|
1
|
Ozturk S, Papageorgis P, Wong CK, Lambert
AW, Abdolmaleky HM, Thiagalingam A, Cohen HT and Thiagalingam S:
SDPR functions as a metastasis suppressor in breast cancer by
promoting apoptosis. Proc Natl Acad Sci USA. 113:638–643. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Tian Y, Yu Y, Hou LK, Chi JR, Mao JF, Xia
L, Wang X, Wang P and Cao XC: Serum deprivation response inhibits
breast cancer progression by blocking transforming growth factor-β
signaling. Cancer Sci. 107:274–280. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Poma AM, Giannini R, Piaggi P, Ugolini C,
Materazzi G, Miccoli P, Vitti P and Basolo F: A six-gene panel to
label follicular adenoma, low- and high-risk follicular thyroid
carcinoma. Endocr Connect. 7:124–132. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Codenotti S, Vezzoli M, Poliani PL,
Cominelli M, Monti E and Fanzani A: Cavin-2 is a specific marker
for detection of well-differentiated liposarcoma. Biochem Biophys
Res Commun. 493:660–665. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Unozawa M, Kasamatsu A, Higo M, Fukumoto
C, Koyama T, Sakazume T, Nakashima D, Ogawara K, Yokoe H, Shiiba M,
et al: Cavin-2 in oral cancer: A potential predictor for tumor
progression. Mol Carcinog. 55:1037–1047. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hansen CG, Bright NA, Howard G and Nichols
BJ: SDPR induces membrane curvature and functions in the formation
of caveolae. Nat Cell Biol. 11:807–814. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Nassar ZD and Parat MO: Cavin family: New
players in the biology of caveolae. Int Rev Cell Mol Biol.
320:235–305. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gustincich S, Vatta P, Goruppi S, Wolf M,
Saccone S, Della Valle G, Baggiolini M and Schneider C: The human
serum deprivation response gene (SDPR) maps to 2q32-q33 and codes
for a phosphatidylserine-binding protein. Genomics. 57:120–129.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Baig A, Bao X, Wolf M and Haslam RJ: The
platelet protein kinase C substrate pleckstrin binds directly to
SDPR protein. Platelets. 20:446–457. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Gupta R, Toufaily C and Annabi B: Caveolin
and cavin family members: Dual roles in cancer. Biochimie.
107:188–202. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li X, Jia Z, Shen Y, Ichikawa H, Jarvik J,
Nagele RG and Goldberg GS: Coordinate suppression of Sdpr and Fhl1
expression in tumors of the breast, kidney, and prostate. Cancer
Sci. 99:1326–1333. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Curtis C, Shah SP, Chin SF, Turashvili G,
Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, et
al: The genomic and transcriptomic architecture of 2,000 breast
tumours reveals novel subgroups. Nature. 486:346–352. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Bai L, Deng X, Li Q, Wang M, An W, Deli A,
Gao Z, Xie Y, Dai Y and Cong YS: Down-regulation of the cavin
family proteins in breast cancer. J Cell Biochem. 113:322–328.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gianazza E, Chinello C, Mainini V,
Cazzaniga M, Squeo V, Albo G, Signorini S, Di Pierro SS, Ferrero S,
Nicolardi S, et al: Alterations of the serum peptidome in renal
cell carcinoma discriminating benign and malignant kidney tumors. J
Proteomics. 76:125–140. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Altintas DM, Allioli N, Decaussin M, de
Bernard S, Ruffion A, Samarut J and Vlaeminck-Guillem V:
Differentially expressed androgen-regulated genes in
androgen-sensitive tissues reveal potential biomarkers of early
prostate cancer. PLoS One. 8:e662782013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Vogelstein B, Papadopoulos N, Velculescu
VE, Zhou S, Diaz LA Jr and Kinzler KW: Cancer genome landscapes.
