Introduction
Lung cancer is a prevalent and fatal malignancy,
with non-small cell lung cancer (NSCLC) accounting for >85% of
cases (1,2). Lung squamous cell carcinoma (LUSC)
accounts for 25–30% of NSCLC, and presents a particularly
challenging clinical scenario, with a <20% 5-year survival rate
and ~400,000 annual deaths globally (3). The low frequency of epidermal growth
factor receptor (EGFR) mutations in LUSC limits the efficacy of
EGFR tyrosine kinase inhibitors, a standard first-line treatment
for NSCLC (4). While immunotherapy
has emerged as a relatively effective strategy, tumor
heterogeneity, microenvironmental complexity and variable immune
infiltration patterns contribute to modest response rates (5,6).
Additionally, severe adverse reactions continue to impede the
widespread application of immunotherapy in patients with LUSC
(7). Therefore, there is an urgent
need to identify novel therapeutic targets and develop innovative
treatment approaches for LUSC.
Acute-phase proteins (APPs) are primarily involved
in host defense and exhibit dynamic changes in expression levels
during inflammatory responses, including those related to cancer
such as tumor cell migration, invasion and anti-apoptotic signaling
(8,9). Current literature has documented a
significant decrease in APP expression during tumor development
(10–12), yet the therapeutic potential of most
APPs remains largely unexplored. Notably, research suggests that
α-2-macroglobulin, encoded by A2M, could be a viable
therapeutic target for NSCLC (13);
however, the roles of other APPs in the progression of LUSC are
still unknown. This present hypothesis that specific APPs may
regulate the growth and advancement of LUSC, potentially offering a
novel therapeutic avenue for managing this condition.
Ferroptosis is an iron-dependent programmed cell
death mechanism that is distinct from apoptosis, necrosis and
autophagy in its morphological, biochemical and genetic
characteristics (14,15). Recent studies have indicated the
potential therapeutic utility of regulating ferroptosis in tumors
such as hepatocellular carcinoma (16) and lung cancer (17) that have developed resistance to
conventional therapies. Disruption of iron metabolism is a key
mechanism that triggers ferroptosis. When intracellular iron
metabolism is imbalanced, excess Fe2+ reacts with
endogenous hydrogen peroxide via the Fenton reaction, generating
substantial quantities of lipid reactive oxygen species (ROS)
(18). These ROS amplify oxidative
stress by activating metabolic enzymes and inducing peroxidation of
polyunsaturated fatty acid phospholipids in the cell membrane
(18,19). The resulting lipid peroxide
production ultimately initiates ferroptosis. Consequently, both
increased cellular Fe2+ uptake and restricted
Fe2+ efflux can lead to iron accumulation and subsequent
cell death (20).
Haptoglobin (HP) is the predominant APP in the body,
with its primary biological function being the high-affinity
binding of free hemoglobin to prevent toxic effects and protect the
body from heme iron loss (21,22).
However, its specific role in LUSC progression and ferroptosis
remains largely unknown. Therefore, the present study aimed to
investigate the expression and potential function of HP in LUSC,
with a particular focus on its involvement in ferroptosis.
Materials and methods
Reagents and antibodies
Phorbol 12-Myristate 13-Acetate (PMA; cat. no.
HY-18739), FerroOrange dye (cat. no. HY-D1913), BODIPY 581/591 C11
dye (cat. no. HY-D1301), tin protoporphyrin IX dichloride (SnPPIX,
cat. no. HY-101194) and deferoxamine (DFO; cat. no. HY-B1625) were
purchased from MedChemExpress. RPMI-1640 medium (cat. no.
11875093), Trypsin-EDTA (cat. no. 25200056), FBS (cat. no.
A5670701), penicillin-streptomycin (cat. no. 15070063),
7-aminoactinomycin D (7-AAD, cat. no. 00-6993), PE-anti-CD206 (cat.
no. 12-2069-42), APC-anti-CD163 (cat. no. 17-1639-42), Alexa Fluor
568-goat anti-rabbit IgG (cat. no. A-11011), CD163 polyclonal
antibody (cat. no. PA5-109340) and formic acid (cat. no. 036617.36)
were purchased from Thermo Fisher Scientific, Inc. Alexa Fluor
647-anti-heme oxygenase 1 (HO-1; cat. no. ab237268) was purchased
from Abcam. The cytokines IL-4 (cat. no. 200-04-5UG) and IL-13
(cat. no. 200-13-10UG) were purchased from PeproTech Inc. (Thermo
Fisher Scientific, Inc.). Hb (cat. no. CSB-NP002401h) was purchased
from Cusabio Technology LLC.
Bioinformatics analysis and
visualization
The Gene Expression Profiling Interactive Analysis
(GEPIA) database (gepia.cancer-pku.cn) was used to examine the
correlation between the expression levels of ALB, A2M, LTF, IL6,
HP, CP, APOA1 and SERPINA1 and the prognosis of patients
with LUSC based on data from The Cancer Genome Atlas (TCGA;
portal.gdc.cancer.gov). Additionally, pan-cancer analysis was
performed on the HP gene. Differentially expressed genes
(DEGs) between LUSC and normal lung tissues were analyzed using
TCGA and Genotype-Tissue Expression data, along with a comparison
of HP expression between tumor and normal tissues. A
GeneCards database (genecards.org) search for ‘acute phase
proteins’ yielded 107 APP genes with relevance scores ≥1. The R
package intersect function (version 1.2.0;
CRAN.R-project.org/package=dplyr) was used to analyze the
intersection between LUSC DEGs and APP genes. Gene interactions
were examined using the STRING database (cn.string-db.org),
focusing on the top 481 genes interacting with HP. Key node
genes were identified using the cytoHubba plugin (version 0.1;
Institute of Information Science at Academia Sinica) with six
topological network algorithms [Maximal Clique Centrality (MCC),
Maximum Neighborhood Component (MNC), radiality, closeness, Edge
Percolated Component (EPC) and degree] in Cytoscape software
(version 3.9.0; Cytoscape Consortium). Pearson tests in the
LinkedOmics database (linkedomics.org) were performed on LUSC
samples from TCGA to statistically analyze HP-associated
genes, identifying 3,763 genes with a false discovery rate (FDR)
<0.001. Using the intersect function from the R package, the
intersection between HP-associated genes and
HP-interacting genes was assessed. The enrichGO and
enrichKEGG functions (version 4.18.4;
bioconductor.org/packages/release/bioc/html/clusterProfiler.html)
in R (version 4.2.1; R Core Team) were used to determine the Gene
Ontology (GO) terms and perform Kyoto Encyclopedia of Genes and
Genomes (KEGG) pathway enrichment analysis associated with the
HP gene in LUSC. Ferroptosis-related genes were obtained
from the KEGG database (genome.jp/kegg). For GO analysis,
significance was defined as q-value <0.001 for BP and q-value
<0.01 for CC and MF after Benjamini-Hochberg correction, while
for KEGG analysis, a q-value <0.05 was considered significant.
The LUSC datasets GSE191881 and GSE74706 from Gene Expression
Omnibus (GEO; ncbi.nlm.nih.gov/geo), along with gene expression
profiles of LUSC tissues and normal lung tissues from TCGA, and
relative HP protein expression levels from The Human Protein Atlas
database (version 25.0;
proteinatlas.org/ENSG00000257017-HP/cancer/lung+cancer#cptac_lung_sqcc),
were obtained for analysis. The LM22 gene signature matrix, which
contains characteristic genes of M2 macrophages, was obtained from
a previous study (23). The
infiltration levels of M2 macrophages in LUSC tissues were
evaluated using the CIBERSORT (https://github.com/Moonerss/CIBERSORT; accessed in
September 2025) algorithm.
Venn diagrams were drawn using the VennDetail
function (version 1.26.0; bioconductor.org/packages/VennDetail) in
R, along with the Venn function (version 1.12;
CRAN.R-project.org/package=venn) and ggvenn function (version
0.1.19; CRAN.R-project.org/package=ggvenn). GO and KEGG enrichment
circle plots were generated using OmicShare Tools (omicshare.com;
accessed in September 2025).
Cell lines and cell culture
Human normal lung epithelial cells BEAS-2B were
obtained from Jennio Biotech Co., Ltd. and were verified using STR
analysis. The lung squamous cell lines (NCI-H226, NCI-H520,
NCI-H1703 and NCI-H2170) and the human pro-monocyte cell line THP-1
were purchased from Procell Life Science & Technology Co., Ltd.
and were verified using STR analysis. BEAS-2B cells were cultured
in bronchial epithelial cell-specific medium (Jennio Biotech Co.),
which was already supplemented with dThe NCI-H226, NCI-H520,
NCI-H1703, NCI-H2170 and THP-1 cells were cultured in RPMI-1640
(Thermo Fisher Scientific, Inc.) medium supplemented with 1%
penicillin-streptomycin and 10% FBS (Thermo Fisher Scientific,
Inc.). All cells were maintained at 37°C and 5% CO2.
THP-1 cells were differentiated into M2 macrophages as described
previously (24). Briefly, THP-1
cells were treated with 150 nM Phorbol 12-Myristate 13-Acetate for
24 h, followed by culture in fresh RPMI-1640 for 24 h. The
macrophages were polarized into M2 macrophages by incubating them
with IL-4 (20 ng/ml) and IL-13 (20 ng/ml) for 24 h. The cells were
stained with APC-anti-CD163 and PE-anti-CD206 for flow cytometry
analysis using the CyFlow Cube 6 system (Sysmex Europe SE) to
confirm M2 macrophage polarization.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from BEAS-2B and LUSC cell
lines (NCI-H226, NCI-H520, NCI-H1703, and NCI-H2170) using
TRIzol® reagent (cat. no. 15596018CN; Thermo Fisher
Scientific, Inc.) and treated with DNase I to eliminate DNA
contamination. Subsequently, single-stranded cDNA was synthesized
from RNA using reverse transcriptase M-MLV (cat. no. 6110B; Takara
Bio, Inc.). The following primer sequences used for amplification
of HP and GAPDH were as follows: HP forward
(F), 5′-CAGCACAGTCCCCGAAAAGAA-3′ and reverse;
5′-CAGTCGCATACCAGGTGTCC-3′; and GAPDH F,
5′-GGAGCGAGATCCCTCCAAAAT-3′ and R, 5′-GGCTGTTGTCATACTTCTCATGG-3′;
primers were synthesized by Takara Bio, Inc. qPCR analysis was
performed using UltraSYBR Green Master Mix (cat. no. CW0957H;
Jiangsu Kangwei Century Biotechnology Co., Ltd.) to measure the
mRNA expression levels of HP. Cq values for HP were
normalized to GAPDH, and quantitative analysis was performed using
the 2−ΔΔCq method (25).
