Introduction
Pancreatic cancer is the fourth leading cause of
cancer-related mortality in Western countries and has the lowest
survival rate of any solid tumor (1). Although patients with pancreatic
cancer are treated with comprehensive therapy, including surgery,
chemotherapy and radiotherapy, their 5-year survival rate is <5%
(2). Approximately 25–30% of these
patients are initially diagnosed with locally advanced pancreatic
cancer (3,4). Although radiotherapy combined with
chemotherapy is the recommended treatment strategy, the
contribution of radiotherapy is unclear (3,4). To
improve the outcome of patients with pancreatic cancer, a greater
understanding of the biological mechanisms that contribute to tumor
refractoriness to radiotherapy is required. Moreover,
identification of markers of radioresistance may contribute to the
identification of agents to treat this fatal disease.
The cDNA microarray technique is a powerful tool for
analyzing comprehensive gene expression in cells (5). To identify genes that contribute to
resistance to radiation therapy, cDNA microarray analysis has
compared radioresistant and radiosensitive cancer cells in various
types of tumor, including uterine cervical, head and neck,
colorectal, breast, esophageal, lung, hepatocellular and pancreatic
cancer, as well as in malignant melanoma (6). Several genes identified by cDNA
microarray analysis encode proteins involved in apoptosis, DNA
repair, signal transduction, cell cycle and cell adhesion, as well
as genes encoding growth factors and growth factor receptors
(6). Although these genes may
contribute to radioresistance, more information is required to
evaluate the mechanism underlying resistance to radiotherapy in
these tumor types.
We therefore exposed pancreatic cancer cell lines to
radiation and compared comprehensive gene expression of irradiated
(IR) and parental cells using cDNA microarray technique. We
selected promising genes and evaluated their contribution to the
radioresistance of pancreatic cancer cell lines.
Materials and methods
Cell lines
Three human pancreatic cancer cell lines, SUIT-2,
PANC-1 and MIA PaCa-2, were obtained from the Japanese Cancer
Resource Bank (Tokyo, Japan) and three other human pancreatic
cancer cell lines, Capan-1, Capan-2 and CFPAC-1, were obtained from
the American Type Culture Collection (Manassas, VA, USA); all 6
cell lines bear mutant p53 (7–9). The
cells were maintained as previously described (10).
Irradiation of pancreatic cancer cell
lines
Two pancreatic cancer cell lines, Capan-1 and
CFPAC-1, were repeatedly exposed to 2 Gy X-ray irradiation for a
total dose of 10 Gy, followed by irradiation with 5 Gy to a total
dose of 60 Gy, with 14-day intervals between treatments. IR cells
were cultured in Dulbecco’s modified Eagle’s medium (DMEM)
supplemented with 10% FBS.
Evaluation of cell rediosensitivity
Radiosensitivity was assessed by colony formation
assays. All pancreatic cell lines were plated at 1×104
cells/well in 6-well plates (Thermo Scientific, Rockford, IL, USA)
for 24 h, followed by exchange of conditioned medium. The cells
were irradiated at 0, 1, 2, 3, 5, 7, 8 and 10 Gy and incubated for
7 days without exchange of medium. The plates were fixed in 70%
ethanol and stained with 0.1% crystal violet and colonies were
counted using a ChemiDoc XRS System (Bio-Rad Laboratories).
Survival rates were determined by comparing the number of colonies
at each radiation dose with that in unirradiated (0 Gy) cells. All
survival curves represent the combined results of three independent
experiments.
Microarray analysis
Microarray analyses were performed using IR-Capan-1
and IR-CFPAC-1 cells and their respective parental cells. The
qualities of the RNA samples were evaluated using a 2100
Bioanalyzer (Agilent Technologies, Waldbronn, Germany) as described
(11). Analyses were performed
using a HumanWG-6 Expression BeadChip and the results were analyzed
using BeadStudio software version 3.2.3 (both from Illumina, San
Diego, CA, USA).
Real-time qRT-PCR
Total RNA was extracted from pellets of cultured
cells using a High Pure RNA Kit (Roche Diagnostics, Mannheim,
Germany) and treated with DNase I (Roche Diagnostics), according to
the manufacturer’s instructions. Specific primers for S100 calcium
binding protein A4 (S100A4) (forward,
5′-atcgccatgatgtgtaccga-3′; reverse, 5′-cccaaccacatcagagg agt-3′)
and β-actin (forward, 5′-tgagcgcggctacagctt-3′; reverse,
5′-tccttaatgtcacgcacgattt-3′) RNAs were designed using primer 3,
with BLAST searches performed to ensure the specificity of each
primer. The extracts were analyzed by qRT-PCR using a QuantiTect
SYBR-Green RT-PCR Kit (Qiagen, Tokyo, Japan) and a Chrom4 Real-Time
PCR Detection System (Bio-Rad Laboratories, Hercules, CA, USA).
Each reaction mixture was initially incubated at 50°C for 30 min to
allow reverse transcription, in which first-strand cDNA was
synthesized by priming the total RNA with the same gene-specific
primer (reverse). PCR was initiated by incubation at 95°C for 15
min to activate the polymerase, followed by 40 cycles of 94°C for
15 sec, 58°C for 30 sec and 72°C for 30 sec. Each of these primer
sets produced a single prominent band of the expected size after
electrophoresis. Each sample was analyzed twice and any samples
showing >10% deviation in qRT-PCR results were tested a third
time. The level of mRNA expression in each sample was calculated by
reference to a standard curve generated using total RNA from the
SUIT-2 human pancreatic cancer cell line. The expression of
S100A4 mRNA was normalized relative to the expression of
β-actin mRNA in the same sample.
