Introduction
Gastric cancer is the most common gastrointestinal
malignant tumor in the world and a large number of factors that
have significant long-term impact have been linked with this
disease (1). The morbidity and
mortality for gastric cancer appears to be 14.1/100,000 and
10.3/100,000 cases, respectively (2). Numerous studies have shown that
Helicobacter pylori infection is a major risk factor for
gastric cancer since persistent Helicobacter pylori
infections may consequently cause damage of the gastric mucosa and
thereby trigger the proliferation of gastric epithelial cells.
Moreover, Helicobacter pylori infection is also associated
with the decrease in gastric acid secretion which may induce
chronic gastritis, gastric precancerous lesions and gastric
carcinogenesis (3). Other factors
including family history, poor diet, smoking, drinking and mental
stress are also involved in the development of gastric cancer
(1,4). Medications are still one of the most
effective treatment options for patients with gastric cancer and
they are usually recommended for patients in advanced stages
(5). However, chemotherapeutic
agents are not capable of effectively controlling the disease in a
large proportion of patients as they present with resistance to
these agents. One of the major limitations of chemotherapy may
arise from the fact that some gastric cancer patients exhibit
resistance to multiple medications and hence chemotherapy may not
achieve its effectiveness (6).
Therefore, a large amount of research has been conducted to
overcome this issue.
MicroRNAs (miRNAs) are a conserved family of small
non-coding RNA molecules that function as pivotal regulators of
gene expression (7). They modulate
gene expression by binding to the 3′-untranslated region (3′UTR) of
their target mRNAs and thereby suppression of protein translation
or mRNA decay can be triggered (8,9).
Furthermore, miRNAs not only participate in tumor proliferation,
invasion and metastasis but also regulate a variety of biological
processes (10–12). For instance, the expression of
miR-200b/c in gastric cancer SGC790l/VCR cells was lower compared
to that in SGC790l cells whereas the expression of anti-apoptotic
protein Bcl-2 was increased in SGC790l/VCR cells and there was a
marked difference in its sensitivity to cisplatin (DDP) and
vincristine (13). MicroRNA-200c
(miR-200c) is part of the miR-200 family in which another four
members are also included: miR-200a, miR-200b, miR-141 and miR-429
(14). In addition, the expression
of miR-200c is significantly downregulated in tumor cells and
tissues and it may play a role in tumor suppression (15,16).
For instance, Chen et al demonstrated that the expression of
miR-200c was significantly lower in SGC790l/DDP cells whereas the
sensitivity of SGC790l/DDP cells to four chemotherapy medications
was significantly enhanced after the upregulation of miR-200c
(17).
Zinc finger E-box binding homeobox 2 (ZEB2) is an
important member of the Snail superfamily and it is commonly
expressed in human and rat tissues. ZEB2 can bind the E-box
sequence in the E-cadherin gene promoter region which inhibits
E-cadherin, cytokeratin, mucin and mac1 protein transcription. More
importantly, the downregulation of the aforementioned proteins
plays an important role in epithelial-to-mesenchymal transition
(EMT) (18). Apart from that, ZEB2
is closely linked with bladder cancer resistance to epidermal
growth factor inhibitor and ovarian cancer resistance to DDP
(19,20).
No current evidence has been revealed with respect
to the hypothesis that miR-200c regulates DDP resistance by
targeting ZEB2 in human gastric cancer cells. As a result of this,
we are motivated to systematically clarify the potential function
of the miR-200c and ZEB2 signaling pathway in gastric cancer and
DDP resistance may be predicted based on the aforementioned
functions.
Materials and methods
Tissue samples
Fifty gastric cancer tissues and paired normal
adjacent tissues were acquired from patients who underwent surgical
treatment in our hospital from 2013 to 2015. There were 28 males
and 22 females, with an average age of 55.6 years (ranging from 30
to 81 years). Clinical TNM stages were determined according to the
newly revised standards of TNM staging for gastric cancer from the
2010 American Joint Committee on Cancer (AJCC) (21). No patients received radiotherapy,
chemotherapy or hormone therapy before surgery. Tissue samples were
immediately frozen in liquid nitrogen at the time of surgery, then
stored at −80°C until the extraction of RNA. Another group of tumor
tissues was fixed with 10% formalin, embedded in paraffin and
immunohistochemistry was performed. Informed consent for this study
was collected from every patient, and this study was approved by
the Research Ethics Committee of Henan Key Laboratory for
Esophageal Cancer Laboratory for Cancer Research.
Cell culture
SGC7901, a human gastric cancer cell line, was
obtained from the Cell Bank of the Chinese Academy of Sciences
(Shanghai, China). The SGC7901/DDP cells with DDP resistance were
purchased from KeyGen Biotech Co., Ltd. (Nanjing, China). Cells
were maintained in RPMI-1640 medium containing 10% fetal bovine
serum (FBS), streptomycin (100 µg/ml) and penicillin (100 U/ml)
(all from Gibco, Grand Island, NY, USA) in a 37°C incubator with 5%
CO2. In order to maintain the DDP resistant phenotype,
DDP (at a final concentration of 1 µg/ml) was added to the culture
medium for the SGC7901/DDP cells. SGC7901/DDP cells were cultured
for one week in medium without DDP before experimentation.
Cell transfection
We transfected miR-200c mimics and ZEB2 siRNA
(purchased from GenePharma, Shanghai, China) into gastric cancer
cells in order to assess the effect of miR-200c and ZEB2 on both
cell apoptosis and chemosensitivity. The scramble group transfected
with scramble miRNA mimics was synthesized by GenePharma and was
used as the negative control. Cells were plated in 6-well plates at
1×105 cells in each well in an antibiotic-free RPMI-1640
medium with 10% FBS. After 12 h, Opti-MEM medium (Gibco) without
antibiotics and serum was used to replace the aforementioned
medium. When cells grew to about 50% confluence, they were
transfected with Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA)
and cultured in a 37°C incubator with 5% CO2. Cells
continued to culture after the complete medium was replaced after
6–8 h. Cells were harvested after a 48-h transfection and used for
western blot analysis and RT-PCR.
Quantitative real time reverse
transcription-polymerase chain reaction (qRT-PCR) analysis
Total RNA extraction from human gastric cancer
tissues and cells was conducted using TRIzol reagent (Invitrogen)
based on the manufacturers instructions. Complementary DNA (cDNA)
was acquired using the Omniscript reverse transcription kit
(Qiagen, Hilden, Germany). After the reaction of reverse
transcription, a real-time quantitative RT-PCR assay was conducted
using an ABI7500 quantitative PCR instrument (Applied Biosystems)
and the relative expression levels of miR-200c and ZEB2 mRNA were
detected. The primers for miR-200c and ZEB2 (Invitrogen) are shown
in Table I. The relative expression
of miR-200c and ZEB2 mRNA was calculated using the
2−ΔΔCt method and normalized to the expression of U6
snRNA and glyceraldehyde-3-phosphate dehydrogenase (GAPDH). All the
aforementioned assays were replicated three times.
 | Table I.Sequence of the primers used for
quantitative RT-PCR. |
Table I.
