Introduction
Alternative splicing is a sophisticated and
ubiquitous nuclear process in eukaryotic cells by which a single
gene may yield various mRNAs, leading to products of multiple
protein isoforms. In humans, more than 70% of genes are
alternatively spliced (1,2), which greatly increases the diversity
of proteins that can be encoded by the surprisingly low number of
human genes. Alternative splicing is known to play a role in the
alteration of the binding properties of proteins, their enzymatic
activity, their intracellular localization, or their stability and
post-translational modifications (3,4).
Recently increasing evidence has revealed that alternative splicing
plays an important role in tumorigenesis and tumor progression, and
splice variants represent potentially useful targets for the
development of novel therapeutic strategies (5,6).
Human protein arginine N-methyltransferase 2 (PRMT2,
HRMT1L1) is a protein that belongs to the arginine
methyltransferase family and has weak methyltransferase activity on
a histone H4 substrate. However, its optimal substrates have not
yet been identified (7,8). Subsequent studies have revealed that
PRMT2 is clearly involved in a variety of cellular processes,
including lung function, the inflammatory response, apoptosis
promotion, and Wnt and leptin signaling regulation (9–13),
suggesting that PRMT2 has diverse roles in transcriptional
regulation through different mechanisms depending on its binding
partners.
Our previous study demonstrated that the PRMT2 gene
produces several isoforms through complex alternative splicing at
the 3′ end of the pre-mRNA, and each has a unique C-terminal
sequence (14). Due to the distinct
gene structure characteristics, each isoform is therefore predicted
to be functionally different. PRMT2β is the most different isoform
compared with the wild-type. Sequence analysis revealed that PRMT2β
deletes exon 7–9 in its coding sequence, and downstream
frame-shifting. As a result, PRMT2β encodes 83 new amino acids at
the C-terminus of the protein. We had previously demonstrated the
negative effect of PRMT2 on breast cancer cell proliferation in
vitro and in vivo. Moreover, cyclin D1 (CCND1) was found
to be downregulated by PRMT2, and PRMT2 was further shown to
suppress the estrogen receptor α (ERα) binding affinity to the
activator protein-1 (AP-1) site in the CCND1 promoter
through indirect binding with the AP-1 site, resulting in the
inhibition of the CCND1 promoter activity in MCF-7 cells
(15). These studies however,
neglected to examine the specific functional contributions of
PRMT2β to breast cancer progression. In the present study, we
revealed that PRMT2β suppressed the cell proliferation and colony
formation of breast cancer MCF7 cells, and induced cell cycle
arrest and apoptosis of MCF7 cells. PRMT2β was revealed to suppress
the transcription activity of the CCND1 promoter, and PRMT2β
was also found to inhibit CCND1 expression via the suppression of
Akt/GSK-3β signaling in breast cancer cells.
Materials and methods
Cell culture
MCF7 and 293T cells were purchased from American
Type Culture Collection (ATCC) (Manassas, VA, USA), and cultured as
previously described (14). Ethics
approval and consent for the use of the human tissue was confirmed
by the Ethics Committee of The First Affiliated Hospital of the
University of South China.
Lentiviral vector construction and
lentivirus infection
Recombined lentiviral vector
pGC-LV-GV308-PRMT2β with the variant PRMT2β
(FJ436411) gene (LV-Tet-on-PRMT2β) and
pGC-LV-GV308-PRMT2 (NM_001535) with the wild-type
PRMT2 (LV-Tet-on-PRMT2) gene were constructed by the
GeneChem Co. (Shanghai, China). The pGC-LV-GV308 vector was
used as a negative control. The packaging plasmid pHelper 1.0 and
pHelper 2.0 were purchased from GeneChem Co.
We co-transfected the pGC-LV-GV308-PRMT2β or
pGC-LV-GV308-PRMT2 vectors with the pHelper 1.0 and pHelper
2.0 packaging plasmid into 293T cells to generate recombinant
lentiviruses. Culture medium was collected 72 h post-transfection,
and MCF7 cells were then infected with the aforementioned
lentiviruses. A total of 5×105 MCF7 cells were seeded
into a 6-well cell plate and further incubated for 12 h to reach
30% confluency, and then infected with LV-Tet-on-PRMT2β (PRMT2β
overexpression group) or LV-Tet-on-PRMT2 (PRMT2 overexpression
group) for 48 h in the presence of 8 µg/ml of Polybrene
(Sigma-Aldrich, St. Louis, MO, USA). Stable cell lines either with
the PRMT2β-3Flag or the PRMT2-3Flag were obtained after infection
with LV-tet-on-PRMT2β or LV-tet-on-PRMT2 cells which were selected
by puromycin for 2 weeks. Western blot analysis was performed to
verify the expression of PRMT2β-3Flag or PRMT2-3Flag induced by 5
µg/ml of Dox in the infected MCF7 cells.
Cell proliferation and colony
formation assays
For cell proliferation assays, MCF7 cells were
seeded in a 96-well plate (2,000 cells/well) and counted using an
automated cell counter (Nexcelom Bioscience, Lawrence, MA, USA).
For the colony formation assay, the cells were seeded in a 6-well
plate (600 cells/well) and maintained for 8 days. The culture
medium with or without 5 µg/ml of Dox was changed every 2 days.
Each experiment was carried out in triplicate and performed at
least twice.
Soft agar colony formation assay
The assay was performed in 6-well plates. A bottom
layer of agar (0.6%) with enriched Dulbeccos modified Eagles medium
(DMEM), [final 10% fetal bovine serum (FBS)] was poured first.
After the bottom agar solidified, MCF7 cells (3.0×103)
were seeded in top agar (0.3%) with enriched DMEM supplemented with
20% FBS and incubated at 37°C for 15 days. The culture medium with
or without 5 µg/ml of Dox was changed twice weekly. Colonies were
visualized by staining for 1 h with 0.005% crystal violet.
Cell cycle analysis by flow
cytometry
After transfection, the MCF7 cells were plated in
6-well plates at 2×105 cells/well. Following the
designated treatments, the cells were harvested by trypsinization
and washed with PBS and fixed in ice-cold 75% ethanol overnight at
−20°C. The fixed cells were washed, and dissolved in RNAse and
subsequently incubated at 37°C for 30 min. Next, the cells were
stained with propidium iodide (PI) for 30 min. The DNA content of
the cells (1×104 cells/experimental group) was
determined using a BD Accuri C6 flow cytometer (BD Biosciences,
Franklin Lakes, NJ, USA).
Transient transfection and luciferase
activity assay
Transient gene delivery was carried out to assess
the effect of PRMT2β and PRMT2 on the CCND1 promoter
activity in 293T cells as previously described (14). A luciferase assay kit (Promega,
Madison, WI, USA) was used to assess the reporter activity
according to the manufacturer's instructions. Luciferase activity
was normalized using a Renilla luciferase internal
control.