Science. 339:1546–1558. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
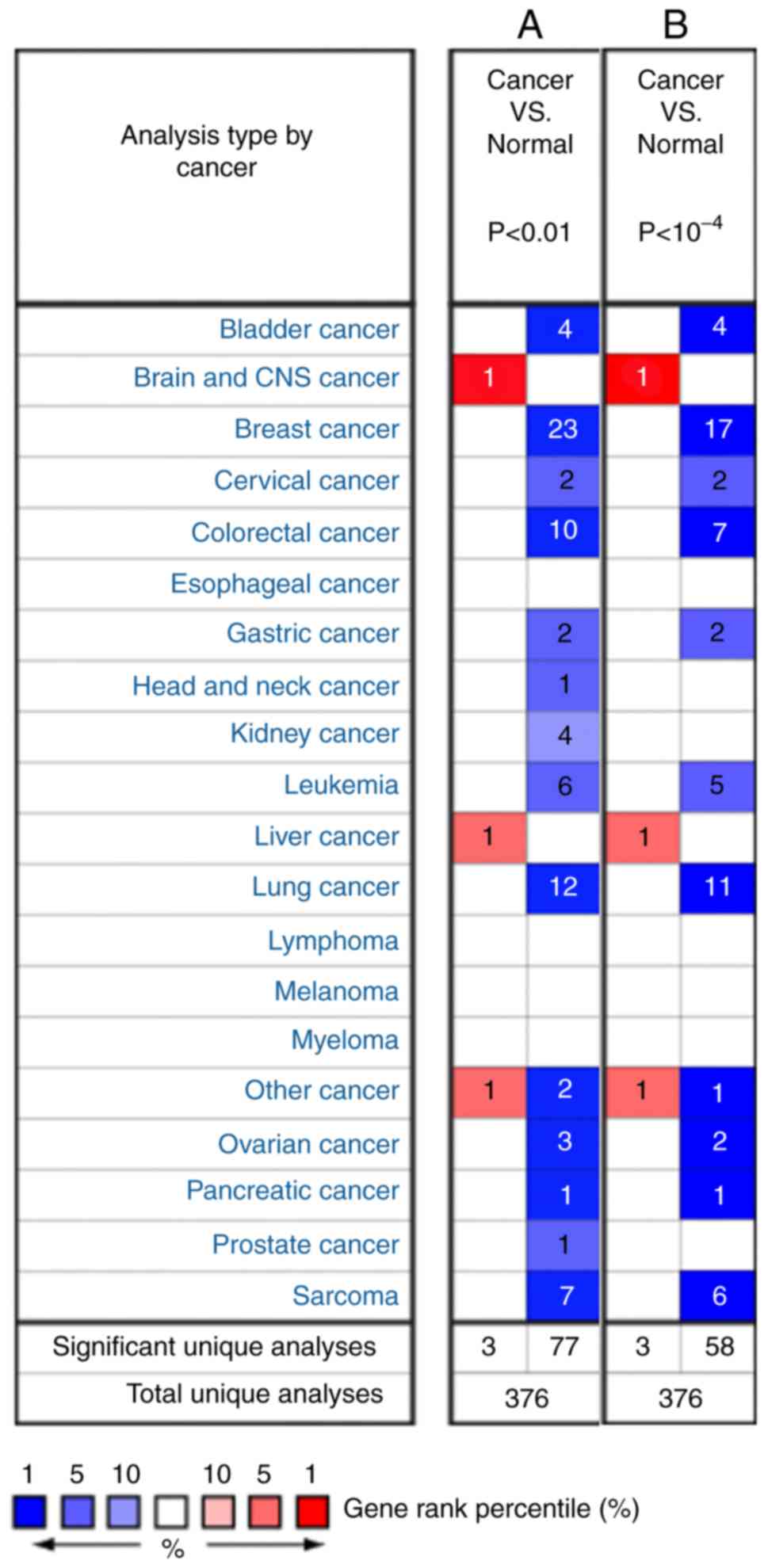
Rhodes DR, Kalyana-Sundaram S, Mahavisno
V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ,
Kincead-Beal C, Kulkarni P, et al: Oncomine 3.0: Genes, pathways,
and networks in a collection of 18,000 cancer gene expression
profiles. Neoplasia. 9:166–180. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Benjamini Y and Hochberg Y: Controlling
the false discovery rate: A practical and powerful approach to
multiple testing. J R Statist Soc B. 57:289–300. 1995.
