Introduction
Lung cancer is one of the most common cancers
worldwide, and it is the leading cause of cancer-related deaths in
both men and women (1). The high
incidence and mortality rate make this malignancy a major public
health issue throughout the world (2). In addition, a recent systematic review
showed that lung cancer has become one of the major causes of years
of life lost (YLLs) worldwide (3).
The development and progression of lung cancer
requires a long period of time, in which a series of molecular
events and alterations may occur (4). It has been shown that adhesion
molecules (5), angiogenic factors
(6), chemokines (7) and growth factors (8,9) appear
to be responsible for the invasion and metastasis of lung cancer
cells. However, the predominant drivers contributing to lung cancer
cell malignancy, migration and invasion, remain to be
determined.
Tumor necrosis factor receptor-associated factors
(TRAFs) are a family of adaptor proteins that couple tumor necrosis
factor receptor family to signaling pathways, and they are also
characterized to be signal transducers of Toll/interleukin-1 (IL-1)
family members, which play an important role in physiological and
pathological processes (10). To
date, 7 TRAFs have been identified, which are termed TRAF1 to 7
(11). As a key activator of
nuclear factor-κB (NF-κB), TRAF6 functions as a signal transducer
in the NF-κB pathway and activates the inhibitor of IκB kinase
(IKK) in response to pro-inflammatory cytokines (11). In addition, TRAF6, which acts as an
E3 ubiquitin ligase, catalyzes K63 polyubiquitination of TAK1 which
is required for IKK activation, and mediates the activation of
other downstream signaling molecules, thereby inducing the
constitutive activation of NF-κB (12).
Elevated TRAF6 expression has been detected in colon
cancer (13), glioma (14), osteosarcoma (15), esophageal squamous cell carcinoma
(16), breast cancer (17) and lung cancer tissues (18,19),
and TRAF has been found to promote the proliferation of these
cancer cells. In addition, inhibition of TRAF6 was reported to
result in suppression of lung cancer cell proliferation and tumor
formation (20,21), and the TRAF6 status was found to
correlate inversely with response to chemotherapy in overall
advanced non-small cell lung cancer patients (22). However, the effect of small
interfering RNA (siRNA)-induced knockdown of TRAF6 on the
biological behaviors of cancer cells remains unknown until now.
Hereby, we knocked down TRAF6 using a specific siRNA
targeting TRAF6, and assessed the effect of TRAF6 knockdown on the
proliferation, migration, invasion, cell cycle and apoptosis of
human lung cancer SPC-A1 cells. In addition, the expression of
proteins associated with tumor migration and invasion was
determined in the SPC-A1 cells transfected with TRAF6 siRNA, so as
to provide new insight into the therapeutic potential of TRAF6
against lung cancer.
Materials and methods
Cell line and culture
The human lung adenocarcinoma A549, non-small cell
lung cancer H1650, human airway epithelial Calu-3 and human lung
cancer SPC-A1 cell lines were purchased from the Cell Bank of the
Chinese Academy of Sciences (Shanghai, China), and were maintained
in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10%
fetal bovine serum (FBS) at 37°C containing 5% CO2.
TRAF6 siRNA and transfection
Log-phase SPC-A1 cells were seeded onto 6-well
plates (Corning, Inc., Corning, NY, USA) at a density of
1×106 cells/well, and incubated at 37°C for 24 h. Four
micrograms of the specific siRNA targeting TRAF6 or control siRNA
(both from Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA) was
added to 250 µl of serum-free DMEM and mixed evenly, while
10 µl of Lipofectamine 2000 (Thermo Fisher Scientific,
Waltham, MA, USA) was added to another 250 µl of serum-free
DMEM, gently mixed evenly and placed at room temperature for 5 min.
The siRNA and Lipofectamine 2000 solutions were mixed evenly, and
placed at room temperature for 20 min. Then, the mixture was
transferred to 6-well plates, mixed evenly and gently and incubated
at 37°C. Following a 6-h transfection, the cell culture solution
was changed, and to each well h 2 ml medium was added. After 48 h
of transfection, the cells were harvested for the subsequent
experiments, and non-transfected cells served as blank controls.
Transfection efficiency was verified using western blot assay.
Determination of SPC-A1 cell
proliferation using a cell proliferation assay
Following a 48-h transfection, the proliferation of
SPC-A1 cells was determined using the CellTiter 96 AQueous One
Solution Cell Proliferation assay (Promega, Madison, WI, USA)
following the manufacturer's instructions. Briefly, TRAF6 and
control siRNA-transfected, and non-transfected SPC-A1 cells were
seeded onto 96-well plates (Corning, Inc.) at a density of
1×105 cells/well, and then 20 µl of Cell Titer 96
AQueous One Solution was added to each well. Three wells containing
DMEM alone were assigned for background subtraction. The cells were
then incubated at 37°C for 30 min. The absorbance at 490 nm in each
well was then determined using a SpectraMax 340 microplate reader
(Molecular Devices Corporation, Sunnyvale, CA, USA). The absorbance
of the transfected cells was normalized to that of the
non-transfected cells (100% viability). All experiments were
replicated in triplicate.
Detection of apoptosis of SPC-A1 cells
with flow cytometry
After 48 h of transfection, TRAF6 and control
siRNA-transfected and non-transfected SPC-A1 cells were harvested,
washed twice in cold PBS and resuspended in 1X Annexin V binding
buffer (Calbiochem, San Diego, CA, USA) at a concentration of
1×106 cells/ml. Then, 500 µl of cell suspension
(4×105 cells) was transferred to a
fluorescence-activated cell sorting (FACS) tube containing 1.25
µl of Annexin V-FITC (Calbiochem). Cells were gently
vortexed and incubated at room temperature for 15 min in the dark.
Subsequently, 500 µl of cold 1X Annexin V binding buffer and
10 µl of propidium iodide (Calbiochem) were transferred and
mixed evenly, and the cells were analyzed on a FACScalibur flow
cytometer (BD Biosciences, Burlington, MA, USA). All experiments
were repeated in triplicate.
Cell cycle analysis of the SPC-A1 cells
using flow cytometry
Following a 48-h transfection, TRAF6 and control
siRNA-transfected and non-transfected SPC-A1 cells were harvested
and washed twice with PBS. Approximately 6×105 cells
were suspended in 150 µl of BD Cytofix/Cytoperm buffer
solution (BD Biosciences) at 4°C for 20 min. Subsequently, the
cells were washed twice with BD Perm/Wash buffer (BD Biosciences)
and incubated in 200 µl of dying buffer containing 0.1 mg/ml
of propidium iodide and 2 mg/ml of RNase A at 37°C for 30 min in
the dark. Cell cycle was then analyzed on a FACScalibur flow
cytometer (BD Biosciences). All procedures were repeated in
triplicate.