The PCR amplification conditions were based on a published report
(26). The thermocycling program
consisted of an initial denaturation at 95°C for 1 min (1 cycle),
followed by 40 cycles of denaturation at 95°C for 5 sec, annealing
at 60°C for 10 sec and extension at 72°C for 15 sec, with a final
extension at 72°C for 10 min (1 cycle). RT-qPCR analysis was
performed using rTaq (Takara Bio, Inc.) in a DNA thermal cycler
(Maxygen).
ELISA
HP levels in BEAS-2B and LUSC cell lines (NCI-H226,
NCI-H520, NCI-H1703, and NCI-H2170) were determined using an HP
assay kit (cat. no. E-EL-H6199; Wuhan Elabscience Biotechnology
Co., Ltd.) according to the manufacturer's protocol. Culture
supernatants from cells in the logarithmic growth phase were
collected and diluted 1:10,000. The diluted test samples, standard
solutions and blank diluent were added into microplate wells at 100
µl per well, and then incubated at 37°C for 90 min. The
intracellular fluid was then discarded, and 100 µl biotinylated
antibody working solution was added to each well, followed by
incubation at 37°C for 60 min. The plate was then washed three
times with detergent, and 100 µl HRP enzyme conjugate working
solution was added to each well. The plate was incubated at 37°C
for 30 min, followed by five washes. Then, 90 µl
3,3′,5,5′-Tetramethylbenzidine substrate solution was added to each
well and incubated at 37°C in the dark for 15 min before adding 50
µl stop solution to terminate the reaction. Finally, the optical
density (OD) value of each well was measured at a wavelength of 450
nm using an ELISA reader (ELX 808IU; Omega Bio-Tek, Inc.), and
protein concentrations were calculated using a standard curve.
Lentiviral infection
Recombinant lentiviral vectors overexpressing
HP (LV-HP; GV492-Ubc-HP-3FLAG) and a control vector
(LV-Ctrl; GV492-Ubc-MCS-3FLAG) were designed and synthesized by
Shanghai GeneChem Co., Ltd. A third-generation self-inactivating
lentiviral packaging system was used. HEK293T cells (Shanghai
GeneChem Co., Ltd.) were co-transfected with 20 µg of GV vector
plasmid, 15 µg of pHelper 1.0 and 10 µg of pHelper 2.0 (mass ratio
of 20:15:10). The transfection mixture was incubated at room
temperature for 15 min and added to the cells. After 6–8 h at 37°C,
the medium was replaced. Lentiviral particles were harvested 48 h
post-transfection, concentrated by ultracentrifugation at 53,000 ×
g for 2 h at 4°C, and resuspended in PBS.
To establish HP overexpression cells, NCI-H226 and
NCI-H520 cells were infected with LV-HP or LV-Ctrl
(multiplicity of infection, 50). The cells were plated at a density
of 5×104 cells/well in a 6-well plate and cultured until
reaching 30% confluence. Subsequently, the cells were infected with
200 µl lentivirus and 40 µl transfection enhancer HiTransG A for 16
h. Following this, the medium was replaced with fresh complete
culture medium for further cultivation. After 72 h post-infection,
the efficiency of HP overexpression was assessed in cells and
culture supernatant using RT-qPCR and ELISA, respectively.
Lactate dehydrogenase release (LDH)
assay
An LDH cytotoxicity assay kit (cat. no. HY-K1090;
MedChemExpress) was used to assess LDH levels in the supernatant of
LUSC cell lines (NCI-H226 and NCI-H520 cells) and M2 macrophages,
according to the manufacturer's protocol. LUSC cells infected with
LV-HP or LV-Ctrl at a concentration of 1×105
cells/ml were seeded in a 96-well plate at 100 µl per well and
incubated with or without Hb treatment for 24 h. Similarly, M2
macrophages at a concentration of 1×105 cells/ml were
plated in a 96-well plate at 100 µl per well and treated with or
without Hb in the presence of conditioned media (CM) from LUSC
cells infected with LV-HP or LV-Ctrl for 24 h. After washing
with PBS, 20 µl 0.4 mol/l lactic acid solution, 20 µl 4 mmol/l
2-p-iodophenyl-3-p-chloronitrobenzenetetrazole solution and 20 µl
reaction solution were added to the cells. The samples were
incubated at room temperature for 30 min, and the OD value was
measured using an ELISA reader with a detection wavelength of 492
nm and a reference wavelength of 650 nm. The fold change for each
sample (including the samples in the control group) was calculated
by dividing the value of each sample by the mean value of the
control group.
Flow cytometry analysis
Flow cytometry was used to quantify 7-AAD-positive
dead cells, intracellular Fe2+ and lipid peroxide
levels, CD163- and CD206-double-positive cells and HO-1-expressing
cells. To assess the proportion of 7-AAD-positive cells, and the
levels of intracellular Fe2+ and lipid peroxides in
NCI-H226 and NCI-H520 cells, cells infected with LV-HP or
LV-Ctrl were seeded at a concentration of 2×105 cells
per well in 6-well plates, with or without the addition of Hb, and
incubated at 37°C for 24 h. To determine the percentage of
7-AAD-positive cells, NCI-H226 and NCI-H520 cells treated as
aforementioned were washed with PBS. Subsequently, the cells were
resuspended in PBS containing 1 µg/ml 7-AAD (cat. no. 00-6993;
Thermo Fisher Scientific, Inc.) and incubated at room temperature
in the dark for 10 min. Following trypsin digestion, the cells were
washed with PBS, centrifuged at 4°C at 200 × g for 5 min to collect
the cells, and the percentage of 7-AAD-positive cells was analyzed
using flow cytometry. To determine the concentration of
Fe2+, NCI-H226 and NCI-H520 cells, treated as
aforementioned, were washed with PBS and then incubated at room
temperature in the dark in 1 µmol/l FerroOrange (cat. no. HY-D1913;
MedChemExpress) working solution for 30 min. Subsequently, the
fluorescence intensity of FerroOrange-stained cells was measured
using flow cytometry. The mean fluorescence intensity of each
sample was then calculated and compared with the control group. To
assess lipid peroxidation levels, NCI-H226 and NCI-H520 cells were
treated with LV-HP or LV-Ctrl, washed with PBS, and then incubated
in the dark at room temperature with fresh culture medium
containing 2 µm BODIPY581/591C11 dye (cat. no. HY-D1301;
MedChemExpress) for 30 min. Following trypsin digestion, cells were
washed with PBS, collected by centrifugation at 4°C, 200 × g for 5
min, and the fluorescence intensity of BODIPY581/591C11-stained
cells was measured using flow cytometry.
To identify the induction of M2 macrophages, THP-1
cells were treated with 150 nM Phorbol 12-Myristate 13-Acetate
(cat. no. HY-18739; MedChemExpress) for 24 h, cultured in fresh
RPMI-1640 for 24 h, and then stimulated with IL-4 (20 ng/ml; cat.
no. 200-04-5UG; Thermo Fisher Scientific, Inc.) and IL-13 (20
ng/ml; cat. no. 200-13-10UG; Thermo Fisher Scientific, Inc.) for an
additional 24 h, all at 37°C. Subsequently, the cells were washed
with PBS and then incubated with APC-anti-CD163 (1:500; cat. no.
17-1639-42; Thermo Fisher Scientific, Inc.) and PE-anti-CD206
(1:500; cat. no. 12-2069-42; Thermo Fisher Scientific, Inc.) at 4°C
in the dark for 30 min. Following a final wash with PBS and
centrifugation, flow cytometry was used to analyze CD163- and
CD206-double-positive cells. To assess the proportion of
7-AAD-positive cells and the levels of intracellular
Fe2+ and lipid peroxides in M2 macrophages, culture
supernatants from NCI-H226 and NCI-H520 cells infected with
LV-HP or LV-Ctrl were used as CM. M2 macrophages were
cultured with the CM in combination with other reagents for 24 h.
Subsequently, the M2 macrophages were stained with 7-AAD,
FerroOrange or BODIPY581/591C11 dyes, followed by flow cytometry
analysis. To assess the expression of HO-1 in M2 macrophages,
2×106 M2 macrophages were fixed in 0.25% cold
paraformaldehyde (cat. no. P0099; Beyotime Biotechnology) at 4°C
for 1 h, followed by lysis with 0.1% Triton X-100 (cat. no. HFH10;
Thermo Fisher Scientific, Inc.) for 10 min at room temperature in
the dark. The cells were incubated with an Alexa Fluor
647-anti-HO-1 antibody (cat. no. ab237268; Abcam) at room
temperature for 1 h prior to flow cytometry analysis. All flow
cytometry analyses were conducted on a CyFlow Cube 6 system. FlowJo
(version 10; BD Biosciences) was used to analyze the data,
employing gating strategies to exclude debris and select single
cells based on the forward- and side-scatter characteristics.
Transmission electron microscopy
M2 macrophages were inoculated into a 6-well plate
and co-cultured with CM and Hb for 24 h. The cells were then fixed
in 2.5% glutaraldehyde in PBS at 4°C for 4 h and subjected to three
15-min washes with PBS. Post-fixation was performed using 1%
OsO4 in PBS at 4°C for 1.5 h, followed by three
additional 15-min washes with PBS. Dehydration was performed using
sequential immersions in ethanol solutions (30, 50, 70 and 80%),
followed by acetone solutions (90 and 95%) for 15 min each.
