Introduction
Colorectal cancer (CRC) is the third most frequently
diagnosed cancer worldwide with an increasingly high incidence rate
in developing countries (1,2). It is also the second leading cause of
cancer-related mortality with very poor prognosis and a high
possibility of metastasis (3,4). To
date, there is no ideal biomarker for CRC detection and therapy,
therefore, development of biomarkers for early detection and
therapy cannot be underrated, as they can help control progression
and mortality of CRC.
MicroRNAs (miRNAs) are a class of non-coding small
RNAs that can block the translation of mRNA targets and affect mRNA
stability (5), thereby regulating
protein-coding gene networks and pathways (6). Aberrant expressions of miRNAs have
been observed in breast (7), brain
(8), lung (9), thyroid (10,11)
and colon (11) cancer cells and
are tightly related to cancer development and progression,
including biological processes such as cellular proliferation,
differentiation, apoptosis and development of metastases (12–14).
miRNAs contribute to oncogenesis as either an oncogene (oncomiR) or
a tumor-suppressor (15). In CRCs,
miR-20a, miR-21, miR-25, miR-31, miR-93, miR-106, miR-183 and
miR-203 were upregulated, suggesting their oncogene functions,
whereas miR-1, miR-126, miR-30, miR-143, miR-145, miR-191 and
miR-192 were downregulated, indicating their tumor suppressor
functions (16). In CRCs, miRNAs
are found to play an important role in cancer risk assessment,
diagnosis, prognosis and drug response (17–19),
therefore, they can serve as potential biomarkers. However, to
date, only ~50 miRNAs have been studied in CRC or CRC cell lines
(16) and their functions are not
well understood. Furthermore, there are few clinical analyses for
the correlation between miRNAs and CRC progression.
MicroRNA-223 (miR-223) has been reported with
opposite functions in different types of cancer. It functions as an
oncomiR in some cancer types, such as recurrent vs. primary ovarian
(20) and bladder cancer (21), adenocarcinoma of esophagus (22) and metastatic gastric cancer
(23,24), whereas in acute lymphoblastic
leukemia (25), primary small cell
lung cancer and hepatocellular carcinoma (26–28),
miR-223 functions as a tumor suppressor. However, the function of
miR-223 in CRC has yet to be characterized.
In the present study, we used miRNA microarray to
compare the miRNA profiles in colon tumors and paired non-tumorous
tissues. We identified a promising candidate, miR-223, and
confirmed its expression pattern in CRC in 90 paired samples using
quantitative real-time PCR. Furthermore, inhibition of miR-223
suppressed colon cancer cell proliferation, migration and invasion.
Taken together, these results suggest that miR-223 plays a critical
role in CRC and can function as a potential molecular biomarker and
therapeutic gene for CRC treatment.
Materials and methods
Patients and samples
Pairs of human colon tumor and adjacent non-tumorous
tissues were obtained from 90 surgical patients from the Department
of Gastroenterology, Peking University Shenzhen Hospital. CRC
samples were collected from patients undergoing bowel resection.
Adjacent normal mucosa samples located at least 2 cm from the
margins of the tumor (polyp or carcinoma) were used as controls.
All tumors were adenocarcinomas. Cases with mucinous carcinomas
(when >50% of the tumor volume was composed of mucin) were
excluded from this study. Collected samples were flash frozen in
liquid nitrogen after surgery. All patients were informed about the
aims of the specimen collection and they provided written consent
in accordance with the ethics guidelines of Peking University. The
present study was approved by the Ethical Committee of Peking
University Shenzhen Hospital.
CRC cell culture and transfection
Human colorectal carcinoma cells LoVo and Colo320
were cultured in Dulbecco’s minimum essential medium (DMEM)
supplemented with 10% fetal bovine serum (FBS; Gibco). Artificial
miR-223 inhibitor (anti-miR-223) and random sequence anti-miR
molecule (anti-miR-NC) were synthesized by Ambion (Austin, TX,
USA). LoVo and Colo320 cells were seeded for 20 h incubation before
transfection followed by transfection with either chemically
synthesized anti-miR-223 or anti-miR-NC using Lipofectamine 2000
(Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s
instructions.
miRNA microarray
The miRNA microarray was performed at the Kangchen
Biotechnology company (Shanghai, China). The LNA-modified
oligonucleotide probes for all annotated miRNAs from human (Homo
sapiens) in miRBase version 11.0 were purchased from Exiqon
(miRCURY™ version 11.0). RNA labeling and hybridization on miRNA
microarray chips was performed as previously described (29). Scanning of microarray was performed
with an Axon GenePix 4000B microarray scanner. GenePix Pro V6.0 was
used for image processing and array data retrieval. The intensity
of green signal was calculated after background subtraction and
replicated spots on the same slide were averaged to get the median
intensity. The median normalization method was used to obtain
‘Normalized Data’. Normalized Data = (Foreground −
Background)/median. Median was 50 percent quantile of miRNA
intensity which was >50 in all samples after background
correction. Unsupervised hierarchical clustering was performed on
the miRNA expression profiling using cluster 3 and Treeview
software.
