Introduction
Salivary adenoid cystic carcinoma (SACC) is a common
type of salivary gland malignancy, and accounts for 25% of
malignant tumors in the major salivary glands (1) and 50% in the minor glands (2). The neoplasm is characterized by
heterogeneous phenotypic features and persistently progressive
biological behavior. The poor long-term prognosis for patients with
adenoid cystic carcinoma is mainly due to local recurrence related
to perineural invasion (PNI) and delayed onset of distant
metastasis, particularly to the lungs (3,4). PNI,
a frequent occurrence in SACC, is difficult to be identified
clinically and this often prevents complete surgical resection
(5). Vrielinck et
al(6) reported the relationship
between PNI and poor prognosis. PNI has also been observed
frequently in other types of cancer such as melanoma, prostate and
pancreatic carcinomas as well as head and neck cancers and is
recognized as one of the most important prognostic factors
(7–11). Due to their predilection for nerves,
these cancers are known as ‘neurotropic cancers’.
Notch signaling is a pathway highly conserved
through evolution which regulates various physiological processes,
including stem cell maintenance, differentiation, proliferation and
apoptosis (12,13). In mammals, key components of the
Notch pathway include four transmembrane receptors (Notch-1,
Notch-2, Notch-3 and Notch-4) and five ligands (Dll1, Dll3, Dll4
and Jagged-1, -2) (14,15). Direct binding of a ligand from a
signaling cell to a Notch receptor on the membrane of the receiving
cell initiates two successive proteolytic cleavages by TACE
(TNF-α-converting enzyme) and the γ-secretase/presenilin complex,
which ultimately results in the release of the intracellular domain
(N-IC). N-IC then translocates into the nucleus and directly
interacts with the DNA binding protein CBF-1/Su(H)/Lag-1 (CSF) that
activates the transcription of target genes including the
hairy/enhancer-of-split (HES-1) (16).
Accumulating evidence strongly indicates that
aberrant Notch signaling has a tumor-promoting function in many
types of tumors, and Notch signaling may be a promising target for
cancer treatment. A role for Notch signaling in salivary gland
adenocarcinoma cells has been suggested which proposes that
5′-nitro-indirubinoxime (5′-NIO) induces
G0/G1 cell cycle arrest and apoptosis by the
downregulation of Notch-1 signaling (17). Notch-1 cross-talk has also been
reported in other major cell growth and apoptotic regulatory
pathways through modulating the activity of the transcription
factor, for example, nuclear factor (NF)-κB and Wnt/β-catenin
signaling (18,19). Notch signaling may contribute to
squamous cell carcinogenesis, and it is considered as a candidate
marker for squamous cell carcinomas of the head and neck (HNSCC)
(20). It was reported that the
Notch signaling pathway also contributes to the drug resistance of
cancer cells. Inhibition of Notch signaling was found to prevent
drug resistance and enhanced chemosensitivity in human myeloma,
breast cancer and HNSCC (21–23).
In SACC, a recent study suggested that Notch-4 activation
contributes to SACC metastasis (24).
In our previous microarray study, Notch-4 was found
to play a potential important role in the pathobiology of SACC
associated with PNI (25). Thus, we
tested our hypothesis on whether knockdown of Notch-4 by short
hairpin RNA (shRNA) inhibits the in vitro proliferation and
PNI in ACC-M cells.
To examine our hypothesis, we silenced Notch-4
expression in a human highly metastatic SACC cell line, ACC-M
(26), by lentiviral
vector-mediated RNA interference (RNAi) technology, and evaluated
the effect of Notch-4 on cell growth, cell cycle distribution and
cell PNI activity in ACC-M cells. Our data showed that Notch-4 RNAi
had antiproliferative activity by modulating
G0/G1 and S cell cycle regulators. The
knockdown of Notch-4 expression by lentiviral vector-mediated RNAi
reduced the PNI activity in vitro in SACC cells. These
results suggest that Notch-4 plays an important role in regulating
the in vitro growth, proliferation and PNI of ACC-M cells.
The suppression of Notch-4 may be a potential therapeutic strategy
for SACC.
Materials and methods
Cell lines and cell culture
condition
ACC-M and 293T cells were kindly provided by the
Department of Oral Biology, School of Stomatology, Fourth Military
Medical University, China. The two types of cells were cultured in
Dulbecco's modified Eagle's medium (DMEM) (Invitrogen Corp.
Carlsbad, CA, USA) supplemented with 10% of fetal bovine serum
(FBS) (Gibco, Invitrogen), 2.05 mM of L-glutamine, 100 g/ml of
penicillin and 100 μg/ml of streptomycin at 37°C in 5%
CO2.
Preparation of Notch-4 lentiviral
vectors
Lentiviral vector system (pLenOR-THM, pMDLg/pRRE,
pRSV-Rev, pMD2.G) was purchased from Innovation Biotechnology Co.,
Ltd. (Shanghai, China). Referencing siRNA design strategy (27,28),
we selected sites of the gene (NM_004557.3) cDNA sequence and
determined the specific sequence by BLAST. Four pairs of siRNA and
one negative control were designed and synthesized. As shown in
Table I, each pair contained a
unique 21-nt (shRNA1 and 2) or 19-nt (shRNA3 and 4) double-stranded
human Notch-4 sequence that is presented as an inverted
complementary repeat and separated by a loop of a 9-nt spacer. DNA
oligonucleotides (Table I)
targeting Notch-4 were synthesized and inserted into the
MIul and CIal site of the linearized lentiviral-shRNA
expression vector according to the manufacturer's instructions.