Sequence of the primers used for
quantitative RT-PCR.
| Genes | Primer pair
sequences |
|---|
| miR-200c | F:
5′-AGCGGTAATACTGCCGGGTA-3′ | R:
5′-GTGCAGGGTCCGAGGT-3′ |
| U6 | F:
5′-GGGTGCTCGCTTCGGCAGC-3′ | R:
5′-CAGTGCAGGGTCCGAGGT-3′ |
| ZEB2 | F:
5′-TGTCATTAGAAGAGGCGTAA-3′ | R:
5′-GCAGAGCAGGTTAGAACT-3′ |
| GAPDH | F:
5′-CATCAGCAATGCCTCCTGCAC-3′ | R:
5′-TGAGTCCTTCCACGATACCAAAGTT-3′ |
Immunohistochemistry
ZEB2 protein expression in gastric cancer specimens
was detected using the universal PV-9000 two-step
immunohistochemical method. In brief, formalin-fixed and
paraffin-embedded tissues from gastric cancer patients were cut
into 4-µm slices. Then, conventional dewaxing, graded ethanol
dehydration, antigen retrieval and the addition of 3% hydrogen
peroxide were performed on tissue slices in order to block
endogenous peroxidase. Primary antibodies (rabbit anti-human ZEB2
polyclonal antibody; BIOSS, Beijing, China) were applied to tissue
slices at 4°C overnight. Then, tissues were incubated for 20 min at
room temperature after the addition of polymerase adjuvants.
Secondary antibodies labeled with horseradish peroxidase (HRP;
BIOSS) were also applied and tissues were incubated for another 30
min at room temperature. Staining was performed using
diaminobenzidine (DAB) and slices were counterstained using
hemalum. Phosphate-buffered solution (PBS) instead of a primary
antibody was considered as the negative control and a known
positive antibody was set as the positive control. The
immunohistochemical score of ZEB2 was calculated by multiplying the
intensity of the staining (0, colorless; 1, light yellow; 2,
yellow; and 3, brown) and the positive cell percentage (0, ≤5%; 1,
6–25%; 2, 26–50%; 3, 51–75%; and 4, >75%). Cells were randomly
selected from 5 high-power fields (x200) in each slice and 100
cells were counted in each field. As suggested by the final scores,
the integral levels of ZEB2 were evaluated as: negative (−), ≤4
points and positive (+), >4 points. Two independent pathologists
were responsible for analyzing these slices.
Dual-luciferase reporter assay
miRNA targeted genes were predicted using TargetScan
(https://www.targetscan.org). Then, the
wild-type and mutant-type of ZEB2 3′UTR luciferase reporter vectors
were constructed. miR-200c mimics were co-transfected with
constructed wild-type or mutant-type luciferase reporter vectors
into SGC7901/DDP cells using Lipofectamine 2000 (Invitrogen). The
pGL3-control vector (Promega, Madison, WI, USA) was transfected as
a control. The dual-luciferase reporter assay system (Promega) was
used to examine the luciferase activity after the cells had been
transfected for 48 h.
Cell viability assay
Cell viability was evaluated using the methyl
thiazolyl tetrazolium (MTT) assay. Briefly, transfected cells were
seeded in a 96-well plate at 5×103 cells in each well
for 48 h. After 12 h, the cells were treated with various
concentrations of DDP (Qilu Pharmaceutical Co., Ltd., Shandong,
China) with final concentrations at 0.01, 0.1, 1 and 10 times those
of the human peak serum doses for DDP, as previously suggested
(13). The peak plasma
concentration of anticancer drugs is 2.0 µg/ml for DDP (22). Approximately 48 h after the addition
of DDP, MTT (20 µl, 5 mg/ml; Sigma-Aldrich, St. Louis, MO, USA) was
added into each well, and the culture was sustained for 4 h in a
37°C incubator with 5% CO2. Subsequently, dimethyl
sulphoxide (DMSO; 150 µl) was added into each well and the cells
were shaken lightly for 10 min to dissolve the crystals. Samples
were read on a microplate reader (SpectraMax Plus; Molecular
Devices, Sunnyvale, CA, USA) at 490 nm. The 50% inhibition
concentration (IC50) of DDP was estimated.
Apoptosis assay
Annexin V-fluorescein isothiocyanate (Annexin
V-FITC) and propidium iodide (PI) apoptosis detection kit
(Becton-Dickinson, Franklin Lakes, NJ, USA) was used to evaluate
the apoptosis of SGC7901/DDP cells. In brief, SGC7901/DDP cells
were treated with DDP at a final concentration of 5 µg/ml after
transfection for 24 h. Cells following a 48-h treatment were washed
two times using cold PBS. Then the cells were re-suspended in
binding buffer and maintained at a concentration of
0.5–1×106/ml. The suspension (100 µl) was incubated with
5 µl of Annexin V-FITC and PI for 15 min at room temperature in the
dark. After the addition of 400 µl binding buffer into each tube,
the cells were assessed using flow cytometry (Beckman FC 500
MCL/MPL; Beckman Coulter, Brea, CA, USA).
Western blot analysis assay
After transfection was sustained for 48 h, cells
were collected and homogenized using RIPA buffer (Beyotime, China).
Cellular proteins were extracted and the protein concentrations
were assessed using a bicinchoninic acid (BCA) protein assay kit
(Boster Biotechnology Co., Ltd., Wuhan, China). Equal amounts of
proteins for each group were loaded and isolated with sodium
dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE),
after which they were transferred onto polyvinylidene fluoride
membranes and blocked with 5% nonfat milk. Membranes were incubated
with the ZEB2, Bcl-2, Bax, caspase-3 primary antibodies or the
GAPDH antibody [Cell Signaling Technology (CST), Beverly, MA, USA]
respectively, at 4°C overnight. Membranes were washed three times
using Tris hydroxymethyl aminoethane (TBST; 10 min each) and then
HRP-linked secondary antibodies were added followed by incubation
at room temperature for 1 h. Membranes were washed again with TBST
three times (10 min each) and signal detection was performed using
a Super ECL Plus Detection reagent (Applygen Technologies, Inc.,
Beijing, China).
Statistical analysis
Statistical analyses were implemented using the SPSS
19.0 software. Differences in continuous variables among groups
(mean ± SD) were compared using the procedure of analysis of
variance (ANOVA) or the Student's t-test. Immunohistochemical
results of the ZEB2 protein were analyzed by the Chi-squared test.
A P-value <0.05 was defined as statistically significant.
Results
Expression levels of miR-200c and ZEB2
in gastric cancer specimens
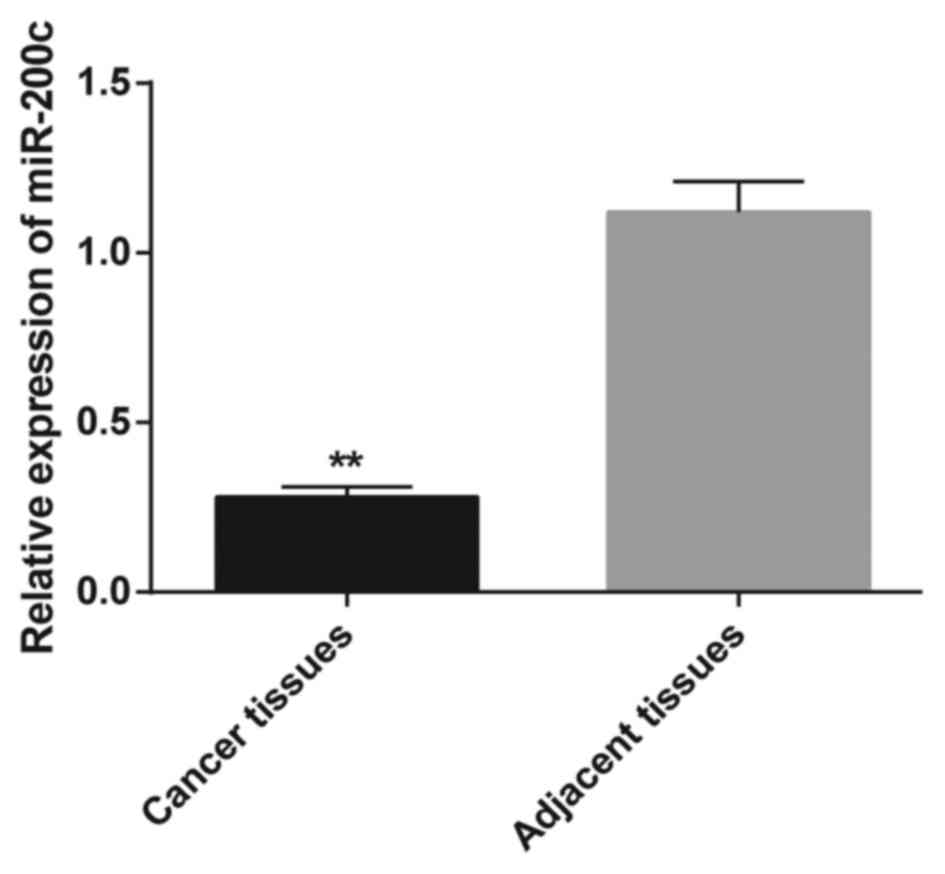
miR-200c expression in 50 gastric cancer tissues and
matched normal adjacent tissues was tested by quantitative
real-time RT-PCR. Gastric cancer tissues showed a significantly
downregulated miR-200c expression level when compared to that noted
in the normal adjacent tissues (P<0.01; Fig. 1). Furthermore, we analyzed the
correlation between the expression of miR-200c and the pathological
characteristics of gastric cancer patients. As shown in Table II, patients with lymph node
metastasis were associated with a marked lower expression level of
miR-200c on average. In addition, patients at TNM III/IV stage had
significantly lower miR-200c expression compared with patients at
TNM I/II stage (all P<0.01). No significant correlation was
found between miR-200c expression and age or gender (both
P>0.05).
