Introduction
Pancreatic neuroendocrine tumors (PNETs) are a rare
(<2%) and biologically diverse class of neoplasms originating
from the endocrine cells of the pancreas, distinguished by a
complex pathological architecture and diverse clinical behavior
(1–5). Despite a rare incidence (2.45 per
100,000) compared with pancreatic ductal adenocarcinoma, the 5-year
survival rate for PNET is up to 40% (6,7).
Particularly, PNETs exhibit a high propensity for liver metastasis
and postoperative recurrence, with the liver being the predominant
site of disease relapse (8–11). Based on hormone secretion, PNETs are
broadly classified into functional and non-functional subtypes
clinically. Functional PNETs (F-PNETs), such as insulinomas and
gastrinomas, are typically diagnosed earlier due to hormone-related
syndromes. By contrast, non-F-PNETs (NF-PNETs), which comprise the
majority of cases, are often asymptomatic until late stages and
indicate an extensive range of malignant potential. A recent study
reported the 3.7-fold increasing incidence rate of NF-PNETs within
the last 30 years (6,12,13).
The World Health Organization (WHO) further stratifies PNETs into
grades G1, G2 and G3 based on mitotic count and Ki-67 proliferation
index (14,15). While G1 and G2 tumors are generally
well-differentiated and may follow a more indolent course, G3
tumors encompass both well-differentiated high-grade PNETs and
poorly differentiated pancreatic neuroendocrine carcinomas, each
with distinct molecular and clinical behavior. Such evolutionary
progression is generally indicative of poor clinical outcomes
(16–19). Defining the evolutionary dynamics
and progression characteristics of PNETs holds potential in
enhancing prognostic assessment. Despite these established clinical
frameworks, the management of PNETs faces challenges. A notable
proportion of patients with well-differentiated tumors still
experience relapse, underscoring the limitations of
histopathological grading alone in predicting clinical trajectory.
Furthermore, therapeutic options for advanced disease, including
chemotherapy and targeted therapy (11,20,21),
are limited by innate and acquired resistance mechanisms. This
markedly highlights the unmet need for a more nuanced, molecularly
driven understanding of PNET pathogenesis.
While the WHO grading system classifies tumors
according to proliferative indices, it overlooks the functional
plasticity and microenvironmental reprogramming that often underlie
aggressive phenotypes in histologically similar tumors. Increasing
evidence suggested that the tumor microenvironment (TME), a dynamic
assembly of stromal cells, immune infiltrates, vasculature and
extracellular matrix (ECM) serves a key role in regulating tumor
progression, metastasis, therapeutic responsiveness and immune
evasion (22–24). In several instances, TME features
have exhibited stronger prognostic and predictive value compared
with tumor-intrinsic factors alone, emphasizing the need to
understand this intricate ecosystem (25–27). A
key unmet need is the development of integrative approaches that
place the TME at the forefront, alongside tumor-intrinsic factors,
to improve prognostic accuracy and optimize therapeutic strategies
in PNETs. Conventional siloed approaches, focusing exclusively on
genomics or the immune compartment, have offered fragmented
insights. Recent advances in multi-omics technologies including
genomic sequencing, transcriptomic sequencing [bulk RNA sequencing
(bulk RNA-seq), single-cell RNA sequencing (scRNA-seq) and spatial
transcriptomics] and proteogenomic sequencing have enabled
comprehensive profiling of both tumor-intrinsic and extrinsic
features. In initial studies, genomic analyses revealed that
multiple endocrine neoplasia type 1 (MEN1), death-associated
protein 6 (DAXX) and α-thalassemia/mental retardation syndrome
X-linked (ATRX) are among the most frequently mutated genes in
PNETs, often associated with poor clinical outcomes and disrupted
chromatin regulation (5,28–30).
Beyond these earlier genomic profiling studies, the newer
multi-omics platforms not only reconstruct the cellular
architecture of PNETs but also reveal functional programs such as
lineage plasticity, immune remodeling, metabolic reprogramming and
tumor-stroma crosstalk that lead to disease progression. In the
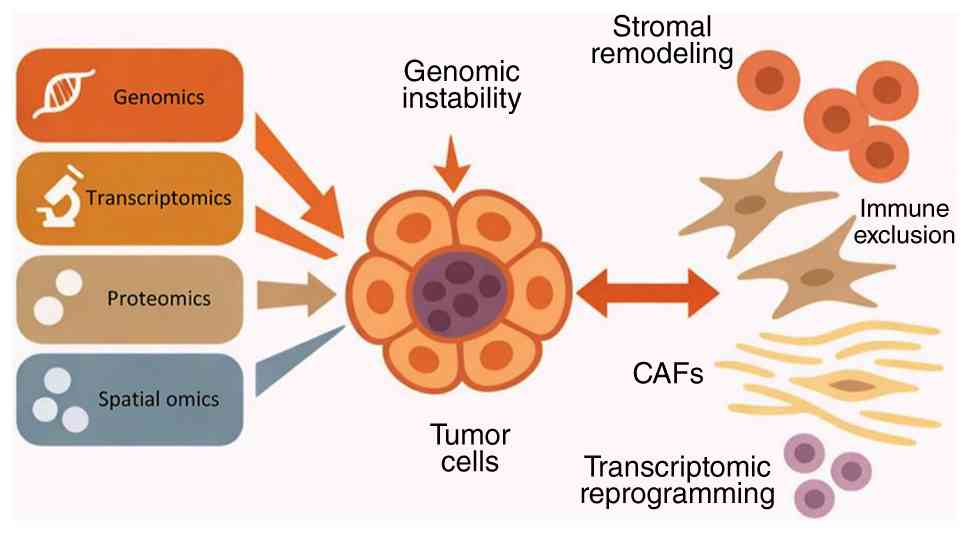
present review, findings from novel multi-omics studies were
integrated to reframe current understanding of PNET biology through
the lens of TME co-evolution (Fig.
1). The present review reveals core mechanisms such as genomic
instability, stromal remodeling, immune modulation and biomarker
identification, with a focus on the translational relevance for
patient stratification and targeted therapeutic development.
Genomic landscape
Genomic instability as a driver of
dedifferentiation and transcriptomic plasticity
Notably, genomic and microenvironmental features
vary markedly across the WHO tumor grades (G1, G2 and G3) as well
as between functional and NF-PNETs, and findings from different
studies should therefore be interpreted within the appropriate
clinical and pathological context. In PNETs, genomic instability
primarily refers to chromosomal instability characterized by copy
number variations (CNVs), large-scale chromosomal gains and losses
and telomere dysfunction associated with ATRX/DAXX alterations,
rather than microsatellite instability or a high mutational burden
(a threshold of >10 mutations/megabase), which are relatively
uncommon in these tumors (5,28,29,31).
Within this framework, genomic instability reflects progressive
structural alterations of the genome that reshape transcriptional
programs and cellular identity.
The transition from well-differentiated,
slow-growing tumors (grade G1-G2) to poorly differentiated,
high-grade PNETs is closely associated with increasing genomic
instability and transcriptomic reprogramming. Cutting-edge
technologies enabled the mapping of this transformation across
multiple molecular layers (32,33).
Low-grade tumors often present low CNV and maintain endocrine
identity through the stable expression of lineage-specific
transcription factors and chromatin regulators. However, as tumors
progress to higher grades, extensive CNVs, chromosomal instability
and altered DNA repair mechanisms emerge. These changes are
accompanied by global shifts in gene expression, including
downregulation of neuroendocrine markers and upregulation of genes
associated with cell cycle progression, epithelial-mesenchymal
transition (EMT), hypoxia and inflammation (34).
Single-cell and bulk transcriptomic analyses
stratified by WHO grade (G1-3) revealed an association between
tumor grade and increased proliferative signaling, loss of
endocrine differentiation and activation of EMT and Myc-driven
transcriptional programs in PNETs (35–37).