Transwell migration assay
The invasion of SPC-A1 cells was evaluated using a
Transwell migration assay. Briefly, SPC-A1 cells were seeded onto
6-well plates at a density of 1×105/well and incubated
at 37°C for 24 h. Then, the cells were transfected with TRAF6 or
control siRNA, while non-transfected cells served as blank
controls. Following a 24-h incubation, the cells were harvested,
digested, centrifuged, and adjusted to a cell density of
4×105 cells/ml. Approximately 0.2 ml of the SPC-A1 cells
were transferred to the Transwell chamber pre-coated with 1
µg/ml of fibronectin (Sigma-Aldrich, St. Louis, MO, USA),
with DMEM containing 1% inactivated FBS in the upper chamber and
DMEM supplemented with 10% inactivated FBS in the lower chamber,
and the Transwell chamber was placed in 24-well cell culture
inserts at 37°C for 24 h. Following a 48-h incubation, the cells in
the chamber were removed, fixed in 90% methanol and stained with
0.1% crystal violet. Five fields of vision were randomly selected,
and cells were counted under a microscope at a magnification of
×200 to determine the mean number in five inserts. All experiments
were repeated in triplicate.
Scratch wound assay
The migration of the SPC-A1 cells was detected using
the scratch wound assay. Briefly, the SPC-A1 cells transfected with
TRAF6 and control siRNA, and non-transfected cells at approximately
80% confluency were seeded onto 6-well plates, and incubated at
37°C for 24 h. Then, a vertical scratch wound was made through the
center of each well using a 10-µl pipette tip. The cells
were then washed three times with PBS to remove the scratched
cells, and fresh serum-free medium was transferred. After 12 h, the
cells were examined by light microscopy at a magnification of ×200
to determine the resealing of the cell monolayer.
Quantitative RT-PCR (qRT-PCR) assay
The relative mRNA expression of TRAF6 was determined
in A549, H1650, Calu-3 and SPC-A1 cells using a qRT-PCR assay.
Total RNA was isolated from the four lung cancer cell lines using
the RNeasy Mini kit (Qiagen Inc., Westborough, MA, USA) following
the manufacturer's protocol, and reverse-transcribed into cDNA
using the Promega Reverse Transcription system A3500 (Promega) at
42°C for 15 min, at 95°C for 5 min, and at 4°C for 5 min. qRT-PCR
assay was run in 20 µl of the reaction system containing 10
µl of 2X PCR Master Mix, 1 µl of each primer, 1
µl of cDNA, and 7 µl of nuclease-free water, on a
LightCycler Roche 480 with DyNAmo Flash SYBR-Green qPCR kit (Thermo
Fisher Scientific, Inc.) using the following primers: TRAF6
(1) forward, 5′-CTA TTC ACC AGT TAG
AGG-3′ and reverse, 5′-GCT CAC TTA CAT ACA TAC T-3′; TRAF6
(2) forward, 5′-GTT GCT GAA ATC GAA
GCA CA-3′ and reverse, 5′-CGG GTT TGC CAG TGT AGA AT-3′; and
β-actin forward, 5′-TGG CAC CAC ACC TTC TAC A-3′ and reverse,
5′-AGC ACA GCC TGG ATA GCA-3′ under the following conditions: at
95°C for 15 min; 40 cycles of at 95°C for 15 sec, at 55°C for 30
sec, and at 72°C for 1 sec; finally at 40°C for 1 min, while
β-actin served as a reference gene. Relative quantity of mRNA
expression was calculated by using the 2−ΔΔCt method.
All experiments were repeated in triplicate.
Western blotting assay
The protein expression of TRAF6, MMP-1, MMP-2,
MMP-9, Twist, TIMP-2, Slug, CD24 and CXCR4 was determined using
western blot analysis. Briefly, total protein was extracted from
the human lung cancer cells, and the protein concentration was
determined using the bicinchoninic acid (BCA) assay. An amount 25
µg of total protein was separated on a 10% SDS-PAGE gel, and
then transferred to the nitrocellulose (NC) membrane for 1 h at 60
V in transfer buffer. The membrane was then blocked with 5% nonfat
milk for 4 h at room temperature, and incubated with 1:1,000
primary rabbit ant-human TRAF6 monoclonal antibody (catalogue no:
8028S), rabbit anti-human MMP-1 monoclonal antibody (catalogue no:
sc-21731), rabbit anti-human MMP-2 polyclonal antibody (catalogue
no: sc-10736), rabbit anti-human MMP-9 polyclonal antibody
(catalogue no: sc-10737), rabbit anti-human Twist polyclonal
antibody (catalogue no: sc-15393), rabbit anti-human TIMP-2
polyclonal antibody (catalogue no: sc-5539), rabbit anti-human Slug
monoclonal antibody (catalogue no: 9585S), rabbit anti-human CD24
polyclonal antibody (catalogue no: sc-11406) (all from Santa Cruz
Biotechnology, Inc.) and rabbit anti-human CXCR4 monoclonal
antibody (catalogue no: sc-53534; Cell Signaling Technology, Inc.,
Beverly, MA, USA) overnight at 4°C on a gentle shaker. Following
washing with TBST (20 mmol/l Tris-HCl, 150 mmol/l NaCl and 0.05%
Tween-20; pH 7.4) three times, for 10 min each time, the membrane
was incubated in 1:4,000 anti-rabbit HRP-conjugated IgG antibody
(catalogue no: 7074S) or anti-mouse HRP-conjugated IgG antibody
(catalogue no: 7076S) (both from Cell Signaling Technology, Inc.)
at room temperature for 1 h. The membrane was then washed three
times for 10 min with TBST buffer (20 mmol/l Tris-HCl, 150 mmol/l
NaCl and 0.05% Tween-20; pH 7.4) and visualized with the
SuperSignal West Pico kit (Thermo Fisher Scientific).
