Introduction
Chronic myelogenous leukemia (CML) is a lethal
malignancy. The increased and unregulated growth of predominantly
myeloid cells in the bone marrow is the main characteristic of CML.
CML has a morbidity of 1–2 cases per 100,000 adults and is
responsible for ~15% of all newly diagnosed cases of leukemia in
adults (1). The majority of
patients with CML fail to respond effectively to the current
regimen of drug therapy due to occurrence of drug resistance. Bone
marrow or allogenic stem cell transplantations are recent and
effective treatments for CML, but they have a high risk of
morbidity and mortality (2). CML
places considerable burden on patients and the majority of
chemotherapeutic drugs have long-term side effects (3,4).
Therefore, it is important to continue research into novel
therapeutic approaches for CML. With regard to the induction of
apoptosis, it is generally believed that drugs cause the
elimination of cancer cells. Apoptosis is a type of cell death used
by multicellular organisms to prevent uncontrolled proliferation
and to dispose of unwanted cells. Apoptosis is originally
characterized by morphological changes, including membrane
blebbing, DNA fragmentation, chromatin condensation, nuclear
fragmentation, cell shrinkage and apoptotic body formation
(5). Resistance to apoptosis is a
reason for concern in developing effective chemotherapeutic agents
in various types of cancer. In addition, differentiation-inducing
therapy has been proven to be a promising strategy. To date,
all-trans retinoic acid-based therapy of acute promyelocytic
leukemia (APL) has been the best clinical application of
differentiation therapy (6,7). However, the development of
differentiation therapy for CML requires further investigation.
Lidamycin (LDM) is a macromolecular enediyne
antitumor antibiotic (8). LDM has
two parts: one is an enediyne chromophore (MW 843 Da) responsible
for the extremely potent bioactivity of the antibiotic and the
other is a non-covalently bound apoprotein (MW 10.5 kDa) (9,10). The
two parts can be dissociated and reconstituted, and the biological
activity of the rebuilt molecule is comparable to that of natural
LDM (11,12). LDM can induce DNA damage (13). LDM is extremely cytotoxic and
induces growth inhibition of transplantable tumors in mice
(14–17). Furthermore, LDM exhibited a strong
inhibitory effect on tumor metastasis and angiogenesis (18). At present, several clinical trials
are in progress to assess the therapeutic efficacy of LDM in
multiple cancer indications (19).
To determine the function of LDM on CML, the present study
investigated the effects of LDM on the K562 cell line as an
experimental model for CML. The present study revealed that LDM was
highly active in inhibiting cell growth and that a low
concentration LDM could induce cell apoptosis by upregulating the
expression of caspase-8 and caspase-3. Furthermore, a low
concentration of LDM increases the cell hemoglobin contents and
induces erythroid differentiation of K562 cells by upregulating the
expression of GATA-1.
Materials and methods
Cell culture
The human K562 cells were cultured in RPMI-1640
medium (Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA)
supplemented with 10% fetal bovine serum (FBS; Gibco; Thermo Fisher
Scientific, Inc.), penicillin (10 U/ml) and streptomycin (100 U/ml)
at 37°C. The cells were accumulated in the metaphase in the
following experiments.
Cell proliferation assay
A cell proliferation assay was performed using a
Cell Counting Kit-8 (CCK-8; Beijing Zoman Biotechnology Co., Ltd.,
Beijing, China). In brief, the K562 cells were plated into 96-well
plates at a density of 5,000 cells/well and incubated at 37°C
overnight. Experimental cells were treated with various
concentrations (10−13 to 10−6 M) of LDM for
48 h at 37°C. Next, 10 µl of CCK-8 reagent was added into each
well. Cells were incubated for 2 h at 37°C and optical density was
measured at 450 nm. The aforementioned experiments were performed
in triplicate.
Morphological analysis of apoptotic
cells
The K562 cells were seeded into 12-well plates at
5×104 cells/well and exposed to LDM at concentrations of
0.01, 0.1 and 1 nM for 48 h. The morphology of apoptosis was
evaluated by acridine orange/ethidium bromide (AO/EB) staining
using a fluorescence microscope (magnification, ×200) (BH2 system;
Olympus Corp., Tokyo, Japan). Cells were washed with cold
phosphate-buffered saline (PBS) and adjusted to a cell density of
5×105 cells/well. A total of 10 µl cells was placed on a
glass slide and mixed with AO/EB solution (1:1, v/v) to a final
concentration of 100 µg/ml. Each sample should be mixed just prior
to microscopy. Acridine orange is a vital dye and will stain both
live and dead cells. Ethidium bromide will stain only cells that
have lost membrane integrity. Live cells will appear uniformly
green. Early apoptotic cells will stain green and contain bright
green dots in the nuclei as a consequence of chromatin condensation
and nuclear fragmentation. Late apoptotic cells will also
incorporate ethidium bromide and therefore stain orange, but, in
contrast to necrotic cells, the late apoptotic cells will show
condensed and often fragmented nuclei. Necrotic cells stain orange,
but have a nuclear morphology resembling that of viable cells, with
no condensed chromatin (5).
In order to distinguish the apoptosis induction and
look for the more effective drug dose, we used a lower
concentration of LDM (0.001 nM) and a higher concentration of LDM
(10 nM) in the following experiments.
Nitro blue tetrazolium (NBT) reduction
experiment
The K562 cells were seeded into 12-well plates at
5×104 cells/well and exposed to LDM for 48 or 72 h.
Next, cells were harvested and washed with PBS for 3 times. After
100 µl 0.2% NBT solution containing TPA (0.2 mg/ml) was added,
cells were incubated in the dark at 37°C for 30 min and 1 ml cold
PBS was added to terminate the reaction. The cells were then placed
onto a glass slide and observed under a light microscope
(magnification, ×100). Three visual fields (each containing 200
cells) were randomly selected, and the percentage of blue punctate
particle-positively stained cells was calculated using Mshot Image
Analysis system (Micro-shot Technology Co., Ltd., Guangzhou,
China).
Morphological analysis of K562
cells
The K562 cells were seeded into 12-well plates at
5×104 cells/well and exposed to LDM for 48 or 72 h.
Morphological changes in the cells were evaluated by Wright-Giemsa
staining using a fluorescence microscope (BH2 system; Olympus
Corp.). Cells were washed with cold PBS and adjusted to a cell
density of 5×105 cells/well. A total of 10 µl cells was
placed onto a glass slide, mixed with Giemsa and observed under a
microscope.