They were incorporated into a pLenOR-THM expression plasmid. The
successful ligation was confirmed by the restrictive cleavage and
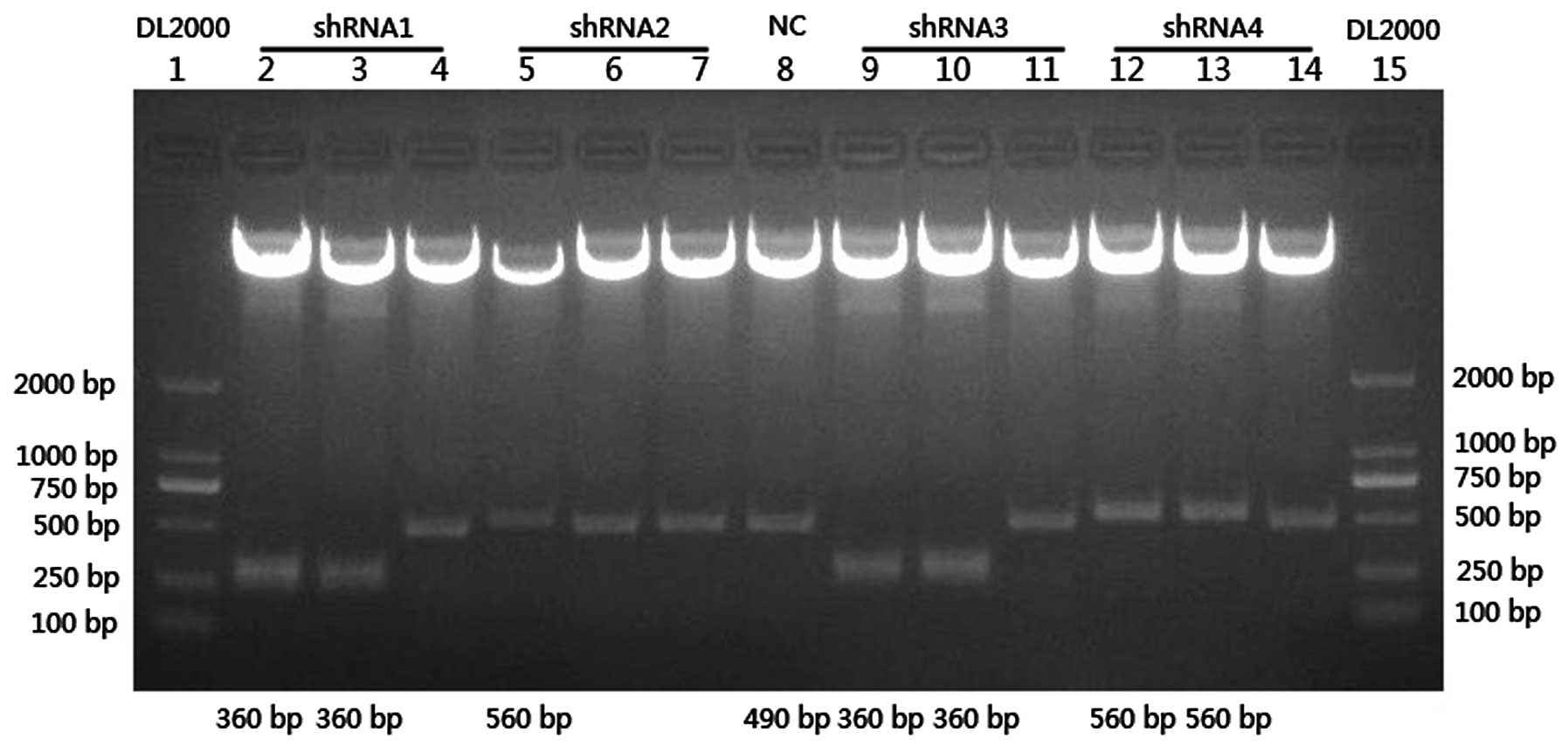
sequenced for an additional verification (Fig. 1). The recombinant vectors were named
pLenOR-Notch-4-shRNA1, 2, 3 and 4.
 | Table IOligonucleotide sequences of
Notch-4-specific shRNAs. |
Table I
Oligonucleotide sequences of
Notch-4-specific shRNAs.
| Name | Base sequence |
|---|
| shRNA1-F |
5′-CGCGTCCCCGCAGATATGTAAGGACCAGAATTCAAGAGATTCTGGTCCTTACATATCTGCTTTTTGGAAAT-3′ |
| shRNA1-R |
5′-CGATTTCCAAAAAGCAGATATGTAAGGACCAGAATCTCTTGAATTCTGGTCCTTACATATCTGCGGGGA-3′ |
| shRNA2-F |
5′-CGCGTCCCCCTGCGATAATGCGAGGAAGATTTCAAGAGAATCTTCCTCGCATTATCGCAGTTTTTGGAAAT-3′ |
| shRNA2-R |
5′-CGATTTCCAAAAACTGCGATAATGCGAGGAAGATTCTCTTGAAATCTTCCTCGCATTATCGCAGGGGGA-3′ |
| shRNA3-F |
5′-CGCGTCCCCAGATATGTAAGGACCAGAATTCAAGAGATTCTGGTCCTTACATATCTTTTTTGGAAAT-3′ |
| shRNA3-R |
5′-CGATTTCCAAAAAAGATATGTAAGGACCAGAATCTCTTGAATTCTGGTCCTTACATATCTGGGGA-3′ |
| shRNA4-F |
5′-CGCGTCCCCCGATGGACAGAACTGCTCATTCAAGAGATGAGCAGTTCTGTCCATCGTTTTTGGAAAT-3′ |
| shRNA4-R |
5′-CGATTTCCAAAAACGATGGACAGAACTGCTCATCTCTTGAATGAGCAGTTCTGTCCATCGGGGGA-3′ |
| NC-F |
5′-CGCGTCCCCTTCTCCGAACGTGTCACGTTTCAAGAGAACGTGACACGTTCGGAGAATTTTTGGAAAT-3′ |
| NC-R |
5′-CGATTTCCAAAAATTCTCCGAACGTGTCACGTTCTCTTGAACGTGACACGTTCGGAGAAGGGGA-3′ |
The recombinant vector was then mixed with virus
packaging mix, which including pMDLg/pRRE (HIV-1 gag/pol
component), pRSV-Rev (a binding site for the Rev protein which
facilitates export of the RNA from the nucleus) and pMD2.G (VSV-G
component), packed and transfected by Lipofectamine™ 2000
(Invitrogen) into 293T cells. Viral supernatant was harvested 48 h
after transfection, filtered through a 0.45-μm cellulose acetate
filter and frozen at −70°C. Virus titer was detected by a 96-well
plate dilution method and flow cytometry.
Lentiviral transfection and construction
of stable silenced cell lines
Approximately 2×105 ACC-M cells/well were
plated in 6-well plates. Twenty-four hours later, ACC-M cells were
transfected with 1×107 specific or negative control
lentiviral vectors (multiplicity of infection of 25) containing 500
μl enhancing transfection solution (Innovation Biotechnology Co.,
Ltd.) and 8 μg/ml Polybrene® (Sigma, St. Louis, MO,
USA). At 24 h post-transfection, the medium was replaced by normal
medium containing 10% FBS and antibiotics. After 96 h
post-transfection, the transfected cells were observed under a
fluorescence microscope (Leica). As the lentiviral vector contains
a GFP expression cassette, the cell transfection rate was directly
observed, which reached 90%.
Quantitative RT-PCR
Quantitative RT-PCR (qRT-PCR) for Notch-4
transcripts (shRNA1, 2, 3 and 4) in the ACC-M cell lines was
performed. ACC-M cells transfected by zero-loaded lentiviral vector
and untreated ACC-M cells were taken as the positive and the
negative control separately. Total RNA was extracted from
1×106 cells with TRIzol reagent (Invitrogen). Samples of
total RNA (1 μg) were reverse-transcribed into cDNA using a kit
according to the manufacturer's instructions (Qiagen, Valencia, CA,
USA). qRT-PCR was performed with QuantiTect SYBR Green PCR Master
Mix (Qiagen) using the Rotor-Gene RG-3000 Real-Time Thermal Cycler
(Corbett Research, Sydney, Australia) and Rotor-Gene software
version 6.0.
