Introduction
Colorectal cancer is one of the major causes for
cancer-associated death in males and females (1). Advancements in treatments involving a
combination of surgical resection, radiation and chemotherapy have
increased the patient’s five-year survival, however, colorectal
cancer remains a major public health concern (2). Therefore, an improved therapeutic
strategy is greatly needed. To our knowledge, the molecular
mechanisms of colorectal cancer is complicated and still poorly
understood. Although tumor-suppressor genes and oncogenes such as
APC, TP53 and K-ras (3,4), have been demonstrated to contribute to
colorectal cancer development, only a few miRNAs have been studied
to determine their roles in colorectal carcinogenesis, such as
miR-21 and miR-145 (5,6).
MicroRNAs (miRNAs) are a broad class of small,
non-coding endogenous single RNA molecules that play important
roles in gene expression through directly binding to the
3′-untranslated region (3′-UTR) of target gene mRNA, leading to
mRNA cleavage or translational repression (7). They are differentially expressed in
human cancers and play essential roles in carcinogenesis. For
instance, number of miRNAs dysregulated in colorectal cancer were
revealed by microarray profiles in colorectal cancer tissues
compared to normal tissues, including miR-100 (8). However, there are fewer studies on the
roles of miR-100 in colorectal cancer.
In this study, we identified a new
anti-proliferative, pro-apoptosis and anti-metastatic miRNA,
miR-100, in colorectal cancer cells that is frequently
downregulated in colorectal cancer tissues compared to normal
tissues. Accordingly, ectopic expression of miR-100 could inhibit
SW620 cell proliferation and invasion, while blockage of miR-100
yielded the reverse phenotype. Moreover, we identified RAP1B, a
putative oncogene in colorectal cancer, as the direct functional
target of miR-100.
Materials and methods
Ethics statement and human colorectal
carcinoma tissues
All specimens were from patients who underwent
surgery at the third affiliated hospital of Nanchang University
Hospital. The protocol had the approval of the Clinical Research
Ethics Committee of Nanchang University School of Medicine, and the
research was carried out according to the provisions of the
Helsinki Declaration of 1975. Written informed consent was obtained
from all participants involved in the study.
Cell lines and transfection
The colorectal carcinoma cell line SW620 was
purchased from the Cell Bank of Type Culture Collection of Chinese
Academy of Sciences, Shanghai Institute of Cell Biology, Chinese
Academy of Sciences. SW620 cells were maintained in RPMI-1640
containing 10% fetal calf serum. Cultures were incubated at 37°C in
standard tissue culture incubators. MiR-100 mimics, miR-100
inhibitor (anti-miR-100) were synthesized by Genepharma, Shanghai,
China. Oligonucleotide transfection was performed with
Lipofectamine 2000 reagents (Invitrogen, Carlsbad, CA, USA). The
final concentration of miR-100 mimics or anti-miR-100 in the
transfection system was 100 nM. Transfection efficiency for the
single and co-transfected studies was determined by fluorescence
microscope.
RNA extraction and real-time PCR
Total RNA was extracted using TRIzol reagent.
Real-time PCR analyses were carried out to detect mRNA expression
using SYBR Premix Ex Taq (Takara, Dalian, China), and GAPDH was
used as an internal control. MiRcute miRNA qPCR detection kit
(Tiangen, Beijing, China) was used to quantitate the expression
levels of mature miR-100 according to the provided protocol, and U6
was used as an internal control.
Open access software
Targetscan was used to predict the putative targets
of miR-100 (http://www.targetscan.org/). miRNA-Map 2.0 was used to
ananlysis the expression of miR-100 in colon cancer tissues and
normal colon tissues (http://mirnamap.mbc.nctu.edu.tw/).
Cell proliferation and colony formation
assay
A cell proliferation assay was performed with the
Cell Counting Kit-8 (Dojindo, Kumamoto, Japan) according to the
manufacturer’s instruction. For the colony formation assay, 1000
cells were placed in each 100-mm cell culture dish and maintained
in media containing 10% FBS for two weeks. Colonies were fixed with
4% paraformaldehyde and stained with coomassie brilliant blue.
Annexin V-FITC and PI staining
Enumeration of apoptotic cells was done by using
FITC conjugated Annexin V and PI (BD Pharmingen, San Jose, CA).
Cells were washed twice in cold 1X PBS and resuspended in Annexin
V-binding buffer (BD Pharmingen) at a concentration of
3×106 per ml. This suspension (100 μl) was stained with
5 μl of Annexin V-FITC and 5 μl PI. These cells were gently
vortexed and incubated for 15 min at room temperature in the dark.
After addition of 400 μl of binding buffer to each tube, cells were
analyzed by flow cytometry.
Cell invasion assay
For invasion assay, the membrane invasion culture
system (Transwell membranes of 6.5-mm diameter and 8-ml pore size;
Costar) was used according to the standard protocol. Briefly,
Harvested cells (1×105) suspended in 100 μl of serum
free RPMI-1640 were added into the upper compartment of the
chamber. A total of 1000 μl conditioned RPMI-1640 medium with 20%
(v/v) fetal bovine serum was used as a source of chemoattractant
and placed in the lower compartment of the chamber. After 48 h, the
non-invasive cells on the upper surface of the membrane were
removed with a cotton swab. The transformed cells that migrated
through the Matrigel matrix and stuck to the lower surface of the
membrane were fixed with 4% paraformaldehyde, stained with 1%
crystal purple. The invasive cells were then counted (five
high-power fields/chamber) using an inverted microscope. Each test
was repeated in triplicate.