Quantitative real-time PCR
Colorectal tumor and adjacent normal mucosa total
RNA was extracted with TRIzol reagent (Invitrogen). The expression
of miR-223 was measured using TaqMan MicroRNA Assays kit (Ambion)
according to the manufacturer’s instructions. Relative expression
was calculated by the 2−ΔΔCt method and normalized to
the expression of RNU6B (Ambion).
Cell proliferation assays
The effect of miR-223 on the cell viability was
measured by WST-1 assay. Cells were counted and plated at a density
of 3×103 cells/well in 96-well plates in triplicates.
Cell viability was determined at 24, 48 and 72 h post-transfection.
Spectrophotometry was performed at λ=450 nm and λref=630
nm after incubation with 10 μl WST-1 (Roche, New York, NY, USA) for
2 h. Proliferation of cells was evaluated using colony formation
assay. For colony formation assays, cells were seeded in 6-well
plates (0.5×103 cells/well) and cultured for two weeks.
Colonies were fixed with methanol for 10 min and stained with 1%
crystal violet (Sigma) for 1 min. Visible colonies in 10 random
view fields were manually counted. Each cell group was measured in
triplicate.
Scratch wound-healing motility
assays
Cells were seeded and grown to confluence in a 60-mm
dish. An artificial wound was created 24 h following transfection
with either anti-miR-223 or anti-miR-NC using a 200-μl pipette tip
on the confluent cell monolayer. To visualize migrated cells and
wound healing, images were captured at 0 and 48 h. The distance
between the two edges of the scratch was measured using Digimizer
software system.
Transwell invasion assay
Cell invasion assay was performed using the Matrigel
Invasion Chamber (BD Biosciences, San Jose, CA, USA) pre-coated
with ECM gel (Sigma, St. Louis, MO, USA). Cells transfected with
either chemically synthesized anti-miR-223 or anti-miR-NC were
seeded into the upper chamber of serum-free medium. The lower
chamber prepared with 10% FBS served as a chemoattractant. After
24-h incubation, the non-invading cells were mechanically removed.
The invasive cells on the lower surface of the membrane were then
washed, fixed and stained with crystal violet. The number of cells
was counted in 10 randomly optical fields under a microscope at a
magnification of ×400.
Statistical analysis
Significance analysis of microarrays (SAM) was used
to identify miRNAs differentially expressed between samples
(FDR=0). miR-223 expressions in CRC tumors and adjacent non-tumor
tissues were compared by the Mann-Whitney U test and in the
evaluation between early and advanced stage tumors. A comparison of
means among two or more groups was performed using one-way analysis
of variance or the Student’s t-test. All numerical data were
expressed as means ± SD. P-value <0.05 was considered to
indicate a statistically significant result. Statistical analyses
were performed using GraphPad Prism 5.0 (GraphPad Software, San
Diego, CA, USA) and SPSS software (version 11).