CNV inference (inferCNV) from scRNA-seq in NF-PNETs demonstrated
that G3 tumors harbor subclonal populations marked by chromosomal
instability and dedifferentiation (38). These subclones preferentially
localize to the invasive tumor margins and express markers of
metastasis and cell cycle progression. Furthermore, in CNV-high
tumors, extensive chromosomal aberrations, elevated tumor
mutational burden (TMB) and activation of DNA repair and oxidative
stress pathways are observed (34).
These molecular features are often accompanied by
microenvironmental alterations, including expansion of
myofibroblastic cancer-associated fibroblasts (CAFs), ECM
deposition and immune exclusion (39,40).
In addition to gene-level mutations, large-scale
chromosomal alterations are a hallmark of PNET progression. CNV
profiling using whole-genome sequencing revealed three major
structural subtypes: CNV-low, -altered and -recurrent (38). CNV-low tumors typically retain
endocrine differentiation and display stable karyotypes. By
contrast, CNV-altered and -recurrent tumors exhibit broad
chromosomal gains and losses. For example, recurrent CNV profiling
by proteomic analysis identified frequent arm-level and focal gains
and losses across the NF-PNET genome, even in tumors classified as
low or intermediate grade (41).
Common gains are observed on chromosomes 4p and q, 7p and q, 13q,
14q, 19p and q, 20p and q, while losses frequently affect 11p and
q. Notably, amplification of CDK6 (chromosome 7) was functionally
associated with enhanced cell cycle progression and defines a
subset of highly proliferative tumors (42). Similarly, amplification of enhancer
of zeste homolog 2, a histone methyltransferase also located on
chromosome 7, may contribute to immune modulation by influencing
interferon signaling and epigenetic silencing (43). By contrast, loss of MEN1 on
chromosome 11, one of the most frequently deleted tumor suppressors
in PNETs, disrupts chromatin regulation and is considered a
fundamental early event in tumor initiation (28,30).
Similarly, ATRX/DAXX mutations, often associated with alternative
lengthening of telomeres, co-occur with chromosomal rearrangements
involving chromosome 21q or 9p (44–46).
The frequent alteration of MEN1, DAXX and ATRX genes underscores
the notable importance of chromatin remodeling in PNET
tumorigenesis. However, beyond the roles in initiation, these
mutations appear to set the stage for later genomic catastrophes
(5,31,47).
The loss of function in these chromatin regulators likely creates a
permissive state for the accumulation of CNVs and structural
variations, ultimately promoting the transition from a
well-differentiated to a poorly differentiated state (48). A notable case study described a
patient with a germline Fanconi anemia group D2 protein (FANCD2)
mutation whose PNET exhibited extensive chromosomal instability and
a dedifferentiated transcriptional profile (49). Whole exome sequencing of this tumor
revealed marked germline mutation of Wnt family member 10A (a Wnt
signaling activator) and deletion of MutY homolog (a connection
with base excision repair deficiency) (50–53).
Furthermore, due to FANCD2 germline mutation, impaired DNA damage
repair and unprotected genome resulted in high mutation rate in
PNETs (54). Although increasing
studies indicated high mutation frequencies (43%) (5,29), the
prognostic impact of specific mutational co-occurrences remains
less defined, suggesting context-dependent roles that warrant
further investigation in larger, prospectively annotated cohorts
(29,49).
Transcriptomic data including bulk RNA-seq,
scRNA-seq, spatial transcriptomics and proteogenomic analysis
further illuminate this process. scRNA-seq of primary tumors
revealed enrichment of gene programs associated with angiogenesis
and stemness (35). Particularly,
transcription factor 4 (TCF4), which was reported to involve in
neural development and neuroendocrine differentiation (55,56),
enhanced activity in primary tumor sites. However, the markedly
enriched cellular functions in metastatic tumor were associated
with biosynthesis, mitosis, cell cycle and cell proliferation.
Proteogenomic profiling revealed that high-grade tumors
downregulate these markers and upregulate genes associated with
hypoxia responses, EMT and an inflammatory signature, which
associated with aggressive phenotype, such as invasion and
metastasis (38,57). Spatial transcriptomics implied that
these proliferative, dedifferentiated states are enriched at
invasive and hypoxic regions with dense CAF infiltration and
limited immune surveillance (44).
In G3 PNETs, tumor cells and CAFs co-express EMT and
invasion-associated genes (for example, MMPs), while CAFs secrete
paracrine inducers of dedifferentiation, notably TGF-β1 (41,44,58,59).
This reciprocal signaling loop may contribute to malignant
transformation and illustrates the coupled evolution of tumor and
stroma. In summary, these data supported that genomic instability
appears to represent a key feature associated with tumor
progression that reshapes cellular identity and primes tumors for
invasion, metastasis and therapy resistance.
Clonal evolution and the framework of
metastatic niches
Longitudinal and cross-sectional integrated omics
studies using pseudotime trajectory inference and CNV profiling has
revealed dynamic clonal transitions during PNET progression. These
transitions are marked by acquisition of metabolic plasticity,
angiogenic signaling and transcriptional programs associated with
immune evasion and proliferative expansion (35,48).
These changes likely reflect selective pressures imposed by
evolving microenvironmental niches. The cross-sectional snapshot
provided by multi-omics data allows us to reconstruct these
evolutionary trajectories, revealing the sequence of molecular
events that culminate in metastatic competence.
Metastatic lesions, particularly in the liver, are
often populated by tumor clones enriched in oxidative
phosphorylation, Myc signaling and resistance to apoptosis depend
on increased early region 2 binding factor family and Bax
expression via scRNA-seq (35).
These environments are also characterized by pro-angiogenic
endothelial cells and M1-like macrophages, forming niches conducive
to metastatic colonization. This metabolic phenotype may not only
support the high energy demands of proliferation and invasion but
also confer resistance to certain therapeutics (for example,
targeted therapy). Spatial transcriptomics demonstrated that
dissemination is often preceded by the emergence of spatially
confined, proliferation-primed subclones at the tumor margin,
frequently associated with hypoxia-driven gene expression (for
example, MMP-9) (44). This finding
suggested that metastasis may be preceded by transcriptional
programs that facilitate dissemination orchestrated by the primary
tumor.
Proteogenomic profiling delineated that EMT-like
subpopulations, enriched in high-grade and metastatic tumors,
indicate TGF-β activation, glycolysis, Myc signaling and
hypoxia-responsive transcription (41,60,61).
Particularly, hypoxia-induced EMT signature exhibits a positive
association with TGF-β signaling pathway (57,62).
scRNA transcriptomics identified that these metastatic clusters
localize to invasive fronts with high CAF density and minimal
T-cell infiltration (38). These
findings suggested that metastatic competency may be preconfigured
within transcriptionally primed subpopulations at the primary tumor
margins. To define the metastatic potential in the high-grade
tumors, scRNA transcriptome analysis in PNETs revealed the role of
a gene signature with notable emergence of two genes proprotein
convertase subtilisin/Kexin type 1 (PCSK1) and secreted modular
calcium-binding protein 1 (SMOC1) (35,38).
The emergence of PCSK1 and SMOC1 as potential markers of metastatic
propensity is of notable translational interest. Validating these
proteins as circulating biomarkers in patient plasma could
potentially provide a non-invasive tool in assessing metastatic
risk and guiding surveillance strategies for patients with
localized disease in the future.
In summary, genomic instability in PNETs promotes
not only tumor-intrinsic transformation but also adaptation to
selective pressures imposed by the microenvironment. The
co-occurrence with dedifferentiation and stromal remodeling of
genomic instability in PNETs underscores its key role in malignant
progression (Fig. 2). These
findings collectively illustrate the need for therapeutic
strategies that account for intratumoral heterogeneity and spatial
dynamics to effectively target the evolving landscape of PNETs.
