Introduction
Neuroblastoma (NB) is an embryonal tumor originating
from the sympathetic nervous system including the adrenal medulla
and paravertebral nerve trunk. NB is the most common extracranial
solid tumor in children and accounts for approximately 15% of all
pediatric cancer deaths (1). NB
displays a wide variety of biological and clinical features with a
heterogeneous prognosis, ranging from spontaneous regression to
rapid tumor progression and death. For example, NB diagnosed at 12
months of age or younger typically regresses and/or spontaneously
differentiates, whereas NB in patients older than 12 months
typically become aggressive and are associated with an unfavorable
prognosis. MYCN gene amplification is often observed in
advanced NB. MYCN encodes a sequence-specific transcription
factor and transactivates its target genes implicated in crucial
cellular processes such as cell cycle progression, proliferation,
apoptosis, differentiation and metabolism (2). In addition to MYCN gene
amplification, a growing body of evidence indicates that gain of
chromosome 17q (3) and deletion of
the distal part of chromosome 1p are tightly associated with poor
prognosis in patients with NB (4).
Unfortunately, despite multimodal therapy such as chemotherapy,
surgical tumor removal, radiation therapy and hematopoietic stem
cell transplantation, the 5-year survival rate of patients with
high-risk NB remains less than 40% (1). Consequently, there is an urgent
clinical need to clarify the precise molecular mechanisms
underlying advanced NB and develop novel treatment strategies.
To identify novel cancer-related genes, we have
screened genome areas based on aberrant methylation status in mouse
skin tumors compared with the normal skin. During the analysis of
mouse skin cancers induced by a 2-stage carcinogenesis protocol
using 7.12-dimethylbenz(a)anthracene (DMBA) and
12-O-tetradecanoylphorbol-13-acetate (TPA), several skin
tumor-specific differentially-methylated regions (ST-DMRs) and
genes differentially expressed in tumor tissues compared with the
normal tissues were identified. Subsequent studies revealed that
some of the ST-DMRs such as zygote arrest 1 (ZAR1), are
aberrantly methylated and genes within these loci are expressed in
numerous types of human tumor tissues including NB (5–8).
TFAP2E, located within one of the ST-DMRs,
has been shown to be highly methylated in SCC tissues. In our
recent study (9), we demonstrated
that the expression levels of TFAP2E are significantly lower
in SCC tissues than in the normal skin. TFAP2E encodes a
nuclear transcription factor activator protein-2 (AP-2) epsilon and
is largely expressed in normal skin tissues (10). Intriguingly, aberrant methylation of
TFAP2E genomic locus and/or its expression is associated
with prognostic outcome or drug resistance in certain human tumors.
Indeed, hypermethylation and lower expression levels of
TFAP2E have been shown to correlate with resistance to
fluorouracil in patients with colon cancer (11). In gastric cancer, hypermethylation
and lower expression levels of TFAP2E were much more
frequently observed in tumors with lower differentiation grades
(12). In addition,
hypermethylation of TFAP2E has been frequently detected in
genomic DNA prepared from urine samples of prostate cancer patients
relative to that of urine samples of normal males (13).
In the present study, we focused on TFAP2E
and extended the findings of our previous study of NB. We
demonstrated that TFAP2E plays a vital role in the
regulation of DNA damage responses in NB.
Materials and methods
Cell lines and culture conditions
Human neuroblastoma-derived NB1 and NB9 cells were
obtained from RIKEN Cell Bank (Ibaraki, Japan). Cells were
maintained in RPMI-1640 (Nacalai Tesque, Kyoto, Japan) supplemented
with 10% (NB1) or 15% (NB9) heat-inactivated fetal bovine serum
(FBS; Nichirei Bioscience, Tokyo, Japan), 100 IU/ml of penicillin
(Life Technologies, Carlsbad, CA, USA) and 100 µl/ml of
streptomycin (Life Technologies).
Cell viability
NB1 cells were seeded in 24-well plates at a density
of 2×104 cells/well and allowed to attach. Cells were
then transfected with control siRNA or with TFAP2E siRNA
using Lipofectamine 3000 (Life Technologies) according to the
manufacturers instructions. Twenty-four hours after transfection,
cells were treated with adriamycin (ADR), cisplatin (CDDP),
hydrogen peroxide (H2O2) or irradiated with
ionizing radiation (IR) (X-ray linear accelerator MBR-1520R-3;
Hitachi Medical, Tokyo, Japan). Twenty-four hours post-treatment,
number of viable cells was determined using a Millipore
Sceptor.
FACS analysis
For the analysis of cell cycle distribution,
floating and attached cells were collected 24 h after ADR exposure.
Cells were washed in phosphate-buffered saline (PBS) and then fixed
in 75% ice-cold ethanol for 2 h. After washing in PBS, cells were
incubated with 0.1% FBS, 25 µg/ml of propidium iodide and 200 µg/ml
of RNase A in PBS for 15 min at room temperature in the dark and
subsequently subjected to the flow cytometric analysis
(FACSCallibur). The analysis was performed 3 times and the flow
cytometry graphs were created by calculating the total data.
The percentage of apoptotic cells was determined 24
h after the ADR exposure by using Annexin V-FITC apoptosis
detection kit (BioVision, Inc., Milpitas, CA, USA) according to the
instructions of the manufacturer. Fluorescence was detected by flow
cytometry. The analysis was conducted 3 times and the average
percentages of apoptotic cells were calculated.