Western blotting
Cell proteins were extracted by PRO-PREP (iNtRON
Biotechnology, Seongnam, Korea) according to the manufacturer’s
instructions. Cell lysates containing 15 μg protein were
fractionated on 12% sodium dodecyl sulfate-polyacrylamide gel
electrophoresis gels and transferred to polyvinylidene difluoride
membranes (Millipore, Billerica, MA, USA). Each membrane was
incubated overnight at 4°C with anti-S100A4 (1:500; DAKO, Glostrup,
Denmark) or anti-β-actin (05-829, 1:1,000; Millipore) and
subsequently with secondary antibodies conjugated to horseradish
peroxidase (Santa Cruz Biotechnology, Inc.). Immunoblots were
detected by enhanced chemiluminescence with a Chemi Doc XRS
System.
Cell proliferation assay
Cell proliferation was evaluated by measuring the
fluorescence intensity of propidium iodide (PI), as previously
described (12). Briefly, cells
were seeded at 2×104 cells/well in 24-well tissue
culture plates (Becton Dickinson Labware, Bedford, MA, USA) and
cultured in DMEM containing 10% FBS for 24 h. Following
confirmation of cellular adhesion to the plates, the medium was
replaced and 30 μM PI and 600 μM digitonin were added
to each well to label the nuclei with PI. The fluorescence
intensity, corresponding to the total cell number, was measured
using an Infinite® 200 PRO multiwell plate reader
(TECAN, Mannedorf, Switzerland) with excitation and emission
wavelengths of 530 and 645 nm, respectively. A separate well
containing the same medium was used to provide a baseline PI signal
as a control. The difference in intensity between each sample well
and the control well was calculated. Cell proliferation was defined
relative to the number of cells counted immediately after
exchanging the medium. All experiments were performed in triplicate
wells and were repeated at least three times.
Statistical analysis
All statistical analyses were performed using JMP
8.0.1 software (SAS Institute, Inc., Cary, NC, USA). Pearson’s test
was measured to evaluate the relationships between S100A4 mRNA
expression and radiation dose associated with 50% cell survival,
S100A4 mRNA expression and relative proliferation of pancreatic
cancer cells and relative proliferation of pancreatic cancer cells
and radiation dose associate with 50% cell survival. All values are
expressed as means ± standard deviation. P<0.05 was considered
to indicate a statistically significant difference.
Results
Evaluation of radiosensitivity of
IR-CFPAC-1 and Capan-1 cells
Two pancreatic cancer cell lines, CFPAC-1 and
Capan-1, were exposed to fractionated irradiation until the total
irradiation dose was 60 Gy. Radiosensitivity was evaluated by the
