Introduction
Pancreatic cancer (PCC) represents one of the
leading causes of cancer-related mortality in industrialized
countries (1). Despite surgical
resection, radiation, and chemotherapy, >94% of patients with
PCC do not survive beyond 5 years. The poor prognosis of this
disease is mainly due to its early systemic metastasis (1–3).
It is well established that the tetraspanin family
of four-pass transmembrane proteins has been implicated in
fundamental biological processes, including cell adhesion,
migration, and proliferation (4).
Tetraspanins interact with various transmembrane proteins,
establishing a network of large multimolecular complexes that
allows specific lateral secondary interactions (5). In animals, the tetraspanins are a
large superfamily of membrane proteins that play important roles in
organizing various cell-cell and matrix-cell interactions and
signal pathways based on such interactions (6,7).
Tetraspanins are a heterogeneous group of
4-transmembrane proteins that segregate into so-called
tetraspanin-enriched microdomains (TEMs) along with other cell
surface proteins such as integrins. TEMs of various types are
reportedly involved in the regulation of cell growth, migration and
invasion of several tumor cell types, both as suppressors or
supporting structures (8).
Tetraspanin 1 (Tspan1), a member of the transmembrane 4 superfamily
of tetraspanins, is overexpressed in high-grade cervical
intraepithelial neoplasia (CIN) and terminal carcinomas (8), lung cancer (9), colon cancer (10), breast cancer (11), as well as squamous cell carcinoma
(12). However, the precise
function of Tspan1 in the context of PCC is not known.
In the present study, quantitative RT-PCR (qRT-PCR)
and western blot analysis were employed to explore the expressions
of Tspan1 in human PCC tissues, adjacent normal tissues and human
AsPC-1 and PANC-1 cell lines. Immunohistochemistry (IHC) was also
used to detect the subcellular locations of Tspan1 in human PCC
tissues. The corrections between Tspan1 expression in PCC and
clinicopathological features were analyzed. Then, virus carrying a
small interference RNA (siRNA) targeting the human Tspan1 gene was
constructed and transfected into PCC cells. After transfection,
Tspan1 expression was detected by qRT-PCR and western blot
analysis. The cell proliferation and apoptosis fractions were also
evaluated by MTT assay and flow cytometry (FCM). Additionally, the
Transwell assays were employed to explore the effects of Tspan1
knockdown on the migration and invasion of PCC cells in
vitro.
Materials and methods
Ethical aspects
This study complied with the International Ethical
Declaration and was approved by the Human Ethics Committee and the
Research Ethics Committee of Shaanxi Province of China. Through the
surgery consent form, patients were informed that the resected
specimens were kept by our hospital and may be used for scientific
research, and that their privacy would be maintained.
Clinical samples collections
Forty-five patients attending the clinic in the
Weinan Central Hospital, between February 2013 to December 2014,
were invited to participate in the study. The patients ranged in
age from 35 to 72 years, with a median age of 51 years. All of them
were operated for the first time, without any antitumor treatment.
The specimens were histopathologically verified as PCC. Then the
tumor samples and matched normal adjacent tissues were obtained,
which were taken at least 0.5 cm distal to tumor margins. The
biopsies obtained were divided into two fragments immediately after
surgery. One fragment was immediately stored at −80°C until nucleic
acid and protein isolation. The remaining fragment was fixed in 4%
formaldehyde for two days and then paraffin-embedded. The paraffin
blocks were sliced and stained with IHC for histological
examination.
IHC staining
Immunohistochemistry was performed to determine the
Tspan1 expression in normal adjacent tissue and PCC tissues.
Briefly, tumor samples and the matched control tissues were fixed
in 4% formaldehyde, and embedded in paraffin wax. Fixed tumor
samples were prepared into 4-µm frozen sections and
incubated with 2% goat serum at 37°C for 20 min, followed by
incubation with rat anti-human Tspan1 (1:400, sc-376551; Santa Cruz
Biotechnology) at 4°C overnight. After washing with PBS, the
sections were incubated with horseradish peroxidase
(HRP)-conjugated rabbit anti-rat IgG (HRP-IgG) at 37°C for 30 min
and colored with 3,3′-diaminobenzidine (DAB) at room temperature.
PBS was substituted for the anti-Tspan1 antibody in negative
control subjects. All sections were then blindly analyzed by three
experienced pathologists under a light microscope. The
clinicopathologic data and patient outcomes were not known by the
pathologists. The results of IHC staining were evaluated as
described (13). The cases were
classified into positive groups by the intensity and proportion of
the immunostained cancer cells or Tspan1. The proportion of
positive cells was assessed as low (5–25% cells stained), medium
(25–50% cells stained), and high (>75% cells stained).
Quantitative reverse-transcription
PCR
The expression of Tspan1 in the PCC specimens and
the cell lines were detected by quantitative real-time PCR
(qRT-PCR). Tumor samples (50 mg) were ground under liquid nitrogen,
lysed with 1 ml of TRIzol (Takara, Japan), and total RNA was
extracted using TRIzol (Invitrogen, Carlsbad, CA, USA). Total RNA
(2 µg) was added to the tumor extract with Moloney Murine
Leukemia Virus Reverse Transcriptase (MMLV-RT; Takara) to
synthesize cDNA, and the reverse transcript was used as the
template for qRT-PCR using a Tower qRT-PCR system (Analytik, Jena,
Germany). The qRT-PCR was conducted using 2X Mix SYBR green I (10
µl; Biosea, USA), primer (0.25 µl, 10 pmol/l),
template DNA (1 µl), and sterile water (8.5 µl). All
PCR reactions included initial denaturation and multiple cycles at
95°C for 3 min; 39 cycles at 95°C for 10 sec, 55°C for 10 sec, and
72°C for 30 sec; followed by 95°C for 10 sec, 65°C for 5 sec, and a
final 95°C for 15 sec. The primer for each gene was synthesized by
Invitrogen. The real-time PCR primers used to quantify GAPDH
expression were: forward, 5′-CGAGATCCCTCCAAAATCAA-3′ and reverse,
5′-TTCACACCCATGACGAACAT-3′; and for Tspan1 were: forward,
5′-TGGGCTGCTATGGTGCTA-3′ and reverse, 5′-T-GCAGGTTTCATTGGCTGT-3′.
Expression of Tspan1 was normalized to endogenous GAPDH
expression.