Detection of hemoglobin contents
The K562 cells were seeded onto 6-well plates at 2
ml/well and exposed to LDM for 48 or 72 h. All the cells were
harvested and lysed, and the concentration of hemoglobin was
determined using a spectrophotometer. The aforementioned
experiments were performed in triplicate.
CD71 expression assay
The expression of CD71 on the surface of K562 cells
was assessed using a flow cytometer. The K562 cells were seeded
into 6-well plates at 2 ml/well and exposed to LDM for 48 or 72 h.
Cells were then harvested and the cell count was adjusted to
1×105 cells/ml. Following washing with PBS, cells were
labeled with 20 µl PE-conjugated anti-CD71 (BD Biosciences, San
Jose, CA, USA) for 15 min in the dark at room temperature. Cells
were washed twice with PBS and resuspended in 500 µl PBS prior to
measurement of CD71 expression. Fluorescence intensity was measured
using FACSCanto II (BD Biosciences) in continuous mode (20).
Western blot analysis
The K562 cells were treated with LDM and harvested
after 48 or 72 h. Harvested cells were washed with PBS and then
lysed using RIPA buffer [25 mM Tris (pH 7.8), 2 mM EDTA, 20%
glycerol, 0.1% Nonidet P-40 (NP-40), 1 mM dithiothreitol] and
protease inhibitors. The protein concentrations of cell
supernatants were determined with the BCA Protein Assay kit from
Hyclone-Pierce. Proteins (30 µg/lane) were equally loaded to and
separated by SDS-PAGE gel (15%). After electrophoresis, proteins
were transferred to polyvinylidene difluoride (PVDF) membrane (EMD
Millipore, Bedford, MA, USA). Blots were blocked for 60 min at room
temperature with 5% non-fat milk powder and 0.1% Tween-20 in PBS
and exposed overnight at 4°C using primary antibodies against
caspase-8 (sc-56071; Santa Cruz Biotechnology, Inc., Dallas, TX,
USA), active caspase-8 (cat. no. 9748; Cell Signaling Technology,
Inc., Danvers, MA, USA), caspase-3 (sc-56052; Santa Cruz
Biotechnology, Inc.), active caspase-3 (cat. no. 9661; Cell
Signaling Technology, Inc.), GATA-1 (sc-266) and β-actin
(sc-130065) (both from Santa Cruz Biotechnology, Inc.). All the
primary antibodies were diluted at 1:500. The membranes were
incubated in horseradish peroxidase-labeled IgG secondary
antibodies (1:1,000; sc-2380 and sc-2379; Santa Cruz Biotechnology,
Inc.) at room temperature for 1 h. The bands were visualized using
enhanced chemiluminescence detection reagents (Santa Cruz
Biotechnology, Inc.). Western blots were quantified via
densitometry scanning using NIH Image software version 1.46
(National Institutes of Health, Bethesda, MD, USA).
Statistical analysis
Results are expressed as the mean ± standard
deviation (SD) of three independent experiments. Treatment effects
were analyzed using the unpaired Student's t-test when comparing
two variables, while inter-group and intra-group comparisons were
conducted using one-way ANOVA test with post hoc contrasts by
Student-Newman-Keuls test. A P-value of ≤0.05 was considered to
indicate a statistically significant difference.
Results
Effects of LDM on the proliferation of
K562 cells
Inhibition of cell proliferation was measured using
CCK-8 assays. K562 cells were treated with different concentrations
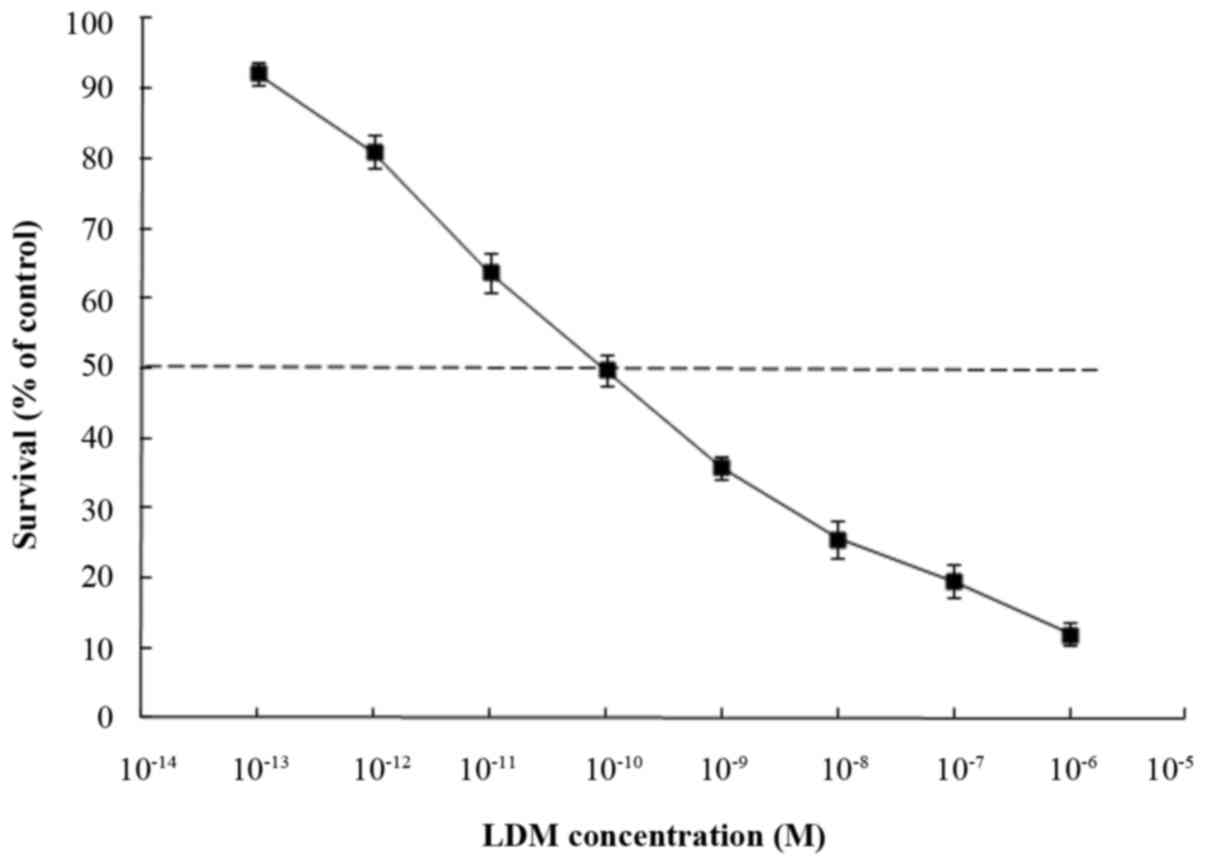
of LDM for 48 h. CCK-8 assays revealed that cell viability was
reduced following treatment with LDM and the IC50 value
was 0.1±3.2 nM which was calculated by GraphPad Prism 7 software
(GraphPad Software, Inc., La Jolla, CA, USA (Fig. 1). Since LDM has extreme cytotoxicity
to many cancer cells as well as normal cells, in order to reduce
its toxicity, we explored the effects of low dose LDM. In our
experiment, we chose 0.01 or 0.001 nM as the low concentration.