Immunoblotting
Cells were lysed in RIPA buffer containing a
protease inhibitor cocktail (Nacalai Tesque). The protein
concentration of lysates was measured using Bio-Rad DC kits
(Bio-Rad Laboratories, Hercules, CA, USA). Cell lysates (20 µg of
protein) were separated by 4–12% SDS-polyacrylamide gel
electrophoresis and then electro-transferred onto Immobilon-P
membrane (Millipore). Membranes were blocked with Blocking One
(Nacalai Tesque) overnight at 4°C and incubated with polyclonal
anti-TFAP2E (ProSci, Inc., Poway, CA, USA), polyclonal anti-caspase
3 (Cell Signaling Technology, Beverly, MA, USA), polyclonal
anti-PARP (Cell Signaling Technology), monoclonal anti-p53 (DO-1;
Santa Cruz Biotechnology, Santa Cruz, CA, USA), polyclonal
anti-phospho-p53 at Ser-15 (Cell Signaling Technology), polyclonal
anti-p21WAF1 (H-164; Santa Cruz Biotechnologies),
monoclonal anti-γ-H2AX (2F3; BioLegend, San Diego, CA, USA) or with
polyclonal anti-β-actin antibody (Sigma-Aldrich, St. Louis, MO,
USA) at 4°C. Twenty-four hours after incubation, membranes were
washed in PBS containing 0.1% Tween-20 (PBS-T) followed by
incubation with horseradish peroxidase-conjugated secondary
antibody (GE Healthcare Life Sciences, Buckinghamshire, UK) for 1 h
at room temperature. The membrane was washed extensively in PBS-T
and then treated with Chemi-Lumi One Super (Nacalai Tesque) to
visualize immunoreactive signals using ImageQant LAS 4000 (Fujifilm
Corp., Tokyo, Japan).
Quantitative real-time PCR (qPCR)
Total RNA was isolated from cells using RNeasy mini
kits (Qiagen, Valencia, CA, USA) according to the manufacturers
instructions. For cDNA synthesis, 500 ng of total RNA were
reverse-transcribed using iScript cDNA synthesis system (Bio-Rad
Laboratories). qPCR was performed using a SYBR Premix Ex Taq™system
(Takara, Shiga, Japan) according to the manufacturers
recommendations. Relative gene expression was expressed as relative
fold-change in mRNA levels compared with reference cDNA. Primer
sets used for qPCR-based amplification were as follows:
TFAP2E, 5-cggttacgtctgtgagacgga-3 (sense) and
5-tgcaaactccttgcagatctgc-3 (antisense); CDKN1A (encoding
p21WAF1), 5-gcagaccagcatgacagattt-3 (sense) and
5-ggattagg gcttcctcttgga-3 (antisense); and 18S rRNA,
5-ggccctgtaattgga atgagtc-3 (sense) and 5-ccaagatccaactacgagctt-3
(antisense). 18S rRNA was used as a reference gene.
Statistical analysis
Statistical analyses were performed using Student's
t-test. Data were presented as means ± SD from at least three
independent experiments. P<0.05 was considered statistically
significant.
Results
A lower expression level of TFAP2E is
closely related to a poor prognosis of the patients with
neuroblastoma (NB)
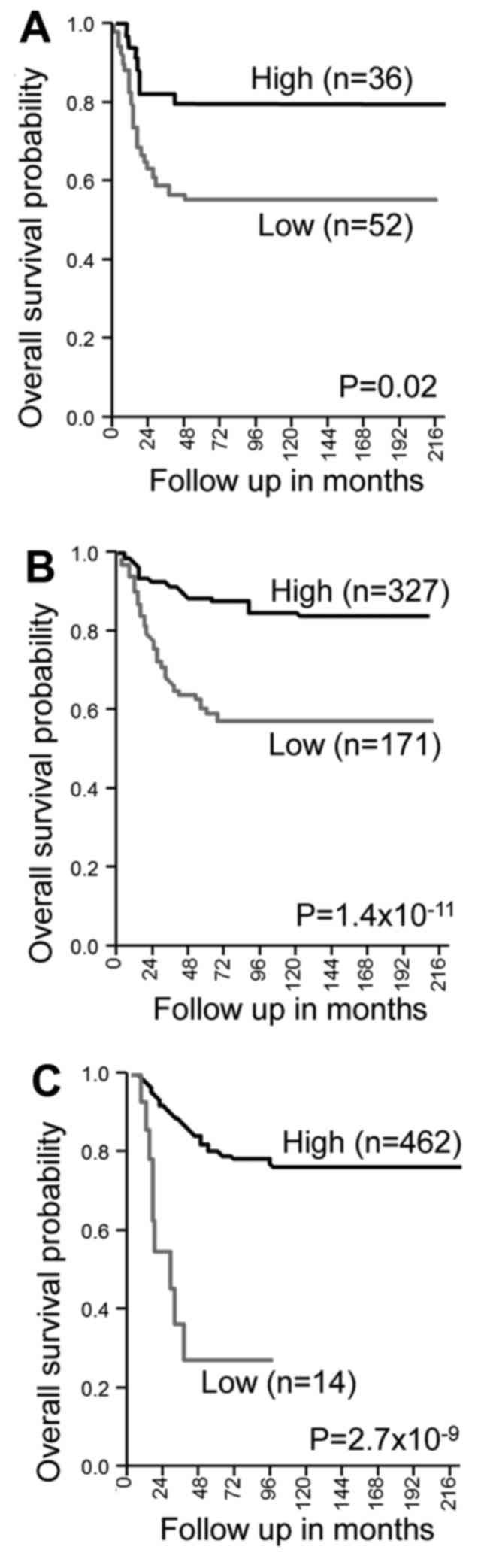
To examine the clinical significance of TFAP2E in
the genesis and/or progression of NB, we performed Kaplan-Meier
survival analysis based on three independent public microarray data
sets. As shown in Fig. 1, a lower
expression level of TFAP2E was closely associated with an
unfavorable clinical outcome of patients with NB. These results
indicate that TFAP2E might have a role in the suppression of
malignant progression of NB such as the acquisition of anticancer
drug resistance.