Western blot analysis
Tspan1 protein levels both in PCC tissues and cell
lines were determined by western blot analysis. Briefly, samples
were lysed for 30 min in CytoBuster Protein Extraction Buffer
(Novagen, USA) and centrifuged at 12,000 rpm. The supernatant was
collected, total protein was measured, and 50 µg was used
for 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis
(SDS-PAGE). The protein was then transferred to a nitrocellulose
(NC) membrane and sealed with Tris-buffered saline Tween-20 (TBST)
containing 5% non-fat milk powder. The membrane was subsequently
incubated with goat anti-human Tspan1 proteins and mouse anti-human
GAPDH (1:500, sc-81545; Santa Cruz Biotechnology) at 4°C overnight.
After washing in TBST, the membrane was incubated with
HRP-conjugated secondary antibodies (1:2,000) at 25°C, and the
protein quantity was determined using electrochemiluminescence
(ECL) technique (BestBio, USA). The results were photographed using
the JS gel Imaging system and the grey density was calculated using
SensiAnsys software (both from Peiqing, China).
Cell culture
Human PCC cell lines AsPC-1 and PANC-1, as well as
normal human pancreatic hTERT-HPNE cell lines were all purchased
from the Cell Bank of the Chinese Academy of Sciences.
All the above cells were cultured in specific medium
supplemented with 10% (v/v) fetal bovine serum (FBS) and 1%
antibiotics at 37°C in a humidified incubator under 5%
Co2 condition (14).
Tspan1 knockdown
According to the CDS of Tspan1 recorded in
neucleopeptide, we predesigned siRNA targeting the human Tspan1
gene (gene ID, 10103) (http://RNAiDesigner.invit-rogen.com). The siRNA
sequences targeting Tspan1 were as follows: si-1,
5′-CCTCAGCAGTTCCCTCTTT-3′; si-2, 5′-GCCTTGGTGTACACCACAA-3′; and
si-3, 5′-GCCTGCCATCAAGAAAGAT-3′. Lentivirus was packaged in 293T
cells using Lipofectamine 2000 (Invitrogen) and virus titers were
determined. Target cells, including AsPC-1 and PANC-1 cells, were
infected with 1×106 recombinant lentivirus-transducing
units in the presence of 6 µg/ml polybrene (Sigma),
respectively. The efficiency of knockdown was tested by qRT-PCR and
western blot analysis. All experiments were performed in
triplicate.
MTT assay
Cell viability was determined using the tetrazolium
salt 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide
(MTT) assay. Briefly, cells were plated into 96-well culture plates
at an optimal density of 5×103 cells/ml in 200 µl
of culture medium/well. After 24–96 h of culture, 20 µl of 5
mg/ml MTT was added to each well and incubated at 37°C for 4 h. The
medium was then gently aspirated and 150 µl of dimethyl
sulfoxide (DMSO) was added to each well to solubilize the formazan
crystals. The optical density of each sample was immediately
measured using a microplate reader (Bio-Rad, Hercules, CA, USA) at
490 nm.
Apoptosis assay
A propidium iodide (PI) and Annexin V-FITC flow
cytometry assay (BD Pharmingen) was used to detect the apoptosis
rate in the cells after Tspan1 transfection. Briefly,
1×106 cells/well were cultured in 6-well plates in the
absence of 10% FBS for 48 h. Adherent cells were detached with
0.25% trypsin without EDTA in 1X PBS. Cells were harvested in
complete RPMI-1640 medium and centrifuged at 1,000 rpm for 5 min.
Each of the cell lines was washed with 1X PBS and stained with 50
µg/ml PI and Annexin V-FITC, following the manufacturer's
instructions.
Cell migration and invasion assay
We employed BioCoat Matrigel invasion chambers (BD
Biosciences, Bedford, MA, USA) to compare the effect of Tspan1
knockdown on in vitro invasion of AsPC-1 and PANC-1 cells as
previously described (15).
Briefly, for the invasion assay, Costar Transwell 8-µm
inserts were coated with 50 µg reduced serum Matrigel (BD
Biosciences). Invasion chambers were coated with Matrigel, and
1×106 cells were added per chamber. Medium supplemented
with 10% FBS was used in the lower chamber. Following incubation
the cells that had invaded through the membrane were fixed and
stained before the membrane was removed and mounted on a slide for
microscopic assessment. Invasive cells were visualized at ×40
magnification and the number of cells in five random fields was
counted and an average calculated. For migration assays, the same
procedure was used excluding the Matrigel. After 12 h, non-invading
cells and media were removed, and cells on the lower surface of the
membrane were fixed with polyoxymethylene and stained with 0.1%
crystal violet (both from Sigma) for 0.5 h. Stained cells were
counted under a microscope in five randomly selected fields, and
the average was used to indicate cell migration and invasion. All
experiments were performed in triplicate (15,16).
Statistical analysis
SPSS v11.5 (SPSS Inc., Chicago, IL, USA) was used
for statistical analysis. Data are presented as means ± standard
deviation. The unpaired t-test was used for comparison between
groups. Association between Tspan1 expression and other
clinicopathological factors of the tumor were assessed by the
Fisher's exact test (two-sided) for categorical variables and
χ2 test were used to compare ordinal variables. The
grading-related data were analyzed by Spearman's test. A P<0.05
was considered statistically significant.
