Introduction
Oral cancer, which is occurring at an increasing
frequency worldwide, is the sixth most common malignancy in humans.
This year alone, it is estimated that ~600,000 new cases will
arise, with only 40–50% of patients with oral squamous cell
carcinoma (OSCC) likely to survive for 5 years. Despite combined
surgery and radiation therapy, long-term survival of patients with
oral cancer has shown no improvement over the past few decades
(1,2). This is primarily due to the poor
clinical prognosis of patients with lymph node metastasis. A deeper
understanding of the molecular basis of the highly malignant
properties of oral cancer combined with patient stratification is
therefore needed.
microRNAs (miRNAs) are non-coding small RNAs (~22
nucleotides) that regulate post-transcriptional gene expression by
interfering with the translation of target mRNAs. A single miRNA
can regulate the expression of several genes, while more than
one-third of all protein-coding genes are thought to be under
translational control of miRNAs. miRNAs are involved in a variety
of cellular processes, including the regulation of cellular
differentiation, proliferation and apoptosis (3,4). In
addition, aberrant expression of miRNAs is known to induce various
human malignancies, clearly classified by their miRNA profiles
(5,6). However, little is known about the
miRNA expression patterns or function in oral cancer (7,8).
Specific overexpression or underexpression of miRNAs
has been correlated with particular types of tumors (5,9,10). It
has also been suggested that miRNA overexpression could result in
downregulation of tumor-suppressor genes, while underexpression may
lead to oncogene upregulation (9).
Most importantly, it has been suggested that miRNA expression
signatures could be used to predict the outcome in several tumor
types, including lung cancer and chronic lymphocytic leukemia, as
well as to predict the response to chemotherapy. miRNAs may
therefore prove to be novel therapeutic targets for a wide range of
diseases, including cancer (11).
In the present study, we examined the expression
profiles of miRNAs in OSCC, revealing differential expression
between normal and cancer tissues. These findings may further
facilitate the potential therapeutic and diagnostic use of miRNAs
in OSCC.
Materials and methods
Clinical samples and cell lines
Twenty-nine OSCC samples and 7 oral mucosal samples
were obtained from oral cancer patients who underwent surgery at
Showa University from April 2004 to November 2008. Written informed
consent was obtained from all patients prior to sampling, and
samples were maintained at −80ºC until use. OSCC samples were
microscopically examined for determination of cancer cell content
by two independent pathologists; they were dissected to enrich
cancer cells when necessary. For molecular analysis, 29 samples
containing ≥50% cancer cells were used. Normal oral mucosa was
extracted at a sufficient margin from the cancer, and the absence
of a muscle layer and cancer cells was confirmed microscopically.
The present study was certified by the Ethics Committee of the
Showa University.
RNA preparation
miRNA was prepared with the Recover All Total
Nucleic Acid Isolation kit (Applied Biosystems, Darmstadt, Germany)
according to the kit protocol. RNA yield and the A260/280 ratio
were monitored with a NanoDrop ND-100 spectrometer (NanoDrop
Technologies, Wilmington, DE, USA).
Reverse transcription
miRNAs were reverse transcribed with the TaqMan
miRNA Reverse Transcription kit (Applied Biosystems), using 60 ng
total RNA and pools of miRNA specific stem-loop primers (Megaplex
RT Primers Pool A and B; Applied Biosystems). After reverse
transcription, the cDNA was preamplified with Megaplex PreAmp
Primers (Pool A and B) according to the recommendations of the
supplier (Applied Biosystems).
TaqMan low density arrays
Differential expression of 768 miRNAs was determined
using Taqman microRNA Array v2.0 (Applied Biosystems), with an A
and B card for each. The array plate also included the RNU48
transcript as a normalization signal. Expression levels of each
mature miRNA were evaluated using the comparative threshold cycle
(Ct) method, with normalization to RNU48 (2−ΔΔCt). The
fold-change in each miRNA was calculated from the difference in
expression levels between tumor tissues and normal tissues.
miRNA target prediction
TargetScan (http://www.targetscan.org/) was used to analyze
potential target genes of the deregulated miRNAs.
