Introduction
Epithelial ovarian cancer remains the fifth most
common cancer in women worldwide and the leading cause of mortality
for female reproductive system tumors (1). The early symptom of ovarian cancer is
obscure and reliable early diagnosis for ovarian cancer is lacking.
Although the survival rates of advanced cancer patients have been
improved over the past decades, such improvements in ovarian cancer
have only been observed in the 5-year survival rate for women with
stage III and IV tumors who are <30 years old (1,2).
Therefore, developing reliable biomarkers for the early diagnosis
for ovarian cancer is crucial.
Common diagnostic methods for ovarian cancer include
transvaginal ultrasonography, bimanual rectovaginal pelvic
examination and the serum level of CA125, an ovarian-related
antigenic determinant. However, routine gynecologic examinations
lack specificity and sensitivity (3,4).
Moreover, although the CA125 serum level is elevated in 80% of
patients with epithelial ovarian cancer, the use of this marker for
the early detection of ovarian cancer is limited, since only 50% of
patients with stage I ovarian cancer show elevated CA125 levels. In
addition, the CA125 serum level is also increased in a significant
number of women with benign ovarian lesions as well as in healthy
women, which limits its specificity as a diagnostic biomarker
(5–7). As yet, no useful biomarker has been
identified for ovarian cancer in urine, probably since the majority
of efforts have focused on blood protein biomarkers, which are
hardly detected in the urine.
MicroRNAs (miRNAs) are small (18–24 nucleotides),
highly conserved, non-coding RNAs that regulate gene expression
post-transcriptionally (8).
Computational studies suggest that >60% of mammalian gene
transcripts are regulated by miRNAs (9). miRNAs play a key regulatory role in
divergent cell processes, such as proliferation, differentiation,
apoptosis, migration, as well as tumorigenesis (10,11).
miRNAs are aberrantly expressed in human cancer, and the miRNA
profiling of poorly differentiated tumors was more accurate than
the mRNA-profiling, suggesting a potential clinical value of miRNAs
in the diagnosis of cancer (12).
On a technical level, urine samples are easier to
obtain and contain fewer proteins than blood-based samples,
reducing the interference of proteins during RNA extraction and the
subsequent analysis. In addition to miRNAs that may be excreted
locally by the urinary system, urine miRNAs may be derived from the
circulating blood via renal elimination (13,14). A
number of miRNAs have been identified in the urine (15–17),
suggesting the potential utility of urine miRNAs for identifying or
qualifying ovarian cancer.
In the present study, we aimed to examine the roles
of urine miRNAs in the diagnosis of ovarian serous adenocarcinoma.
In addition, we preliminarily explored the underlying mechanisms
involved.
Materials and methods
Ethics statement
The present study was approved by the Clinical
Research Ethics Committee of Central South University, and the EC
(S) ID no. 201406374. In addition, the patients involved in the
present study provided written informed consent.
Urine and tissue samples
Morning whole-stream urine samples were collected at
the time of initial consultation prior to therapy at the Xiangya
Hospital, Central South University (Changsha, China). As shown in
Table I, urine samples were
collected from 39 ovarian serous adenocarcinoma patients, 26
patients with benign gynecological disease and 30 age-matched
healthy women used as normal controls. Women with benign
gynecologic diseases had a variety of pathological diagnoses,
including benign ovarian cyst adenoma, uterine fibroids,
endometriosis and physiological ovarian cysts (Table I). Urine samples from 20 gastric
cancer and 20 colon carcinoma patients were also collected
(Table I). Moreover, a total of 51
tissue specimens were collected, including 21 benign gynecologic
diseases specimens and 30 ovarian serous adenocarcinoma tissues
(Table I).
 | Table ISummary description of study
subjects. |
Table I
Summary description of study
subjects.
| Variables | Urine miRNA
microarray (n) | Urine RT-qPCR
(n) | Tissue RT-qPCR
(n) | Urine total
(n) |
|---|
| Healthy woman | 5 | 25 | 20 | 30 |
| Benign gynecologic
disease | 0 | 26a | 21d | 26a |
| Ovarian serous
adenocarcinoma | 5f | 34e | 30e | 39b |
| Gastric cancer | 0 | 20c | 0 | 20c |
| Colon
carcinoma | 0 | 20c | 0 | 20c |
Cell lines
The SKOV-3, SKOV-3ip, HO-8910, HO-8910pm human
ovarian carcinoma cell lines and the 293-T and U-937 non-ovarian
cell lines were obtained from the American Type Culture Collection
(ATCC; Manassas, VA, USA). The normal OSE ovarian surface
epithelium cell lines was used as a non-cancerous control in the
present study. The cell lines were maintained at 37°C in a
humidified atmosphere of 5% CO2. OSE, SKOV-3, SKOV-3ip
and HO-8910 were cultured in RPMI-1640 supplemented with 10% fetal
bovine serum (FBS) (both from Life Technologies, Carlsbad, CA,
USA). HO-8910pm, 293-T and U-937 cell lines were cultured in
Dulbecco’s modified Eagle’s medium (DMEM; Life Technologies) with
10% FBS.
Preparation of RNA
The mirVana miRNA Isolation kit (Ambion, Austin, TX,
USA) was used for the extraction of total RNA from the urine sample
cell supernatants and urine exosomes according to the
manufacturer’s instructions. For each subject, 600 μl of
urine was used for miRNA extraction. Total RNA content was
extracted from tissues and cells using TRIzol reagent (Ambion).
miRNA microarray analysis
miRNA microarray was performed at the Beijing
CapitalBio Corporation. miRNA (200 ng) isolated from each urine
sample was labeled with cyanine 3-pCp using Agilent miRNA labeling
and hybridization kits, hybridized to the Agilent Human miRNA
Microarray (miRBase release 19.0), and then scanned. The
fluorescence intensities were converted into digital data and Log2
transformed using Feature Extraction (version 10.7). Genes that
were differentially expressed between ovarian cancer patients and
healthy controls were analyzed using GeneSpring software version
12.0 (Agilent). miRNAs that fulfilled the criteria of a P-value (%)
<0.05 and a fold-change ≥2 between the groups were considered to
be significantly different. Heat maps were generated using the
Cluster 3.0 package software. Our data was uploaded in the Gene
Expression Omnibus (GEO) of the National Center for Biotechnology
Information (NCBI) (accession no. GSE58517).
Supervised clustering analysis of
microarray miRNAs
Prior to hierarchical clustering, miRNA profiles
were standardized to a mean of zero and a standard deviation of
1.0. The clustering analysis was performed with average linkage and
the Pearson’s correlation. Briefly, the purpose of this algorithm
is to assemble all elements into a single tree by calculating a
dendrogram. For any set of miRNA genes, an upper-diagonal
similarity matrix was computed. All the pairs of genes were
assigned similarity scores. The highest value was scanned to
identify the matrix (representing the most similar miRNAs). The
paired genes formed a node that was used to compute a gene
expression profile for the joined elements by averaging
observation: after the missing values were omitted, the gene
expression of the two joined elements was determined by the number
of miRNAs they contained.
