase assay (Fig. 6D).
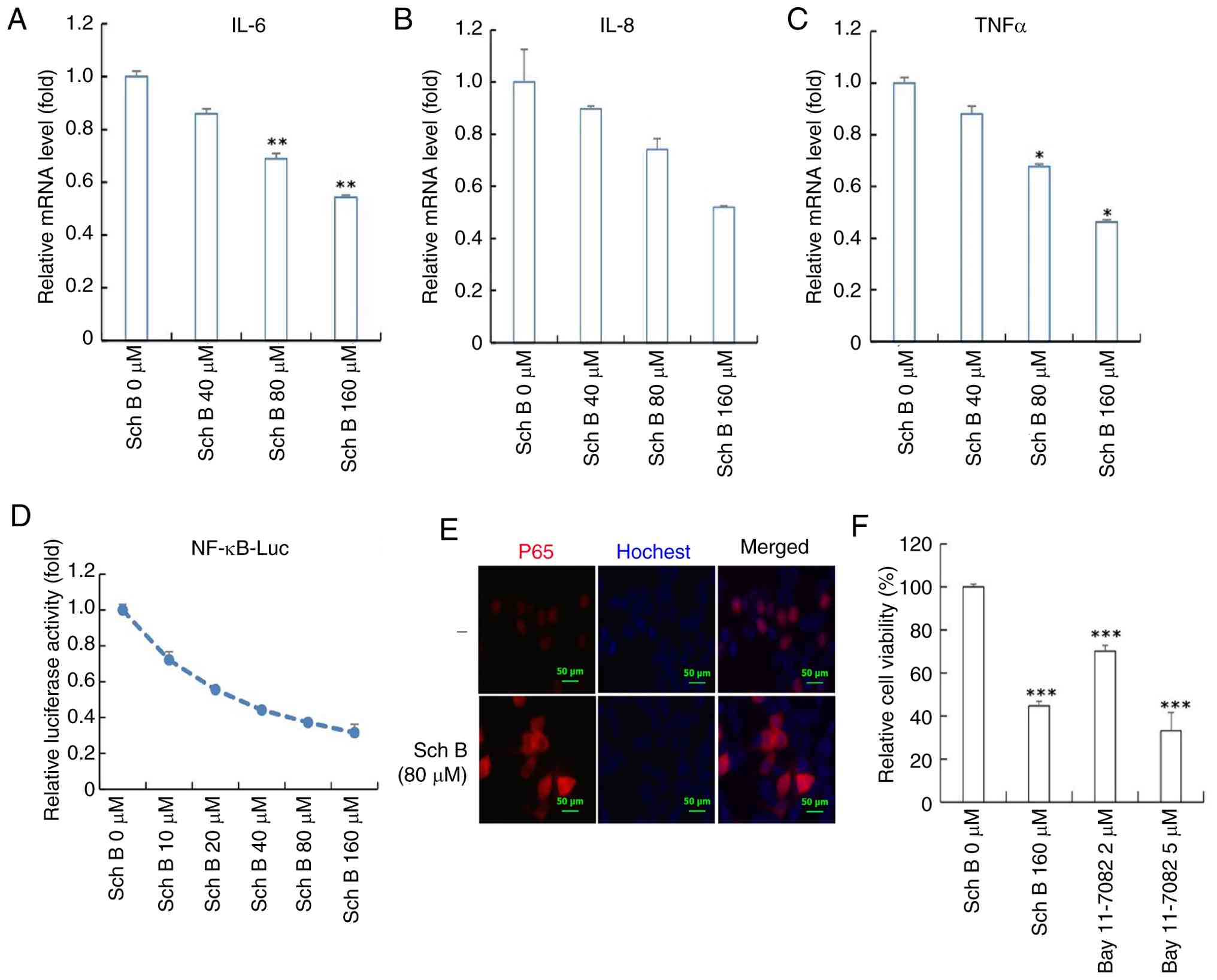 | Figure 6.Sch B inhibits NF-κB signaling
pathway expression. mRNA expression levels of (A) IL-6, (B) IL-8
(C) and TNF-α were measured in CCA cells treated with different
concentrations of Sch B (0, 40, 80, 160 µmol/l). Statistical
analysis was performed using Welch's ANOVA followed by Dunnett's T3
post hoc test. (D) NF-κB expression in CCA cells was evaluated
using a double luciferase assay with different concentrations of
Sch B (0, 10, 20, 40, 80, 160 µmol/l). (E) Hoechst
immunofluorescence staining of p65 expression in CCA cells. (F) CCA
cell activity was assessed after treatment with Sch B and Bay
11–7082, a targeted NF-κB inhibitor. Statistical analysis was
performed using ANOVA and Dunnett's post hoc test. *P<0.05,
**P<0.01, ***P<0.001. Sch B, Schisandrin B; CCA,
cholangiocarcinoma. |
In the experimental group, 160 µmol/l Sch B notably
reduced CCA cell activity. To directly and visually assess the
effect of Sch B on p65 translocation, the present study performed
immunofluorescence staining for p65. The results demonstrate
nuclear accumulation of p65 in control cells (without Sch B
treatment), whereas in Sch B-treated cells, p65 was predominantly
retained in the cytoplasm, with correspondingly lower nuclear
levels (Fig. 6E). Bay 11–7082 is a
selective NF-κB inhibitor that suppresses IκBα phosphorylation
(26), thereby preventing IκBα
degradation and NF-κB nuclear translocation, which ultimately
inhibits downstream NF-κB-dependent gene transcription.
Accordingly, CCA cell activity was significantly reduced following
the addition of Bay 11–7082 in a dose-dependent manner, with both 2
and 5 µM concentrations showing significant decreases compared with
the control group (Fig. 6F). These
results highlight the effectiveness of Sch B in inhibiting the
NF-κB signaling pathway, resulting in the suppression of CCA cell
activity.
Discussion
According to reports, over the past few decades, the
incidence of CCA, a rare but highly malignant disease, has rapidly
increased (1,2). It has become the second most common
primary liver cancer in humans, following hepatocellular carcinoma,
and is emerging as a major global health issue (3). Currently, surgical resection remains
the primary treatment for CCA. However, due to the insidious onset
of the disease and tendency to metastasize, only 20–40% of patients