siRNA-mediated knockdown of TFAP2E
attenuates ADR-dependent cell death of NB-derived NB1 cells
To determine whether TFAP2E could affect
anticancer drug sensitivity of NB cells, we performed
siRNA-mediated knockdown of TFAP2E in NB-derived NB1 cells.
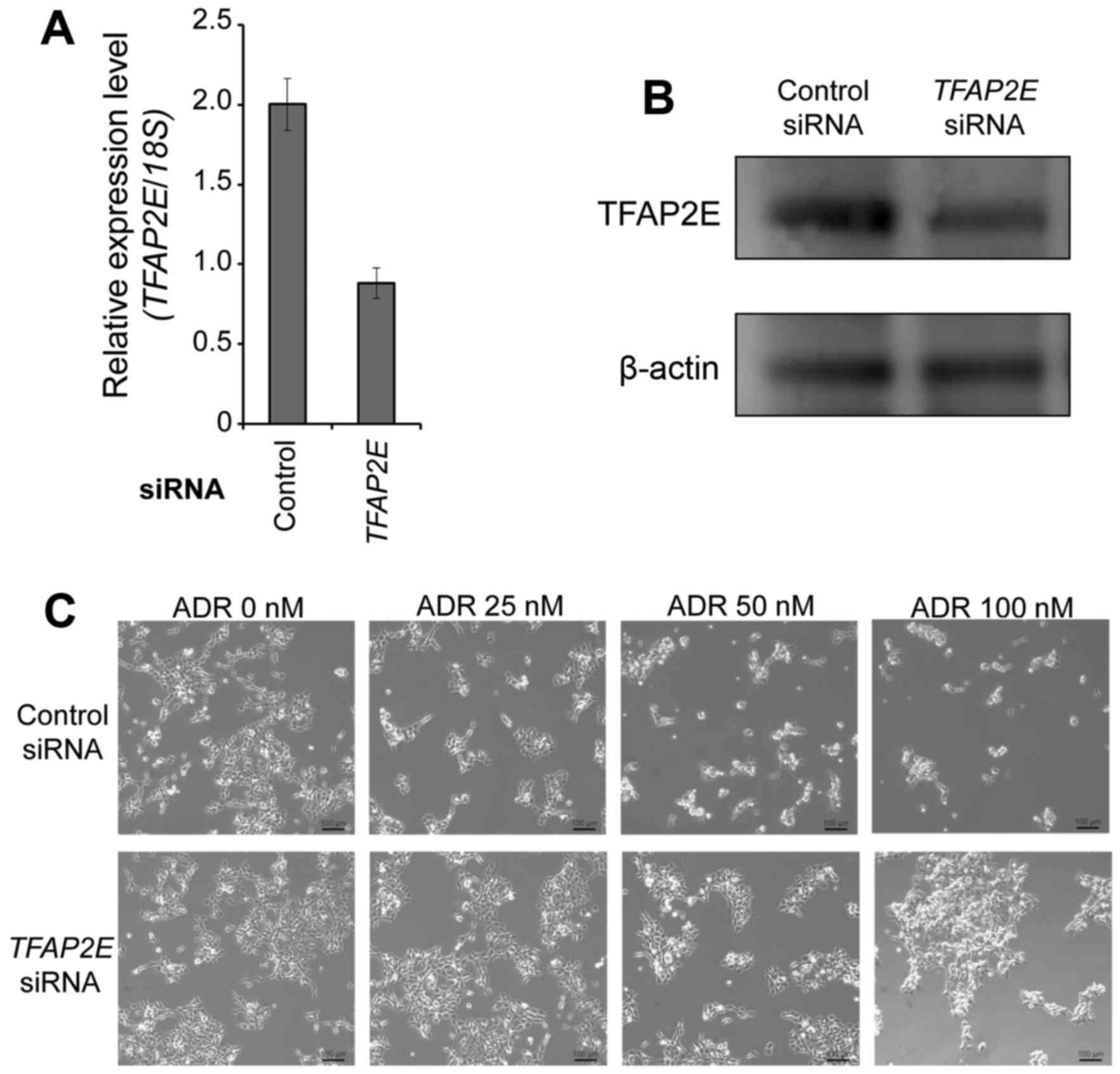
Forty-eight hours after transfection, total RNA and cell lysates
were prepared and analyzed to determine the expression level of
TFAP2E by real-time PCR and immunoblotting, respectively. As
expected, TFAP2E expression was significantly reduced in
TFAP2E-depleted cells at both mRNA and protein levels
(Fig. 2A and B). Under the same
experimental conditions, non-depleted and TFAP2E-depleted
cells were treated with the indicated concentrations of the