Results
Upregulation of Tspan1 expression in
human PCC tissues
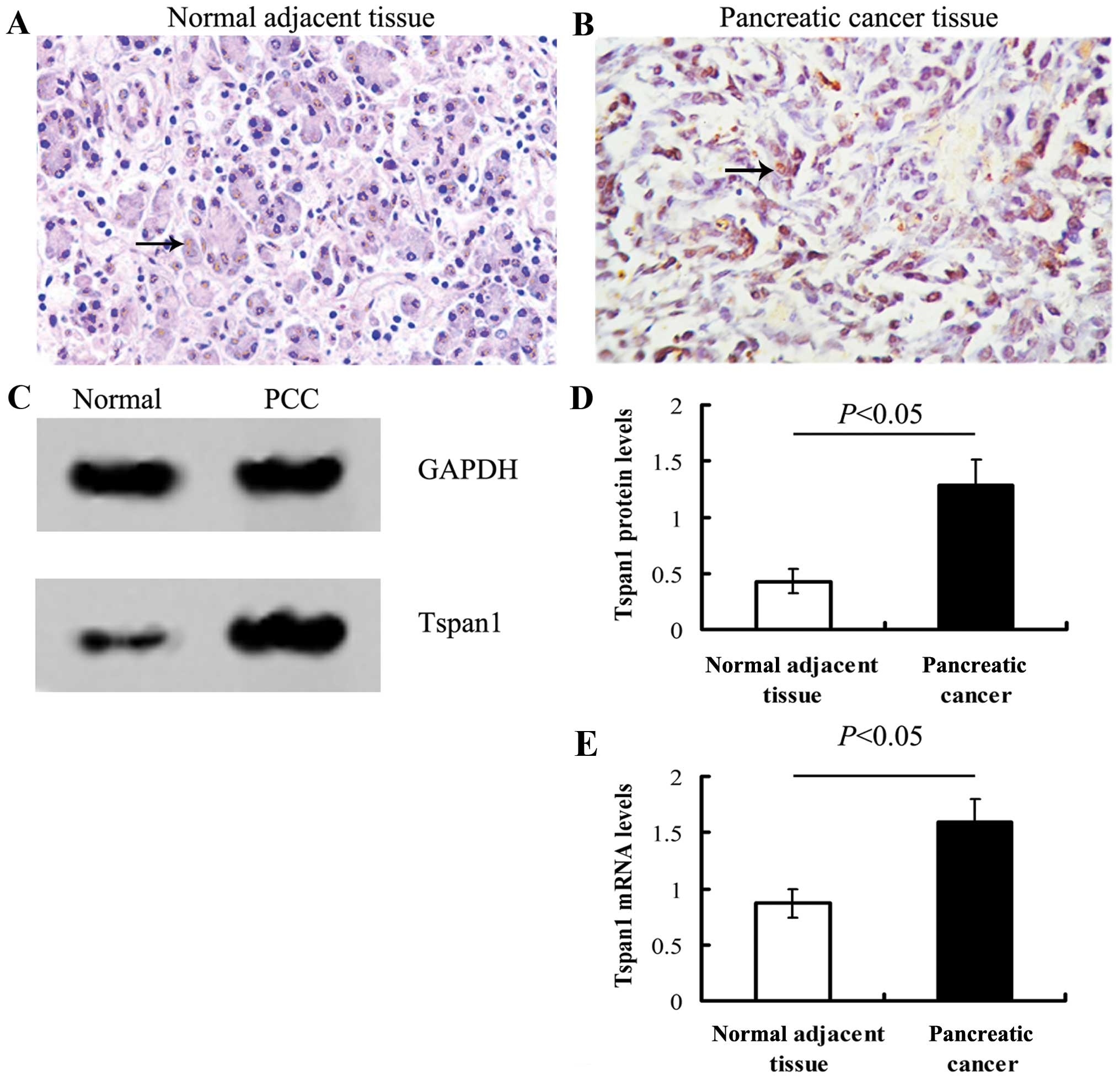
Tspan1 staining in normal adjacent tissue was weak
relative to PCC tissues. The IHC positive files of Tspan1 exhibited
light yellow to brown staining (Fig.
1).
Either qRT-PCR or western blot analysis showed that
the expression of Tspan1 in PCC tissue was significantly stronger
than that of normal tissues (P<0.05, Fig. 1).
Spearman's analysis showed that the Tspan1 level was
correlated with the lymph node metastasis (r=0.311, P<0.05) and
the pathological tumor node metastasis (pTNM) stages (r=0.295,
P<0.05) in these 45 cancer samples (Table I). Within a period of 60 Smonths of
the follow-up, 15 cancer-related deaths occurred, all of the deaths
come from patients with Tspan1 positive tumors. The 5-year survival
rate of pTNM stage I is >95%, while it is <10% in patients
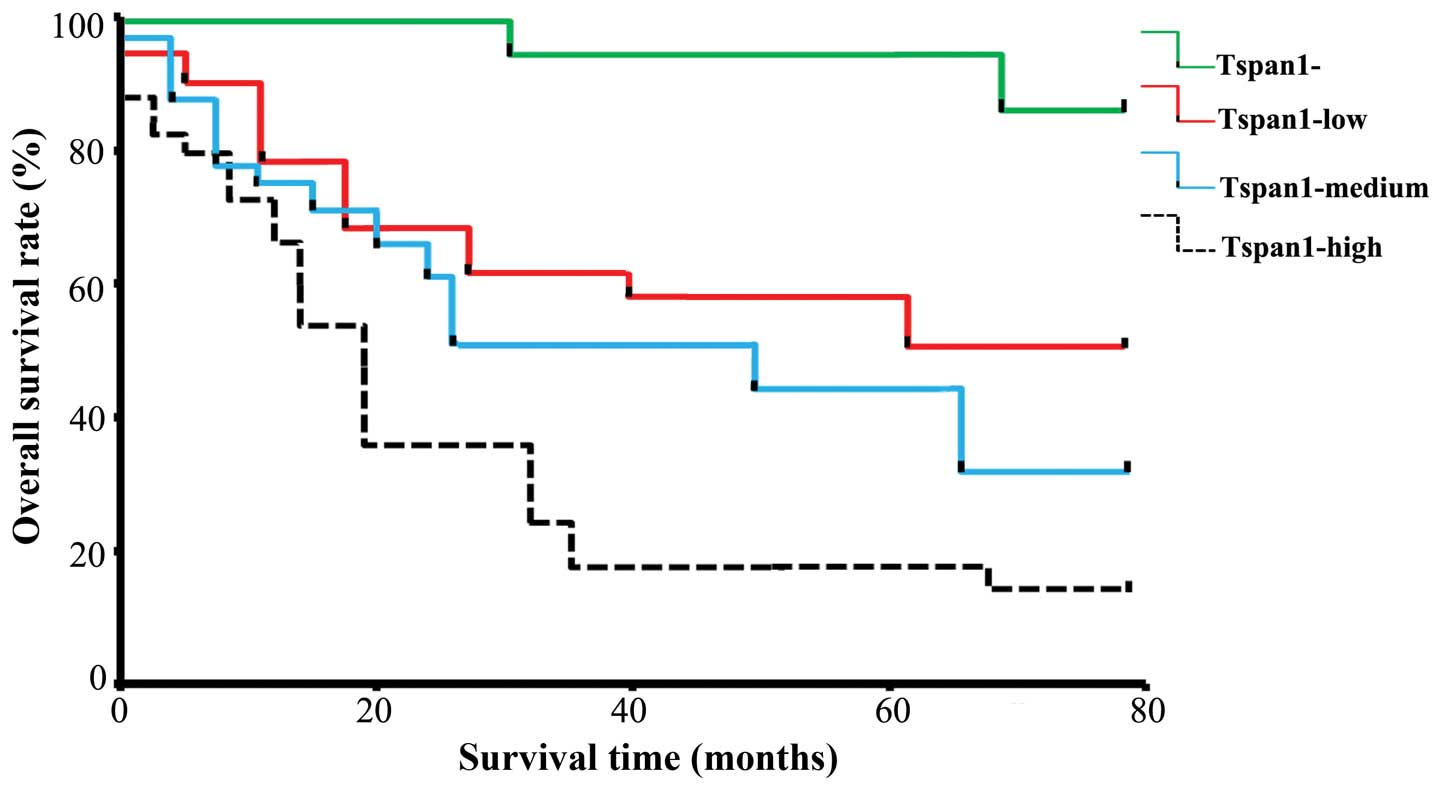
with pTNM stage III-IV. The Kaplan-Meier estimates of overall
survival rate were based on cell Tspan1 expression in the patients
with a follow-up period of 60 months (Fig. 2). In the entire cohort, the overall
survival rate of patients with Tspan1-negative tumors was
significantly higher than that in Tspan1-positive tumors (85.71 vs.