Reverse transcription-quantitative
real-time polymerase chain reaction
The TaqMan microRNA reverse transcription kit
(Applied Biosystems, Foster City, CA, USA) and High Capacity cDNA
Reverse Transcription kit (Applied Biosystems) were used for
reverse transcription. For miRNA amplification, 5 μl total
RNA was mixed with 0.15 μl of 100 mm dNTPs (with dTTP), 1
μl (50 units) MultiScribe reverse transcriptase, 1.5
μl of 10X reverse transcription buffer, 0.19 μl RNase
inhibitor (20 U/μl), and 3 μl 1X gene-specific
primers. Nuclease-free H2O was added to obtain a final
volume of 15 μl. Reverse transcription was performed at 16°C
for 30 min, followed by an incubation step at 42°C for 30 min and
enzyme inactivation at 85°C for 5 min. The resulting cDNA was
stored at −20°C until use. The cDNA was quantified by reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
using the ABI PRISM 7500 Sequence Detection System (Applied
Biosystems). For RT-qPCR, 0.33 μl cDNA solution, 10
μl of 1X TaqMan universal PCR master mix, 2 μl of 1X
gene specific primer/probe set (all from Applied Biosystems) and
7.67 μl RNase-free H2O were mixed to obtain a
final reaction volume of 20 μl. Each sample was run in
triplicate. The RT-qPCR steps were 50°C for 2 min, 95°C for 10 min
and 40 cycles of denaturation at 95°C for 15 sec, followed by an
annealing/elongation step at 60°C for 60 sec. The miR-30a-5p and
miR-6076 were detected in triplicate via RT-qPCR, and RNU6 (Applied
Biosystems) was used as the housekeeping gene to normalize the
miRNA expression in the urine. The data were analyzed with the SDS
relative quantification software version 2.2.2 (Applied
Biosystems). The same baseline and cycle threshold (CT) were set
for each target. The 2−ΔΔCt method was used for relative
quantization of the differences of the expression levels of each
target in the urine samples. The absolute amount of cell
supernatant and urine exosome miR-30a-5p was calculated by software
based on the sample RT-qPCR numbers and the standard curve, and
were expressed as pmol/l.
Exosome isolation from the cell
supernatant or urine
Exosomes from the cell supernatant and urine were
isolated by ultracentrifugation. Briefly, the urine was centrifuged
at 2,000 × g for 20 min to eliminate the cells and debris and then
at 13,500 × g for 20 min, followed by ultracentrifugation (Beckman
L-80XP 70Ti) at 200,000 × g for 1 h to pellet the exosomes. The
exosome pellets were resuspended in phosphate-buffered saline (PBS)
prior to experimentation.
Electron microscopy of exosomes
Exosomes were isolated from the cell supernatant and
urine. The exosomes were washed in PBS, filtered and
ultracentrifugation was performed at 200,000 × g for 1 h to
re-pellet the exosomes. The exosome pellet was resuspended and
fixed in PBS containing 2% glutaraldehyde and then loaded onto
formvar/carbon-coated electron microscopy grids. The samples were
contrasted with uranyl acetate to visualize the membrane and viewed
with a FEI Tecnai G2 Spirit electron microscope.
Transfection
Lipofectamine 2000 (Life Technologies) was used to
perform cell transfection following the manufacturer’s
instructions. Briefly, ovarian cancer cells were transfected with
scramble miRNA (NC) or miR-30a-5p inhibitor (Nlunbio Co., Changsha,
China).
Cell proliferation assay
The MTT assay was used to measure the cell
proliferation of ovarian cancer cells transfected with scramble
miRNA (NC) or miR-30a-5p inhibitor. At 48 h post-transfection, the
transfection medium in each well was replaced with 100 μl
fresh serum-free medium containing 0.5 g/l MTT. Subsequent to
incubation at 37°C for 4 h, the MTT medium was removed by
aspiration and 50 μl dimeth-ylsulfoxide was added to each
well. Following incubation at 37°C for an additional 10 min, the
optical density at 570 nm was measured using the Bio-Tek™ ELx800™
absorbance microplate reader (Bio-Tek, Winooski, VT, USA). This
experiment was repeated three times.
Cell migration assay
A wound-healing assay was performed to evaluate the
cell migratory capacity of ovarian cancer cells transfected with
scramble miR (NC) or miR-30a-5p inhibitor. Briefly, the cells were
cultured to full confluence. Wounds of ~1 mm width were created
with a plastic scriber, and the cells were washed and incubated in
serum-free medium. After wounding for 24 h, the cells were
incubated in a medium including 10% FBS. After further culturing
for 0 and 48 h, the cells were fixed and observed under a
microscope.
Statistical analysis
Values are presented as the mean ± SD. Statistical
analysis was performed using SPSS 16.0 (SPSS, Inc., IBM, Armonk,
NY, USA). An unpaired Student’s t-test was used to compare the
miRNA expression in different groups. P<0.05 indicated
significant differences.