Electrophoretic mobility shift assay
(EMSA)
Nuclear protein was extracted as described
previously, and all nuclear extractions contained 1X Roche Mini
Complete protease inhibitor cocktail solution (Roche Applied
Science, Burgess Hill, UK). A 22-mer NF-κB consensus
oligonucleotide (Promega) was labeled according to the
manufacturer's instructions using the DIG gel-shift kit (Roche
Applied Science). An amount of 10 mg of nuclear extract was used
per binding reaction as detailed in the manufacturer's protocol,
with some control reactions using NF-κB unlabeled oligonucleotides
or nonspecific negative control OCT1 consensus oligonucleotides
(Promega) for competition or the NF-κB p65 antibody (Thermo Fisher
Scientific) for supershift reactions. Reactions were run on 6%
(v/v) DNA retardation gels (Invitrogen, Paisley, UK), and then
blotted and cross-linked onto positively charged nylon membranes
(Roche Applied Science) before immunodetection.
Statistical analysis
All data are expressed as mean ± standard deviation
(SD), and all statistical analyses were performed using the
statistical software SPSS version 16.0 (SPSS Inc., Chicago, IL,
USA). Differences in means were tested for statistical significance
with the Student's t-test, with a p-value <0.05 indicative of
statistical significance.
Results
TRAF6 expression and K63-linked
ubiquitination of TRAF6
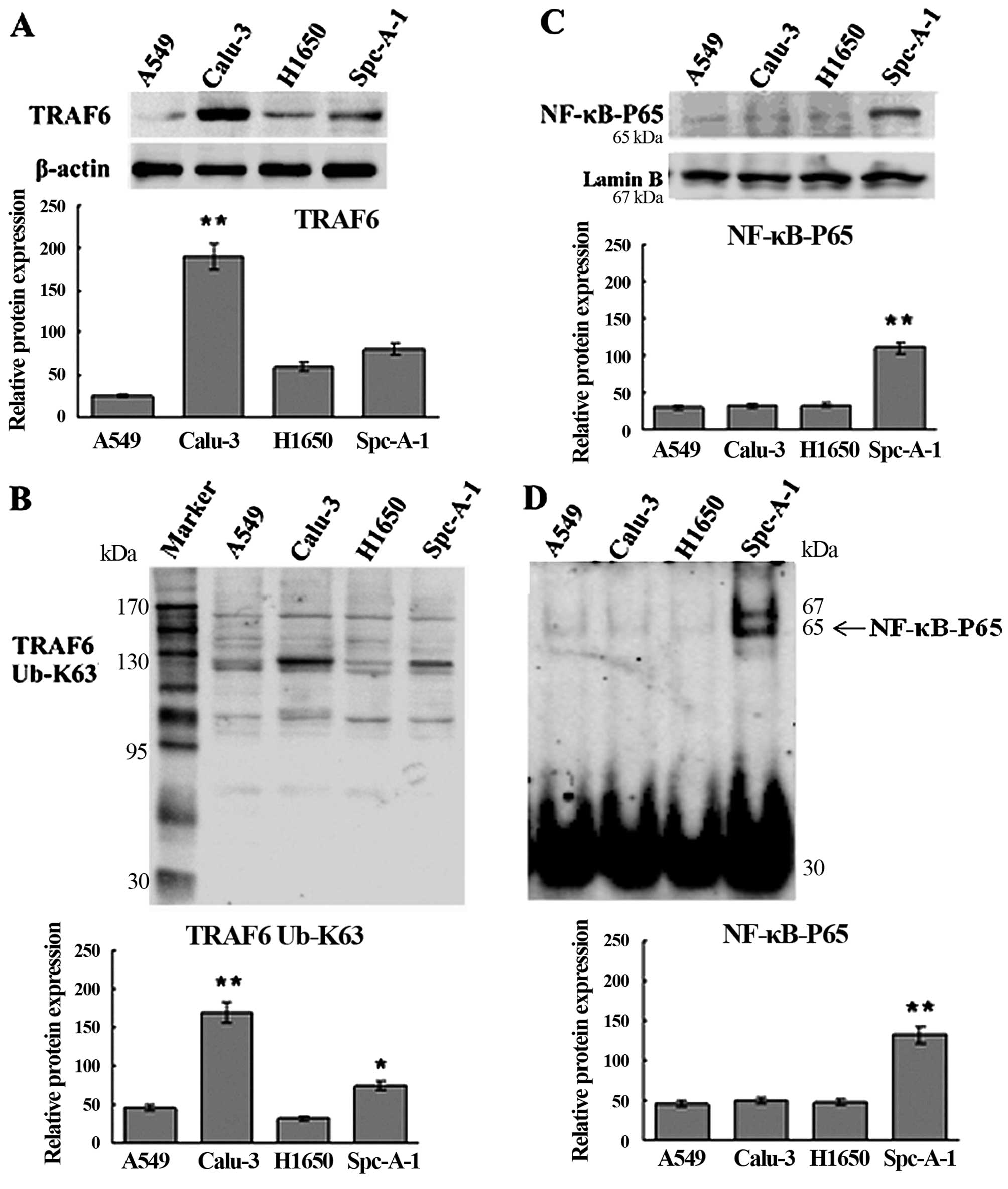
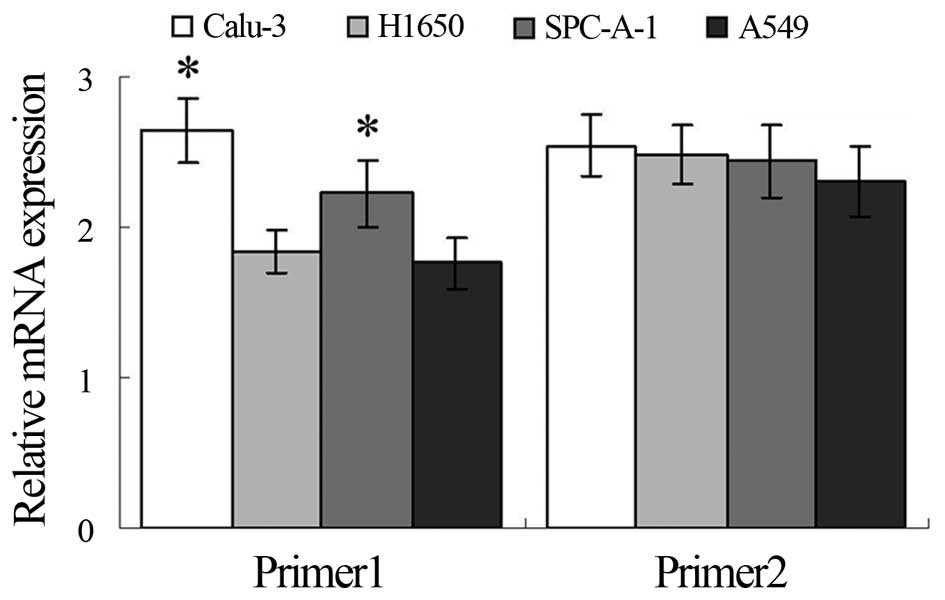
Western blotting and qRT-PCR assays showed higher
expression of TRAF6 in the Calu-3 and SPC-A1 cells than the
expression level in the A549 and H1650 cells at both the
translational and transcriptional levels (Figs. 1A and 2), and K63-linked ubiquitination of TRAF6
was detected in the SPC-A1 cells, as revealed by western blot
analysis (Fig. 1B).