 | Table II.Expression of miR-200c in gastric
cancer specimens and the correlation with the clinicopathological
features. |
Table II.
Expression of miR-200c in gastric
cancer specimens and the correlation with the clinicopathological
features.
| Clinicopathological
factors | N | miR-200c | P-value |
|---|
| Age (years) |
|
≥55 | 26 | 0.29±0.03 | 0.245 |
|
<55 | 24 | 0.28±0.03 |
|
| Gender |
|
Male | 28 | 0.28±0.02 | 0.254 |
|
Female | 22 | 0.29±0.04 |
|
| Lymph node
metastasis |
|
Yes | 34 | 0.27±0.02 |
<0.001 |
| No | 16 | 0.31±0.02 |
|
| TNM stage |
|
I/II | 19 | 0.30±0.03 |
<0.001 |
|
III/IV | 31 | 0.27±0.02 |
|
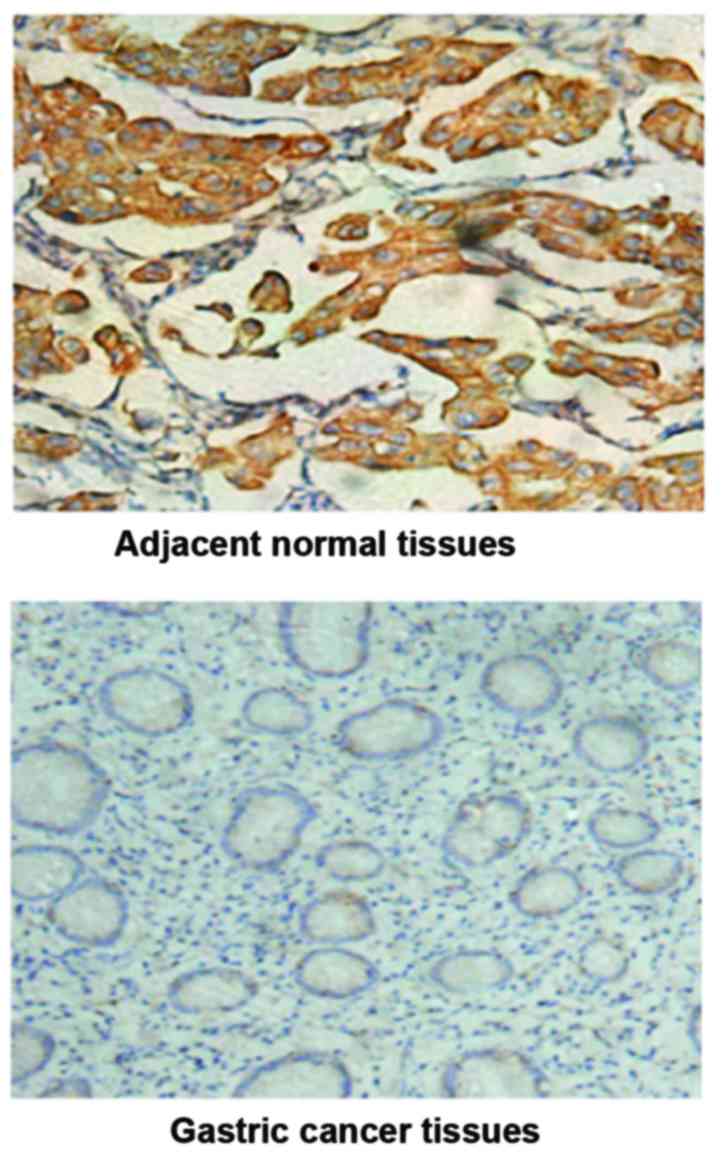
The protein expression of ZEB2 in gastric cancer
specimens was detected by immunohistochemical method (Fig. 2). The association between the
expression of ZEB2 and the clinical characteristics of the gastric
cancer patients is shown in Table
III. The expression of ZEB2 in patients with lymph node
metastasis was significantly higher than that of patients without
lymph node metastasis; the expression of ZEB2 in patients at a TNM
III/IV stage was also significantly higher than that of patients at
a TNM I/II stage (both P<0.05). No significant association
between the expression of ZEB2 and age/gender was suggested (both
P>0.05).
 | Table III.Correlation between the expression of
ZEB2 protein and the clinicopathological features of gastric
cancer. |
Table III.
Correlation between the expression of
ZEB2 protein and the clinicopathological features of gastric
cancer.
|
|
| ZEB2 |
|
|---|
|
|
|
|
|
|---|
| Clinicopathological
factors | N | Positive | Negative | P-value |
|---|
| Age (years) |
|
≥55 | 26 | 17 | 9 | 0.825 |
|
<55 | 24 | 14 | 10 |
|
| Gender |
|
Male | 28 | 18 | 10 | 0.803 |
|
Female | 22 | 14 | 8 |
|
| Lymph node
metastasis |
|
Yes | 34 | 23 | 11 | 0.035 |
| No | 16 | 5 | 11 |
|
| TNM stage |
|
I/II | 19 | 7 | 12 | 0.020 |
|
III/IV | 31 | 23 | 8 |
|
Expression of miR-200c and ZEB2 in
gastric cancer SGC7901 and SGC7901/DDP cells
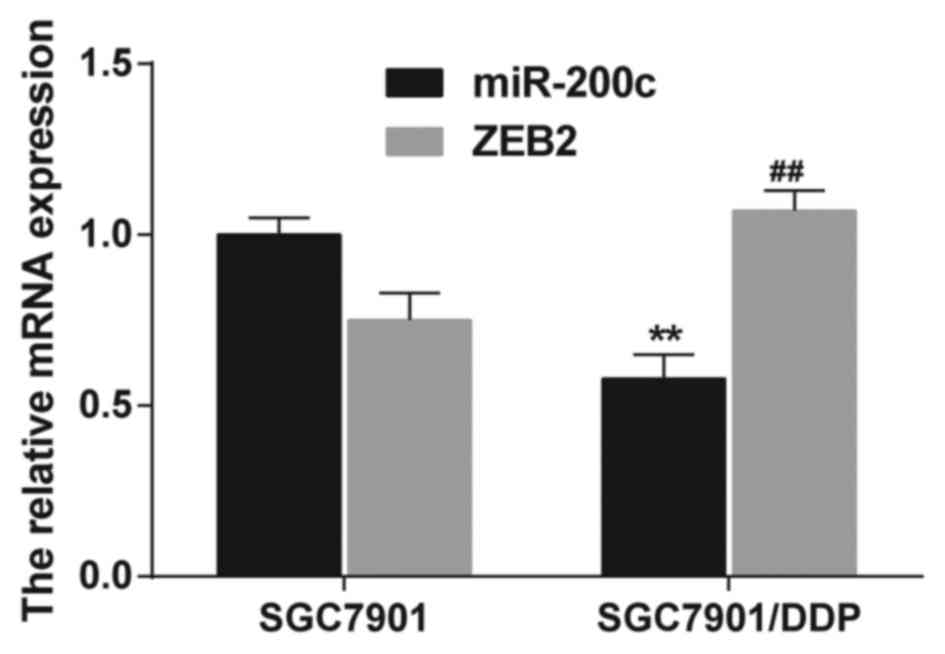
Results from RT-PCR revealed that the expression of
miR-200c was markedly downregulated in SGC7901/DDP cells with DDP
resistance compared with SGC7901 cells (P<0.01). Meanwhile, the
expression of ZEB2 in SGC7901/DDP cells was increased in comparison
to SGC7901 cells (P<0.01; Fig.