Western blot analysis
Total cell or tissue lysates were lysed on ice for
30 min. Soluble proteins (20 µg) were probed with anti-PRMT2,
anti-Flag and anti-CCND1 antibodies (1:500; Abcam, Cambridge, MA,
USA) and anti-p-AKT, anti-AKT, anti-p-GSK-3β and anti-GSK-3β
(1:800; Cell Signaling Technology, Danvers MA, USA). Loading
variations were normalized against β-actin, which was identified by
anti-β-actin monoclonal Ab (1:1,000; Abcam).
Tissue microarray and
immunohistochemical analysis
The tissue microarray (BR1503b; US Biomax, Inc.,
Rockville, MD, USA) consisting of 138 breast malignant tumor cases,
6 normal cases and 6 benign tumor cases was utilized, and was
histologically interpretable and analyzed for the correlation with
clinicopathological parameters. IHC staining was performed as
detailed in our previous study (14). The rabbit polyclonal PRMT2 (1:50;
Aviva Systems Biology, Beijing, China) was used.
We developed affinity-purified rabbit polyclonal
anti-PRMT2β Abs as a custom service from Boster (Wuhan, China). The
Abs were raised against a synthetic peptide antigen corresponding
to the unique C-terminal 15 aa of PRMT2β (HLEMTVDALFGKQCA). The
specificity of the Ab was tested in the PRMT2β expression
vector-transfected 293T cells that do not express endogenous
PRMT2β. Immunofluorescence was used to demonstrate immunoreactive
signals only in transfectants with the PRMT2β-expressing vectors,
but not in transfectants harboring an empty expression vector.
Statistical analysis
All experiments were performed using 3 replicates
and the results are expressed as the mean ± SEM or mean ± SD.
Statistical analysis was carried out using SPSS, version 13.0
(SPSS, Inc., Chicago, IL, USA). A statistical association between
clinicopathological and molecular parameters was tested, using the
non-parametrically two-tailed Mann-Whitney U test. p-values
<0.05 were considered significant. Spearman's rank correlation
coefficients were used to assess the correlation of PRMT2β and
PRMT2 expression with clinicopathological parameters.
Results
PRMT2β suppresses the cell
proliferation and colony formation of breast cancer MCF7 cells
Our previous study reported that suppression of
PRMT2 expression promoted the proliferation of breast cancer cells
(15). To address the cellular and
molecular mechanisms of the alternatively spliced PRMT2 in breast
cancer cells, a tetracycline (doxycycline hyclate, Dox)-inducible
lentiviral system was established to overexpress PRMT2β or PRMT2,
and the pGC-LV-GV308 vector was used as a negative control.
Stable cell lines either with the PRMT2β-3Flag or the PRMT2-3Flag
were obtained after selection with puromycin for 2 weeks. Western
blot analysis was performed to verify the expression of
PRMT2β-3Flag or PRMT2-3Flag induced by Dox in the infected MCF7
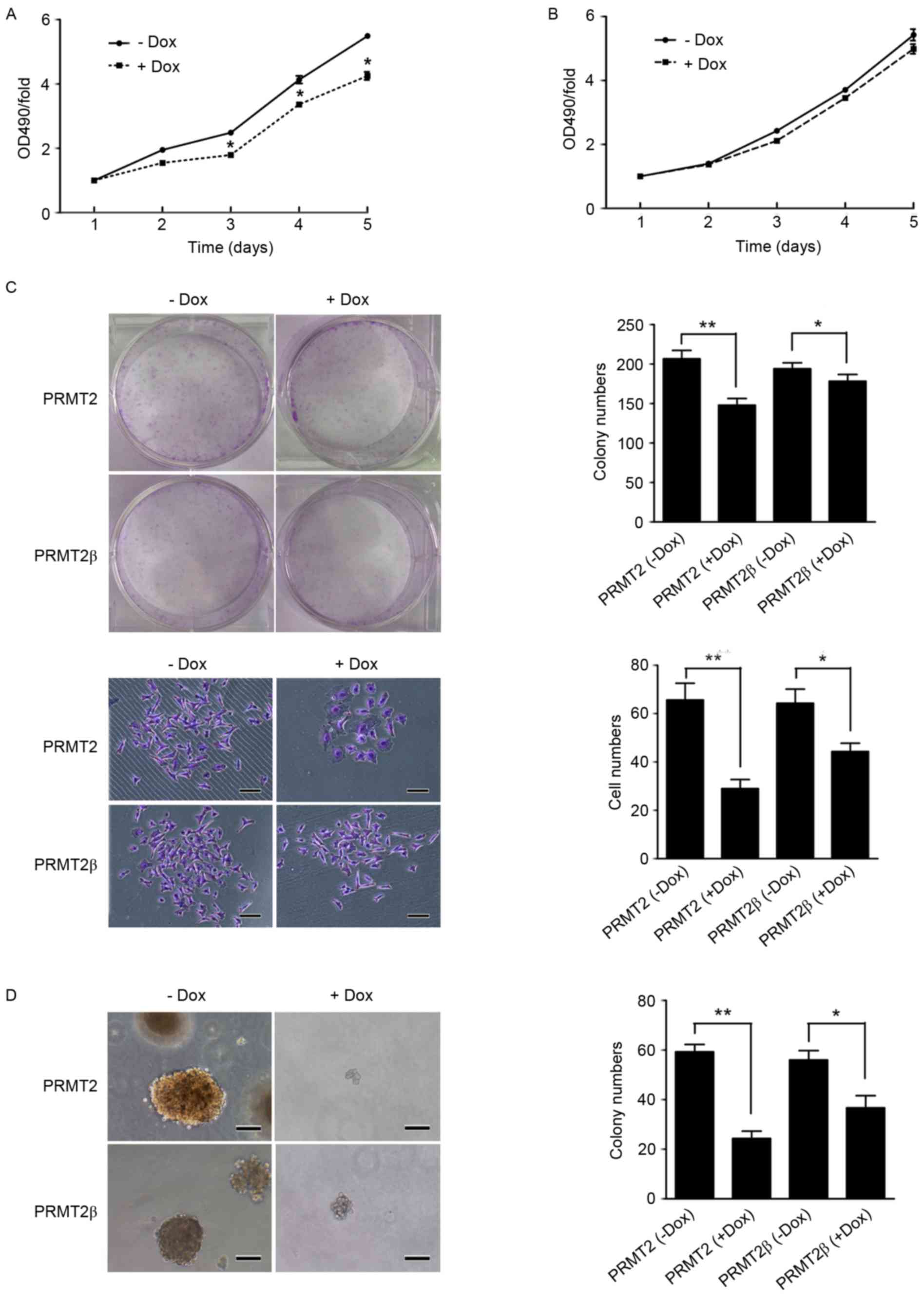
cells (data not shown). As shown in Fig. 1A, with treatment of 5 µg/ml of Dox,
MCF7 cells carrying lentivirus PRMT2 expression exhibited a
markedly decreased proliferative ability compared to those without
treatment of 5 µg/ml of Dox at the indicated time-points
(p<0.05; Fig. 1A). Whereas, MCF7
cells with lentivirus PRMT2β expression exhibited a weakly
decreased proliferative ability compared to those without treatment
of 5 µg/ml of Dox (Fig. 1B). The
proliferative ability exhibited no difference in the negative
control group when treated with or without 5 µg/ml of Dox (data not
shown).