Luciferase assay
SW620 cells were cultured in 24-well plates and
cotransfected with 100 nM of miR-100 mimics or anti-miR-100, 100 ng
reporters, and 10 ng pGL3-CMV Renilla luciferase reporter using
Lipofectamine 2000. After 24 h of transfection, firefly and Renilla
luciferase activities were measured using the dual-luciferase
reporter assay system (Promega, Madison, WI, USA).
Western blotting
Proteins were separated on 15% SDS-PAGE gel and then
transferred to PVDF membrane. After blocked with 5% nonfat milk,
the membrane was incubated with rabbit anti-RAP1B polyclonal
antibody (1:1000, Proteintech, Chicago, IL, USA) and anti-GAPDH
antibody (Abcam, San Francisco, CA, USA, 1:1000 dilution). The
secondary antibody was goat anti-rabbit IgG conjugated with HRP
(horseradish Peroxidase) with a dilution of 1:1000. The bound
antibodies were detected using ECL Plus Western blotting detection
system (GE Healthcare). GAPDH was used as an internal control to
normalize RAP1B expression level.
Tumor formation assay in a nude mouse
model
The SW620 cells (5×106) were injected
into the flanks of athymic nude mice. One week after the
injections, mice with comparably sized tumors were treated for 4
weeks with miR-NC and miR-100 mimics. Tumor growth was examined
twice a week. After 4 weeks, the mice were sacrificed and examined
for the growth of subcutaneous tumors.
Results
miR-100 is downregulated in human
colorectal cancer specimen
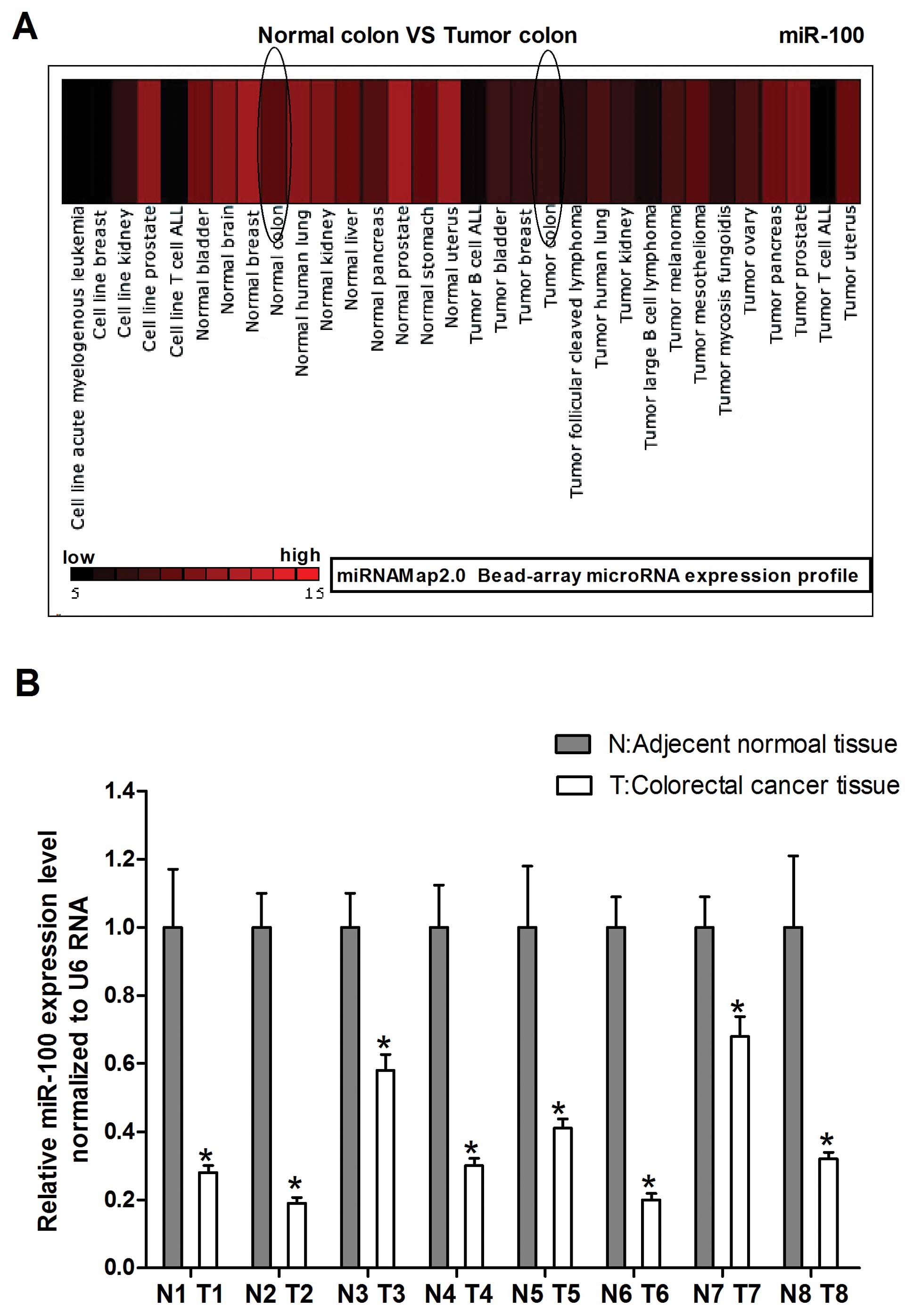
MiRNA-Map-2.0 was used to investigate miR-100
expression in normal colon compared with tumor colon. As shown in
Fig. 1A we found that miR-100
levels were frequently downregulated in tumor human colon compared
to normal colon. To explore the possible role of miR-100 in human
colorectal cancer development, we detected miR-100 expression in
human colorectal cancer specimen obtained from 8 patients by
real-time PCR. The carcinoma tissues showed reduced miR-100
expression with respect to normal counterparts, which is consistent
with the miRNAMap-2.0 gene chip results (Fig. 1B). Together, these results suggest
that miR-100 plays an important role in colorectal cancer
development.