|
|
19
|
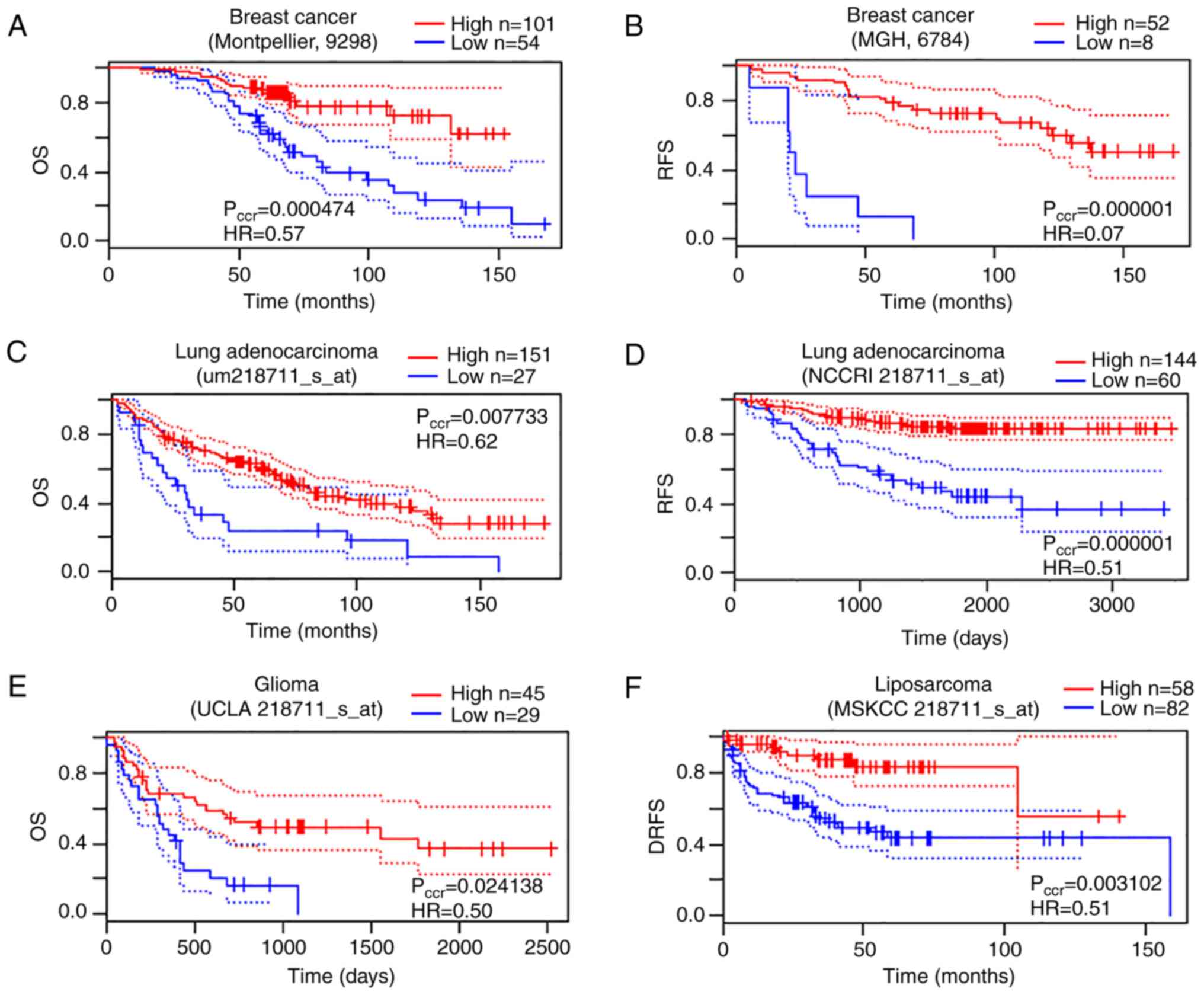
Mizuno H, Kitada K, Nakai K and Sarai A:
PrognoScan: A new database for meta-analysis of the prognostic
value of genes. BMC Med Genomics. 2:182009. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Miller R and Siegmund D: Maximally
selected chi square statistics. Biometrics. 38:1011–1016. 1982.
