Introduction
microRNAs (miRNAs) are a class of approximately
20–25 nucleotide-non-coding RNAs that can regulate
post-transcriptional gene expression. These miRNAs may degrade the
target mRNAs or the translational repression of encoded proteins by
partially binding to complementary target sites in messenger RNA
(mRNA) 3′ untranslated regions (UTRs) (1).
Bioinformatic analyses have estimated that miRNAs
may regulate as many as 30% of the human protein coding genes
(2). In general, one gene can be
repressed by multiple miRNAs, and one miRNA may repress multiple
target genes. Using miRNA target prediction algorithms, hundreds of
potential target mRNAs for a specific miRNA can be identified.
miRNA target binding sites have been predicted to be present in
almost any transcript. Many miRNAs are expressed in a
tissue-specific manner and play pivotal roles in the control of
proliferation and differentiation of different cell types (3–5).
Many miRNAs have been shown to be downregulated or
upregulated in different tumors. miRNA expression profiles have
been reported to undergo changes in papillary thyroid cancer
(6), breast cancer (7), lung cancer (8), hepatocellular carcinoma (9), pancreatic adenocarcinoma (10) and colorectal cancer (11). In addition, deregulation of miRNAs
may lead to oncogenesis and cancer progression. Studies have shown
that miRNAs affect the expression of genes and pathways involved in
cancer pathogenesis from initiation to metastatic disease (12,13).
miRNAs are also involved in important homeostatic processes such as
cellular proliferation and cell apoptosis (14,15).
In our previous study, we found that miR-1301
expression was lower in the HepG2 cell line than in the Qsg7701
cell line which is a normal liver cell line. In this study, we
compared mRNA transcription, cell proliferation, migration ability,
invasion ability and apoptosis rate in the HepG2 cell line before
and after transfection with miR-1301 mimics to understand the role
of miR-1301 in the control of HepG2 cell apoptosis.
Materials and methods
Cell line, culture and transfection
The human cell line HepG2 (Cell Bank of Chinese
Academy of Sciences, Shanghai, China) were assigned to miR-1301
group and control group. The cells were maintained in monolayer
cultures in high glucose Dulbecco’s modified Eagle’s medium (DMEM)
containing 10% (v/v) fetal bovine serum (FBS) and 1% (v/v)
penicillin at 37°C in a humidified atmosphere of 5% CO2
in air.
Cells at the log phase were harvested and a cell
suspension of 3.0×104 cells/ml was prepared in 96-well
tissue culture plates. miR-1301 mimics (GenePharma Co., Ltd.,
Shanghai, China) were introduced into cells by the procedure
according to the manufacturer’s instructions. miR-1301 mimics were
mixed with Lipofectamine transfection reagent (GenePharma Co.,
Ltd.), incubated for 10 min, and diluted to a concentration of 50
nmol before transfection. The transfected cells were incubated for
24 and 48 h before harvest, and the samples were assayed. All
transfections were carried out in triplicate. The primers for
miR-1301 mimics were forward, UUGCAGCUGCCUGGGAGUGACUUC and reverse,
AGUCACUCCCAGGCAGCUGCAAUU; negative control forward,
UUCUCCGAACGUGUCACGUTT and reverse, ACGUGACACGUUCGGAGAATT.
Reverse transcription-polymerase chain
reaction (RT-PCR) analysis
Total RNA was extracted from human HepG2 cell lines
using TRIzol reagent (Takara Biotechnology Co., Ltd.). Reverse
transcription was performed using 1 μg of total RNA as a template
and random hexamer as a primer. cDNA was amplified by PCR using
specific paired primers. Expression of p65, β-catenin, p53, Tg737,
Bcl-2, Bcl-xL, caspase-3, and caspase-8 in the cell line was
detected by TaqMan stem-loop RT-PCR. TaqMan probes were used to
quantify the levels of miRNA. All PCR reactions were run in
triplicate.
Western blotting
Cellular lysates were prepared as previously
described (16). Total cell
extracts were separated by SDS-polyacrylamide gel electrophoresis
(SDS-PAGE) and transferred to nitrocellulose membranes. After
saturation overnight in phosphate-buffered saline (PBS) containing
10% (w/v) skim milk with constant shaking, the nitrocellulose
membrane was cut into strips and individually incubated with 1 ml
of human serum diluted 1:500 in PBS/10% (w/v) skim milk at 37°C for
2 h. Each strip was washed three times with PBS/0.1% (v/v) Tween-20
and incubated with peroxidase-conjugated anti-human IgG (diluted
1:2000) for 2 h at room temperature. After washing, the reaction
was developed with 0.5 mg/ml diaminobenzidine in PBS. The reaction
was terminated with water. Positive and negative control sera were
included in each experiment.
Wound healing assay
For in vitro scratch assays, HepG2 cells were
seeded and grown in 6-well plates at a density of 3×104
cells/well in growth medium until they reached a confluence of
~80%. A scratch was made through each well using a sterile pipette
tip, and cells were monitored under the microscope (magnification,
×150) for 0, 12, 24 and 48 h after wounding at 37°C in 5%
CO2. Images of cells were captured at the same position
before and after incubation to document the repair process. The
experiments were repeated twice and representative pictures are
shown.
Transwell chamber migration assay
The Transwell migration assay was performed in a
24-well Transwell chamber system. The filter was washed with the
same medium and placed between the lower and the upper chambers.