3). Therefore, we concluded that there is a potential
relationship between miR-200c and ZEB2.
ZEB2 is a target gene of miR-200c
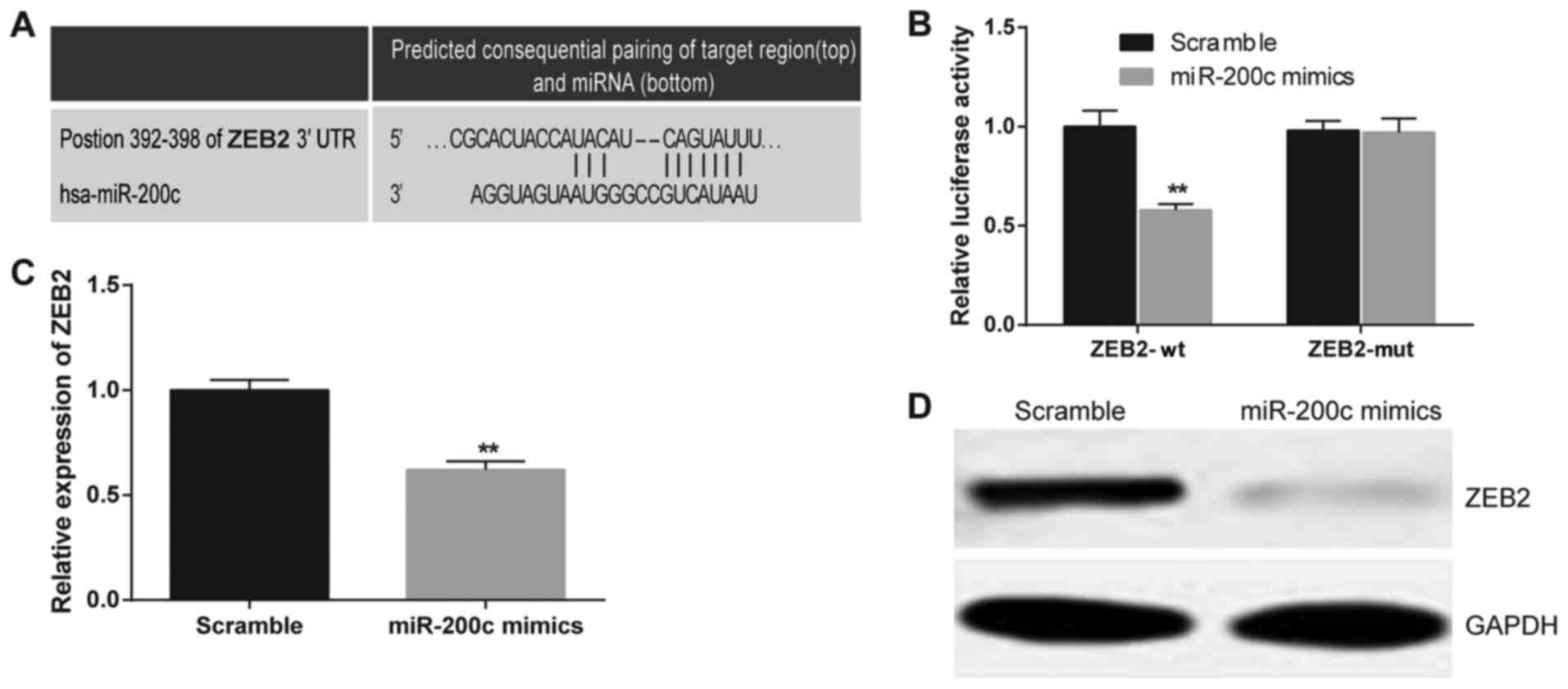
For the purpose of clarifying the potential
relationship between miR-200c and ZEB2, a putative conserved
binding site of miR-200c at nucleotide position 392–398 of human
ZEB2 3′UTR was predicted by the TargetScan database. Perfect base
pairing is shown between the seed sequence of mature miR-200c and
the 3′UTR of ZEB2 mRNA (Fig. 4A).
The results of the dual luciferase reporter gene assays revealed
that miR-200c decreased the luciferase activity of ZEB2 wild-type
by 42% (P<0.01). However, the effect of miR-200c on the
luciferase activity of ZEB2 with mutant-type 3′UTR was not
significant (Fig. 4B). As suggested
by RT-PCR and western blot analysis assays, the expression levels
of both ZEB2 mRNA and protein were inhibited by the miR-200c mimics
compared to the scramble group (P<0.01; Fig. 4C and D). The aforementioned findings
indicate that ZEB2 is a direct target of miR-200c.
Upregulation of miR-200c and knockdown
of ZEB2 enhance the sensitivity of SGC7901/DDP cells to DDP
To explore the association between miR-200c and DDP
resistance in SGC7901/DDP cells, the impact of miR-200c
overexpression and downregulation of ZEB2 on the DDP sensitivity of
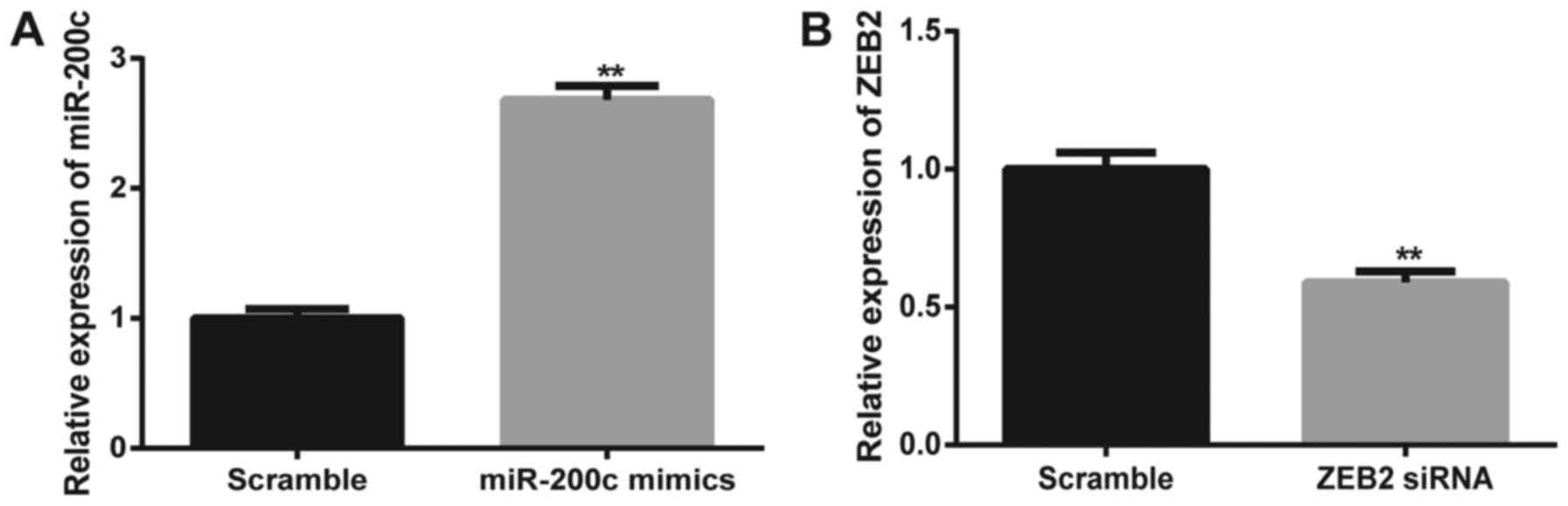
cells was assessed. The data from qRT-PCR showed that miR-200c
mimics significantly increased the expression level of miR-200c,
suggesting that miR-200c was efficiently transfected into the
SGC7901/DDP cells (Fig. 5A).