Absolute acetone dehydration was then performed twice for 20 min
each. Subsequently, the cells were incubated in a 1:1 mixture of
absolute acetone and the final Spurr resin for 1 h at room
temperature, followed by a 1:3 mixture for 2 h, and finally in a
mixture of the final Spurr resin and absolute acetone overnight.
Following embedding, ultrathin sections, sectioned at 60 nm
thickness were prepared, stained and imaged using a transmission
electron microscope (cat. no. H-7650; Hitachi, Ltd.).
Determination of intracellular heme
content
A total of 5×103 M2 macrophages per well
was inoculated into a 96-well plate and co-cultured with CM and Hb,
either alone or in combination with a CD163 polyclonal antibody
(1:50-100; cat. no. PA5-109340; Thermo Fisher Scientific, Inc.),
for 24 h. Intracellular heme content was determined following
established protocols (27).
Briefly, cells were washed three times with PBS, centrifuged at 200
× g for 5 min and then resuspended in 180 µl concentrated formic
acid. The heme concentration of the formic acid solution was
determined spectrophotometrically at 400 nm.
Statistical analysis
Experimental data were analyzed using GraphPad Prism
software version 6.0 (Dotmatics). Survival curves were generated
using the Kaplan-Meier method, and differences between groups were
assessed by the log-rank test. Correlation analysis was performed
using the Pearson correlation coefficient. A one- or two-way ANOVA
with a Tukey's post hoc test was used to compare differences among
multiple groups. To determine the difference between two groups, an
unpaired t-test was used. All data are presented as the mean ± SD.
P<0.05 was considered to indicate a statistically significant
difference.
Results
HP is a hub APP gene in LUSC
To investigate the expression of APPs in LUSC and
its correlation with prognosis, a total of 107 APP genes were
initially identified with a relevance score of ≥1 from the
GeneCards database (Table SI).
From the GEPIA database, 5,944 DEGs were identified between LUSC
tumors and normal lung tissues (Table
SII). Intersection analysis revealed 46 overlapping APP and DEG
genes (Fig. 1A). STRING database
analysis demonstrated significant network interactions among 44 of
these 46 APP genes (Fig. 1B). Using
six topological network algorithms, including MCC, MNC, degree,
EPC, closeness and radiality from the Cytoscape plugin cytoHubba,
eight consistently top-ranked genes were identified: ALB, A2M,
LTF, IL6, HP, CP, APOA1 and SERPINA1 (Table SIII and Fig. 1C). Among the eight genes,
Kaplan-Meier survival analysis using GEPIA data showed that only
HP (P=0.049) and A2M (P=0.0043) were significantly
correlated with the prognosis of patients with LUSC, while the
remaining six genes did not demonstrate statistically significant
prognostic associations (Fig. 1D).
A previous study identified A2M as a potential therapeutic
target for NSCLC (13). However,
the specific role of the HP gene in predicting prognosis in
NSCLC remains unexplored. Through pan-cancer analysis using the
GEPIA database, it was found that HP was associated with the
prognosis of LUSC and five other cancer types: Stomach
adenocarcinoma (STAD), skin cutaneous melanoma (SKCM), brain lower
grade glioma (LGG), kidney renal papillary cell carcinoma (KIRP)
and kidney renal clear cell carcinoma (KIRC) (Fig. 1E). Analysis of the GEPIA database
revealed a significant differential expression in HP gene
expression in LUSC tissues compared with that of normal lung
tissues, with no significant differences observed in the other five
tumor types (Fig. 1F). These
findings collectively indicated that HP was a hub gene in
LUSC progression.
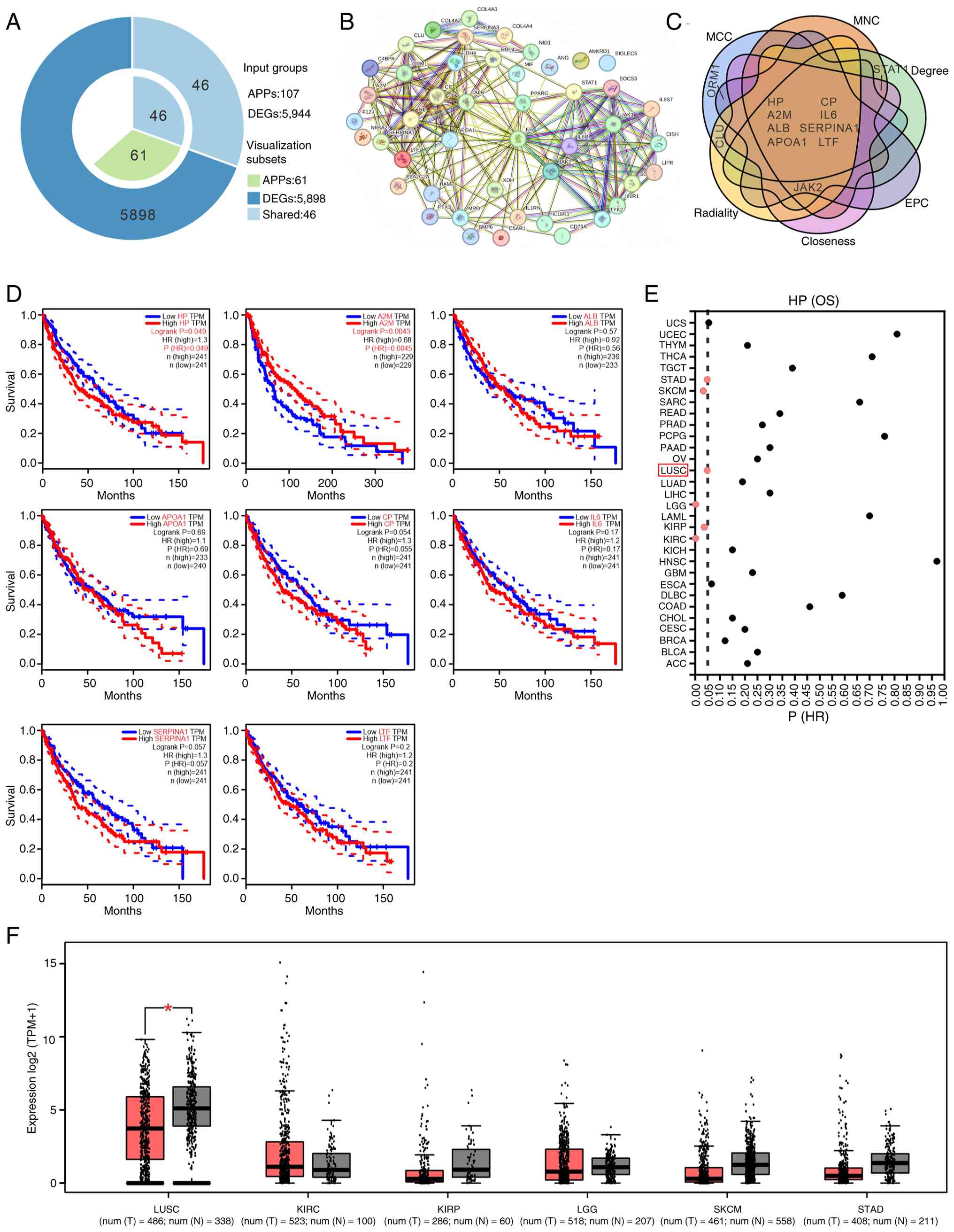 | Figure 1.Identification of HP as a hub
gene in LUSC. (A) A total of 107 acute-phase proteins with a
relevance score of ≥1 were selected from the GeneCards database and
intersected with 5,944 genes showing differential expression
between LUSC and normal lung tissues in the GEPIA database,
yielding 46 overlapping genes. (B) The 46 genes were then input
into the STRING database, where 44 genes formed a significant
interaction network. (C) The 44 genes within the network were then
assessed using six topological network algorithms (MCC, MNC,
radiality, closeness, EPC and degree) in the Cytoscape software
with the cytoHubba plugin. This analysis yielded the top 10 genes
ranked by each algorithm, with eight genes (HP, A2M, ALB, APOA1,
CP, IL6, SERPINA1 and LTF) consistently ranking highly.
(D) Survival analysis conducted on the GEPIA database revealed the
association between the expression levels of HP, A2M, ALB,
APOA1, CP, IL6, SERPINA1 and LTF and the overall
survival of patients with LUSC. (E) Data analysis from the GEPIA
database indicated a statistically significant association
(P<0.05) between HP expression in KIRC, KIRP, LGG, LUSC,
SKCM and STAD and patient OS rates. The red box is used
to highlight LUSC, the focus of this study. (F) Gene expression
data from the GEPIA database demonstrated a significant statistical
difference in HP expression between LUSC tissues and normal
tissues, while the changes in its expression in KIRC, KIRP, LGG,
SKCM and STAD did not reach statistical significance.
*P<0.05. HP, haptoglobin; LUSC, lung squamous cell
carcinoma; APP, acute-phase proteins; DEG, differentially expressed
genes; HR, hazard ratio; OS, overall survival; TPM, transcripts per
million. |
HP is significantly downregulated in
LUSC
To investigate HP expression in LUSC, gene
expression profiles were obtained from GEO and TCGA. Findings from
GEO (Fig. 2A) and TCGA (Fig. 2B) revealed a significant
downregulation of HP gene expression in LUSC tumor tissues
compared with that of normal lung tissues. Subsequent RT-qPCR
analysis of HP gene expression in the normal lung epithelial
cell line BEAS-2B and LUSC cell lines (NCI-H226, NCI-H520,
NCI-H1703, NCI-H2170) showed significant decreases in the LUSC cell
lines compared with that of BEAS-2B (Fig. 2C). Examination of the Human Protein
Atlas database showed a notable decrease in HP protein expression
in LUSC tumor tissues compared with that of normal lung tissues
(Fig. 2D). ELISA analysis also
confirmed a decrease in HP protein secretion levels in LUSC cell
culture supernatant compared with that from the normal lung
epithelial cell line BEAS-2B (Fig.