anticancer drug ADR. Twenty-four hours after treatment,
representative pictures were taken. As shown in Fig. 2C, TFAP2E-knocked down cells
became much more resistant to ADR compared with non-depleted cells
exposed to ADR. Consistent with these observations, silencing of
TFAP2E significantly increased number of viable cells as
compared to the control transfection (Fig. 2D). TFAP2E knockdown cells
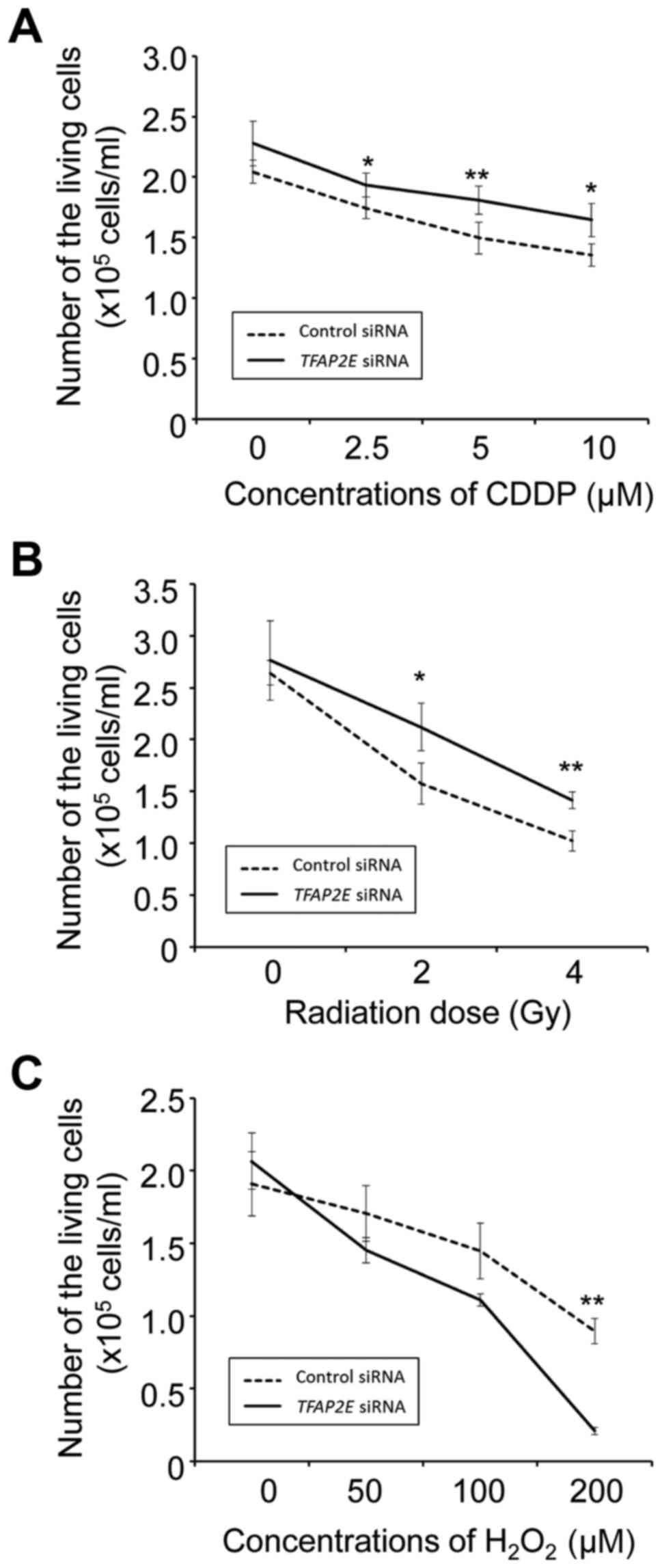
also became resistant to CDDP and IR but not to
H2O2 (Fig.
3).
FACS analysis demonstrated that the relative number
of cells with sub-G1 DNA content markedly declined by TFAP2E
depletion in the presence of ADR (Fig.
2E). On the other hand, the cell population in G2/M phase was
increased by TFAP2E depletion depending on the dose of ADR.