Results
Differential expression of miRNAs between
normal oral mucosa and oral cancer tissues
To define the role of miRNAs in OSCC, we
investigated the expression profiles of 29 OSCC tumors and 7 normal
oral mucosal samples. The 29 OSCC tissue samples were of various
stages and histological grades (Table
I). TaqMan-based microRNA assays of 768 different miRNA targets
were performed, revealing a total of 177 detectable miRNAs. Volcano
plots show differential expression between normal and cancer
tissues (Fig. 1). We subsequently
calculated the degree of differential expression, revealing that
miRNAs >4-fold upregulated or downregulated could be considered
as candidate miRNAs. miR-31*, miR-31, miR-135b, miR-193a-5p,
miR-103, miR-224, miR-93, miR-200c, miR-183, miR203, miR-21 and
miR-223 were shown to be upregulated, while miR-133a, miR-376c,
miR-411, miR-30a-3p, miR-489, miR-139-5p, miR-483-5p, miR-30e-3p,
miR-409-3p, let-7c and miR-486-5p were shown to be downregulated in
OSCC (Tables II and III).
 | Table IDemographic and clinical features of
the OSCC patients. |
Table I
Demographic and clinical features of
the OSCC patients.
| Characteristic | n (%) |
|---|
| Gender |
| Male | 20 (69) |
| Female | 9 (31) |
| Smoking habit |
| Positive | 8 (28) |
| Negative | 21 (72) |
| Drinking habit |
| Positive | 9 (31) |
| Negative | 20 (69) |
| WHO
classification |
| Well | 10 (34) |
| Moderate | 8 (28) |
| Poor | 11 (38) |
| YK grade |
| 1 | 1 (4) |
| 2 | 3 (10) |
| 3 | 13 (45) |
| 4 | 12 (41) |
| Local recurrence |
| Positive | 10 (34) |
| Negative | 19 (66) |
| Age (years) |
| >65 | 21 (72) |
| ≤65 | 8 (28) |
| Stage |
| I | 6 (21) |
| II | 9 (31) |
| III | 3 (10) |
| IV | 11 (38) |
| T status |
| I | 6 (21) |
| II | 14 (48) |
| III | 2 (7) |
| IV | 7 (24) |
| N status |
| N0 | 15 (51) |
| N1 | 4 (13) |
| N2 | 10 (36) |
 | Table IIDifferentially expressed miRNAs
showing increased expression in OSCC compared to normal mucosa. |
Table II
Differentially expressed miRNAs
showing increased expression in OSCC compared to normal mucosa.
| microRNA | Fold-change
(Ct-Ca/Ct-N) | FDR | Chromosome
location | Putative target |
|---|
| hsa-miR-31* | 0.60 | 0.008 | 9p21.3 | RSBN1, ARHGEF2,
IDE, NR5A2, SH2D1A |
| hsa-miR-31 | 0.29 | 0.020 | 9p21.3 | RSBN1, ARHGEF2,
IDE, NR5A2, SH2D1A |
| hsa-miR-135b | 0.68 | 0.004 | 1q32.1 | ANGPT2, GK5, NR3C2,
GULP1, LOC221710 |
|
hsa-miR-193a-5p | 0.77 | 0.013 | 17q11.2 | HSPB6, ZNF385C,
ITSN1, OLIG3, USO1 |
| hsa-miR-103 | 0.76 | 0.007 | 5q34 | DICER1, TMEM16C,
NF1, FOXP1, HRB |
| hsa-miR-224 | 0.71 | 0.029 | Xq28 | ZDHHC20, AFF3,
U2SURP, C8orf44, TTC3 |
| hsa-miR-93 | 0.64 | 0.003 | 7q22.1 | FGD4, PKD2, MAP3K2,
ZNFX1, PDCD1LG |
| hsa-miR-200c | 0.27 | 0.004 | 12p13.31 | ZEB1, FAM122C,
ZEB2, LRP1B, WIPF1 |
| hsa-miR-183 | 0.82 | 0.001 | 7q32.2 | ABAT, AKAP12, PIGX,
PTPN4, REV1 |
| hsa-miR-203 | 0.22 | 0.040 | 14q32.33 | ZNF281, CAMTA1,
B3GNT5, LIFR, ABCE1 |
| hsa-miR-21 | 0.54 | 0.014 | 17q23.1 | ANF367, GPR64,
YOD1, PHF14, PLEKHA1 |
| hsa-miR-223 | 0.27 | 0.003 | Xq12 | FBXW7, SP3, PAX6,
C13orf31, PURB |
 | Table IIIDifferentially expressed miRNAs
showing decreased expression in OSCC compared to normal mucosa. |
Table III
Differentially expressed miRNAs
showing decreased expression in OSCC compared to normal mucosa.