2E). These results indicated a significant downregulation of
the HP gene and protein in LUSC tumor tissues and cells.
HP enrichment in LUSC suggests
potential involvement in the ferroptosis signaling pathway
An examination of potential signaling pathways
influenced by HP in LUSC was conducted using TCGA data in
the LinkedOmics database. The analysis identified 3,763 genes
linked to HP with an FDR threshold of <0.001 (Table SIV). In the STRING database, by
setting the maximum number of genes in the interactome to 500 and
filtering for interactions with a score >0.4, 481 genes,
directly or indirectly, interacted with HP (Table SV). When comparing the 3,763
HP-associated genes with the 481 interacting genes, it was
found that 191 genes were both associated with and interacted with
HP (Fig. 3A). GO enrichment
analyses (Table SVI), visualized
using the OmicShare platform, revealed 20 enriched GO terms across
biological processes, cellular components and molecular functions
(Fig. 3B, upper panel). The
biological processes included lipid metabolism (‘neutral lipid
metabolic process’, ‘triglyceride metabolic process’ and
‘arachidonic acid secretion process’) and oxidation processes
(‘response to oxidative stress’ and ‘ROS metabolic process’).
Cellular component enrichment was observed in lipid components,
specifically the outer mitochondrial membrane. Molecular functions
included ‘oxygen binding’, enzyme activities (‘oxidoreductase’ and
‘peroxidase’), ‘lipoprotein lipase activator activity’ and
‘antioxidant activity’. The GO enrichment analysis indicated that
the 191 identified genes were predominantly involved in oxidative
stress responses associated with ferroptosis. This finding was
corroborated by the KEGG pathway enrichment analysis, which showed
a significant enrichment of these genes in the ferroptosis pathway
(Fig. 3B, lower panel). Analysis of
the KEGG database using the keyword ‘Ferroptosis’ identified 33
genes involved in the ferroptosis pathway (Table SVII). By comparing the 191 genes
associated with and interacted with HP and the 33
ferroptosis pathway genes, four intersecting genes were identified:
HMOX1, SLC40A1, CP and TF (Fig. 3C). Pearson correlation analysis
revealed significant associations between HP and these four
genes, with correlation coefficients of 0.23
(P=2.19×10−7), 0.31 (P=1.47×10−12), 0.48
(P=3.98×10−30) and 0.27 (P=6.87×10−10),
respectively. These results suggested that these four genes may
serve essential roles as regulators in the HP-induced
ferroptosis pathway.
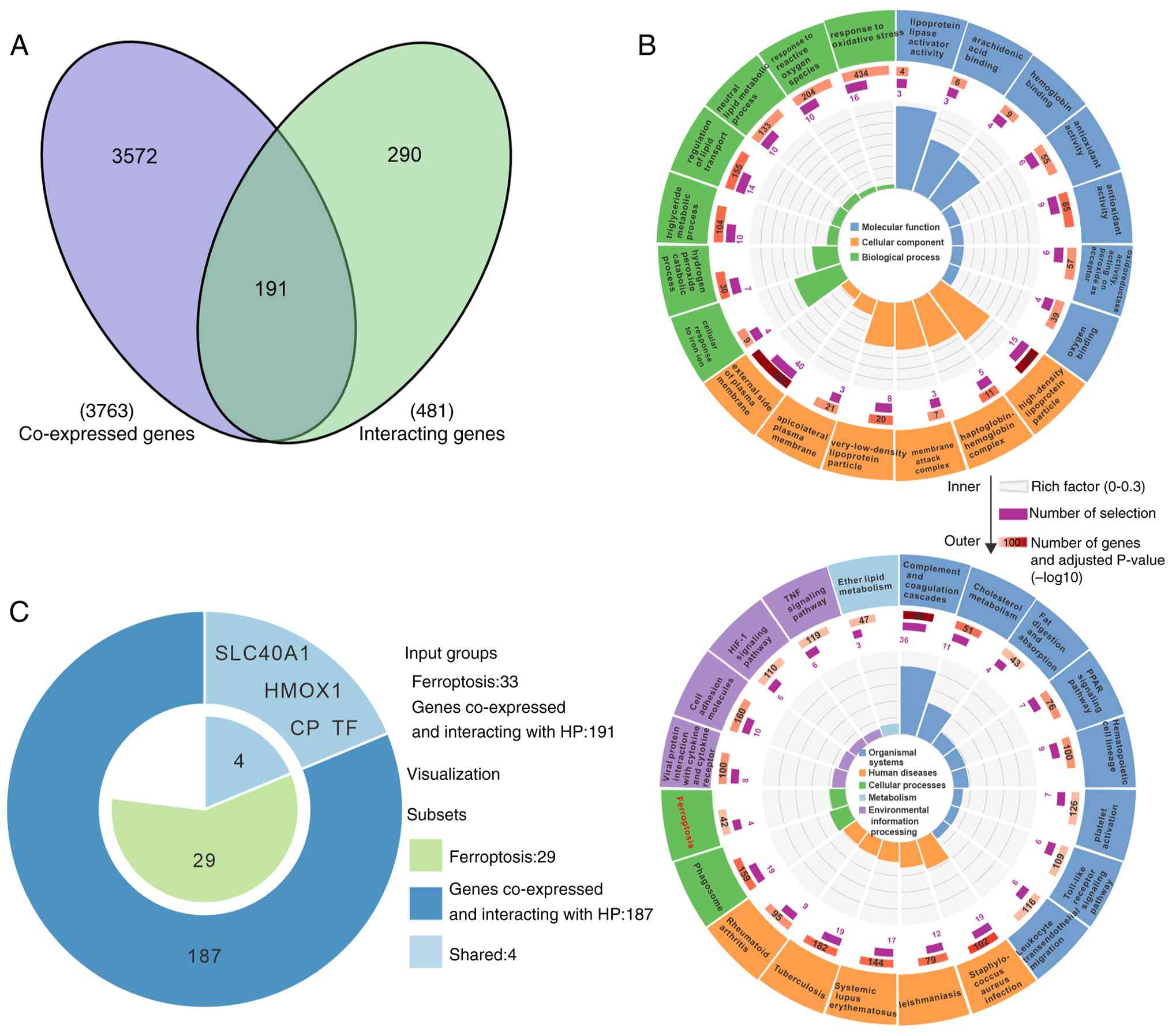 | Figure 3.HP enrichment in LUSC suggests
potential involvement in the ferroptosis signaling pathway. (A) In
TCGA samples from the LinkedOmics database, 3,763
HP-associated genes were identified (FDR threshold
<0.001), along with 481 genes interacting with HP sourced
from the STRING database. An intersection analysis of the 3,763
HP-associated genes and the 481 HP-interacting genes
yielded 191 overlapping genes. (B) The 191 genes associated with
HP interactions were subjected to GO enrichment analysis and
KEGG pathway enrichment analysis using R, followed by the
generation of enrichment circle plots using OmicShare tools. The GO
enrichment circle plot displayed seven biological processes, six
cellular components and seven molecular function entries (upper);
the KEGG enrichment circle plot showed a total of 20 entries
(bottom). (C) The keyword ‘Ferroptosis’ was used to search the KEGG
database, resulting in the retrieval of 33 genes relevant to
ferroptosis. An intersection analysis of the 33 ferroptosis-related
genes with the 191 genes from (A) identified four intersecting
genes: HMOX1, SLC40A1, CP and TF. HP,
haptoglobin; LUSC, lung squamous cell carcinoma; GO, Gene Ontology;
KEGG, Kyoto Encyclopedia of Genes and Genomes; FDR, false-discovery
rate. |
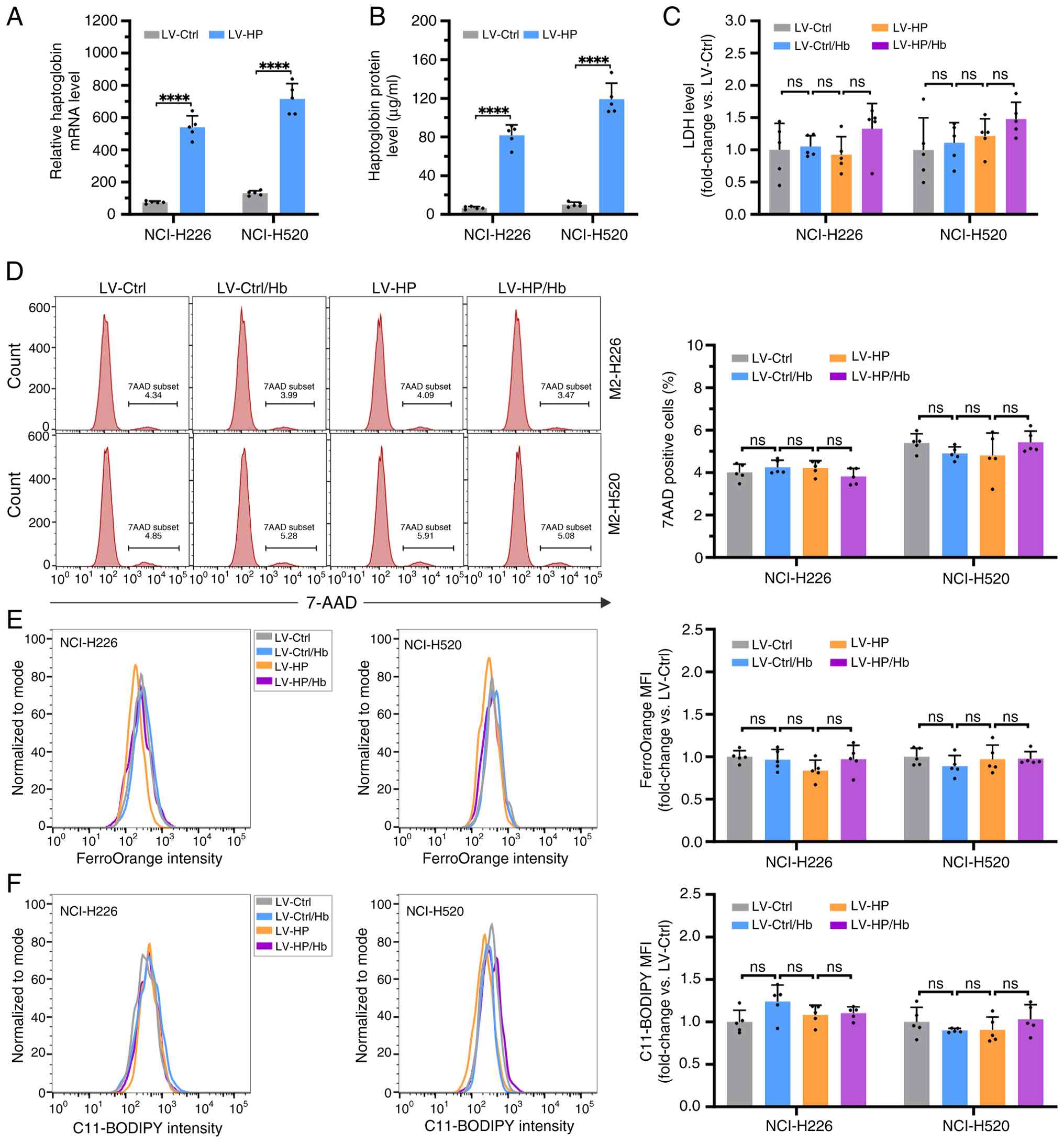
Overexpression of HP in LUSC cells
does not induce ferroptosis
Given the low expression of HP in LUSC cells,
lentiviral vectors were constructed to overexpress HP in the
NCI-H226 and NCI-H520 LUSC cell lines. After 72 h, cells and
culture supernatants were collected to quantify HP mRNA and
protein expression using RT-qPCR and ELISA, respectively. Compared
with the LV-Ctrl group, LV-HP infection significantly
increased HP mRNA (Fig. 4A)
and protein (Fig. 4B) expression
levels, confirming the successful generation of lentiviral vectors