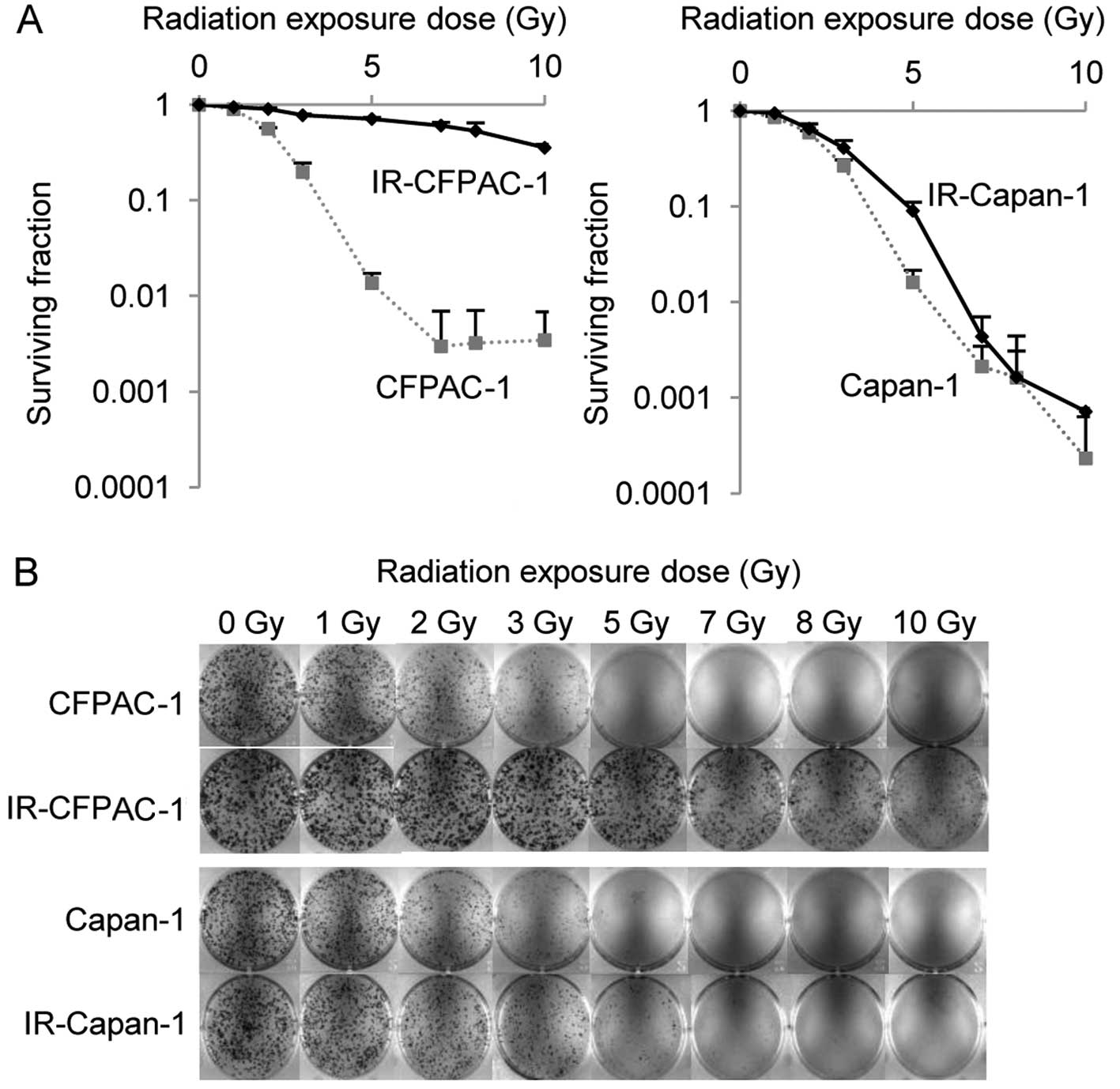
colony formation assay. IR-CFPAC-1 cells showed significantly
greater radioresistance compared to parental cells. The radiation
dose resulting in a 50% survival rate was significantly higher in
IR compared to parental CFPAC-1 cells (8.31±0.85 Gy vs. 2.14±0.04
Gy, P<0.0001; Fig. 1A and B),
but the difference was less pronounced in IR compared to parental
Capan-1 cells (2.66±0.24 Gy vs. 2.25±0.03 Gy, P=0.04).
mRNA expression profiling of IR and
parental pancreatic cancer cell lines
We performed global microarray expression analysis
of 48,803 genes to compare two IR pancreatic cancer cell lines
(IR-CFPAC-1 and IR-Capan-1) with those of their respective parental
cell lines (CFPAC-1 and Capan-1). A difference was defined as
significant if the ratio of expression between IR and parental
cells was >2- or <0.33-fold. We found that 4 genes were
upregulated (Table I) and 23 were
downregulated (Table II) in IR
compared with parental cells. One of the overexpressed genes was
S100A4, the increased expression of which has been associated with
radiation exposure in lung cancer cell lines (13). Of the 23 downregulated genes, none
of which has been reported to be associated with radioresistance, 7
were potentially involved in apoptosis, cell attachment and
inhibition of cell proliferation and tumor suppression.
 | Table IAverage ratios of expression of
upregulated genes in radioresistant compared with parental cells
(fold-change >2). |
Table I
Average ratios of expression of
upregulated genes in radioresistant compared with parental cells
(fold-change >2).
| | Fold-change | | |
|---|
| |
| | |
|---|
| Rank | Gene name | CFPAC-1 | Capan-1 | Function | Sequence code |
|---|
| 1 | S100 calcium
binding protein A4 (S100A4) | 138.3 | 8.5 | Regulation of a
number of cellular processes such as cell cycle progression and
differentiation | NM_019554.2 |
| 2 | Transmembrane
protein 158 (TMEM158) | 18.1 | 3.2 | Upregulated in
response to activation of the Ras pathway, but not under other
conditions that induce senescence | NM_015444.2 |
| 3 | Caveolin 2
(CAV2) | 16.3 | 2.4 | Signal
transduction, lipid metabolism, cellular growth control and
apoptosis | NM_001233.3 |
| 4 | Phosphoprotein
enriched in astrocytes 15 (PEA15) | 5.5 | 2.1 | Regulation of
apoptosis | NM_003768.2 |
 | Table IIIdentification of the 23 genes
downregulated in radioresistant CFPAC-1 and Capan-1 cells compared
with their respective parental cells (fold-change <0.33). |
Table II
Identification of the 23 genes
downregulated in radioresistant CFPAC-1 and Capan-1 cells compared
with their respective parental cells (fold-change <0.33).