Colony formation assay shown in Fig. 1C, with treatment of 5 µg/ml of Dox,
revealed that both the number of formed colonies and the cell
number of each colony from the MCF7 cells carrying lentivirus PRMT2
expression were markedly decreased compared to the cells without
treatment of 5 µg/ml of Dox (p<0.01; Fig. 1C). Whereas, with treatment of Dox,
both the number of colonies formed and the cell number of each
colony from the MCF7 cells carrying lentivirus PRMT2β expression
were weakly decreased compared to the cells without treatment of 5
µg/ml of Dox (p<0.05; Fig. 1C).
Meanwhile, with soft agar colony formation assay we found that both
the number and the size of formed colonies from the MCF7 cells
carrying lentivirus PRMT2β expression were significantly decreased
compared to the cells without treatment of 5 µg/ml of Dox
(p<0.05; Fig. 1D). Similar
results were also observed in the cells carrying lentivirus of
PRMT2 expression, and the proliferation inhibition effect PRMT2 was
shown to be more profound than that of PRMT2β (p<0.01; Fig. 1D). These results indicated that
PRMT2β suppressed the cell proliferation and colony formation of
breast cancer MCF7 cells.
PRMT2β induces cell cycle arrest and
apoptosis of MCF7 cells
We then carried out cell cycle analysis to further
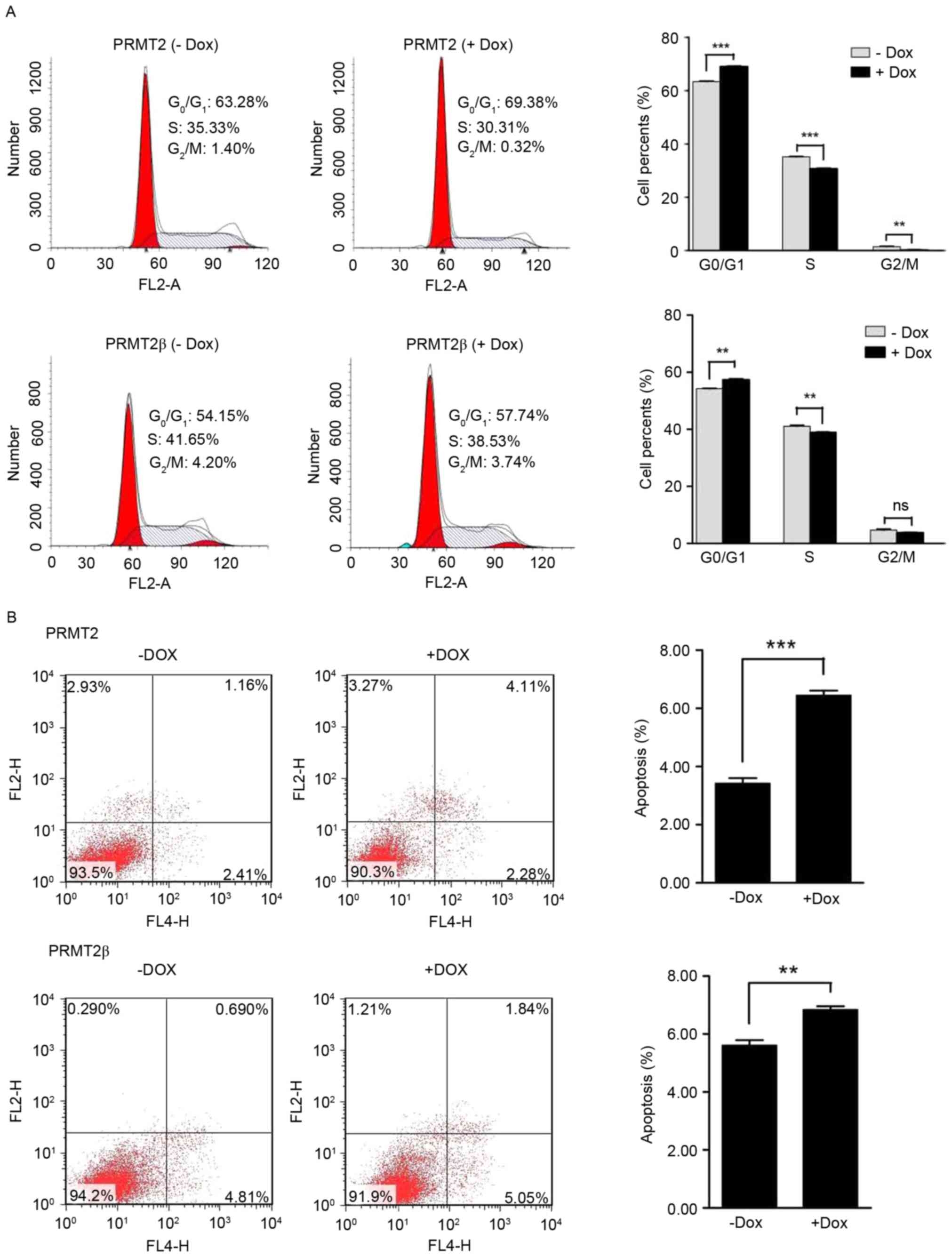
characterize the suppressive effects of PRMT2β on the cell cycle.
MCF7 cell populations in the G0-G1 and S
phases were 57.74 and 38.53% in the PRMT2β (+ Dox) group, while
they were 54.15 and 41.65% in the PRMT2β (− Dox) group. After 3
independent experiments, in the PRMT2β (+ Dox) group, the
population of cells in the G0-G1 phase
increased (vs. - Dox, p<0.01; Fig.
2A) and the population of cells in the S phase significantly
decreased (vs. - Dox, p<0.01; Fig.
2A). Moreover, MCF7 cell populations in the
G0-G1 and S phases were 69.38% and 30.31% in
the PRMT2 (+ Dox) group, while they were 63.28% and 35.33% in the
PRMT2 (− Dox) group. After 3 independent experiments, in the PRMT2
(+ Dox) group, the population of cells in the
G0-G1 phase increased (vs. - Dox, p<0.001;
Fig. 2A) and the population of
cells in the S phase significantly decreased (vs. - Dox,
p<0.001; Fig. 2A). These results
revealed that PRMT2β significantly induced
G0-G1 phase arrest in MCF7 cells.