Fig. 5B shows that ZEB2 was
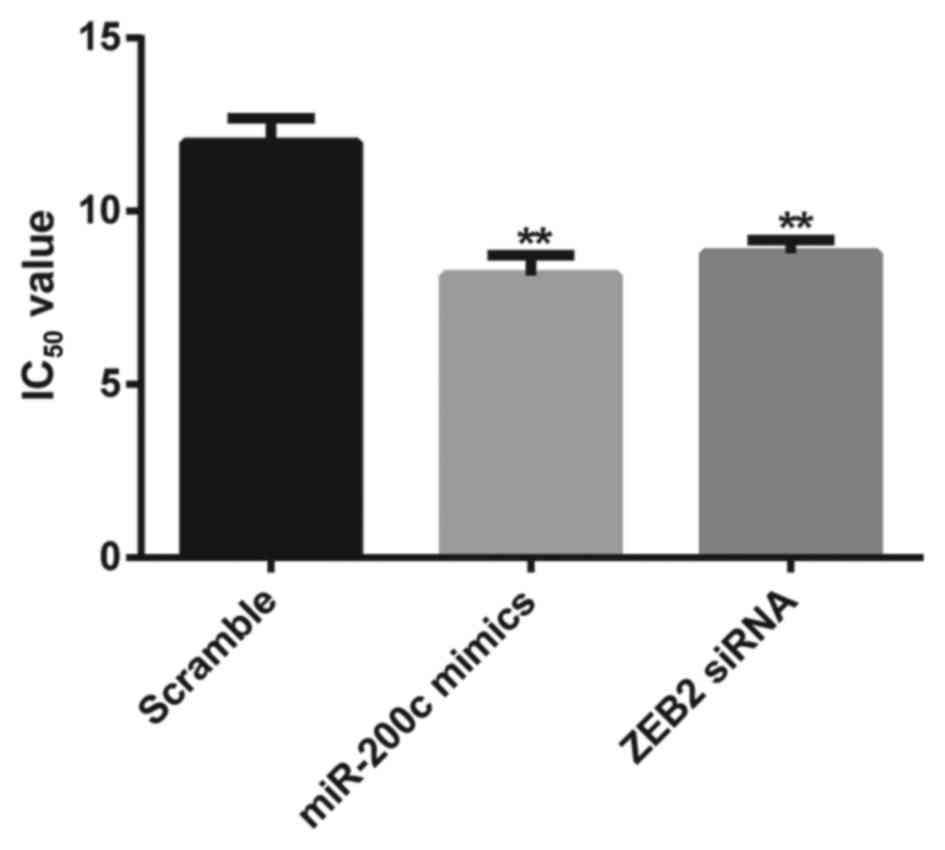
successfully knocked down by the transfection of ZEB2 siRNA. MTT
assay revealed that the SGC7901/DDP cells transfected with miR-200c
mimics exhibited a significantly lower survival status than the
scramble group (IC50, 8.14±0.59 vs. 11.97±0.71 µg/ml,
P<0.01). The sensitivity of DDP was significantly enhanced in
cells transfected with ZEB2 siRNA compared to the scramble group,
with the IC50 of DDP at 8.78±0.39 µg/ml in the ZEB2
siRNA group (P<0.01; Fig.
6).
Overexpression of miR-200c and
downregulation of ZEB2 sensitize cells to DDP-induced
apoptosis
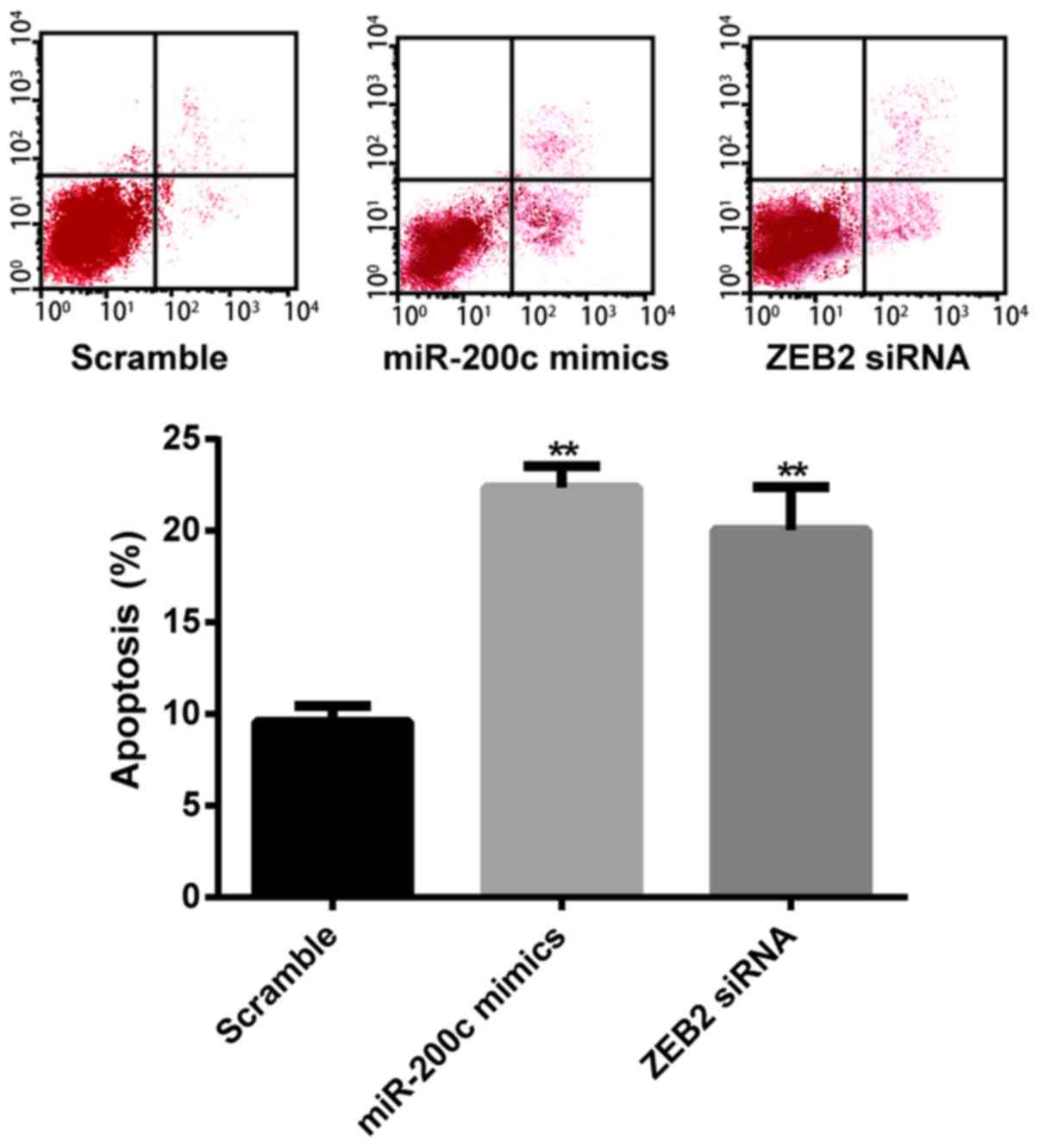
The results of flow cytometry showed that the
SGC7901/DDP cells transfected with miR-200c mimics had a
significantly higher apoptosis rate (22.35±1.16, P<0.01)
compared with the scramble group (9.58±0.86). The apoptosis rate of
cells transfected with ZEB2 siRNA was also significantly increased
(20.03±2.36, P<0.01), while any differences between the miR-200c
mimics and the ZEB2 siRNA group were insignificant (P>0.05;
Fig. 7). These findings suggest
that overexpression of miR-200c and knockdown of ZEB2 enhanced cell
sensitivity to DDP and increased cell apoptosis.