Induction of apoptosis by LDM in K562
cells
AO/EB staining was used to detect the apoptosis of
K562 cells. AO can inset the DNA of whole cells and show green
fluorescence, while EB only penetrates the impaired cell membrane
and shows orange red fluorescence. Fluorescence increases in an
apoptotic cell. Under a fluorescence microscope, the nuclei of the
control cells were large and round without condensation or
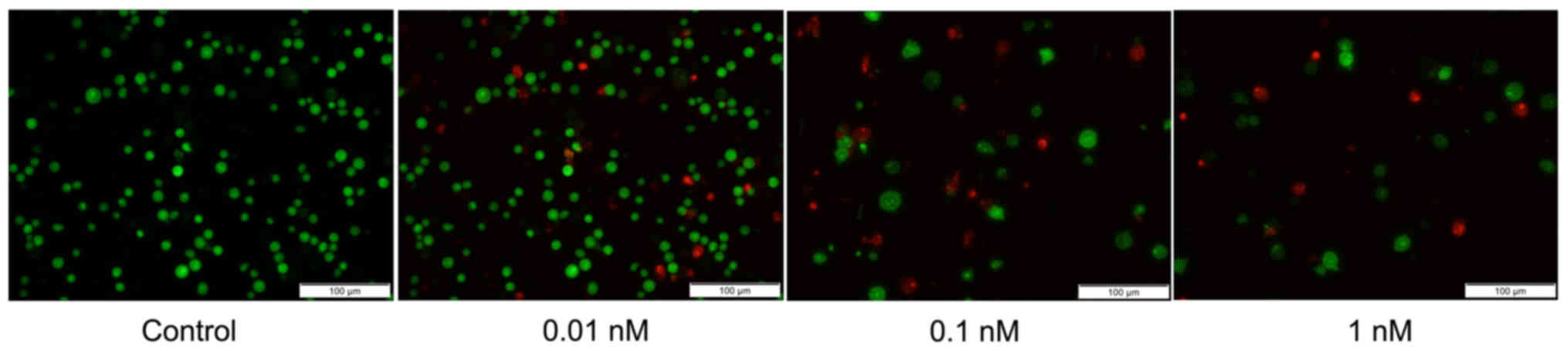
fragmentation. By contrast, following exposure to LDM for 48 h, the
majority of cells presented with typical morphological changes of
apoptosis, including chromatin condensation or shrunken nuclei
(Fig. 2). In the group treated with
higher concentrations of LDM, condensed nuclei were observed. The
live cells appeared green in color with undamaged nuclei, while the
early apoptotic cells exhibited condensed nuclei and were green in
color with bright green dots in their nuclei, and the late
apoptotic cells were orange in color. In addition, the number of
cells also decreased in the 0.1 and 1 nM LDM groups,
respectively.
In addition, flow cytometry (FACS) analysis was
carried out to determine the percentage of apoptosis.
Unfortunately, the results were unsatisfactory and this will be
repeated in future studies.
Effects of LDM on the activation of
apoptosis-related proteins in K562 cells
Apoptosis is induced by the activation of a group of
cysteine proteases called ‘caspases’, which cleave proteins during
cell death. According to the function of caspases, they are grouped
into two different subfamilies. One subfamily includes the
initiation (caspase-8 and −9) and execution (caspase-3, −6 and −7)