35.71%; log-rank test: χ2=19.08, P=0.0001). The
relationship between Tspan1 expression pattern (IHC staining) and
survival rate was also determined. Results revealed that Tspan1
higher-expression group had significantly shorter survival than the
Tspan1 lower-expression group (Tspan1-medium vs. Tspan1-high,
P<0.05; Tspan1-negative vs. Tspan1-high, P<0.05;
Tspan1-medium vs. Tspan1-negative, P<0.05; Fig. 2).
 | Table ICorrelations between Tspan1 expression
in PCC and clinicopathological features. |
Table I
Correlations between Tspan1 expression
in PCC and clinicopathological features.
| Tspan1 expression
| rs | P-values |
|---|
| Negative | Low | Medium | High |
|---|
| Gender |
| Male | 4 | 8 | 5 | 8 | 0.127 | 0.61 |
| Female | 3 | 5 | 7 | 6 | 0.119 | 0.59 |
| Tumor size (cm) | 2.208±0.25 | 3.109±1.13 | 4.027±1.98 | 5.924±1.72 | 9.561 | 0.000326 |
| Histological
grade | | | | | 0.103 | 0.34 |
| G1 | 5 | 3 | 4 | 5 | | |
| G2 | 1 | 6 | 3 | 4 | | |
| G3 | 1 | 4 | 5 | 5 | | |
| LN metastasis | | | | | 0.311 | 0.003 |
| N0 | 6 | 9 | 4 | 4 | | |
| N1 | 1 | 3 | 8 | 10 | | |
| pTNM stage | | | | | 0.295 | 0.007 |
| I, II | 6 | 7 | 2 | 2 | | |
| III | 1 | 3 | 4 | 5 | | |
| IV | 0 | 2 | 6 | 7 | | |
Interference expression of Tspan1 by
siRNA transfection
The PCC cell lines, AsPC-1 and PANC-1 cells, as well
as the hTERT-HPNE cell stably transfected with
Tspan1-siRNA-expressing vector (named as AsPC-1-Tspan1-si-1/2/3,
PANC-1-Tspan1-si-1/2/3 and normal-Tspan1-si1/2/3, respectively).
Control AsPC-1, PANC-1 and hTERT-HPNE cells were transfected with
empty vectors. They were recorded as AsPC-1-Empty, PANC-1-Empty and
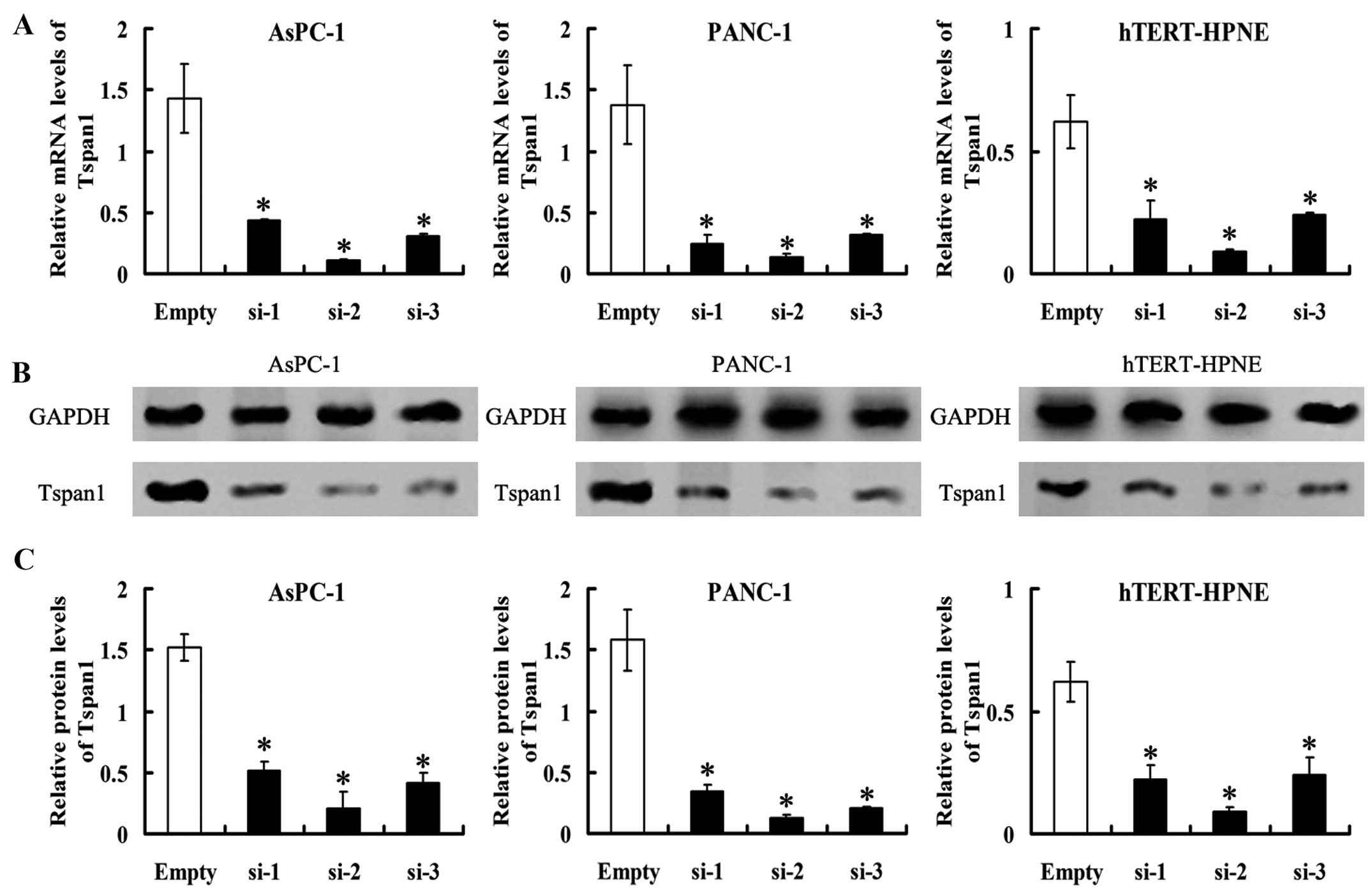
normal-Empty, respectively. Tspan1 mRNA levels detected by RT-PCR
were significantly lower in Tspan1-siRNA expressed cells than the
matched control (P<0.05, Fig.