expressing HP for subsequent experiments. Ferroptosis is a
form of cell death triggered by iron-dependent lipid peroxidation,
which increases membrane permeability (28). To study the influence of HP on
membrane permeability in LUSC NCI-H226 and NCI-H520 cells, LDH
release and 7-AAD assays were performed in these cells treated with
or without LV-HP in the presence or absence of Hb. It was
observed that there were no significant differences in LDH release
(Fig. 4C) and 7-AAD positive cells
(Fig. 4D) among the LV-Ctrl,
LV-HP, LV-Ctrl/Hb, and LV-HP/Hb groups. Flow
cytometry analysis showed no significant differences in
intracellular Fe2+ (Fig.
4E) and lipid peroxides (Fig.
4F) among the cells in the LV-Ctrl, LV-HP, LV-Ctrl/Hb
and LV-HP/Hb groups. Thus, it was demonstrated that the
overexpression of HP did not induce ferroptosis in the LUSC
cells.
CM from LUSC cells overexpressing HP
leads to ferroptosis in M2 macrophages
Previous studies have elucidated the roles of HP,
HMOX1, SLC40A1, CP and TF in regulating iron recycling
in M2 macrophages, with CD163 serving as the receptor (29,30).
To assess the correlation between HP expression and M2
macrophage abundance, LUSC samples from the TCGA database were
stratified into high- and low-HP expression groups based on
median HP expression levels (cut-off value, 50th
percentile). An analysis of immune infiltration in the LUSC gene
expression profile was then conducted to assess M2 macrophage
levels in these groups. The high-HP expression group
exhibited a significantly higher abundance of M2 macrophages
compared with that of the low-HP expression group (Fig. S1), indicating a positive
correlation between HP expression and M2 macrophage
abundance. Furthermore, LM22 data containing characteristic genes
of M2 macrophages from a previous study was obtained (23) and analyzed the correlation between
HP and characteristic genes of M2 macrophages. The results
showed significant associations between HP and marker genes
of M2 macrophages, such as ACP5, AIF1 and C3AR1, with
a Pearson correlation of 0.30 (P=8.16×10−12), 0.28
(P=8.15×10−11), and 0.21 (P=1.37×10−6),
respectively. These results demonstrated a positive correlation
between HP gene expression and the abundance of M2
macrophages.
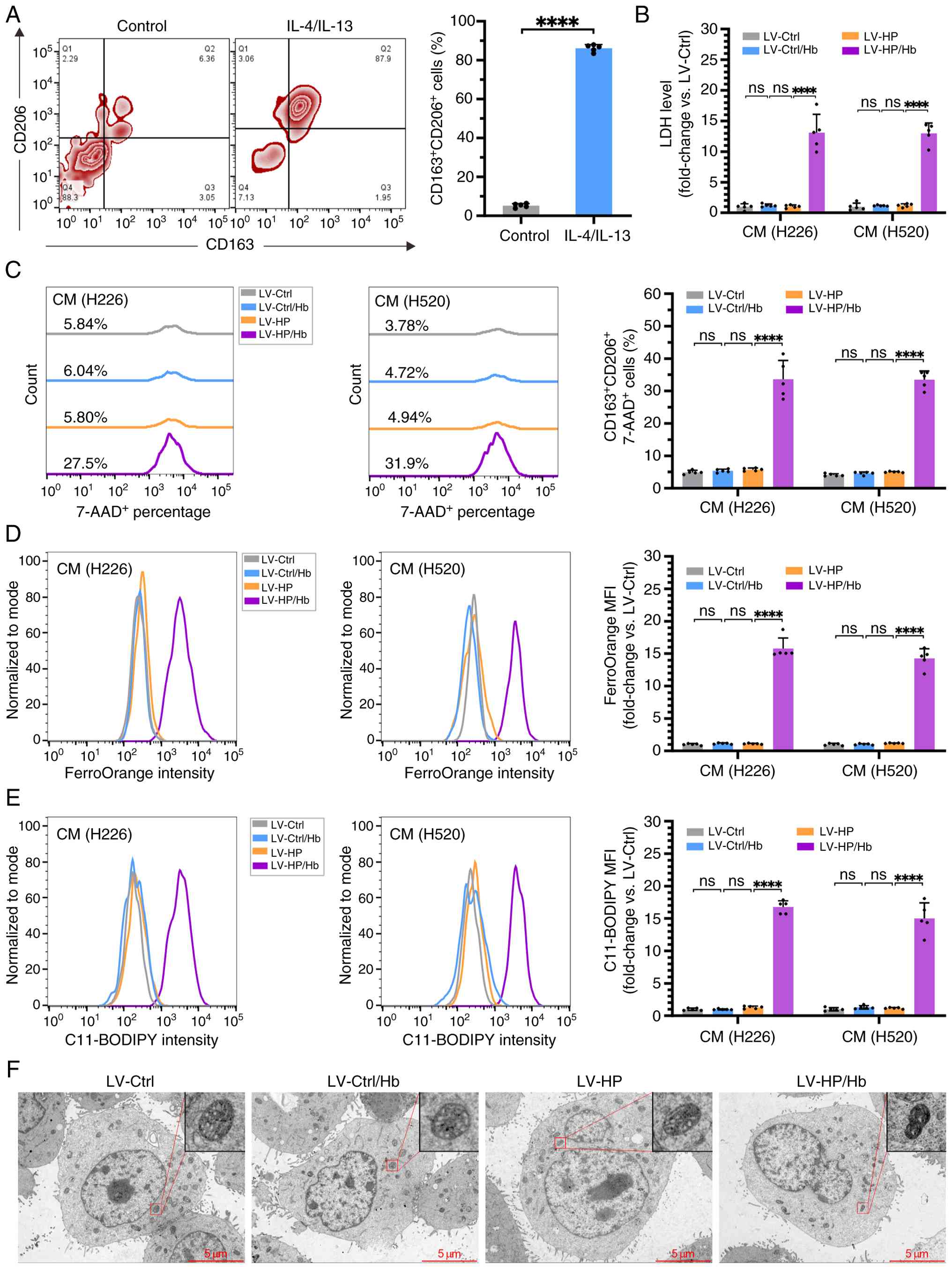
It was next investigated whether the overexpression
of HP in LUSC cells induced ferroptosis in M2 macrophages. THP-1
cells were differentiated into M2 macrophages, as confirmed by flow
cytometry demonstrating a CD206+CD163+
phenotype in ~80% of cells (Fig.
5A). Subsequently, NCI-H226 and NCI-H520 cells were transfected
with lentivirus LV-HP or LV-Ctrl, and the CM was collected.
M2 macrophages were exposed to the CM with or without Hb added for
24 h to assess induction of ferroptosis, marked by enhanced
membrane permeability attributed to iron-dependent lipid
peroxidation. The LDH release assay revealed that the LDH levels
were not significantly altered in the supernatant of M2 macrophages
exposed to CM from LV-HP-transfected tumor cells,
LV-Ctrl-transfected cells and LV-Ctrl-transfected cells with Hb
(LV-Ctrl/Hb) (Fig. 5B). Conversely,
M2 macrophages cultured in CM from LV-HP with Hb
(LV-HP/Hb) exhibited a significant increase in LDH release
levels compared with that of the other three groups. Flow cytometry
analysis of 7-AAD-positive cells indicated no statistically
significant variances among the LV-Ctrl, LV-Ctrl/Hb and
LV-HP groups, but a significantly higher percentage of
7-AAD-positive cells in the LV-HP/Hb group (Fig. 5C). Intracellular Fe2+ and
lipid peroxide levels in M2 macrophages, measured using FerroOrange
(Fig. 5D) and C11-BODIPY
respectively (Fig. 5E), showed no
significant differences among the LV-Ctrl, LV-Ctrl/Hb and
LV-HP groups. However, the LV-HP/Hb group
demonstrated a significantly higher fluorescence intensity for both
markers, indicating increased free Fe2+ (Fig. 5D) and lipid peroxide (Fig. 5E) levels. Transmission electron
microscopy showed ferroptotic characteristics in M2 macrophages
from the LV-HP/Hb group, such as decreased mitochondrial
size, increased mitochondrial membrane density and a reduction in
mitochondrial cristae, with no similar morphological alterations
observed in the other three groups (Fig. 5F). Taken together, these findings
suggest the CM from lung squamous carcinoma cells overexpressing HP
effectively triggered ferroptosis in M2 macrophages in the presence
of Hb.