High expression of TRAF6 is associated
with constitutive activation of NF-κB in the SPC-A1 cells
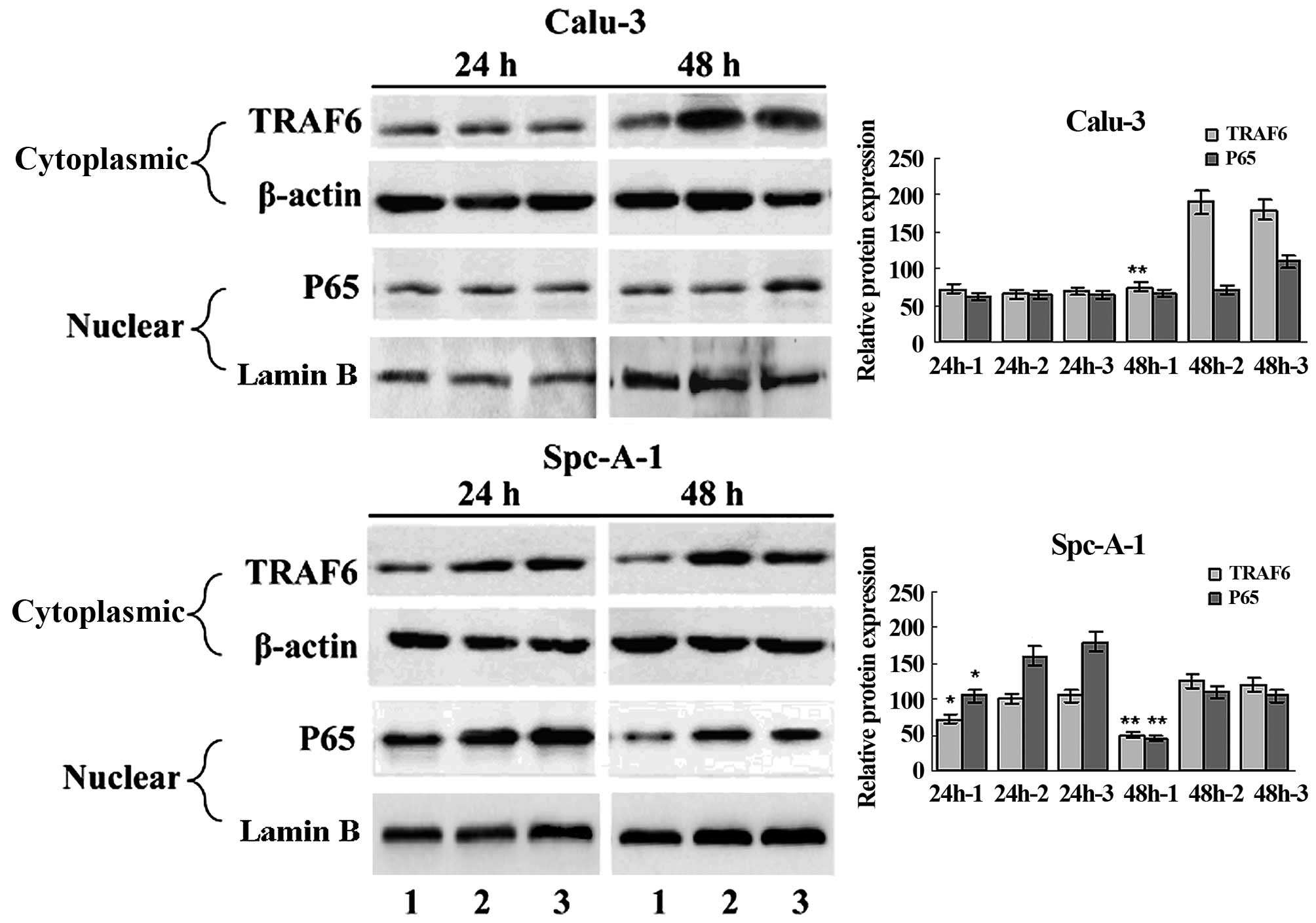
EMSA and western blot analysis showed constitutive
NF-κB activation in the SPC-A1 cells (Fig. 1C and D), and a significant
inhibition of NF-κB activation in the SPC-A1 cells 48 h
post-transfection with TRAF6 siRNA (Fig. 3). These findings indicate that high
expression of TRAF6 was associated with constitutive activation of
NF-κB in the SPC-A1 cells.