HepG2 cells were trypsinized, resuspended in DMEM with 15% (v/v)
FBS and transferred to the upper chambers. The chambers were
incubated at 37°C in 5% CO2. After 48 h, the filter was
removed. Cells were stained with eosin and then with thiazide dye.
The upper surface of the filter containing non-migrating cells was
cleared using a wet cotton swab. Five fields of each well were
randomly gated and counted.
MTT assay
The MTT assay was performed in HepG2 cells that were
seeded and grown in 6-well plates at a density of 5×104
cells/well. Cells were grown in growth medium until they reached a
confluence of ~60%. miR-1301 mimics were diluted to concentrations
of 10, 20, 40, 60, 80 and 100 nmol for transfection. The
transfected cells were incubated in 96-well microculture plates at
37°C and maintained in humidified air with 5% CO2. After
24-, 48- and 72-h of transfection, MTT was added to all wells. The
optical density (OD) of each well was measured with a microplate
spectrophotometer at 490 nm. The inhibitor rate (IR) of HepG2 cell
proliferation was calculated by the equation: IR =
(1-ODtreated well/mean ODcontrol well) ×
100%.
Effect of miR-1301 transfection on HepG2
cell apoptosis
After removing the medium, HepG2 cells were washed
with PBS. The reaction product was prepared according to the
manufacturer’s instructions (Trevigen, Inc., Gaithersburg, MD, USA)
and added. Cells were viewed under a fluorescence microscope
through a dual pass filter allowing to visualize the Annexin-V-FITC
positive and the propidium iodide positive cells in the same field
according to the manufacturer’s instructions (Trevigen, Inc.).
Statistical analysis
All data are presented as the mean ± SD. Statistical
significance was calculated using the unpaired Student’s t-test.
Significance was accepted at P≤0.05. All analyses were performed
using SPSS version 13.0 (SPSS Inc., Chicago, IL, USA).
Results
miR-1301 regulates p53, Bcl-2 and Bcl-xL
gene expression
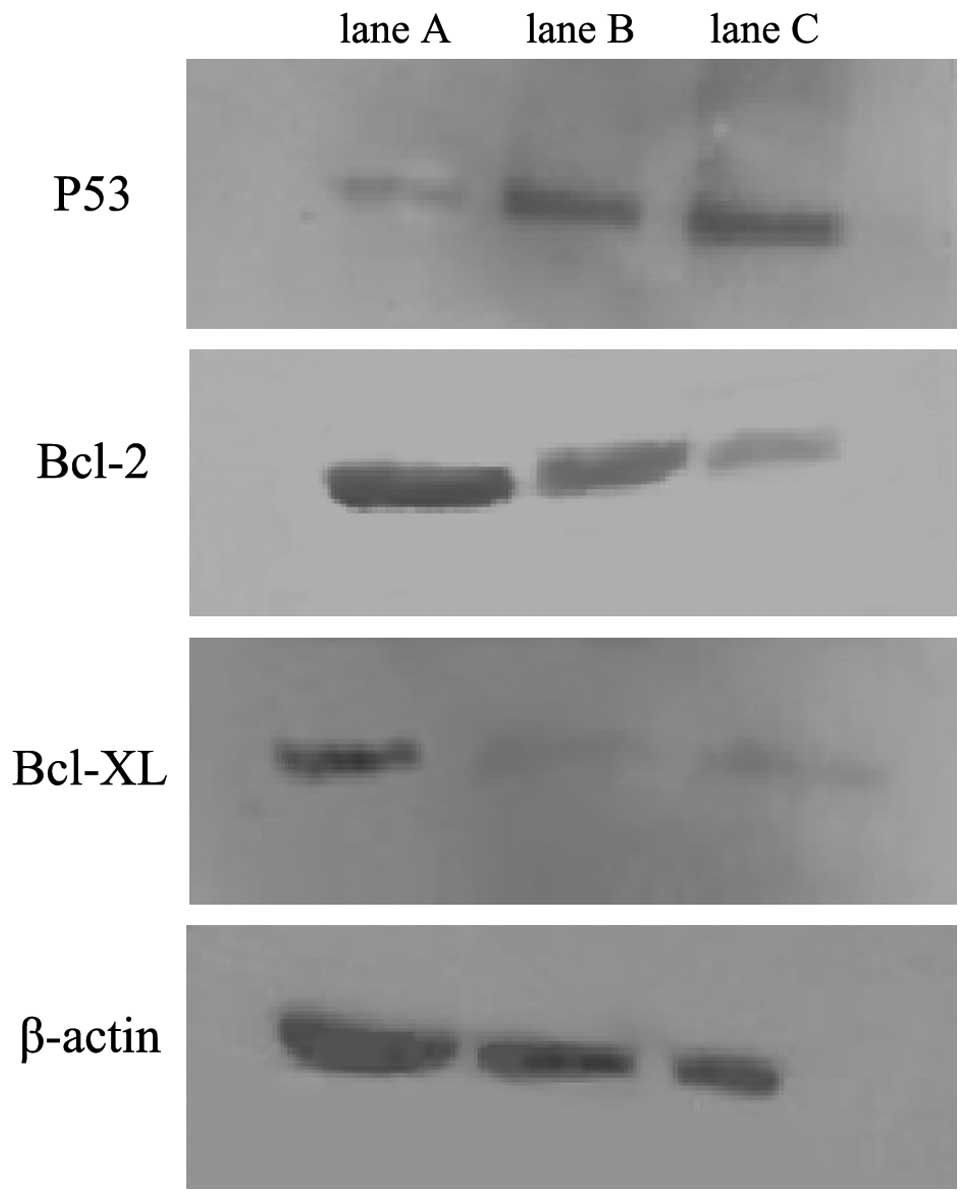
In this study, we found that p53 gene expression was
upregulated in the miR-1301 group, and that Bcl-2 gene and Bcl-xL
gene expression was downregulated when compared with the
untransfected control group (Table
I). Western blot analysis showed that levels of p53 protein
increased, while levels of the Bcl-2 and Bcl-xL proteins decreased
in the miR-1301 group (Fig. 1),
indicating that miR-1301 may regulate HepG2 cell
proliferation/apoptosis through apoptotic gene expression or tumor
suppressor gene expression. There were no significant differences
in p65, β-catenin, Tg737, caspase-3 and caspase-8 expression levels
between the miR-1301 group and the control group.