Results
Expression profiling of miRNAs in human
CRC and adjacent non-tumor colorectal tissues
To investigate the miRNA expression patterns in
human CRC, we used miRNA microarrays to compare their expression in
human CRC and adjacent non-tumor colorectal tissues. Experiments
were conducted with miRNA isolated from three paired normal and
tumor specimens. Microarray expression profiling revealed a series
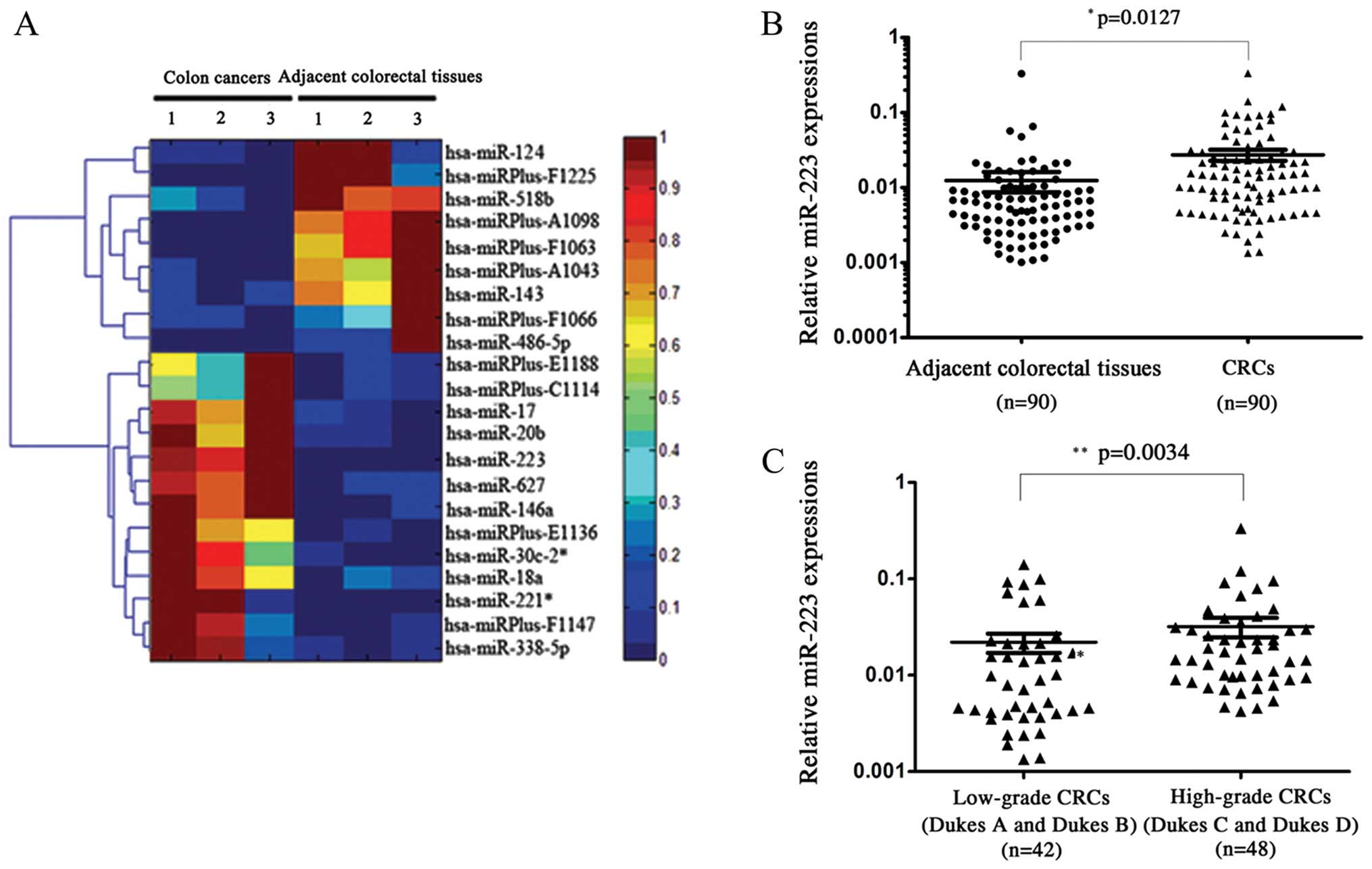
of miRNAs that showed altered expressions in CRC. Scatter plot and
hierarchical clustering analysis are shown in Fig. 1A. Compared with adjacent non-tumor
tissues, 13 miRNAs were upregulated for at least 2-fold, and 9
miRNAs were downregulated for at least 2-fold in CRCs. Upregulated
miRNAs were miR-338-5p, miRPlus-C1114, miR-146a, miR-18a, miR-20b,
miR-30c-2*, miRPlus-F1147, miR-17, miRPlus-E1136, miRPlus-E1188,
miR-627, miR-221* and miR-223. Downregulated miRNAs were
miR-486-5p, miRPlus-F1225, miR-124, miR-143, miR-518b,
miRPlus-A1043, miRPlus-F1063, miRPlus-A1098 and miRPlus-F1066.
miR-223 is upregulated in human CRC
Since an interesting miRNA, miR-223, has been
reported to be overexpressed in metastatic gastric cancer cells
(30), we sought to analyze the
function of miR-223 in CRC. To further confirm the upregulation of
miR-223 in CRC, we performed quantitative real-time PCR in 90
paired CRC samples. The overexpression of miR-223 in CRC was
identified with a significant P-value (P=0.0127; Fig. 1B). Furthermore, compared with CRC
tissues from patients with low-grade CRC (Dukes A and B),
high-grade (Dukes C and D) patient tissues presented an even higher
miR-223 (P=0.0034; Fig. 1C). These
results suggest that miR-223 plays an important role in CRC
progression and functions as a potential biomarker for early
screening and diagnosis.