siRNA-induced knockdown of TRAF6 promotes
the apoptosis of SPC-A1 cells but does not affect cell
proliferation and cell cycle
Following a 48-h transfection, the cell
proliferation assay revealed no significant difference in the
proliferation of the TRAF6 siRNA-transfected, control
siRNA-transfected and non-transfected SPC-A1 cells. Similarly,
siRNA-induced knockdown of TRAF6 showed no clear-cut effect on the
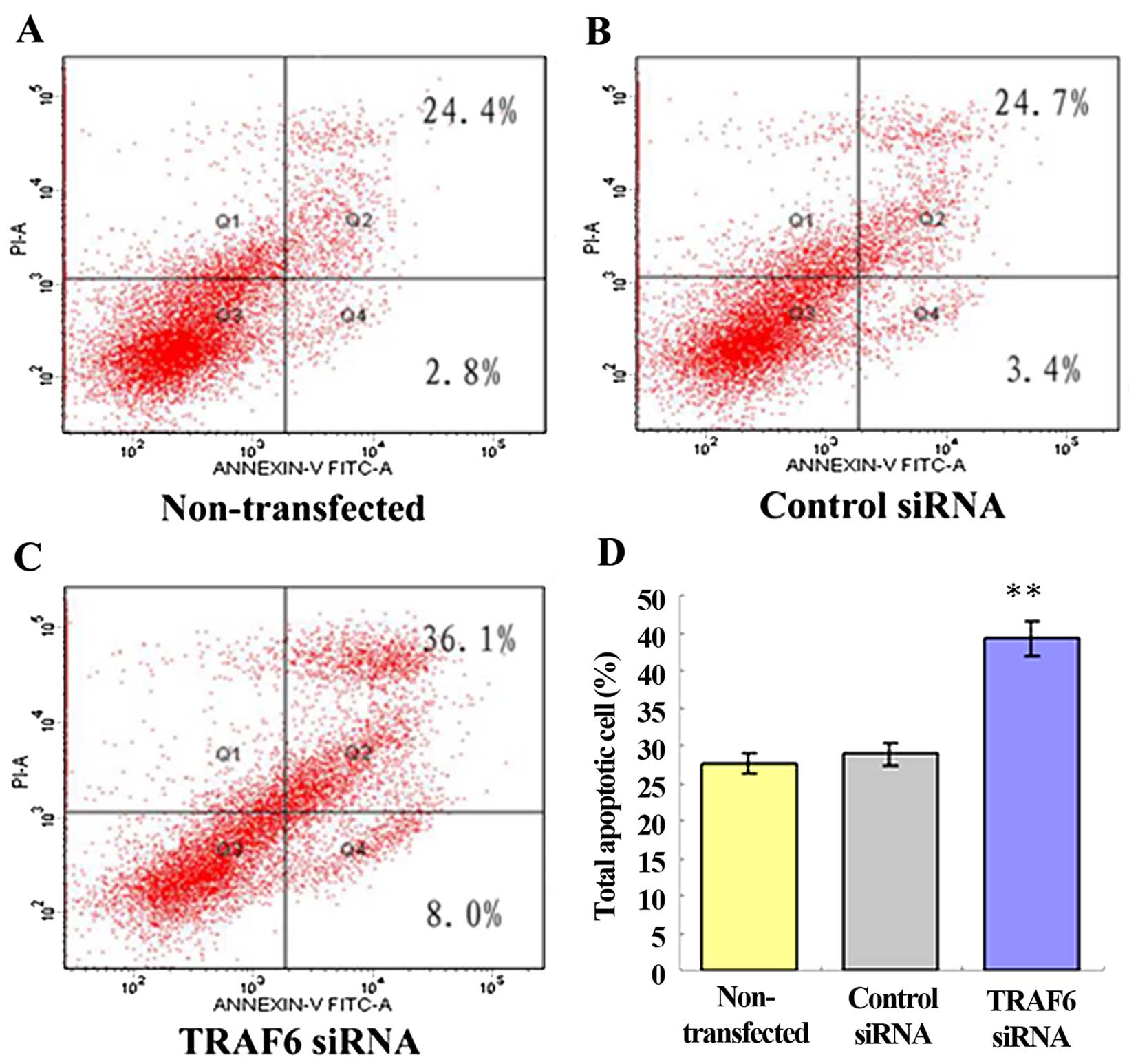
cell cycle of the SPC-A1 cells (data not shown). After 48 h of
transfection, flow cytometry showed a 44.0±0.98% apoptosis rate in
the TRAF6 siRNA-transfected SPC-A1 cells, a 27.2±1.86% apoptosis
rate in the control siRNA-transfected cells, and a 28.1±1.45%
apoptosis rate in the non-transfected cells (Fig. 4). The results demonstrated that
siRNA-induced knockdown of TRAF6 resulted in an ~1.5-fold increase
in the apoptosis of the SPC-A1 cells (P<0.05).
siRNA-induced knockdown of TRAF6
suppresses cell migration and invasion and reduces CD24 and CXCR4
protein expression in SPC-A1 cells
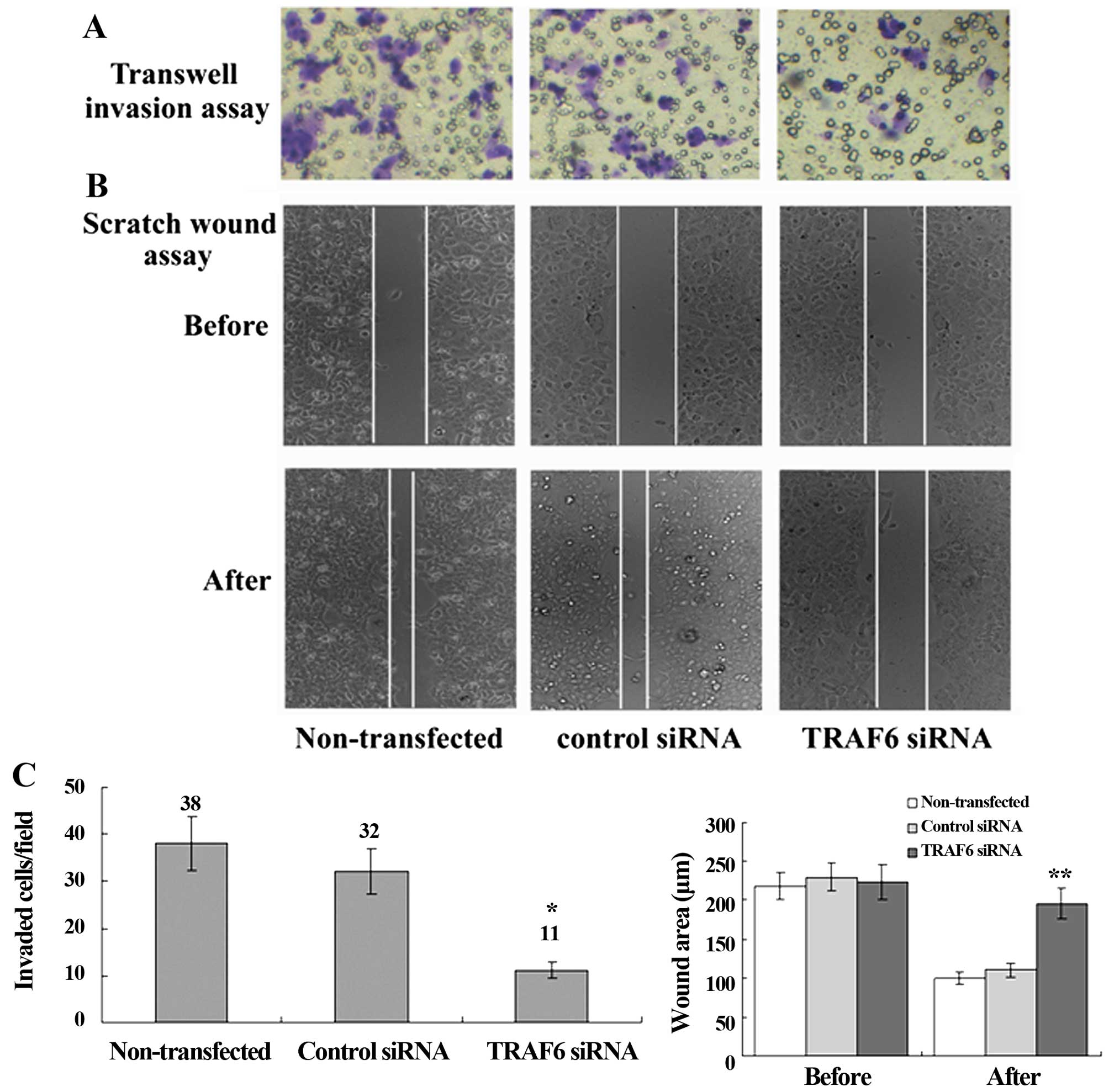