with potentially resectable disease undergo surgery. Consequently,
the majority of patients require systemic treatment, including
systemic chemotherapy with drugs such as gemcitabine, cisplatin,
5-fluorouracil and capecitabine (27). However, the development of drug
resistance to chemotherapy poses an additional challenge to the
treatment of CCA. Therefore, it is important to find new and safe
drugs for treating CCA.
Currently, an increasing number of researchers are
focusing on traditional Chinese medicine extracts, which are
considered to contain various active ingredients capable of acting
on multiple molecular targets and signaling pathways simultaneously
(28–30). These extracts have relatively low
side effects and are suitable for long-term use. Among these
compounds, Sch B has been demonstrated to exert therapeutic effects
in various malignant tumors (7–10). One
report indicates that Sch B may inhibit the viability and
proliferation of gallbladder cancer cells and promote apoptosis,
highlighting its potential as a therapeutic agent for gallbladder
cancer (8).
The present study first employed network
pharmacology to predict the potential targets of Sch B in CCA.
Functional enrichment analysis revealed significant enrichment of
these targets in cancer-related pathways and the MAPK signaling
pathway. Molecular docking further confirmed the strong binding
affinity between Sch B and core targets, including MAPK.
The present in vitro study demonstrated that
Sch B exerts its anti-CCA effects, at least in part, by modulating
ROS levels. ROS play a dual role in tumorigenesis, acting as
signaling molecules at low to moderate concentrations to promote
tumor survival via pathways such as MAPK/ERK1/2, p38MAPK, JNK and
PI3K/Akt, leading to activation of downstream effectors including
NF-κB, MMP and vascular endothelial growth factor. By contrast,
high concentrations of ROS trigger apoptotic pathways (31–33).
Results of the present study revealed that Sch B treatment
dose-dependently upregulated the expression of endoplasmic
reticulum stress-related molecules (BIP, CHOP and XBP1s) and
significantly increased ROS levels in CCA cells, while also
inhibiting proliferation and promoting the expression of the
pro-apoptotic protein Bax. The ROS scavenger NAC reversed these
effects, restoring cell viability and reducing apoptosis,
confirming that Sch B primarily exerts its anti-CCA activity
through ROS elevation.