of the apoptotic program (initiator and executioner caspases).
Caspase-8 and caspase-9 are caspases upstream of the apoptosis
signal transduction process, while caspase-3 is downstream, all of
which are effector molecules of cell apoptosis. The activation of
initiator caspase-8 will, in turn, activate caspase-3. Caspase-3 is
an executioner caspase in the last and irreversible phase of the
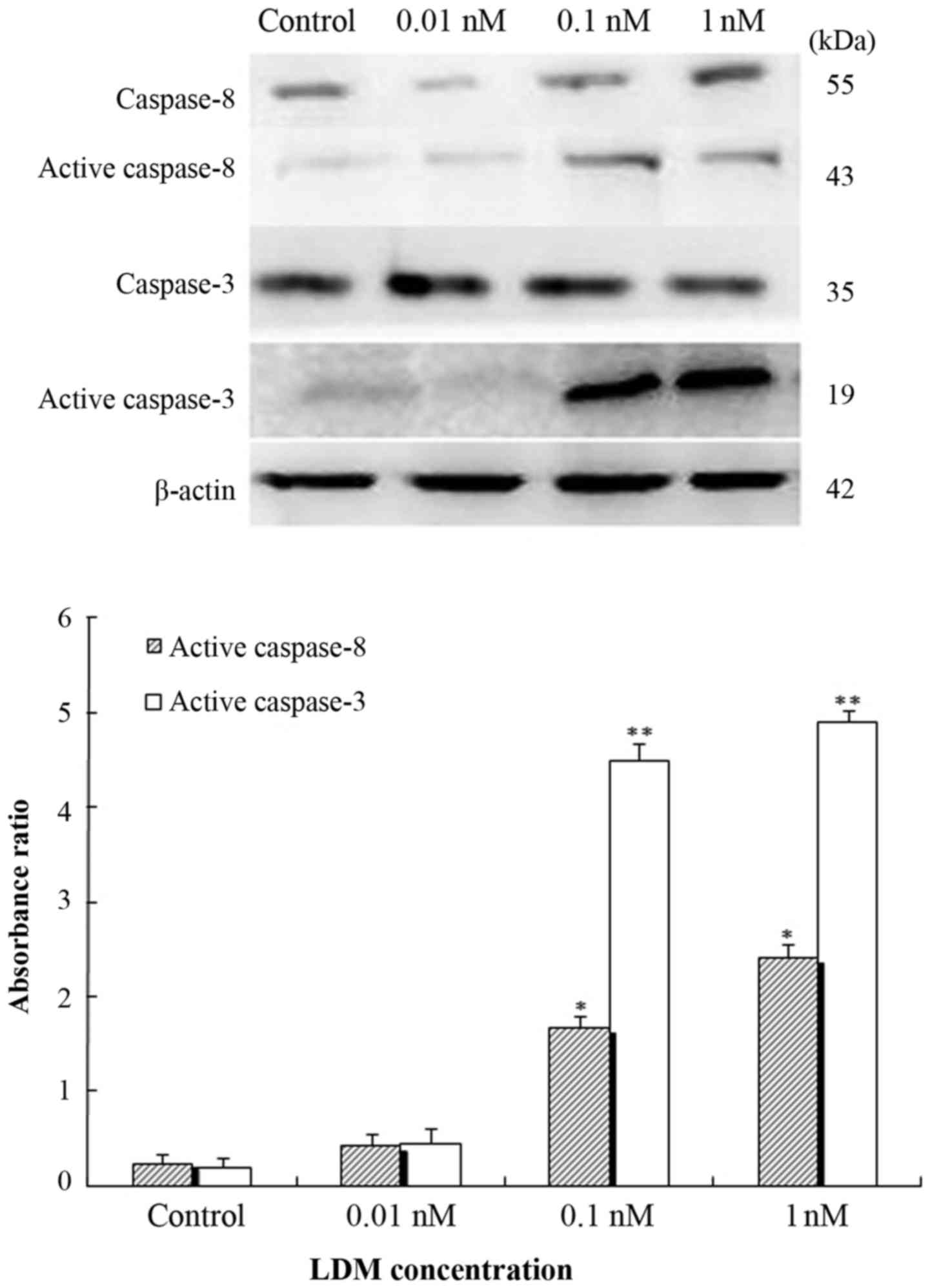
apoptotic caspase-dependent pathway. The expression of caspase-8
and caspase-3 in K562 cells treated with 0.01, 0.1 and 1 nM LDM was
investigated by western blot analysis. As shown in Fig. 3, treatment with LDM for 48 h led to
upregulation and activation of caspase-8 and caspase-3 in K562
cells compared to controls.
Effect of LDM at low concentrations on
the differentiation induction of K562 cells
To assess whether a low concentration of LDM could
induce differentiation of K562 cells, the NBT reduction experiment
was performed. Differentiated K562 cells can reduce NBT to dark
blue diformazan particles, which can easily be observed under a
light microscope. K562 cells were treated with 0.001, 0.1 and 10 nM
LDM for 48 or 72 h. The NBT-positive percentages of the control
group were 7.6±1.5 and 10.3±1.7% at 48 and 72 h, respectively,
while the NBT-positive percentages of K562 cells were 45.7±4.2,
52.5±2.7 and 28.4±5.6% in the 0.001, 0.1 and 10 nM groups,
respectively, at 48 h; and 59.4±3.1, 73.1±1.9 and 32.8±2.8% in the
0.001, 0.1 and 10 nM groups, respectively, at 72 h. To note, most
cells died in the 10 nM group, thus the NBT reduction rate was
decreased (Fig. 4).