| | Fold-change | | |
|---|
| |
| | |
|---|
| Rank | Gene name | CFPAC-1 | Capan-1 | Function | Sequence code |
|---|
| 1 | Inhibitor of DNA
binding 3 (ID3) | 0.02 | 0.27 | Negative regulation
of transcription factor activity | NM_002167.2 |
| 2 | Odontogenic,
ameloblast associated (ODAM) | 0.03 | 0.15 | Odontogenesis | NM_017855.3 |
| 3 | Rho-related BTB
domain containing 3 (RHOBTB3) | 0.03 | 0.19 | Small
GTPase-mediated signal transduction and the organization of the
actin filament system | NM_014899.3 |
| 4 | Eukaryotic
translation elongation factor 1 α2 (EEF1A2) | 0.04 | 0.07 | Protein
biosynthesis, translational elongation | NM_001958.2 |
| 5 | Naked cuticle
homolog 2 (Drosophila) (NKD2) | 0.07 | 0.30 | Negative regulator
of the Wnt in the mouse | NM_033120.2 |
| 6 | Quinolinate
phosphoribosyl-transferase (QPRT) | 0.08 | 0.04 | Generation of
precursor metabolites and energy, NAD biosynthesis, synaptic
transmission | NM_014298.3 |
| 7 | Imprinted
maternally expressed transcript (H19) | 0.08 | 0.05 | Non-coding RNA and
functions as a tumor suppressor | NR_002196.1 |
| 8 | Homo sapiens
laminin, α2 (LAMA2) | 0.10 | 0.09 | Regulation of
embryonic development, muscle development, cell migration and
adhesion | NM_001079823.1 |
| 9 | Vasorin (VASN) | 0.13 | 0.28 | Vasorin was
reported to contribute to neointimal formation after vascular
injury | NM_138440.2 |
| 10 | AT rich interactive
domain 5B (MRF1-like) (ARID5B) | 0.15 | 0.32 | Negative regulation
of transcription | NM_032199.1 |
| 11 | G protein-coupled
receptor 56 (GPR56) | 0.15 | 0.26 | Cell adhesion,
cell-cell signaling, signal transduction | NM_005682.4 |
| 12 | Vaccinia related
kinase 2 (VRK2) | 0.17 | 0.33 | Protein amino acid
phosphorylation | NM_006296.3 |
| 13 | Similar to
CG14853-PB (LOC285141) | 0.17 | 0.26 | | XM_939141.1 |
| 14 | Growth
arrest-specific 6 (GAS6) | 0.17 | 0.32 | Cell
proliferation | NM_000820.1 |
| 15 | T-box 3 (ulnar
mammary syndrome) (TBX3) | 0.17 | 0.29 | Morphogenesis,
skeletal development, regulation of transcription from RNA
polymerase II promoter | NM_016569.3 |
| 16 | Collagen, type VII,
α1 (COL7A1) | 0.21 | 0.20 | Cell adhesion,
phosphate transport, epidermis development | NM_000094.2 |
| 17 | Cystic fibrosis
transmembrane conductance regulator (CFTR) | 0.21 | 0.20 | Ion transport,
respiratory gaseous exchange | NM_000492.3 |
| 18 | Synaptotagmin-like
2 (SYTL2) | 0.22 | 0.15 | Vesicle-mediated