Tumor-stromal-immune microenvironment
Integrated omics deconstructing the
PNET microenvironment
While tumor-intrinsic alterations are key to
understanding PNET progression, a complete understanding requires
dissecting the dynamic interactions of PNETs with the surrounding
stroma and immune compartments. Traditional bulk sequencing
approaches averaged signals across all cellular constituents,
obscuring the unique contributions of rare but functionally key
cell states. scRNA-seq and spatial transcriptomics revolutionized
the ability to deconstruct the cellular heterogeneity of the PNET
microenvironment. These high-resolution technologies exposed a
complex ecosystem composed of malignant neuroendocrine cells, CAFs,
endothelial cells and immune subsets including T and B cells,
monocytic cell, tumor-associated macrophages and mast cells each
exhibiting context-dependent differentiation states and functional
programs (35,38). However, single-cell dissociation
serves the key spatial context that defines cellular function.
Spatial transcriptomic mapping demonstrated that these cellular
populations exhibit non-random, spatially organized distributions
(44,58). Proliferative tumor regions are
typically bordered by dense fibroblast-rich stroma, which serves
both as a physical barrier and a source of immunomodulatory cues.
Proteogenomic profiling revealed that hypoxic and fibrotic zones
often co-localize with reduced immune infiltration, highlighting
spatial regulation of immune exclusion and therapeutic resistance
(41). These technologies anchor
the molecular profiles provided by scRNA-seq to a precise
geographical location within the tumor architecture. These
integrated omics approaches revealed that the spatial
compartmentalization of the TME has notable clinical implications,
which can explain the frequent failure of therapies that target
ubiquitous molecules but fail to penetrate specific anatomical
niches. Therefore, mapping the spatial TME may potentially provide
a key foundation for the development of spatially informed
combination therapies in the future.
CAF-driven niches promote
dedifferentiation and immune exclusion
Recent transcriptomic studies have identified
distinct CAF subtypes in PNETs with specialized roles in shaping
functional tumor niches (35,38,44).
The functional diversification of CAFs appears to be closely
associated with tumor grade progression (63–65).
Spatial profiling of sorted α-smooth muscle actin+
stromal cells exhibited that CAFs evolve with tumor grade: G1/2
tumors exhibit ECM-remodeling CAFs (often termed myofibroblastic
CAFs), while G3 tumors harbor immunoregulatory and pro-fibrotic
CAFs (reminiscent of inflammatory CAFs) expressing TGF-β1,
fibronectin 1 and fibroblast growth factor 8 (44,58).
These changes contribute to immune exclusion, enhanced invasiveness
and hypoxia (66). Furthermore,
single-cell transcriptomic and spatial data reported that
dedifferentiated tumor cells localize to CAF-enriched fibrotic
zones, where EMT and cell cycle genes (for example, MMPs and Myc)
are upregulated (38,44,58).
These data illustrated how functional state transitions, such as
the loss of neuroendocrine identity and acquisition of mesenchymal
or progenitor-like traits, are not solely driven by intrinsic
oncogenic signaling but are spatially reinforced by CAF-derived
cues within the fibrotic tumor margins (58). This shift from a desmoplastic to an
immunosuppressive CAF phenotype represents a key transition in PNET
pathogenesis. Pseudotime trajectory analyses elucidated that
proliferative, dedifferentiated subclones emerge along a continuum
from endocrine to mesenchymal transcriptional states, particularly
at hypoxic and fibrotic margins (35). Single-cell and proteomic profiling
elucidated that tumor cells from patients with PNET within these
micro-niches frequently express exhaustion markers (for example
programmed cell death 1, lymphocyte-activation gene 3 and cytotoxic
T-lymphocyte antigen 4), further reinforcing immune evasion
(35,38,41,60).
Therefore, these observations underscored a model in
which tumor cell state plasticity, stromal remodeling and immune
exclusion co-evolve within spatially defined microenvironments.
Dissecting these coupled networks at single-cell and spatial
resolution is key to identifying actionable targets and designing
therapies that disrupt both malignant progression and its
supportive niche.
Immunological dimension
Spectrum of immune engagement in
PNETs
To fully understand the immunological consequences
of stromal remodeling, the preset review focuses on the spatial and
functional characterization of immune infiltration within the PNET
microenvironment. The immune contexture of PNETs appears to be a
key determinant of disease progression, metastatic potential and
therapeutic responsiveness (67–69).
Multi-omics approaches including bulk RNA-seq, scRNA-seq and immune
deconvolution enabled the classification of PNETs into distinct
immunophenotypes: Immune-cold, -active and -suppressive (41,60,70).
According to low immune enrichment score, immune-cold tumors
displayed sparse lymphocyte infiltration and diminished expression
of antigen-presentation machinery. By contrast, the gene signature
of the immune-hot tumor subgroup, defined by the nearest template
prediction algorithm, demonstrated a notable association between
activated inflammatory stromal response and immune suppression
(60,71–73).
Immune-active tumors, a subgroup of immune-hot tumor, are
characterized by abundant cytotoxic T cells and M1-like
macrophages, elevated interferon-γ signaling and upregulation of
costimulatory molecules, features indicative of a partially
activated or exhausted immune response (60,74).
Immune-suppressive tumors with activated stroma display high
infiltration of regulatory T cells, M2-polarized macrophages and
elevated expression levels of immune checkpoint ligands such as
programmed cell death-ligand 1 (75), indoleamine 2,3 dioxygenase 1 (IDO1)
and T cell immunoglobulin and mucin-domain containing-3 (TIM-3),
along with TGF-β pathway activation (41,60).
Therefore, the immune-cold phenotype presents a unique clinical
challenge; these tumors may be inherently less immunogenic due to
low TMB or defects in antigen presentation machinery [for example,
downregulation of major histocompatibility complex (MHC) class I],
making them resistant to current immunotherapies (41). Furthermore, multiplex spatial
profiles validated that these immune profiles are spatially
compartmentalized within the TME of PNETs (70). The metastasis-like primary subtype
defined by micro RNA profiles, presenting increased macrophages
under hypoxia and necroptosis, represents a key axis of immune
suppression (36,76). This creates a malignant cycle where
hypoxia induces immunosuppression, which further promotes tumor
growth and aggravates hypoxia. Although the clinical relevance of
immunophenotypes cannot be overstated, these phenotypes are
strongly associated with tumor grade and microenvironmental
features, providing a key framework for prognostic stratification
and identifies distinct patient subsets requiring tailored
therapeutic approaches.
Spatial and molecular determinants of
immune modulation
The stromal barrier may serve a notable role in
limiting immune infiltration (77).
Spatially resolved transcriptomic and proteo-transcriptomic
profiling of PNETs further clarified how architectural and
molecular features of the TME regulate immune infiltration and
function (44,60,78).
Immune-active regions are often segregated by fibrotic stroma
enriched in CAFs and immunosuppressive cytokines, which limit
lymphocyte trafficking and activation. These barriers are
maintained by signaling pathways such as TGF-β and
Hippo-yes-associated protein (YAP)/transcriptional coactivator with
PDZ-binding motif (TAZ), which promote ECM deposition and immune
exclusion (59,61,79–81).
By contrast, immune-permissive zones display higher expression
levels of MHC class I and II molecules, chemokines and
costimulatory ligands that facilitate effector T-cell recruitment
and retention (41,82). The juxtaposition of immune-active
and -silent regions within the same tumor reflected the spatial
heterogeneity of immunological pressure and underscored the
limitations of bulk profiling in capturing clinically relevant
immune dynamics. Overall, these findings underscored the necessity
of spatially integrated multi-omics approaches to fully resolve the
immunological landscape of PNETs. The spatial compartmentalization
of the immune response suggests that effective immunotherapy should
overcome dual challenges: Boosting immune activation in permissive
zones and breaking through immune suppression in excluded zones.
This necessitates spatially informed combination strategies.