Each reaction mixture contained 10 μl SYBR-Green
Master Mix, 0.5 μl of each sense and antisense primer, 0.5 μl cDNA
template supplemented with water to a final volume of 20 μl. The
specific primers of Notch-4 (forward, 5′-TCAACACT
CCTGGCTCCTTCAACT-3′, and reverse, 5′-AGAGGCAC TCATTGTGATCAGCCT-3′)
were amplified as follows: 94°C for 3 min and 40 cycles at 94°C for
30 sec, followed by 61.1°C for 30 sec, 72°C for 20 sec, then ended
with 72°C for 10 min for elongation. Human 18S gene was amplified
as the internal control (forward, 5′-CGGCTACCACATCCAAGGAA-3′ and
reverse, 5′-GCTGGAATTACCGCGGCT-3′). Target genes and the 18S gene
were amplified in the same reaction. Each sample was performed in
triplicate. Comparative quantification was determined using the
2−ΔΔCt method.
Western blot analysis
Cells were washed twice with cold phosphate-buffered
saline and lysed on ice in buffer containing protease inhibitors.
Equal amounts of protein (20 μg/lane) from the cell lysates were
electrophoresed on 10% acrylamide gels. After SDS-PAGE, proteins
were transferred to a polyvinylidene difluoride membrane. The
membrane was incubated for 2 h in PBS plus 0.1% Tween 20 and 5%
non-fat skim milk to block non-specific binding. Subsequently, the
membrane was incubated for 2 h with an antibody against Notch-4
(R&D Systems, Minneapolis, MN, USA). After washing, proteins
were visualized using an ECL detection kit with the appropriate
HRP-conjugated secondary antibody (Amersham Pharmacia Biotech,
Piscataway, NJ, USA). The membranes were stripped and probed with
monoclonal antibodies for GAPDH for loading control as per standard
protocols. The experiment was repeated three times to confirm the
results.
Proliferation assay
The MTT
(3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide)
colorimetric assay was performed to assess the cell proliferation
of the transfected cells. Briefly, the cells were plated in 96-well
plates at a density of 103 cells/well. Then for 8 days,
every 24 h, a batch of cells were stained with 20 μl sterile MTT
dye (5 mg/ml; Sigma-Aldrich) at 37°C for 4 h. The culture medium
was then removed, and 150 μl of dimethyl sulphoxide (DMSO) was
added and thoroughly mixed in for 10 min. Spectrometric absorbance
at 490 nm was measured using a microplate reader. Each group
consisted of three wells.
Flow cytometric analysis
Different cell cycle phases
(G0/G1, S or G2/M phase) are
characterized by different DNA contents. Fluorescence dye propidium
iodide (PI) binds with DNA strongly at a ratio of 1:1, and hence
the DNA contents of cell cycle phases are reflected by varying PI
fluorescent intensities. Stable transfected ACC-M cells,
1×106, were harvested by trypsinization and fixed in 70%
ice-cold (4°C) ethanol for 2 h. Cell pellets were resuspended in 1
mg/ml RNase solution (Sigma-Aldrich) for 30 min at 37°C and
subsequently in 0.1 mg/ml PI solution (Sigma-Aldrich) at 4°C for 1
h in the dark. Cell cycle analysis was performed on a flow
cytometer (Beckman Coulter, Inc., Fullerton, CA, USA).
In vitro perineural invasion assay
The inhibitory effect of RNAi on the PNI ability of
ACC-M cells in vitro was demonstrated in modified Boyden
chambers. Transwell invasion chambers containing polycarbonate
filters (8-μm) (Millipore Corp., Billerica, MA, USA) were coated on
the upper surface with Matrigel basement membrane (BD Biosciences,
San Diego, CA, USA). Cells (1×105) were suspended in
DMEM supplemented with 1% fetal bovine serum and added to the upper
chamber. The lower chamber contained 600 μl supernatant of 24
h-cultured RSC96 cells (a rat Schwann cell line, purchased from the
Cell Bank for Type Culture Collection, Chinese Academy of Sciences)
as a chemoattractant to simulate the perineural surrounding
environment (29,30). Cells were incubated for 12 h at 37°
in 5% CO2 incubator. At the end of the incubation, the
cells on the upper surface of the filter were completely removed by
wiping with a cotton swab. The filters were then fixed in methanol
and stained with hematoxylin and eosin. Cells that had invaded the
Matrigel and had reached the lower surface of the filter were
counted under a light microscope at a magnification of ×400. We
chose five fields of vision and counted the numbers of the invaded
cells on the lower surface of the filter. The assay was performed
in triplicate.
Statistical analysis
Results are expressed as means ± standard deviation
(SD). Statistical analysis was performed using SPSS 17.0
statistical software (SPSS Inc., Chicago, IL, USA). Data were
tested for statistical significance using analysis of variance
(ANOVA). Normally distributed, continuous variables were compared
using one-way ANOVA. When ANOVA produced a significant difference
between groups, multiple comparisons of group means were performed
using the Bonferroni procedure with a type I error adjustment. All
P-values were two-sided, and significance was defined as
P<0.05.
Results
Lentiviral vector-mediated RNAi of
Notch-4 causes effective and specific downregulation of Notch-4
expression
The knockdown efficiencies of different
Notch-4-specific shRNAs in ACC-M cells were first evaluated using
qRT-PCR. Relative Notch-4 mRNA levels in individual stable
transfectants were normalized against mRNA levels of an internal
control gene, human 18S, performed in the same run. As shown in
Fig. 2A, cells transfected with
pLenOR-Notch-4-shRNA2, 3 and 4 showed a significantly reduced
transcription of Notch-4 mRNA when compared with that of the
negative control ACC-M cells and the positive control vector ACC-M
Mock, respectively (P<0.01), but there was no significant mRNA
transcription reduction in cells transfected with
pLenOR-Notch-4-shRNA1. The cells transfected with
pLenOR-Notch-4-shRNA2 showed the most significant inhibition of
Notch-4 mRNA levels.