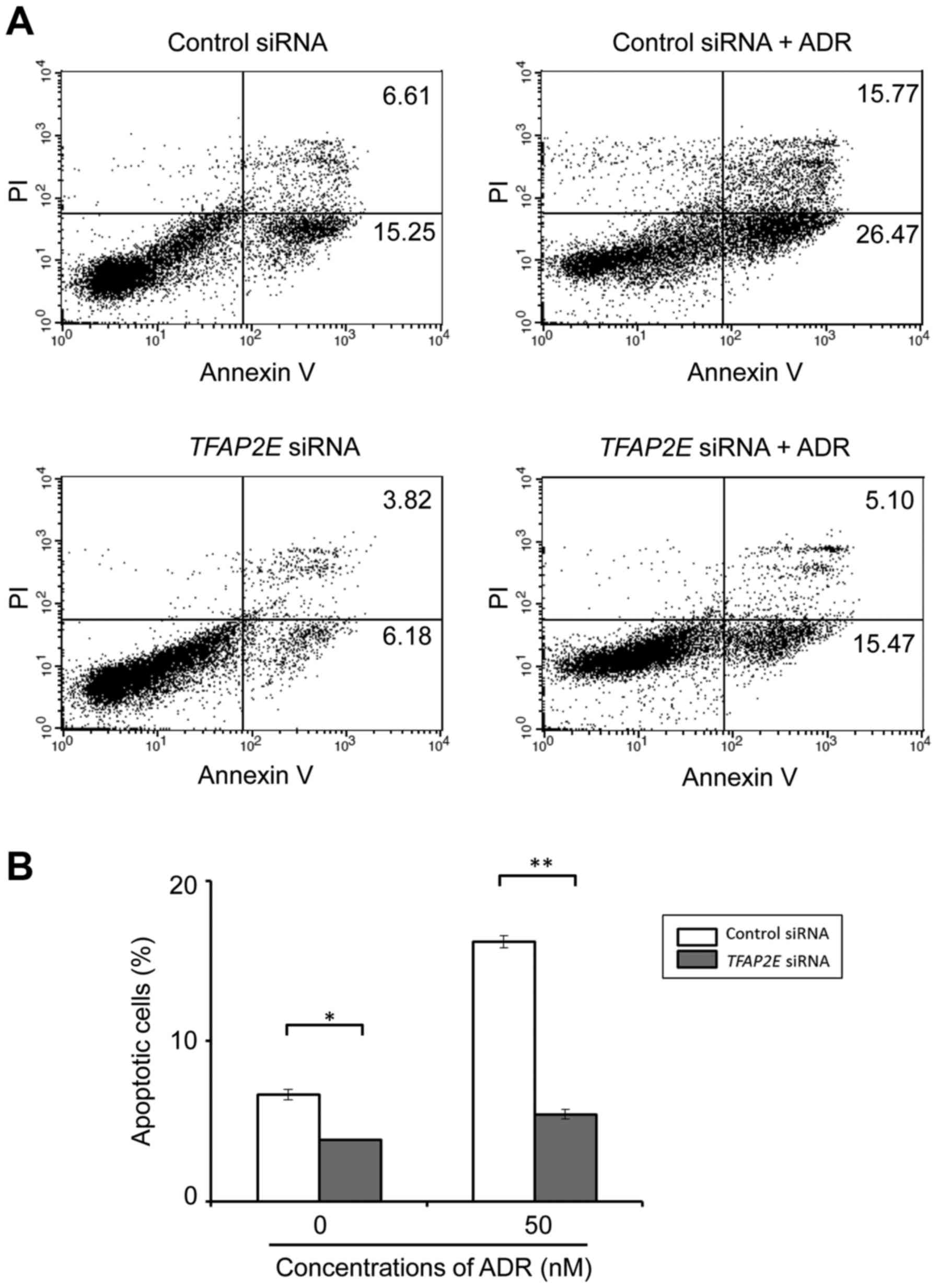
Moreover, FACS analysis after Annexin V/PI staining revealed that
number of apoptotic cells in response to 50 nM of ADR was markedly
suppressed in TFAP2E-knocked down cells compared with the
control cells (Fig. 4). Late
apoptotic cell population (Annexin V, PI-double positive cells) of
control and knocked down cells in the absence of ADR were 6.65±0.31
and 3.82±0.03%, respectively. Upon ADR treatment, number of control
cells with late apoptotic property was markedly increased
(16.19±0.38%), whereas TFAP2E depletion had a negligible
effect on ADR-mediated apoptosis (5.42±0.302%) (Fig. 4). Similar results were also obtained
in NB-derived NB9 cells (data not shown). Together, these findings
indicate that TFAP2E plays a vital role in the regulation of
DNA damage response of NB cells.
ADR-dependent induction of
p21WAF1 is further augmented in TFAP2E-knocked down NB1
cells
To gain insight into understanding the molecular
mechanisms behind TFAP2E depletion-mediated ADR resistance,
we sought to examine the tumor suppressor p53-dependent cell death
pathway under our experimental conditions. According to the IARC