We further examined the effect of PRMT2β on
apoptosis in MCF7 cells. As shown in Fig. 2B, the percentage of apoptotic cells
was 6.89% in the PRMT2β (+ Dox) group, and 5.5% in the PRMT2β (−
Dox) group. After 3 independent experiments, in the PRMT2β (+ Dox)
group, the apoptosis of the MCF7 cells increased (vs. - Dox,
p<0.01; Fig. 2B). Furthermore,
the percentage of apoptotic MCF7 cells in the PRMT2 (+ Dox) group
was 6.39%, and 3.57% in the PRMT2 (− Dox) group. After 3
independent experiments, in the PRMT2 (+ Dox) group, the apoptosis
of the MCF7 cells increased (vs. - Dox, p<0.001; Fig. 2B). These results revealed that
PRMT2β induced the apoptosis in MCF7 cells.
PRMT2β antagonizes the transcription
suppressive activity of PRMT2 on the CCND1 promoter
In a previous study, we identified that PRMT2
suppresses the CCND1 promoter activity through indirect
binding with the AP-1 site in the CCND1 promoter and
functions as a transcription coregulator (15). To understand the involvement of
PRMT2β in gene transcription regulation, 293T cells were
co-transfected with pcDNA3.1-PRMT2 and/or pcDNA3.1-PRMT2β and
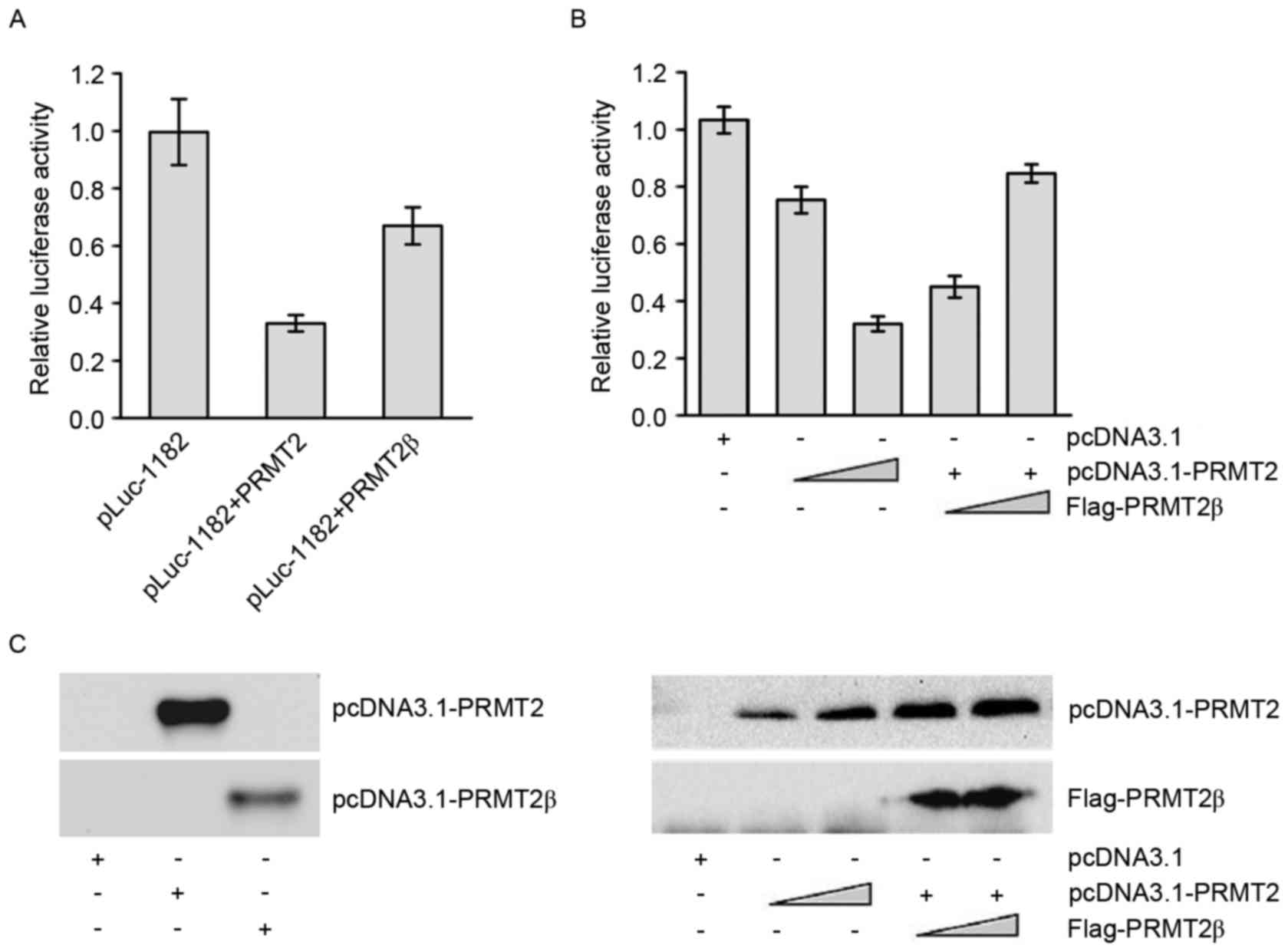
CCND1 luciferase reporter (pLuc-1182). The results indicated
that transfection of either pcDNA3.1-PRMT2 or pcDNA3.1-PRMT2β
resulted in the suppression of the reporter activity (Fig. 3A), and the suppressive activity of
pcDNA3.1-PRMT2β was weaker compared with that of pcDNA3.1-PRMT2.
Co-transfection of FLAG-PRMT2β with pcDNA3.1-PRMT2 led to a
relieved suppression of the reporter activity by PRMT2 (Fig. 3B), suggesting that PRMT2β could
antagonize the transcription suppression activity of PRMT2. The
protein expression of pcDNA3.1-PRMT2, pcDNA3.1-PRMT2β and
FLAG-PRMT2β was examined by western blotting with anti-PRMT2 and
anti-FLAG antibodies (Fig. 3C).