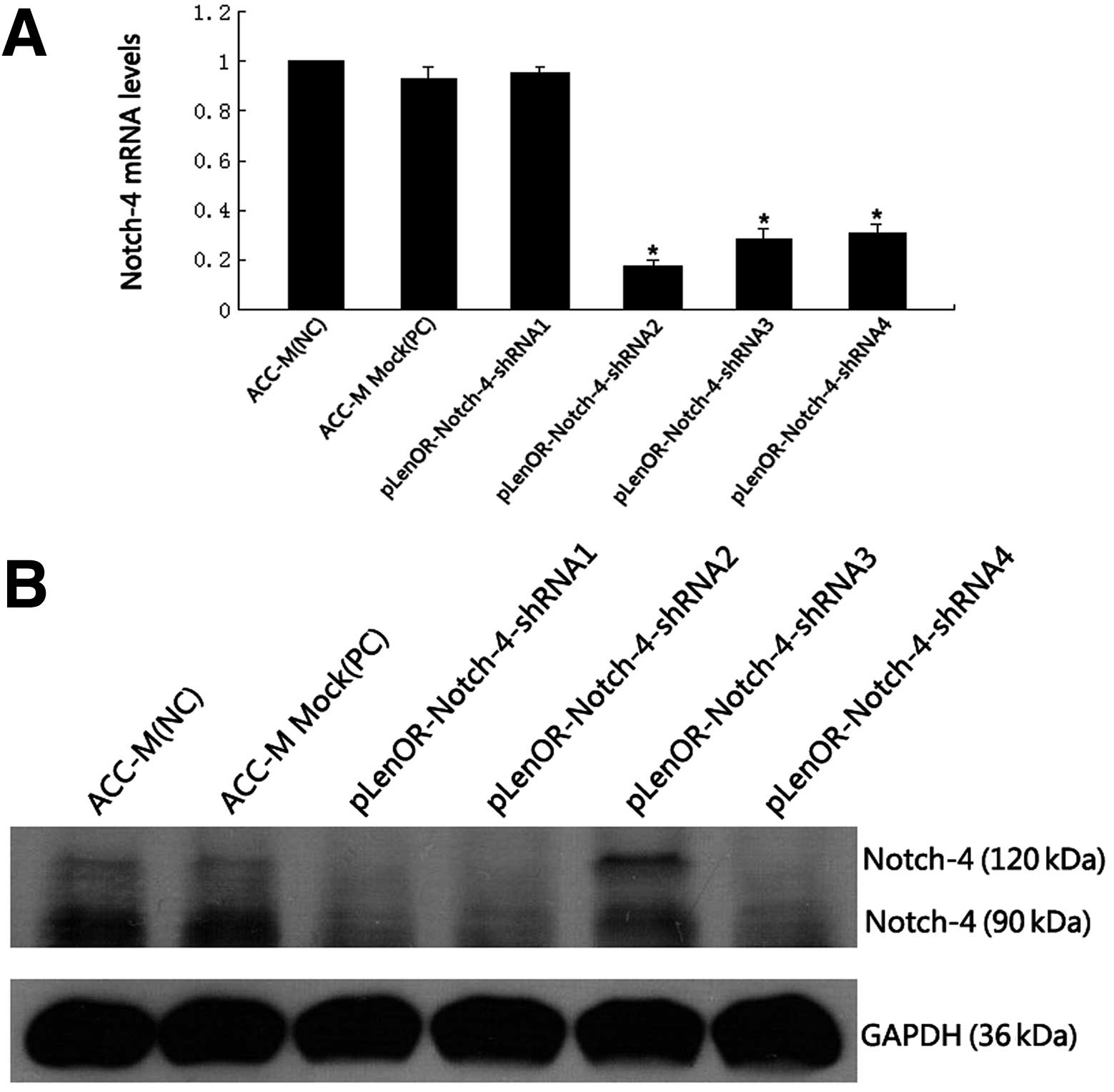 | Figure 2Confirmation of Notch-4 expression in
different clones by qRT-PCR and western blot analysis in ACC-M
cells. (A) Notch-4 mRNA levels were determined by qRT-PCR. Relative
fold induction for the Notch-4 mRNA (means ± SD) in mock and
Notch-4 siRNA-transfected cells is presented relative to the
expression in parental ACC-M cells. Cells transfected with
pLenOR-Notch-4-shRNA2, 3, 4 showed a significantly reduced
transcription of Notch-4 mRNA when compared with that of the
negative control ACC-M cells and the positive control vector ACC-M
Mock, respectively, while there was no significant reduction in
mRNA transcription in cells transfected with pLenOR-Notch-4-shRNA1
(*P<0.01 compared with ACC-M). (B) Western blot
analysis for Notch-4 protein expression in the indicated cell
lines. GAPDH was used as a loading control. The Notch-4 protein is
a ~210 kDa heterodimer. The Notch-4 protein was split by the
lysate, so therefore the western blot analysis detected the
intracellular and extracellular domain of Notch-4 protein in the
cells. Western blot analysis revealed a decreased expression of
Notch-4 protein in the ACC-M cells transfected with
pLenOR-Notch-4-shRNA1, 2, 4, while cells transduced by
pLenOR-Notch-4-shRNA3 showed no notable reduced expression of
Notch-4 protein compared with that of the negative control ACC-M
cells and the positive control vector ACC-M Mock. These results
revealed that the most effective vector was pLenOR-Notch-4-shRNA2.
ACC-M, high metastatic potential control used as a negative
control; ACC-M Mock, mock transfection control used as the positive
control) pLenOR-Notch-4-shRNA1, 2, 3 and 4 represent the four
different clones, respectively. |
The Notch-4 protein is a heterodimer ~210 kDa, and
the Notch-4 protein was split by the lysate, thus our western blot
analysis detected the intracellular and extracellular domain of
Notch-4 protein in the cells. In addition, western blot analysis
(Fig. 2B) showed reduced expression
of Notch-4 protein in the ACC-M cells transfected with
pLenOR-Notch-4-shRNA1, 2, 4, while cells transduced by
pLenOR-Notch-4-shRNA3 showed no notable decreased expression of
Notch-4 protein compared with that of the negative control ACC-M
cells and the positive control vector ACC-M Mock. The above results
demonstrated that the expression of Notch-4 was specifically and
effectively downregulated by Notch-4 RNAi, allowing its application
for the subsequent experiment. For the sake of convenience, we
chose the most effective vector (pLenOR-Notch-4-shRNA2)-transfected
cells, positive and negative control groups to study the cell
biological behavior.