Effect of LDM at low concentrations on
the induction of erythroid differentiation of K562 cells
Morphological changes of K562
cells
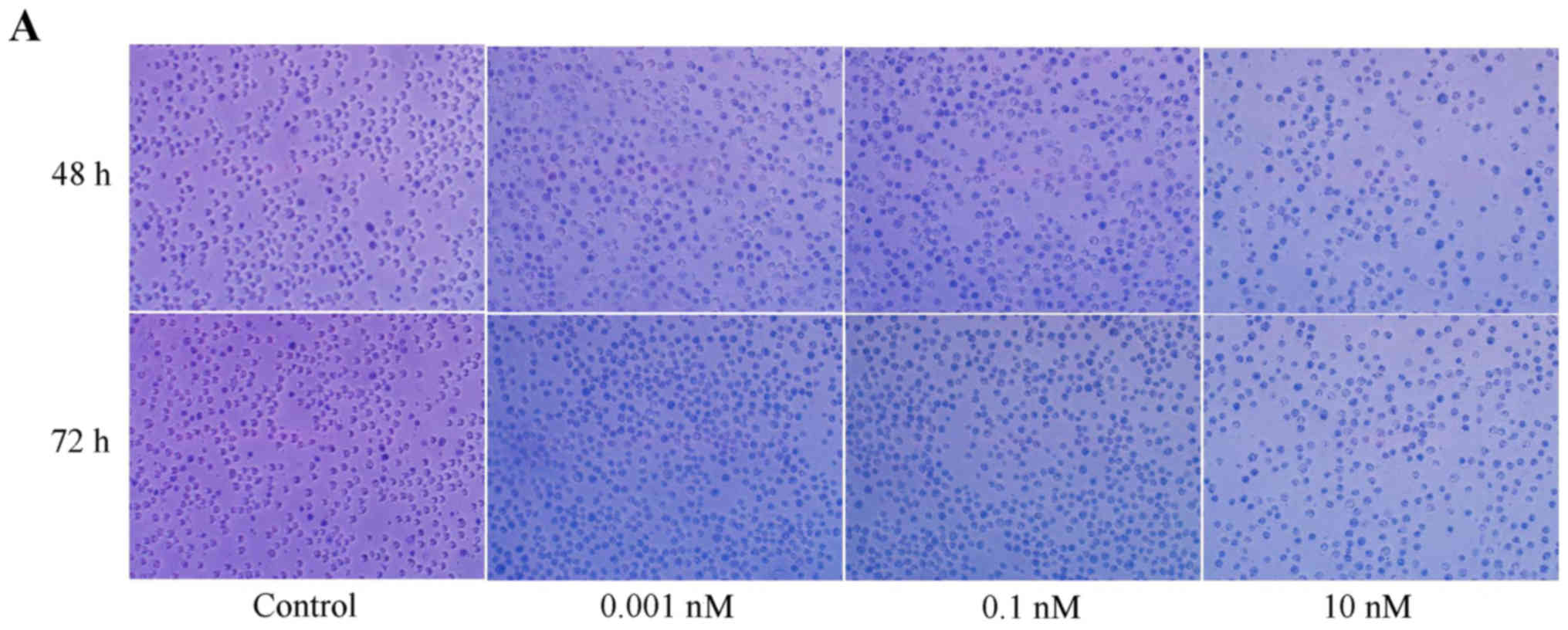
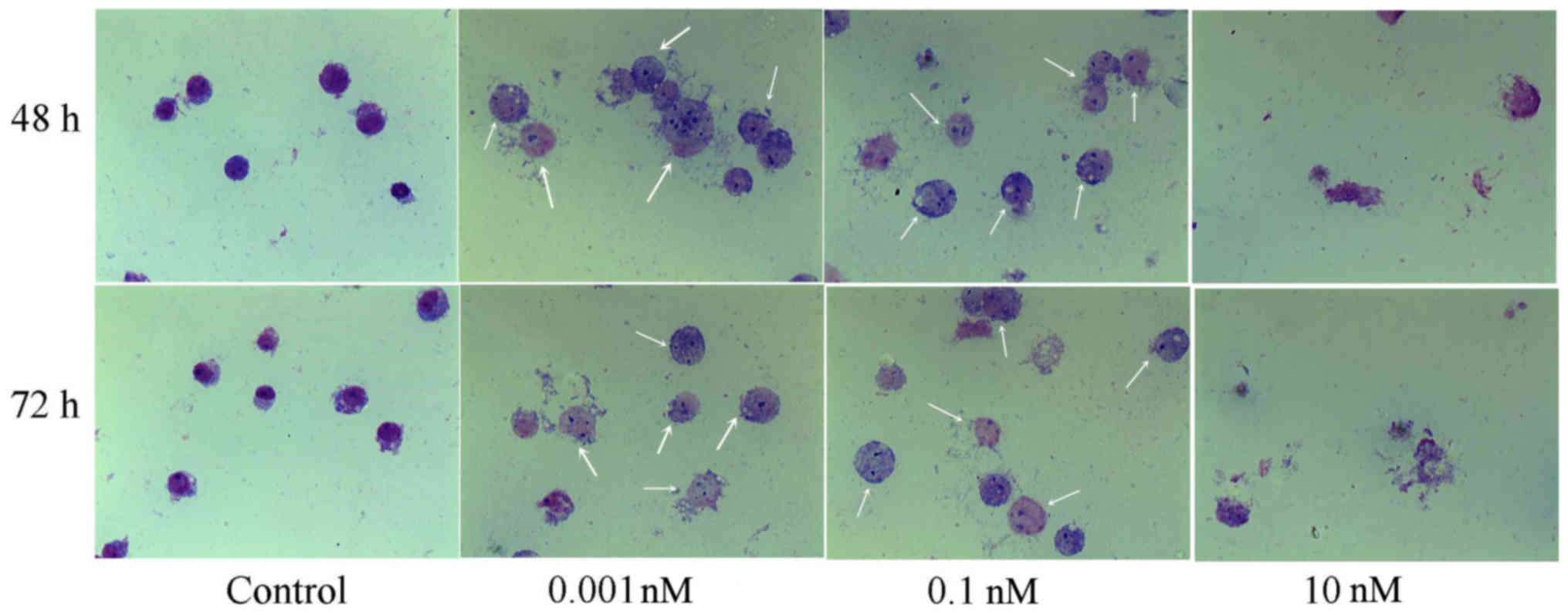
Wright-Giemsa staining revealed that a low
concentration of LDM induced distinct morphological changes in K562
cells. In control groups, K562 cells showed blue purple staining.
While some cells showed pale pink in the 0.001 and 0.1 nM LDM
groups. This reflected some changes in cellular components. In
addition, the cell volume also changed considerably. Most cells
died in the 10 nM LDM group. The cytoplasm of K562 cells following
treatment with 0.1 and 0.001 nM LDM for 48 or 72 h became more
ample than that of the control group cells. These typical
morphological changes showed a differentiation tendency for K562
cells (Fig. 5).
LDM increases the hemoglobin contents
of K562 cells
The results of the hemoglobin content assay
demonstrated that the level of hemoglobin in K562 cells was
increased following treatment with LDM for 48 or 72 h. In the 0.001
and 0.1 nM LDM groups, the hemoglobin contents of K562 cells were
markedly increased, compared with those in the control group.
Additionally, most cells died in the 10 nM group, thus there was a
decrease in the hemoglobin content (Table I).
 | Table I.Influence of LDM on the hemoglobin
content of K562 cells. |
Table I.
Influence of LDM on the hemoglobin
content of K562 cells.
|
|
| Hemoglobin content
(µg/l) |
|---|
|
|
|
|
|---|
| Concentration of LDM
(nmol/l) | n | 48 h | 72 h |
|---|
| 0 | 3 | 0.273±0.024 | 0.341±0.052 |
| 0.001 | 3 |
0.703±0.204a |
0.827±0.174a |