Expression of apoptosis-related
molecules in SGC7901/DDP cells
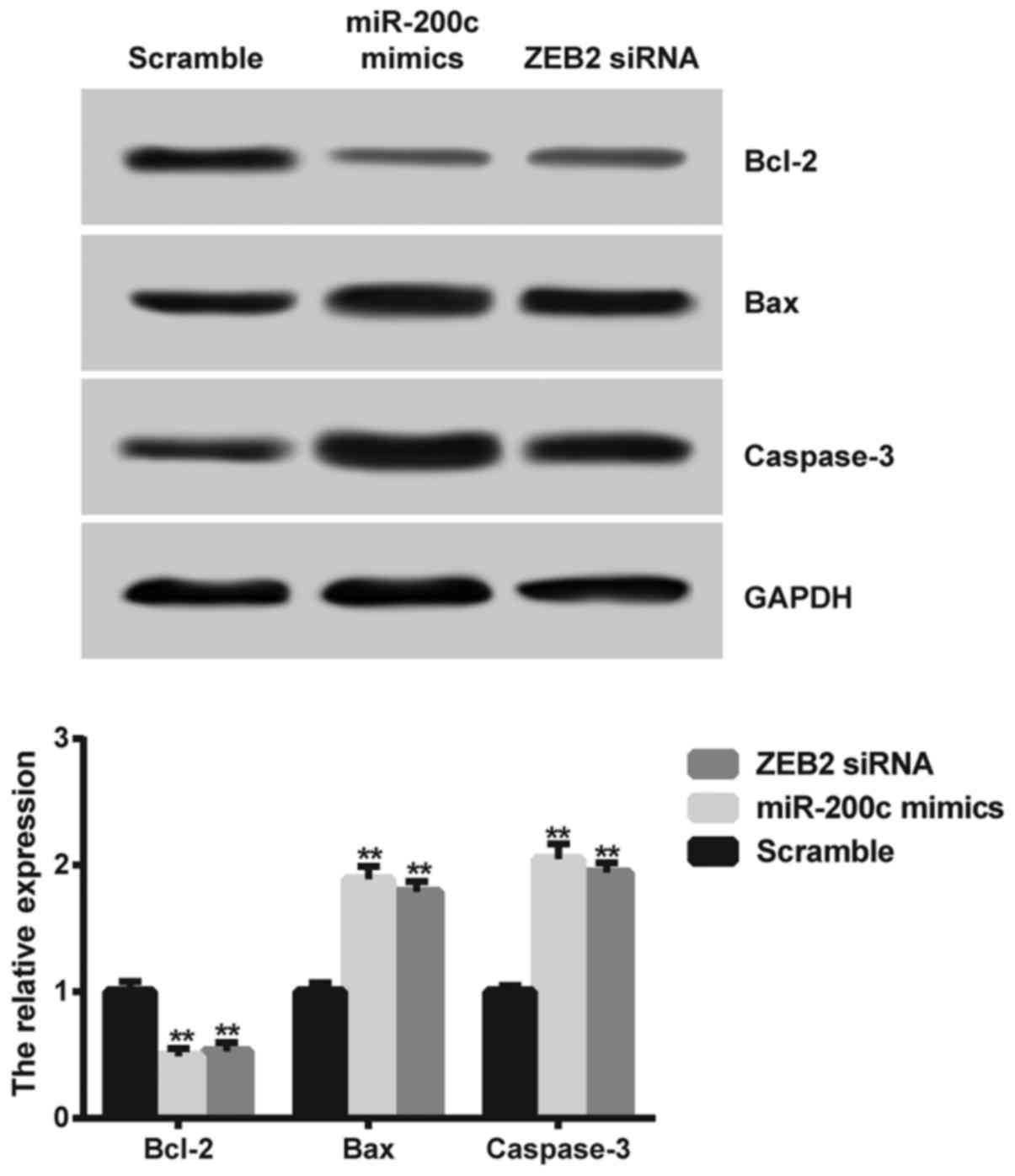
Western blot analysis was carried out in SGC7901/DDP
cells to further clarify the mechanism of miR-200c and ZEB2 in
regulating DDP resistance in gastric cancer. The expression levels
of Bax and caspase-3 were increased in the miR-200c mimic- and the
ZEB2 siRNA-transfected cells, while the expression level of Bcl-2
was markedly decreased compared to the scramble group (P<0.01;
Fig. 8).
Discussion
Gastric cancer is one of the most common malignant
tumors and remains the second largest threat to individuals in the
world (23). The incidence of
gastric cancer is higher in Eastern countries, including China,
Korea, and Japan (24,25). Gastric cancer is often diagnosed at
late stages in which surgical procedures may not be effective
(26,27). On the other hand, resistance to
chemotherapy is another challenge in clinical practice (28–30).
DDP is a popular chemotherapeutic medication which induces tumor
cell death by DNA damage (31).
Therefore, finding the related factors that may affect the
resistance to DDP may improve the survival status of gastric cancer
patients.
miRNAs are a class of endogenous ~22-nucleotide
single-strand and highly conserved non-coding RNAs, which are
forecasted to modulate about 30% of gene expression through the
interference with mRNA translation (32). It has been reported that miRNAs play
a multifunctional role in many biological processes including cell
differentiation, apoptosis, proliferation, tumorigenesis, tumor
development and tumor chemoresistance (33–38).
miR-200c, which belongs to the miR-200 family, is involved in the
inhibition of EMT, tumor invasion, and metastasis (39,40).
Many studies have shown that miR-200c can increase the sensitivity
of cells to antitumor medications in a variety of cancers,
including breast (41,42), ovarian (43) and non-small cell lung cancer
(43). In this study, miR-200c was
downregulated in gastric cancer and human gastric cancer cell line
SGC7901 with DDP-resistance (SGC7901/DDP cells), which is
consistent with the corresponding results from a previous study
(34). Therefore, all of this
evidence suggests that miR-200c may be involved in regulating the
chemoresistance of gastric cancer patients.
Generally, miRNAs mediate a series of biological
processes through different target sites and they also regulate the
expression of their downstream target mRNAs (44–46).