3). Western blot analysis found that the level of
immunoreactive protein was significantly downregulated in
Tspan1-siRNA-transfected cells relative to the controls cells
(P<0.05, Fig. 3).
Stable expression of three Tspan1-siRNA (si-1, si-2
and si-3) in AsPC-1, PANC-1 and hTERT-HPNE cells resulted in
>60% decrease in Tspan1 expression (Fig. 3). Considering the highest rates of
inhibition of expression in Tspan1, AsPC-1-Tspan1-si-2 and
PANC-1-Tspan1-si-2 were chosen as the target for further
investigation.
Effects of Tspan1 on PCC cells
proliferation
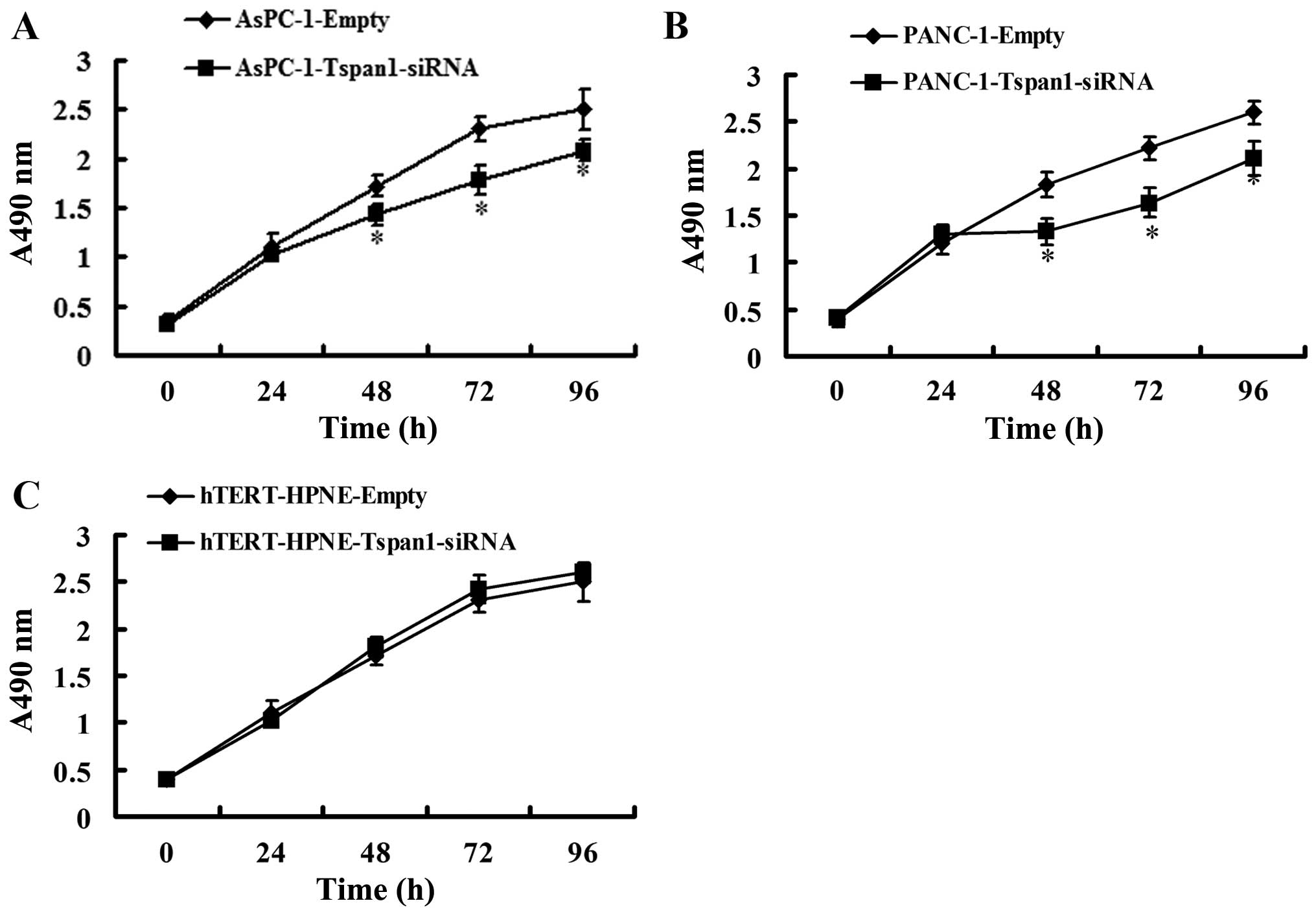
We assessed the effect of Tspan1 expression
silencing on the regulation of PCC cells viability. MTT assay
showed that silencing of Tspan1 expression caused significant
decrease in cell viability in AsPC-1 and PANC-1 cells, but not in
hTERT-HPNE cells (Fig. 4).