transport, intracellular protein transport | NM_032943.2 |
| 19 | POM and ZP3 fusion
(POMZP3) | 0.25 | 0.18 | | NM_152992.2 |
| 20 | Prominin 1
(PROM1) | 0.28 | 0.06 | Visual
perception | NM_006017.1 |
| 21 | Suppressor of
cytokine signaling 2 (SOCS2) | 0.32 | 0.19 | JAK-STAT cascade,
negative regulation of signal transduction, anti-apoptosis,
regulation of cell growth | NM_003877.3 |
| 22 | Shroom family
member 4 (SHROOM4) | 0.32 | 0.16 | To play a role in
cytoskeletal architecture | NM_020717.2 |
| 23 | Angiopoietin 1
(ANGPT1) | 0.33 | 0.16 | Angiogenesis, cell
differentiation, signal transduction | NM_001146.3 |
S100A4 expression in cultured pancreatic
cancer cell lines
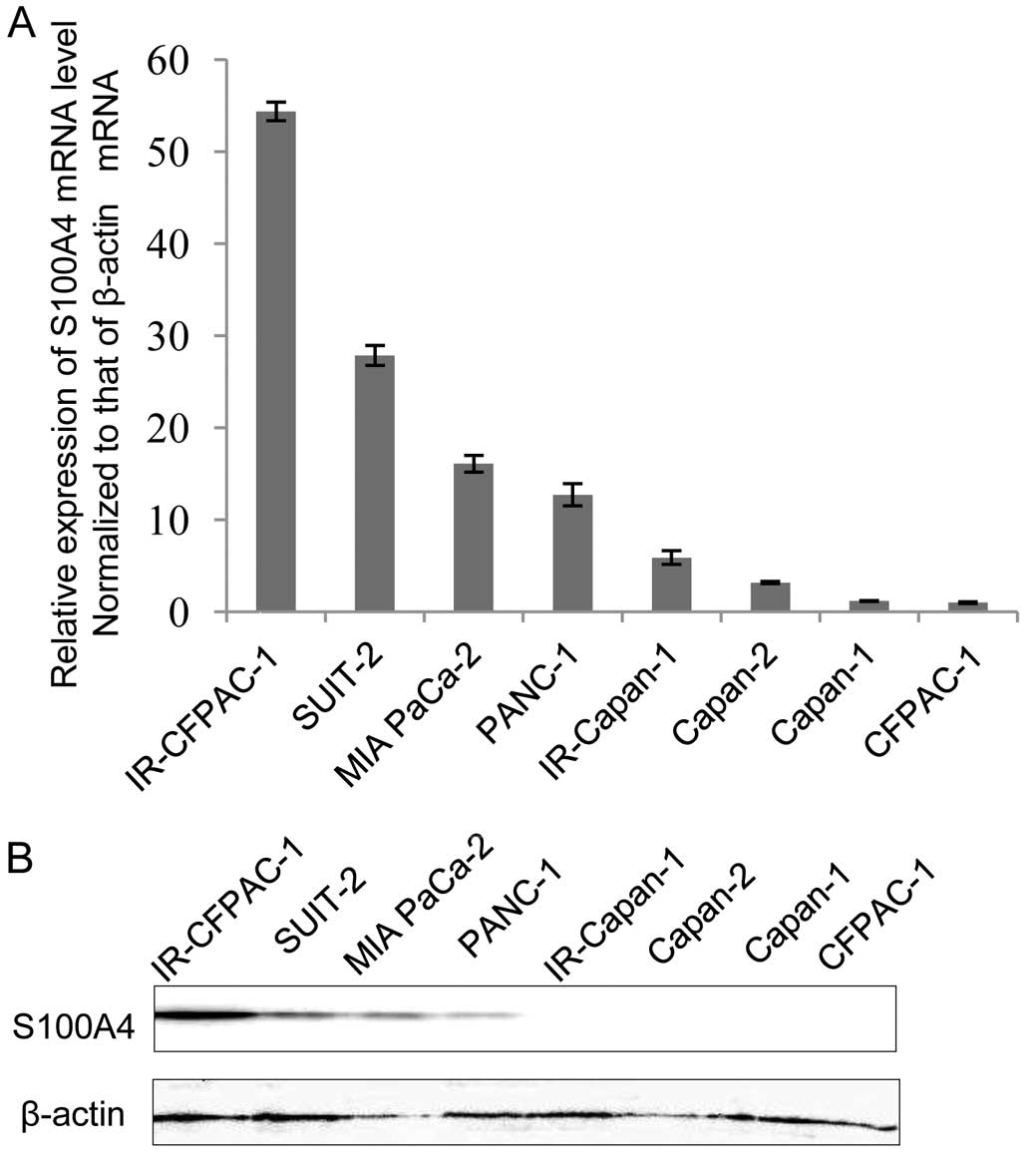
We investigated the levels of expression of S100A4
mRNA in cultures of 6 unirradiated and 2 IR pancreatic cancer cell
lines. S100A4 mRNA expression was higher in both IR cell lines
compared to the unirradiated cells, a finding consistent with our
microarray data (Fig. 2A). Compared
with their respective parental cell lines, the levels of expression
of S100A4 mRNA were 54-fold higher in IR-CFPAC-1 compared to
CFPAC-1 cells and 5.0-fold higher in IR-Capan-1 compared to Capan-1
cells, with both differences being statistically significant. The
level of S100A4 protein was significantly higher in IR-CFPAC-1
compared to IR-Capan-1 cells, the latter of which, as well as
parental Capan-1 cells, expressed no S100A4 protein (Fig. 2B). S100A4 protein and mRNA
expression were closely correlated in these cell lines. These
findings suggest that the difference in radiosensitivity between
IR-CFPAC-1 and IR-Capan-1 cells may be due to differences in S100A4
expression.
S100A4 mRNA expression is significantly
correlated with radiation dose associated with 50% survival
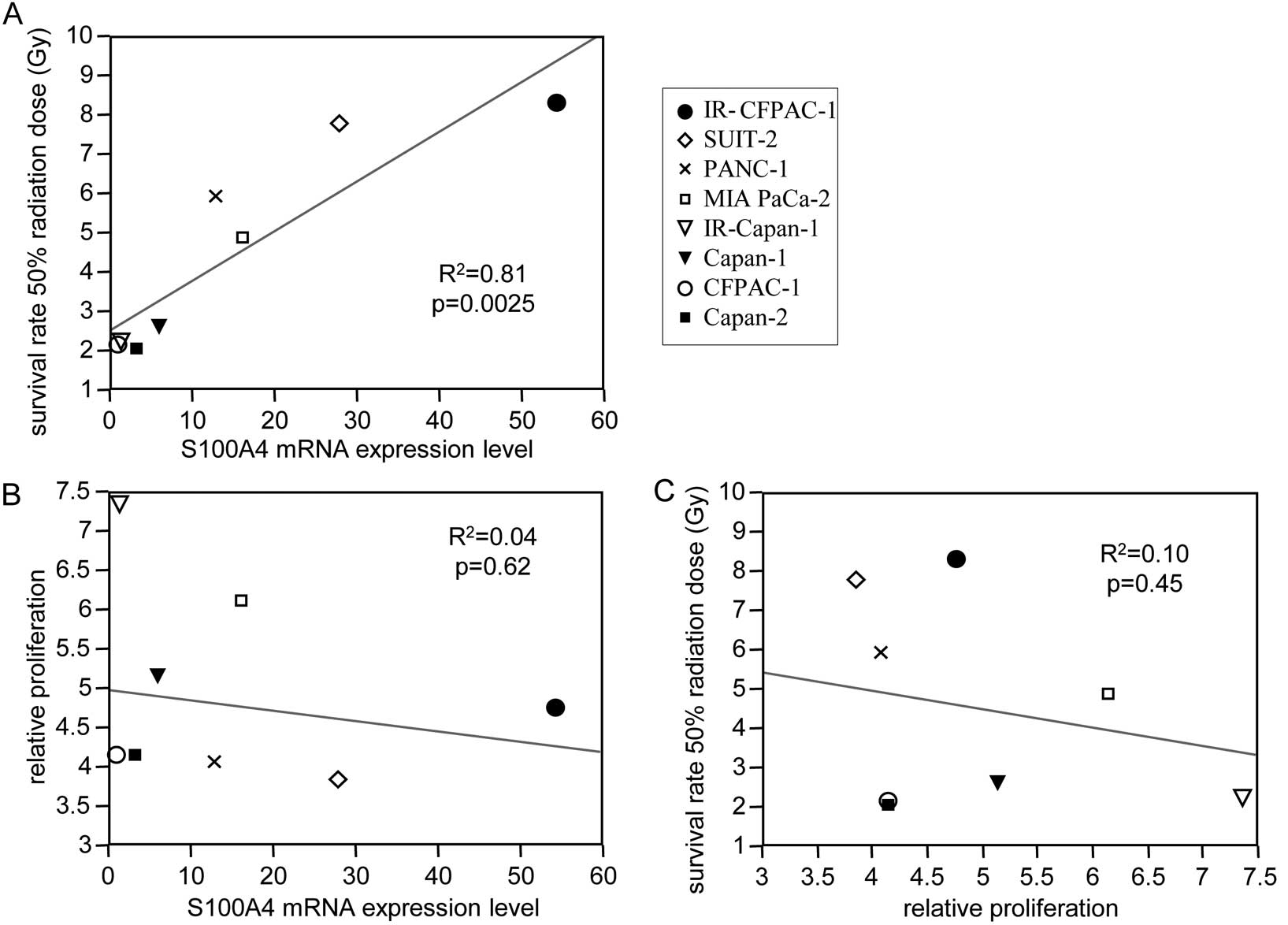
To analyze the radiosensitivity of pancreatic cell
lines, we calculated the radiation dose associated with 50%
survival rates (Fig. 3A and B,
Table III). We found that S100A4
mRNA expression was significantly correlated with this radiation
dose (Pearson’s test, R2=0.81, P=0.0025) (Fig. 4A) but not with cell proliferation
(Pearson’s test, R2=0.04, P=0.62) (Fig. 4B). In addition, radiation dose
associated with 50% cell survival was not correlated with cell
proliferation (Pearson’s test, R2=0.10, P=0.45)
(Fig. 4C). These findings indicate
that S100A4 mRNA expression was closely associated with the
radioresistance of pancreatic cancer cell lines.