Transwell migration assay revealed a significant
reduction in the invasion of the TRAF6 siRNA-transfected SPC-A1
cells as compared to the control siRNA-transfected or
non-transfected cells, and scratch wound assay showed a slower
migration of the TRAF6 siRNA-transfected cells than the control
siRNA-transfected or non-transfected cells (Fig. 5). Western blot analysis revealed a
marked reduction in the protein expression of CD24 and CXCR4 in the
TRAF6 siRNA-transfected SPC-A1 cells as compared to that in the
control siRNA-transfected or the non-transfected cells, while no
significant difference was detected in the protein expression of
MMP-1, MMP-2, MMP-9, Twist, TIMP-2 or Slug (Fig. 6). These findings demonstrated that
siRNA-induced knockdown of TRAF6 reduced CD24 and CXCR4 protein
expression in the SPC-A1 cells.
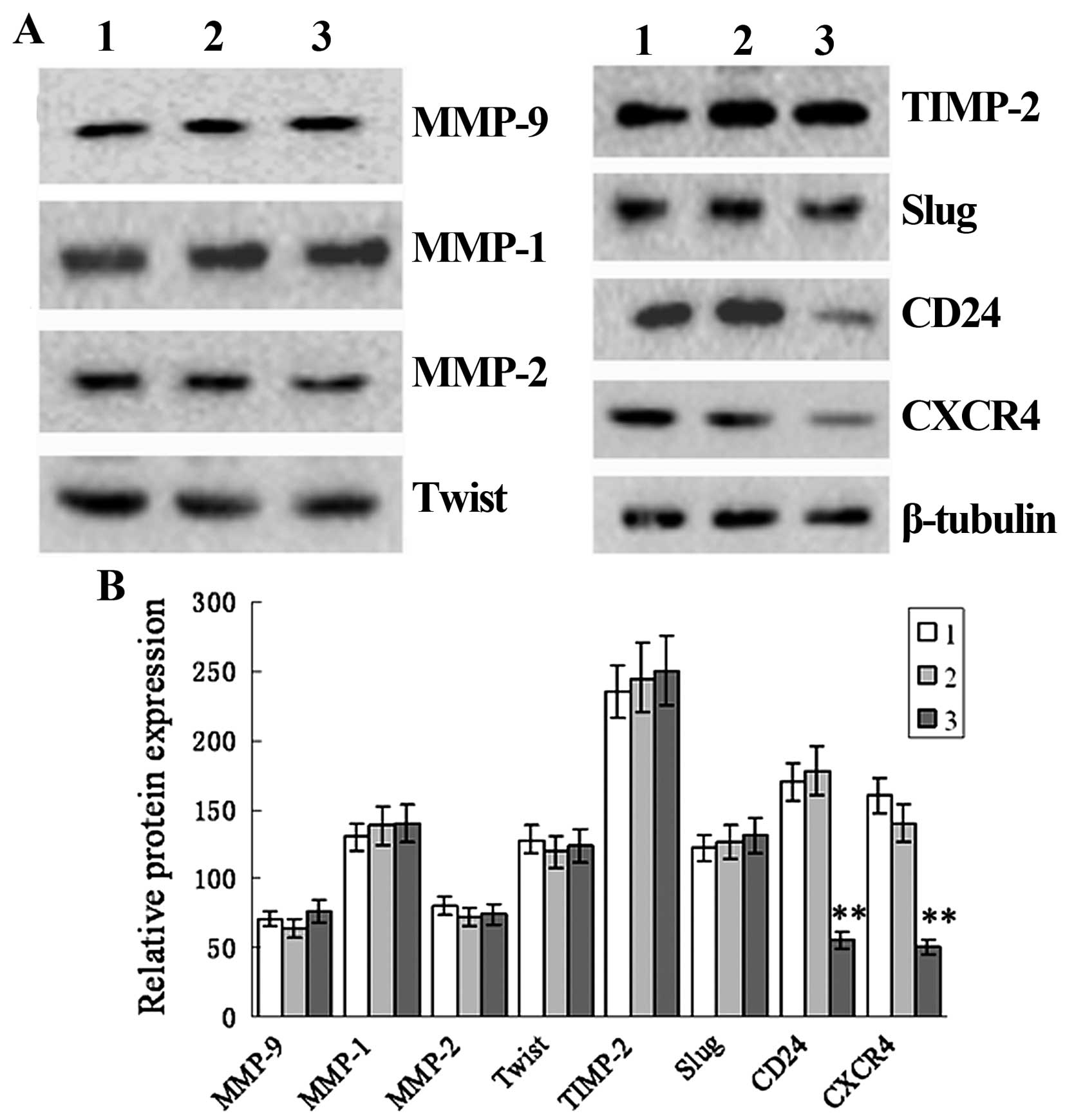 | Figure 6siRNA-induced TRAF6 knockdown
decreases CD24 and CXCR4 expression in the SPC-A1 cells. SPC-A1
cells were transfected by TRAF6 siRNA or control siRNA. After 48 h,
cell extracts were immunoblotted to detect the indicated protein.