 | Table IΔCt values of expression genes between
the miR-1301 and the control groups. |
Table I
ΔCt values of expression genes between
the miR-1301 and the control groups.
| Group | miR-1301 | p65 | β-catenin | p53 | Tg737 | Bcl-2 | Bcl-xL | Caspase-3 | Caspase-8 |
|---|
| miR-1301 | −1.25±0.32a | 5.26±0.26 | 4.60±0.30 | 5.05±0.14a | 3.27±3.69 | 3.54±0.40a | −1.17±0.30a | 3.35±0.52 | 4.26±0.46 |
| Control | 3.62±0.11 | 5.10±0.20 | 3.94±0.26 | 5.95±0.05 | 0.82±0.23 | 0.98±0.11 | −3.70±0.23 | 2.75±0.25 | 4.68±0.02 |
| t-value | −21.141 | 0.829 | 3.508 | −16.043 | 1.945 | 11.227 | 20.221 | 2.651 | −1.482 |
miR-1301 inhibits the migration and
invasion ability of HepG2 cells
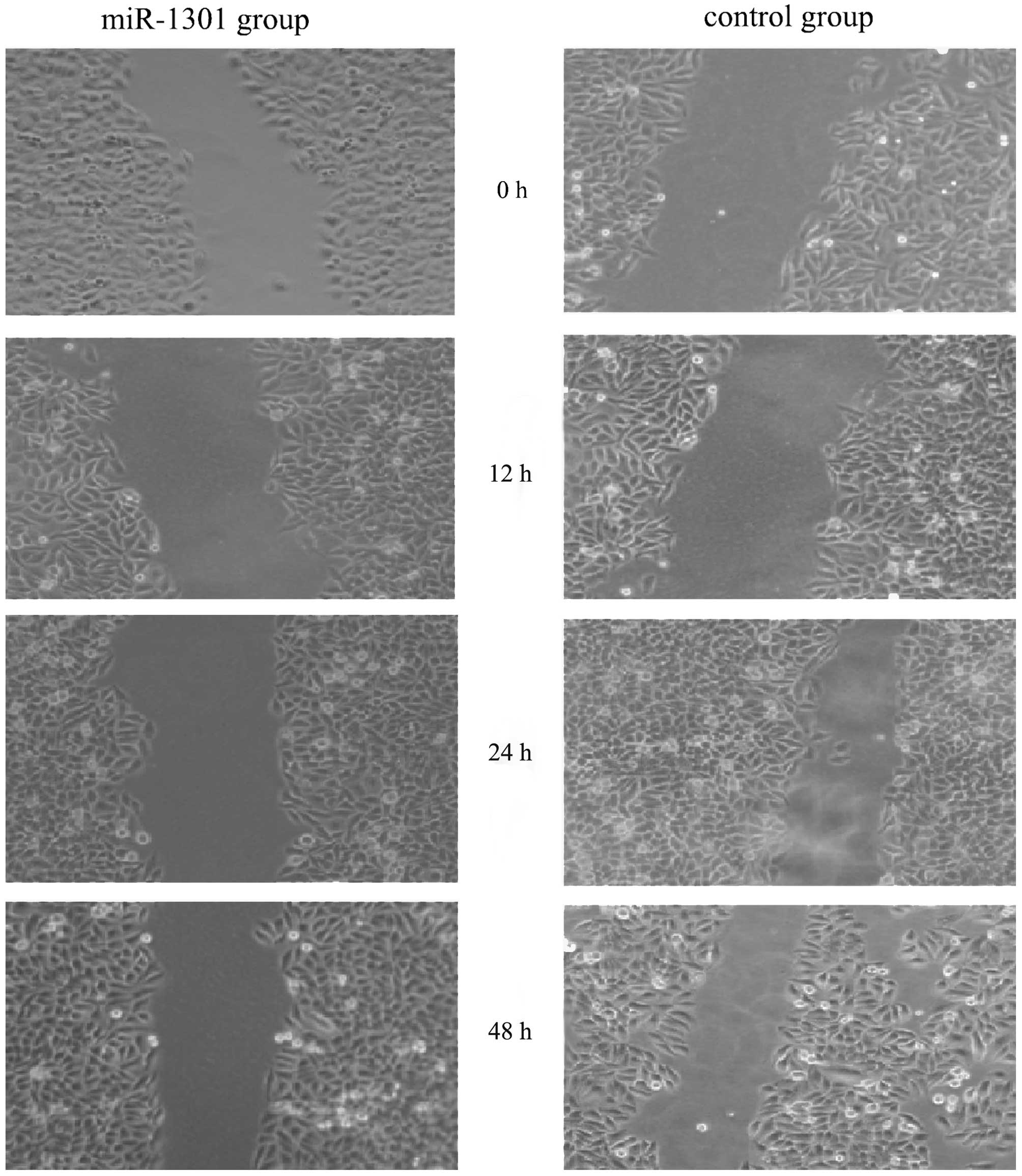
The scratch test is a useful method to investigate
wound healing ability. Our results showed that the capacity for
proliferation and migration of HepG2 cells into the wounded area
was reduced in the miR-1301 group after a 24 and 48 h repair period
(Fig. 2). There was no difference
between the miR-1301 group and the control group at 12 h. The
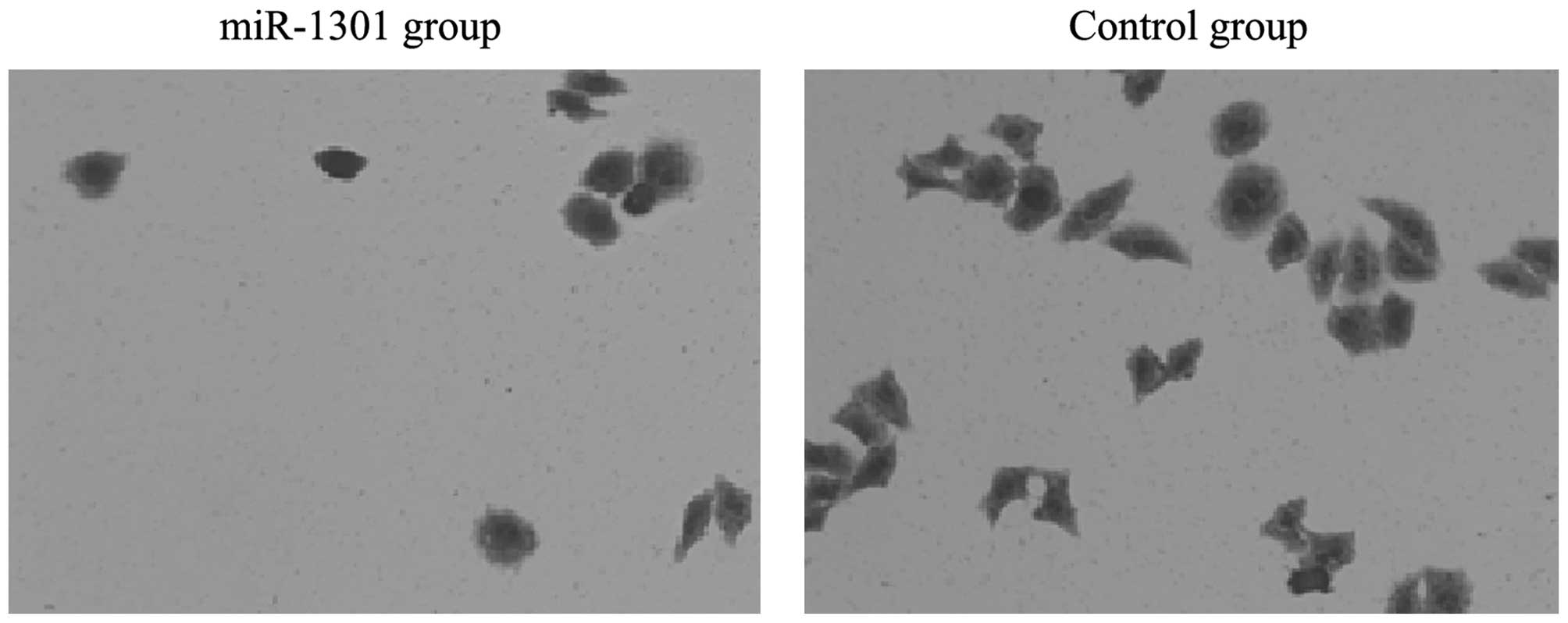
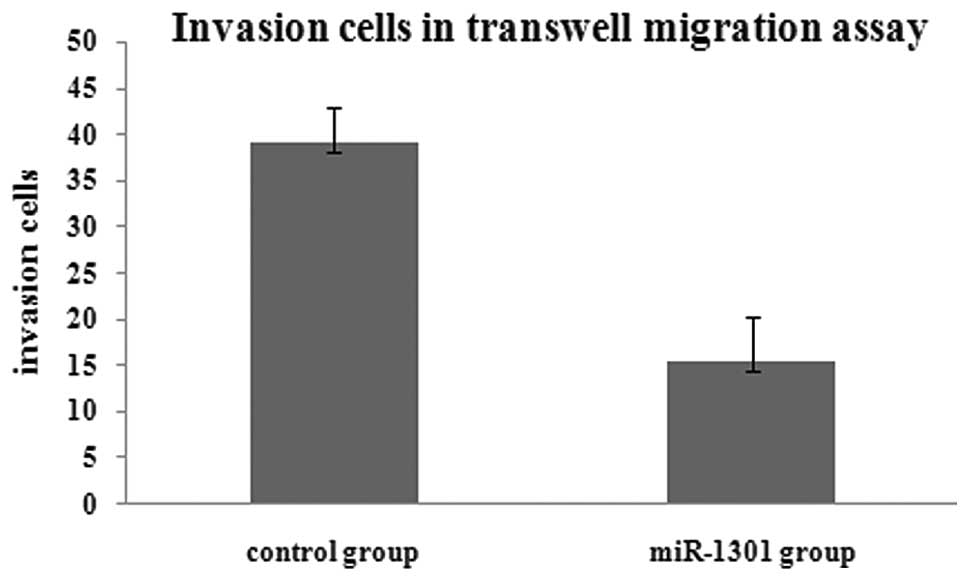
results of the transwell migration assay showed that the number of
migration cells in the miR-1301 group was less than that in the
control group. This indicated that the invasion ability of HepG2
cells might be inhibited by miR-1301 (Figs. 3 and 4).