Results
Microarray identification of
differentially expressed miRNAs in urine samples from patients with
ovarian serous adenocarcinoma
To identify differentially expressed miRNAs in urine
from ovarian serous adenocarcinoma patients, urine samples were
collected from 5 ovarian serous adenocarcinoma patients and 5
age-matched healthy female control subjects. miRNA expression
profiling was conducted using the Agilent Human miRNA Microarray
(miRBase release 19.0), which contains probes for 2,006 human
miRNAs. miRNAs with a mean change >2.0-fold and P<0.05
between ovarian cancer and healthy control samples were considered
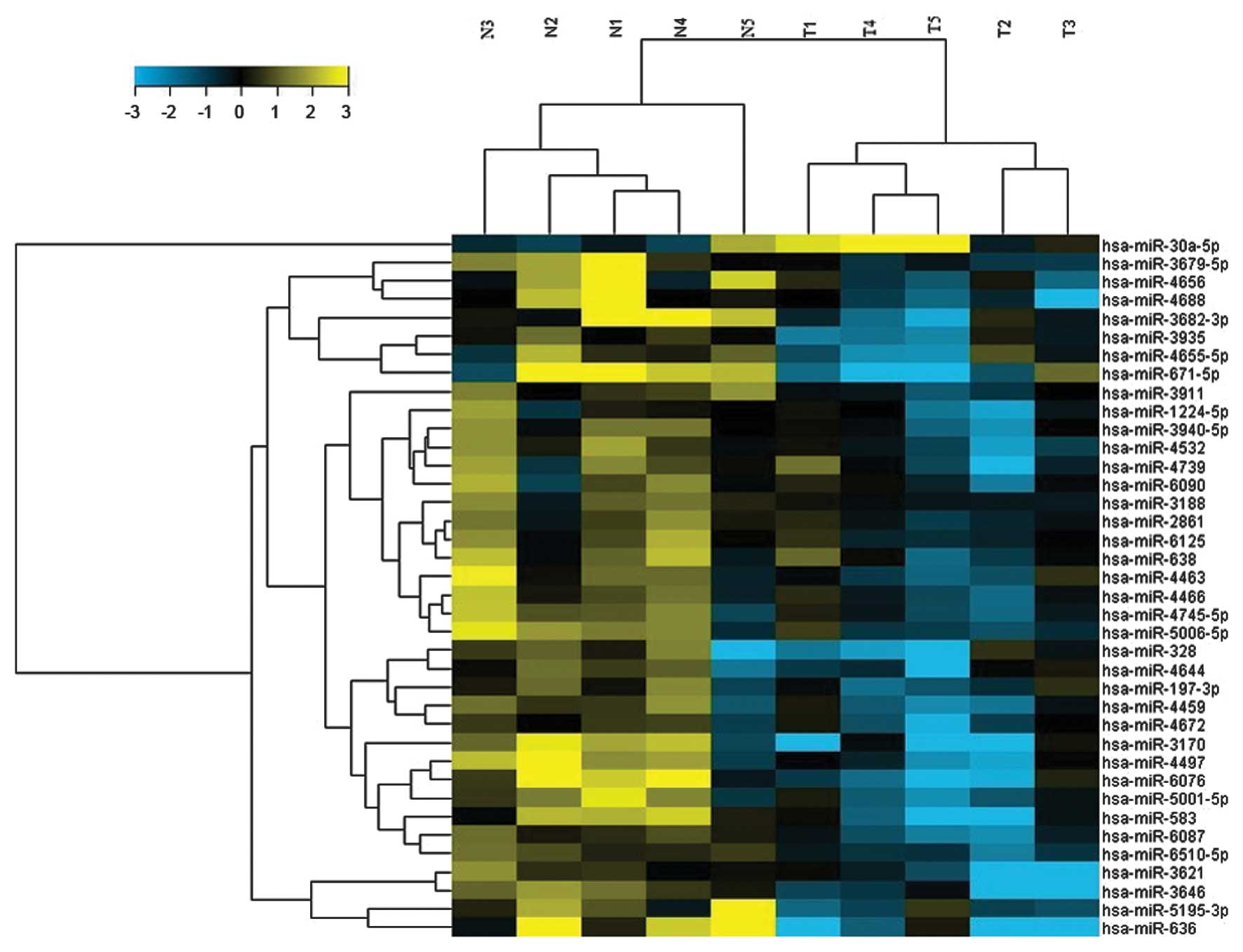
significantly different. As shown in Fig. 1 and Table II, one miRNA was upregulated and 37
miRNAs were downregulated in the ovarian serous adenocarcinoma
samples compared to the healthy controls. To validate the
microarray analysis, we compared the relative level of
representative miRNAs in the urine samples from an independent set
of 34 ovarian serous adenocarcinoma patients, 26 patients with
benign gynecological disease and 25 healthy controls. The
upregulated miRNA identified by microarray analysis (miR-30a-5p)
and a representative downregulated miRNA (miR-6076) were assessed.
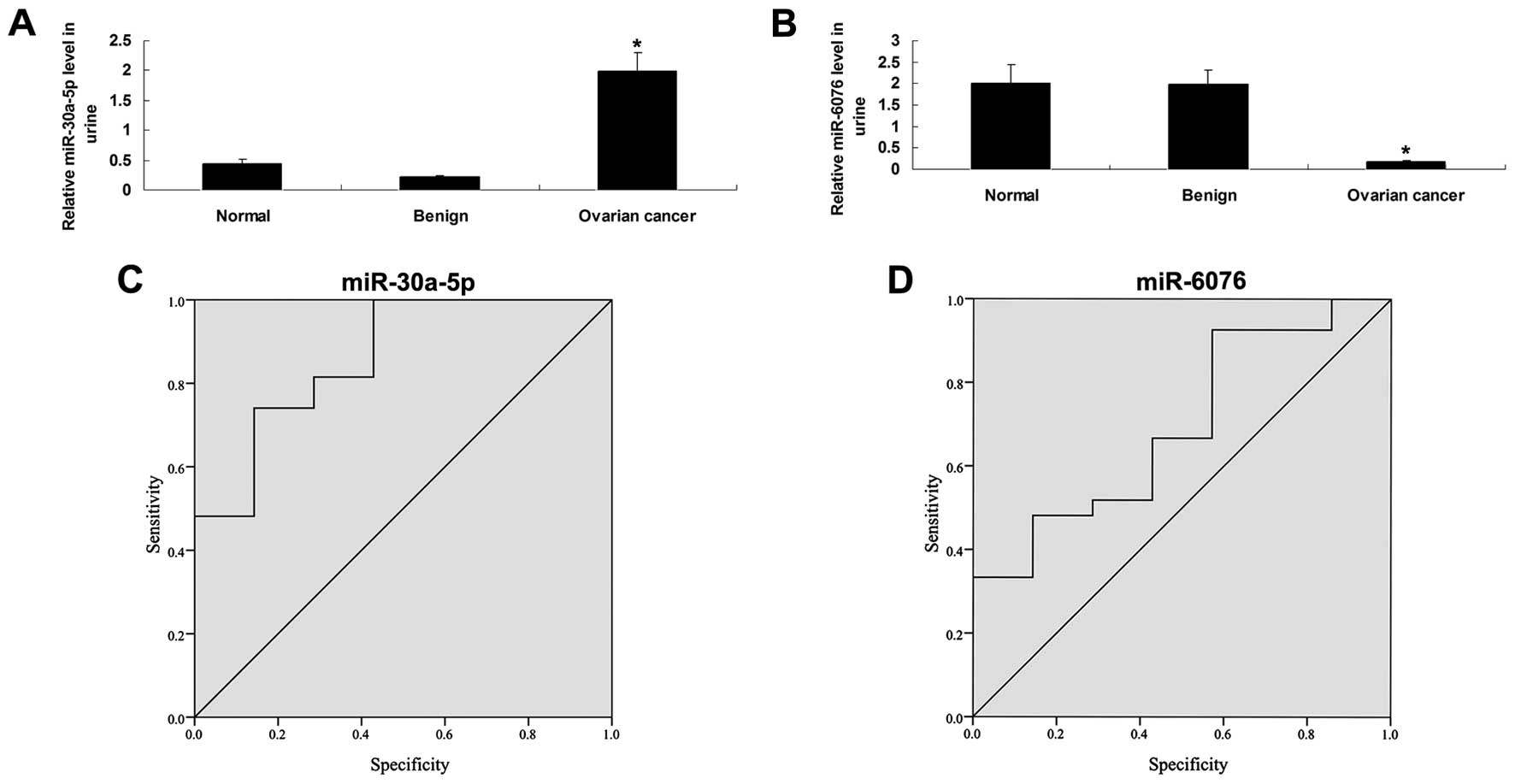
As shown in Fig. 2A, the relative
miR-30a-5p level in ovarian serous adenocarcinoma specimens was
4.6-fold higher when compared with the healthy controls
(P<0.05), and 9.0-fold higher when compared with the benign
ovarian specimens (P<0.05). Additionally, the relative
miR-30a-5p level in the benign ovarian specimens was lower than
that in the healthy controls, although this difference was not
statistically significant (P>0.05; Fig. 2A). Moreover, the results suggested
that the upregulation of urinary miR-30a-5p was closely associated
with the early stage of ovarian serous adenocarcinoma, since its
expression level was higher in stage I-II ovarian serous
adenocarcinoma samples than that in stage III-IV samples
(P<0.01), and higher in well or well-moderately-differentiated
ovarian serous adenocarcinoma samples than in poorly-differentiated
samples (P<0.05, Table III).
