Introduction
Head and neck squamous cell carcinoma (HNSCC) is the
sixth-most common malignancy worldwide, with 890,000 new cases and
450,000 deaths in 2018 (1,2). Despite progress in treatment strategies,
patient survival remains poor, with a 5-year survival rate of
<50% (3). Recurrence and
metastasis are frequent events leading to HNSCC mortality (4). Therefore, it is important to identify
novel biomarkers and understand the molecular mechanisms of HNSCC
progression.
Branched-chain amino acid (BCAA) transaminase 1
(BCAT1) is a BCAA aminotransferase enzyme that catalyzes
transamination of glutamate and BCAAs (5,6). Cytosolic
BCAT1 expression is restricted to a limited number of tissues,
including the brain, ovaries and testes (7). The involvement of BCAT1 in tumor
development has been implicated in glioblastoma (8), breast cancer (9) and leukemia (10,11). BCAT1
promotes the proliferation of cancer cells by regulating energy
metabolism (10). BCAT1 can promote
epithelial-mesenchymal transition in liver cancer (12). BCAT1 promotes the proliferation of
breast cancer cells by enhancing mTOR-mediated mitochondrial
biogenesis and function (9). These
reports thus suggested that BCAT1 is an onco-protein in human
cancers. At present, however, its expression pattern and biological
characteristics in human HNSCC remain unexplored.
In the present study, BCAT1 expression was examined
in HNSCC tissues via immunohistochemistry, and its relationship
with clinicopathological factors was analyzed. The effects of BCAT1
on cell proliferation, invasion, chemosensitivity, mitochondrial
function and glucose metabolism were evaluated, and the underlying
mechanism was explored via chromatin immunoprecipitation (ChIP).
The present findings provided novel evidence implicating BCAT1 as
an oncogene in HNSCC progression.
Materials and methods
Samples
The present study was approved by the Institutional
Review Board and Ethics Committee of the First Affiliated Hospital
of China Medical University (approval no. AF-SOP-07-1.0-01). A
total of 106 cases of HNSCC and 23 cases of normal head and neck
tissues were obtained from the Pathology Archives of the First
Affiliated Hospital of China Medical University (Shenyang, China),
which were collected between January 2012 and December 2016. HNSCC
samples included 83 male and 23 female patients. The age of the
patients ranged between 36–79 years, with an average age of 57.6
years. The normal control group enrolled 10 male and 13 female
patients, with an average age of 58.5 years (range, 41–77 years).
The patients agreed to the use of their samples in scientific
research and informed consent was obtained from the
participants.
Immunohistochemistry
Paraffin sections (4 µm) were deparaffinized using
xylene and a graded alcohol series. H2O2 (3%)
was applied to the slides to block peroxidase activity at 37°C for
15 min, and the sections were incubated with normal goat serum
(Fuzhou Maixin Biotech Co., Ltd.) at 37°C for 30 min to reduce
nonspecific binding. Then, the sections were incubated with BCAT1
antibody (1:500; cat. no. 28622-1-AP; ProteinTech Group, Inc.)
overnight at 4°C. Immunohistochemical staining was performed using
an Elivision Plus kit (Fuzhou Maixin Biotech Co., Ltd.). Briefly,
the sections were incubated with polymer enhancer at 37°C for 20
min and then with horseradish peroxidase (HRP)-conjugated
anti-mouse/rabbit IgG. Staining was developed with a DAB kit
(Fuzhou Maixin Biotech Co., Ltd.) at room temperature for 3 min.
Hematoxylin was used for counterstaining at room temperature for 5
min.
Immunostaining of BCAT1 was evaluated using a
semiquantitative scale by assessing both the intensity of staining
and the percentage of cells stained. Five fields were examined per
slide under a BX53 light microscope (magnification, ×400; Olympus
Corporation). BCAT1 intensity was scored as 0 (negative), 1 (weak)
or 2 (strong). The percentage of cells stained was scored as 1
(1–25%), 2 (26–50%), 3 (51–75%) or 4 (76–100%). The intensity and
percentage scores were multiplied to obtain a final score of 0–8.
Samples were considered to have low BCAT1 expression with final
scores <4. Samples with scores ≥4 were considered to have high
BCAT1 expression.
Bioinformatics analysis
BCAT1 gene expression profiles for patients with
HNSCC were obtained from The Cancer Genome Atlas (TCGA; http://tcga-data.nci.nih.gov/tcga/). Clinical
data including survival and clinical T stage were also downloaded
from TCGA (13). The expression of
BCAT1 in various types of HNSCC was identified in the Oncomine
database (https://www.oncomine.org/resource/login.html)
(14). Data were analyzed using
GraphPad Prism 7.00 software (GraphPad Software, Inc.).