Gene silencing of Notch-4 reduces cell
proliferation in vitro
The proliferative activity of tumor cells is
important in the invasion/metastasis of tumors. To examine whether
the knockdown of Notch-4 expression has any effect on cell growth,
an MTT cell proliferation assay was performed. Under the same cell
culture conditions, the proliferative activity of the
pLenOR-Notch-4-shRNA2-transfected cells, negative and positive
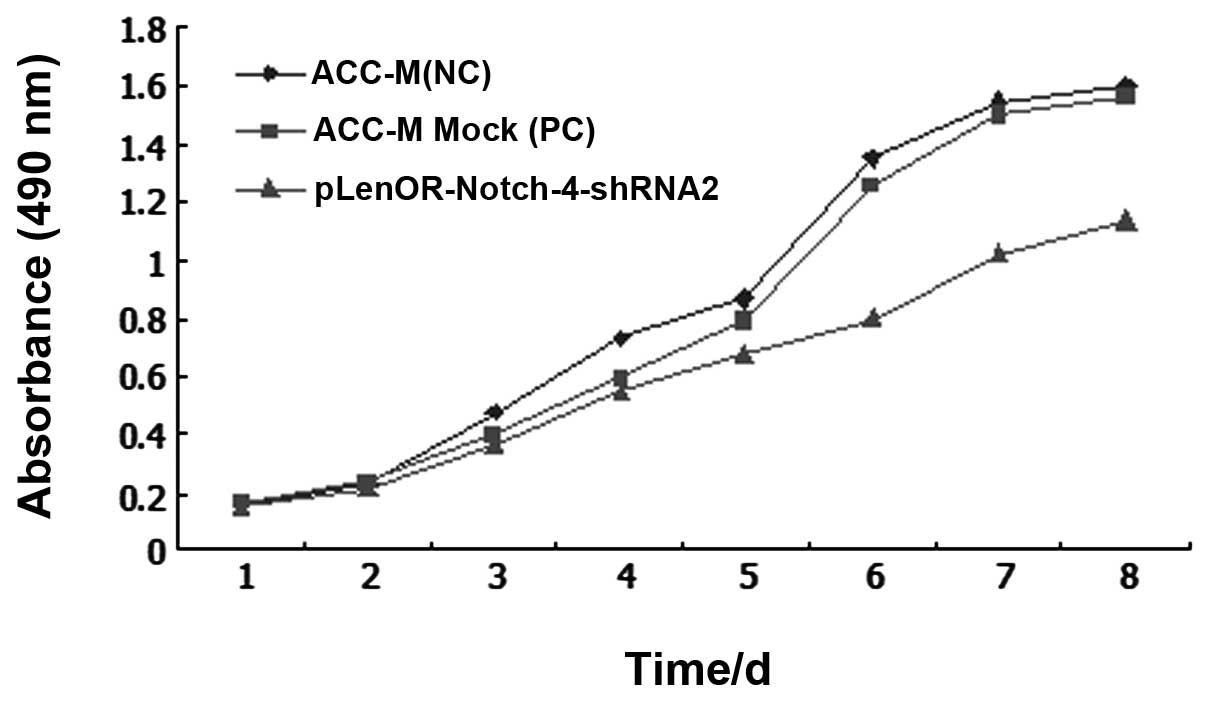
control cells was almost similar for the first 24 h. With
time-lapse, the cells with Notch-4 gene silencing grew more slowly
when compared with the control cells. Among the three groups,
Notch-4 RNAi (pLenOR-Notch-4-shRNA2) cells showed decreased cell
proliferation, when compared to the negative control (ACC-M) and
mock-transfected (ACC-M Mock) cells, supporting the role of Notch-4
in the cell growth of ACC-M cells (P<0.01) (Fig. 3).
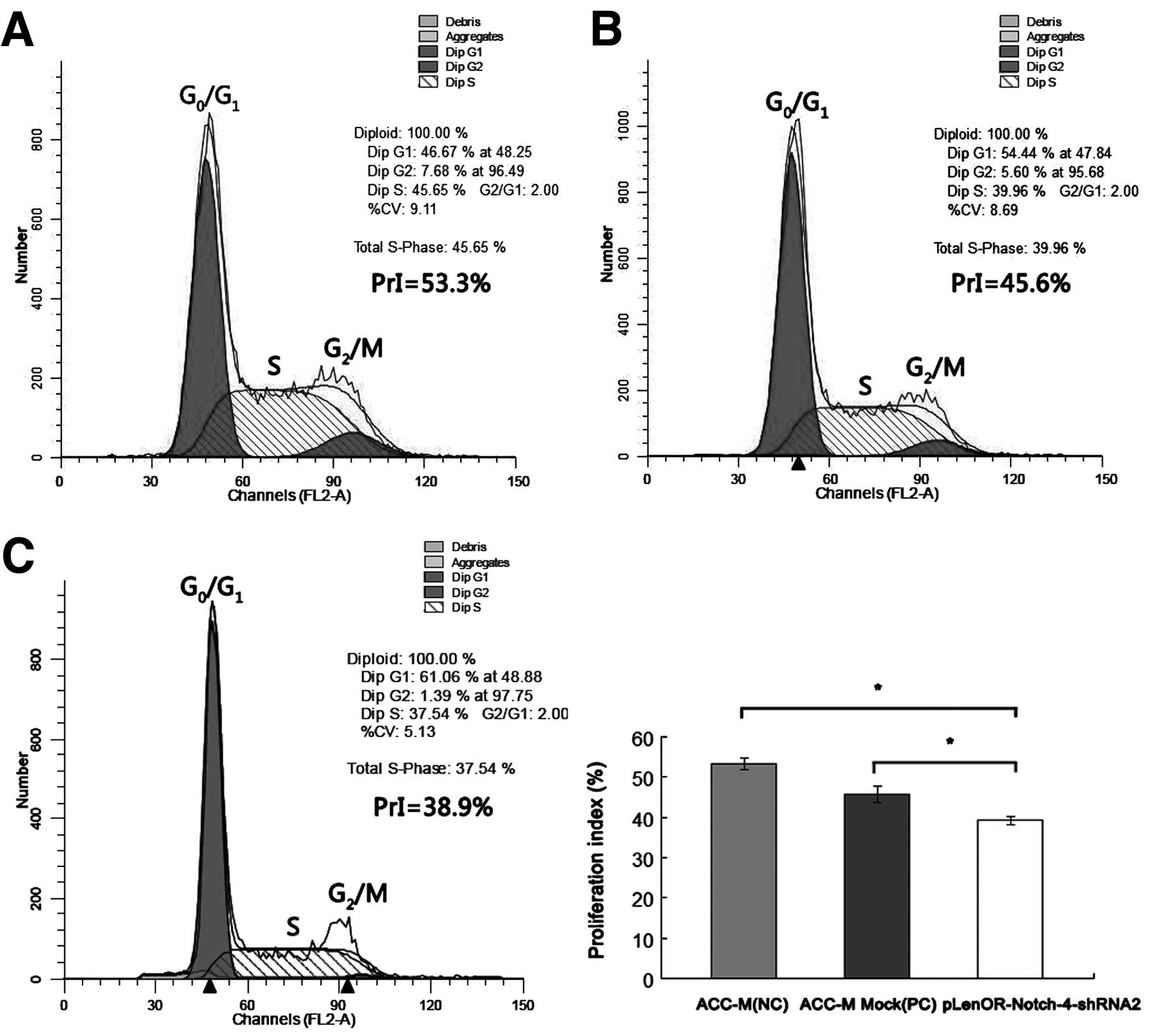
We next used flow cytometry (FCM) to study the
effect of Notch-4-specific shRNA on the cell cycle distribution in
ACC-M cells. The negative control group (ACC-M) and positive
control group (ACC-M Mock) resulted in a cell cycle distribution of
~53 and 46% of the cells in the S and G2/M phases. In
the pLenOR-Notch-4-shRNA2 group, the proliferation index value
(PrI) of cycling cells (combined total number of cells in the S and
G2/M phases) was decreased to ~39%, with a concomitant
increase in the number of cells in the G0/G1
phase (Table II) (Fig. 4). Significant decreases in PrI were
found in the pLenOR-Notch-4-shRNA2-transfected cells compared with
negative and positive control cells (P<0.01). There was no
significant difference in PrI between that of the negative and
positive control transfectant cells (P>0.05). These findings
indicate that knockdown of Notch-4 expression may inhibit
proliferation of ACC-M cells by modulating
G0/G1 and S cell cycle regulators.