View Article : Google Scholar
|
|
21
|
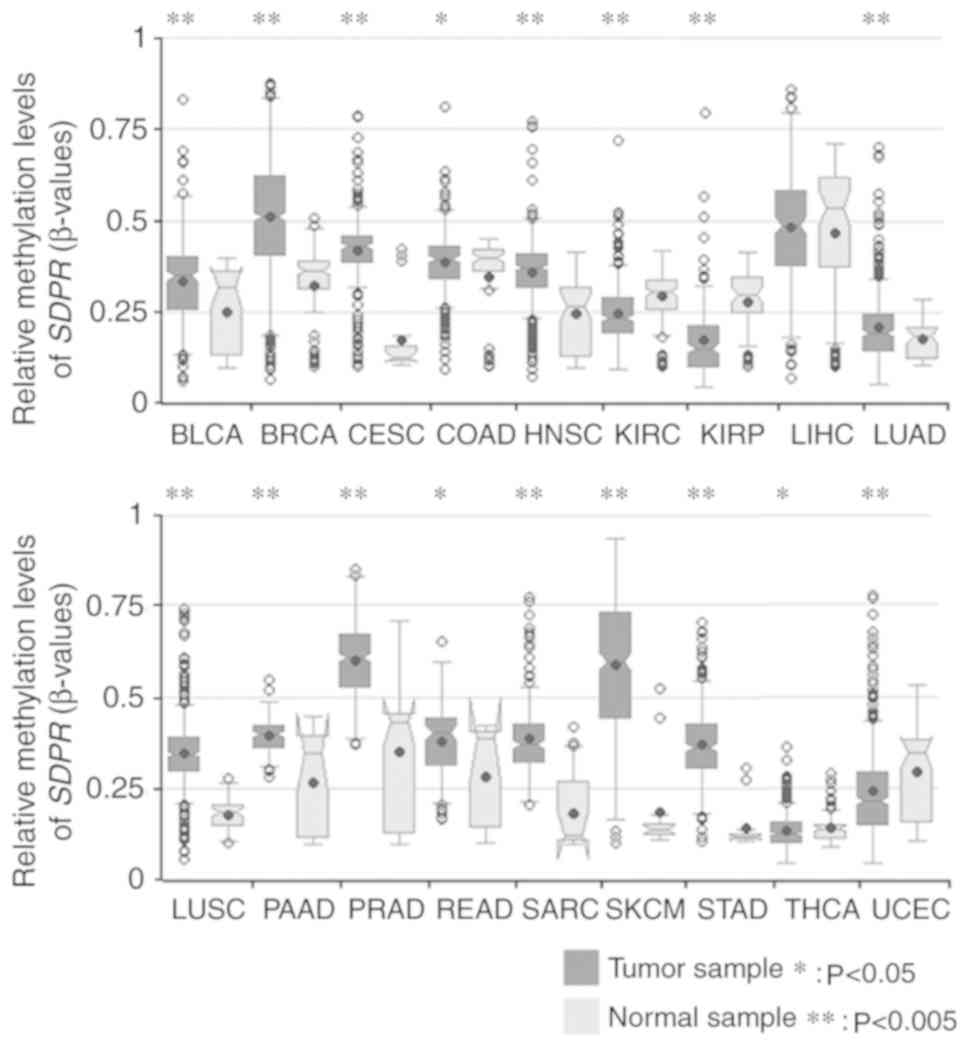
Huang WY, Hsu SD, Huang HY, Sun YM, Chou
CH, Weng SL and Huang HD: MethHC: A database of DNA methylation and
gene expression in human cancer. Nucleic Acids Res 43 (Database
Issue). D856–D861. 2015. View Article : Google Scholar
|
|
22
|
Broad Institute TCGA Genome Data Analysis
Center (2016), . Firehose stddata__2016_01_28 run. Broad Institute
of MIT and Harvard. Doi: 10.7908/C11G0KM9.