The results suggested that the upregulation of urinary miR-30a-5p
was associated with lymphatic metastasis (P<0.01, Table III).
 | Table IIDifferently expressed miRNAS in the
urine of ovarian serous adenocarcinoma patients vs. healthy control
subjects. |
Table II
Differently expressed miRNAS in the
urine of ovarian serous adenocarcinoma patients vs. healthy control
subjects.
| miRNA | P-value | Fold-change |
|---|
| Upregulated |
|
hsa-miRNA-30a-5p | 0.02159635 | 4.668867 |
| Downregulated |
| hsa-miR-636 | 0.0202967 | 64.08348 |
| hsa-miR-3170 | 0.0248452 | 52.002758 |
|
hsa-miR-671-5p | 0.0176856 | 20.946095 |
| hsa-miR-583 | 0.0356045 | 14.240913 |
| hsa-miR-6076 | 0.0146871 | 12.56148 |
|
hsa-miR-3682-3p | 0.0465591 | 11.362884 |
| hsa-miR-4688 | 0.0284103 | 10.914472 |
| hsa-miR-3646 | 0.0430764 | 10.0874815 |
| hsa-miR-3621 | 0.0498169 | 8.299675 |
| hsa-miR-4497 | 0.0360293 | 6.8510914 |
|
hsa-miR-5195-3p | 0.0154927 | 5.6532087 |
| hsa-miR-4656 | 0.029555 | 4.497623 |
|
hsa-miR-5001-5p | 0.0007499 | 4.3971524 |
|
hsa-miR-3679-5p | 0.0225566 | 4.358498 |
|
hsa-miR-5006-5p | 0.0363791 | 3.9775543 |
| hsa-miR-6087 | 0.0063278 | 3.7868533 |
| hsa-miR-4532 | 0.004153 | 3.615911 |
|
hsa-miR-6510-5p | 0.0111118 | 3.306896 |
|
hsa-miR-4655-5p | 0.0250463 | 3.2763517 |
| hsa-miR-4459 | 0.0215316 | 3.2297115 |
| hsa-miR-328 | 0.0284044 | 3.2181585 |
| hsa-miR-4463 | 0.0395245 | 3.191302 |
| hsa-miR-3935 | 0.002951 | 3.1714096 |
|
hsa-miR-3940-5p | 0.0007361 | 3.157188 |
|
hsa-miR-4745-5p | 0.0373854 | 3.0417292 |
| hsa-miR-3911 | 0.0342525 | 2.7798533 |
| hsa-miR-4466 | 0.0265112 | 2.7150042 |
| hsa-miR-4739 | 0.0150427 | 2.6702106 |
| hsa-miR-638 | 0.0410605 | 2.4692678 |
|
hsa-miR-1224-5p | 0.0308876 | 2.3942 |
| hsa-miR-4672 | 0.0159597 | 2.3273127 |
| hsa-miR-6125 | 0.0495394 | 2.2485054 |
| hsa-miR-4644 | 0.0017442 | 2.2420375 |
|
hsa-miR-197-3p | 0.0437008 | 2.1905556 |
| hsa-miR-6090 | 0.0250247 | 2.1693227 |
| hsa-miR-2861 | 0.0368039 | 2.080642 |
| hsa-miR-3188 | 0.0407896 | 2.0413487 |
 | Table IIIClinicopathological characteristics
of ovarian serous adenocarcinoma in urine. |
Table III
Clinicopathological characteristics
of ovarian serous adenocarcinoma in urine.
| Characteristic | No. | Relative miR-30a-5p
expression | P-value |
|---|
| Mean age
(years) |
| <50 | 21 | 1.84±0.41 | >0.05 |
| ≥50 | 13 | 2.24±0.49 | |
| Regional lymph node
involvement |
| No | 6 | 1.73±0.31 | <0.001 |
| Yes | 28 | 3.21±0.91 | |
| Clinical stage |
| I–II | 16 | 2.68±0.49 | <0.001 |
| III–IV | 18 | 1.38±0.34 | |
| Histological
grade |
|
Well-moderately-differentiated | 12 | 2.80±0.56 | <0.05 |
|
Poorly-differentiated | 22 | 1.55±0.35 | |
Furthermore, the relative miR-6076 level was
significantly reduced (>10-fold reduction) in the urine of
ovarian serous adenocarcinoma patients, when compared to the
healthy controls and the patients with benign gynecological
disease, respectively (P<0.05; Fig.
2B), These findings are consistent with the miRNA microarray
data, and suggest that an increased miR-30a-5p and a decreased
miR-6076 may serve as promising biomarkers that distinguish ovarian
serous adenocarcinoma from other gynecological diseases.
Assessment of the diagnostic value of
miR-30a-5p and miR-6076 for ovarian serous adenocarcinoma
To assess the diagnostic potential of urinary
miR-30a-5p and miR-6076 levels for ovarian serous adenocarcinoma,
we performed a receiver operator characteristic (ROC) curve
analysis of specificity and sensitivity of the 34 ovarian serous
adenocarcinoma and 25 healthy control samples. The ROC curve for
miR-30a-5p demonstrated high specificity and sensitivity, with an
area under the ROC curve (AUC) of 0.862 (95% confidence interval,
0.709–1.016), which suggests a high discrimination potential
(Fig. 2C). However, miR-6076 showed
a lower specificity and sensitivity, with an AUC of only 0.693 (95%
confidence interval, 0.482–0.904) (Fig.
2D). These results suggested that miR-30a-5p provides a useful
diagnostic biomarker for ovarian serous adenocarcinoma, in contrast
to miR-6076.