 | Table IIIRadiation dose associated with 50%
survival rates of 6 pancreatic cancer cell lines and IR cell
lines. |
Table III
Radiation dose associated with 50%
survival rates of 6 pancreatic cancer cell lines and IR cell
lines.
| Cell line | Radiation dose
(Gy) |
|---|
| IR-CFPAC-1 | 8.31±0.85 |
| SUIT-2 | 7.80±1.07 |
| PANC-1 | 5.94±0.66 |
| MIA PaCa-2 | 4.90±0.64 |
| IR-Capan-1 | 2.66±0.24 |
| Capan-1 | 2.25±0.03 |
| CFPAC-1 | 2.14±0.04 |
| Capan-2 | 2.04±0.32 |
Discussion
Radiotherapy is among the key treatment strategies
for patients with various types of cancer. Some patients benefit
from radiotherapy, whereas others do not. In particular, the
survival benefits of radiotherapy in patients with pancreatic
cancer remain unclear (3,4). The molecular mechanisms associated
with tumor response to radiotherapy have been assessed in several
types of tumor by cDNA microarray analysis, comparing
radiosensitive and radioresistant cancer. These include esophageal
cancer (14), head and neck cancer
(15), cervical cancer (16–18),
breast cancer (19), pancreatic
cancer (6), glioblastoma (20), hepatocellular carcinoma (21), malignant melanoma (22) and lung cancer (23,24).
Most of the genes identified in these cDNA arrays were associated
with DNA-repair, apoptosis, growth factors, signal transduction,
cell cycle and cell adhesion. Similarly, our cDNA microarray
analysis also identified several genes involved in these signaling
pathways. However, none of these genes has been identified in
previous cDNA microarrays and some tumor types have few genes in
common that are up- or downregulated. Thus, the mechanism of
radioresistance in cancer cells is very complicated.
In analyzing pancreatic cancer cells, we found that
S100A4 mRNA expression was markedly upregulated following
irradiation, with the degree of upregulation differing
significantly in strongly radioresistant IR-CFPAC-1 cells and
weakly radioresistant IR-Capan-1 cells. Irradiation of lung cancer
cell lines has been reported to increase the expression of S100A4
and S100A6 proteins (13). To our
knowledge, our study is the first to show that the level of
expression of S100A4 mRNA is directly correlated with
radioresistance of pancreatic cancer cells.
S100A4 is a member of the S100 family of calcium
binding proteins, which are characterized by two distinct EF-hand
structural motifs (25,26). S100A4 is overexpressed in a number
of solid tumors, including breast, esophageal, gastric, colorectal
and pancreatic cancer and is associated with poor prognosis in
patients with these types of cancer (27,28).
S100A4 has been reported to promote cell motility and invasion in
cancer and to be associated with tumor metastasis (29,30).
Although we observed an association between S100A4 expression and
radioresistance of pancreatic cancer cell lines, the mechanism by
which S100A4 induces radioresistance remains unclear.
Wild-type p53 has been shown to play a crucial role
in cellular responses to radiation-induced DNA damage through cell
cycle arrest, apoptosis and DNA repair (31,32).
By contrast, mutant p53 has been shown to be involved in resistance
to radiotherapy (33–36) and to have oncogenic properties by
inducing the expression of sets of genes that activate cell
proliferation, cell survival and angiogenesis (35). S100A4 has been reported to interact
with wild-type p53 through the C-terminal domain of the latter and
may regulate p53 function, inducing cell apoptosis (37). Moreover, S100A4 may interact with
mutant p53 (37). Activation of
c-myc gene expression by mutant p53 has been reported to require
the C-terminal domain of the latter (38). The pancreatic cancer cells utilized
in this study contain mutant p53 (7–9). These
findings suggest that upregulation of S100A4 may increase its
opportunities to interact with mutant p53, enhancing resistance to
radiation.