Biomarker identification and translational
implications
The integration of multi-omics technologies
accelerated the identification of clinically relevant biomarkers
embedded within the PNET microenvironment. In addition to biomarker
identification, the multi-omics studies discussed in the present
review also highlight several molecular events and translationally
relevant targets in PNETs, as summarized in Table I. These biomarkers may hold notable
promise in refining prognosis, predicting therapeutic response and
guiding personalized interventions, which can be categorized as
prognostic, predictive and pharmacodynamic. Among the emerging
candidates, versican (VCAN), a stromal proteoglycan enriched in
mesenchymal subtypes was proposed as a prognostic biomarker and
detectable in both tumor tissue and plasma of PNETs (60). For example, a patient with a
localized G2 tumor but a VCAN-high/immune-cold profile might be
considered for more frequent imaging or adjuvant therapy trials.
Also, transcriptional co-activators YAP1 and TAZ, central mediators
of Hippo pathway signaling, are preferentially activated in
stromal-rich, immune-excluded tumors and represent potential
therapeutic targets for patients with PNET (78). Predictive biomarkers are used to
select patients for specific therapies. From an immunological
perspective, the expression levels of immune checkpoint molecules
including PD-L1, TIM-3 and IDO1 aligns with suppressive immune
phenotypes and may identify patients most likely to benefit from
immune checkpoint blockade (60).
Furthermore, detecting specific mutations (for example, MEN1 loss)
or activation signatures (for example, Myc and TGF-β) can guide the
use of targeted agents, such as epigenetic modulators or kinase
inhibitors. Lastly, pharmacodynamic biomarkers (for example,
changes in circulating VCAN levels or immune cell subsets in
peripheral blood) can be used in early-phase clinical trials to
confirm that a drug is acting on its intended target and modulating
the TME as expected (83,84).
 | Table I.Key molecular events and
translationally relevant targets discussed in PNETs. |
Table I.
Key molecular events and
translationally relevant targets discussed in PNETs.
| Biological
processes | Target/pathway | Study type | Mechanistic roles
in PNETs | Potential
translational implications | (Refs.) |
|---|
| Chromatin
remodeling | MEN1 | Genomic
studies | Loss of MEN1
disrupts chromatin regulation and transcriptional control,
contributing to tumor initiation | Molecular
stratification; rationale for epigenetic susceptibility
studies | (5,31,101) |
| Telomere
maintenance/chromatin remodeling | ATRX/DAXX | Genomic
studies | Mutations
associated with ALT and chromosomal instability | Biomarker of
aggressive biology; potential DNA repair or ALT-related
susceptibility | (5,29,45) |
| Cell cycle
regulation | CDK6 | Proteogenomic
analysis | Amplification
promotes proliferation and cell cycle progression in aggressive
tumors | Candidate target
for cell-cycle-directed therapies | (42) |
| Epigenetic
regulation | EZH2 | Transcriptomic and
proteomic studies | Histone
methyltransferase involved in transcriptional silencing and immune
modulation | Candidate target
for epigenetic therapy | (43) |
| Stromal
signaling | TGF-β1/TGF-β
pathway | Spatial
transcriptomics and proteomic studies | Promotes stromal
remodeling, immune suppression and EMT-like phenotypes | Candidate target
for stromal reprogramming and combination therapy | (41,44) |
| Hippo
signaling | YAP1/TAZ | Multi-omics
profiling | Activated in
stromal-rich and immune-excluded tumors | Candidate target in
stroma-rich PNETs | (78) |
| Immune evasion | PD-L1, TIM-3 and
IDO1 | Multi-omics
profiling | Associated with
immune-suppressive microenvironment and T-cell dysfunction | Biomarker-guided
selection for immunotherapy-based approaches | (60) |
| Stromal
biomarker | VCAN | Multi-omics
biomarker studies | Enriched in
mesenchymal/stromal subtype and detectable in tumor tissue and
plasma | Prognostic
biomarker and disease-monitoring candidate |
|
| DNA damage
response | FANCD2-associated
DNA repair axis | Multi-omics
biomarker studies | Stromal
proteoglycan enriched in mesenchymal subtype; potential prognostic
biomarker | Precision oncology
rationale in selected genomically unstable cases | (49) |
Spatial and proteogenomic profiling further refines
these associations by capturing context-specific expression
patterns within distinct tumor-stroma compartments (58,60).
These findings support a potential combination therapeutic strategy
for stroma-rich PNETs. The proposed combination strategy (YAP/TAZ
inhibition, immune checkpoint blockade and TGF-β suppression)
warrants evaluation in rationally designed clinical trials, for
example in patients with advanced, stroma-rich PNETs (78). The strategy emphasizes the value of
multi-omics-informed stratification for personalized therapy.
Furthermore, a previous case study of FANCD2 exemplified the
potential of multi-omics for precision oncology; however, its
extensive implementation faces notable hurdles (49). The high cost, complex data analysis
and longer turnaround time of techniques like scRNA-seq currently
preclude the use of multi-omics in routine diagnostics. The
ultimate goal is a future where the treatment of each patient with
PNET is guided by enhanced understanding of the unique ecosystem of
the tumor, moving from a one-size-fits-all approach to personalized
ecosystem-based therapy.
Artificial intelligence (AI) and
computational integration in PNET research
The rapid expansion of multi-omics datasets in PNETs
has generated novel opportunities in computational modeling and
AI-driven identification. In recent years, machine learning and
deep learning approaches have demonstrated growing utility in
cancer diagnosis, grading and risk stratification across multiple
tumor types, including breast and pancreatic cancer (85,86).
In the context of PNETs, AI-assisted digital pathology and
radiomics analyses may improve diagnostic accuracy, particularly in
distinguishing well-differentiated G3 tumors from poorly
differentiated neuroendocrine carcinomas, which often present
overlapping morphological features (87–91).
Image-based deep learning algorithms have the potential to
integrate histological architecture with molecular signatures,
thereby enhancing grading precision and predicting metastatic
potential (88).
Beyond diagnostic applications, AI offers key tools
in integrating complex multi-layered omics datasets. PNET biology
is characterized by heterogeneous genomic alterations (for example,
MEN1 and ATRX/DAXX mutations), metabolic reprogramming, stromal
remodeling and immune heterogeneity (5,38,45).
Computational frameworks such as graph-based neural networks,
multi-modal data fusion algorithms and representation learning
models may enable simultaneous integration of genomic,
transcriptomic, proteomic and spatial transcriptomic data (92–95).
These approaches could reveal latent molecular patterns that are
not apparent through single-layer analysis and may improve the
identification of clinically actionable subtypes. Notably,
AI-driven modeling may facilitate prediction of TME interactions,
vascular niche activation and potential immune evasion mechanisms
(96).
Although AI applications in PNETs remain in early
development and are not yet standardized for clinical practice,
future directions are promising. Prospective integration of spatial
transcriptomics with AI-based spatial modeling could enable
reconstruction of tumor-stroma co-evolution dynamics (97,98).
Furthermore, predictive modeling of therapeutic
response-particularly to targeted inhibitors, anti-angiogenic
agents or immune checkpoint blockade may improve patient
stratification (97). However,
challenges remain, including limited cohort sizes in rare tumors
such as PNETs, data harmonization issues and the need for
interpretability of AI models. Addressing these limitations through
collaborative consortia and standardized pipelines will be key to
translating AI-driven insights into precision oncology for patients
with PNET.
Limitations and challenges of multi-omics
approaches in PNET research
Despite the transformative potential of multi-omics
technologies, several limitations should be considered when
interpreting findings in PNETs. First, numerous multi-omics studies
rely on relatively small patient cohorts due to the rarity of
PNETs, which may limit statistical power and reduce the
generalizability of findings (35,38,49).
Second, technical variability arising from sample processing,
sequencing platforms and data normalization strategies can
introduce batch effects and complicate cross-study comparisons. In
single-cell and spatial transcriptomic analyses, tissue
dissociation procedures may alter cellular states or lead to
selective loss of fragile cell populations, potentially biasing the
representation of the TME (38,44,49).