β-actin was used as a loading control. Lane 1, TRAF6 siRNA; lane 2,
non-transfected; lane 3, control siRNA. (A) Western blot analysis
shows the protein expression of MMP-9, MMP-1, MMP-2, Twist, TIMP-2,
Slug, CD24, CXCR4 and β-tubulin. (B) Relative protein expression of
CD24 and CXCR4 in non-transfected cells and cells transfected with
control siRNA and TRAF6 siRNA. 1, TRAF6 siRNA; 2, non-transfected;
3, control siRNA. **P<0.01 vs. the non-transfected
group. |
Discussion
As a member of the TRAF family, TRAF6 expression has
been shown to be upregulated in cancer cells (14–16).
Tumor tissue assay analysis demonstrated that TRAF6 was upregulated
in colon cancers when compared with that in adjacent non-cancerous
tissues (13); western blotting
revealed upregulated TRAF6 protein expression in glioma U251,
U-87MG, LN-18, and U373 cell lines as compared to that in the
noncancerous human glial SVG p12 cell line (14). Using RT-PCR and western blot
analysis, TRAF6 expression was found to be markedly upregulated in
osteosarcoma tissues than that in normal bone tissues at both the
transcriptional and translational levels (P<0.05) (15), and ELISA assay showed significantly
higher serum TRAF6 expression in triple-negative breast cancer
patients when compared with that in an obese control group (0.90
vs. 0.73 ng/ml; P=0.033) (17). In
the present study, we determined the mRNA and protein expression of
TRAF6 in human lung adenocarcinoma A549, non-small cell lung cancer
H1650, human airway epithelial Calu-3 and human lung cancer SPC-A1
cell lines, and western blotting and qRT-PCR assays showed
clear-cut upregulation of TRAF6 mRNA and protein expression in the
Calu-3 and SPC-A1 cells when compared with that in the A549 and
H1650 cells. The results are consistent with a previous study
reporting significantly higher TRAF6 expression in lung cancer
tissues than that in normal lung tissues, as revealed by
immunohistochemical detection (18). Our findings further validated the
important role of TRAF6 in the development of lung cancer.
TRAF6 contains an amino terminal RING domain that
comprises the core of the ubiquitin ligase catalytic domain
(10). As signal transducers of
Toll/IL-1 family members, TRAF6 is recruited to the receptor
complexes and forms oligomers upon ligand binding to IL-1R or
Toll-like receptors (TLRs), and TRAF6 oligomerization activates its
ligase activity, leading to K63-linked polyubiquitination of
targets including TRAF6 itself (23). TRAF6 appears to participate in the
activation of NF-κB (24), which
plays an important role in the regulation of many genes involved in
inflammation, the immune response, cellular proliferation,
apoptosis, tumorigenesis and invasion (25–27).
It has been shown that TRAF6 is overexpressed and bridges RAS and
NF-κB signaling in lung cancer tumors, which is important for
RAS-mediated oncogenesis (21). In
the present study, K63-linked ubiquitination of TRAF6 was detected
in human lung cancer SPC-A1 cells, and EMSA and western blot
analysis revealed constitutive NF-κB activation in the SPC-A1
cells, and a marked inhibition of NF-κB activation in the SPC-A1
cells 48 h post-transfection with TRAF6 siRNA, indicating that high
expression of TRAF6 is associated with constitutive activation of
NF-κB in SPC-A1 cells. Our findings further confirm the oncogenic
role of TRAF6 in human lung cancer.
It has been demonstrated that TRAF6 promotes cancer
cell proliferation, while knockdown of TRAF6 may decrease cell
viability, suppress cell proliferation, invasion and migration, and
promote cell apoptosis (14,15).
To assess the role of TRAF6 in proliferation, apoptosis, cell
cycle, invasion and migration of lung cancer cells, we generated a
human lung cancer SPC-A1 cell line in which TRAF6 was depleted by
using the technique of RNAi. Subsequently, the effects of TRAF6
knockdown on cell viability, apoptosis, cell cycle, invasion and
migration of the SPC-A1 cells were determined using a cell
proliferation assay, flow cytometry, Transwell invasion assay and
scratch wound assay. Our findings showed that siRNA-induced
knockdown of TRAF6 promoted the apoptosis of SPC-A1 cells, but had
little effect on cell proliferation or cell cycle. Our results are
different from a previous study reporting that depletion of TRAF6
expression with short-hairpin RNA (shRNA) decreased viability,
suppressed proliferation and invasion, and promoted apoptosis of
human lung adenocarcinoma A549 cells (20). Such a difference may be attributed
to the following factors: i) the different cell lines used; and ii)
experimental errors. Thus, further studies are required to validate
TRAF6 knockdown on the biological behaviors of lung cancers.
However, Sun and colleagues (13)
also observed little effect of TRAF6 knockdown on the survival of
colon cancer cells. In the present study, we found that knockdown
of TRAF6 inhibited the migration and invasion of SPC-A1 cells,
which was in agreement with the findings reported in human lung
adenocarcinoma A549 cells (20).
Similar results were observed in colon cancer (13), glioma (14), osteosarcoma (15) and esophageal squamous cells
(16).
To address the mechanisms underlying the promotion
of SPC-A1 cell apoptosis and inhibition of cell migration and
invasion by TRAF6 knockdown, we determined the expression of
proteins associated with tumor migration and invasion in TRAF6
siRNA-transfected SPC-A1 cells. Western blot analysis revealed that
knockdown of TRAF6 caused a marked reduction in the protein
expression of CD24 and CXCR4, but had little effect on the protein
expression of MMP-1, MMP-2, MMP-9, Twist, TIMP-2 or Slug. CD24 and
CXCR4 are known targets of the NF-κB signaling pathway (12), and NF-κB signaling has been
demonstrated to promote cancer cell apoptosis by mediating CD24 and
CXCR4 expression (28–30). It is therefore postulated that
siRNA-induced knockdown of TRAF6 may affect the apoptosis of SPC-A1
cells by mediating CD24 and CXCR4 expression; however, further
studies are required to investigate the exact mechanisms
responsible for the promotion of SPC-A1 cell apoptosis induced by
TRAF6 knockdown.
In conclusion, the results of the present study
demonstrated that TRAF6 is upregulated in human lung cancer cells,
and TRAF6 may be involved in the apoptosis, migration and invasion
of SPC-A1 cells. In addition, siRNA-induced TRAF6 knockdown
inhibits the invasion of lung cancer cells and promotes apoptosis
through the NF-κB-CD24/CXCR4 signaling pathway. These findings
suggest that TRAF6 may be a promising target for the therapy of
lung cancer. However, the present study was designed using an in
vitro cell assay. Further studies are required in lung cancer
animal models to validate the findings and hypotheses from the
present study.