 | Table IIFCM analysis of the cell cycle of
ACC-M cells after Notch-4-specific inhibition. |
Table II
FCM analysis of the cell cycle of
ACC-M cells after Notch-4-specific inhibition.
| Cell group |
G0/G1 (%) | S (%) | G2/M
(%) | Proliferation index
(%) |
|---|
| ACC-M (NC) | 46.7±1.45 | 45.7±0.96 | 7.6±0.50 | 53.3±1.45 |
| ACC-M Mock
(PC) | 54.2±2.06 | 40.2±1.27 | 5.5±0.80 | 45.8±2.06 |
|
pLenOR-Notch-4-shRNA2 | 60.8±1.12 | 37.8±1.16 | 1.5±0.10 | 39.2±1.12a |
Gene silencing of Notch-4 inhibits in
vitro perineural invasion ability of ACC-M cells
PNI activity of the effective Notch-4-specific shRNA
transfectant (pLenOR-Notch-4-shRNA2) was assayed in vitro by
modified Boyden chambers. Cells transfected with
pLenOR-Notch-4-shRNA2 showed much lower PNI activities compared
with the negative and the positive controls (P<0.01). There was
no significant difference in in vitro PNI ability between
the negative and positive control transfectant cells (P>0.05)
(Fig. 5). These data confirm that
the knockdown of Notch-4 expression inhibits in vitro PNI
activity of ACC-M cells.
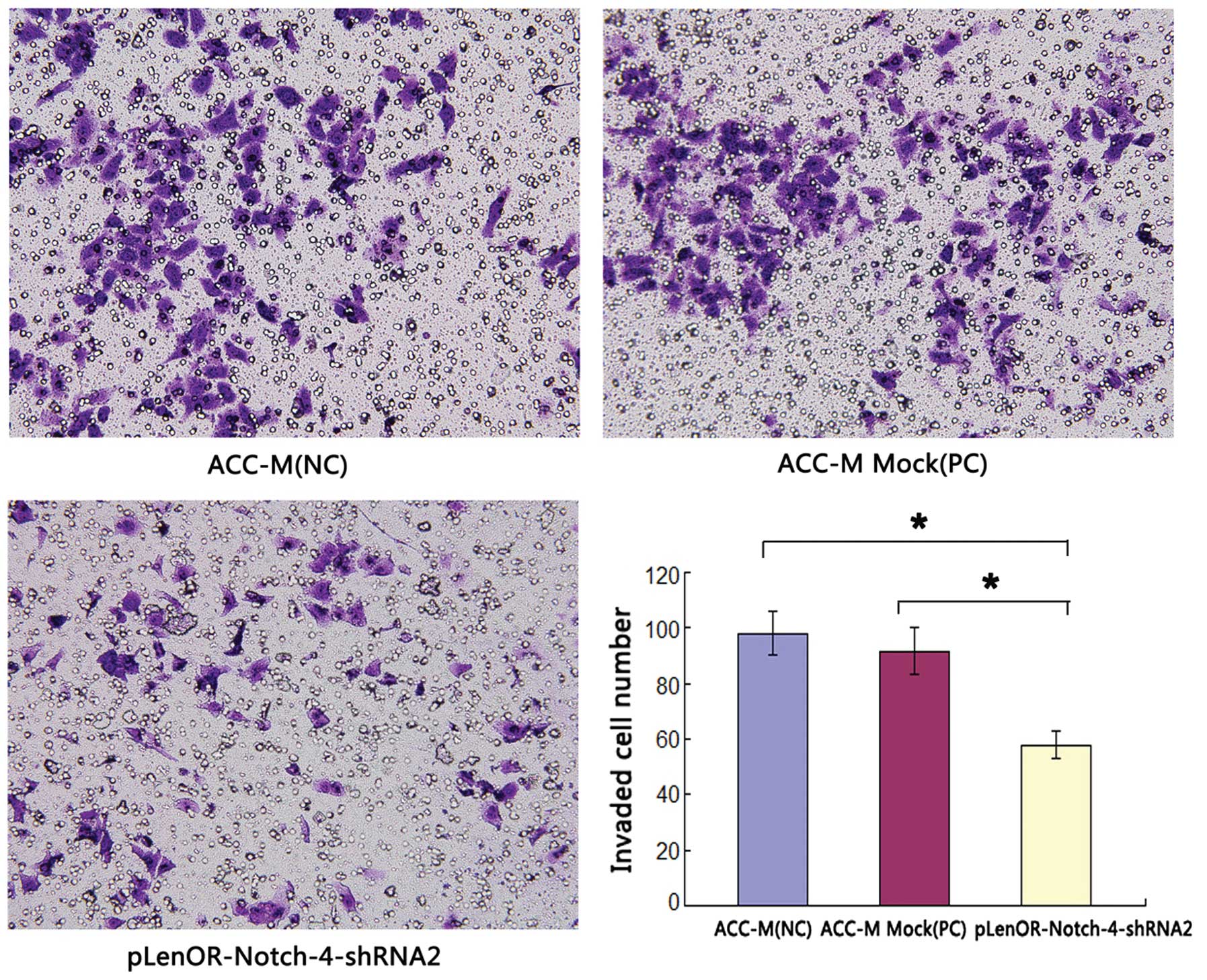 | Figure 5Effects of Notch-4-specific shRNA on
Matrigel perineural invasion of ACC-M cells. Matrigel PNI was
evaluated using modified Boyden chambers as described in Materials
and methods. The numbers of cells migrated were evaluated in three
fields for each experimental group and averaged. The average
invaded cell number of the groups ACC-M (NC), ACC-M Mock (PC) and
pLenOR-Notch-4-shRNA2 were 98.0±7.98, 91.5±8.46 and 57.8±4.95,
respectively. Statistical analysis was performed with one-way
ANOVA. P1,3<0.01, P2,3<0.01,
P1,2>0.05. 1, ACC-M (NC); 2, ACC-M Mock (PC); 3,
pLenOR-Notch-4-shRNA2 (magnification, ×400). |
Discussion
The gene encoding the Notch receptor was discovered
almost 90 years ago, and gained its name because partial loss of
Notch function resulted in notches in the wing margins of
Drosophila(31). It only
became apparent some years later that the Notch signaling pathway
has been conserved throughout evolution. The Notch signaling
pathway plays a pivotal role in several cell functions, such as
cell fate decisions, cell proliferation, differentiation and cell
death during development and postnatal life in species as diverse
as Drosophila, worms and vertebrates (32–36).
In the mammalian system, there are four Notch receptors (Notch-1,
Notch-2, Notch-3 and Notch-4) and five known ligands (Dll1, Dll3,
Dll4 and Jagged-1, -2) (14,15).