Furthermore, integrating heterogeneous data types, including
genomics, transcriptomics, proteomics and spatial data remains
computationally challenging and lacks standardized analytical
pipelines (99). Lastly, while
multi-omics studies have generated notable descriptive insights
into TME interactions (35,58,78),
translating these findings into clinically actionable biomarkers or
therapeutic strategies requires further functional validation and
prospective clinical studies. Addressing these challenges will be
key to improving the reproducibility and clinical applicability of
multi-omics research in PNETs in the future.
Future outlook and conclusions
The advent of multi-omics technologies markedly
transformed current understanding of PNETs. Once classified
primarily by morphology and proliferation indices, these tumors are
now recognized as dynamic, spatially structured ecosystems shaped
by genomic instability, transcriptional diversity and
microenvironmental remodeling (Fig.
3). The integrated evidence synthesized in the present review
established that malignant progression in PNETs is not solely
driven by cell-intrinsic mechanisms or extrinsic microenvironmental
pressures alone, but by the continuous, bidirectional co-evolution
of PNETs. Genomic instability may act as a key contributor to tumor
evolution, not only promoting tumor cell dedifferentiation and
heterogeneity but also actively remodeling the surrounding niche
(38). By contrast, a
microenvironment dominated by immunosuppressive CAFs and myeloid
cells exerts potent selective pressures, fostering the expansion of
subclones equipped with adaptive traits such as immune evasion and
metabolic plasticity (66,100). This ‘genome-microenvironment’
co-evolution framework may provide a novel paradigm in
understanding the aggressive and therapy-resistant nature of
advanced PNETs.
 | Figure 3.Chromosomal alterations orchestrate
stromal remodeling and immune modulation in PNETs. Large-scale
chromosomal gains, structural variants and ECM deposition
contribute to stromal remodeling in PNETs. These stromal changes
are associated with molecular drivers such as TGF-β signaling and
hypoxia, which promote CAF activation and ECM remodeling. CAF
expansion and activation, marked by α-SMA expression, contribute to
ECM accumulation and fibrosis. The remodeled stroma shapes the
immune landscape by polarizing macrophages (M1-like vs. M2),
recruiting Treg cells and excluding CD8+ T cells,
collectively establishing an immune-evasive microenvironment. ECM,
extracellular matrix; CAF, cancer-associated fibroblast; PNETs,
pancreatic neuroendocrine tumors; Treg, regulatory T cells; α-SMA,
α-smooth muscle actin. |
Despite recent progress, several key questions
remain unanswered, which may be explored in future research. The
precise cell(s) of origin for different PNET subtypes remain to be
elucidated. Furthermore, a key unanswered question is how
therapeutic interventions (for example, targeted therapy and
chemotherapy) reshape the spatiotemporal evolutionary trajectory of
the PNET ecosystem. Prospective longitudinal studies incorporating
multi-omics analysis of paired pre- and post-treatment samples
(including liquid biopsies) will provide an enhanced view of
resistance evolution. The metabolic coupling between tumor cells
and stromal components is an underexplored area. Understanding how
specific signals guide the formation of immune niches and how
tumor-stroma metabolic interactions evolve under therapy is key to
therapeutic reprogramming. Addressing spatial heterogeneity remains
a major hurdle, as well as the identification of reliable
biomarkers and the development of physiologically relevant
preclinical models. Several strategies are warranted to progress
research in this field. Advanced organoid and microfluidic systems
will be key to modeling tumor-immune-stroma interactions. Humanized
mouse models that preserve immune function will be key to
validating novel therapies in the future.
In summary, validating the prognostic utility of
multi-omics signatures warrants integration of multi-omics
signatures into prospective clinical trials. For instance, a
‘window-of-opportunity’ trial design could be employed: Patients
scheduled for resection of locally advanced PNETs receive a short
course of a candidate agent (for example, a YAP/TAZ inhibitor or
TGF-β blocker) prior to surgery. Paired pre- and post-treatment
multi-omics analyses (scRNA-seq and spatial transcriptomics) of the
resected specimens would then directly assess the impact of the
drug on both tumor cell states and the TME, providing key
mechanistic pharmacodynamic data. Furthermore, overcoming the
technical and computational barriers to implementing these
technologies in routine pathological workflows is a prerequisite
for clinical translation. This will warrant the development of
streamlined cost-effective assays and automated bioinformatics
pipelines in the future.
Acknowledgements
Not applicable.
Funding
The present review was partly supported by the National
Institutes of Health/National Cancer Institute Cancer Center
Support (grant no. P30CA022453) and the Cancer Research Horizons
Fund Grant at the Karmanos Cancer Institute.
Availability of data and materials
Not applicable.
Authors' contributions
YW, AA and YS conceptualized the present review. YW
and YS devised the methodology. YW conducted the formal analysis
and investigation, and obtained resources. YW, YZ and YS prepared
the original draft of the manuscript. AA, PP, HC and BE reviewed
and edited the manuscript. AA and YS supervised the present review,
participated in project administration and obtained funding. All
authors read and approved the final manuscript. Data authentication
is not applicable.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Reid MD, Balci S, Saka B and Adsay NV:
Neuroendocrine tumors of the pancreas: Current concepts and
controversies. Endocr Pathol. 25:65–79. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Lawrence B, Gustafsson BI, Chan A, Svejda
B, Kidd M and Modlin IM: The epidemiology of gastroenteropancreatic
neuroendocrine tumors. Endocrinol Metab Clin North Am. 40:1–18.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Cives M and Strosberg JR:
Gastroenteropancreatic neuroendocrine tumors. CA Cancer J Clin.
68:471–487. 2018.PubMed/NCBI
|
|
4
|
Simon T, Riemer P, Jarosch A, Detjen K, Di
Domenico A, Bormann F, Menne A, Khouja S, Monjé N, Childs LH, et
al: DNA methylation reveals distinct cells of origin for pancreatic
neuroendocrine carcinomas and pancreatic neuroendocrine tumors.
Genome Med. 14:242022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chan CS, Laddha SV, Lewis PW, Koletsky MS,
Robzyk K, Da Silva E, Torres PJ, Untch BR, Li J, Bose P, et al:
ATRX, DAXX or MEN1 mutant pancreatic neuroendocrine tumors are a
distinct alpha-cell signature subgroup. Nat Commun. 9:41582018.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Halfdanarson TR, Rubin J, Farnell MB,
Grant CS and Petersen GM: Pancreatic endocrine neoplasms:
Epidemiology and prognosis of pancreatic endocrine tumors. Endocr
Relat Cancer. 15:409–427. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Hopper AD, Jalal M and Munir A: Recent
advances in the diagnosis and management of pancreatic
neuroendocrine tumours. Frontline Gastroenterol. 10:269–274. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Garcia-Carbonero R, Capdevila J,
Crespo-Herrero G, Díaz-Pérez JA, Martínez Del Prado MP, Orduña VA,
Sevilla-García I, Villabona-Artero C, Beguiristain-Gómez A,
Llanos-Muñoz M, et al: Incidence, patterns of care and prognostic
factors for outcome of gastroenteropancreatic neuroendocrine tumors
(GEP-NETs): Results from the national cancer registry of Spain
(RGETNE). Ann Oncol. 21:1794–1803. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Singh S, Chan DL, Moody L, Liu N, Fischer
HD, Austin PC and Segelov E: Recurrence in resected
gastroenteropancreatic neuroendocrine tumors. JAMA Oncol.
4:583–585. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Frilling A, Modlin IM, Kidd M, Russell C,
Breitenstein S, Salem R, Kwekkeboom D, Lau WY, Klersy C, Vilgrain
V, et al: Recommendations for management of patients with
neuroendocrine liver metastases. Lancet Oncol. 15:e8–e21. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Raymond E, Dahan L, Raoul JL, Bang YJ,
Borbath I, Lombard-Bohas C, Valle J, Metrakos P, Smith D, Vinik A,
et al: Sunitinib malate for the treatment of pancreatic
neuroendocrine tumors. N Engl J Med. 364:501–513. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Vagefi PA, Razo O, Deshpande V, McGrath
DJ, Lauwers GY, Thayer SP, Warshaw AL and Fernández-Del Castillo C:
Evolving patterns in the detection and outcomes of pancreatic
neuroendocrine neoplasms: The Massachusetts General Hospital
experience from 1977 to 2005. Arch Surg. 142:347–354. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
McKenna LR and Edil BH: Update on
pancreatic neuroendocrine tumors. Gland Surg. 3:258–275.