PRMT2β inhibits CCND1 expression via
the suppression of Akt/GSK-3β signaling in breast cancer cells
Our previous study reported that PRMT2 suppressed
the CCND1 expression in MCF-7 cells partially via the suppression
of Akt/GSK-3β signaling. We therefore explored whether PRMT2β was
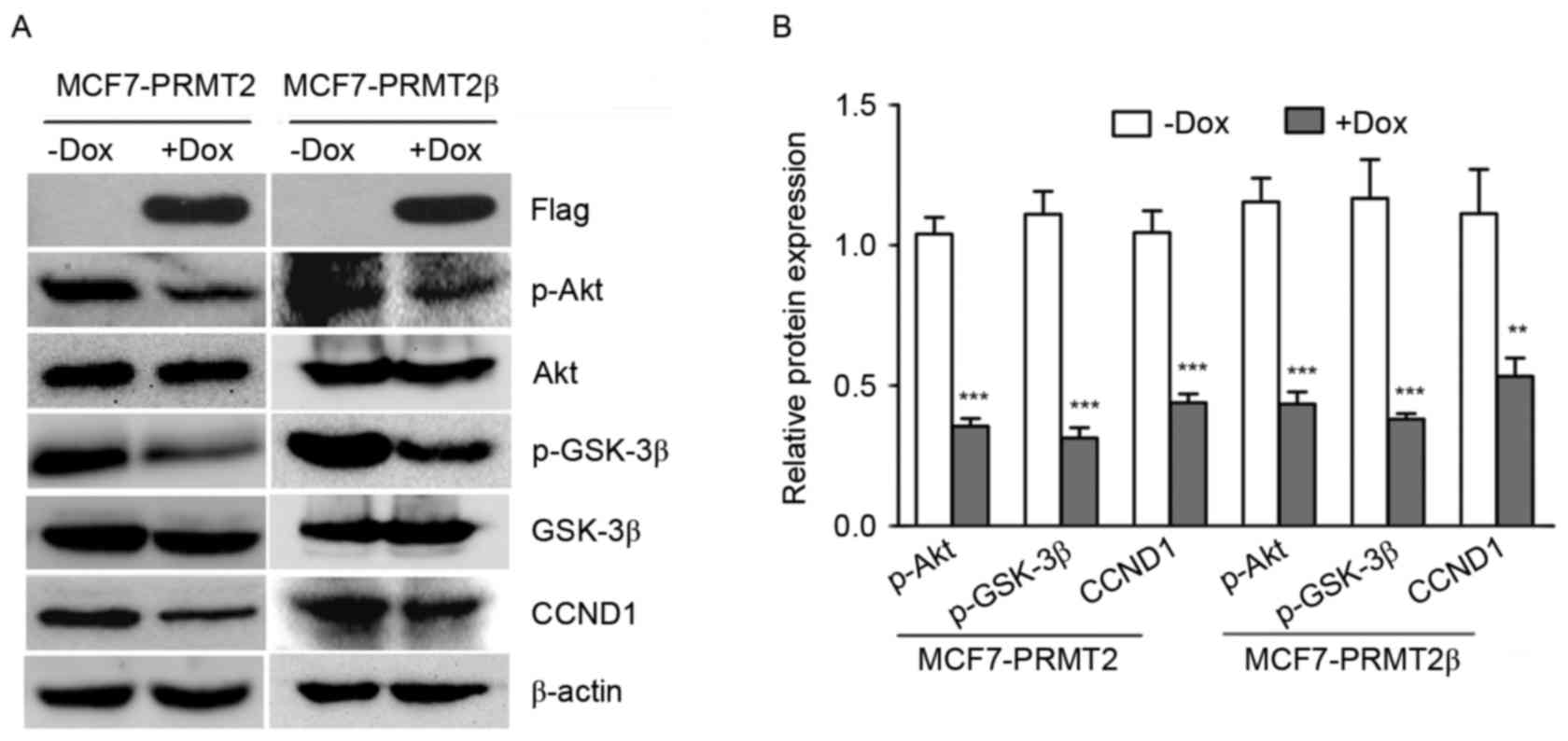
involved in the Akt/GSK-3β/CCND1 axis in MCF7 cells. As shown in
Fig. 4, with treatment of 5 µg/ml
of Dox, the expression of p-Akt and p-GSK3β in MCF7 cells carrying
lentivirus PRMT2β expression was markedly decreased compared to the
cells without treatment of 5 µg/ml of Dox. Lentivirus-mediated
PRMT2β overexpression also markedly suppressed the expression of
CCND1, suggesting that both the overexpression of PRMT2β and PRMT2
suppressed Akt/GSK-3β/CCND1 signaling. These results support the
notion that PRMT2β suppressed the CCND1 expression in MCF-7 cells
partially via the suppression of Akt/GSK-3β signaling.
PRMT2β expression is negatively
correlated with human epidermal growth factor receptor 2
(HER2)
To further identify the association of PRMT2β with
breast cancer progression, a tissue microarray (BR1503b; US Biomax,
Inc.) consisting of 138 breast malignant tumor cases, 6 normal
cases and 6 benign tumor cases was used. The tissue microarray
analysis revealed that the PRMT2β protein expression exhibited a
decreased tendency in different tissue types: 66.7% in normal
breast tissue, 50.0% in breast benign tumors and 29.0% in breast
malignant tumors, although the statistical analysis revealed no
significant difference (p=0.096; Table
I). Conversely, the percentage of cases with PRMT2 protein
expression increased from normal breast tissue to breast malignant
tumors: 33.3% in normal breast tissue, 83.3% in breast benign
tumors and 95.7% in breast malignant tumors, and the difference was
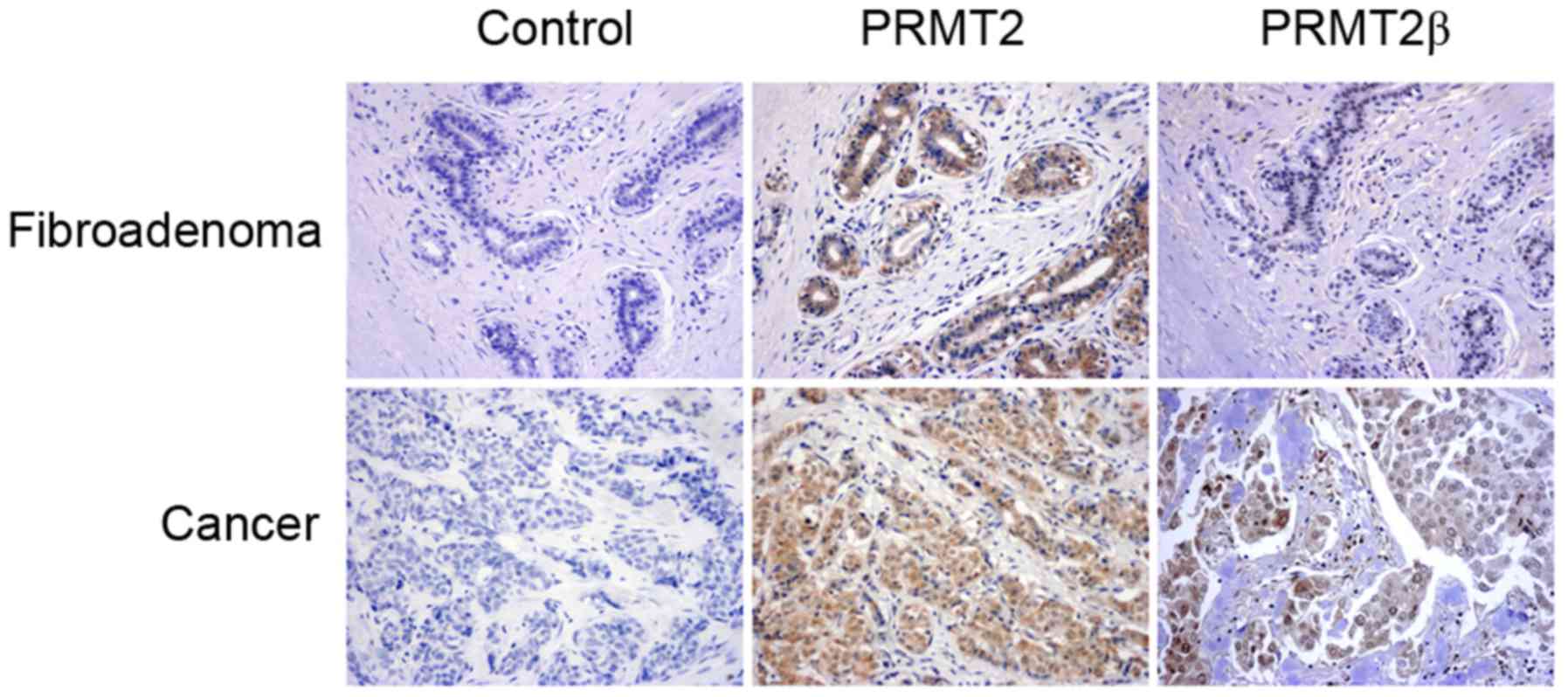
significant (p=0.000; Table I). The
immunostainings of breast fibroadenoma and carcinoma tissue of
representative cases are shown in Fig.