HP promotes ferroptosis in M2
macrophages by inducing intracellular Fe2+ generation
via a CD163/HO-1 signaling pathway
To investigate the role of HP in intracellular
Fe2+ generation and ferroptosis induction in M2
macrophages, NCI-H226 and NCI-H520 cells were transfected with
LV-HP or LV-Ctrl. The resulting supernatants were used as CM. M2
macrophages were exposed to different combinations of CM, Hb and an
anti-CD163 antibody. After 24 h, heme and HO-1 protein levels were
measured, along with markers of ferroptosis. Spectrophotometry
analysis showed that in the absence of HP, there was no significant
difference in heme levels between the LV-Ctrl/Hb and
LV-Ctrl/Hb/anti-CD163 groups. However, in the presence of HP, heme
levels in the LV-HP/Hb/anti-CD163 group were significantly
decreased when compared with that of the LV-HP/Hb group
(Fig. 6A). These findings suggested
that blocking CD163 receptors with an anti-CD163 antibody impeded
Hb entry into M2 macrophages, thereby reducing intracellular heme
production.
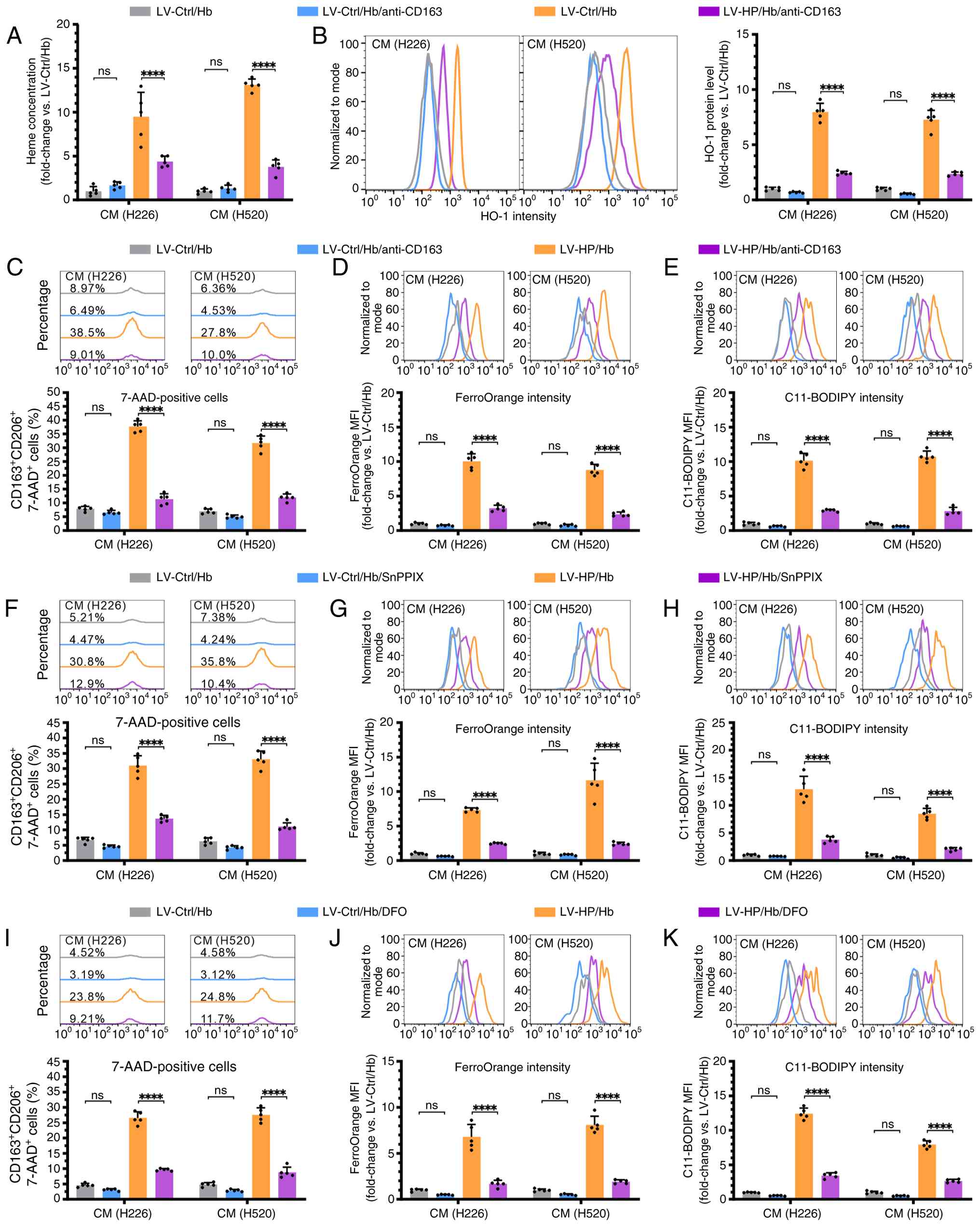 | Figure 6.HP promotes ferroptosis in M2
macrophage by inducing intracellular Fe2+ generation via
the CD163/HO-1 signaling pathway. (A and B) M2 macrophages were
cultured with CM from NCI-H226 and NCI-H520 cells infected with
LV-HP or LV-Ctrl, in the presence of Hb and an anti-CD163
antibody. The heme levels in M2 macrophages were measured using (A)
spectrophotometry and (B) HO-1 protein expression levels were
examined using flow cytometry. (C) M2 macrophages were co-cultured
with CM containing Hb, with or without the addition of an
anti-CD163 antibody, the HO-1 inhibitor SnPPIX, or the
Fe2+ chelator DFO for 24 h. (D) Following treatment with
the anti-CD163 antibody, the percentage of 7-AAD+ cells,
and the mean fluorescence intensity of FerroOrange and (E)
C11-BODIPY were analyzed by flow cytometry. After SnPPIX treatment,
(F) 7-AAD+ cells, (G) FerroOrange intensity (H) and
C11-BODIPY intensity were assessed. Following DFO treatment, (I)
7-AAD+ cells, (J) FerroOrange intensity, and (K)
C11-BODIPY intensity were measured. Data are presented as the mean
± SD from five independent experiments and were analyzed using an
unpaired t-test or a two-way ANOVA followed by a Tukey's post-hoc
multiple comparison analysis. ****P<0.0001. HP, haptoglobin; CM,
conditioned media; Hb, hemoglobin; 7-AAD, 7-aminoactinomycin D;
HO-1, heme oxygenase-1; DFO, deferoxamine. |
The results of flow cytometry analysis of
intracellular HO-1 protein levels were similar to those of heme
detection (27,31). There was no significant difference
in HO-1 levels between the LV-Ctrl/Hb and LV-Ctrl/Hb/anti-CD163
groups; however, the HO-1 levels in the LV-HP/Hb/anti-CD163
group were significantly lower compared with those in the
LV-HP/Hb group (Fig. 6B),
consistent with previous research findings (27,31).
To confirm the impact of CD163, HO-1, and Fe2+ on M2
macrophage ferroptosis, M2 macrophages were treated with an
anti-CD163 antibody, the HO-1 inhibitor SnPPIX or the
Fe2+ chelator DFO. Ferroptosis markers were then
assessed using flow cytometry, including 7-AAD-positive
membrane-permeable dead cells, Fe2+ and lipid peroxide.
In the absence of HP, the addition of the anti-CD163 antibody did
not affect 7-AAD-positive death cells, Fe2+ levels and
lipid peroxide in the LV-Ctrl/Hb and LV-Ctrl/Hb/anti-CD163 groups
(Fig. 6C-E). However, in the
presence of both HP and Hb, treatment with the anti-CD163 antibody
significantly reduced the levels of 7-AAD-positive dead cells,
Fe2+ and lipid peroxide compared to the groups without
antibody treatment (Fig. 6C-E).
Similar results were found in the corresponding M2 macrophages
treated with SnPPIX (Fig. 6F-H) and
DFO (Fig. 6I-K). Taken together,
these findings indicated that HP promoted ferroptosis in M2
macrophages by inducing intracellular Fe2+ generation
through the CD163/HO-1 signaling pathway.
Discussion
LUSC remains a leading cause of cancer-related
mortality, with the majority of patients being diagnosed with
advanced-stage disease, resulting in high mortality rates (32). Due to the lack of targeted therapies
for LUSC, treatment options for patients with advanced-stage
disease are extremely limited. Currently, immune checkpoint
inhibitors have emerged as a relatively successful therapeutic
strategy for patients with LUSC (5). However, the lower response rates and
severe adverse reactions have hindered the application of
immunotherapy in these patients (6). Therefore, there is an urgent need to
explore novel strategies for the management of LUSC. In the present
study, the HP gene was identified as a hub gene in LUSC via
bioinformatics analysis. It was demonstrated that HP expression
levels were lower in LUSC cells compared with that of normal lung
tissue cells, and were significantly associated with patient
prognosis. Further bioinformatics analysis suggested that HP
enrichment in LUSC indicated potential involvement in the
ferroptosis signaling pathway. Mechanistic investigations using
in vitro cell experiments revealed that overexpression of HP
in LUSC cells did not induce ferroptosis in the cells themselves,
but rather promoted intracellular Fe2+ accumulation in
M2 macrophages via the CD163/HO-1 signaling pathway, ultimately
inducing ferroptosis in M2 macrophages. These findings suggest a
potential mechanism by which LUSC downregulates HP expression to
inhibit M2 macrophage ferroptosis.