Upregulation of urinary miR-30a-5p level
may be specific for ovarian cancer
We investigated whether urinary miR-30a-5p is
elevated in other types of cancer or is specific for ovarian
cancer. The urinary miR-30a-5p level was examined in 34 ovarian
serous adenocarcinoma patients, 25 healthy controls, 20 gastric
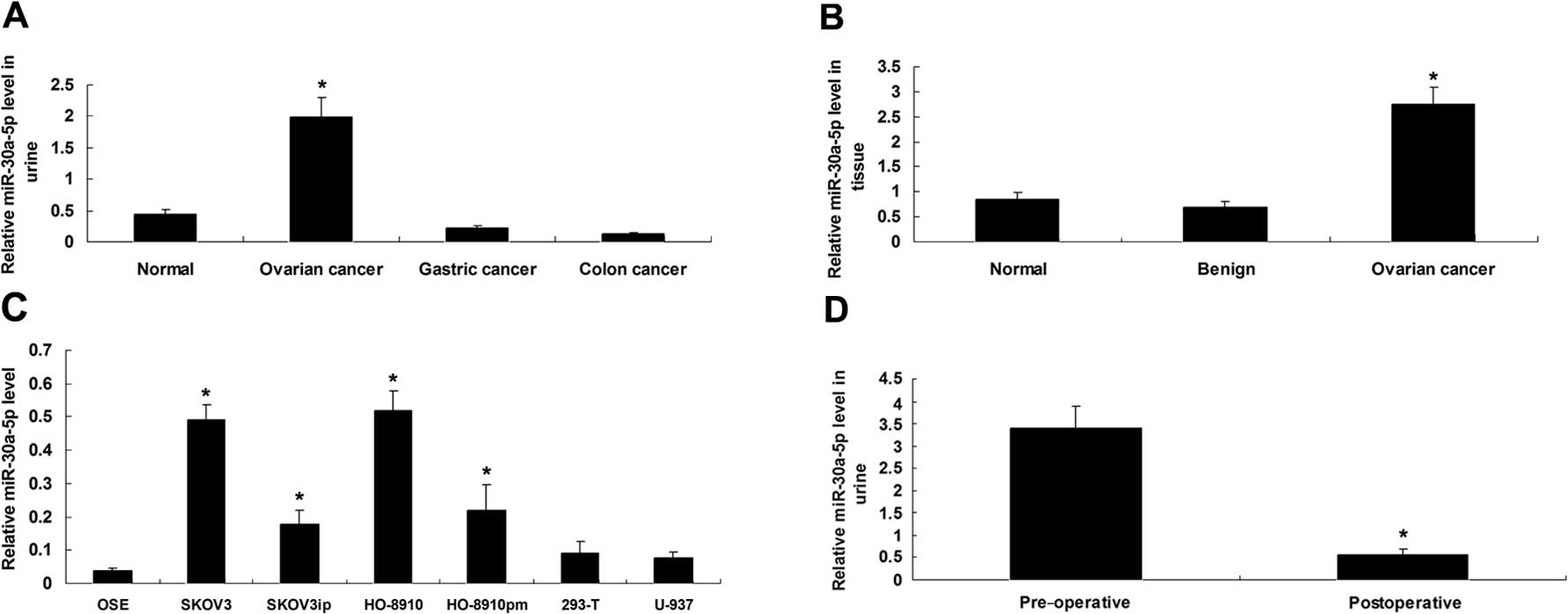
cancer patients and 20 colon carcinoma patients. Whereas the
ovarian serous adenocarcinoma samples showed an elevated miR-30a-5p
level compared with the healthy controls, the urine samples from
the gastric cancer patients and colon carcinoma patients showed a
reduced miR-30a-5p level (Fig. 3A).
The urinary miR-30a-5p levels of the gastric cancer and colon
carcinoma samples were 0.5-and 0.3-fold that of the healthy control
samples, respectively, although this difference was not
statistically significant (P>0.05) (Fig. 3A). These results suggested that
upregulation of the urinary miR-30a-5p level may be specific for
ovarian serous adenocarcinoma.
Urinary miR-30a-5p is dervied from the
ovarian tumor tissue
To determine whether the urinary miR-30a-5p was
derived from ovarian serous adenocarcinoma tissues, we first
determined the miR-30a-5p level in tissue samples from 30 ovarian
serous adenocarcinoma patients, 21 patients with benign
gynecological disease and 20 healthy controls. As shown in Fig. 3B, the relative miR-30a-5p level in
ovarian serous adenocarcinoma tissues was 3.2-fold higher than that
in the healthy controls (P<0.05) and 4.0-fold higher than that
in the benign ovarian specimens (P<0.05). Although the relative
miR-30a-5p expression level in the benign ovarian cancer specimens
was slightly lower than that in the healthy controls, this
difference was not statistically significant (P>0.05) (Fig. 3B). Similar to the expression
differences of miR-30a-5p among the urine samples, these data
confirmed that miR-30a-5p was also upregulated in ovarian serous
adenocarcinoma tissues. Subsequently, the miR-30a-5p level were
determined in normal human ovarian surface epithelial OSE cells,
low-invasion human ovarian cancer SKOV-3 and HO-8910 cells,
high-invasion human ovarian cancer SKOV-3ip and HO-8910pm cells,
human embryonic kidney 293-T cells and human U-937 macrophages.
Notably, miR-30a-5p was most highly expressed in all four ovarian
cancer cell lines, when compared to other cell types (Fig. 3C), confirming the association of
miR-30a-5p upregulation with ovarian cancer.
To confirm that miR-30a-5p in the urine was derived
from ovarian cancer cells, we determined whether the urinary
miR-30a-5p level was affected by the surgical removal of ovarian
serous adenocarcinoma. Assessment of the relative pre-operative and
postoperative miR-30a-5p levels for 15 ovarian serous
adenocarcinoma patients demonstrated that miR-30a-5p levels were
6-fold higher prior to surgery, than after surgery (Fig. 3D). The reduced urine level of
miR-30a-5p after surgery provided evidence that the urinary
miR-30a-5p level is associated with the presence of ovarian serous
adenocarcinoma, further supporting that the urinary miR-30a-5p is
derived from ovarian serous adenocarcinoma tissues.