| microRNA | Fold-change
(Ct-Ca/Ct-N) | FDR | Chromosome
location | Putative
target |
|---|
| hsa-miR-133a | 3.13 | 0.019 | 18q.11.2 | SYT2, LHFP, CCBL2,
BRUNOL4, TTPAL |
| hsa-miR-376c | 1.55 | <1e-03 | 14q32.31 | ARFGEF1, PAPSS2,
GABRG2, SYF2, ARFGEF2 |
| hsa-miR-411 | 1.52 | <1e-03 | 14q32.31 | ELFN1, SLC4A7,
C16orf52, C21orf91, SPRY4 |
| hsa-miR-30a-3p | 1.75 | <1e-03 | 6q13 | NUFIP2, ZNF85,
RUNDC2B, FIGN, POU4F1 |
| hsa-miR-489 | 1.36 | 0.010 | 7q21.3 | ETNK1, PARM1,
CWC25, NRIP1, PNISR |
| hsa-miR-139-5p | 1.65 | <1e-03 | 11q13.4 | TMF1, USP6NL, TBX1,
SCAPER, NDRG2 |
| hsa-miR-483-5p | 1.33 | 0.010 | 11p15.5 | SELO, MPZ, SRSF4,
SLC12A5, RNF165 |
| hsa-miR-30e-3p | 1.54 | 0.006 | 1p34.2 | NUFIP2, ZNF85,
RUNDC2B, POU4F1, DSN |
| hsa-miR-409-3p | 1.42 | 0.006 | 14q32.31 | MRPL35, LRRN4CL,
POMP, MTF2, TMEM65 |
| hsa-let-7c | 1.29 | 0.025 | 21q21.1 | C14orf28, FIGNL2,
HMGA2, LIN28B, TRIM71 |
| hsa-miR-486-5p | 1.49 | 0.025 | 8p11.21 | FOXO1, GPX8, PTEN,
TRAPPC6B, TWF1 |
Differential expression of miRNAs between
non-metastatic (-) and metastatic (+) OSCC
As cervical lymph node metastasis is directly
correlated with oral cancer prognosis, we subsequently aimed to
determine whether miRNA expression was associated with outcome in
OSCC patients. Volcano plots show the differential expression of
miRNAs in non-metastatic (-) and metastatic (+) OSCC (Fig. 2). We subsequently calculated the
level of significance of the differential expression between these
groups. miR-489, miR-483-5p and miR-1291 were shown to be
downregulated in OSCC (Table IV).
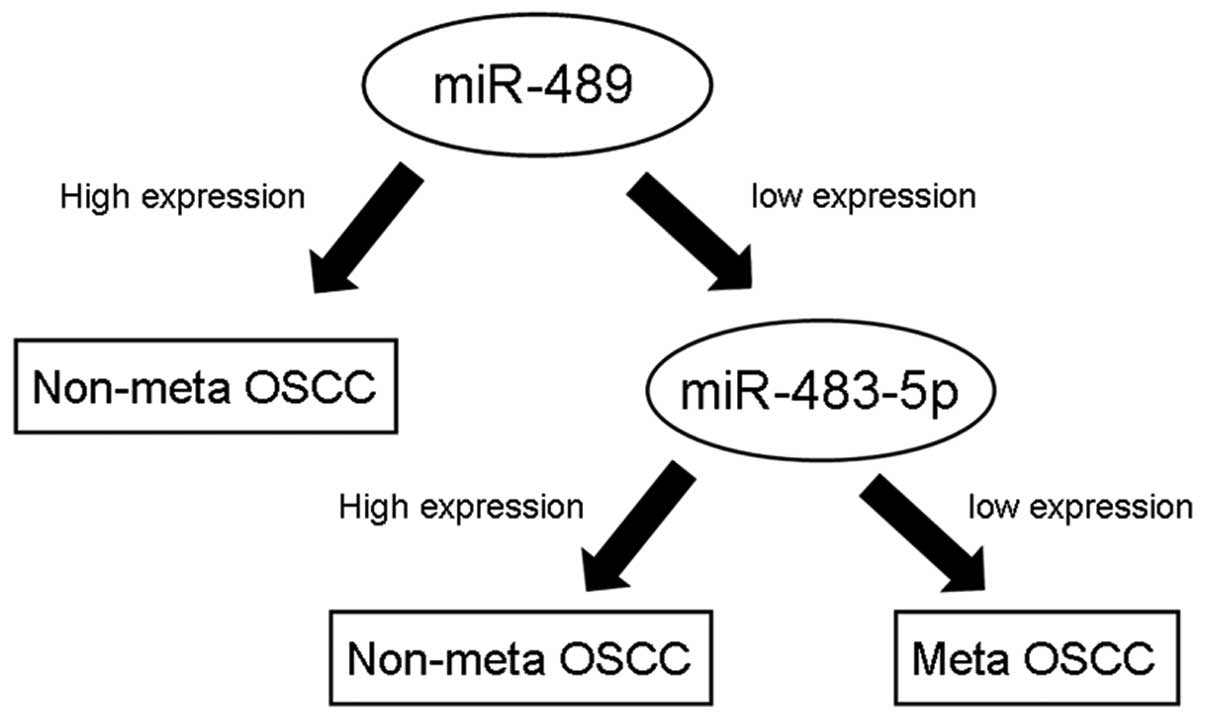
In addition, a decision tree model was used to classify the two
groups (Fig. 3), doing so with 82%
accuracy.
 | Table IVDifferentially expressed miRNAs
showing decreased expression in non-metastatic and metastatic
OSCC. |
Table IV
Differentially expressed miRNAs
showing decreased expression in non-metastatic and metastatic
OSCC.
| microRNA | Fold-change
(Ct-Ca/Ct-N) | FDR | Chromosome
location | Putative
target |
|---|
| hsa-miR-489 | 1.27 | 0.004 | 7q21.3 | ETNK1, ALS2CR13,
HRH4, LONRF2, SFRS7 |
| hsa-miR-1291 | 1.18 | 0.017 | 12q13.11 | AQP1, ARID3B,