5. Furthermore, statistical analysis confirmed that the PRMT2β
expression was negatively correlated with HER2 (p=0.033; Table II), when analyzed regardless of
breast tumor types. PRMT2β expression in breast carcinoma did not
exhibit a statistically significant correlation with androgen
receptor (AR), estrogen receptor (ER), progesterone receptor (PR),
Ki-67 and the p53 status of the tumor (p>0.05; Table II). PRMT2 expression in breast
carcinoma also exhibited a statistically significant correlation
with the PR and HER2 status (p=0.039 and 0.019, respectively;
Table II).
 | Table I.Expression of PRMT2β and PRMT2 in
normal breast tissue and tumors. |
Table I.
Expression of PRMT2β and PRMT2 in
normal breast tissue and tumors.
|
|
| PRMT2 |
| PRMT2β |
|
|---|
|
|
|
|
|
|
|
|---|
|
| n | Neg. | Pos. | P-value | Neg. | Pos. | P-value |
|---|
| Tissue types |
|
|
| 0.000 |
|
| 0.096 |
|
Normal |
6 | 4 | 2 |
| 2 | 4 |
|
|
Benign |
6 | 1 | 5 |
| 3 | 3 |
|
|
Malignant | 138 | 6 | 132 |
| 98 | 40 |
|
 | Table II.Correlation of PRMT2β and PRMT2
expression with clinicopathological parameters in breast
carcinoma. |
Table II.
Correlation of PRMT2β and PRMT2
expression with clinicopathological parameters in breast
carcinoma.
|
|
| PRMT2 |
| PRMT2β |
|
|---|
|
|
|
|
|
|
|
|---|
|
| n | − | + | ++ | +++ | P-value | − | + | ++ | P-value |
|---|
| AR |
|
|
|
|
| 0.641 |
|
|
| 0.430 |
| − | 95 | 5 | 62 | 25 | 3 |
| 65 | 25 | 5 |
|
| + | 21 | 1 | 18 | 2 | 0 |
| 15 | 6 | 0 |
|
| ++ | 21 | 0 | 11 | 9 | 1 |
| 16 | 4 | 1 |
|
|
+++ | 1 | 0 | 1 | 0 | 0 |
| 1 | 0 | 0 |
|
| ER |
|
|
|
|
| 0.714 |
|
|
| 0.844 |
| − | 91 | 5 | 59 | 22 | 5 |
| 65 | 20 | 6 |
|
| + | 11 | 0 | 10 | 1 | 0 |
| 8 | 3 | 0 |
|
| ++ | 10 | 0 | 7 | 3 | 0 |
| 8 | 2 | 0 |
|
|
+++ | 26 | 1 | 15 | 10 | 0 |
| 16 | 10 | 0 |
|
| PR |
|
|
|
|
| 0.039 |
|
|
| 0.766 |
| − | 100 | 4 | 72 | 19 | 5 |
| 71 | 25 | 4 |
|
| + | 15 | 1 | 8 | 6 | 0 |
| 12 | 3 | 0 |
|
| ++ | 9 | 0 | 4 | 5 | 0 |
| 8 | 1 | 0 |
|
|
+++ | 14 | 0 | 8 | 6 | 0 |
| 7 | 5 | 2 |
|
| HER2 |
|
|
|
|
| 0.019 |
|
|
| 0.033 |
| − | 113 | 3 | 82 | 28 | 0 |
| 76 | 31 | 6 |
|
| + | 17 | 2 | 7 | 5 | 3 |
| 15 | 2 | 0 |
|
| ++ | 7 | 1 | 1 | 3 | 2 |
| 6 | 1 | 0 |
|
|
+++ | 1 | 0 | 1 | 0 | 0 |
| 1 | 0 | 0 |
|
| Ki-67 |
|
|
|
|
| 0.400 |
|
|
| 0.738 |
| − | 83 | 4 | 57 | 19 | 3 |
| 59 | 20 | 4 |
|
| + | 19 | 2 | 9 | 8 | 0 |
| 14 | 4 | 1 |
|
| ++ | 24 | 0 | 16 | 6 | 2 |
| 19 | 4 | 1 |
|
|
+++ | 12 | 0 | 9 | 3 | 0 |
| 5 | 7 | 0 |
|
| p53 |
|
|
|
|
| 0.648 |
|
|
| 0.296 |
| − | 50 | 2 | 32 | 16 | 0 |
| 35 | 13 | 2 |
|
| + | 38 | 1 | 30 | 7 | 0 |
| 24 | 12 | 2 |
|
| ++ | 14 | 1 | 7 | 5 | 1 |
| 10 | 2 | 2 |
|
|
+++ | 36 | 2 | 22 | 8 | 4 |
| 29 | 7 | 0 |
|
Discussion
Protein arginine methyltransferases (PRMTs) catalyze
the methylation of a variety of protein substrates, and there are 9
PRMTs encoded in mammalian genomes. PRMTs are generally
ubiquitously expressed and involved in multiple cellular processes
including cell growth (16),
differentiation (17),
embryogenesis (18),
nuclear/cytoplasmic protein shuttling (19), RNA splicing and transport (20,21),
and post-transcriptional regulation (22). Dysregulation of PRMTs may probably
be implicated in the pathogenesis of some different diseases,
particularly cancer. Accumulating evidence has revealed that
overexpression of these enzymes is often associated with various
cancers (23–26), which may make some of them viable
targets for therapeutic strategies (6). Recently, the existence of
alternatively spliced isoforms of PRMTs provides an additional
layer of complexity. Specifically, PRMT1, PRMT2, CARM1 and PRMT7
have been shown to have alternative isoforms and most of them play
an essential role on the progression and aggressiveness of cancer
(24,27–30).