2014.PubMed/NCBI
|
|
14
|
Inzani F, Petrone G and Rindi G: The new
world health organization classification for pancreatic
neuroendocrine neoplasia. Endocrinol Metab Clin North Am.
47:463–470. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Nagtegaal ID, Odze RD, Klimstra D, Paradis
V, Rugge M, Schirmacher P, Washington KM, Carneiro F and Cree IA;
WHO Classification of Tumours Editorial Board, : The 2019 WHO
classification of tumours of the digestive system. Histopathology.
76:182–188. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tang LH, Untch BR, Reidy DL, O'Reilly E,
Dhall D, Jih L, Basturk O, Allen PJ and Klimstra DS:
Well-differentiated neuroendocrine tumors with a morphologically
apparent high-grade component: A pathway distinct from poorly
differentiated neuroendocrine carcinomas. Clin Cancer Res.
22:1011–1017. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Crona J, Norlén O, Antonodimitrakis P,
Welin S, Stålberg P and Eriksson B: Multiple and secondary hormone
secretion in patients with metastatic pancreatic neuroendocrine
tumours. J Clin Endocrinol Metab. 101:445–452. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
de Mestier L, Hentic O, Cros J, Walter T,
Roquin G, Brixi H, Lombard-Bohas C, Hammel P, Diebold MD, Couvelard
A, et al: Metachronous hormonal syndromes in patients with
pancreatic neuroendocrine tumors: A case-series study. Ann Intern
Med. 162:682–689. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Botling J, Lamarca A, Bajic D, Norlén O,
Lönngren V, Kjaer J, Eriksson B, Welin S, Hellman P, Rindi G, et
al: High-grade progression confers poor survival in pancreatic
neuroendocrine tumors. Neuroendocrinology. 110:891–898. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Moertel CG, Kvols LK, O'Connell MJ and
Rubin J: Treatment of neuroendocrine carcinomas with combined
etoposide and cisplatin. Evidence of major therapeutic activity in
the anaplastic variants of these neoplasms. Cancer. 68:227–232.
1991. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yao JC, Shah MH, Ito T, Bohas CL, Wolin
EM, Van Cutsem E, Hobday TJ, Okusaka T, Capdevila J, de Vries EG,
et al: Everolimus for advanced pancreatic neuroendocrine tumors. N
Engl J Med. 364:514–523. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
de Visser KE and Joyce JA: The evolving
tumor microenvironment: From cancer initiation to metastatic
outgrowth. Cancer Cell. 41:374–403. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xiao Y and Yu D: Tumor microenvironment as
a therapeutic target in cancer. Pharmacol Ther. 221:1077532021.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Shi H, Jiang C, Zhang Q, Qi C, Yao H and
Lin R: Clinicopathological heterogeneity between primary and
metastatic sites of gastroenteropancreatic neuroendocrine neoplasm.
Diagn Pathol. 15:1082020. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang J, Huang D, Saw PE and Song E:
Turning cold tumors hot: From molecular mechanisms to clinical
applications. Trends Immunol. 43:523–545. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang L, Geng H, Liu Y, Liu L, Chen Y, Wu
F, Liu Z, Ling S, Wang Y and Zhou L: Hot and cold tumors:
Immunological features and the therapeutic strategies. MedComm
(2020). 4:e3432023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Baghban R, Roshangar L, Jahanban-Esfahlan
R, Seidi K, Ebrahimi-Kalan A, Jaymand M, Kolahian S, Javaheri T and
Zare P: Tumor microenvironment complexity and therapeutic
implications at a glance. Cell Commun Signal. 18:592020. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Scarpa A, Chang DK, Nones K, Corbo V,
Patch AM, Bailey P, Lawlor RT, Johns AL, Miller DK, Mafficini A, et
al: Whole-genome landscape of pancreatic neuroendocrine tumours.
Nature. 543:65–71. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Jiao Y, Shi C, Edil BH, de Wilde RF,
Klimstra DS, Maitra A, Schulick RD, Tang LH, Wolfgang CL, Choti MA,
et al: DAXX/ATRX, MEN1, and mTOR pathway genes are frequently
altered in pancreatic neuroendocrine tumors. Science.
331:1199–1203. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hong X, Qiao S, Li F, Wang W, Jiang R, Wu
H, Chen H, Liu L, Peng J, Wang J, et al: Whole-genome sequencing
reveals distinct genetic bases for insulinomas and non-functional
pancreatic neuroendocrine tumours: Leading to a new classification
system. Gut. 69:877–887. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Park JK, Paik WH, Lee K, Ryu JK, Lee SH
and Kim YT: DAXX/ATRX and MEN1 genes are strong prognostic markers
in pancreatic neuroendocrine tumors. Oncotarget. 8:49796–49806.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zhou W, Han X, Ji Y, Wang D, Xie D, Qiu Z
and Lou W: Targeted deep sequencing reveals the genetic
heterogeneity in well-differentiated pancreatic neuroendocrine
tumors with liver metastasis. Hepatobiliary Surg Nutr. 12:302–313.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Abbas T, Keaton MA and Dutta A: Genomic
instability in cancer. Cold Spring Harb Perspect Biol.
5:a0129142013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ciriello G, Miller ML, Aksoy BA,
Senbabaoglu Y, Schultz N and Sander C: Emerging landscape of
oncogenic signatures across human cancers. Nat Genet. 45:1127–1133.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhou Y, Liu S, Liu C, Yang J, Lin Q, Zheng
S, Chen C, Zhou Q and Chen R: Single-cell RNA sequencing reveals
spatiotemporal heterogeneity and malignant progression in
pancreatic neuroendocrine tumor. Int J Biol Sci. 17:3760–3775.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Sadanandam A, Wullschleger S, Lyssiotis
CA, Grötzinger C, Barbi S, Bersani S, Körner J, Wafy I, Mafficini
A, Lawlor RT, et al: A cross-species analysis in pancreatic
neuroendocrine tumors reveals molecular subtypes with distinctive
clinical, metastatic, developmental, and metabolic characteristics.
Cancer Discov. 5:1296–1313. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Miki M, Oono T, Fujimori N, Takaoka T,
Kawabe K, Miyasaka Y, Ohtsuka T, Saito D, Nakamura M, Ohkawa Y, et
al: CLEC3A, MMP7, and LCN2 as novel markers for predicting
recurrence in resected G1 and G2 pancreatic neuroendocrine tumors.
Cancer Med. 8:3748–3760. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ye Z, Zhou Y, Hu Y, Li Q, Xu Z, Lou X,
Zhang W, Zhu D, Xie C, Zhou Q, et al: Single-cell sequencing
reveals the heterogeneity of pancreatic neuroendocrine tumors under
genomic instability and histological grading. iScience.
27:1108362024. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhao Y, Shen M, Wu L, Yang H, Yao Y, Yang
Q, Du J, Liu L, Li Y and Bai Y: Stromal cells in the tumor
microenvironment: Accomplices of tumor progression? Cell Death Dis.
14:5872023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Li M, Gao X and Wang X: Identification of
tumor mutation burden-associated molecular and clinical features in
cancer by analyzing multi-omics data. Front Immunol.