Notch is a cellular fate determinant and can induce cell
proliferation and/or differentiation, depending on the cellular
environment (37). A recent study
showed that targeting Notch-1 and Notch-4 may provide a new
therapeutic strategy for triple-negative and possibly other breast
cancer subtypes (38). In SACC,
Notch-4 may play a key role in SACC metastasis, and inhibition of
Notch-4 gene expression may have potential therapeutic application
in treating metastatic patients (24).
Perineural invasion (PNI), is a typical biological
behavior of SACC, which may prevent a complete surgical resection
(5). PNI is associated with poor
prognosis in SACC patients (6). In
our previous study, we established the gene expression profile of
SACC associated with PNI by combining the use of laser capture
microdissection (LCM) and cDNA microarray. In the profile, Notch-4
was notably overexpressed in the PNI cell group, and this was
verified by qRT-PCR (25). Thus, we
hypothesized that inhibition of Notch-4 gene expression may reduce
in vitro proliferation and PNI in ACC-M cells.
To understand the biological function of Notch-4 in
SACC, we examined the effects of a decreased expression of Notch-4
in a human highly metastatic SACC cell line, ACC-M, using a
lentiviral-mediated RNAi system. RNAi uses the phenomenon by which
double-strand RNA induces potent and specific inhibition of
eukaryotic gene expression through the degradation of complementary
messenger RNA (mRNA) and is functionally similar to the processes
of post-transcriptional gene silencing (39). In the past few years, siRNA and
shRNA have been widely used to silence the expression of many
target genes, and both methods have had great achievement, but the
silencing effect lasts for less than 2 weeks. New systems based on
lentiviral vectors have provided new solutions to achieving stable
shRNA-mediated knockdown (40,41).
In the present study, we used a lentiviral-mediated
RNAi method to obtain effective knockdown of the Notch-4 gene in
ACC-M cells and constructed stable silencing clones.
Here, we designed four shRNAs targeted at the
Notch-4 gene and successfully transfected them into ACC-M cells.
Among the four designed shRNAs, the cells transfected with
pLenOR-Notch-4-shRNA2 showed the most significant inhibitory effect
as determined by qRT-PCR and western blot analysis. The results
indicated that lentiviral-mediated RNAi of Notch-4 silenced the
expression of Notch-4 effectively and specifically in ACC-M
cells.
We next examined the consequence of ACC-M cells
transfected with Notch-4-specific shRNA. The proliferation of the
ACC-M cells in which the Notch-4 gene was knocked down was
inhibited compared with that of the positive or negative control.
In the FCM study, the Notch-4-knockdown ACC-M cells showed an
arrest in G0/G1-to-S transition, suggesting
growth inhibition of these cells (Table II). Therefore, Notch-4 may be a
positive regulator of cell growth to promote a mitogenic signal,
which then enhances cell proliferation of ACC-M cells.
In the present study, the silencing of Notch-4 in
ACC-M cells inhibited the cell PNI activity in vitro. This
result was consistent with the results of the MTT assay and FCM
analysis which revealed that silencing of Notch-4 by
lentiviral-mediated RNAi inhibited the cell proliferation in
vitro by modulating G0/G1 and S cell
cycle regulators. The finding may also be associated with the
metastasic ability of Notch-4 in SACC and breast cancer (24,38).
SACC is a common subtype of salivary gland
malignancy, and it has an important biological behavior for PNI. It
is urgent to develop new therapeutic strategies for SACC. In this
report, the knockdown of Notch-4 expression by lentiviral-mediated
RNAi successfully inhibited the malignant behaviors of ACC-M cells,
particularly PNI ability in vitro, implicating that Notch-4
may be a new candidate target gene for the treatment of SACC.
Acknowledgements
This study was supported by grants from the National
Natural Science Foundation of China (Project no. 81102051), the
Natural Science Foundation of Jiangsu Province (Project no.
BK2011659) and the Youth Foundation of Nanjing Jinling Hospital
(Project no. 2010Q009).
Abbreviations:
|
SACC
|
salivary adenoid cystic carcinoma
|
|
PNI
|
perineural invasion
|
|
siRNA
|
small interfering RNA
|
|
shRNA
|
short hairpin RNA
|
|
TACE
|
TNF-α-converting enzyme
|
|
NF-κB
|
nuclear factor-κB
|
|
FCM
|
flow cytometry
|
References
|
1
|
Renehan A, Gleave EN, Hancock BD, Smith P
and McGurk M: Long-term follow-up of over 1000 patients with
salivary gland tumours treated in a single centre. Br J Surg.
83:1750–1754. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Anderson JN Jr, Beenken SW, Crowe R, Soong
SJ, Peters G, Maddox WA and Urist MM: Prognostic factors in minor
salivary gland cancer. Head Neck. 17:480–486. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Van der Wal JE, Becking AG, Snow GB and
van der Waal I: Distant metastases of adenoid cystic carcinoma of
the salivary glands and the value of diagnostic examinations during
follow-up. Head Neck. 24:779–783. 2002.PubMed/NCBI
|
|
4
|
Ramer N, Wu H, Sabo E, Ramer Y, Emanuel P,
Orta L and Burstein DE: Prognostic value of quantitative p63
immuno- staining in adenoid cystic carcinoma of salivary gland
assessed by computerized image analysis. Cancer. 116:77–83.
2010.PubMed/NCBI
|
|
5
|
Van der Wal JE, Snow GB and van der Waal
I: Intraoral adenoid cystic carcinoma. The presence of perineural
spread in relation to site, size, local extension, and metastatic
spread in 22 cases. Cancer. 66:2031–2033. 1990.PubMed/NCBI
|
|
6
|
Vrielinck L, Ostyn F, van Damme B, van den
Boqaert W and Fossion E: The significance of perineural spread in
adenoid cystic carcinoma of the major and minor salivary glands.
Int J Oral Maxillofac Surg. 17:190–193. 1988. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Newlin HE, Morris CG, Amdur RJ and
Mendenhall WM: Neurotropic melanoma of the head and neck with
clinical perineural invasion. Am J Clin Oncol. 28:399–402. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ayala GE, Dai H, Ittmann M, Li R, Powell
M, Frolov A, Wheeler TM, Thompson TC and Rowley D: Growth and
survival mechanisms associated with perineural invasion in prostate
cancer. Cancer Res. 64:6082–6090. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Pour PM, Bell RH and Batra SK: Neural
invasion in the staging of pancreatic cancer. Pancreas. 26:322–325.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Faqan JJ, Collins B, Barnes L, D'Amico F,
Myers EN and Johnson JT: Perineural invasion in squamous cell
carcinoma of the head and neck. Arch Otolaryngol Head Neck Surg.
124:637–640. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Rahima B, Shingaki S, Nagata M and Saito
C: Prognostic significance of perineural invasion in oral and
oropharyngeal carcinoma. Oral Surg Oral Med Oral Pathol Radiol
Endod. 97:423–431. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Go MJ, Eastman DS and Artavanis-Tsakonas
S: Cell proliferation control by Notch signaling in
Drosophila development. Development. 125:2031–2040.