Our previous study found several alternative
splicing variants of PRMT2, which are missing different exons in
the C-terminals of the wild-type PRMT2 gene, and produce different
isoforms. Particularly, due to frame-shifting, PRMT2β loses
original structure, and encodes a novel amino acid sequence at the
C-terminus of the protein (14).
PRMT2 suppressed E2F1 transcriptional activity in an RB-dependent
manner (31). PRMT2 also inhibited
NF-κB function and promoted cell apoptosis (11). Our previous study found that
suppression of PRMT2 expression promoted the cell proliferation of
breast cancer MCF7 cells (15). To
explore the role of PRMT2β in the cell proliferation, apoptosis and
cell cycle in breast cancer cells, stable cell lines with
PRMT2β-3Flag or PRMT2-3Flag were established. We found that the
expression of PRMT2β suppressed the cell proliferation and colony
formation of MCF7 cells. We also found that PRMT2β expression
induced cell cycle arrest and apoptosis of MCF7 cells. In agreement
with our previous observations, the wild-type PRMT2 also inhibited
breast cancer cell growth. Furthermore, the function of PRMT2β was
less efficient than that of the wild-type PRMT2. This may be due to
the loss of exons 7–10 in PRMT2β, which affects the location of
some nuclear receptor-binding interfaces in the C-terminus.
However, the mechanism requires further exploration (32).
In a previous study, we identified that PRMT2
suppresses the CCND1 promoter activity through indirect
binding with the AP-1 site in the CCND1 promoter, and
functions as a transcription coregulator (15). To understand the cellular function
of PRMT2β, we further investigated the role of PRMT2β in
transcription regulation with 293T cells. We found that both PRMT2β
and PRMT2 suppressed the activity of the CCND1 promoter, and that
the suppressive activity of PRMT2β was weaker compared with that of
PRMT2. With coexpression of PRMT2β and PRMT2, the bigger the amount
of PRMT2β was, the weaker the inhibition of the CCND1 promoter
activity was. We speculated that PRMT2β antagonizes the
transcription suppressive activity of PRMT2 by competitively
binding to the AP-l site of the CCND1 promoter, attenuating
the transcriptional suppression of PRMT2. It is well known that
CCND1 overexpression is a common event in cancer and usually leads
to defective modulation at the post-translational level (33,34).
Therefore, regulation of the CCND1 protein level is one of the
critical aspects in cell proliferation and tumor progression. We
had previously demonstrated that miRNA-mediated PRMT2
downregulation promotes CCND1 expression via activation of
Akt/GSK-3β/CCND1 signaling in breast cancer cells (15). In the present study, we confirmed
that lentivirus-mediated PRMT2β expression also significantly
decreased Akt/GSK-3β/CCND1 signaling, consistent with the results
of wild-type PRMT2, indicating that PRMT2β functions as another
negative modulator of the Akt/GSK-3β/CCND1 axis leading to the
suppression of proliferation of breast cancer cells.
The tissue microarray analysis revealed that the
PRMT2β protein expression exhibited a decreased expression in
breast malignant tissue, although the statistical analysis revealed
that this difference was not significant, indicating that PRMT2β
plays a role in normal breast development and that the loss of
PRMT2β expression may be related with the progression of breast
cancer, consistent with the results from a cell model. Further
analysis of the tissue microarray data revealed that PRMT2β
expression was negatively correlated with HER2 (p=0.033), although
the molecular mechanism remains unknown. Further investigations are
required to clarify the alternative molecular mechanisms of PRMT2β
in the progression of breast cancer. In summary, our results
revealed that PRMT2β, a novel splice variant of PRMT2, has a
potential antitumor effect through the suppression of CCND1
expression and inhibition of Akt signaling activity. This also
opens a new avenue for treating breast cancer.
Acknowledgements
The present study was supported by projects from the
National Natural Science Foundation of China (grant nos. 31200573,
81272355 and 81472608), the Key Project of the Education Department
of Hunan Province (16A189), the Natural Science Foundation of Hunan
Province (2016JJ4077) and the Young Talents Program of the
University of South China.
Glossary
Abbreviations
Abbreviations:
|
PRMT2
|
arginine N-methyltransferase 2
|
|
AR
|
androgen receptor
|
|
ER
|
estrogen receptor
|
|
PR
|
progesterone receptor
|
|
HER2
|
human epidermal growth factor receptor
2
|
|
AP-1
|
activator protein-1
|
|
Dox
|
doxycycline hyclate
|
|
FBS
|
fetal bovine serum
|
|
GFP
|
green fluorescent protein
|
References
|
1
|
Modrek B and Lee C: A genomic view of
alternative splicing. Nat Genet. 30:13–19. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Johnson JM, Castle J, Garrett-Engele P,
Kan Z, Loerch PM, Armour CD, Santos R, Schadt EE, Stoughton R and
Shoemaker DD: Genome-wide survey of human alternative pre-mRNA
splicing with exon junction microarrays. Science. 302:2141–2144.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Resch A, Xing Y, Modrek B, Gorlick M,
Riley R and Lee C: Assessing the impact of alternative splicing on
domain interactions in the human proteome. J Proteome Res. 3:76–83.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sorek R, Shamir R and Ast G: How prevalent
is functional alternative splicing in the human genome? Trends
Genet. 20:68–71. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Baldwin RM, Morettin A and Côté J: Role of
PRMTs in cancer: Could minor isoforms be leaving a mark? World J
Biol Chem. 5:115–129. 2014.PubMed/NCBI
|
|
6
|
Yang Y and Bedford MT: Protein arginine
methyltransferases and cancer. Nat Rev Cancer. 13:37–50. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Herrmann F, Pably P, Eckerich C, Bedford
MT and Fackelmayer FO: Human protein arginine methyltransferases in
vivo - distinct properties of eight canonical members of the PRMT
family. J Cell Sci. 122:667–677. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lakowski TM and Frankel A: Kinetic
analysis of human protein arginine N-methyltransferase 2: Formation
of monomethyl- and asymmetric dimethyl-arginine residues on histone
H4. Biochem J. 421:253–261. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Besson V, Brault V, Duchon A, Togbe D,
Bizot JC, Quesniaux VF, Ryffel B and Hérault Y: Modeling the
monosomy for the telomeric part of human chromosome 21 reveals
haploinsufficient genes modulating the inflammatory and airway
responses. Hum Mol Genet. 16:2040–2052. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Blythe SA, Cha SW, Tadjuidje E, Heasman J
and Klein PS: beta-Catenin primes organizer gene expression by
recruiting a histone H3 arginine 8 methyltransferase, Prmt2. Dev
Cell. 19:220–231. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ganesh L, Yoshimoto T, Moorthy NC, Akahata
W, Boehm M, Nabel EG and Nabel GJ: Protein methyltransferase 2
inhibits NF-kappaB function and promotes apoptosis. Mol Cell Biol.