miR-100 regulates proliferation of human
colorectal cancer SW620 cells in vitro and in vivo
To determine the role of miR-100 in proliferation of
colorectal cancer cells in vitro, MTT assays and colony
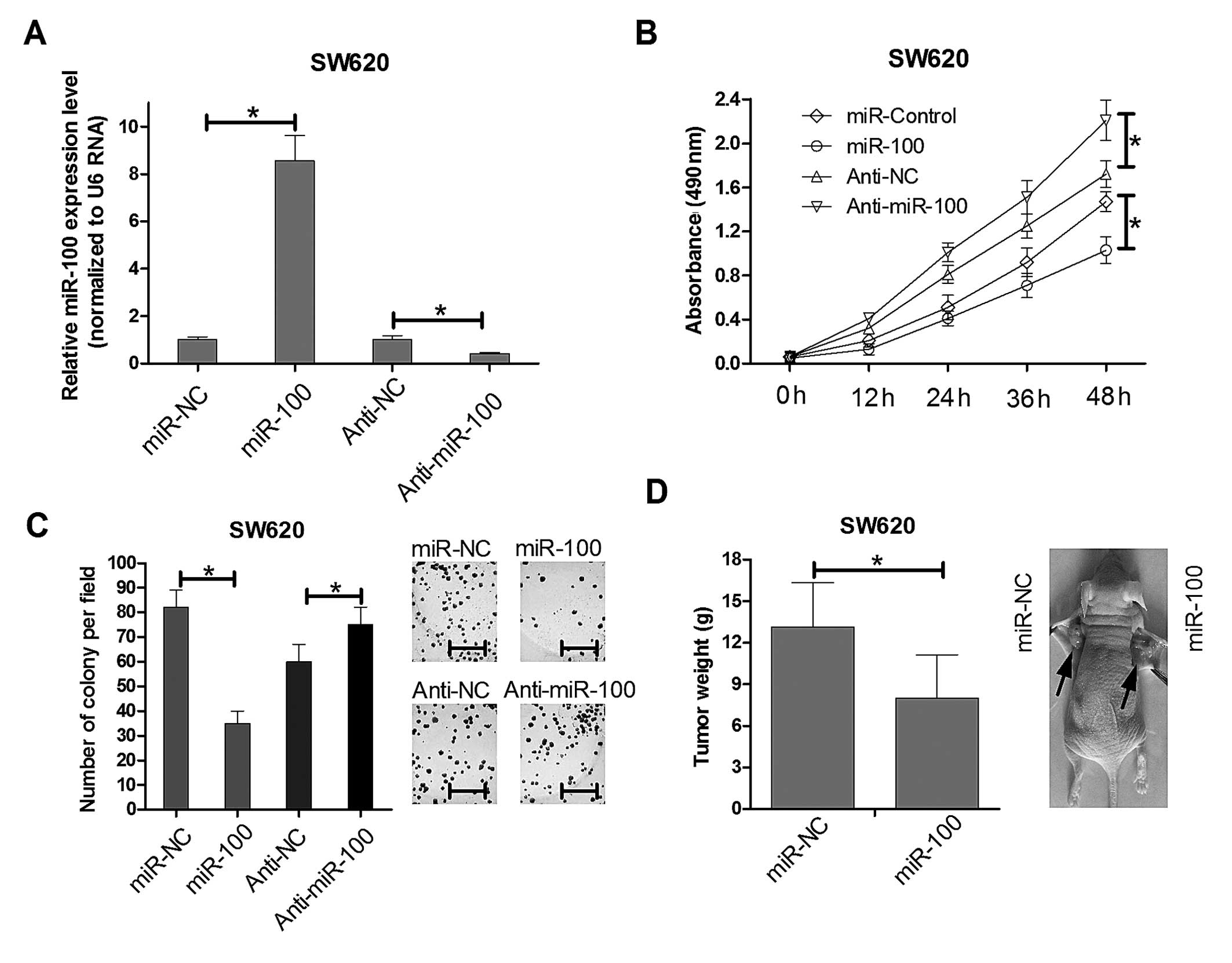
formation assays were performed. Real-time PCR results showed the
miR-100 mimics could significant increased the endogenous miR-100
expression level while anti-miR-100 mimics greatly reduced its
expression in SW620 as showed in Fig.