Urinary miR-30a-5p is derived from the
exosomes secreted by ovarian cancer cells
Aberrant activity of cellular export machinery
contributes to cancer development. To determine whether the urinary
miR-30a-5p was derived from exosomes, which are cell-derived
vesicles that are present in urine and other body fluids, we
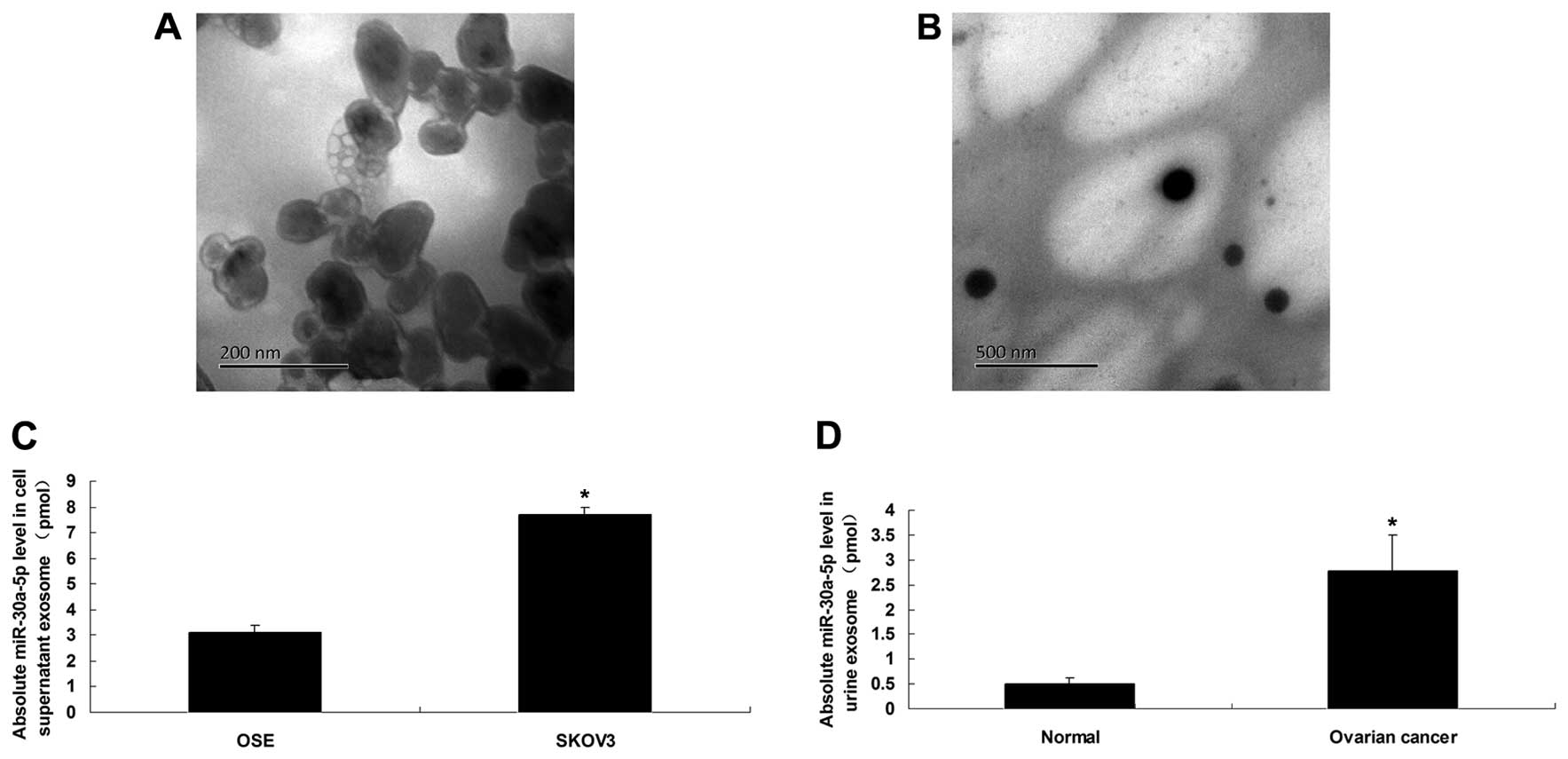
compared the miR-30a-5p levels in exosomes isolated from the
supernatant of SKOV-3 (Fig. 4A) vs.
OSE cells, and urine samples (Fig.
4B) from 6 ovarian serous adenocarcinoma patients vs. 6 healthy
controls. The miR-30a-5p level in SKOV-3 exosomes was 2.5-fold
higher than that in OSE exosomes (P<0.05; Fig. 4C). Furthermore, the miR-30a-5p level
in the urine exosomes from ovarian serous adenocarcinoma patients
was 3.3-fold higher than that from the healthy controls (Fig. 4D). The results suggested that the
increased miR-30a-5p level in the urine of ovarian cancer patients
may be due to the secretion of exosomes from the ovarian cancer
cells.
Knockdown of miR-30a-5p significantly
inhibits cell prolife ration and migration of ovarian cancer
cells
We preliminarily investigated the role of miR-30a-5p
in the regulation of malignant phenotypes of ovarian cancer in
vitro. We transfected SKOV-3ip and HO-8910pm cells with
scrambled miRNA or miR-30a-5p inhibitor, respectively. After
transfection, RT-qPCR was conducted to detect the expression level
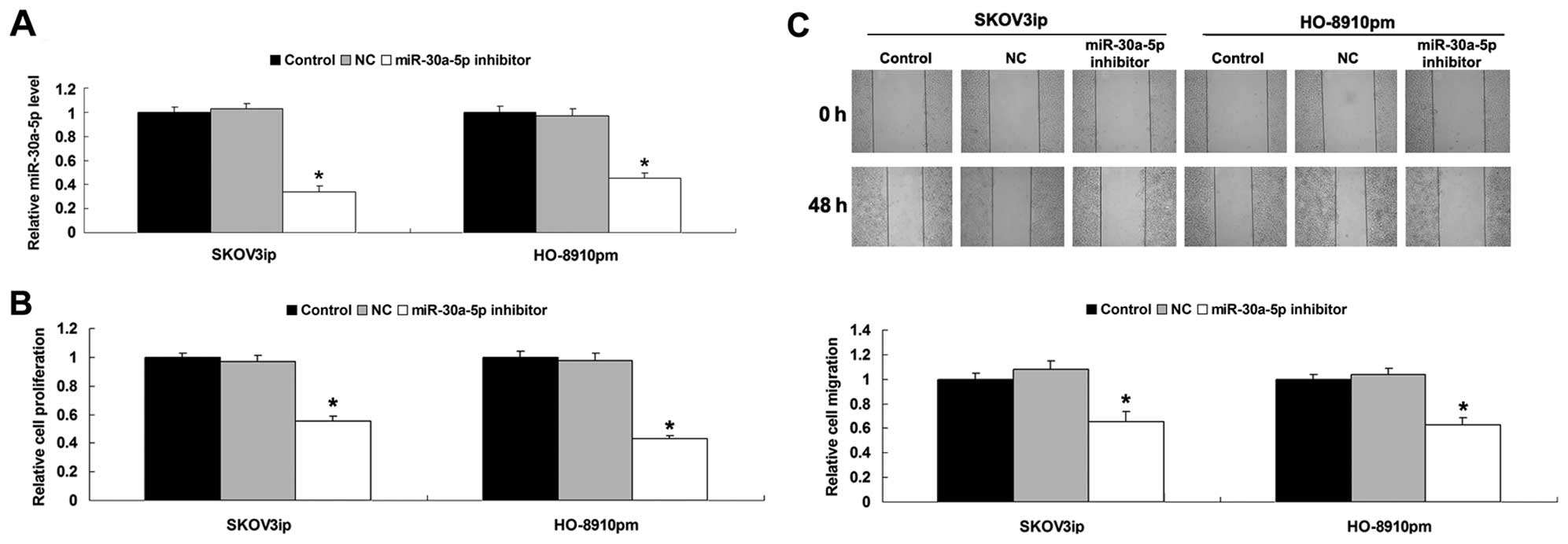
of miR-30a-5p in each group. As shown in Fig. 5A, the expression level of miR-30a-5p
was notably downregulated in ovarian cancer cells transfected with
miR-30a-5p inhibitor compared to the control group (P<0.05).