APPs are critical regulatory proteins that modulate
their expression in response to inflammatory processes (33). Emerging research indicates that
inflammatory mechanisms significantly contribute to tumor
development, with APPs serving a pivotal role in tumor cell
migration, invasion and anti-apoptotic inflammatory responses
(9), suggesting their potential as
tumor growth inhibitors. At present, the APP gene A2M has been
identified as a potential therapeutic target for NSCLC (13). However, the therapeutic implications
of other APPs in tumor management, particularly in LUSC, remain
unexplored. The present study initially identified a total of 107
APP genes from the GeneCards database. To focus on APP genes
affecting LUSC growth, the overlap between these 107 APP genes and
DEGs in LUSC were analyzed using protein-protein interaction and
cytoHubba algorithms. As a result, 8 APP genes were pinpointed as
hub genes for LUSC. Subsequent survival analysis of these 8 hub
genes revealed that only HP and A2M were significant for the
prognosis of patients with LUSC. Given that A2M has been reported
as a potential therapeutic target for NSCLC, HP was selected to
explore its role in inducing ferroptosis in LUSC. Further
bioinformatics analysis and in vitro cell experiments
demonstrated a decreased expression of HP in LUSC tumor tissues and
various cell lines (NCI-H226, NCI-H520, NCI-H1703 and NCI-H2170).
Enrichment analysis suggested that HP is potentially involved in
the ferroptosis signaling pathway in LUSC. Consequently, lentiviral
vectors were employed to overexpress HP in LUSC cells, followed by
a series of in vitro cellular experiments. The results
indicated that overexpression of HP in LUSC cells did not directly
trigger ferroptosis, as evidenced by the lack of significant
differences in LDH release, the frequency of 7-AAD-positive cells,
Fe2+ levels and lipid peroxidation across the LV-Ctrl,
LV-HP, LV-Ctrl/Hb and LV-HP/Hb groups. Instead, the CM from
HP-overexpressing LUSC cells in the presence of Hb (LV-HP/Hb group)
increased ferroptosis in M2 macrophages compared with that of the
three control groups. This was supported by an increase in LDH
release, the frequency of 7-AAD-positive cells, Fe2+
levels, lipid peroxidation and mitochondrial abnormalities observed
under transmission electron microscopy. These findings indicated
that overexpression of HP in in vitro cell models of LUSC
cells effectively induce M2 macrophage ferroptosis in the presence
of Hb.
Ferroptosis is a form of cell death characterized by
increased iron levels, which drive the generation of numerous lipid
peroxides, leading to increased cell membrane permeability and
ultimately cell death (28). Iron
metabolism serve a crucial regulatory role in the regulation of
ferroptosis. Previous studies have demonstrated that HP, HMOX1,
SLC40A1, CP and TF facilitate the retrieval of iron from
free Hb via M2 macrophages (29,30).
M2 macrophages serve a crucial regulatory role in hemoglobin-iron
recycling by internalizing the HP/Hb complex via CD163. HP binds to
Hb from damaged red blood cells and enters M2 macrophages via
CD163, where the Hb moiety is catabolized to heme within lysosomes
(29,34). HO-1, encoded by HMOX1,
converts heme to free iron (Fe2+) (29). Ferroportin, produced by
SLC40A1, transports most free iron extracellularly (29). Ceruloplasmin, encoded by CP,
oxidizes extracellular iron to Fe3+, which subsequently
binds to transferrin, encoded by TF, and is internalized by
iron-requiring cells (30). This
suggests that the ferroptosis mechanism involving HP may
occur in M2 macrophages. M2 macrophages are the predominant
tumor-associated macrophages in the tumor microenvironment, serving
a key role in promoting tumor growth through interactions with the
tumor immune microenvironment (35,36).
It was previously reported that patients with high levels of M2
macrophage infiltration in LUSC tend to have a poorer prognosis
(37). Therefore, targeting M2
macrophages represents a potential strategy for cancer management.
The present study demonstrated a significant amount of Hp-Hb
complexes promote the generation of Fe2+ within M2
macrophages via the CD163/HO-1 pathway, reflecting the iron uptake
process by M2 macrophages. While the iron metabolism of M2
macrophages involves both iron uptake and iron efflux, the specific
pathway of iron efflux was not elucidated in the present study. The
regulatory mechanisms of this process may exhibit specificity in
LUSC, thereby limiting the direct extrapolation of the functional
changes identified in the present study to other cancer types.
Therefore, future research needs to systematically map the complete
iron metabolism profile of M2 macrophages in LUSC, particularly
focusing on their iron efflux pathway.
The present in vitro experiments showed that
overexpression of HP in LUSC induced M2 macrophage
ferroptosis, indicating its potential in eliminating
immune-suppressive cells. However, prognosis and immune
infiltration analysis using patient data from the TCGA database for
LUSC revealed that patients with LUSC with high HP expression have
worse outcomes compared with those with low HP expression, with
significantly higher levels of M2 macrophage infiltration, a trend
consistent with previous reports associating elevated serum HP
levels with adverse prognosis in NSCLC (38). These findings suggested that the
role of HP in the complex tumor microenvironment in
vivo is more intricate than a singular mechanism observed in
vitro. It could be considered that in a complete tumor
ecosystem, the direct cytotoxic effect of HP on M2
macrophages may trigger a more intense compensatory immune
reshaping. The process of remodeling may involve the generation of
ROS from ferroptosis, potentially enhancing the sustained
recruitment of tumor-associated macrophages and their
differentiation into M2 macrophages under the influence of local
tumor microenvironment factors (39), thereby reinforcing the
immune-suppressive milieu and ultimately associating with malignant
progression and poor prognosis of tumors. HP may be a key
player in the tumor immune microenvironment, operating not through
a simple linear clearance mechanism but embedded within a dynamic
network of regulation. Future research should further validate the
existence of this ‘compensatory recruitment’ in in vivo
models and explore the feasibility of targeting HP as a
therapeutic approach. For instance, concurrent targeting of the
HP pathway and the key chemokine receptor CCR2 (40) for monocyte recruitment may
synergistically disrupt this detrimental immune homeostasis,
offering new avenues for immune combination therapy for LUSC.
However, there were still several limitations in
the present study. It was found through bioinformatics that low HP
expression in LUSC is negatively correlated with patient prognosis,
and four ferroptosis-related genes (HMOX1, SLC40A1, CP and
TF) were identified as being associated with and interacting
with HP. Among these, three genes (HMOX1, SLC40A1 and
TF) showed relatively weak correlations with HP
(r<0.3), warranting further validation of their roles in
HP-mediated ferroptosis. In vitro cell experiments also
showed that high expression of HP in LUSC cells promoted M2
macrophage ferroptosis via the hemoglobin-dependent CD163/HO-1
pathway, thereby indirectly validating that low expression of HP in
LUSC cells may inhibit M2 macrophage ferroptosis. However, this
mechanism has not been further validated in vivo, and the
relationship or mechanism between M2 macrophage ferroptosis and
LUSC prognosis has not been comprehensively studied. The potential
reason for this is that, in contrast to human HP, mouse HP does not
promote high-affinity binding of mouse Hb to CD163 (41), which limits the ability to validate
the function of the identified hemoglobin-dependent CD163/HO-1
pathway in vivo using mouse tumor models. Additionally, due
to limitations in available resources, clinical samples for in
vivo validation could not be conducted. Furthermore, the
simplification of in vitro experimental systems fails to
replicate the complex tumor microenvironment and its compensatory
feedback networks in vivo, making single intervention
strategies ineffective in vivo. Even with sufficient
clinical samples from patients with LUSC, verification of the
mechanism of downregulating haptoglobin expression to inhibit M2
macrophage ferroptosis via the hemoglobin-dependent CD163/HO-1
pathway remains challenging. Therefore, future studies should
utilize appropriate in vivo models and systematic combined
interventions to investigate whether LUSC regulates the tumor
immune microenvironment by downregulating HP expression to inhibit
M2 macrophage ferroptosis. In summary, the present study
demonstrated that the HP gene served a central role in LUSC.
HP was significantly downregulated in LUSC and was
associated with patient prognosis. Overexpression of HP in LUSC
cells led to exogenous Hb binding, triggering M2 macrophage
ferroptosis via the CD163/HO-1 pathway. These findings suggested
that LUSC may decrease HP expression to modulate ferroptosis
in M2 macrophages, which may provide a promising direction for
future research on LUSC.
Supplementary Material
Supporting Data
Supporting Data
Supporting Data
Supporting Data
Supporting Data
Supporting Data
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was funded by the National Natural Science
Foundation of China (grant no. 32460190) and the Hainan Provincial
Natural Science Foundation (grant no. ZDKJ202003).
Availability of data and materials
The datasets used and/or analyzed during the
present study are available from the corresponding author on
reasonable request.
Authors' contributions
FYH and GHT designed the study, and drafted and
revised the manuscript. WX, YYL and FYH performed the experiments
and analyzed the data. FYH and GHT oversaw the manuscript and gave
approval for the final submitted version. FYH and WX confirm the
authenticity of all the raw data. All authors have read and
approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Brody H: Lung cancer. Nature. 587 (Suppl
1):S72020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Herbst RS, Morgensztern D and Boshoff C:
The biology and management of non-small cell lung cancer. Nature.
553:446–454. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Siegel RL, Giaquinto AN and Jemal A:
Cancer statistics, 2024. CA Cancer J Clin. 74:12–49.