2A. Then we utilized the miR-100 mimics and anti-miR-100 mimics
compared with their relative control mimics to infect the SW620
cells. MTT detection showed that overexpression of miR-100
decreased cell viability in SW620 cells lines at 48 h after
transfection. The inhibition of miR-100 increased SW620 cell
proliferation (Fig. 2B). Colony
formation assays were used to further elucidate the effect of
miR-100 on the growth of SW620 cells. The colony formation rate of
SW620 cells transfected with miR-100 was reduced ~65% over that of
the control group (Fig. 2C). The
opposite phenomenon was observed in SW620 cells transfected with
anti-miR-100. These results revealed that miR-100 regulates the
proliferation of human colorectal cancer SW620 cell lines in
vitro. Moreover, to determine whether miR-100 is involved in
tumorigenesis of colorectal cancer in vivo, we injected
SW620 cells into the flanks of nude mice. One week after the
injections, mice with comparably sized tumors were treated for 4
weeks with miR-NC and miR-100 mimics, and the tumor growth activity
was measured. When tumors were harvested, the average weight of
tumors derived from the miR-100 group was lower than that of the
control group (Fig. 2D). These
results were consistent with the effects of miR-100 in vitro
and strongly suggest that miR-100 regulated proliferation of human
colorectal cancer SW620 cells.
miR-100 regulates apoptosis and invasion
of human colorectal cancer SW620 cells in vitro
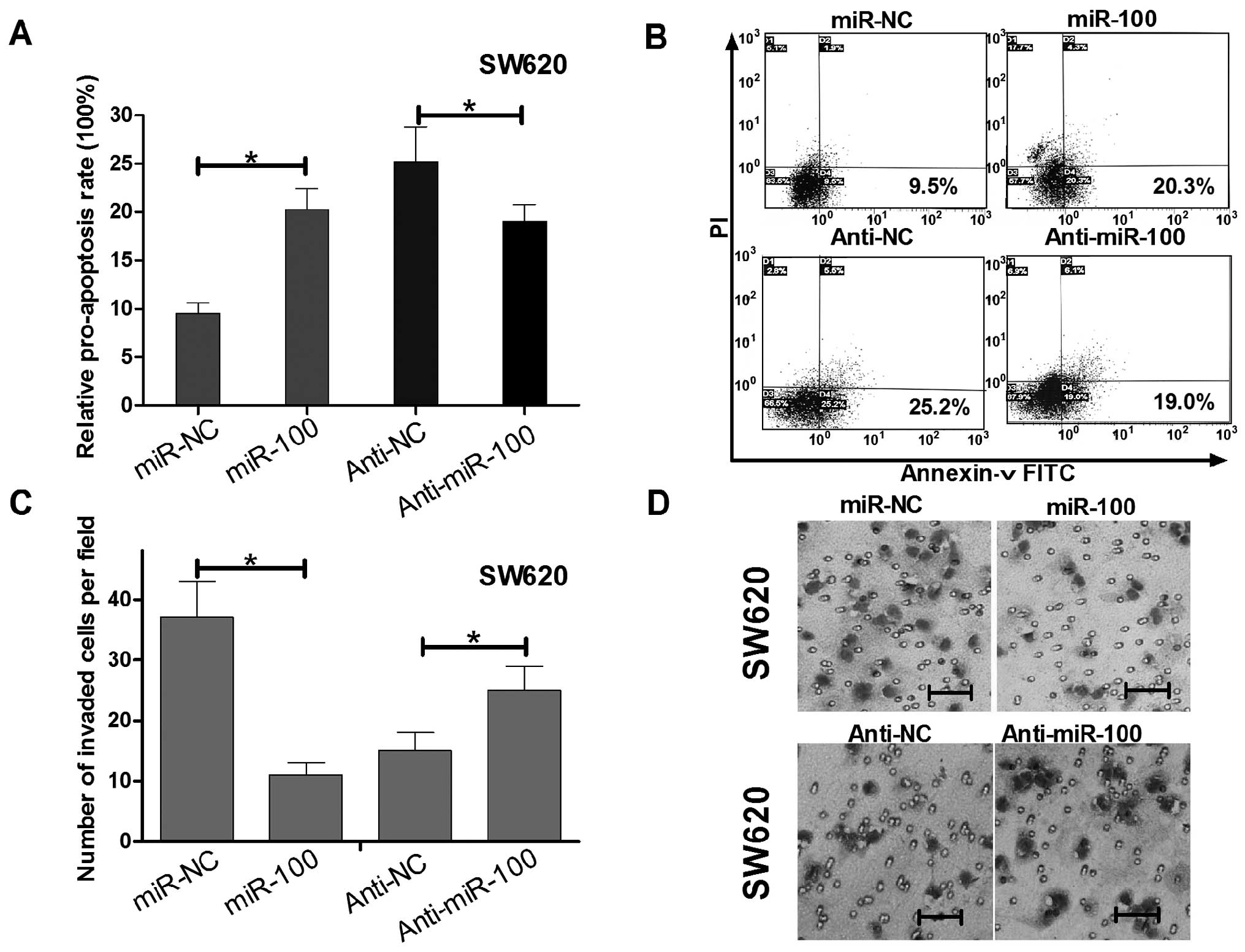
To determine whether the SW620 cell growth
regulation was attributed to apoptosis, we performed flow
cytometric analysis of SW620 cells after transfection of miR-100
mimics, anti-miR-100 or their relative controls. In miR-100 or
anti-miR-100 transfected SW620 cells, the rates of early
apoptosis/necrosis (D2 quadrant) were 20.3 and 19.0%, while the
rates in miR-control and anti-miR-100 control transfected cells
were 9.6 and 25.2%, respectively (Fig.