1998.PubMed/NCBI
|
|
13
|
Shelly LL, Fuchs C and Miele L: Notch-1
inhibits apoptosis in murine erythroleukemia cells and is necessary
for differentiation induced by hybrid polar compounds. J Cell
Biochem. 73:164–175. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bigas A, Martin DI and Milner LA: Notch1
and Notch2 inhibit myeloid differentiation in response to different
cytokines. Mol Cell Biol. 18:2324–2333. 1998.PubMed/NCBI
|
|
15
|
Mumm JS and Kopan R: Notch signaling: from
the outside in. Dev Biol. 228:151–165. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Stockhausen MT, Sjölund J and Axelson H:
Regulation of the Notch target gene Hes-1 by TGFalpha induced
Ras/MAPK signaling in human neuroblastoma cells. Exp Cell Res.
310:218–228. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yoon JH, Kim SA, Kwon SM, Park JH, Park
HS, Kim YC, Yoon JH and Ahn SG: 5′-Nitro-indirubinoxime induces G1
cell cycle arrest and apoptosis in salivary gland adenocarcinoma
cells through the inhibition of Notch-1 signaling. Biochim Biophys
Acta. 1800:352–358. 2010.
|
|
18
|
Yao J, Duan L, Fan M, Yuan J and Wu X:
Notch1 induces cell cycle arrest and apoptosis in human cervical
cancer cells: involvement of nuclear factor kappa B inhibition. Int
J Gynecol Cancer. 17:502–510. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Duan L, Yao J, Wu X and Fan M: Growth
suppression induced by Notch1 activation involves Wnt-β-catenin
down-regulation in human tongue carcinoma cells. Biol Cell.
98:479–490. 2006.PubMed/NCBI
|
|
20
|
Leethanakul C, Patel V, Gillespie J,
Pallente M, Ensley JF, Koontongkaew S, Liotta LA, Emmert-Buck M and
Gutkind JS: Distinct pattern of expression of differentiation and
growth-related genes in squamous cell carcinomas of the head and
neck revealed by the use of laser capture microdissection and cDNA
arrays. Oncogene. 19:3220–3224. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Nefedova Y, Sullivan DM, Bolick SC, Dalton
WS and Gabrilovich DI: Inhibition of Notch signaling induces
apoptosis of myeloma cells and enhances sensitivity to
chemotherapy. Blood. 111:2220–2229. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zang S, Chen F, Dai J, Guo D, Tse W, Qu X,
Ma D and Ji C: RNAi-mediated knockdown of Notch-1 leads to cell
growth inhibition and enhanced chemosensitivity in human breast
cancer. Oncol Rep. 23:893–899. 2010.PubMed/NCBI
|
|
23
|
Gu F, Ma Y, Zhang Z, Zhao J, Kobayashi H,
Zhang L and Fu L: Expression of Stat3 and Notch1 is associated with
cisplatin resistance in head and neck squamous cell carcinoma.
Oncol Rep. 23:671–676. 2010.PubMed/NCBI
|
|
24
|
Ding LC, She L, Zheng DL, Huang QL, Wang
JF, Zheng FF and Lu YG: Notch-4 contributes to the metastasis of
salivary adenoid cystic carcinoma. Oncol Rep. 24:363–368.
2010.PubMed/NCBI
|
|
25
|
Chen W, Zhang HL, Shao XJ, Jiang YG, Zhao
XG, Gao X, Li JH, Yang J, Zhang YF, Liu BL and Sun MY: Gene
expression profile of salivary adenoid cystic carcinoma associated
with perineural invasion. Tohoku J Exp Med. 212:319–334. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Guan XF, Qiu WL, He RG and Zhou XJ:
Selection of adenoid cystic carcinoma cell clone highly metastatic
to the lung: an experimental study. Int J Oral Maxillofac Surg.
26:116–119. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tuschl T: Expanding small RNA
interference. Nat Biotechnol. 20:446–448. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Reynolds A, Leake D, Boese Q, Scaringe S,
Marshall WS and Khvorova A: Rational siRNA design for RNA
interference. Nat Biotechnol. 22:326–330. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wang L, Sun M, Jiang Y, Yang L, Lei D, Lu
C, Zhao Y, Zhang P, Yang Y and Li J: Nerve growth factor and
tyrosine kinase A in human salivary adenoid cystic carcinoma:
expression patterns and effects on in vitro invasive behavior. J
Oral Maxillofac Surg. 64:636–641. 2006. View Article : Google Scholar
|
|
30
|
Chen W, Zhang HL, Jiang YG, Li JH, Liu BL
and Sun MY: Inhibition of CD146 gene expression via RNA
interference reduces in vitro perineural invasion on ACC-M cell. J
Oral Pathol Med. 38:198–205. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Mohr OL: Character changes caused by
mutation of an entire region of a chromosome in Drosophila.
Genetics. 4:275–282. 1919.PubMed/NCBI
|
|
32
|
Miele L and Osborne B: Arbiter of
differentiation and death: Notch signaling meets apoptosis. J Cell
Physiol. 181:393–409. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Cheng P and Gabrilovich D: Notch signaling
in differentiation and function of dendritic cells. Immunol Res.
41:1–14. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Baker NE: Notch signaling in the nervous
system. Pieces still missing from the puzzle. Bioessays.
22:264–273. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Artavanis-Tsakonas S, Rand MD and Lake RJ:
Notch signaling: cell fate control and signal integration in
development. Science. 284:770–776. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Milner LA and Bigas A: Notch as a mediator
of cell fate determination in hematopoiesis: evidence and
speculation. Blood. 93:2431–2448. 1999.PubMed/NCBI
|
|
37
|
Kopan R and Ilagan MX: The canonical Notch
signaling pathway: unfolding the activation mechanism. Cell.
137:216–233. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Clementz AG, Rogowski A, Pandya K, Miele L
and Osipo C: Notch-1 and Notch-4 are novel gene targets of PEA3 in
breast cancer: novel therapeutic implications. Breast Cancer Res.
13:R632011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zamore PD, Tuschl T, Sharp PA and Bartel
DP: RNAi: double-stranded RNA directs the ATP-dependent cleavage of
mRNA at 21 to 23 nucleotide intervals. Cell. 101:25–33. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Yamamoto T, Miyoshi H, Yamamoto N,
Yamamoto N, Inoue J and Tsunetsugu-Yokota Y: Lentivirus vectors
expressing short hairpin RNAs against the U3-overlapping region of
HIV nef inhibit HIV replication and infectivity in primary
macrophages. Blood. 108:3305–3312. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Desclaux M, Teigell M, Amar L, Vogel R,
Gimenez Y, Ribotta M, Privat A and Mallet J: A novel and efficient
gene transfer strategy reduces glial reactivity and improves
neuronal survival and axonal growth in vitro. PLoS One.
4:e62272009. View Article : Google Scholar : PubMed/NCBI
|