14:10908382023. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Tanaka A, Ogawa M, Zhou Y, Otani Y,
Hendrickson RC, Miele MM, Li Z, Klimstra DS, Wang JY and Roehrl MH:
Proteogenomic characterization of pancreatic neuroendocrine tumors
uncovers hypoxia and immune signatures in clinically aggressive
subtypes. iScience. 27:1105442024. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Shi Y, Qian ZR, Zhang S, Li W, Masugi Y,
Li T, Chan JA, Yang J, Da Silva A, Gu M, et al: Cell cycle protein
expression in neuroendocrine tumors: Association of CDK4/CDK6,
CCND1, and phosphorylated retinoblastoma protein with proliferative
index. Pancreas. 46:1347–1353. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
April-Monn SL, Andreasi V, Lena MS,
Sadowski MC, Kim-Fuchs C, Buri MC, Ketkar A, Maire R, Di Domenico
A, Schrader J, et al: EZH2 inhibition as new epigenetic treatment
option for pancreatic neuroendocrine neoplasms (PanNENs). Cancers
(Basel). 13:50142021. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Niedra H, Peculis R, Saksis R, Mandrika I,
Vilisova S, Nazarovs J, Breiksa A, Gerina A, Earl J, Ruz-Caracuel
I, et al: Tumor and α-SMA-expressing stromal cells in pancreatic
neuroendocrine tumors have a distinct RNA profile depending on
tumor grade. Mol Oncol. 19:659–681. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Gisder DM, Overheu O, Keller J,
Nöpel-Dünnebacke S, Uhl W, Reinacher-Schick A, Tannapfel A and
Tischoff I: DAXX, ATRX, and MSI in PanNET and their metastases:
Correlation with histopathological data and prognosis.
Pathobiology. 90:71–80. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Heaphy CM and Singhi AD: Reprint of: The
diagnostic and prognostic utility of incorporating DAXX, ATRX, and
alternative lengthening of telomeres (ALT) to the Evaluation of
pancreatic neuroendocrine tumors (PanNETs). Hum Pathol. 132:1–11.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Marinoni I, Kurrer AS, Vassella E, Dettmer
M, Rudolph T, Banz V, Hunger F, Pasquinelli S, Speel EJ and Perren
A: Loss of DAXX and ATRX are associated with chromosome instability
and reduced survival of patients with pancreatic neuroendocrine
tumors. Gastroenterology. 146:453–460. e52014. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Backman S, Botling J, Nord H, Ghosal S,
Stålberg P, Juhlin CC, Almlöf J, Sundin A, Zhang L, Moens L, et al:
The evolutionary history of metastatic pancreatic neuroendocrine
tumours reveals a therapy driven route to high-grade
transformation. J Pathol. 264:357–370. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Avsievich E, Salimgereeva D, Maluchenko A,
Antysheva Z, Voloshin M, Feidorov I, Glazova O, Abramov I, Maksimov
D and Kaziakhmedova S: Pancreatic neuroendocrine tumor: The case
report of a patient with germline FANCD2 mutation and tumor
analysis using single-cell RNA sequencing. J Clin Med. 13:76212024.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Pilati C, Shinde J, Alexandrov LB, Assié
G, André T, Hélias-Rodzewicz Z, Ducoudray R, Le Corre D,
Zucman-Rossi J, Emile JF, et al: Mutational signature analysis
identifies MUTYH deficiency in colorectal cancers and
adrenocortical carcinomas. J Pathol. 242:10–15. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Zhang JH, Tasaki T, Tsukamoto M, Wang KY
and Azuma K: Deficiency of Wnt10a causes female infertility via the
beta-catenin/Cyp19a1 pathway in mice. Int J Med Sci. 19:701–710.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zhang JH, Tasaki T, Tsukamoto M, Wang KY,
Kubo KY and Azuma K: Deletion of Wnt10a is implicated in
hippocampal neurodegeneration in mice. Biomedicines. 10:15002022.
View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Li P, Liu W, Xu Q and Wang C: Clinical
significance and biological role of Wnt10a in ovarian cancer. Oncol
Lett. 14:6611–6617. 2017.PubMed/NCBI
|
|
54
|
Liu W, Polaczek P, Roubal I, Meng Y, Choe
WC, Caron MC, Sedgeman CA, Xi Y, Liu C, Wu Q, et al: FANCD2 and
RAD51 recombinase directly inhibit DNA2 nuclease at stalled
replication forks and FANCD2 acts as a novel RAD51 mediator in
strand exchange to promote genome stability. Nucleic Acids Res.
51:9144–9165. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Lee GT, Rosenfeld JA, Kim WT, Kwon YS,
Palapattu G, Mehra R, Kim WJ and Kim IY: TCF4 induces enzalutamide
resistance via neuroendocrine differentiation in prostate cancer.
PLoS One. 14:e02134882019. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Forrest MP, Hill MJ, Quantock AJ,
Martin-Rendon E and Blake DJ: The emerging roles of TCF4 in disease
and development. Trends Mol Med. 20:322–331. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Ruan K, Song G and Ouyang G: Role of
hypoxia in the hallmarks of human cancer. J Cell Biochem.
107:1053–1062. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Ye Z, Li Q, Hu Y, Hu H, Xu J, Guo M, Zhang
W, Lou X, Wang Y, Gao H, et al: The stromal microenvironment endows
pancreatic neuroendocrine tumors with spatially specific invasive
and metastatic phenotypes. Cancer Lett. 588:2167692024. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Lambert AW, Pattabiraman DR and Weinberg
RA: Emerging biological principles of metastasis. Cell.
168:670–691. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Ji S, Cao L, Gao J, Du Y, Ye Z, Lou X, Liu
F, Zhang Y, Xu J, Shi X, et al: Proteogenomic characterization of
non-functional pancreatic neuroendocrine tumors unravels clinically
relevant subgroups. Cancer Cell. 43:776–796. e142025. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Mittal V: Epithelial Mesenchymal
transition in tumor metastasis. Annu Rev Pathol. 13:395–412. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Deshmukh AP, Vasaikar SV, Tomczak K,
Tripathi S, den Hollander P, Arslan E, Chakraborty P, Soundararajan
R, Jolly MK, Rai K, et al: Identification of EMT signaling
cross-talk and gene regulatory networks by single-cell RNA
sequencing. Proc Natl Acad Sci USA. 118:e21020501182021. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Ohlund D, Handly-Santana A, Biffi G,
Elyada E, Almeida AS, Ponz-Sarvise M, Corbo V, Oni TE, Hearn SA,
Lee EJ, et al: Distinct populations of inflammatory fibroblasts and
myofibroblasts in pancreatic cancer. J Exp Med. 214:579–596. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Helms EJ, Berry MW, Chaw RC, DuFort CC,
Sun D, Onate MK, Oon C, Bhattacharyya S, Sanford-Crane H, Horton W,
et al: Mesenchymal lineage heterogeneity underlies nonredundant
functions of pancreatic cancer-associated fibroblasts. Cancer
Discov. 12:484–501. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Glabman RA, Choyke PL and Sato N:
Cancer-associated fibroblasts: Tumorigenicity and targeting for
cancer therapy. Cancers (Basel). 14:39062022. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Caligiuri G and Tuveson DA: Activated
fibroblasts in cancer: Perspectives and challenges. Cancer Cell.
41:434–449. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Bruni D, Angell HK and Galon J: The immune
contexture and Immunoscore in cancer prognosis and therapeutic
efficacy. Nat Rev Cancer. 20:662–680. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Greenberg J, Limberg J, Verma A, Kim D,
Chen X, Lee YJ, Moore MD, Ullmann TM, Thiesmeyer JW, Loewenstein Z,
et al: Metastatic pancreatic neuroendocrine tumors feature elevated
T cell infiltration. JCI Insight. 7:e1601302022. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Meng D, Zhao L, Liu J, Ge C and Zhang C:
Identification of the immune subtypes for the prediction of
metastasis in pancreatic neuroendocrine tumors. Neuroendocrinology.
113:719–735. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Young K, Lawlor RT, Ragulan C, Patil Y,
Mafficini A, Bersani S, Antonello D, Mansfield D, Cingarlini S,
Landoni L, et al: Immune landscape, evolution, hypoxia-mediated
viral mimicry pathways and therapeutic potential in molecular
subtypes of pancreatic neuroendocrine tumours. Gut. 70:1904–1913.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Sia D, Jiao Y, Martinez-Quetglas I, Kuchuk
O, Villacorta-Martin C, de Moura MC, Putra J, Camprecios G,
Bassaganyas L, Akers N, et al: Identification of an immune-specific
class of hepatocellular carcinoma, based on molecular features.