miR-1301 inhibits cell proliferation
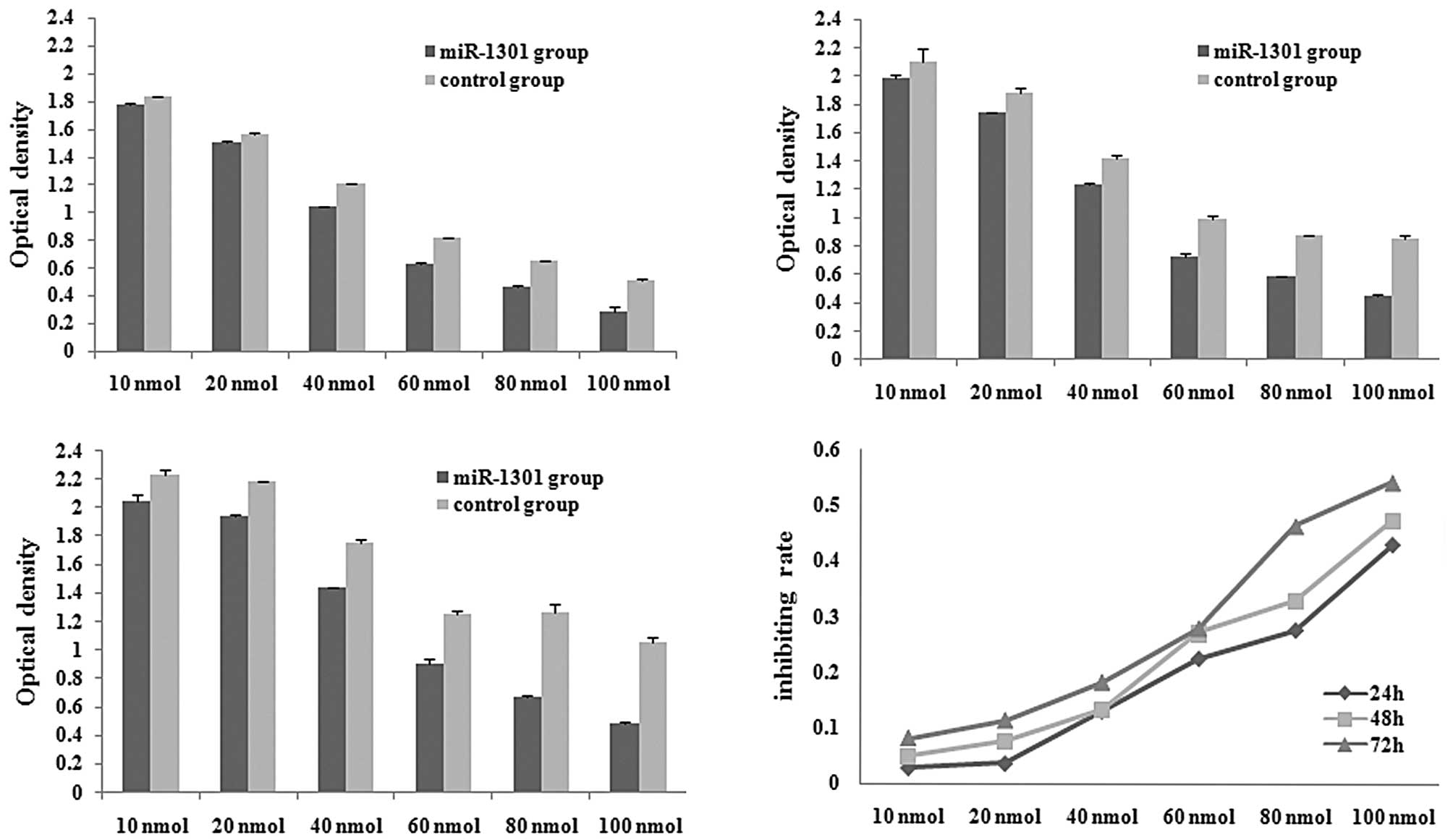
The MTT assay was performed to monitor the
proliferation rate of HepG2 cells after transfection with varying
concentrations of miR-1301 (10, 20, 40, 60, 80 and 100 nmol). The
optical density of each well was measured with a microplate
spectrophotometer at 490 nm at 24 h (Fig. 5A), 48 h (Fig. 5B), and 72 h (Fig. 5C) after transfection. Compared with
the control group, the optical density of the miR-1301 group
decreased at 24, 48 and 72 h. The proliferation rate of HepG2 cells
was dependent on the concentration and time of transfection, with
higher concentrations of miR-1301 and longer transfection times
inhibiting cell proliferation (Fig.
5D).