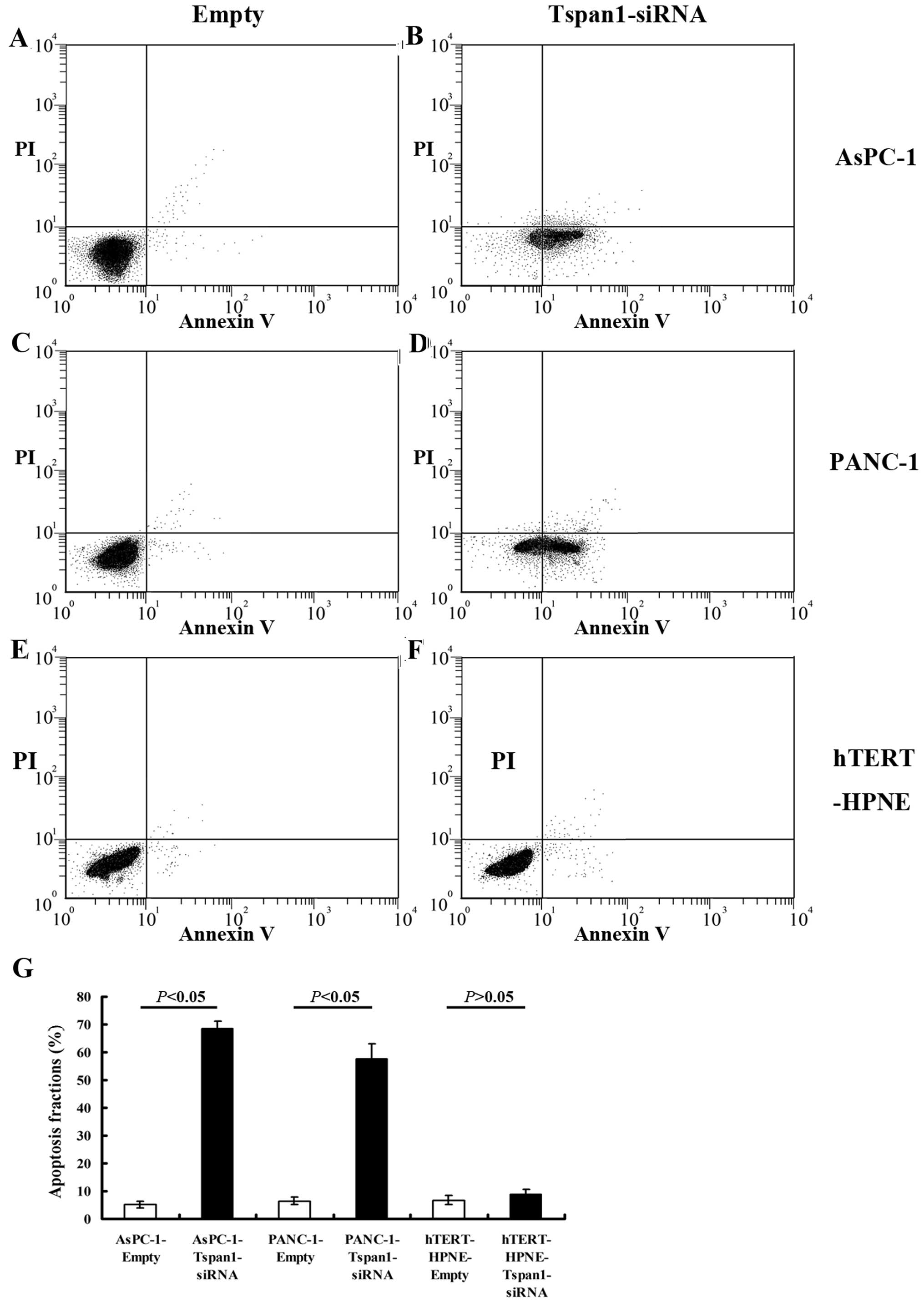
Downregulation of Tspan1 induces
increased apoptosis of PCC cells
There was a significant increase in the apoptosis
rate in Tspan1-siRNA-infected cells relative to empty-infected ones
(Fig. 5). There were more apoptotic
PCC cells in AsPC-1-Tspan1-sh2 and PANC-1-Tspan1-sh1 groups, when
compared with that of AsPC-1-Empty and PANC-1-Empty groups,
respectively (P<0.05, Fig. 5).
However, Tspan1-siRNA showed no significant effects on the
apoptosis of human normal hTERT-HPNE cells (P>0.05).
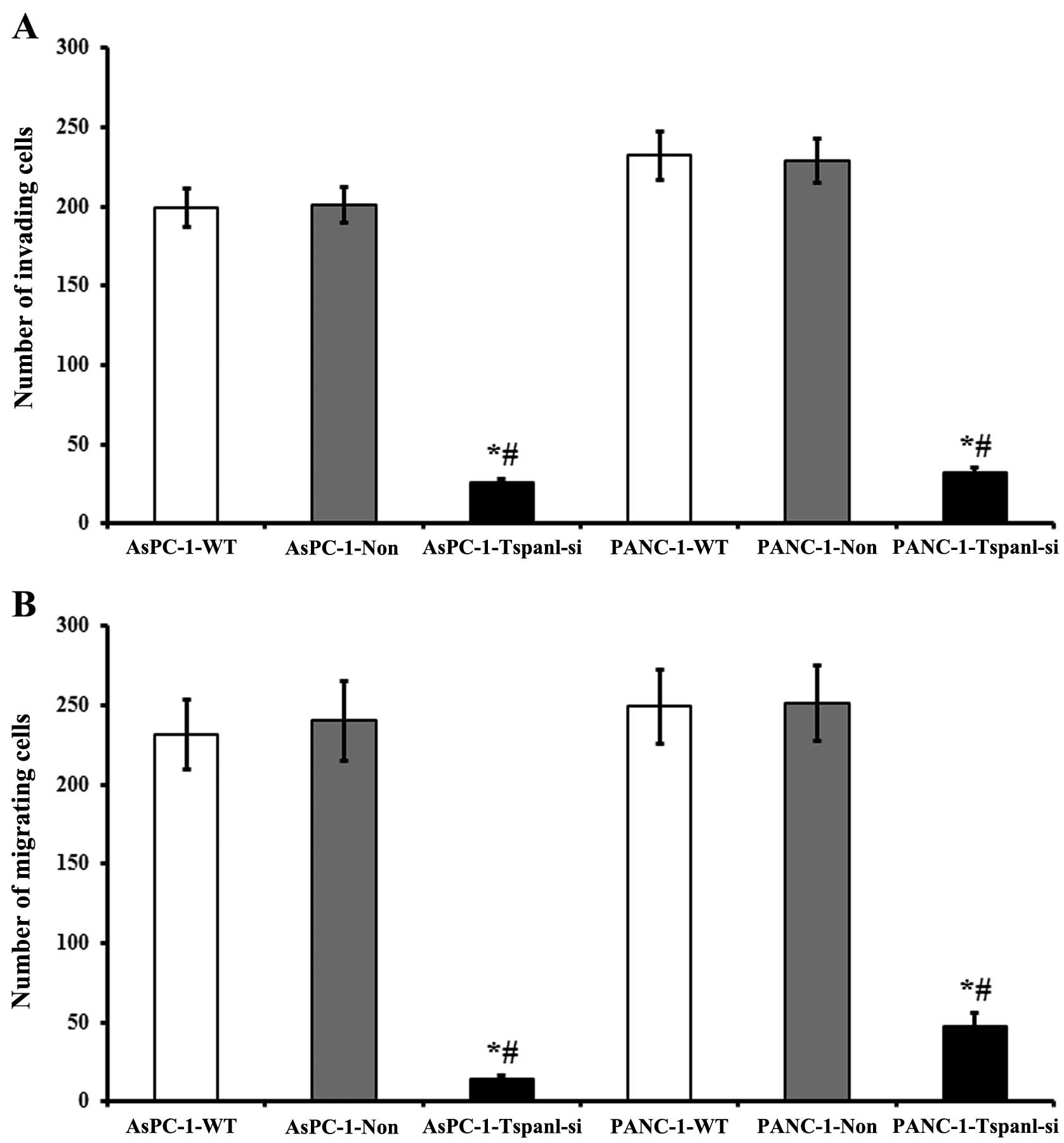
Effect of Tspan1 knockdown on PDAC cell
migration and invasion
Following knockdown, we compared the migration of
control non-transduced innocent cells (blank), non-targeting
scrambled siRNA-transduced cells (negative control), as well as the
Tspan1 knockdown cells transduced with Tspan1 targeting siRNA. The
migration assay showed that the crystal violet stained cells
significantly decreased in the Tspan1-siRNA-treated cells, compared
with that of the matched WT and non-targeting siRNA control groups
(P<0.01, Fig. 6A). The Tspan1
knockdown treatment significantly decreased the migration of the
two cell types compared to the negative and blank control. Through
the whole experimental duration, migration was not significantly
different between AsPC-1 and PANC-1 blank cells and the negative
control cells transduced with non-targeting scrambled siRNA, as
shown in Fig. 6A.
There were significant reductions in the invasion of
AsPC-1 and PANC-1 cells following Tspan1 knockdown, in comparison
with that of the control cells, respectively (P<0.005, Fig. 6B). The invasion of control cells
transduced with non-targeting siRNA, which had unchanged levels of
Tspan1, was not significantly different from the non-transduced WT
PCC cells.
Discussion
The present study focused on the possible roles of
Tspan1 involved in the tumorigenic process of PCC. Our results
revealed that Tspan1 was elevated in human PCC tissues and cell
lines. The increased Tspan1 in PCC tissues was associated with the
clinicopathological features and survival rate. Interference of
Tspan1 expression by special siRNA introduction induced significant
decline in proliferative capacity, increase in apoptosis and
reduced migration and invasion of AsPC-1 and PANC-1 cells, compared
with that of hTERT-HPNE cells. These data indicated that Tspan1 may
be involved in the pathological changes and development of PCC.
Our results revealed that expression of Tspan1 in
PCC cells displayed cytoplasmic patterns, which showed the
distribution and functional sites of the Tspan1 molecule in cells
(12). The molecule may accept
extracellular signals when located on the membrane and carry out
functions in the cytoplasm, like other tetraspanins such as CD9,
CD82 and CD63 (12,17,18).