3A and B). These results suggest miR-100 induced while
anti-miR-100 reduced apoptosis of SW620 cells.
To determine whether miR-100 could regulate invasion
of SW620 cells, we performed Transwell invasion assays. As shown in
Fig. C and D, SW620 cells
transfected with miR-100 mimics displayed invasion ability
inhibition when compared with the control group, while transfected
with anti-miR-100 have the opposite effect. The above data
indicated that miR-100 not only could regulate SW620 cell growth
and apoptosis, but also impaired SW620 cell invasion.
RAP1B directly targeted by miR-100 is
inversely expressed with miR-100 in human colorectal cancer
Given that miR-100 has pivotal function in human
colorectal cancer SW620 cells, the question how the miRNA exerts
its role in colorectal cancer needs to be investigated. TargetScan
prediction algorithm was used for computational screen of genes
with complementary sites of miR-100 in their 3′-UTR. We found that
RAP1B, a member of RAS oncogene family, was a putative target of
miR-100 in the top 100 predicted targets (http://www.targetscan.org/cgi-bin/targetscan/vert_61/targetscan.cgi?species=Human&gid=&mir_c=&mir_sc=miR-99ab/100&mir_nc=&mirg=&sortType=&allTxs=&incl_nc=100).
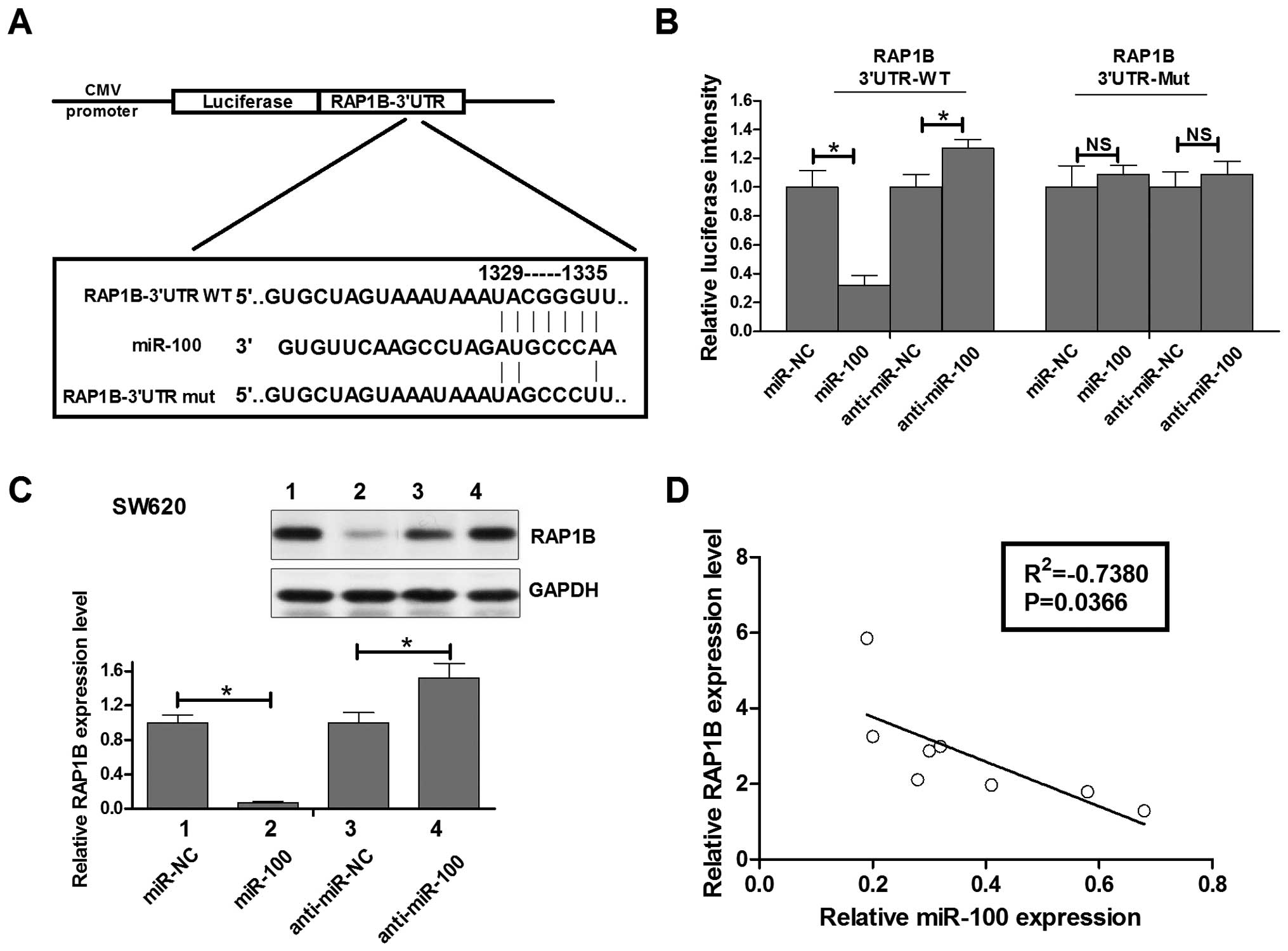
To confirm this possibility, the miR-100 binding sequences present
at the 3′-UTR of RAP1B mRNA (WT-3′-UTR), its mutant site
(RAP1B-3′UTR-mut) were subcloned downstream of the luciferase
reporter gene in pGL3 vector (Fig.