|
|
23
|
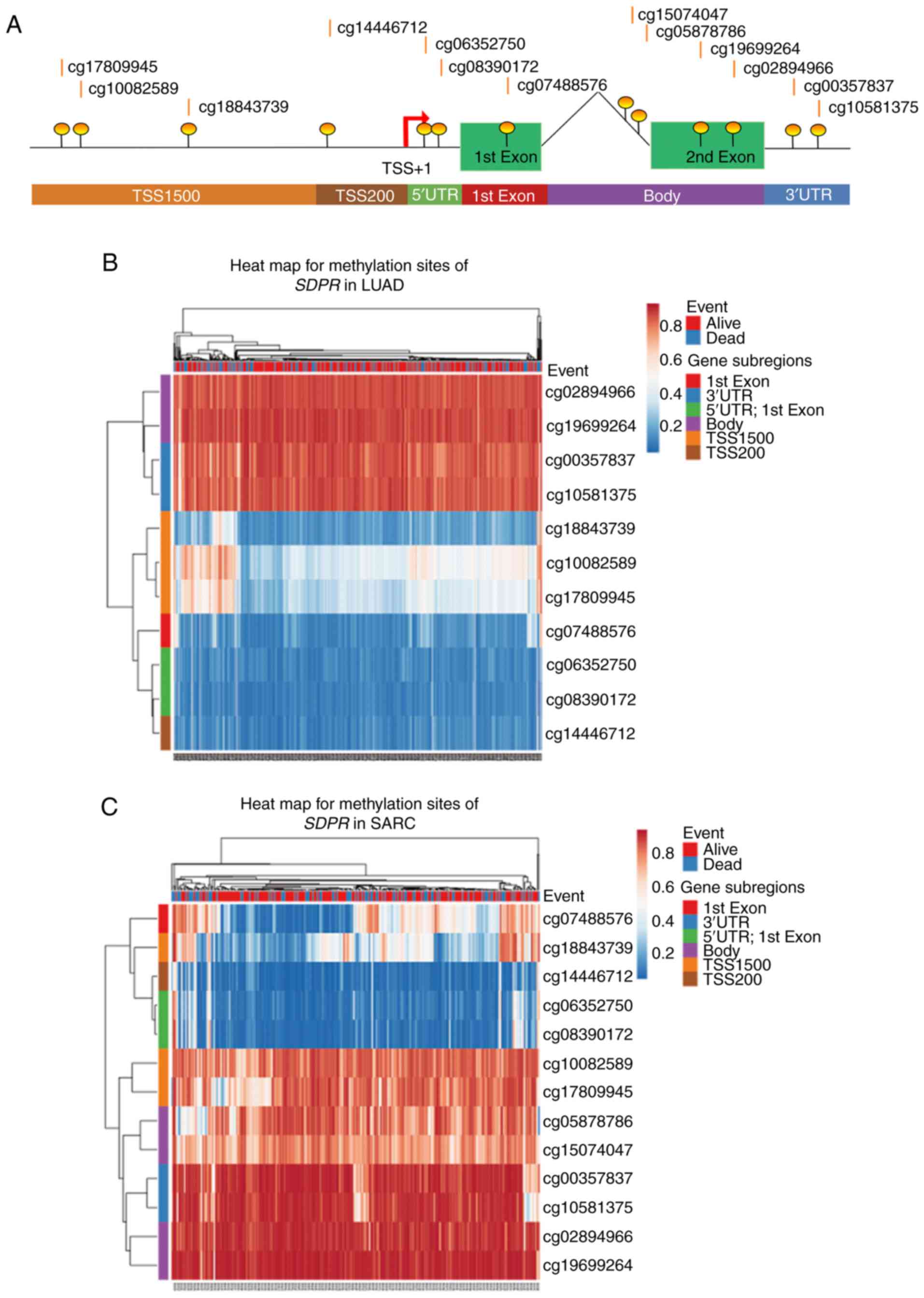
Modhukur V, Iljasenko T, Metsalu T, Lokk
K, Laisk-Podar T and Vilo J: MethSurv: A web tool to perform
multivariable survival analysis using DNA methylation data.
Epigenomics. 10:277–288. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Metsalu T and Vilo J: ClustVis: A web tool
for visualizing clustering of multivariate data using principal
component analysis and heatmap. Nucleic Acids Res 43W. W566–W570.
2015. View Article : Google Scholar
|
|
25
|
Farré D, Roset R, Huerta M, Adsuara JE,
Roselló L, Albà MM and Messeguer X: Identification of patterns in
biological sequences at the ALGGEN server: PROMO and MALGEN.