MECP2, MAP3K9, PGM5 |
| hsa-miR-483-5p | 1.22 | 0.005 | 11p15.5 | RAB3IP, ZBTB26,
PGAM1, C5orf42, RBAK |
Discussion
miRNAs are a new class of non-coding small RNAs that
regulate cell proliferation and various cellular functions by
interfering with the translation of the target mRNAs (3,4,12,13).
Recent studies have demonstrated that altered expression of miRNAs
induces various human malignancies (14,15).
However, little is known about the role of miRNAs in OSCC. In this
study, we investigated the miRNA expression profiles of normal oral
mucosa and OSCC tissues. miRNA profiling in B cell lymphoma
(16), prostate (17), and colon cancer (18) was previously performed; however, to
the best of our knowledge, this is the first comprehensive miRNA
profiling study of oral cancer through exhaustive analysis.
miRNA analysis was performed using TaqMan microRNA
Array v2.0, which consists of 768 candidate miRNA sites. A total of
177 miRNAs were detected in the oral cancer tissues and normal oral
mucosa. Volcano plot analysis subsequently revealed aberrant
expression of miRNAs in OSCC (Fig.
2). This result suggests that miRNA expression profiles differ
between normal and abnormal cells.
To identify candidate cancer-related miRNAs, we
calculated the degree of differential expression of miRNAs between
normal oral mucosa and OSCC tissues. Twelve miRNAs (miR-31*,
miR-31, miR135b, miR-193a-5p, miR-103, miR-224, miR-93, miR-200c,
miR-183, miR-203, miR-21 and miR-223) were significantly
upregulated in most of the cancer tissue samples compared with the
normal oral mucosa. Those showing >4-fold upregulation were
considered candidate miRNAs; namely, miR-31*, miR-31, miR-135b,
miR-193a-5p, miR-103, miR-224, miR-93, miR-200c, miR-183, miR-203,
miR-21 and miR-223.
miR-21 has been investigated widely in various human
malignancies including hematological malignancies and glioblastoma
as a putative oncogenic miRNA (5,19–26).
Moreover, aberrantly expressed miR-203 has been detected in colon
(27), breast (28) and ovarian cancer (29–31),
while miR-31 is also known to have oncogenic functions in
esophageal cancer (32).