| 0.1 | 3 |
0.833±0.314a |
1.157±0.162b |
| 10 | 3 |
0.562±0.116a |
0.614±0.332a |
LDM increases the expression level of
CD71 in K562 cells
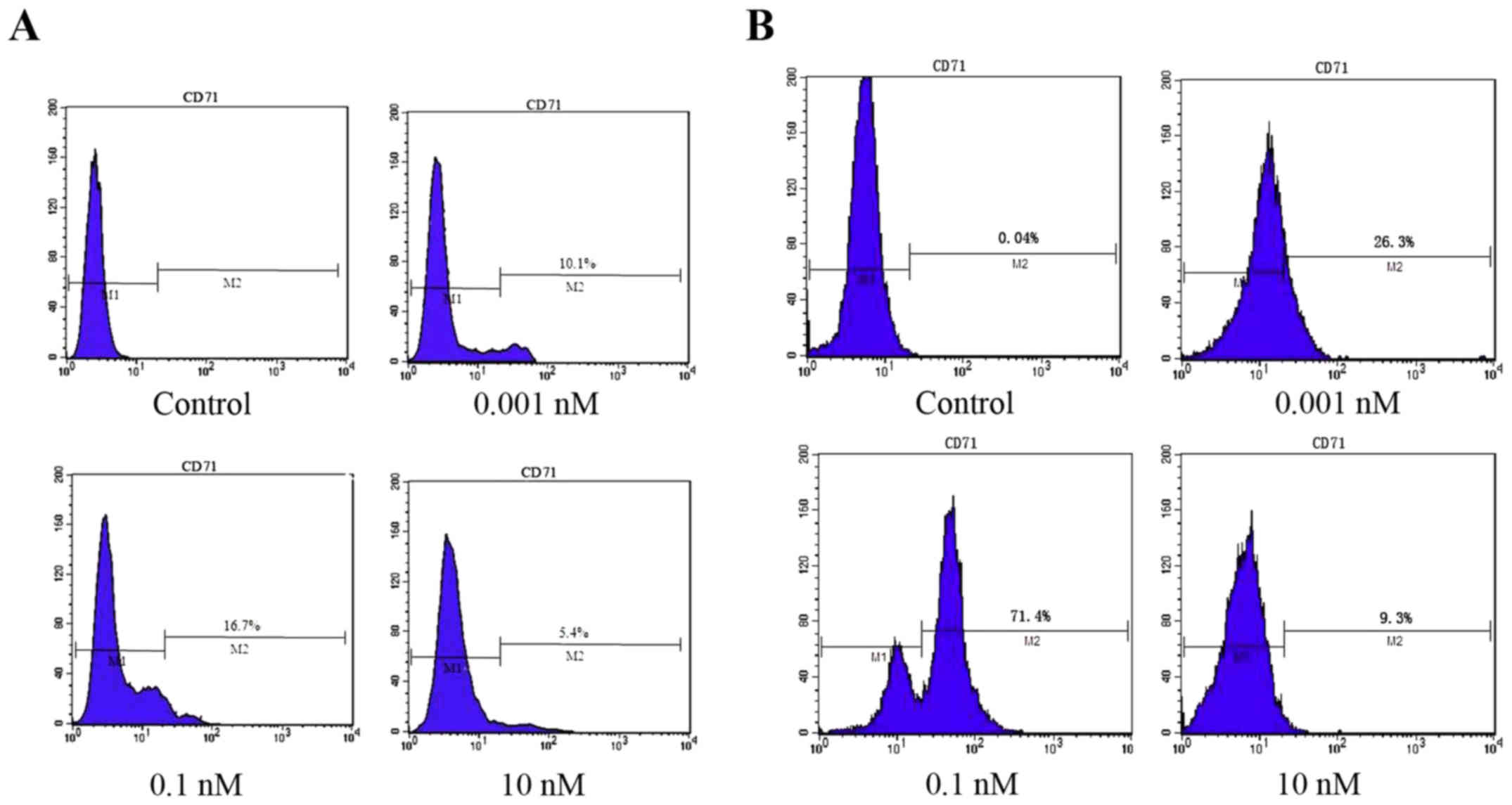
The CD71 antigen is a classical erythroid
differentiation marker expressed on the cell surface. Flow
cytometric analysis demonstrated that LDM increased the expression
level of CD71 in K562 cells. Compared with the control cells, the
expression of CD71 was 10.1, 16.7 and 5.4% in the 0.001, 0.1 and 10
nM groups, respectively, at 48 h; and 26.3, 71.4 and 9.3% in the
0.001, 0.1 and 10 nM groups, respectively, at 72 h. The expression
of CD71 in the 0.1 nM LDM-treated K562 cells was significantly
higher than that in the other group cells (Fig. 6). These results clearly demonstrated
that low concentration LDM could induce the differentiation of K562
cells into erythroid lineage.
Activation of GATA-1 is associated
with the erythroid-inducing differentiation mechanism in K562 cells
by LDM at a low concentration
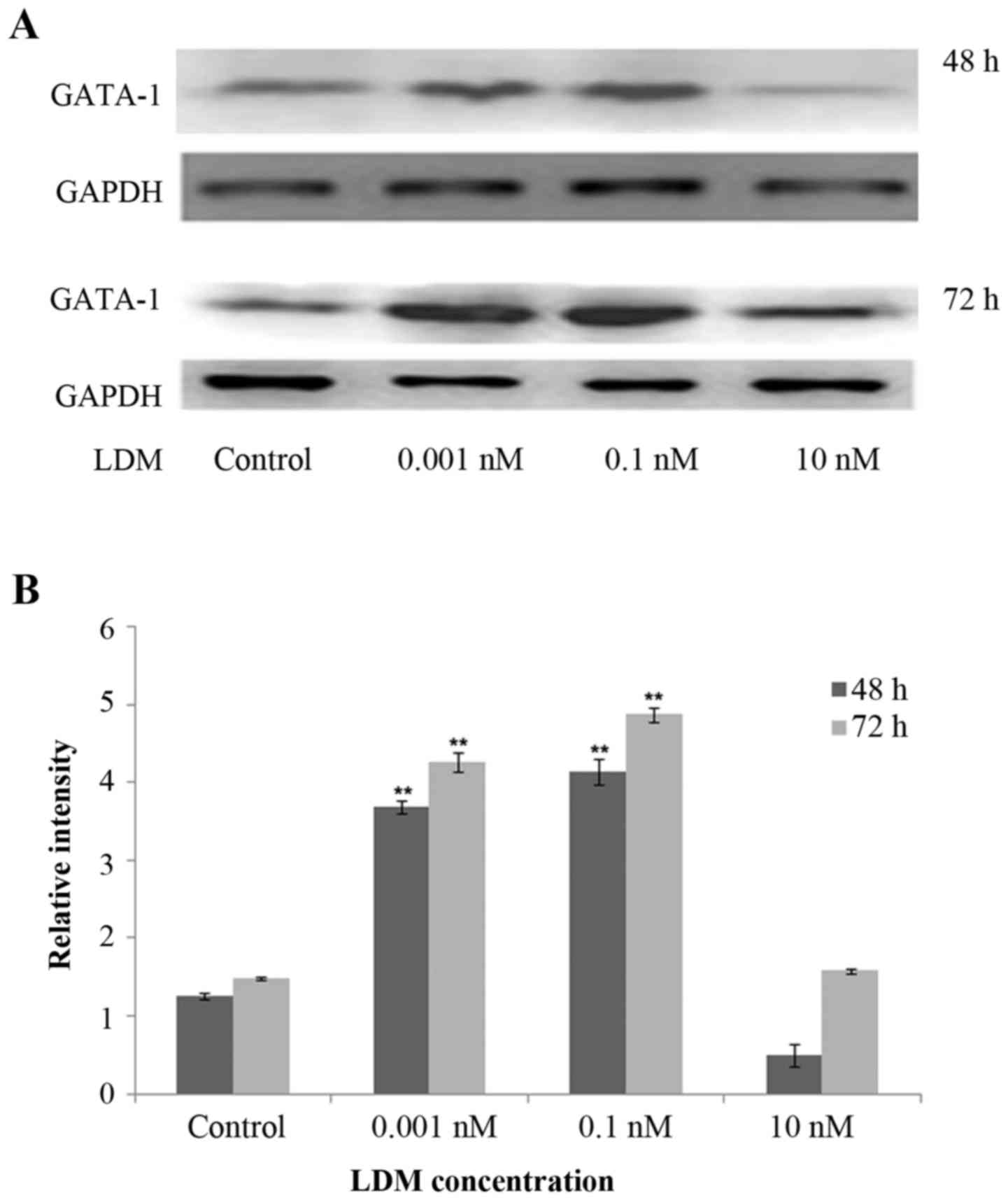
GATA-1 is a member of the GATA transcription factor
family and is a key mediator of the development of specific types
of blood cells from their precursor cells. Based on the
erythroid-inducing differentiation effects of LDM on K562 cells and
the significant role of GATA-1 in erythropoiesis, GATA-1 protein
expression was detected by western blot analysis. The results
demonstrated that following treatment with LDM for 48 and 72 h, the
GATA-1 protein expression in the K562 cells was increased compared
with that in the control cells, while it was decreased in the 10 nM
group as most of the cells died (Fig.