Studies have established that ROS influence multiple
signaling pathways, notably the MAPKs and NF-κB transduction
cascades (34,35). ROS play a key role in activating the
MAPK signaling pathway (34) and
are involved in p38 activation (35). Upon activation by oxidative stress,
the MAPK signaling pathway compromises cellular anti-apoptotic
capacity, leading to caspase activation and apoptosis induction.
Additionally, p38MAPK has been shown to modulate the
transcriptional activity of NF-κB, enabling NF-κB p65 nuclear
translocation and subsequent activation of downstream signaling
pathways (36,37). The NF-κB signaling pathway promotes
CCA cell proliferation, metastasis and invasion (38).
In the present study, increasing concentrations of
Sch B led to decreased p-p38 expression and reduced AP-1 activity,
accompanied by inhibited cell proliferation. Concurrently,
expression of p-p65 and p-IκBα increased, while downstream
inflammatory factors IL-6, IL-8 and TNF-α decreased.
Immunofluorescence staining revealed reduced nuclear translocation
of p65, indicating suppression of the NF-κB pathway. The use of the
p38MAPK inhibitor SB203580 and the NF-κB inhibitor Bay 11–7082
further suppressed CCA cell viability, suggesting a role for these
pathways in the mechanism of Sch B.
The in vitro findings of the present study
provide valuable mechanistic insights, however, several important
limitations must be acknowledged. Firstly, the inherent constraints
of cell-based models cannot fully replicate the complex in
vivo tumor microenvironment, including key factors such as
cell-cell interactions, tissue architecture and pharmacokinetic
processes such as drug metabolism, distribution and clearance, all
of which may substantially influence therapeutic outcomes.
Secondly, and more specifically to the present study, the
mechanistic conclusions are derived primarily from experiments
using a single human CCA cell line (HuccT1). This approach does not
capture the known heterogeneity of CCA among patients and the
absence of a parallel normal cholangiocyte control line precludes a
definitive assessment of the selective toxicity of Sch B against
cancer cells. Consequently, the generalizability of the present
findings is currently limited. To address these points, future work
will extend these investigations to other representative CCA cell
lines (for example, RBE and TFK-1) and include normal human
intrahepatic biliary epithelial cells (for example, HIBEC) for a
thorough evaluation of broader applicability and selectivity.
Ultimately, in vivo validation using appropriate animal
models will be essential to confirm the anti-tumor efficacy and
biosafety of Sch B and to pave the way for its potential clinical
translation.
In conclusion, the present findings demonstrate that
Sch B may suppress proliferation and induces apoptosis in CCA cells
by elevating intracellular ROS levels, mechanistically associated
with the modulation of the p38MAPK/NF-κB signaling pathway. As a
natural compound, Sch B exhibits multi-target intervention
characteristics, underscoring its potential for clinical
translation. Future studies will focus on in vivo validation
of its antitumor efficacy and biosafety, and explore combination
strategies with conventional cytotoxic agents to elucidate
synergistic mechanisms. These efforts will provide a stronger
experimental foundation for developing Sch B as a promising
therapeutic candidate against CCA.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
All data generated in the present study may be
requested from the corresponding author.
Authors' contributions
JY and SW confirmed the authenticity of all raw
data. The conception and design of the study were carried out by
JY, SW, WL and XW. Data acquisition was performed by JY, WL and XY.
Data analysis and interpretation were conducted by WL, XY and QH.
Manuscript writing and/or revision were undertaken by JY and WL.
Administrative, technical or material support was provided by SW
and XY. Study supervision was managed by SW. All authors read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Li X, Guan R and Zhang S: Factors
contributing to the high malignancy level of cholangiocarcinoma and
its epidemiology: Literature review and data. Biology (Basel).
14:3512025.PubMed/NCBI
|
|
2
|
Ilyas SI, Khan SA, Hallemeier CL, Kelley
RK and Gores GJ: Cholangiocarcinoma-evolving concepts and
therapeutic strategies. Nat Rev Clin Oncol. 15:95–111. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bridgewater J, Galle PR, Khan SA, Llovet
JM, Park JW, Patel T, Pawlik TM and Gores GJ: Guidelines for the
diagnosis and management of intrahepatic cholangiocarcinoma. J
Hepatol. 60:1268–1289. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Peery AF, Crockett SD, Murphy CC, Lund JL,
Dellon ES, Williams JL, Jensen ET, Shaheen NJ, Barritt AS, Lieber
SR, et al: Burden and cost of gastrointestinal, liver, and
pancreatic diseases in the united states: Update 2018.