TP53 database (http://p53.iarc.fr/CellLines.aspx), NB1 cells carry
wild-type p53. NB1 cells were transfected with control siRNA
or with siRNA against TFAP2E and then incubated in the
presence of ADR. At the indicated time-points after ADR exposure,
the cell lysates were prepared and subjected to immunoblotting. As
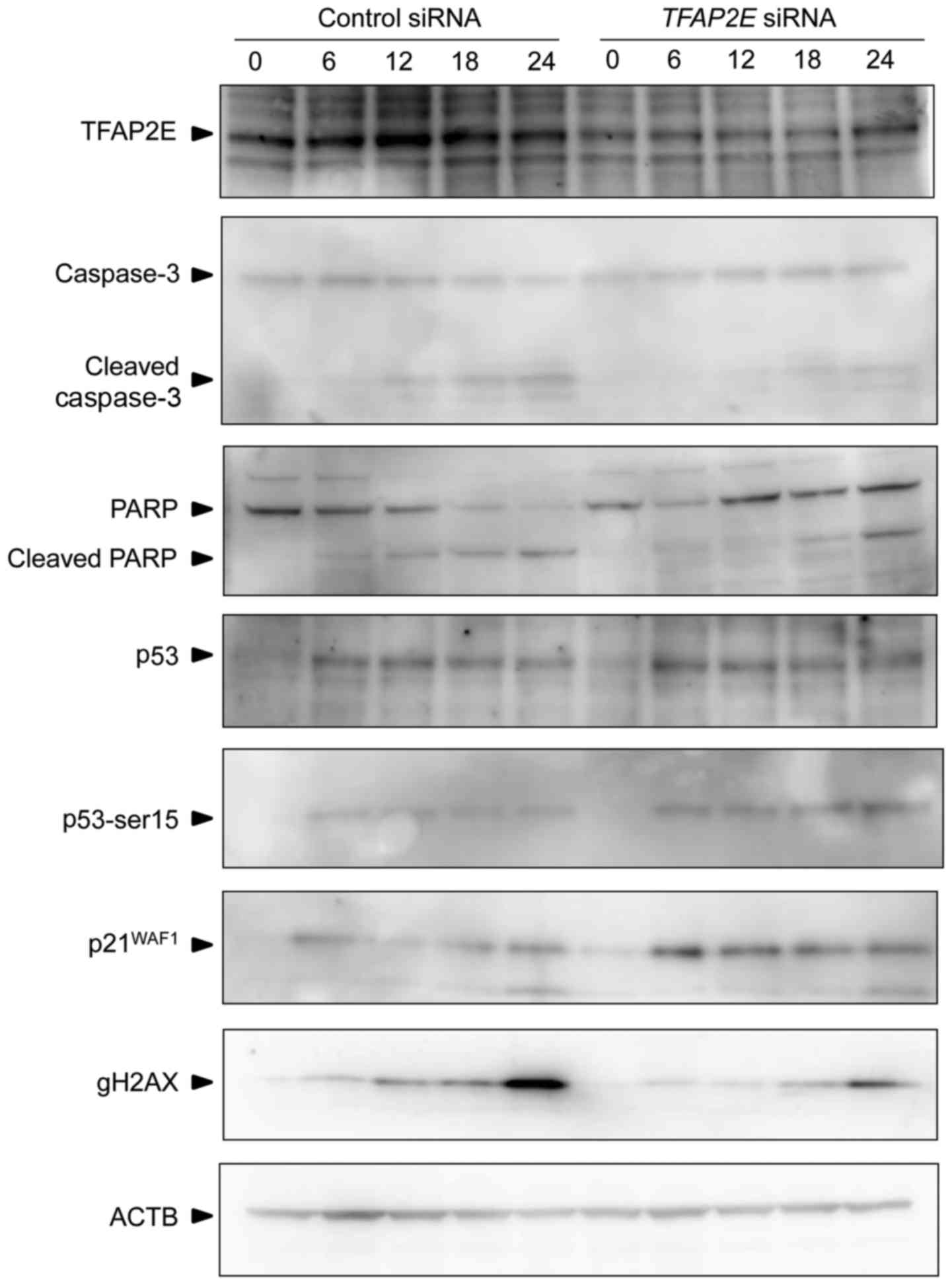
shown in Fig. 5, ADR-induced
accumulation and phosphorylation of p53 at Ser-15 were basically
unchanged regardless of TFAP2E depletion. In accordance with
the results shown in Fig. 3,
ADR-mediated proteolytic cleavage of caspase-3 and its substrate,
PARP, was substantially downregulated in TFAP2E-knocked down
cells compared with that in the non-depleted cells. Of note, an
obvious reduction of γ-H2AX, which has been considered to be a
reliable DNA damage marker, was detectable in
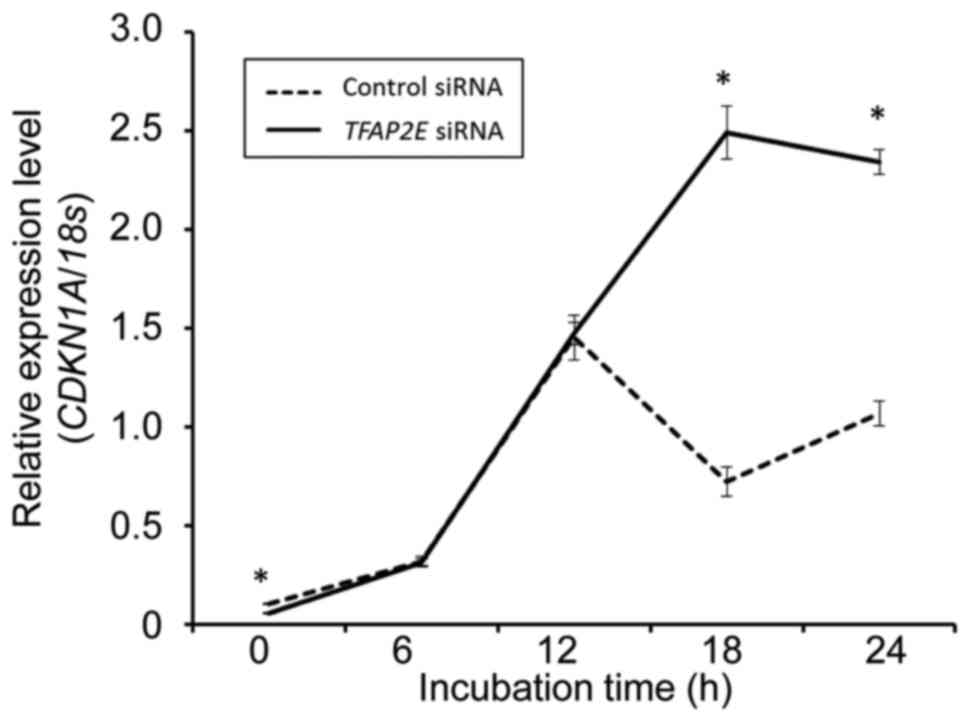
TFAP2E-depleted cells exposed to ADR. Furthermore, silencing
of TFAP2E stimulated ADR-dependent induction of cell
cycle-related p21WAF1. Real-time PCR analysis revealed
that the expression of p21WAF1 is regulated at mRNA
level (Fig. 6).