4A) and then co-transfected into SW620 cells. The relative
luciferase activity of the reporter that contained wild-type 3′-UTR
was decreased by 70% when miR-100 was co-transfected and it was
increased by 28% when anti-miR-100 was co-transfected, but the
luciferase activity of RAP1B-3′UTR-mut reporter was unaffected by
simultaneous transfection of miR-100 or anti-miR-100. These results
suggesting that miR-100 might suppress RAB14 expression through the
putative binding site in its 3′-UTR.
Next, real-time PCR and western blot assays were
performed to check whether miR-100 expression affects the
expression of endogenous RAP1B at both transcriptional and
translational levels. Consistent with the results of Luciferase
report assay, the levels of RAB14 mRNA showed a significant
decrease between miR-100 mimic-transfected SW620 cells and
miR-control-transfected cells or between anti-miR-100 transfected
SW620 cells and anti-miR-control transfected cells (data not
shown). Moreover, western blot analysis showed that the level of
RAP1B protein expression in miR-100 transfected SW620 cells was
inhibited by 90% compared with that in miR-control transfected
cells, while the level of RAP1B protein expression in anti-miR-100
transfected SW620 cells was upregulated by 45% compared with that
in anti-miR-control transfected cells (Fig. 4C). Having identified RAP1B as a
target of miR-100, we assessed the relationship of the expression
of miR-100 and RAP1B in human colorectal cancer tissues. Fig. 4D shows the RAP1B mRNA was inversely
expressed with miR-100 in human colorectal cancer and statistically
significant in the Pearson relation mean (R2=−0.7380,
P=0.0366). These data indicated that miR-100 could negatively
regulate RAP1B expression in human colorectal cancer SW620 cells by
interaction with the putative binding site in 3′-UTR of RAP1B mRNA,
which may partly explained the miR-100 induced growth, apoptosis
and invasion regulation mechanism of SW620 cells.
Discussion
It has been widely demonstrated that miRNAs regulate
diverse biological processes, including tumorigenesis. The role of
miR-100 is reported to be frequently downregulated in human cancer,
such as acute lymphoblastic leukaemia, hepatocellular carcinoma,
human esophageal squamous cell carcinoma, human bladder urothelial
carcinoma, non-small cell lung cancer and breast cancer (9–14).
However, its expression patterns in tumors are controversial. Wang
et al reported that miR-100 overexpression strongly
associates with advanced tumor progression and unfavorable clinical
outcome of patients with renal cell carcinoma (RCC) (15). Herein, we focused on the regulation
of miR-100 in colorectal cancer. First, we utilized the miR-map2.0
software to test the expression of miR-100 in normal colon tissues
and tumor colon tissues. Next, we examined miR-100 expression in
human colorectal cancer tissues and matched normal tissues by
real-time RT-PCR assay. We discovered that the levels of miR-100
were downregulated in tumor tissues, compared with the matched
normal tissues in eight pairs of matched specimens. Therefore, we
hypothesized that the downregulated miR-100 may function as a tumor
suppressor gene in colorectal cancer, which was consistent with
most of other human solid tumor types.
Using the MTT and colony formation assays to detect
the effect of miR-100 on the growth capacity of colorectal cancer
cell lines in vitro, we found that SW620 cells transfected
with the miR-100 mimics exhibited decreased growth compared with
the control cells while transfected with the miR-100 inhibitor
exhibited increased growth. The in vivo study also
demonstrated miR-100 could reduce SW620 cell proliferation. Thus,
we inferred that miR-100 may be a growth inhibition factor in
colorectal cancer.
To further reveal the exact role of miR-100 in
colorectal cancer, we tested the effect of miR-100 on apoptosis and
invasion by up- and downregulating the expression level of miR-100.