Previous studies have identified many downstream target mRNAs of
miRNA200c, such a RhoE and ZEB1/2 (34,47).
Accordingly, the dual luciferase reporter gene assay in our
experiments suggests that miR-200c could specifically bind with the
3′UTR of ZEB2 and significantly suppress the luciferase activity,
implying that ZEB2 is a direct downstream target gene of miR-200c
in SGC7901/DDP cells.
ZEB2, as a key member of the Snail gene family, is
closely associated with the biological processes of numerous tumors
(48,49). Additionally, it has been reported
that ZEB2 plays a major role in EMT by combining the E-box sequence
of E-cadherin and then suppressing the transcription of numerous
genes (e.g., bridge grain protein, cytokeratin and E-calcium sticky
protein) (50). In our study, ZEB2
was markedly upregulated in gastric cancer tissues and SGC7901/DDP
cells whereas miR-200c exhibited the opposite trend. Furthermore,
our results showed that the upregulation of miR-200c or
downregulation of ZEB2 could increase the sensitivity of
SGC7901/DDP cells to DDP. Previous studies also suggested that
upregulated miR-200c enhanced the sensitivity to chemotherapy in
patients with gastric cancer (17,34).
Meanwhile, downregulation of ZEB2 also enhanced the sensitivity to
chemotherapy in small-cell lung cancer (51). Our results also confirmed that the
upregulation of miR-200c or downregulation of ZEB2 could enhance
the sensitivity of SGC7901/DDP cells to DDP-induced apoptosis and
therefore miR-200c and ZEB2 can potentially regulate chemotherapy
sensitivity through the apoptotic signaling pathway in gastric
cancer patients. Accordingly, it has been reported that miR-200c
not only regulates the induction of apoptosis by targeting FAP-1
(52), but can also mediate the
resistance to breast cancer medications by regulating a series of
apoptosis-related genes (53).
Moreover, knockdown of ZEB2 enhanced the sensitivity of lung cancer
patients to chemotherapy since chemotherapy-induced apoptosis is
potentially stimulated (51).
In the present study, we demonstrated that miR-200c
enhanced the sensitivity of gastric cancer to DDP by directly
targeting ZEB2. However, there are still some limitations in this
study. For example, only the SGC7901/DDP cell line was used in this
study and the detailed molecular mechanism involved remains
unclear. The molecular mechanism of miR-200c and ZEB2 with respect
to DDP-induced apoptosis needs further analysis in the future. In
conclusion, we assessed and reported the effect of miR-200c and its
target gene ZEB2 on DDP resistance in gastric cancer. Our findings
illustrated that miR-200c expression was significantly
downregulated in gastric cancer, while ZEB2 expression exhibited
the opposite trend. Moreover, ZEB2 was found to be a direct target
of miR-200c. Our data further demonstrated that upregulation of
miR-200c and downregulation of ZEB2 could increase the
chemotherapeutic sensitivity of gastric cancer and
chemotherapy-induced apoptosis. Collectively, all of these data
suggest that miR-200c enhanced the sensitivity of gastric cancer to
chemotherapy by directly targeting and regulating the expression of
ZEB2. Thus, both miR-200c and ZEB2 exhibit great potential to serve
as effective therapeutic targets for increasing the sensitivity of
gastric cancer to DDP.
References
|
1
|
Giordano A and Cito L: Advances in gastric
cancer prevention. World J Clin Oncol. 3:128–136. 2012.PubMed/NCBI
|
|
2
|
Ferlay J, Shin HR, Bray F, Forman D,
Mathers C and Parkin DM: Estimates of worldwide burden of cancer in
2008: GLOBOCAN 2008. Int J Cancer. 127:2893–2917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Mbulaiteye SM, Hisada M and El-Omar EM:
Helicobacter pylori associated global gastric cancer burden. Front
Biosci (Landmark Ed). 14:1490–1504. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lee YY and Derakhshan MH: Environmental
and lifestyle risk factors of gastric cancer. Arch Iran Med.
16:358–365. 2013.PubMed/NCBI
|
|
5
|
Schuhmacher C, Reim D and Novotny A:
Neoadjuvant treatment for gastric cancer. J Gastric Cancer.
13:73–78. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lordick F, Lorenzen S, Yamada Y and Ilson
D: Optimal chemotherapy for advanced gastric cancer: Is there a
global consensus? Gastric Cancer. 17:213–225. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Liu YR, Jiang YZ, Zuo WJ, Yu KD and Shao
ZM: PIK3CA mutations define favorable prognostic biomarkers in
operable breast cancer: A systematic review and meta-analysis. Onco
Targets Ther. 7:543–552. 2014.PubMed/NCBI
|
|
8
|
Fabian MR, Sonenberg N and Filipowicz W:
Regulation of mRNA translation and stability by microRNAs. Annu Rev
Biochem. 79:351–379. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bartels CL and Tsongalis GJ: MicroRNAs:
Novel biomarkers for human cancer. Clin Chem. 55:623–631. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ke TW, Wei PL, Yeh KT, Chen WT and Cheng
YW: MiR-92a promotes cell metastasis of colorectal cancer through
PTEN- mediated PI3K/AKT pathway. Ann Surg Oncol. 22:2649–2655.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Krol J, Loedige I and Filipowicz W: The
widespread regulation of microRNA biogenesis, function and decay.
Nat Rev Genet. 11:597–610. 2010.PubMed/NCBI
|
|
12
|
Slabý O, Krekác D, Hrstka R, Svoboda M and
Vyzula R: Involvement of microRNAs in cancer biology and
possibilities of their application to diagnostic and predictive
oncology. Cas Lek Cesk. 147:25–31. 2008.(In Czech). PubMed/NCBI
|
|
13
|
Zhu W, Xu H, Zhu D, Zhi H, Wang T, Wang J,
Jiang B, Shu Y and Liu P: miR-200bc/429 cluster modulates multidrug
resistance of human cancer cell lines by targeting BCL2 and XIAP.
Cancer Chemother Pharmacol. 69:723–731. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Sossey-Alaoui K, Bialkowska K and Plow EF:
The miR200 family of microRNAs regulates WAVE3-dependent cancer
cell invasion. J Biol Chem. 284:33019–33029. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Smith CM, Watson DI, Leong MP, Mayne GC,
Michael MZ, Wijnhoven BP and Hussey DJ: miR-200 family expression
is downregulated upon neoplastic progression of Barrett's
esophagus. World J Gastroenterol. 17:1036–1044. 2011.PubMed/NCBI
|
|
16
|
Zhang Y, Li M, Wang H, Fisher WE, Lin PH,
Yao Q and Chen C: Profiling of 95 microRNAs in pancreatic cancer
cell lines and surgical specimens by real-time PCR analysis. World
J Surg. 33:698–709. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chen Y, Zuo J, Liu Y, Gao H and Liu W:
Inhibitory effects of miRNA-200c on chemotherapy-resistance and
cell proliferation of gastric cancer SGC7901/DDP cells. Chin J
Cancer. 29:1006–1011. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hill L, Browne G and Tulchinsky E:
ZEB/miR-200 feedback loop: At the crossroads of signal transduction
in cancer. Int J Cancer. 132:745–754. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Haslehurst AM, Koti M, Dharsee M, Nuin P,
Evans K, Geraci J, Childs T, Chen J, Li J, Weberpals J, et al: EMT
transcription factors snail and slug directly contribute to
cisplatin resistance in ovarian cancer. BMC Cancer. 12:912012.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Adam L, Zhong M, Choi W, Qi W, Nicoloso M,
Arora A, Calin G, Wang H, Siefker-Radtke A, McConkey D, et al:
miR-200 expression regulates epithelial-to-mesenchymal transition
in bladder cancer cells and reverses resistance to epidermal growth
factor receptor therapy. Clin Cancer Res. 15:5060–5072. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Washington K: 7th edition of the AJCC
cancer staging manual: stomach. Ann Surg Oncol. 17:3077–3079. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yamaue H, Tanimura H, Noguchi K, Hotta T,
Tani M, Tsunoda T, Iwahashi M, Tamai M and Iwakura S:
Chemosensitivity testing of fresh human gastric cancer with highly
purified tumour cells using the MTT assay. Br J Cancer. 66:794–799.