Discussion
In the present study, we demonstrated that depletion
of TFAP2E attenuates ADR-dependent apoptosis but promotes
mitotic arrest in NB cells. In addition to ADR, TFAP2E gene
silencing prohibited apoptosis induced by the other DNA damaging
agents such as CDDP and IR, suggesting that TFAP2E might act as a
tumor suppressor of NB.
TFAP2E belongs to the AP-2 transcription factor
family, which consists of five members including TFAP2A, TFAP2B,
TFAP2C, TFAP2D and TFAP2E. All of AP-2 family proteins share a
highly conserved structure such as a helix-span-helix motif at the
carboxyl terminus and act as transcription factors. According to
the previous studies (14), AP-2
proteins affect the transcription of numerous number of genes
involved in the crucial biological processes including cell
proliferation and differentiation. The possible functional roles of
AP-2 family members in carcinogenesis vary among individual
proteins. For example, reduced expression levels of TFAP2A are
closely associated with unfavorable phenotypes of many cancers such
as gastric adenocarcinoma, prostate cancer and melanoma (15–17).
Overexpression of TFAP2B has been shown to contribute to poor
prognosis of lung adenocarcinoma (18). In contrast, low expression level of
TFAP2B was related to unfavorable prognostic markers in
neuroblastoma (19). Additionally,
elevated expression of TFAP2C has been found in testicular
carcinoma, advanced-stage of ovarian carcinoma and advanced grade
of breast cancer (20–22). Collectively, it is likely that
TFAP2A is a potent tumor suppressor, whereas TFAP2C has a
tumor-promoting function.
Recently, it has been reported that hypermethylation
of TFAP2E genome locus and lower expression of its
transcript are associated with unfavorable outcome and
non-responsiveness to chemotherapy in colorectal cancer and gastric
cancer (11,12). Analysis of the public database
revealed that a lower expression of TFAP2E is also related
to a shorter survival of neuroblastoma patients. Although these
findings indicate that TFAP2E is a potent tumor suppressor,
it remains elusive how TFAP2E could regulate the expression of
cancer-related genes. It has been described that TFAP2E exerts its
tumor suppressive function through the downregulation of Dickkopf
WNT signaling pathway inhibitor 4 (DKK4) in CRC (11). However, we were unable to detect
DKK4 expression in NB1 cells under our experimental
conditions (data not shown).
According to our results, ADR-dependent stimulation
of cell cycle-related p21WAF1 was further augmented by
TFAP2E depletion. Although it is well known that
p21WAF1 inhibits CDK activity of cyclin A-CDK2 and
cyclin E-CDK2 complexes and thereby functions as a tumor
suppressor, p21WAF1 also has an anti-apoptotic potential
(23). Thus, it is possible that
TFAP2E depletion-mediated upregulation of p21WAF1
prohibits ADR-dependent apoptosis and induces mitotic arrest of NB
cells. Since p21WAF1 is one of p53-target gene products,
it is suggestive that TFAP2E might participate in p53-dependent DNA
damage response of NB cells. However, it was not the case. Firstly,
TFAP2E gene silencing had an undetectable effects on
ADR-mediated induction of p53 and accumulation of phosphorylated
p53 at Ser-15. Secondary, the expression level of p53-target
14-3-3σ implicated in mitotic arrest (24) was unaffected by TFAP2E
depletion (data not shown). Thirdly, the complex formation between
TFAP2E and p53 was not detectable in the presence or absence
of ADR as examined by co-immunoprecipitation experiments (data not
shown). Therefore, it is indicative that TFAP2E
depletion-dependent mitotic arrest is regulated in a
p53-independent manner.
In conclusion, our present findings suggest that
TFAP2E acts as a tumor suppressor and potentiates proper DNA damage
response in NB.
Acknowledgements
We thank Ms. A. Oguni for her excellent technical
support and Ms. K. Takgata for her secretarial assistance. The
present study was supported in part by the JSPS KAKENHI Grant
Number 24591637 to K.F and 24791893 to K.O.
References
|
1
|
Maris JM, Hogarty MD, Bagatell R and Cohn
SL: Neuroblastoma. Lancet. 369:2106–2120. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Dang CV: MYC on the path to cancer. Cell.
149:22–35. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Caron H: Allelic loss of chromosome 1 and
additional chromosome 17 material are both unfavourable prognostic
markers in neuroblastoma. Med Pediatr Oncol. 24:215–221. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lampert F, Rudolph B, Christiansen H and
Franke F: Identical chromosome 1p breakpoint abnormality in both
the tumor and the constitutional karyotype of a patient with
neuroblastoma. Cancer Genet Cytogenet. 34:235–239. 1988. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Takagi K, Fujiwara K, Takayama T, Mamiya
T, Soma M and Nagase H: DNA hypermethylation of zygote arrest 1
(ZAR1) in hepatitis C virus positive related hepatocellular
carcinoma. Springerplus. 2:1502013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Shinojima Y, Terui T, Hara H, Kimura M,
Igarashi J, Wang X, Kawashima H, Kobayashi Y, Muroi S, Hayakawa S,
et al: Identification and analysis of an early diagnostic marker
for malignant melanoma: ZAR1 intra-genic differential methylation.