Upregulation of both miR-31 and miR-31* by delivery of pre-miR-31
was also shown to enhance OSCC oncogenicity (33). Our results also indicated that
miR-31 and miR-31* have a biological function in cancer
development. However, a role in oral cancer has yet to be
documented, and the function of these candidates in human cancer
remains unclear. miR-133a, miR-376c, miR-411, miR-30a-3p, miR-489,
miR-139-5p, miR-483-5p, miR-30e-3p, miR-409-3p, let-7c and
miR-486-5p showed a >4-fold downregulation (Table III). Previously, miR-133a was
shown to be downregulated in breast cancer tissues and correlated
with poor prognosis (34), while
let-7c is reportedly related to tumor growth inhibition in prostate
cancer (35).
Cervical lymph node metastasis has a large impact on
the prognosis of OSCC. We, therefore, performed volcano plot
analysis according to the metastatic state. Three downregulated
miRNAs (niR-489, miR-1291 and miR-483-5p) (Table IV) were detected, and subsequently,
a decision tree model was constructed to classify non-metastatic
and metastatic OSCC using Weka software. Our model divided the two
groups with 82% accuracy (Fig. 3).
This model may therefore have potential in determining the
prognosis of OSCC patients, acting as a predictive marker of
metastasis.
In summary, our findings identifying cancer-related
miRNAs in OSCC suggest that oral cancer may have a unique miRNA
expression pattern at the individual level. Further investigations
are now required to determine the molecular functions and
mechanisms of these miRNAs as well as their potential use as
prognostic and/or diagnostic markers in oral cancer.
Acknowledgements
We thank Tomohide Isobe and Tetsuhiko Tachikawa for
the diagnostic clinical oral cancer tissues.
References
|
1
|
Bernier J, Bentzen SM and Vermorken JB:
Molecular therapy in head and neck oncology. Nat Rev Clin Oncol.
6:266–277. 2009. View Article : Google Scholar
|
|
2
|
Bozec A, Peyrade F, Fischel JL and Milano
G: Emerging molecular targeted therapies in the treatment of head
and neck cancer. Expert Opin Emerg Drugs. 14:299–310. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hummel R, Maurer J and Haier J: MicroRNA:
biogenesis, function and role in rancer. Curr Genomics. 11:537–561.
2010. View Article : Google Scholar
|
|
4
|
Wiemer EA: The role of microRNAs in
cancer: no small matter. Eur J Cancer. 43:1529–1544. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lu J, Getz G, Miska EA, et al: MicroRNA
expression profiles classify human cancers. Nature. 435:834–838.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wu BH, Xiong XP, Jia J and Zhang WF:
MicroRNAs: new actors in the oral cancer scene. Oral Oncol.
47:314–319. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Reis PP, Tomenson M, Cervigne NK, Machado
J, Jurisica I and Pintilie M: Programmed cell death 4 loss
increases tumor cell invasion and is regulated by miR-21 in oral
squamous cell carcinoma. Mol Cancer. 10:2382010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Gottardo F, Liu CG, Ferracin M, et al:
Micro-RNA profiling in kidney and bladder cancers. Urol Oncol.
25:387–392. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Volinia S, Calin GA, Liu CG, et al: A
microRNA expression signature of human solid tumors defines cancer
gene targets. Proc Natl Acad Sci USA. 103:2257–2261. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hammond SM: MicroRNA therapeutics: a new
niche for antisense nucleic acids. Trends Mol Med. 12:99–101. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lagos-Quintana M, Rauhut R, Lendeckel W
and Tuschl T: Identification of novel genes coding for small
expressed RNAs. Science. 294:853–858. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Bartel DP: MicroRNAs: genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ruan K, Fang X and Ouyang G: MicroRNAs:
novel regulators in the hallmarks of human cancer. Cancer Lett.
285:116–126. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Farazi TA, Spitzer JI, Morozov P and
Tuschl T: miRNAs in human cancer. J Pathol. 223:102–115. 2011.