The NF-κB pathway has been reported to play an
important role in the development of resistance to radiotherapy
(39–41). Inhibition of the NF-κB pathway
decreased resistance to radiotherapy. Moreover, release of S100A4
into the extracellular space has been found to activate the NF-κB
pathway and induce specific gene transcription (42). We have shown here that higher
expression of S100A4 was directly correlated with stronger
radioresistance of pancreatic cancer cell lines. These findings
suggest that upregulation of S100A4 expression may contribute to
the activation of the NF-κB pathway more strongly in pancreatic
cancer cells, leading to a sequential increase in radioresistance
via the transcription of genes associated with radioresistance.
Taken together, these findings indicate that S100A4 increases
radioresistance by interacting with transcription factors that
induce radioresistance. Further investigations of the mechanisms
underlying these interactions are required.
In summary, we showed that S100A4 expression in
pancreatic cancer cell lines was augmented by continuous
irradiation and that assaying S100A4 mRNA expression could predict
the radioresistance of pancreatic cancer cell lines. Based on these
findings, we conclude that S100A4 is involved in the
radioresistance of pancreatic cancer cell lines and may be involved
in the mechanism by which pancreatic cancer acquires
radioresistance. Treatment with agents targeting S100A4, along with
radiation, may improve the prognosis of patients with pancreatic
cancer undergoing radiotherapy.
Acknowledgements
This study was supported in part by the Ministry of
Education, Culture, Sports, Science and Technology of Japan (MEXT)
(MEXT KAKENHI Grants 23390327, 24659613, 24390319, 23659654,
24390318 and 23659655). The authors thank Emiko Manabe, Miyuki
Omori and Makiko Masuda (Department of Surgery and Oncology, Kyushu
University) for their technical assistance, and the Research
Support Center, Graduate School of Medical Sciences, Kyushu
University, for the technical support.
References
|
1
|
Hariharan D, Saied A and Kocher HM:
Analysis of mortality rates for pancreatic cancer across the world.
HPB (Oxford). 10:58–62. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel R, Ward E, Brawley O and Jemal A:
Cancer statistics, 2011: the impact of eliminating socioeconomic
and racial disparities on premature cancer deaths. CA Cancer J
Clin. 61:212–236. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Johung K, Saif MW and Chang BW: Treatment
of locally advanced pancreatic cancer: the role of radiation
therapy. Int J Radiat Oncol Biol Phys. 82:508–518. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wang F and Kumar P: The role of
radiotherapy in management of pancreatic cancer. J Gastrointest
Oncol. 2:157–167. 2011.PubMed/NCBI
|
|
5
|
Golub TR, Slonim DK, Tamayo P, et al:
Molecular classification of cancer: class discovery and class
prediction by gene expression monitoring. Science. 286:531–537.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ogawa K, Utsunomiya T, Mimori K, et al:
Differential gene expression profiles of radioresistant pancreatic
cancer cell lines established by fractionated irradiation. Int J
Oncol. 28:705–713. 2006.PubMed/NCBI
|
|
7
|
Moore PS, Sipos B, Orlandini S, et al:
Genetic profile of 22 pancreatic carcinoma cell lines. Analysis of
K-ras, p53, p16 and DPC4/Smad4. Virchows Arch. 439:798–802. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gardner-Thorpe J, Ito H, Ashley SW and
Whang EE: Differential display of expressed genes in pancreatic
cancer cells. Biochem Biophys Res Commun. 293:391–395. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Gupta S, Sathishkumar S and Ahmed MM:
Influence of cell cycle checkpoints and p53 function on the
toxicity of temozolomide in human pancreatic cancer cells.
Pancreatology. 10:565–579. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ohuchida K, Mizumoto K, Murakami M, et al:
Radiation to stromal fibroblasts increases invasiveness of
pancreatic cancer cells through tumor-stromal interactions. Cancer
Res. 64:3215–3222. 2004. View Article : Google Scholar
|
|
11
|
Antonov J, Goldstein DR, Oberli A, et al:
Reliable gene expression measurements from degraded RNA by
quantitative real-time PCR depend on short amplicons and a proper
normalization. Lab Invest. 85:1040–1050. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhang L, Mizumoto K, Sato N, et al:
Quantitative determination of apoptotic death in cultured human
pancreatic cancer cells by propidium iodide and digitonin. Cancer
Lett. 142:129–137. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Orre LM, Pernemalm M, Lengqvist J,
Lewensohn R and Lehtiö J: Up-regulation, modification, and
translocation of S100A6 induced by exposure to ionizing radiation
revealed by proteomics profiling. Mol Cell Proteomics. 6:2122–2131.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Fukuda K, Sakakura C, Miyagawa K, et al:
Differential gene expression profiles of radioresistant oesophageal
cancer cell lines established by continuous fractionated
irradiation. Br J Cancer. 91:1543–1550. 2004. View Article : Google Scholar
|
|
15
|
Hanna E, Shrieve DC, Ratanatharathorn V,
et al: A novel alternative approach for prediction of radiation
response of squamous cell carcinoma of head and neck. Cancer Res.
61:2376–2380. 2001.PubMed/NCBI
|
|
16
|
Harima Y, Togashi A, Horikoshi K, et al:
Prediction of outcome of advanced cervical cancer to
thermoradiotherapy according to expression profiles of 35 genes
selected by cDNA microarray analysis. Int J Radiat Oncol Biol Phys.