Gastroenterology. 156:254–272.e11. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Razumilava N and Gores GJ:
Cholangiocarcinoma. Lancet. 383:2168–2179. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Qi L, Yu HQ, Li YQ, Jin H, Zhao DH and Xu
Y: Schidandrin B kills tumor cells by initiating apoptosis in
glioma SHG-44 cells. Chin J Integr Med. Aug 2–2016.doi:
10.1007/s11655-015-2406-9 (Epub ahead of print). View Article : Google Scholar
|
|
7
|
Li Q, Lu XH, Wang CD, Cai L, Lu JL, Wu JS,
Zhuge QC, Zheng WM and Su ZP: Antiproliferative and
apoptosis-inducing activity of schisandrin B against human glioma
cells. Cancer Cell Int. 15:122015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Xiang SS, Wang XA, Li HF, Shu YJ, Bao RF,
Zhang F, Cao Y, Ye YY, Weng H, Wu WG, et al: RETRACTED: Schisandrin
B induces apoptosis and cell cycle arrest of gallbladder cancer
cells. Molecules. 19:13235–13250. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Dai X, Yin C, Guo G, Zhang Y, Zhao C, Qian
J, Wang O, Zhang X and Liang G: Schisandrin B exhibits potent
anticancer activity in triple negative breast cancer by inhibiting
STAT3. Toxicol Appl Pharmacol. 358:110–119. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Nasser MI, Han T, Adlat S, Tian Y and
Jiang N: Inhibitory effects of Schisandrin B on human prostate
cancer cells. Oncol Rep. 41:677–685. 2019.PubMed/NCBI
|
|
11
|
Zhang H, Chen Q, Dahan A, Xue J, Wei L,
Tan W and Zhang G: Transcriptomic analyses reveal the molecular
mechanisms of schisandrin B alleviates CCl4-induced liver fibrosis
in rats by RNA-sequencing. Chem Biol Interact. 309:1086752019.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sabharwal SS and Schumacker PT:
Mitochondrial ROS in cancer: Initiators, amplifiers or an Achilles'
heel? Nat Rev Cancer. 14:709–721. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Trinh VH, Nguyen Huu T, Sah DK, Choi JM,
Yoon HJ, Park SC, Jung YS and Lee SR: Redox regulation of PTEN by
reactive oxygen species: Its role in physiological processes.
Antioxidants (Basel). 13:1992024. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Caligiuri A, Becatti M, Porro N, Borghi S,
Marra F, Pastore M, Taddei N, Fiorillo C and Gentilini A: Oxidative
stress and Redox-dependent pathways in cholangiocarcinoma.
Antioxidants (Basel). 13:282023. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Srinivas US, Tan BWQ, Vellayappan BA and
Jeyasekharan AD: ROS and the DNA damage response in cancer. Redox
Biol. 25:1010842019. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Marchi S, Guilbaud E, Tait SWG, Yamazaki T
and Galluzzi L: Mitochondrial control of inflammation. Nat Rev
Immunol. 23:159–173. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Glover HL, Schreiner A, Dewson G and Tait
SWG: Mitochondria and cell death. Nat Cell Biol. 26:1434–1446.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wandee J, Srinontong P, Prawan A,
Senggunprai L, Kongpetch S, Yenjai C and Kukongviriyapan V:
Derrischalcone suppresses cholangiocarcinoma cells through
targeting ROS-mediated mitochondrial cell death, Akt/mTOR, and FAK
pathways. Naunyn Schmiedebergs Arch Pharmacol. 394:1929–1940. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Nazim UM, Yin H and Park SY: Autophagy
flux inhibition mediated by celastrol sensitized lung cancer cells
to TRAIL-induced apoptosis via regulation of mitochondrial
transmembrane potential and reactive oxygen species. Mol Med Rep.
19:984–993. 2019.PubMed/NCBI
|
|
20
|
Nogales C, Mamdouh ZM, List M, Kiel C,
Casas AI and Schmidt H: Network pharmacology: Curing causal
mechanisms instead of treating symptoms. Trends Pharmacol Sci.