However, transfection with scrambled miRNA did not affect the
miR-30a-5p level in ovarian cancer cells. These results suggested
that our transfection was successful. Subsequently, we determined
the proliferation and migration abilities of ovarian cancer cells
in each group. As shown in Fig. 5B and
C, knockdown of miR-30a-5p significantly suppressed the cell
proliferation and migration of ovarian cancer cells, when compared
to the control group (P<0.05). These results suggested that
miR-30a-5p acts as an oncogenic miRNA in ovarian cancer.
Discussion
MicroRNAs (miRNAs) can be detected in blood, urine,
breast milk, tears, colostrum, bronchial lavage, seminal and
pleural fluids. The diagnostic and prognostic value of miRNAs in
the urine has been suggested for a variety of diseases (16,17).
Previous findings have shown that the different expression patterns
of miRNAs in tissues can be used to distinguish ovarian cancers
from normal tissue (18). However,
the miRNA profiling from tissues is only available to patients
undergoing surgery. Accumulating evidence has demonstrated the
imbalanced production of some miRNAs in the circulation (19,20).
However, blood tests was invasive, limiting its clinical utility.
Collection of urine is more convenient, cost-effective and
non-invasive, when compared to serum, plasma and tissue specimens.
Therefore, we examined whether miRNAs in the urine served as
non-invasive biomarkers for ovarian cancer.
Our miRNA microarray results showed that 38 miRNAs
were significantly differentially expressed in the urine from
ovarian serous adenocarcinoma patients compared to healthy
controls, including one (miR-30a-5p) upregulated miRNA and 37
downregulated miRNAs. Further investigation found that the urinary
miR-30a-5p in gastric cancer and colon carcinoma patients was
comparable or less than that in the normal controls. However, the
miR-30a-5p level in the urine of patients with ovarian serous
adenocarcinoma was significantly elevated compared to the healthy
controls. This tissue-specific expression of miR-30a-5p makes it
particularly suitable as a biomarker for ovarian serous
adenocarcinoma. Recently, Jiang et al showed that miR-30a-5p
was among novel non-invasive biomarkers for the diagnosis and
recurrence of bladder cancer (21).
Wang et al found that miR-30a-5p was upregulated in glioma
samples as compared to the normal brain tissues, and its expression
level was positively correlated with tumor grade of malignancy
(22). Notably, we found that the
urinary miR-30a-5p level was higher in stage I-II ovarian serous
adenocarcinoma samples than that in stage III-IV samples, and also
higher in well or moderately differentiated ovarian serous
adenocarcinoma samples than in poorly differentiated samples. We
hypothesized that the upregulation of miR-30a-5p may be associated
with earlier stages or low-invasive features of ovarian serous
adenocarcinoma. Consistent with these data, the low-invasion
ovarian cell lines expressed higher levels of miR-30a-5p than the
high-invasion cell lines. However, we found that the upregulated
miR-30a-5p level in the urine was associated with lymphatic
metastasis.
We also showed that the expression pattern of
miR-30a-5p in tissues was similar to that in urine, demonstrated by
a significantly higher level of miR-30a-5p in ovarian serous
adenocarcinoma tissues, when compared with that in healthy controls
or patients with benign gynecological diseases. Additionally, the
expression of miR-30a-5p in ovarian cancer cell lines was
upregulated when compared with that in normal human ovarian
epithelial cells or non-ovarian cells. These results suggest that
the urinary miR-30a-5p may be derived from the ovarian cancer cells
within the tumor tissues. To further verify the ovarian serous
adenocarcinoma origin of miR-30-5p in the urine, we compared
miR-30a-5p levels in the urine of 15 ovarian serous adenocarcinoma
patients before and after surgical resection of tumor tissues. The
pre-operative miR-30a-5p level in the urine was significantly
higher than the postoperative miR-30a-5p level, indicating that the
ovarian serous adenocarcinoma is the original source of miR-30a-5p
in the urine.
Exosomes are small (30–120 nm) membrane vesicles
that may be released from different cell types including cancer
cells to the urine (23). It has
been demonstrated that exosomes can be isolated from various body
fluids including plasma, malignant ascites, amniotic fluid, breast
milk, saliva and urine (24,25).
Furthermore, extracellular miRNAs in exosomes are hypothesized to
play a role in cell-to-cell communication and to serve as
biomarkers for human diseases (25). Taylor and Gercel-Taylor showed that
miRNAs in exosomes from ovarian cancer patients exhibited similar
profiles with miRNAs in tumor cells from the same ovarian cancer
patients (19), which is in
agreement with our findings that the miR-30a-5p level in exosomes
and cells showed a similar expression pattern.
As the expression level of miR-30a-5p was
significantly upregulated in urine and tumor tissues from ovarian
serous adenocarcinoma patients, it plays an oncogenic role in
ovarian cancer. To investigate the exact role of miR-30a-5p in
ovarian cancer, we inhibited its expression in ovarian cancer cell
lines, and showed that knockdown of miR-30a-5p significantly
suppressed cell proliferation and migration of ovarian cancer
cells. miR-30a-5p has been found to be deregulated in several other
types of human cancers, such as glioma, colon cancer, thyroid
anaplastic carcinomas and gastric cancer (22,26–28).
Moreover, the exact role of miR-30a-5p is tumor-specific. For
instance, miR-30a-5p acts as a tumor suppressor in colon carcinoma
by targeting DTL (29). Knockdown
of miR-30a-5p using antisense oligonucleotide suppresses glioma
cell growth by targeting SEPT7, suggesting that it acts as an
oncogenic miRNA in glioma (30).
To the best of our knowledge, this is the first use
of micro-array technology to investigate the urine miRNA profiling
of ovarian serous adenocarcinoma. The extraction of miRNA and the
identification of miRNAs from the urine are feasible and
practicable for the diagnosis of serous epithelial ovarian cancer.