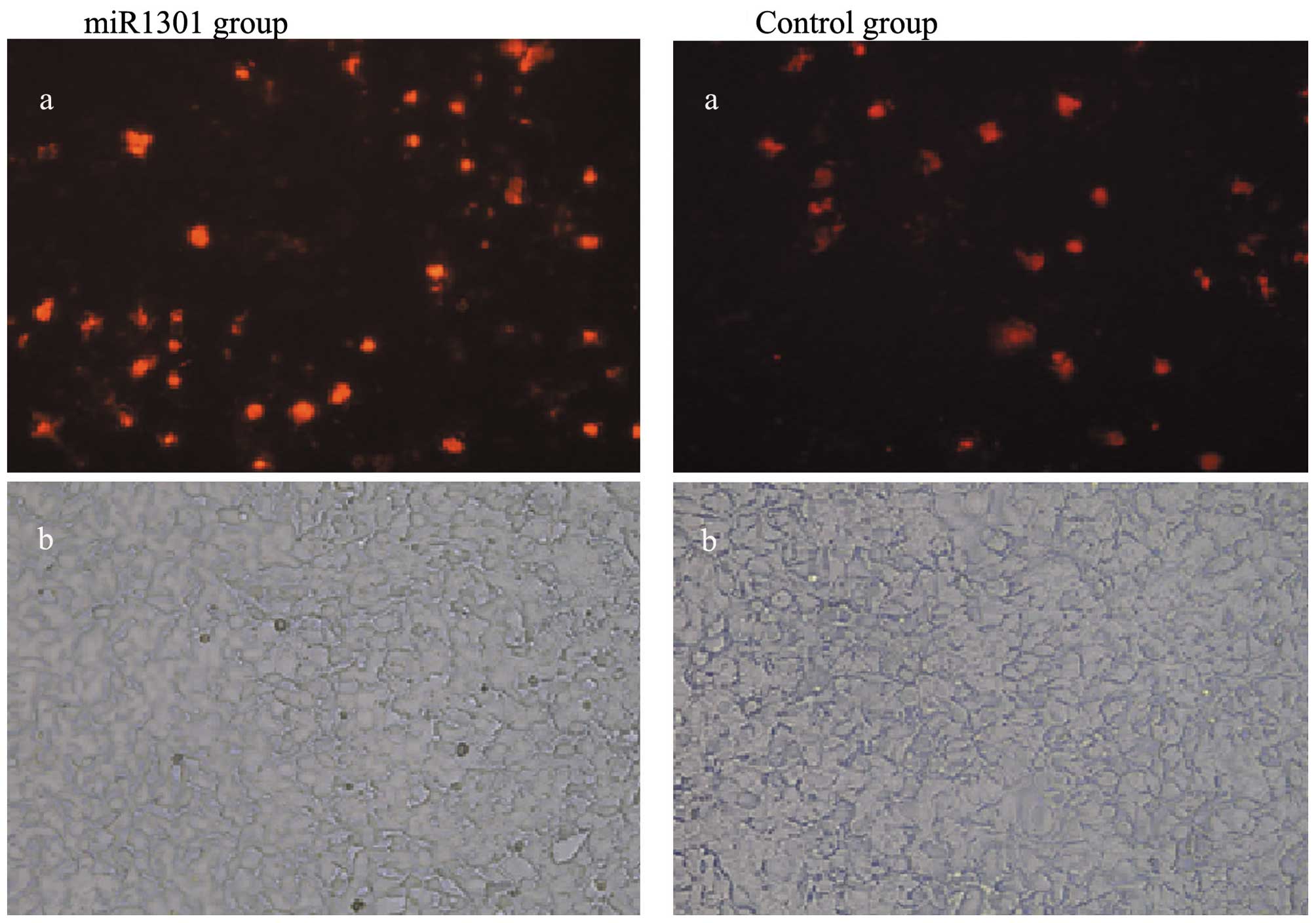
miR-1301 promotes cell apoptosis
HepG2 cell apoptosis was observed under a
fluorescence microscopy. The results showed that HepG2 cell
apoptosis increased in the miR-1301 group 48 h after transfection
with miR-1301 mimics when compared with the control group (Fig. 6).
Discussion
In this study, we found that miR-1301 expression was
low in HepG2 cells. In addition, cells transfected with miR-1301
exhibited an upregulation of p53 gene expression, a downregulation
of Bcl-2 and Bcl-xL gene expression, and increased apoptotic cell
death. Moreover, miR-1301 reduced the cell migratory and invasive
function of HepG2 cells. We therefore speculate that miR-1301 may
be a tumor inhibitor.
The function of miRNAs has attracted much attention
and research interest. Recent studies have shown that some miRNAs
may behave as oncomiRs or tumor suppressors (17–19).
These differing expressions of miRNAs could lead to different human
cancers. miRNAs that are downregulated in cancers are usually
termed tumor suppressors, while miRNAs that are upregulated in
cancers are classified as oncogenes.