Nucleic Acids Res. 31:3651–3653. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Barretina J, Taylor BS, Banerji S, Ramos
AH, Lagos-Quintana M, Decarolis PL, Shah K, Socci ND, Weir BA, Ho
A, et al: Subtype-specific genomic alterations define new targets
for soft-tissue sarcoma therapy. Nat Genet. 42:715–721. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tessema M, Yingling CM, Snider AM, Do K,
Juri DE, Picchi MA, Zhang X, Liu Y, Leng S, Tellez CS and Belinsky
SA: GATA2 is epigenetically repressed in human and mouse lung
tumors and is not requisite for survival of KRAS mutant lung
cancer. J Thorac Oncol. 9:784–793. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Borup R, Rossing M, Henao R, Yamamoto Y,
Krogdahl A, Godballe C, Winther O, Kiss K, Christensen L, Høgdall
E, et al: Molecular signatures of thyroid follicular neoplasia.
Endocr Relat Cancer. 17:691–708. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Krendl C, Shaposhnikov D, Rishko V, Ori C,
Ziegenhain C, Sass S, Simon L, Müller NS, Straub T, Brooks KE, et
al: GATA2/3-TFAP2A/C transcription factor network couples human
pluripotent stem cell differentiation to trophectoderm with
repression of pluripotency. Proc Natl Acad Sci USA.
114:E9579–E9588. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
de Pater E, Kaimakis P, Vink CS, Yokomizo
T, Yamada-Inagawa T, van der Linden R, Kartalaei PS, Camper SA,
Speck N and Dzierzak E: Gata2 is required for HSC generation and
survival. J Exp Med. 210:2843–2850. 2013. View Article : Google Scholar
|
|
31
|
Kazenwadel J, Betterman KL, Chong CE,
Stokes PH, Lee YK, Secker GA, Agalarov Y, Demir CS, Lawrence DM,
Sutton DL, et al: GATA2 is required for lymphatic vessel valve
development and maintenance. J Clin Invest. 125:2979–2994. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Jones PA: Functions of DNA methylation:
Islands, start sites, gene bodies and beyond. Nat Rev Genet.
13:484–492. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ni W, Song E, Gong M, Li Y, Yao J and An
R: Downregulation of lncRNA SDPR-AS is associated with poor
prognosis in renal cell carcinoma. Onco Targets Ther. 10:3039–3047.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Sanchez-Carbayo M, Socci ND, Lozano J,
Saint F and Cordon-Cardo C: Defining molecular profiles of poor
outcome in patients with invasive bladder cancer using
oligonucleotide microarrays. J Clin Oncol. 24:778–789. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Kim WJ, Kim EJ, Kim SK, Kim YJ, Ha YS,
Jeong P, Kim MJ, Yun SJ, Lee KM, Moon SK, et al: Predictive value
of progression-related gene classifier in primary non-muscle
invasive bladder cancer. Mol Cancer. 9:32010. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Biewenga P, Buist MR, Moerland PD, Ver
Loren van Themaat E, van Kampen AH, ten Kate FJ and Baas F: Gene
expression in early stage cervical cancer. Gynecol Oncol.
108:520–526. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Scotto L, Narayan G, Nandula SV,
Arias-Pulido H, Subramaniyam S, Schneider A, Kaufmann AM, Wright
JD, Pothuri B, Mansukhani M and Murty VV: Identification of copy
number gain and overexpressed genes on chromosome arm 20q by an
integrative genomic approach in cervical cancer: Potential role in
progression. Genes Chromosomes Cancer. 47:755–765. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Skrzypczak M, Goryca K, Rubel T, Paziewska
A, Mikula M, Jarosz D, Pachlewski J, Oledzki J and Ostrowski J:
Modeling oncogenic signaling in colon tumors by multidirectional
analyses of microarray data directed for maximization of analytical
reliability. PLoS One. 5(pii): e130912010. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Ki DH, Jeung HC, Park CH, Kang SH, Lee GY,
Lee WS, Kim NK, Chung HC and Rha SY: Whole genome analysis for
liver metastasis gene signatures in colorectal cancer. Int J
Cancer. 121:2005–2012. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
D'Errico M, de Rinaldis E, Blasi MF, Viti
V, Falchetti M, Calcagnile A, Sera F, Saieva C, Ottini L, Palli D,
et al: Genome-wide expression profile of sporadic gastric cancers
with microsatellite instability. Eur J Cancer. 45:461–469. 2009.