7).
Discussion
Chronic myelogenous leukemia (CML) is a lethal
hematological disorder originating from a small number of leukemia
stem cells. The properties of CML cells are as follows:
uncontrolled growth, escape of apoptosis and differentiation
disorder. Traditional chemotherapeutic agents inevitably act on
healthy tissue. Therefore, induction of apoptosis or terminal
differentiation is an attractive approach for the therapy of human
leukemia. Lidamycin (LDM) is an antitumor antibiotic that is
produced by a streptomyces globisporus C1027, which was
isolated in China. LDM displays extreme cytotoxic and
anti-angiogenic activity, as well as distinct growth inhibition
against transplantable tumors in mice (16). The cytotoxicity of LDM is more
powerful than that of mitomycin or doxorubicin (21). Currently, several phase II and phase
III clinical trials are underway to evaluate the therapeutic
efficacy of LDM in breast, non-small cell lung, colon cancer and
lymphoma (19). The present study
investigated the anticancer effect of LDM on the K562 cell line as
an experimental model for CML. In the present study, it was
observed that LDM decreased the viability of K562 cells in a
dose-dependent manner and the IC50 value of lidamycin
was 0.1±3.2 nM. Since the inhibition of apoptosis is a hallmark of
cancer, induction of apoptosis in cancer cells is known to be an
efficient strategy for the treatment of cancer. In the present
study, the induction of apoptosis was investigated morphologically
by AO/EB staining. The results revealed that LDM at low
concentrations could induce the apoptosis of K562 cells. The most
important factor in the mechanism of apoptosis is the activation of
caspases. Caspases are proteases that participate in the control of
cell growth and apoptosis (22).
Caspase-8 and caspase-9 are caspases upstream of the apoptosis
signal transduction process, while caspase-3 is downstream, and all
of these are effector molecules of cell apoptosis (23). The present study analyzed the
protein expression level of caspase-8 and caspase-3 by western blot
analysis. The results demonstrated that a low concentration of LDM
could increase the expression of caspase-8 and caspase-3 in K562
cells. Furthermore, the fact that LDM at a low concentration
increased NBT reduction capacity indicated that it could induce the
differentiation of K562 cells. Subsequently, morphological changes,
including decreased nuclear to cytoplasm ratio following LDM
treatment, were observed under a light microscope. The hemoglobin
concentration in LDM-treated K562 cells was further evidenced by a
colorimetry assay. Flow cytometric analysis revealed that LDM could
increase the expression level of CD71 in K562 cells. These
phenotypic changes are frequently taken as indications for cellular
erythroid differentiation.
Malignant hematopoiesis is characterized by a lack
of differentiation capacity. Cell differentiation is a process of
gene selective expression. The presence of hemoglobin in a cell
indicates that the cell gene is selectively expressed, indicating
that the cell has differentiated. The effect of cell
differentiation is often associated with the dysfunction of the
transcription factors in the signal transduction pathway.
GATA-binding factor 1 (GATA-1), NF-E2, T-cell acute lymphocytic
leukemia protein 1 (TAL1) and Kruppel like factor 1 (KLF1) are
major erythroid-specific transcription activators that bind to the
β-globin locus and regulate transcription of the globin gene
(24). GATA-1 is a major regulator
of the expression of other erythroid-specific activators in human
erythroid K562 cells. During red blood cell maturation, GATA-1
activates nearly all erythroid-specific genes, while silencing
genes associated with the immature proliferative red blood cell
precursor cells (25,26). Hemoglobin is a specific protein. The
presence of hemoglobin indicates that cells have differentiated
into red blood cells. In order to investigate the mechanism of
erythroid differentiation of LDM on K562 cells, the present study
detected the hemoglobin contents and the GATA-1 protein expression
in LDM-treated K562 cells. The results revealed that LDM at a low
concentration could increase the hemoglobin contents and upregulate
the GATA-1 protein expression in K562 cells.
In conclusion, a major finding of the present study
was that low concentrations of lidamycin could upregulate and
activate the caspase-8 and caspase-3 signaling pathway, which in
turn may have contributed toward the induction of apoptosis and
cancer cell growth inhibition. On the other hand, LDM at low
concentrations upregulated the GATA-1 protein expression of K562
cells, which in turn may have contributed toward the induction of
K562 cell erythroid differentiation. These results indicate that
lidamycin may serve a positive role in challenging the
differentiation-inducing therapy of chronic myelogenous
leukemia.
Acknowledgements
Not applicable.
Funding
The present study was supported by the National
Science Foundation of Hebei Province (no. H2013209040) and the
Chinese Academy of Medical Sciences (CAMS) Innovation Fund for
Medical Sciences (no. 2017-I2M-1-010).
Availability of data and materials
The datasets used during the present study are
available from the corresponding author upon reasonable
request.
Authors' contributions
JC and JHG conceived and designed the study. CZ and
LYG performed the experiments and prepared the manuscript. DM
performed the image and data analysis. JC and JHG revised the
manuscript. All authors read and approved the manuscript and agree
to be accountable for all aspects of the research in ensuring that
the accuracy or integrity of any part of the work are appropriately
investigated and resolved.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests
References
|
1
|
Jabbour E and Kantarjian H: Chronic
myeloid leukemia: 2014 update on diagnosis, monitoring, and
management. Am J Hematol. 89:547–556. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Dickinson AM, Pearce KF, Norden J, O'Brien
SG, Holler E, Bickeböller H, Balavarca Y, Rocha V, Kolb HJ,
Hromadnikova I, et al: Impact of genomic risk factors on outcome
after hematopoietic stem cell transplantation for patients with
chronic myeloid leukemia. Haematologica. 95:922–927. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Rousselot P, Charbonnier A, Cony-Makhoul
P, Agape P, Nicolini FE, Varet B, Gardembas M, Etienne G, Réa D,
Roy L, et al: Loss of major molecular response as a trigger for
restarting tyrosine kinase inhibitor therapy in patients with
chronic-phase chronic myelogenous leukemia who have stopped
imatinib after durable undetectable disease. J Clin Oncol.