Suppressing miR-223 expression reduces
CRC cell proliferation and viability
To further test the function of miR-223 in CRC
progression, we use chemically synthesized miR-223 inhibitor
(anti-miR-223) to downregulate miR-223 expression in human CRC cell
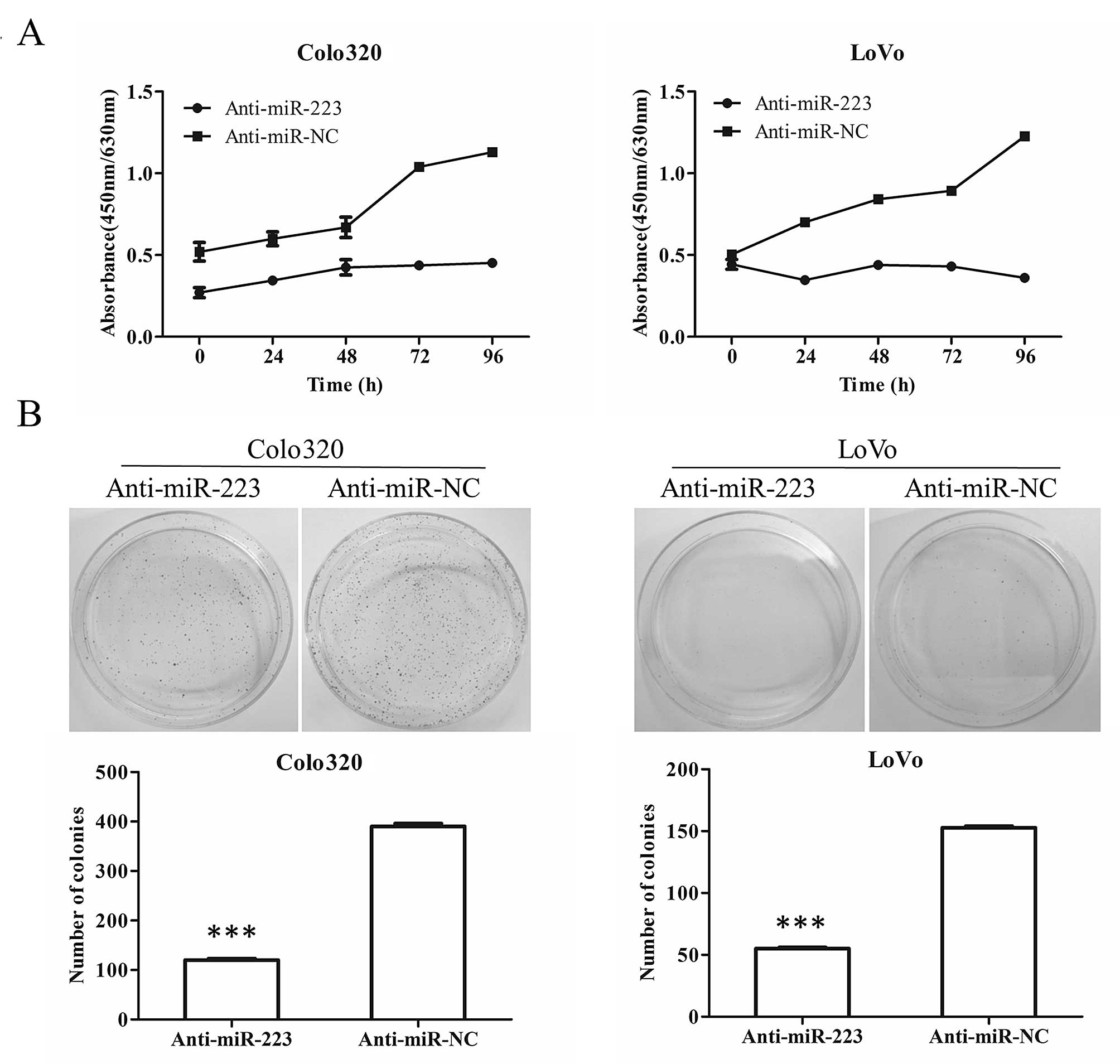
lines, Colo320 and LoVo. Over 96 h after transfection of
anti-miR-223 in both Colo320 and LoVo cells, the WST-1 absorbance
at 450 nm subtracting background absorbance at 630 nm did not
increase and almost remained unchanged whereas over 2-fold increase
was observed in the anti-miR-NC transfected cells (Fig. 2A), suggesting a decreased
proliferation and viability upon suppression of miR-223. Consistent
with the results in WST-1 assay, cells transfected with
anti-miR-223 showed significantly fewer colonies than cells
transfected with anti-miR-NC (P<0.001) (Fig. 2B).