The results showed that increased miR-100 induced apoptosis of
SW620 cells, while decreased miR-100 inhibited apoptosis,
suggesting that miR-100 suppressed the ability of SW620 cells to
proliferate by inducing apoptosis in colorectal cancer. Since
metastasis is an important feature of colorectal cancer, we
examined the implication of miR-100 in SW620 cell invasion, and
data showed that miR-100 had a negative effect on invasion
suggesting that downregulation of miR-100 in colorectal cancer
cells may play roles in the development of colorectal cancer
through inhibiting cell proliferation, inducing apoptosis, and
decreasing cell invasion. However, the inhibition of invasion
ability by miR-100 in colorectal cancer should be confirmed with
further in vivo experiments.
As we known, miRNAs are recognized as important
regulators of gene expression, suppressing the expression of target
genes through translational repression or degradation of a target
transcript. We integrated bioinformatics-based predictions and the
resulting candidate functions and found that the RAP1B gene had the
highest recurrence rate as a potential target gene of miR-100.
miRNAs are believed to bind partially to the homologous sequence of
a target gene in the 3′-UTR. Accordingly, we constructed a
luciferase reporter plasmid bearing the wild-type 3′-UTR of RAP1B
mRNA for in vitro analysis. We found that inhibition or
overexpression of miR-100 could significantly enhance or reduce
luciferase expression, respectively. Furthermore, we constructed
another luciferase reporter vector containing a mutated miR-100
‘seed region’ binding site, and no significant difference was
detected when miR-100 was either overexpressed or inhibited. These
results suggested that miR-100 can directly and negatively regulate
RAP1B gene expression by binding to the 3′-UTR of RAP1B mRNA. It
was suggested that highly expressed miRNAs can suppress target gene
expression, whereas inhibition of an endogenous miRNA can protect
mRNA targets from increased degradation. Accordingly, we utilized
western blot assays to confirm the hypothesis that RAP1B was
regulated by miR-100. We found that when miR-100 was blocked, RAP1B
expression was enhanced, while miR-100 was overexpressed, RAP1B
protein expression levels were reduced. Thus, we concluded that
miR-100 negatively regulated the expression of RAP1B.
RAP1B was a member of the RAS-like small GTP-binding
protein superfamily. Members of this family regulate multiple
cellular processes including cell adhesion and growth and
differentiation. RAP1B localizes to cellular membranes and has been
shown to regulate integrin-mediated cell signaling. It also plays a
role in regulating outside-in signaling in platelets (16–20).
Recently, Rap1B was reported to be regulated by miR-139 in
colorectal cancer and regulate by miR-518b in esophageal squamous
cell carcinoma (21,22). However, RAP1B induced mechanism of
tumor cell malignant behavior is complex and multiple. Besides, it
has been proposed that a single miRNA can target several genes and
multiple miRNAs can target a single gene in a comprehensive manner
(23,24). In the present study, we identified
that miR-100 negatively regulates the expression of RAP1B possibly
supporting the hypothesis that the overexpression of RAP1B in
colorectal cancer, at least partly, result from not only the
underexpression of miR-139 but also miR-100. How many miRNAs
participate in the RAP1B pathway on colorectal cancer need to be
further elucidated.
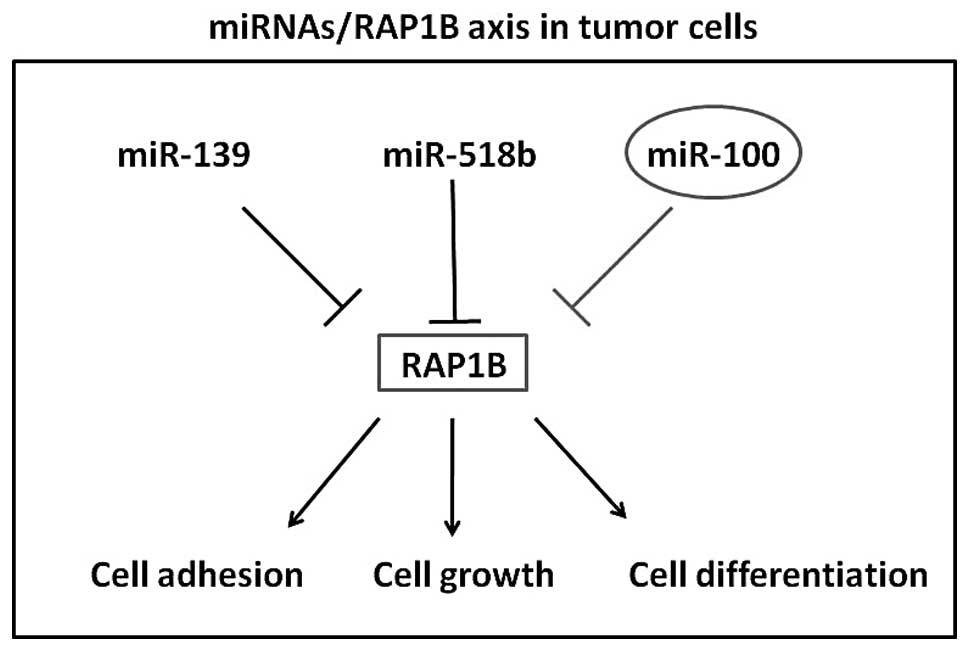
Collectively, the present study provides evidence
that miR-100 is downregulated in colorectal cancer tissues and that
it functions as tumor suppressor inhibiting cell proliferation,
invasion and promoting apoptosis. Moreover, its novel target gene,
RAP1B, was identified and found to be negatively expressed with
miR-100 in colorectal cancer tissues. Our results (Fig. 5) strongly support and supplement the
mechanism of miR/RAP1B axis in tumor cell growth and invasion.