Gastroenterology. 153:812–826. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Moffitt RA, Marayati R, Flate EL, Volmar
KE, Loeza SG, Hoadley KA, Rashid NU, Williams LA, Eaton SC, Chung
AH, et al: Virtual microdissection identifies distinct tumor- and
stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat
Genet. 47:1168–1178. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Yoshihara K, Shahmoradgoli M, Martínez E,
Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW,
Levine DA, et al: Inferring tumour purity and stromal and immune
cell admixture from expression data. Nat Commun. 4:26122013.
View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Flavell RA, Sanjabi S, Wrzesinski SH and
Licona-Limón P: The polarization of immune cells in the tumour
environment by TGFbeta. Nat Rev Immunol. 10:554–567. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Cheung A, Chenoweth AM, Quist J, Sow HS,
Malaktou C, Ferro R, Hoffmann RM, Osborn G, Sachouli E, French E,
et al: CDK inhibition primes for anti-PD-L1 treatment in
triple-negative breast cancer models. Cancers (Basel). 14:33612022.
View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Olson P, Lu J, Zhang H, Shai A, Chun MG,
Wang Y, Libutti SK, Nakakura EK, Golub TR and Hanahan D: MicroRNA
dynamics in the stages of tumorigenesis correlate with hallmark
capabilities of cancer. Genes Dev. 23:2152–2165. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Mhaidly R and Mechta-Grigoriou F: Role of
cancer-associated fibroblast subpopulations in immune infiltration,
as a new means of treatment in cancer. Immunol Rev. 302:259–272.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Yang KC, Kalloger SE, Aird JJ, Lee MKC,
Rushton C, Mungall KL, Mungall AJ, Gao D, Chow C, Xu J, et al:
Proteotranscriptomic classification and characterization of
pancreatic neuroendocrine neoplasms. Cell Rep. 37:1098172021.
View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Tseng PC, Chen CL, Lee KY, Feng PH, Wang
YC, Satria RD and Lin CF: Epithelial-to-mesenchymal transition
hinders interferon-gamma-dependent immunosurveillance in lung
cancer cells. Cancer Lett. 539:2157122022. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Wang X, Eichhorn PJA and Thiery JP: TGF-β,
EMT, and resistance to anti-cancer treatment. Semin Cancer Biol.
97:1–11. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Guo Z, Ashrafizadeh M, Zhang W, Zou R,
Sethi G and Zhang X: Molecular profile of metastasis, cell
plasticity and EMT in pancreatic cancer: A pre-clinical connection
to aggressiveness and drug resistance. Cancer Metastasis Rev.
43:29–53. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Kraehenbuehl L, Weng CH, Eghbali S,
Wolchok JD and Merghoub T: Enhancing immunotherapy in cancer by
targeting emerging immunomodulatory pathways. Nat Rev Clin Oncol.
19:37–50. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Jo A, Green A, Medina JE, Iyer S, Ohman
AW, McCarthy ET, Reinhardt F, Gerton T, Demehin D, Mishra R, et al:
Inaugurating High-Throughput profiling of extracellular vesicles
for earlier ovarian cancer detection. Adv Sci (Weinh).
10:e23019302023. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Hoshino A, Kim HS, Bojmar L, Gyan KE,
Cioffi M, Hernandez J, Zambirinis CP, Rodrigues G, Molina H,
Heissel S, et al: Extracellular vesicle and particle biomarkers
define multiple human cancers. Cell. 182:1044–1061. e182020.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Kourou K, Exarchos TP, Exarchos KP,
Karamouzis MV and Fotiadis DI: Machine learning applications in
cancer prognosis and prediction. Comput Struct Biotechnol J.
13:8–17. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Tran KA, Kondrashova O, Bradley A,
Williams ED, Pearson JV and Waddell N: Deep learning in cancer
diagnosis, prognosis and treatment selection. Genome Med.
13:1522021. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Zhou RQ, Ji HC, Liu Q, Zhu CY and Liu R:
Leveraging machine learning techniques for predicting pancreatic
neuroendocrine tumor grades using biochemical and tumor markers.
World J Clin Cases. 7:1611–1622. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Ye JY, Fang P, Peng ZP, Huang XT, Xie JZ
and Yin XY: A radiomics-based interpretable model to predict the
pathological grade of pancreatic neuroendocrine tumors. Eur Radiol.
34:1994–2005. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Li J, Huang L, Liao C, Liu G, Tian Y and
Chen S: Two machine learning-based nomogram to predict risk and
prognostic factors for liver metastasis from pancreatic
neuroendocrine tumors: A multicenter study. BMC Cancer. 23:5292023.
View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Kiremitci S, Seven G, Silahtaroglu G,
Kochan K, Tosun SD and Senturk H: The role of artificial
intelligence and deep learning in determining the histopathological
grade of pancreatic neuroendocrine tumors by using EUS images.
Endosc Ultrasound. 14:48–56. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Momeni-Boroujeni A, Yousefi E and Somma J:
Computer-assisted cytologic diagnosis in pancreatic FNA: An
application of neural networks to image analysis. Cancer
Cytopathol. 125:926–933. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Xuan P, Sun H, Wang X, Zhang T and Pan S:
Inferring the disease-associated miRNAs Based on network
representation learning and convolutional neural networks. Int J
Mol Sci. 20:36482019. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Cristiano S, Leal A, Phallen J, Fiksel J,
Adleff V, Bruhm DC, Jensen SØ, Medina JE, Hruban C, White JR, et
al: Genome-wide cell-free DNA fragmentation in patients with
cancer. Nature. 570:385–389. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Gao H, Zheng Z, Yue Z, Liu F, Zhou L and
Zhao X: Evaluation of serum diagnosis of pancreatic cancer by using
surface-enhanced laser desorption/ionization time-of-flight mass
spectrometry. Int J Mol Med. 30:1061–1068. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Huang B, Huang H, Zhang S, Zhang D, Shi Q,
Liu J and Guo J: Artificial intelligence in pancreatic cancer.
Theranostics. 12:6931–6954. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Valanarasu JMJ, Xu H, Usuyama N, Kim C,
Wong C, Argaw P, Shimol RB, Crabtree A, Matlock K, Bartlett AQ, et
al: Multimodal AI generates virtual population for tumor
microenvironment modeling. Cell. 189:386–400. e192026. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Li Z, Li Y, Xiang J, Wang X, Yang S, Zhang
X, Eweje F, Chen Y, Luo X, Li Y, et al: AI-enabled virtual spatial
proteomics from histopathology for interpretable biomarker
discovery in lung cancer. Nat Med. 32:231–244. 2026. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Li Y, Stanojevic S and Garmire LX:
Emerging artificial intelligence applications in Spatial
Transcriptomics analysis. Comput Struct Biotechnol J. 20:2895–2908.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Hemme CL, Atoyan J, Cai A and Liu C:
Challenges and opportunities in multi-omics data acquisition and
analysis: Toward integrative solutions. Biomolecules. 16:2712026.
View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Costa A, Kieffer Y, Scholer-Dahirel A,
Pelon F, Bourachot B, Cardon M, Sirven P, Magagna I, Fuhrmann L,
Bernard C, et al: Fibroblast heterogeneity and immunosuppressive
environment in human breast cancer. Cancer Cell. 33:463–479.
e102018. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Corbo V, Dalai I, Scardoni M, Barbi S,
Beghelli S, Bersani S, Albarello L, Doglioni C, Schott C, Capelli
P, et al: MEN1 in pancreatic endocrine tumors: analysis of gene and
protein status in 169 sporadic neoplasms reveals alterations in the
vast majority of cases. Endocr Relat Cancer. 17:771–783. 2010.
View Article : Google Scholar : PubMed/NCBI
|