Suppression of miR-223 inhibits migration
and invasion of CRC cells
Since miR-223 has a higher level of tissues from
high grade patients (Duke C and D) (Fig. 1C), suggesting a higher chance of
metastasis, we used scratch wound healing assay and Transwell assay
to evaluate cell migration and invasion ability.
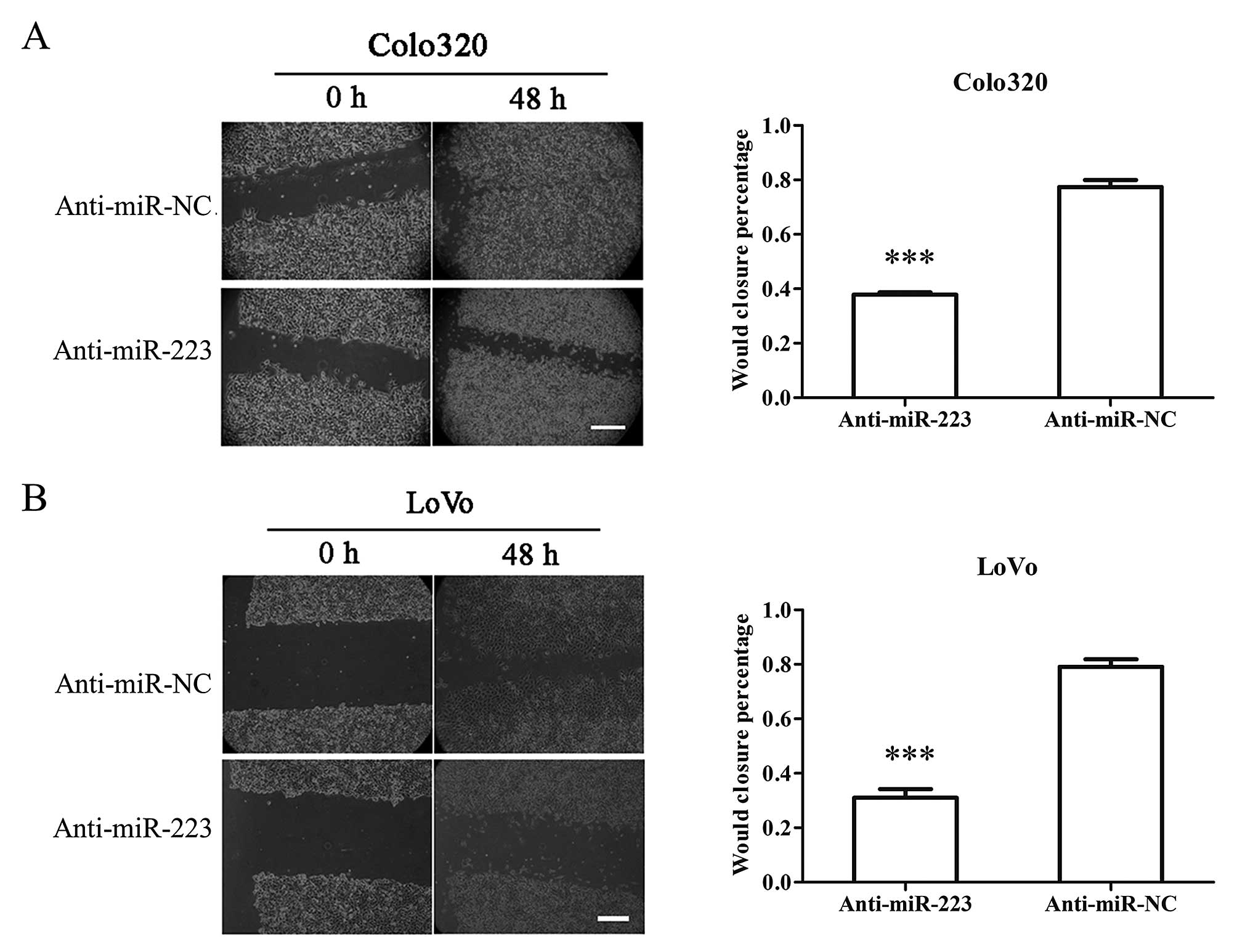
Upon suppression of miR-223, both Colo320 and LoVo
cells showed largely decreased motility and could not efficiently
close the wound as anti-miR-NC transfected cells did over 48 h
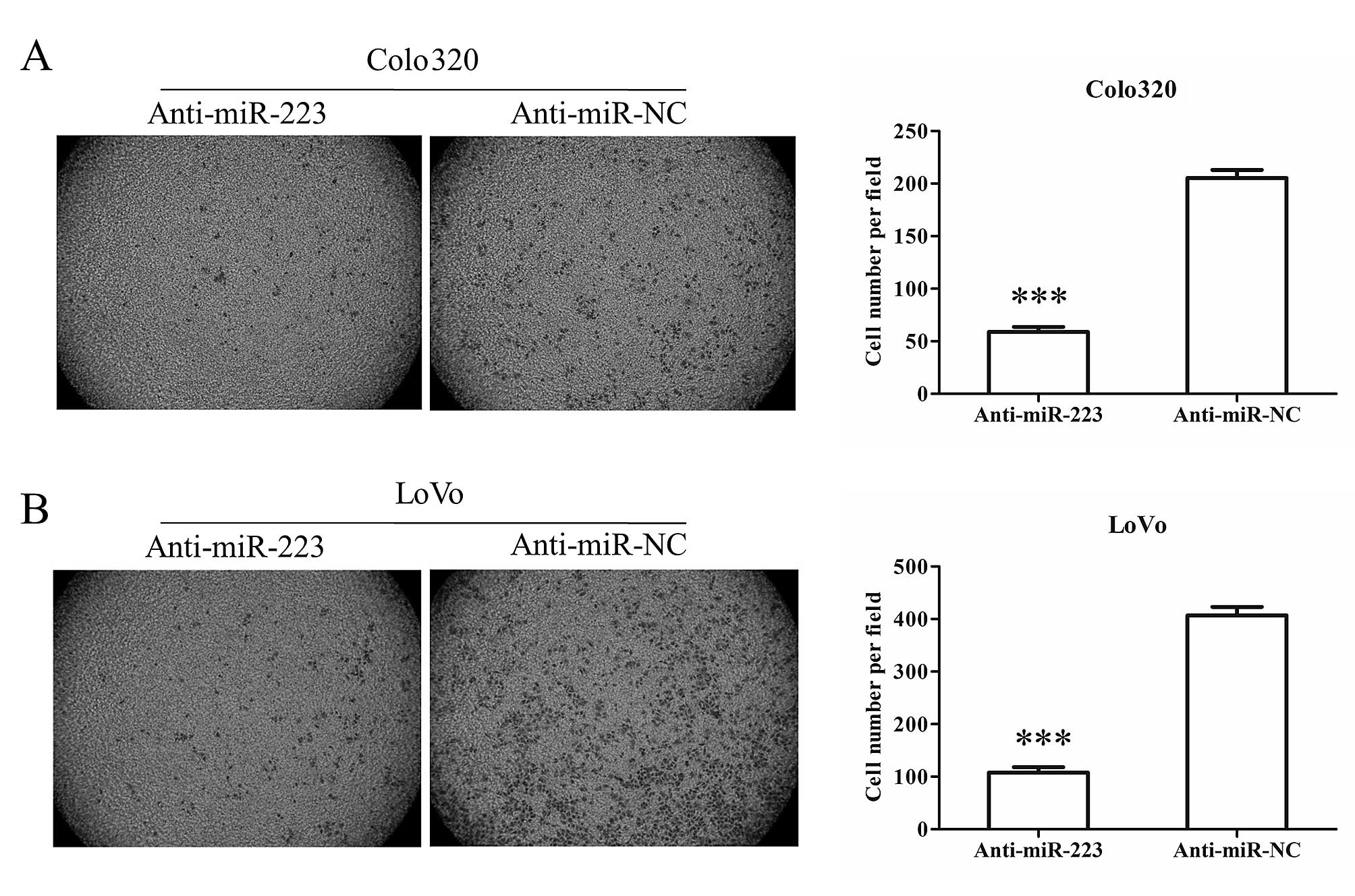
(Fig. 3). In addition to motility,
after transfection of anti-miR-223, both Colo320 and LoVo cells
showed less invasion ability through Matrigel coated Transwell
compared with anti-miR-NC transfected cells (Fig. 4). These results suggest that
suppression of miR-223 inhibits colon cancer cell migration and
invasion.
Discussion
At present, although with a high mortality rate,
upon detection of preneoplastic lesions with modern screening
methods, the mortality and morbidity of CRC have modestly decreased
and it has become a preventable disease (31). Yet, it is still insufficient to use
present clinical and pathological parameters for early diagnosis or
personalized treatment. As assessment of the microRNA (miRNA)
genetic profile of colorectal cancer (CRC) and increasingly more
miRNAs have been identified and characterized, they could serve as
potential clinical biomarkers (14,32).