60:237–248. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Tewari D, Monk BJ, Al-Ghazi MS, et al:
Gene expression profiling of in vitro radiation resistance in
cervical carcinoma: a feasibility study. Gynecol Oncol. 99:84–91.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wong YF, Sahota DS, Cheung TH, et al: Gene
expression pattern associated with radiotherapy sensitivity in
cervical cancer. Cancer J. 12:189–193. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Helland A, Johnsen H, Frøyland C, et al:
Radiation-induced effects on gene expression: an in vivo study on
breast cancer. Radiother Oncol. 80:230–235. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Otomo T, Hishii M, Arai H, Sato K and
Sasai K: Microarray analysis of temporal gene responses to ionizing
radiation in two glioblastoma cell lines: up-regulation of DNA
repair genes. J Radiat Res. 45:53–60. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Jeong J, Hong SJ, Ju YJ, et al: Temporal
cDNA microarray analysis of gene expression in human hepatocellular
carcinoma upon radiation exposure. Oncol Rep. 15:33–48.
2006.PubMed/NCBI
|
|
22
|
Kumagai K, Nimura Y, Mizota A, et al:
Arpc1b gene is a candidate prediction marker for choroidal
malignant melanomas sensitive to radiotherapy. Invest Ophthalmol
Vis Sci. 47:2300–2304. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xu QY, Gao Y, Liu Y, Yang WZ and Xu XY:
Identification of differential gene expression profiles of
radioresistant lung cancer cell line established by fractionated
ionizing radiation in vitro. Chin Med J (Engl). 121:1830–1837.
2008.PubMed/NCBI
|
|
24
|
Lee YS, Oh JH, Yoon S, et al: Differential
gene expression profiles of radioresistant non-small-cell lung
cancer cell lines established by fractionated irradiation: tumor
protein p53-inducible protein 3 confers sensitivity to ionizing
radiation. Int J Radiat Oncol Biol Phys. 77:858–866. 2010.
View Article : Google Scholar
|
|
25
|
Donato R: S100: a multigenic family of
calcium-modulated proteins of the EF-hand type with intracellular
and extracellular functional roles. Int J Biochem Cell Biol.
33:637–668. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Tarabykina S, Kriajevska M, Scott DJ, et
al: Heterocomplex formation between metastasis-related protein
S100A4 (Mts1) and S100A1 as revealed by the yeast two-hybrid
system. FEBS Lett. 475:187–191. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Helfman DM, Kim EJ, Lukanidin E and
Grigorian M: The metastasis associated protein S100A4: role in
tumour progression and metastasis. Br J Cancer. 92:1955–1958. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ikenaga N, Ohuchida K, Mizumoto K, et al:
S100A4 mRNA is a diagnostic and prognostic marker in pancreatic
carcinoma. J Gastrointest Surg. 13:1852–1858. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Boye K and Maelandsmo GM: S100A4 and
metastasis: a small actor playing many roles. Am J Pathol.
176:528–535. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Mishra SK, Siddique HR and Saleem M:
S100A4 calcium-binding protein is key player in tumor progression
and metastasis: preclinical and clinical evidence. Cancer
Metastasis Rev. 31:163–172. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
El-Deiry WS: The role of p53 in
chemosensitivity and radiosensitivity. Oncogene. 22:7486–7495.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Bristow RG, Benchimol S and Hill RP: The
p53 gene as a modifier of intrinsic radiosensitivity: implications
for radiotherapy. Radiother Oncol. 40:197–223. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Li R, Sutphin PD, Schwartz D, et al:
Mutant p53 protein expression interferes with p53-independent
apoptotic pathways. Oncogene. 16:3269–3277. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Saintigny Y, Rouillard D, Chaput B, Soussi
T and Lopez BS: Mutant p53 proteins stimulate spontaneous and
radiation-induced intrachromosomal homologous recombination
independently of the alteration of the transactivation activity and
of the G1 checkpoint. Oncogene. 18:3553–3563. 1999. View Article : Google Scholar
|
|
35
|
Cadwell C and Zambetti GP: The effects of
wild-type p53 tumor suppressor activity and mutant p53
gain-of-function on cell growth. Gene. 277:15–30. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Williams JR, Zhang Y, Zhou H, et al: A
quantitative overview of radiosensitivity of human tumor cells
across histological type and TP53 status. Int J Radiat Biol.
84:253–264. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Grigorian M, Andresen S, Tulchinsky E, et
al: Tumor suppressor p53 protein is a new target for the
metastasis-associated Mts1/S100A4 protein: functional consequences
of their interaction. J Biol Chem. 276:22699–22708. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Frazier MW, He X, Wang J, Gu Z, Cleveland
JL and Zambetti GP: Activation of c-myc gene expression by
tumor-derived p53 mutants requires a discrete C-terminal domain.
Mol Cell Biol. 18:3735–3743. 1998.PubMed/NCBI
|
|
39
|
Ahmed KM and Li JJ: NF-κB-mediated
adaptive resistance to ionizing radiation. Free Radic Biol Med.
44:1–13. 2008.
|
|
40
|
Dai Y, Lawrence TS and Xu L: Overcoming
cancer therapy resistance by targeting inhibitors of apoptosis
proteins and nuclear factor-kappa B. Am J Transl Res. 1:1–15.
2009.PubMed/NCBI
|
|
41
|
Li F and Sethi G: Targeting transcription
factor NF-κB to overcome chemoresistance and radioresistance in
cancer therapy. Biochim Biophys Acta. 1805:167–180. 2010.
|
|
42
|
Boye K, Grotterød I, Aasheim HC, Hovig E
and Maelandsmo GM: Activation of NF-κB by extracellular S100A4:
analysis of signal transduction mechanisms and identification of
target genes. Int J Cancer. 123:1301–1310. 2008.
|