32:424–430. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Song P, Ye L, Fan J, Li Y, Zeng X, Wang Z,
Wang S, Zhang G, Yang P, Cao Z, et al: Asparaginase induces
apoptosis and cytoprotective autophagy in chronic myeloid leukemia
cells. Oncotarget. 6:3861–3873. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Rahimi R, Mahdavi M, Pejman S, Zare P and
Balalaei S: Inhibition of cell proliferation and induction of
apoptosis in K562 human leukemia cells by the derivative (3-NpC)
from dihydro-pyranochromenes family. Acta Biochim Pol. 62:83–88.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Marchwicka A, Cebrat M, Sampath P,
Snieżewski L and Marcinkowska E: Perspectives of differentiation
therapies of acute myeloid leukemia: The search for the molecular
basis of patients' variable responses to 1,25-dihydroxyvitamin d
and vitamin d analogs. Front Oncol. 4:125–137. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Takahashi H, Hatta Y, Iriyama N, Hasegawa
Y, Uchida H, Nakagawa M, Makishima M, Takeuchi J and Takei M:
Induced differentiation of human myeloid leukemia cells into M2
acrophages by combined treatment with retinoic acid and
1α,25-dihydroxyvitamin D3. PLoS One. 9:e1137222014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hu JL, Xue YC, Xie MY, Zhang R, Otani T,
Minami Y, Yamada Y and Marunaka T: A new macromolecular antitumor
antibiotic, C-1027. I. Discovery, taxonomy of producing organism,
fermentation and biological activity. J Antibiot (Tokyo).
41:1575–1579. 1988. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sakata N, Ikeno S, Hori M, Hamada M and
Otani T: Cloning and nucleotide sequencing of the antitumor
antibiotic C-1027 apoprotein gene. Biosci Biotechnol Biochem.
56:1592–1595. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Tanaka T, Fukuda-Ishisaka S, Hirama M and
Otani T: Solution structures of C-1027 apoprotein and its complex
with the aromatized chromophore. J Mol Biol. 309:267–283. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Shao RG and Zhen YS: Relationship between
the molecular composition of C1027, a new macromolecular antibiotic
with enediyne chromophore, and its antitumor activity. Yao Xue Xue
Bao. 30:336–342. 1995.(In Chinese). PubMed/NCBI
|
|
12
|
Jiang W, Shang B, Li L, Zhang S and Zhen
Y: Construction of a genetically engineered chimeric apoprotein
consisting of sequences derived from lidamycin and
neocarzinostatin. Anticancer Drugs. 27:24–28. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Dziegielewski J and Beerman TA: Cellular
responses to the DNA strand-scission enediyne C-1027 can be
independent of ATM, ATR, and DNA-PK kinases. J Biol Chem.
277:20549–20554. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhen YS, Ming XY, Yu B, Otani T, Saito H
and Yamada Y: A new macromolecular antitumor antibiotic, C-1027.
III. Antitumor activity. J Antibiot (Tokyo). 42:1294–1298. 1989.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Xu YJ, Zhen YS and Goldberg IH: C1027
chromophore, a potent new enediyne antitumor antibiotic, induces
sequence-specific double-strand DNA cleavage. Biochemistry.
33:5947–5954. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhen H, Xue Y and Zhen Y: Inhibition of
angiogenesis by antitumor antibiotic C1027 and its effect on tumor
metastasis. Zhonghua Yi Xue Za Zhi. 77:657–660. 1997.(In Chinese).
PubMed/NCBI
|
|
17
|
Huang YH, Shang BY and Zhen YS: Antitumor
efficacy of lidamycin on hepatoma and active moiety of its
molecule. World J Gastroenterol. 11:3980–3984. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ding LL, Liu M, Zhang SH, Zhao XZ, Wu N,
Chen L, Wang GJ and Lin XK: Lidamycin inhibits angiogenesis of
zebrafish embryo via down-regulation of VEGF. Yao Xue Xue Bao.
45:456–461. 2010.(In Chinese). PubMed/NCBI
|
|
19
|
Shao RG and Zhen YS: Enediyne anticancer
antibiotic lidamycin: Chemistry, biology and pharmacology.
Anticancer Agents Med Chem. 8:123–131. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Suzuki M, Tanaka H, Tanimura A, Tanabe K,
Oe N, Rai S, Kon S, Fukumoto M, Takei K, Abe T, et al: The clathrin
assembly protein PICALM is required for erythroid maturation and
transferrin internalization in mice. PLoS One. 7:e318542012.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Xin C, Ye S, Ming Y, Shenghua Z, Qingfang
M, Hongxing G, Xu S, Yuanfu X, Yuan Z, Dongmei F, et al: Efficient
inhibition of B-cell lymphoma xenografts with a novel recombinant
fusion protein: anti-CD20Fab-LDM. Gene Ther. 17:1234–1243. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Morgan CW, Julien O, Unger EK, Shah NM and
Wells JA: Turning on caspases with genetics and small molecules.
Methods Enzymol. 544:179–213. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhao Y, Lei M, Wang Z, Qiao G, Yang T and
Zhang J: TC R-induced, PKC-θ-mediated NF-κB activation is regulated
by a caspase-8-caspase-9-caspase-3 cascade. Biochem Biophys Res
Commun. 450:526–531. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kang Y, Kim YW, Yun J, Shin J and Kim A:
KLF1 stabilizes GATA-1 and TAL1 occupancy in the human β-globin
locus. Biochim Biophys Acta. 1849:282–289. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Welch JJ, Watts JA, Vakoc CR, Yao Y, Wang
H, Hardison RC, Blobel GA, Chodosh LA and Weiss MJ: Global
regulation of erythroid gene expression by transcription factor
GATA-1. Blood. 104:3136–3147. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Cheng Y, Wu W, Kumar SA, Yu D, Deng W,
Tripic T, King DC, Chen KB, Zhang Y, Drautz D, et al: Erythroid
GATA1 function revealed by genome-wide analysis of transcription
factor occupancy, histone modifications, and mRNA expression.
Genome Res. 19:2172–2184. 2009. View Article : Google Scholar : PubMed/NCBI
|