26:3864–3874. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Iwasaki H, Kovacic JC, Olive M, Beers JK,
Yoshimoto T, Crook MF, Tonelli LH and Nabel EG: Disruption of
protein arginine N-methyltransferase 2 regulates leptin
signaling and produces leanness in vivo through loss of STAT3
methylation. Circ Res. 107:992–1001. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yildirim AO, Bulau P, Zakrzewicz D,
Kitowska KE, Weissmann N, Grimminger F, Morty RE and Eickelberg O:
Increased protein arginine methylation in chronic hypoxia: Role of
protein arginine methyltransferases. Am J Respir Cell Mol Biol.
35:436–443. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhong J, Cao RX, Zu XY, Hong T, Yang J,
Liu L, Xiao XH, Ding WJ, Zhao Q, Liu JH, et al: Identification and
characterization of novel spliced variants of PRMT2 in breast
carcinoma. FEBS J. 279:316–335. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhong J, Cao RX, Liu JH, Liu YB, Wang J,
Liu LP, Chen YJ, Yang J, Zhang QH, Wu Y, et al: Nuclear loss of
protein arginine N-methyltransferase 2 in breast carcinoma is
associated with tumor grade and overexpression of cyclin D1
protein. Oncogene. 33:5546–5558. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lin WJ, Gary JD, Yang MC, Clarke S and
Herschman HR: The mammalian immediate-early TIS21 protein and the
leukemia-associated BTG1 protein interact with a protein-arginine
N-methyltransferase. J Biol Chem. 271:15034–15044. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chen SL, Loffler KA, Chen D, Stallcup MR
and Muscat GEO: The coactivator-associated arginine
methyltransferase is necessary for muscle differentiation: CARM1
coactivates myocyte enhancer factor-2. J Biol Chem. 277:4324–4333.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Torres-Padilla ME, Parfitt DE, Kouzarides
T and Zernicka-Goetz M: Histone arginine methylation regulates
pluripotency in the early mouse embryo. Nature. 445:214–218. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
McBride AE and Silver PA: State of the
arg: Protein methylation at arginine comes of age. Cell. 106:5–8.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lukong KE and Richard S: Arginine
methylation signals mRNA export. Nat Struct Mol Biol. 11:914–915.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Meister G and Fischer U: Assisted RNP
assembly: SMN and PRMT5 complexes cooperate in the formation of
spliceosomal UsnRNPs. EMBO J. 21:5853–5863. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Li H, Park S, Kilburn B, Jelinek MA,
Henschen-Edman A, Aswad DW, Stallcup MR and Laird-Offringa IA:
Lipopolysaccharide-induced methylation of HuR, an mRNA-stabilizing
protein, by CARM1. Coactivator-associated arginine
methyltransferase. J Biol Chem. 277:44623–44630. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang B, Dong S, Li Z, Lu L, Zhang S, Chen
X, Cen X and Wu Y: Targeting protein arginine methyltransferase 5
inhibits human hepatocellular carcinoma growth via the
downregulation of beta-catenin. J Transl Med. 13:3492015.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Goulet I, Gauvin G, Boisvenue S and Côté
J: Alternative splicing yields protein arginine methyltransferase 1
isoforms with distinct activity, substrate specificity, and
subcellular localization. J Biol Chem. 282:33009–33021. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
El Messaoudi S, Fabbrizio E, Rodriguez C,
Chuchana P, Fauquier L, Cheng D, Theillet C, Vandel L, Bedford MT
and Sardet C: Coactivator-associated arginine methyltransferase 1
(CARM1) is a positive regulator of the Cyclin E1 gene. Proc
Natl Acad Sci USA. 103:13351–13356. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Thomassen M, Tan Q and Kruse TA: Gene
expression meta-analysis identifies chromosomal regions and
candidate genes involved in breast cancer metastasis. Breast Cancer
Res Treat. 113:239–249. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Baldwin RM, Morettin A, Paris G, Goulet I
and Côté J: Alternatively spliced protein arginine
methyltransferase 1 isoform PRMT1v2 promotes the survival and
invasiveness of breast cancer cells. Cell Cycle. 11:4597–4612.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zhong J, Cao RX, Hong T, Yang J, Zu XY,
Xiao XH, Liu JH and Wen GB: Identification and expression analysis
of a novel transcript of the human PRMT2 gene resulted from
alternative polyadenylation in breast cancer. Gene. 487:1–9. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ohkura N, Takahashi M, Yaguchi H, Nagamura
Y and Tsukada T: Coactivator-associated arginine methyltransferase
1, CARM1, affects pre-mRNA splicing in an isoform-specific manner.
J Biol Chem. 280:28927–28935. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Gros L, Renodon-Cornière A, de Saint
Vincent BR, Feder M, Bujnicki JM and Jacquemin-Sablon A:
Characterization of prmt7alpha and beta isozymes from Chinese
hamster cells sensitive and resistant to topoisomerase II
inhibitors. Biochim Biophys Acta. 1760:1646–1656. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yoshimoto T, Boehm M, Olive M, Crook MF,
San H, Langenickel T and Nabel EG: The arginine methyltransferase
PRMT2 binds RB and regulates E2F function. Exp Cell Res.
312:2040–2053. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Meyer R, Wolf SS and Obendorf M: PRMT2, a
member of the protein arginine methyltransferase family, is a
coactivator of the androgen receptor. J Steroid Biochem Mol Biol.
107:1–14. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Tashiro E, Tsuchiya A and Imoto M:
Functions of cyclin D1 as an oncogene and regulation of cyclin D1
expression. Cancer Sci. 98:629–635. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Kim JK and Diehl JA: Nuclear cyclin D1: An
oncogenic driver in human cancer. J Cell Physiol. 220:292–296.
2009. View Article : Google Scholar : PubMed/NCBI
|