The present study, for the first time, identified
miR-223 and verified its oncogenetic function in CRC tissue. Using
miRNA array and quantitative real-time PCR, we identified miR-223
as a highly upregulated miRNA in CRC compared to adjacent
non-tumorous tissue (Fig. 1A and
B). In addition, in tissues from higher grade (Duke C and D)
patients, miR-223 levels were significantly higher than from lower
grade (Duke A and B) patients (Fig.
1C). These findings led us to explore the functional analysis
of miR-223. Downregulation of miR-223 was able to significantly
suppress proliferation (Fig. 2),
migration (Fig. 3) and invasion
(Fig. 4) of colon cancer cells.
Other than miR-223, several other miRNAs with
significant different expression between CRC and normal tissue were
also identified by miRNA array (Fig.
1). Consistent with previous reports, the expression of miR-17,
miR-18 and miR-20 was significantly higher in CRC (33–35).
Functional analysis showed miR-17 increases proliferation, invasion
and migration of colon cancer cells (34,35).
On the other hand, miR-143 is downregulated in CRC tissue, and
functions as an inhibitor for the cancer progression of CRC through
multiple targets (33,35–39).
By identifying these miRNAs, we could use miR-223
and other miRNAs as clinical biomarkers either by themselves or in
combination with each other. It is possible that combining
different miRNAs will give a more accurate clinical diagnosis.
Acknowledgements
We thank our colleagues for their insight and
technical support. This study was supported by the National Natural
Scientific Foundation of China (grant no. 81171447), and the
Natural Science Foundation of Guangdong Province, China (grant no.
8104518036002006310).
References
|
1
|
Siegel R, Ward E, Brawley O and Jemal A:
Cancer statistics, 2011: the impact of eliminating socioeconomic
and racial disparities on premature cancer deaths. CA Cancer J
Clin. 61:212–236. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Center MM, Jemal A and Ward E:
International trends in colorectal cancer incidence rates. Cancer
Epidemiol Biomarkers Prev. 18:1688–1694. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ferlay J, Shin HR, Bray F, Forman D,
Mathers C and Parkin DM: Estimates of worldwide burden of cancer in
2008: GLOBOCAN 2008. Int J Cancer. 127:2893–2917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Edwards BK, Ward E, Kohler BA, et al:
Annual report to the nation on the status of cancer, 1975–2006,
featuring colorectal cancer trends and impact of interventions
(risk factors, screening, and treatment) to reduce future rates.
Cancer. 116:544–573. 2010.
|
|
5
|
Winter J, Jung S, Keller S, Gregory RI and
Diederichs S: Many roads to maturity: microRNA biogenesis pathways
and their regulation. Nat Cell Biol. 11:228–234. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bartel DP: MicroRNAs: genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Qu J, Zhao L, Zhang P, et al: MicroRNA-195
chemosensitizes colon cancer cells to the chemotherapeutic drug
doxorubicin by targeting the first binding site of BCL2L2 mRNA. J
Cell Physiol. View Article : Google Scholar : 2013.[Epub ahead of
print].
|
|
8
|
Catania A, Maira F, Skarmoutsou E, D’Amico
F, Abounader R and Mazzarino MC: Insight into the role of microRNAs
in brain tumors (Review). Int J Oncol. 40:605–624. 2012.PubMed/NCBI
|
|
9
|
Cherni I and Weiss GJ: miRNAs in lung
cancer: large roles for small players. Future Oncol. 7:1045–1055.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Le Pennec S and Savagner F: MiRNAs in
follicular thyroid tumors. Presse Med. 40:683–689. 2011.(In
French).
|
|
11
|
Agostini M, Pucciarelli S, Calore F, Bedin
C, Enzo M and Nitti D: miRNAs in colon and rectal cancer: a
consensus for their true clinical value. Clin Chim Acta.
411:1181–1186. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Liu X, Zhang Z, Sun L, et al:
microRNA-499-5p promotes cellular invasion and tumor metastasis in
colorectal cancer by targeting FOXO4 and PDCD4. Carcinogenesis.
32:1798–1805. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
He L and Hannon GJ: MicroRNAs: small RNAs
with a big role in gene regulation. Nat Rev Genet. 5:522–531. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Croce CM: Causes and consequences of
microRNA dysregulation in cancer. Nat Rev Genet. 10:704–714. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lu J, Getz G, Miska EA, et al: MicroRNA
expression profiles classify human cancers. Nature. 435:834–838.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Corte H, Manceau G, Blons H and
Laurent-Puig P: MicroRNA and colorectal cancer. Dig Liver Dis.
44:195–200. 2012. View Article : Google Scholar
|
|
17
|
Schetter AJ, Leung SY, Sohn JJ, et al:
MicroRNA expression profiles associated with prognosis and
therapeutic outcome in colon adenocarcinoma. JAMA. 299:425–436.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Boni V, Zarate R, Villa JC, et al: Role of
primary miRNA polymorphic variants in metastatic colon cancer
patients treated with 5-fluorouracil and irinotecan.