In this study, miR-1301 inhibits the proliferation
of HepG2 cells. A number of miRNAs involved in regulating
proliferation and growth have been identified through linkage with
cancer phenotypes (20,21). Let-7 is known to control the timing
of proliferation and differentiation in C. elegans (22). Let-7 has also been shown to repress
Ras and c-Myc expression. Akao and colleagues (23) reported that Let-7 was greatly
reduced in lung cancer and may facilitate high levels of Ras
expression, and hence may act as a potent growth suppressor and
tumor suppressor in normal cells. miR-372 and miR-373 have been
shown to interfere with p53 function. They can also act as a
promoter of tumors, and as oncogenes in testicular germ cell tumors
(24). miR-125a has been shown to
act as a regulator of the p53 gene, and may be add to the growing
list of miRNA with oncogenic targets (25). The results of our study show that
miR-1301 can inhibit the proliferation of HepG2 cells. The p53 gene
was upregulated after transfection with miR-1301 mimics. We presume
that miR-1301 may also interfere with p53 function, hence affecting
cell proliferation. We also found that miR-1301 interfered with the
migration and invasion ability of HepG2 cells, but the reason for
this action remains unclear.
In this study, miR-1301 promoted apoptosis in HepG2
cells, and miRNAs have been shown to induce tumor apoptosis
(26). Cimmino and colleagues
(27) found that miR-15 and miR-16
could induce apoptosis by targeting Bcl-2 mRNA. Other studies
(28,29) showed that there was a correlation
between the expression of Bcl-2 and an absence of miR-15 and
miR-16, which is important in the regulation of apoptosis in
chronic lymphocytic leukemia. The Let-7 family of microRNAs
inhibits Bcl-xL expression and potentially induces apoptosis in
human hepatocellular carcinoma (30). Bcl-2 and Bcl-xL are anti-apoptotic
proteins, which oppose the progression of apoptosis. These proteins
inhibit apoptosis through the maintenance of mitochondrial membrane
integrity and by binding to pro-apoptotic Bcl family members
(31). Our study showed that
miR-1301 promotes apoptosis, and that Bcl-2 and Bcl-xL genes were
downregulated by miR-1301 at the same time. In our study, Bcl-2 and
Bcl-xL gene expression were inhibited by miR-1301. We therefore
conclude that miR-1301 may affect cell apoptosis through Bcl-2 and
Bcl-xL.
Although we observed that miR-1301 could affect the
proliferation and invasion ability and promote apoptosis in HepG2
cells, the true reason for this function remains unclear. First, it
is unclear which genes are controlled by miR-1301 and how these
genes are regulated. In addition, the mechanisms of cellular
apoptosis and invasion remain unclear. Finally, our study was only
performed in HepG2 cells, and we did not perform experiments to
find target genes of miR-1301, which we are currently
investigating.
In conclusion, our study indicates that miR-1301
reduces cellular proliferation, migration and invasion, and
promotes cell apoptosis. We speculate that miR-1301 may be a tumor
inhibitor. Further studies are needed to find target genes of
miR-1301 and mechanisms of cellular apoptosis that are regulated by
miR-1301, with the aim of preventing and treating tumors with
related miRNAs.
Acknowledgements
This study was partly supported by grants from the
National Natural Science Foundation of China (NSFC) (30940034).
References
|
1
|
Baetel DP: MicroRNAs: genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dar AA, Majid S, de Semir D, Nosrati M,
Bezrookove V and Kashani-Sabet M: miRNA-205 suppresses melanoma
cell proliferation and induces senescence via regulation of E2F1
protein. J Biol Chem. 286:16606–16614. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Osaki M, Takeshita F, Sugimoto Y, Kosaka
N, Yamamoto Y, Yoshioka Y, Kobayashi E, Yamada T, Kawai A, Inoue T,
et al: MicroRNA-143 regulates human osteosarcoma metastasis by
regulating matrix metalloprotease-13 expression. Mol Ther.
19:1123–1130. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Leucht C, Stigloher C, Wizenmann A, Klafke
R, Folchert A and Bally-Cuif L: MicroRNA-9 directs late organizer
activity of the midbrain-hindbrain boundary. Nat Neurosci.
11:641–648. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mazeh H, Mizrahi I, Halle D, Ilyayev N,
Stojadinovic A, Trink B, Mitrani-Rosenbaum S, Roistacher M, Ariel
I, Eid A, et al: Development of a microRNA-based molecular assay
for the detection of papillary thyroid carcinoma in aspiration
biopsy samples. Thyroid. 21:111–118. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Dedes KJ, Natrajan R, Lambros MB, Geyer
FC, Lopez-Garcia MA, Savage K, Jones RL and Reis-Filho JS:
Down-regulation of the miRNA master regulators Drosha and Dicer is
associated with specific subgroups of breast cancer. Eur J Cancer.
47:138–150. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang XC, Du LQ, Tian LL, Wu HL, Jiang XY,
Zhang H, Li DG, Wang YY, Wu HY, She Y, et al: Expression and
function of miRNA in postoperative radiotherapy sensitive and
resistant patients of non-small cell lung cancer. Lung Cancer.
72:92–99. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Murakami Y, Yasuda T, Saigo K, Urashima T,
Toyoda H, Okanoue T and Shimotohno K: Comprehensive analysis of
microRNA expression patterns in hepatocellular carcinoma and
non-tumorous tissues. Oncogene. 25:2537–2545. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bloomston M, Frankel WL, Petrocca F,
Volinia S, Alder H, Hagan JP, Liu CG, Bhatt D, Taccioli C and Croce
CM: MicroRNA expression patterns to differentiate pancreatic
adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA.