Acknowledgements
This study was financially supported by research
grants from the National Natural Science Foundation of China (No.
1360337) and National Natural Science Foundation of Guangdong
Province (No. S2012010009082).
References
|
1
|
Jemal A, Siegel R, Ward E, Murray T, Xu J
and Thun MJ: Cancer statistics, 2007. CA Cancer J Clin. 57:43–66.
2007. View Article : Google Scholar
|
|
2
|
Cunningham D, Atkin W, Lenz HJ, et al:
Colorectal cancer. Lancet. 375:1030–1047. 2010. View Article : Google Scholar
|
|
3
|
Cho KR and Vogelstein B: Genetic
alterations in the adenoma - carcinoma sequence. Cancer. 70(Suppl
6): 1727–1731. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Fearon ER and Vogelstein B: A genetic
model for colorectal tumorigenesis. Cell. 61:759–767. 1990.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhang J, Guo H, Zhang H, et al: Putative
tumor suppressor miR-145 inhibits colon cancer cell growth by
targeting oncogene Friend leukemia virus integration 1 gene.
Cancer. 117:86–95. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lin Y, Wu J, Chen H, et al:
Cyclin-dependent kinase 4 is a novel target in micoRNA-195-mediated
cell cycle arrest in bladder cancer cells. FEBS Lett. 586:442–447.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Hutvagner G and Zamore PD: A microRNA in a
multiple-turnover RNAi enzyme complex. Science. 297:2056–2060.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ma Y, Zhang P, Yang J, Liu Z, Yang Z and
Qin H: Candidate microRNA biomarkers in human colorectal cancer:
systematic review profiling studies and experimental validation.
Int J Cancer. 130:2077–2087. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Liu J, Lu KH, Liu ZL, Sun M, De W and Wang
ZX: MicroRNA-100 is a potential molecular marker of non-small cell
lung cancer and functions as a tumor suppressor by targeting
polo-like kinase 1. BMC Cancer. 12:5192012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li XJ, Luo XQ, Han BW, Duan FT, Wei PP and
Chen YQ: MicroRNA-100/99a, deregulated in acute lymphoblastic
leukaemia, suppress proliferation and promote apoptosis by
regulating the FKBP51 and IGF1R/mTOR signalling pathways. Br J
Cancer. 109:2189–2198. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sun J, Chen Z, Tan X, et al:
MicroRNA-99a/100 promotes apoptosis by targeting mTOR in human
esophageal squamous cell carcinoma. Med Oncol. 30:4112013.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xu C, Zeng Q, Xu W, et al: miRNA-100
inhibits human bladder urothelial carcinogenesis by directly
targeting mTOR. Mol Cancer Ther. 12:207–219. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen P, Zhao X and Ma L: Downregulation of
microRNA-100 correlates with tumor progression and poor prognosis
in hepatocellular carcinoma. Mol Cell Biochem. 383:49–58. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gebeshuber CA and Martinez J: miR-100
suppresses IGF2 and inhibits breast tumorigenesis by interfering
with proliferation and survival signaling. Oncogene. 32:3306–3310.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang G, Chen L, Meng J, Chen M, Zhuang L
and Zhang L: Overexpression of microRNA-100 predicts an unfavorable
prognosis in renal cell carcinoma. Int Urol Nephrol. 45:373–379.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bernardi B, Guidetti GF, Campus F, et al:
The small GTPase Rap1b regulates the cross talk between platelet
integrin alpha2beta1 and integrin alphaIIbbeta3. Blood.
107:2728–2735. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Carmona G, Gottig S, Orlandi A, et al:
Role of the small GTPase Rap1 for integrin activity regulation in
endothelial cells and angiogenesis. Blood. 113:488–497. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Matsuse M, Mitsutake N, Rogounovitch T, et
al: Mutation analysis of RAP1 gene in papillary thyroid carcinomas.
Endocr J. 56:161–164. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Edreira MM, Li S, Hochbaum D, et al:
Phosphorylation-induced conformational changes in Rap1b: allosteric
effects on switch domains and effector loop. J Biol Chem.
284:27480–27486. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wittchen ES, Aghajanian A and Burridge K:
Isoform-specific differences between Rap1A and Rap1B GTPases in the
formation of endothelial cell junctions. Small GTPases. 2:65–76.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang M, Zhou S, Zhang L, et al: miR-518b
is down-regulated, and involved in cell proliferation and invasion
by targeting Rap1b in esophageal squamous cell carcinoma. FEBS
Lett. 586:3508–3521. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Guo H, Hu X, Ge S, Qian G and Zhang J:
Regulation of RAP1B by miR-139 suppresses human colorectal
carcinoma cell proliferation. Int J Biochem Cell Biol.
44:1465–1472. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hobert O: miRNAs play a tune. Cell.
131:22–24. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Peter ME: Targeting of mRNAs by multiple
miRNAs: the next step. Oncogene. 29:2161–2164. 2010. View Article : Google Scholar : PubMed/NCBI
|