In the present study, we found that the expression of Tspan1 in PCC
tissues was significantly higher than the normal adjacent
tissues.
Tetraspanins, a large family of ubiquitously
expressed membrane proteins, have been identified and implicated in
the regulation of cell development, differentiation, proliferation,
motility and tumor cell invasion (19–21).
In many human cancers, tumor progression was found to be associated
with an altered expression of tetraspanins (22). Tspan1, a new member of the
tetraspanins group, was found to be elevated in some tumors
(12,23–25).
Recent studies also suggested Tspan1 gene may play a role in the
proliferation of cancer cells and be associated with cancer cell
motility, implying a function of the gene in the development of
various cancer (12,23,26).
In this study, our results revealed that Tspan1 immunopositive
staining was significantly correlated with the lymph node
metastasis, pTNM stages and poor prognosis of PCC. Our data also
showed that there was a significant correlation between the Tspan1
level and overall survival rate. Similarly, other reports show that
there was a significant correlation between the Tspan1 expression
and overall survival rate, disease stages as well as the
pathological features of other various tumors, such as colorectal
cancer and cervical carcinoma (13,23).
The present data also demonstrated that
siRNA-mediated Tspan1 expression knockdown significantly inhibiting
the growth, proliferation, migration and invasion of PCC cells
in vitro, which was supported by earkier reports in
different cancer cells (10,12).
We speculated that siRNA-mediated downregulation of Tspan1
inhibited the proliferation of PCC cells in vitro by
inhibiting cell cycle progression from G1 to S phase (26). Therefore, we postulated that: i),
Tspan1 overexpression status may yield poor prognosis for PCC; and
ii), Tspan1 may play a critical role in the progression of tumor
growth and proliferation in human PCC.
In conclusion, our results show that Tspan1 was
elevated in human PCC tissues and cell lines. Interference of
Tspan1 expressions by siRNA introduction induced significant
decline in proliferative capacity and increase in apoptosis of
AsPC-1 and PANC-1 cells. This finding suggests that Tspan1 plays an
important role in PCC progression, and siRNA targeting of Tspan1
may be a potential therapeutic strategy for the treatment of PCC.
Identifying the patients with high-risk PCC by Tspan1 expression
detection would be of great benefit for improving treatment
strategies.
Acknowledgments
We thank the Labreal Bioscience and Technology Ltd.,
Company, Kunming, China for valuable contribution to parts of the
experimental design.
References
|
1
|
Bosetti C, Bertuccio P, Negri E, La
Vecchia C, Zeegers MP and Boffetta P: Pancreatic cancer: overview
of descriptive epidemiology. Mol Carcinog. 51:3–13. 2012.