297:1901–1908. 2007. View Article : Google Scholar
|
|
11
|
Schetter A, Leung SY, Sohn JJ, Zanetti KA,
Bowman ED, Yanaihara N, Yuen ST, Chan TL, Kwong DL, Au GK, et al:
MicroRNA expression profiles associated with prognosis and
therapeutic outcome in colon adenocarcinoma. JAMA. 299:425–436.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ma L, Teruya-Feldstein J and Weinberg RA:
Tumour invasion and metastasis initiated by microRNA-10b in breast
cancer. Nature. 449:682–688. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhu S, Wu H, Wu F, Nie D, Sheng S and Mo
YY: MicroRNA-21 targets tumor suppressor genes in invasion and
metastasis. Cell Res. 18:350–359. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Cheng AM, Byrom MW, Shelton J and Ford LP:
Antisense inhibition of human miRNAs and indications for an
involvement of miRNA in cell growth and apoptosis. Nucleic Acids
Res. 33:1290–1297. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lima RT, Busacca S, Almeida GM, Gaudino G,
Fennell DA and Vasconcelos MH: MicroRNA regulation of core
apoptosis pathways in cancer. Eur J Cancer. 47:163–174. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lai TH, Fong YC, Fu WM, Yang RS and Tang
CH: Osteoblasts derived BMP-2 enhances the motility of prostate
cancer cells via activation of integrins. Prostate. 68:1341–1353.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Pallante P, Visone R, Ferracin M, Ferraro
A, Berlingieri MT, Troncone G, Chiappetta G, Liu CG, Santoro M,
Negrini M, et al: MicroRNA deregulation in human thyroid papillary
carcinomas. Endocr Rel Cancer. 13:497–508. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Volinia S, Calin GA, Liu CG, Ambs S,
Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M, et
al: A microRNA expression signature of human solid tumors defines
cancer gene targets. Proc Natl Acad Sci USA. 103:2257–2261. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Noguchi S, Mori T, Hoshino Y, Maruo K,
Yamada N, Kitade Y, Naoe T and Akao Y: MicroRNA-143 functions as a
tumor suppressor in human bladder cancer T24 cells. Cancer Lett.
307:211–220. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Esquela-Kerscher A and Slack FJ:
Oncomirs-microRNAs with a role in cancer. Nat Rev Cancer.
6:259–269. 2006. View
Article : Google Scholar
|
|
21
|
Liu WH, Yeh SH, Lu CC, Yu SL, Chen HY, Lin
CY, Chen DS and Chen PJ: MicroRNA-18a prevents estrogen receptor-α
expression, promoting proliferation of hepatocellular carcinoma
cells. Gastroenterology. 136:683–693. 2009.PubMed/NCBI
|
|
22
|
Pasquinelli AE, Reinhart BJ, Slack F,
Martindale MQ, Kuroda MI, Maller B, Hayward DC, Ball EE, Degnan B,
Muller P, et al: Conservation of the sequence and temporal
expression of let-7 heterochronic regulatory RNA. Nature.
408:86–89. 2000. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Akao Y, Nakagawa Y and Naoe T: let-7
microRNA functions as a potential growth suppressor in human colon
cancer cells. Biol Pharm Bull. 29:903–906. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Voorhoeve PM, le Sage C, Schrier M, Gillis
AJ, Stoop H, Nagel R, Liu YP, van Duijse J, Drost J, Griekspoor A,
et al: A genetic screen implicates miRNA-372 and miRNA-373 as
oncogenes in testicular germ cell tumors. Cell. 124:1169–1181.
2006. View Article : Google Scholar
|
|
25
|
YZ, Gao JS, Tang XL, Tucker LD,
Quesenberry P, Rigoutsos I and Ramratnam B: MicroRNA 125a and its
regulation of the p53 tumor suppressor gene. FEBS Lett.
583:3725–3730. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Babashah S and Soleimani M: The oncogenic
and tumour suppressive roles of microRNAs in cancer and apoptosis.
Eur J Cancer. 47:1127–1137. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Cimmino A, Calin GA, Fabbri M, Iorio MV,
Ferracin M, Shimizu M, Wojcik SE, Aqeilan RI, Zupo S, Dono M,
Rassenti L, Alder H, et al: MiR-15 and miR-16 induce apoptosis by
targeting BCL2. Proc Natl Acad Sci USA. 102:13944–13949. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Calin GA, Dumitru CD, Shimizu M, Bichi R,
Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K, et al:
Frequent deletions and down-regulation of micro-RNA genes mir15 and
mir16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci
USA. 99:15524–15529. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Calin GA, Liu CG, Sevignani C, Ferracin M,
Felli N, Dumitru CD, Shimizu M, Cimmino A, Zupo S, Dono M, et al:
MicroRNA profiling reveals distinct signatures in B cell chronic
lymphocytic leukemias. Proc Natl Acad Sci USA. 101:11755–11760.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Shimizu S, Takehara T, Hikita H, Kodama T,
Miyagi T, Hosui A, Tatsumi T, Ishida H, Noda T, Nagano H, et al:
The let-7 family of microRNAs inhibits Bcl-xL expression and
potentiates sorafenib-induced apoptosis in human hepatocellular
carcinoma. J Hepatol. 52:698–704. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Sun XM, Bratton SB, Butterworth M,
Macfarlane M and Cohen GM: Bcl-2 and Bcl-xl inhibit CD95- mediated
apoptosis by preventing mitochondrial release of Smac/DIABLO and
subsequent inactivation of X-linked inhibitor-of-apoptosis protein.
J Biol Chem. 277:11345–11351. 2002. View Article : Google Scholar
|