2024.PubMed/NCBI
|
|
4
|
Bonomi PD, Gandara D, Hirsch FR, Kerr KM,
Obasaju C, Paz-Ares L, Bellomo C, Bradley JD, Bunn PA Jr, Culligan
M, et al: Predictive biomarkers for response to EGFR-directed
monoclonal antibodies for advanced squamous cell lung cancer. Ann
Oncol. 29:1701–1709. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Herbst RS, Baas P, Kim DW, Felip E,
Pérez-Gracia JL, Han JY, Molina J, Kim JH, Arvis CD, Ahn MJ, et al:
Pembrolizumab versus docetaxel for previously treated,
PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010):
A randomised controlled trial. Lancet. 387:1540–1550. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Karachaliou N, Fernandez-Bruno M and
Rosell R: Strategies for first-line immunotherapy in squamous cell
lung cancer: Are combinations a game changer? Transl Lung Cancer
Res. 7 (Suppl 3):S198–S201. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Marusyk A, Janiszewska M and Polyak K:
Intratumor heterogeneity: The Rosetta stone of therapy resistance.
Cancer Cell. 37:471–484. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cray C, Zaias J and Altman NH: Acute phase
response in animals: A review. Comp Med. 59:517–526.
2009.PubMed/NCBI
|
|
9
|
Nickoloff BJ, Ben-Neriah Y and Pikarsky E:
Inflammation and cancer: Is the link as simple as we think? J
Invest Dermatol. 124:1275–1284. 2005. View Article : Google Scholar
|
|
10
|
Chen L, Wei WH, Sun J, Sun BC and Deng R:
Cordycepin enhances anti-tumor immunity in breast cancer by
enhanceing ALB expression. Heliyon. 10:e299032024. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang GR, Liu XY, Sun ZY, Feng XN, Wang
HY, Hao J and Zhang XL: A2M is a potential core gene in
intrahepatic cholangiocarcinoma. BMC Cancer. 22:52022. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zheng TZ, Zheng ZW, Zhou HX, Guo YQ and Li
SK: The multifaceted roles of COL4A4 in lung adenocarcinoma: An
integrated bioinformatics and experimental study. Comput Biol Med.
170:1078962024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhang NN: Screening and identification of
diagnostic and therapeutic targets of NSCLC and exploring the
expression and clinical significance of A2M in NSCLC. China
National Knowledge Infrastructure. Lanzhou University; 2019
|
|
14
|
Lei G, Zhuang L and Gan BY: Targeting
ferroptosis as a vulnerability in cancer. Nat Rev Cancer.
22:381–396. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Stockwell BR, Friedmann Angeli JP, Bayir
H, Bush AI, Conrad M, Dixon SJ, Fulda S, Gascón S, Hatzios SK,
Kagan VE, et al: Ferroptosis: A regulated cell death nexus linking
metabolism, redox biology, and disease. Cell. 171:273–285. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tang D, Kroemer G and Kang R: Ferroptosis
in hepatocellular carcinoma: From bench to bedside. Hepatology.
80:721–739. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lei Y, Jiang S, Kong C, Pang P and Shan H:
Ferroptosis: Therapeutic potential and strategies in Non-Small cell
lung cancer. Biology (Basel). 14:5452025.PubMed/NCBI
|
|
18
|
Chen X, Comish PB, Tang DL and Kang R:
Characteristics and biomarkers of ferroptosis. Front Cell Dev Biol.
9:6371622021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Koppula P, Zhuang L and Gan BY: Cytochrome
P450 reductase (POR) as a ferroptosis fuel. Protein cell.
12:675–679. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen X, Yu CH, Kang R and Tang DL: Iron
metabolism in ferroptosis. Front Cell Dev Biol. 8:5902262020.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
di Masi A, De Simone G, Ciaccio C, D'Orso
S, Coletta M and Ascenzi P: Haptoglobin: From hemoglobin scavenging
to human health. Mol Aspects Med. 73:1008512020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kristiansen M, Graversen JH, Jacobsen C,
Sonne O, Hoffman HJ, Law SK and Moestrup SK: Identification of the
haemoglobin scavenger receptor. Nature. 409:198–201. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Newman AM, Liu CL, Green MR, Gentles AJ,
Feng WG, Xu Y, Hoang CD, Diehn M and Alizadeh AA: Robust
enumeration of cell subsets from tissue expression profiles. Nat
Methods. 12:453–457. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kawase A, Takashima O, Tanaka S, Shimada H
and Iwaki M: Diclofenac-induced cytotoxicity in direct and indirect
co-culture of HepG2 cells with differentiated THP-1 cells. Int J
Mol Sci. 23:86602022. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Huang FY, Dai SZ, Xu WT, Xiong W, Sun Y,
Huang YH, Wang JY, Lin YY, Chen H, Tan GH and Zheng WP:
3′-epi-12β-hydroxyfroside-mediated autophagy degradation of
RIPK1/RIPK3 necrosomes leads to anergy of immunogenic cell death in
triple-negative breast cancer cells. Pharmacol Res. 187:1066132023.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Delaby C, Pilard N, Puy H and
Canonne-Hergaux F: Sequential regulation of ferroportin expression
after erythrophagocytosis in murine macrophages: Early mRNA
induction by haem, followed by iron-dependent protein expression.
Biochem J. 411:123–131. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Li SX and Huang Y: Ferroptosis: An
iron-dependent cell death form linking metabolism, diseases, immune
cell and targeted therapy. Clin Transl Oncol. 24:1–12. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Cairo G, Recalcati S, Mantovani A and
Locati M: Iron trafficking and metabolism in macrophages:
Contribution to the polarized phenotype. Trends Immunol.
32:241–247. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Fonseca Ó, Ramos AS, Gomes LTS, Gomes MS
and Moreira AC: New perspectives on circulating ferritin: Its role
in health and disease. Molecules. 28:77072023. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Alam J, Stewart D, Touchard C, Boinapally
S, Choi AM and Cook JL: Nrf2, a Cap'n'Collar transcription factor,
regulates induction of the heme oxygenase-1 gene. J Biol Chem.
274:26071–26078. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Lau SCM, Pan YW, Velcheti V and Wong KK:
Squamous cell lung cancer: Current landscape and future therapeutic
options. Cancer Cell. 40:1279–1293. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kushner I and Mackiewicz A: Acute phase
proteins as disease markers. Dis Markers. 5:1–11. 1987.PubMed/NCBI
|
|
34
|
Thomsen JH, Etzerodt A, Svendsen P and
Moestrup SK: The haptoglobin-CD163-heme oxygenase-1 pathway for
hemoglobin scavenging. Oxid Med Cell Longev. 2013:5236522013.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Dai EY, Han L, Liu J, Xie YC, Kroemer G,
Klionsky DJ, Zeh HJ, Kang R, Wang J and Tang DL:
Autophagy-dependent ferroptosis drives tumor-associated macrophage
polarization via release and uptake of oncogenic KRAS protein.
Autophagy. 16:2069–2083. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Qian Y, Qiao S, Dai YF, Xu GQ, Dai BL, Lu
LS, Yu X, Luo QM and Zhang ZH: Molecular-Targeted immunotherapeutic
strategy for melanoma via Dual-targeting nanoparticles delivering
small interfering RNA to Tumor-associated macrophages. ACS Nano.
11:9536–9549. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Han YS and Li YX: Comprehensive
exploration of M2 macrophages and its related genes for predicting
clinical outcomes and drug sensitivity in lung squamous cell
carcinoma. J Oncol. 2022:11639242022. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Lu JJ, Wang YH, Yan MS, Feng PN, Yuan LJ,
Cai YS, Xia X, Liu M, Luo JM and Li LS: High serum haptoglobin
level is associated with tumor progression and predicts poor
prognosis in non-small cell lung cancer. Oncotarget. 7:41758–41766.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Liang XS, Weng JD, You ZY, Wang Y, Wen J,
Xia ZW, Huang SR, Luo P and Cheng Q: Oxidative stress in cancer:
From tumor and microenvironment remodeling to therapeutic
frontiers. Mol Cancer. 24:2192025. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Muscat S, Nichols AEC, Gira E and Loiselle
AE: CCR2 is expressed by tendon resident macrophage and T cells,
while CCR2 deficiency impairs tendon healing via blunted
involvement of tendon-resident and circulating
monocytes/macrophages. FASEB J. 36:e226072022. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Etzerodt A, Kjolby M, Nielsen MJ, Maniecki
M, Svendsen P and Moestrup SK: Plasma clearance of hemoglobin and
haptoglobin in mice and effect of CD163 gene targeting disruption.
Antioxid Redox Signal. 18:2254–2263. 2013. View Article : Google Scholar : PubMed/NCBI
|
















![Significant downregulation of
HP in LUSC. (A) The LUSC datasets GSE19188 and GSE74706 from
the Gene Expression Omnibus database were examined. These datasets
included 35 LUSC samples and 73 normal lung tissue samples. The
objective was to evaluate HP gene expression in normal lung
and LUSC tissues (unit: log2_count). (B) The TCGA database provided
the expression profiles of the HP gene in normal lung and
LUSC samples [unit: norm_log2(TPM+1)]. (C) Reverse
transcription-quantitative PCR analysis was performed to examine
HP mRNA expression in the normal lung epithelial cell line
BEAS-2B and LUSC cell lines (NCI-H226, NCI-H520, NCI-H1703 and
NCI-H2170). (D) Relative HP protein expression levels in LUSC
tissues and normal lung tissues were obtained from Human Protein
Atlas database. (E) ELISA was performed to assess HP secretion in
the normal lung epithelial cell line BEAS-2B and the LUSC cell
lines. Data are presented as the mean ± SD from five independent
experiments and were analyzed using an unpaired t-test or a two-way
ANOVA followed by a Tukey's post-hoc multiple comparison analysis.
**P<0.01, ***P<0.001, ****P<0.0001. HP, haptoglobin; LUSC,
lung squamous cell carcinoma; TPM, transcripts per million; ns, not
significant.](/article_images/ol/31/5/ol-31-05-15558-g01.jpg)