View Article : Google Scholar
|
|
16
|
Calin GA, Liu CG, Sevignani C, et al:
MicroRNA profiling reveals distinct signatures in B cell chronic
lymphocytic leukemias. Proc Natl Acad Sci USA. 101:11755–11760.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Schaefer A, Jung M, Mollenkopf HJ, et al:
Diagnostic and prognostic implications of microRNA profiling in
prostate carcinoma. Int J Cancer. 126:1166–1176. 2010.PubMed/NCBI
|
|
18
|
Schepeler T, Reinert JT, Ostenfeld MS, et
al: Diagnostic and prognostic microRNAs in stage II colon cancer.
Clin Chem. 54:1696–1704. 2008.
|
|
19
|
Zhang B, Pan X, Cobb GP and Anderson TA:
microRNAs as oncogenes and tumor suppressors. Dev Biol. 302:1–12.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Papagiannakopoulos T, Shapiro A and Kosik
KS: MicroRNA-21 targets a network of key tumor-suppressive pathways
in glioblastoma cells. Cancer Res. 68:8164–8172. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Rossi L, Bonmassar E and Faraoni I:
Modification of miR gene expression pattern in human colon cancer
cells following exposure to 5-fluorouracil in vitro. Pharmacol Res.
56:248–253. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Cheng Y, Ji R, Yue J, et al: MicroRNAs are
aberrantly expressed in hypertrophic heart: do they play a role in
cardiac hypertrophy? Am J Pathol. 170:1831–1840. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Liu X, Cheng Y, Zhang S, Lin Y, Yang J and
Zhang C: A necessary role of miR-221 and miR-222 in vascular smooth
muscle cell proliferation and neointimal hyperplasia. Circ Res.
104:476–487. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhu S, Wu H, Wu F, Nie D, Sheng S and Mo
YY: MicroRNA-21 targets tumor suppressor genes in invasion and
metastasis. Cell Res. 18:350–359. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Meng F, Henson R, Wehbe-Janek H, Ghoshal
K, Jacob ST and Patel T: MicroRNA-21 regulates expression of the
PTEN tumor suppressor gene in human hepatocellular cancer.
Gastroenterology. 133:647–658. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Löffler D, Brocke-Heidrich K, Pfeifer G,
et al: Interleukin-6 dependent survival of multiple myeloma cells
involves the Stat3-mediated induction of microRNA-21 through a
highly conserved enhancer. Blood. 15:1330–1333. 2007.PubMed/NCBI
|
|
27
|
Schetter AJ, Leung SY, Sohn JJ, Zanetti
KA, Bowman ED and Yanaihara N: MicroRNA expression profiles
associated with prognosis and therapeutic outcome in colon
adenocarcinoma. JAMA. 299:425–436. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Iorio MV, Ferracin M, Liu CG, et al:
MicroRNA gene expression deregulation in human breast cancer.
Cancer Res. 65:7065–7070. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Nam EJ, Yoon H, Kim SW, Kim H, Kim YT and
Kim JH: MicroRNA expression profiles in serous ovarian carcinoma.
Clin Cancer Res. 14:2690–2695. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Saini S, Majid S, Yamamura S, et al:
Regulatory role of mir-203 in prostate cancer progression and
metastasis. Clin Cancer Res. 17:5287–5298. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wellner U, Schubert J, Burk UC, et al: The
EMT-activator ZEB1 promotes tumorigenicity by repressing
stemness-inhibiting microRNAs. Nat Cell Biol. 11:1487–1495. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Alder H, Taccioli C, Chen H, et al:
Dysregulation of miR-31 and miR-21 induced by zinc deficiency
promotes esophageal cancer. Carcinogenesis. 33:1736–1744. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Chang KW, Kao SY, Wu YH, et al: Passenger
strand miRNA miR-31(*) regulates the phenotypes of oral cancer
cells by targeting RhoA. Oral Oncol. 49:27–33. 2013.PubMed/NCBI
|
|
34
|
Wu ZS, Wang CQ, Xiang R, et al: Loss of
miR-133a expression associated with poor survival of breast cancer
and restoration of miR-133a expression inhibited breast cancer cell
growth and invasion. BMC Cancer. 12:512012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Sugimura K, Miyata H, Tanaka K, et al:
Let-7 expression is a significant determinant of response to
chemotherapy through the regulation of IL-6/STAT3 pathway in
esophageal squamous cell carcinoma. Clin Cancer Res. 18:5144–5153.
2012. View Article : Google Scholar : PubMed/NCBI
|