View Article : Google Scholar
|
|
41
|
Cho JY, Lim JY, Cheong JH, Park YY, Yoon
SL, Kim SM, Kim SB, Kim H, Hong SW, Park YN, et al: Gene expression
signature-based prognostic risk score in gastric cancer. Clin
Cancer Res. 17:1850–1857. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Haferlach T, Kohlmann A, Wieczorek L,
Basso G, Kronnie GT, Béné MC, De Vos J, Hernández JM, Hofmann WK,
Mills KI, et al: Clinical utility of microarray-based gene
expression profiling in the diagnosis and subclassification of
leukemia: Report from the international microarray innovations in
leukemia study group. J Clin Oncol. 28:2529–2537. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Garber ME, Troyanskaya OG, Schluens K,
Petersen S, Thaesler Z, Pacyna-Gengelbach M, van de Rijn M, Rosen
GD, Perou CM, Whyte RI, et al: Diversity of gene expression in
adenocarcinoma of the lung. Proc Natl Acad Sci USA. 98:13784–13789.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Okayama H, Kohno T, Ishii Y, Shimada Y,
Shiraishi K, Iwakawa R, Furuta K, Tsuta K, Shibata T, Yamamoto S,
et al: Identification of genes upregulated in ALK-positive and
EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res.
72:100–111. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Selamat SA, Chung BS, Girard L, Zhang W,
Zhang Y, Campan M, Siegmund KD, Koss MN, Hagen JA, Lam WL, et al:
Genome-scale analysis of DNA methylation in lung adenocarcinoma and
integration with mRNA expression. Genome Res. 22:1197–1211. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Su LJ, Chang CW, Wu YC, Chen KC, Lin CJ,
Liang SC, Lin CH, Whang-Peng J, Hsu SL, Chen CH and Huang CY:
Selection of DDX5 as a novel internal control for Q-RT-PCR from
microarray data using a block bootstrap re-sampling scheme. BMC
Genomics. 8:1402007. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Landi MT, Dracheva T, Rotunno M, Figueroa
JD, Liu H, Dasgupta A, Mann FE, Fukuoka J, Hames M, Bergen AW, et
al: Gene expression signature of cigarette smoking and its role in
lung adenocarcinoma development and survival. PLoS One.
3:e16512008. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Hou J, Aerts J, den Hamer B, van Ijcken W,
den Bakker M, Riegman P, van der Leest C, van der Spek P, Foekens
JA, Hoogsteden HC, et al: Gene expression-based classification of
non-small cell lung carcinomas and survival prediction. PLoS One.
5:e103122010. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Wachi S, Yoneda K and Wu R:
Interactome-transcriptome analysis reveals the high centrality of
genes differentially expressed in lung cancer tissues.
Bioinformatics. 21:4205–4208. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Crabtree JS, Jelinsky SA, Harris HA, Choe
SE, Cotreau MM, Kimberland ML, Wilson E, Saraf KA, Liu W,
McCampbell AS, et al: Comparison of human and rat uterine
leiomyomata: Identification of a dysregulated mammalian target of
rapamycin pathway. Cancer Res. 69:6171–6178. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Yoshihara K, Tajima A, Komata D, Yamamoto
T, Kodama S, Fujiwara H, Suzuki M, Onishi Y, Hatae M, Sueyoshi K,
et al: Gene expression profiling of advanced-stage serous ovarian
cancers distinguishes novel subclasses and implicates ZEB2 in tumor
progression and prognosis. Cancer Sci. 100:1421–1428. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Pei H, Li L, Fridley BL, Jenkins GD,
Kalari KR, Lingle W, Petersen G, Lou Z and Wang L: FKBP51 affects
cancer cell response to chemotherapy by negatively regulating Akt.
Cancer Cell. 16:259–266. 2009. View Article : Google Scholar : PubMed/NCBI
|