Moreover, we have demonstrated for the first time that miR-30a-5p
was increased in the urine of ovarian serous adenocarcinoma
patients, and inhibition of miR-30a-5p suppressed the malignant
phenotypes of ovarian cancer in vitro. Therefore, miR-30a-5p
serves as a promising diagnostic and therapeutic target for ovarian
serous adenocarcinoma.
Acknowledgments
This study was financially supported by the
Technology Plan of Science and Technology Bureau Hunan Province
2014FJ3090, and the National Natural Science Foundation of China
nos. 81172469 and 81102009.
References
|
1
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2012. CA Cancer J Clin. 62:10–29. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Wright JD, Shah M, Mathew L, Burke WM,
Culhane J, Goldman N, Schiff PB and Herzog TJ: Fertility
preservation in young women with epithelial ovarian cancer. Cancer.
115:4118–4126. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
van Nagell JR Jr and Hoff JT: Transvaginal
ultrasonography in ovarian cancer screening: Current perspectives.
Int J Womens Health. 6:25–33. 2013. View Article : Google Scholar :
|
|
4
|
Leung F, Diamandis EP and Kulasingam V:
Ovarian cancer biomarkers: Current state and future implications
from high-throughput technologies. Adv Clin Chem. 66:25–77. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Jacobs I and Bast RC Jr: The CA 125
tumour-associated antigen: A review of the literature. Hum Reprod.
4:1–12. 1989.PubMed/NCBI
|
|
6
|
Cohen JG, White M, Cruz A and
Farias-Eisner R: In 2014, can we do better than CA125 in the early
detection of ovarian cancer? World J Biol Chem. 5:286–300. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Berek JS and Bast RC Jr: Ovarian cancer
screening. The use of serial complementary tumor markers to improve
sensitivity and specificity for early detection. Cancer. 76(Suppl
10): S2092–S2096. 1995. View Article : Google Scholar
|
|
8
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lu J, Getz G, Miska EA, Alvarez-Saavedra
E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA,
et al: MicroRNA expression profiles classify human cancers. Nature.
435:834–838. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Neal CS, Michael MZ, Pimlott LK, Yong TY,
Li JY and Gleadle JM: Circulating microRNA expression is reduced in
chronic kidney disease. Nephrol Dial Transplant. 26:3794–3802.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Lorenzen JM, Volkmann I, Fiedler J,
Schmidt M, Scheffner I, Haller H, Gwinner W and Thum T: Urinary
miR-210 as a mediator of acute T-cell mediated rejection in renal
allograft recipients. Am J Transplant. 11:2221–2227. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hanke M, Hoefig K, Merz H, Feller AC,
Kausch I, Jocham D, Warnecke JM and Sczakiel G: A robust
methodology to study urine microRNA as tumor marker: microRNA-126
and microRNA-182 are related to urinary bladder cancer. Urol Oncol.
28:655–661. 2010. View Article : Google Scholar
|
|
16
|
Wang G, Tam LS, Li EK, Kwan BC, Chow KM,
Luk CC, Li PK and Szeto CC: Serum and urinary cell-free miR-146a
and miR-155 in patients with systemic lupus erythematosus. J
Rheumatol. 37:2516–2522. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Luo Y, Wang C, Chen X, Zhong T, Cai X,
Chen S, Shi Y, Hu J, Guan X, Xia Z, et al: Increased serum and
urinary microRNAs in children with idiopathic nephrotic syndrome.
Clin Chem. 59:658–666. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Iorio MV, Visone R, Di Leva G, Donati V,
Petrocca F, Casalini P, Taccioli C, Volinia S, Liu CG, Alder H, et
al: MicroRNA signatures in human ovarian cancer. Cancer Res.
67:8699–8707. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Taylor DD and Gercel-Taylor C: MicroRNA
signatures of tumor-derived exosomes as diagnostic biomarkers of
ovarian cancer. Gynecol Oncol. 110:13–21. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Resnick KE, Alder H, Hagan JP, Richardson
DL, Croce CM and Cohn DE: The detection of differentially expressed
microRNAs from the serum of ovarian cancer patients using a novel
real-time PCR platform. Gynecol Oncol. 112:55–59. 2009. View Article : Google Scholar
|
|
21
|
Jiang X, Du L, Wang L, Li J, Liu Y, Zheng
G, Qu A, Zhang X, Pan H, Yang Y, et al: Serum microRNA expression
signatures identified from genome-wide microRNA profiling serve as
novel noninvasive biomarkers for diagnosis and recurrence of
bladder cancer. Int J Cancer. 136:854–862. 2015. View Article : Google Scholar
|
|
22
|
Wang K, Jia Z, Zou J, Zhang A, Wang G, Hao
J, Wang Y, Yang S and Pu P: Analysis of hsa-miR-30a-5p expression
in human gliomas. Pathol Oncol Res. 19:405–411. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Vlassov AV, Magdaleno S, Setterquist R and
Conrad R: Exosomes: Current knowledge of their composition,
biological functions, and diagnostic and therapeutic potentials.
Biochim Biophys Acta. 1820:940–948. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gu Y, Li M, Wang T, Liang Y, Zhong Z, Wang
X, Zhou Q, Chen L, Lang Q, He Z, et al: Lactation-related microRNA
expression profiles of porcine breast milk exosomes. PLoS One.
7:e436912012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Keller S, Ridinger J, Rupp AK, Janssen JW
and Altevogt P: Body fluid derived exosomes as a novel template for
clinical diagnostics. J Transl Med. 9:862011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Xi Y, Formentini A, Chien M, Weir DB,
Russo JJ and Ju J, Kornmann M and Ju J: Prognostic values of
microRNAs in colorectal cancer. Biomark Insights. 2:113–121.
2006.
|
|
27
|
Visone R, Pallante P, Vecchione A,
Cirombella R, Ferracin M, Ferraro A, Volinia S, Coluzzi S, Leone V,
Borbone E, et al: Specific microRNAs are downregulated in human
thyroid anaplastic carcinomas. Oncogene. 26:7590–7595. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Li X, Zhang Y, Zhang Y, Ding J, Wu K and
Fan D: Survival prediction of gastric cancer by a seven-microRNA
signature. Gut. 59:579–585. 2010. View Article : Google Scholar
|
|
29
|
Baraniskin A, Birkenkamp-Demtroder K,
Maghnouj A, Zöllner H, Munding J, Klein-Scory S, Reinacher-Schick
A, Schwarte-Waldhoff I, Schmiegel W and Hahn SA: MiR-30a-5p
suppresses tumor growth in colon carcinoma by targeting DTL.
Carcinogenesis. 33:732–739. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Jia Z, Wang K, Wang G, Zhang A and Pu P:
MiR-30a-5p antisense oligonucleotide suppresses glioma cell growth
by targeting SEPT7. PLoS One. 8:e550082013. View Article : Google Scholar : PubMed/NCBI
|