J Dermatol Sci. 59:98–106. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Watanabe T, Yachi K, Ohta T, Fukushima T,
Yoshino A, Katayama Y, Shinojima Y, Terui T and Nagase H: Aberrant
hypermethylation of non-promoter zygote arrest 1 (ZAR1) in human
brain tumors. Neurol Med Chir (Tokyo). 50:1062–1069. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sugito K, Kawashima H, Yoshizawa S, Uekusa
S, Hoshi R, Furuya T, Kaneda H, Hosoda T, Konuma N, Masuko T, et
al: Non-promoter DNA hypermethylation of zygote arrest 1 (ZAR1) in
neuroblastomas. J Pediatr Surg. 48:782–788. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Fujiwara K, Ghosh S, Liang P, Morien E,
Soma M and Nagase H: Genome-wide screening of aberrant DNA
methylation which associated with gene expression in mouse skin
cancers. Mol Carcinog. 54:178–188. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Tummala R, Romano RA, Fuchs E and Sinha S:
Molecular cloning and characterization of AP-2 epsilon, a fifth
member of the AP-2 family. Gene. 321:93–102. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ebert MP, Tänzer M, Balluff B,
Burgermeister E, Kretzschmar AK, Hughes DJ, Tetzner R, Lofton-Day
C, Rosenberg R, Reinacher-Schick AC, et al: TFAP2E-DKK4 and
chemoresistance in colorectal cancer. N Engl J Med. 366:44–53.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sun J, Du N, Li J, Zhou J, Tao G, Sun S
and He J: Transcription factor AP2ε: A potential predictor of
chemoresistance in patients with gastric cancer. Technol Cancer Res
Treat. 15:285–295. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Payne SR, Serth J, Schostak M, Kamradt J,
Strauss A, Thelen P, Model F, Day JK, Liebenberg V, Morotti A, et
al: DNA methylation biomarkers of prostate cancer: Confirmation of
candidates and evidence urine is the most sensitive body fluid for
non-invasive detection. Prostate. 69:1257–1269. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Eckert D, Buhl S, Weber S, Jäger R and
Schorle H: The AP-2 family of transcription factors. Genome Biol.
6:2462005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang W, Lv L, Pan K, Zhang Y, Zhao JJ,
Chen JG, Chen YB, Li YQ, Wang QJ, He J, et al: Reduced expression
of transcription factor AP-2α is associated with gastric
adenocarcinoma prognosis. PLoS One. 6:e248972011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lipponen P, Aaltomaa S, Kellokoski J,
Ala-Opas M and Kosma V: Expression of activator protein 2 in
prostate cancer is related to tumor differentiation and cell
proliferation. Eur Urol. 37:573–578. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Karjalainen JM, Kellokoski JK, Eskelinen
MJ, Alhava EM and Kosma VM: Downregulation of transcription factor
AP-2 predicts poor survival in stage I cutaneous malignant
melanoma. J Clin Oncol. 16:3584–3591. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Fu L, Shi K, Wang J, Chen W, Shi D, Tian
Y, Guo W, Yu W, Xiao X, Kang T, et al: TFAP2B overexpression
contributes to tumor growth and a poor prognosis of human lung
adenocarcinoma through modulation of ERK and VEGF/PEDF signaling.
Mol Cancer. 13:892014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ikram F, Ackermann S, Kahlert Y, Volland
R, Roels F, Engesser A, Hertwig F, Kocak H, Hero B, Dreidax D, et
al: Transcription factor activating protein 2 beta (TFAP2B)
mediates noradrenergic neuronal differentiation in neuroblastoma.
Mol Oncol. 10:344–359. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Hoei-Hansen CE, Nielsen JE, Almstrup K,
Sonne SB, Graem N, Skakkebaek NE, Leffers H and Rajpert-De Meyts E:
Transcription factor AP-2gamma is a developmentally regulated
marker of testicular carcinoma in situ and germ cell tumors. Clin
Cancer Res. 10:8521–8530. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ødegaard E, Staff AC, Kaern J, Flørenes
VA, Kopolovic J, Tropé CG, Abeler VM, Reich R and Davidson B: The
AP-2gamma transcription factor is upregulated in advanced-stage
ovarian carcinoma. Gynecol Oncol. 100:462–468. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Sotiriou C, Wirapati P, Loi S, Harris A,
Fox S, Smeds J, Nordgren H, Farmer P, Praz V, Haibe-Kains B, et al:
Gene expression profiling in breast cancer: Understanding the
molecular basis of histologic grade to improve prognosis. J Natl
Cancer Inst. 98:262–272. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Dutto I, Tillhon M, Cazzalini O, Stivala
LA and Prosperi E: Biology of the cell cycle inhibitor
p21CDKN1A: Molecular mechanisms and relevance in
chemical toxicology. Arch Toxicol. 89:155–178. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Hermeking H, Lengauer C, Polyak K, He TC,
Zhang L, Thiagalingam S, Kinzler KW and Vogelstein B: 14-3-3sigma
is a p53-regulated inhibitor of G2/M progression. Mol Cell. 1:3–11.
1997. View Article : Google Scholar : PubMed/NCBI
|