View Article : Google Scholar
|
|
2
|
Vincent A, Herman J, Schulick R, Hruban RH
and Goggins M: Pancreatic cancer. Lancet. 378:607–620. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Loos M, Kleeff J, Friess H and Büchler MW:
Surgical treatment of pancreatic cancer. Ann NY Acad Sci.
1138:169–180. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Veenbergen S and van Spriel AB:
Tetraspanins in the immune response against cancer. Immunol Lett.
138:129–136. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yamamoto Y, Grubisic K and Oelgeschläger
M: Xenopus tetraspanin-1 regulates gastrulation movements and
neural differentiation in the early Xenopus embryo.
Differentiation. 75:235–45. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Huang S, Yuan S, Dong M, Su J, Yu C, Shen
Y, Xie X, Yu Y, Yu X, Chen S, et al: The phylogenetic analysis of
tetraspanins projects the evolution of cell-cell interactions from
unicellular to multicellular organisms. Genomics. 86:674–684. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
André M, Chambrion C, Charrin S, Soave S,
Chaker J, Boucheix C, Rubinstein E and Le Naour F: In situ chemical
cross-linking on living cells reveals CD9P-1 cisoligomer at cell
surface. J Proteomics. 73:93–102. 2009. View Article : Google Scholar
|
|
8
|
Hölters S, Anacker J, Jansen L,
Beer-grondke K, Dürst M and Rubio I: Tetraspanin 1 promotes
invasiveness of cervical cancer cells. Int J oncol. 43:503–512.
2013.PubMed/NCBI
|
|
9
|
Chen Y, Peng W, Lu Y, Chen J, Zhu YY and
Xi T: miR-200a enhances the migrations of A549 and SK-MES-1 cells
by regulating the expression of TSPAN1. J Biosci. 38:523–532. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Chen L, Yuan D, Zhao R, Li H and Zhu J:
Suppression of TSPAN1 by RNA interference inhibits proliferation
and invasion of colon cancer cells in vitro. Tumori. 96:744–750.
2010.
|
|
11
|
Desouki MM, Liao S, Huang H, Conroy J,
Nowak NJ, Shepherd L, Gaile DP and Geradts J: Identification of
metastasis-associated breast cancer genes using a high-resolution
whole genome profiling approach. J Cancer Res Clin oncol.
137:795–809. 2011. View Article : Google Scholar
|
|
12
|
Chen L, Zhu Y, Li H, Wang GL, Wu YY, Lu
YX, Qin J, Tuo J, Wang JL and Zhu J: Knockdown of TSPAN1 by RNA
silencing and antisense technique inhibits proliferation and
infiltration of human skin squamous carcinoma cells. Tumori.
96:289–295. 2010.PubMed/NCBI
|
|
13
|
Chen L, Zhu YY, Zhang XJ, Wang GL, Li XY,
He S, Zhang JB and Zhu JW: TSPAN1 protein expression: A significant
prognostic indicator for patients with colorectal adenocarcinoma.
World J gastroenterol. 15:2270–2276. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Jiang SH, He P, Ma MZ, Wang Y, Li RK, Fang
F, Fu Y, Tian GA, Qin WX and Zhang ZG: PNMA1 promotes cell growth
in human pancreatic ductal adenocarcinoma. Int J Clin Exp Pathol.
7:3827–3835. 2014.PubMed/NCBI
|
|
15
|
Sun GG, Wei CD, Jing SW and Hu WN:
Interactions between filamin A and MMP-9 regulate proliferation and
invasion in renal cell carcinoma. Asian Pac J Cancer Prev.
15:3789–3795. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Kramer N, Walzl A, Unger C, Rosner M,
Krupitza G, Hengstschläger M and Dolznig H: In vitro cell migration
and invasion assays. Mutat Res. 752:10–24. 2013. View Article : Google Scholar
|
|
17
|
Hemler ME: Tetraspanin functions and
associated micro-domains. Nat Rev Mol Cell Biol. 6:801–811. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zöller M: Tetraspanins: Push and pull in
suppressing and promoting metastasis. Nat Rev Cancer. 9:40–55.
2009. View
Article : Google Scholar
|
|
19
|
Mazzocca A, Carloni V, Sciammetta S,
Cordella C, Pantaleo P, Caldini A, Gentilini P and Pinzani M:
Expression of transmembrane 4 superfamily (TM4SF) proteins and
their role in hepatic stellate cell motility and wound healing
migration. J Hepatol. 37:322–330. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Furuya M, Kato H, Nishimura N, Ishiwata I,
Ikeda H, Ito R, Yoshiki T and Ishikura H: Downregulation of CD9 in
human ovarian carcinoma cell might contribute to peritoneal
dissemination: Morphologic alteration and reduced expression of
beta1 integrin subsets. Cancer Res. 65:2617–2625. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Chen Z, Mustafa T, Trojanowicz B,
Brauckhoff M, Gimm O, Schmutzler C, Köhrle J, Holzhausen HJ, Kehlen
A, Klonisch T, et al: CD82, and CD63 in thyroid cancer. Int J Mol
Med. 14:517–527. 2004.PubMed/NCBI
|
|
22
|
Klosek SK, Nakashiro K, Hara S, Goda H,
Hasegawa H and Hamakawa H: CD151 regulates HGF-stimulated
morphogenesis of human breast cancer cells. Biochem Biophys Res
Commun. 379:1097–1100. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wollscheid V, Kühne-Heid R, Stein I,
Jansen L, Köllner S, Schneider A and Dürst M: Identification of a
new proliferation-associated protein NET-1/C4.8 characteristic for
a subset of high-grade cervical intraepithelial neoplasia and
cervical carcinomas. Int J Cancer. 99:771–775. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Chen L, Wang Z, Zhan X, Li DC, Zhu YY and
Zhu J: Association of NET-1 gene expression with human
hepatocellular carcinoma. Int J Surg Pathol. 15:346–353. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Scholz CJ, Kurzeder C, Koretz K, Windisch
J, Kreienberg R, Sauer G and Deissler H: Tspan-1 is a tetraspanin
preferentially expressed by mucinous and endometrioid subtypes of
human ovarian carcinomas. Cancer Lett. 275:198–203. 2009.
View Article : Google Scholar
|
|
26
|
Leyden J, Murray D, Moss A, Arumuguma M,
Doyle E, McEntee G, O'Keane C, Doran P and MacMathuna P: Net1 and
Myeov: Computationally identified mediators of gastric cancer. Br J
Cancer. 94:1204–1212. 2006. View Article : Google Scholar : PubMed/NCBI
|