43:136–150. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang P, Zhang D, Zhou W, Wang L, Wang B,
Zhang T and Li S: Network pharmacology: Towards the artificial
intelligence-based precision traditional Chinese medicine. Brief
Bioinform. 25:bbad5182023. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhao L, Zhang H, Li N, Chen J, Xu H, Wang
Y and Liang Q: Network pharmacology, a promising approach to reveal
the pharmacology mechanism of Chinese medicine formula. J
Ethnopharmacol. 309:1163062023. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Esmat A, El-Demerdash E, El-Mesallamy H
and Abdel-Naim AB: Toxicity and oxidative stress of acrylonitrile
in rat primary glial cells: Preventive effects of N-acetylcysteine.
Toxicol Lett. 171:111–118. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Du L, Empey PE, Ji J, Chao H, Kochanek PM,
Bayır H and Clark RS: Probenecid and N-Acetylcysteine prevent loss
of intracellular glutathione and inhibit neuronal death after
mechanical stretch injury in vitro. J Neurotrauma. 33:1913–1917.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Strickson S, Campbell DG, Emmerich CH,
Knebel A, Plater L, Ritorto MS, Shpiro N and Cohen P: The
anti-inflammatory drug BAY 11–7082 suppresses the MyD88-dependent
signalling network by targeting the ubiquitin system. Biochem J.
451:427–437. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wilbur HC, Soares HP and Azad NS:
Neoadjuvant and adjuvant therapy for biliary tract cancer: Advances
and limitations. Hepatology. 82:1287–1302. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang JX, Zhang MX, Yu CH, Wang SJ and
Zhang H: Targeting DNA damage: A natural product-based strategy for
inhibiting cancer progression. J Ethnopharmacol. 355:1206432026.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ma J, Zhang Y, Du J, Chen J, Sun J, Ma X,
Zeng J and Efferth T: High-fat Diet-associated digestive cancers:
Mechanisms, natural Product-based therapies, and drug development.
Phytomedicine. 150:1575742026. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
He Z, Wang Y, Han L, Hu Y and Cong X: The
mechanism and application of traditional Chinese medicine extracts
in the treatment of lung cancer and other lung-related diseases.
Front Pharmacol. 14:13305182023. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ismail T, Kim Y, Lee H, Lee DS and Lee HS:
Interplay between mitochondrial peroxiredoxins and ROS in cancer
development and progression. Int J Mol Sci. 20:44072019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li J, Lim JYS, Eu JQ, Chan AKMH, Goh BC,
Wang L and Wong AL: Reactive oxygen species modulation in the
current landscape of anticancer therapies. Antioxid Redox Signal.
41:322–341. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ebrahimi SO, Reiisi S and Shareef S:
miRNAs, oxidative stress, and cancer: A comprehensive and updated
review. J Cell Physiol. 235:8812–8825. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Imran M, Saeed F, Gilani SA, Shariati MA,
Imran A, Afzaal M, Atif M, Tufail T and Anjum FM: Fisetin: An
anticancer perspective. Food Sci Nutr. 9:3–16. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Huang M and Xin W: Matrine inhibiting
pancreatic cells epithelial-mesenchymal transition and invasion
through ROS/NF-κB/MMPs pathway. Life Sci. 192:55–61. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Lin K, Zhang Y, Shen Y, Xu Y, Huang M and
Liu X: Hydrogen sulfide can scavenge free radicals to improve
spinal cord injury by inhibiting the p38MAPK/mTOR/NF-κB signaling
pathway. Neuromolecular Med. 26:262024. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Montal R, Sia D, Montironi C, Leow WQ,
Esteban-Fabró R, Pinyol R, Torres-Martin M, Bassaganyas L, Moeini
A, Peix J, et al: Molecular classification and therapeutic targets
in extrahepatic cholangiocarcinoma. J Hepatol. 73:315–327. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Pan S, Hu Y, Hu M, Xu Y, Chen M, Du C, Cui
J, Zheng P, Lai J, Zhang Y, et al: S100A8 facilitates
cholangiocarcinoma metastasis via upregulation of VEGF through
TLR4/NF-κB pathway activation. Int J Oncol. 56:101–112.
2020.PubMed/NCBI
|