1992. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kelley JR and Duggan JM: Gastric cancer
epidemiology and risk factors. J Clin Epidemiol. 56:1–9. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lee JH, Kim KM, Cheong JH and Noh SH:
Current management and future strategies of gastric cancer. Yonsei
Med J. 53:248–257. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hudler P: Genetic aspects of gastric
cancer instability. ScientificWorldJournal. 2012:7619092012.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yan LH, Wei WY, Cao WL, Zhang XS, Xie YB
and Xiao Q: Overexpression of E2F1 in human gastric carcinoma is
involved in anti-cancer drug resistance. BMC Cancer. 14:9042014.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kang JH, Lee SI, Lim DH, Park KW, Oh SY,
Kwon HC, Hwang IG, Lee SC, Nam E, Shin DB, et al: Salvage
chemotherapy for pretreated gastric cancer: A randomized phase III
trial comparing chemotherapy plus best supportive care with best
supportive care alone. J Clin Oncol. 30:1513–1518. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sun F, Lu X, Li H, Peng Z, Wu K, Wang G
and Tong Q: Special AT-rich sequence binding protein 1 regulates
the multidrug resistance and invasion of human gastric cancer
cells. Oncol Lett. 4:156–162. 2012.PubMed/NCBI
|
|
29
|
Zhang H, Sun LL, Meng YL, Song GY, Hu JJ,
Lu P and Ji B: Survival trends in gastric cancer patients of
Northeast China. World J Gastroenterol. 17:3257–3262.
2011.PubMed/NCBI
|
|
30
|
Maconi G, Manes G and Porro GB: Role of
symptoms in diagnosis and outcome of gastric cancer. World J
Gastroenterol. 14:1149–1155. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Montagnani F, Turrisi G, Marinozzi C,
Aliberti C and Fiorentini G: Effectiveness and safety of
oxaliplatin compared to cisplatin for advanced, unresectable
gastric cancer: a systematic review and meta-analysis. Gastric
Cancer. 14:50–55. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Guo H, Ingolia NT, Weissman JS and Bartel
DP: Mammalian microRNAs predominantly act to decrease target mRNA
levels. Nature. 466:835–840. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang J, Yang M, Li Y and Han B: The role
of microRNAs in the chemoresistance of breast cancer. Drug Dev Res.
76:368–374. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Chang L, Guo F, Wang Y, Lv Y, Huo B, Wang
L and Liu W: MicroRNA-200c regulates the sensitivity of
chemotherapy of gastric cancer SGC7901/DDP cells by directly
targeting RhoE. Pathol Oncol Res. 20:93–98. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Issabekova A, Berillo O, Regnier M and
Anatoly I: Interactions of intergenic microRNAs with mRNAs of genes
involved in carcinogenesis. Bioinformation. 8:513–518. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Hong L, Han Y, Lu Q, Zhang H, Zhao Q, Wu K
and Fan D: Drug resistance-related microRNAs in esophageal cancer.
Expert Opin Biol Ther. 12:1487–1494. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Sato F, Tsuchiya S, Meltzer SJ and Shimizu
K: MicroRNAs and epigenetics. FEBS J. 278:1598–1609. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Huang Y, Shen XJ, Zou Q, Wang SP, Tang SM
and Zhang GZ: Biological functions of microRNAs: A review. J
Physiol Biochem. 67:129–139. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Korpal M and Kang Y: The emerging role of
miR-200 family of microRNAs in epithelial-mesenchymal transition
and cancer metastasis. RNA Biol. 5:115–119. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Hurteau GJ, Carlson JA, Spivack SD and
Brock GJ: Overexpression of the microRNA hsa-miR-200c leads to
reduced expression of transcription factor 8 and increased
expression of E-cadherin. Cancer Res. 67:7972–7976. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Knezevic J, Pfefferle AD, Petrovic I,
Greene SB, Perou CM and Rosen JM: Expression of miR-200c in
claudin-low breast cancer alters stem cell functionality, enhances
chemosensitivity and reduces metastatic potential. Oncogene.
34:5997–6006. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Bai WD, Ye XM, Zhang MY, Zhu HY, Xi WJ,
Huang X, Zhao J, Gu B, Zheng GX, Yang AG, et al: MiR-200c
suppresses TGF-β signaling and counteracts trastuzumab resistance
and metastasis by targeting ZNF217 and ZEB1 in breast cancer. Int J
Cancer. 135:1356–1368. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Liu N, Zeng J, Zhang X, Yang Q, Liao D,
Chen G and Wang Y: Involvement of miR-200a in chemosensitivity
regulation of ovarian cancer. Zhonghua Yi Xue Za Zhi. 94:2148–2151.
2014.(In Chinese). PubMed/NCBI
|
|
44
|
Zheng L, Zhang Y, Liu Y, Zhou M, Lu Y,
Yuan L, Zhang C, Hong M, Wang S and Li X: MiR-106b induces cell
radioresistance via the PTEN/PI3K/AKT pathways and p21 in
colorectal cancer. J Transl Med. 13:2522015. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Ritchie W, Rasko JE and Flamant S:
MicroRNA target prediction and validation. Adv Exp Med Biol.
774:39–53. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Witkos TM, Koscianska E and Krzyzosiak WJ:
Practical aspects of microRNA target prediction. Curr Mol Med.
11:93–109. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Panda H, Pelakh L, Chuang TD, Luo X,
Bukulmez O and Chegini N: Endometrial miR-200c is altered during
transformation into cancerous states and targets the expression of
ZEBsVEGFAFLT1IKKβKLF9, and FBLN5. Reprod Sci. 19:786–796. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Lee JY, Park MK, Park JH, Lee HJ, Shin DH,
Kang Y, Lee CH and Kong G: Loss of the polycomb protein Mel-18
enhances the epithelial-mesenchymal transition by ZEB1 and ZEB2
expression through the downregulation of miR-205 in breast cancer.
Oncogene. 33:1325–1335. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Nam EH, Lee Y, Park YK, Lee JW and Kim S:
ZEB2 upregulates integrin α5 expression through cooperation with
Sp1 to induce invasion during epithelial-mesenchymal transition of
human cancer cells. Carcinogenesis. 33:563–571. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Kakihana M, Ohira T, Chan D, Webster RB,
Kato H, Drabkin HA and Gemmill RM: Induction of E-cadherin in lung
cancer and interaction with growth suppression by histone
deacetylase inhibition. J Thorac Oncol. 4:1455–1465. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Fang S, Zeng X, Zhu W, Tang R, Chao Y and
Guo L: Zinc finger E-box-binding homeobox 2 (ZEB2) regulated by
miR-200b contributes to multi-drug resistance of small cell lung
cancer. Exp Mol Pathol. 96:438–444. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Schickel R, Park SM, Murmann AE and Peter
ME: miR-200c regulates induction of apoptosis through CD95 by
targeting FAP-1. Mol Cell. 38:908–915. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Tryndyak VP, Beland FA and Pogribny IP:
E-cadherin transcriptional down-regulation by epigenetic and
microRNA-200 family alterations is related to mesenchymal and
drug-resistant phenotypes in human breast cancer cells. Int J
Cancer. 126:2575–2583. 2010.PubMed/NCBI
|