Acknowledgments
The present study was funded by the National
Clinical Key Specialty Construction Program, the Medical Innovation
Foundation of Fujian Province (grant no. 2011-cxb-16) and the
Natural Science Foundation of Fujian Province (grant no.
2013J01284).
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Herbst RS, Heymach JV and Lippman SM: Lung
cancer. N Engl J Med. 359:1367–1380. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
GBD 2013 Mortality and Causes of Death
Collaborators: Global, regional, and national age-sex specific
all-cause and cause-specific mortality for 240 causes of death,
1990–2013: A systematic analysis for the Global Burden of Disease
Study 2013. Lancet. 385:117–171. 2015. View Article : Google Scholar
|
|
4
|
Miller YE: Pathogenesis of lung cancer:
100 year report. Am J Respir Cell Mol Biol. 33:216–223. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Syrigos KN, Katirtzoglou N, Kotteas E and
Harrington K: Adhesion molecules in lung cancer: Implications in
the pathogenesis and management. Curr Pharm Des. 14:2173–2183.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Gazdar AF and Minna JD: Angiogenesis and
the multistage development of lung cancers. Clin Cancer Res.
6:1611–1612. 2000.PubMed/NCBI
|
|
7
|
Balkwill F: Cancer and the chemokine
network. Nat Rev Cancer. 4:540–550. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hodkinson PS, Mackinnon A and Sethi T:
Targeting growth factors in lung cancer. Chest. 133:1209–1216.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sethi T and Woll PJ: Growth factors and
lung cancer. Cancer Treat Res. 72:111–130. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bradley JR and Pober JS: Tumor necrosis
factor receptor-associated factors (TRAFs). Oncogene. 20:6482–6491.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chung JY, Park YC, Ye H and Wu H: All
TRAFs are not created equal: Common and distinct molecular
mechanisms of TRAF-mediated signal transduction. J Cell Sci.
115:679–688. 2002.PubMed/NCBI
|
|
12
|
Hayden MS and Ghosh S: Shared principles
in NF-kappaB signaling. Cell. 132:344–362. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Sun H, Li X, Fan L, Wu G, Li M and Fang J:
TRAF6 is upregulated in colon cancer and promotes proliferation of
colon cancer cells. Int J Biochem Cell Biol. 53:195–201. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Peng Z, Shuangzhu Y, Yongjie J, Xinjun Z
and Ying L: TNF receptor-associated factor 6 regulates
proliferation, apoptosis, and invasion of glioma cells. Mol Cell
Biochem. 377:87–96. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Meng Q, Zheng M, Liu H, Song C, Zhang W,
Yan J, Qin L and Liu X: TRAF6 regulates proliferation, apoptosis,
and invasion of osteosarcoma cell. Mol Cell Biochem. 371:177–186.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Han Q, Yao F, Zhong C and Zhao H: TRAF6
promoted the metastasis of esophageal squamous cell carcinoma.
Tumour Biol. 35:715–721. 2014. View Article : Google Scholar
|
|
17
|
Bilir C, Engin H, Can M, Likhan S,
Demirtas D, Kuzu F and Bayraktaroglu T: Increased serum tumor
necrosis factor receptor-associated factor-6 expression in patients
with non-metastatic triple-negative breast cancer. Oncol Lett.
9:2819–2824. 2015.PubMed/NCBI
|
|
18
|
Zhang XL, Dang YW, Li P, Rong MH, Hou XX,
Luo DZ and Chen G: Expression of tumor necrosis factor
receptor-associated factor 6 in lung cancer tissues. Asian Pac J
Cancer Prev. 15:10591–10596. 2014. View Article : Google Scholar
|
|
19
|
Lin G and Huang C, Su G, Hu H, Xu H and
Huang C: Effect of TRAF6 downregulation on malignant biological
behavior of lung cancer cell lines. Zhongguo Fei Ai Za Zhi.
18:661–667. 2015.In Chinese. PubMed/NCBI
|
|
20
|
Zhong L, Cao F and You Q: Effect of TRAF6
on the biological behavior of human lung adenocarcinoma cell.
Tumour Biol. 34:231–239. 2013. View Article : Google Scholar
|
|
21
|
Starczynowski DT, Lockwood WW, Deléhouzée
S, Chari R, Wegrzyn J, Fuller M, Tsao MS, Lam S, Gazdar AF, Lam WL,
et al: TRAF6 is an amplified oncogene bridging the RAS and NF-κB
pathways in human lung cancer. J Clin Invest. 121:4095–4105. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Liu H, Zhang T, Ye J, Li H, Huang J, Li X,
Wu B, Huang X and Hou J: TNF receptor-associated factor 6 in
advanced non-small cell lung cancer: Clinical and prognostic
implications. J Cancer Res Clin Oncol. 138:1853–1863. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Chen ZJ: Ubiquitin signalling in the
NF-kappaB pathway. Nat Cell Biol. 7:758–765. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Hoesel B and Schmid JA: The complexity of
NF-κB signaling in inflammation and cancer. Mol Cancer. 12:862013.
View Article : Google Scholar
|
|
25
|
DiDonato JA, Mercurio F and Karin M: NF-κB
and the link between inflammation and cancer. Immunol Rev.
246:379–400. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Karin M: Nuclear factor-kappaB in cancer
development and progression. Nature. 441:431–436. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Karin M: NF-kappaB as a critical link
between inflammation and cancer. Cold Spring Harb Perspect Biol.
1:a0001412009. View Article : Google Scholar
|
|
28
|
Esencay M, Newcomb EW and Zagzag D: HGF
upregulates CXCR4 expression in gliomas via NF-kappaB: Implications
for glioma cell migration. J Neurooncol. 99:33–40. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ju JH, Jang K, Lee KM, Kim M, Kim J, Yi
JY, Noh DY and Shin I: CD24 enhances DNA damage-induced apoptosis
by modulating NF-κB signaling in CD44-expressing breast cancer
cells. Carcinogenesis. 32:1474–1483. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
LaRocca TJ, Fabris F, Chen J, Benhayon D,
Zhang S, McCollum L, Schecter AD, Cheung JY, Sobie EA, Hajjar RJ,
et al: Na+/Ca2+ exchanger-1 protects against
systolic failure in the Akitains2 model of diabetic
cardiomyopathy via a CXCR4/NF-κB pathway. Am J Physiol Heart Circ
Physiol. 303:H353–H367. 2012. View Article : Google Scholar : PubMed/NCBI
|