Pharmacogenomics J. 11:429–436. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Landi D, Gemignani F, Naccarati A, et al:
Polymorphisms within micro-RNA-binding sites and risk of sporadic
colorectal cancer. Carcinogenesis. 29:579–584. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Laios A, O’Toole S, Flavin R, et al:
Potential role of miR-9 and miR-223 in recurrent ovarian cancer.
Mol Cancer. 7:352008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Gottardo F, Liu CG, Ferracin M, et al:
Micro-RNA profiling in kidney and bladder cancers. Urol Oncol.
25:387–392. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Mathé EA, Nguyen GH, Bowman ED, et al:
MicroRNA expression in squamous cell carcinoma and adenocarcinoma
of the esophagus: associations with survival. Clin Cancer Res.
15:6192–6200. 2009.PubMed/NCBI
|
|
23
|
Li X, Zhang Y, Zhang H, et al: miRNA-223
promotes gastric cancer invasion and metastasis by targeting tumor
suppressor EPB41L3. Mol Cancer Res. 9:824–833. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kang W, Tong JH, Chan AW, et al: Stathmin1
plays oncogenic role and is a target of microRNA-223 in gastric
cancer. PLoS One. 7:e339192012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Mi S, Lu J, Sun M, et al: MicroRNA
expression signatures accurately discriminate acute lymphoblastic
leukemia from acute myeloid leukemia. Proc Natl Acad Sci USA.
104:19971–19976. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Miko E, Czimmerer Z, Csánky E, et al:
Differentially expressed microRNAs in small cell lung cancer. Exp
Lung Res. 35:646–664. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Karakatsanis A, Papaconstantinou I,
Gazouli M, Lyberopoulou A, Polymeneas G and Voros D: Expression of
microRNAs, miR-21, miR-31, miR-122, miR-145, miR-146a, miR-200c,
miR-221, miR-222, and miR-223 in patients with hepatocellular
carcinoma or intrahepatic cholangiocarcinoma and its prognostic
significance. Mol Carcinog. 52:297–303. 2013. View Article : Google Scholar
|
|
28
|
Wong QW, Lung RW, Law PT, et al:
MicroRNA-223 is commonly repressed in hepatocellular
carcinoma and potentiates expression of Stathmin1.
Gastroenterology. 135:257–269. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sempere LF, Christensen M, Silahtaroglu A,
et al: Altered MicroRNA expression confined to specific epithelial
cell subpopulations in breast cancer. Cancer Res. 67:11612–11620.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Xing J, Wan S, Zhou F, et al: Genetic
polymorphisms in pre-microRNA genes as prognostic markers of
colorectal cancer. Cancer Epidemiol Biomarkers Prev. 21:217–227.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Lieberman DA: Clinical practice. Screening
for colorectal cancer. N Engl J Med. 361:1179–1187. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Iorio MV and Croce CM: MicroRNA
dysregulation in cancer: diagnostics, monitoring and therapeutics.
A comprehensive review. EMBO Mol Med. 4:143–159. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Motoyama K, Inoue H, Takatsuno Y, et al:
Over- and under-expressed microRNAs in human colorectal cancer. Int
J Oncol. 34:1069–1075. 2009.PubMed/NCBI
|
|
34
|
Zhang J, Xiao Z, Lai D, et al: miR-21,
miR-17 and miR-19a induced by phosphatase of regenerating liver-3
promote the proliferation and metastasis of colon cancer. Br J
Cancer. 107:352–359. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Monzo M, Navarro A, Bandres E, et al:
Overlapping expression of microRNAs in human embryonic colon and
colorectal cancer. Cell Res. 18:823–833. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Liu H, Zhu L, Liu B, et al: Genome-wide
microRNA profiles identify miR-378 as a serum biomarker for early
detection of gastric cancer. Cancer Lett. 316:196–203. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Slaby O, Svoboda M, Fabian P, et al:
Altered expression of miR-21, miR-31, miR-143 and miR-145 is
related to clinicopathologic features of colorectal cancer.
Oncology. 72:397–402. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ng EK, Tsang WP, Ng SS, et al:
MicroRNA-143 targets DNA methyltransferases 3A in colorectal
cancer. Br J Cancer. 101:699–706. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Chen X, Guo X, Zhang H, et al: Role of
miR-143 targeting KRAS in colorectal tumorigenesis. Oncogene.
28:1385–1392. 2009. View Article : Google Scholar : PubMed/NCBI
|