Cell culture and transfection
The FaDu cell line was obtained from the Shanghai
Cell Bank of the Chinese Academy of Sciences. Cells were cultured
with RPMI-1640 medium (Gibco; Thermo Fisher Scientific, Inc.) with
10% FBS (Gibco; Thermo Fisher Scientific, Inc.) under 5%
CO2 at 37°C. pCMV6-BCAT1 plasmid (1 µg) and control
pCMV6 plasmid (OriGene Technologies, Inc.) were transfected into
FaDu cells (50% confluence) using Lipofectamine® 3000
reagent (Thermo Fisher Scientific, Inc.) for 72 h. BCAT1 small
interfering (si)RNA (target sequence, 5′-GUACAAAGGCGAGACAAUA-3′)
and non-targeting siRNA (target sequence,
5′-UAGCGACUAAACACAUCAA-3′; both GE Healthcare Dharmacon, Inc.) were
transfected using DharmaFECT1 reagent (GE Healthcare Dharmacon,
Inc.). The final siRNA concentration used for transfection was
0.025 µM.
Reverse transcription-quantitative
(RT-q)PCR
TRIzol reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) was used to extract RNA from cells. RT was
performed with an PrimeScript RT Master Mix kit (Takara Bio, Inc.)
at 85°C for 2 min and 37°C for 30 min. qPCR was performed using
SYBR Green Master Mix (Thermo Fisher Scientific, Inc.) on a 7500
Real-Time PCR System (Thermo Fisher Scientific, Inc.) at 50°C for 2
min, 95°C for 2 min, and 40 cycles of 95°C for 15 sec and 60°C for
40 sec. qPCR was quantified using the 2−ΔΔCq method
(15) and GAPDH was used as a
reference control. The following primers were used: BCAT1 forward,
5′-TGCTAGTCTGTATATTCGTCCT-3′ and reverse,
5′-CCAAGAGAAGGCTCAGTTCC-3′; and GAPDH forward,
5′-AACGACCACTTTGTCAAGCTC-3′ and reverse,
5′-AGCCAAATTCGTTGTCATACCAG-3′.
Cell Counting Kit-8 (CCK-8) and colony
formation assays
Cell proliferation was examined using CCK-8 reagent
(Dojindo Molecular Technologies, Inc.). Cells were seeded into
96-well plates at 3,000 cells/well and cultured at 37°C for 24, 48,
72 or 96 h. CCK-8 reagent (10 µl) was added to each well. After
incubation for 2 h, absorbance was examined at a wavelength of 450
nm using a microplate reader.
Colony formation assays were performed using 2,000
cells/culture plate. Cells were cultured at 37°C for 2 weeks and
then stained using Giemsa at room temperature for 10 min.
Western blotting
Total protein was extracted using RIPA lysis and
extraction buffer (Thermo Fisher Scientific, Inc.) and quantified
via the Bradford method. Proteins (50 µg/lane) were separated via
10% SDS-PAGE and transferred to PVDF membranes. The membranes were
blocked with 5% milk at room temperature for 1 h and then incubated
with antibodies against BCAT1 (1:1,500; cat. no. 13640-1-AP;
ProteinTech Group, Inc.), GLUT1 (1:1,000; cat. no. 12939; Cell
Signaling Technology, Inc.), c-Myc (1:1,000; cat. no. 5605; Cell
Signaling Technology, Inc.), and GAPDH (1:3,000; cat. no. 5174;
Cell Signaling Technology, Inc.). After incubation with HRP-coupled
secondary antibodies (1:3,000; cat. no. 7074; Cell Signaling
Technology, Inc.), protein bands were visualized using a Pierce™
Fast Western Blot kit ECL Substrate (Thermo Fisher Scientific,
Inc.). Images were captured using a DNR bio-imaging system (DNR
Bio-Imaging Systems, Ltd.).
Invasion assay
Transwell invasion assays were performed using
Transwell chambers (Corning, Inc.) coated with Matrigel (BD
Biosciences) at 37°C for 4 h. The treated cells were washed gently
with medium without serum, and then placed in the upper chamber at
a density of 1×104 cells/well. RPMI-1640 medium with 10%
FBS was added to the lower chamber. The Transwell chambers were
placed in an incubator at 37°C for 24 h. The cells which invaded to
the lower wells were fixed in paraformaldehyde at room temperature
for 20 min and then stained with hematoxylin for 5 min at room
temperature. The number of invading cells in random three fields
were counted under a BX53 microscope (magnification, ×200).
Apoptosis assay and JC-1 staining
FaDu cells were transfected with pCMV6-BCAT1, BCAT1
siRNA or corresponding negative controls. At 48 h later, the cells
were treated with 2 µM cisplatin (cat. no. P4394; Sigma-Aldrich;
Merck KGaA) at 37°C for 24 h. Then the cells were stained with
Annexin V-FITC/PI or JC-1 respectively. The rate of apoptosis was
determined using Annexin V-FITC/propidium iodide (PI) staining (BD
Biosciences). Cells were incubated with 5 µl FITC Annexin V and 5
µl PI for 15 min at room temperature in the dark. The apoptotic
rate was calculated as the percentage of early + late apoptotic
cells. JC-1 staining was used to determine the mitochondrial
membrane potential (Δψm). The cells were harvested, washed with PBS
and incubated with 20 µM JC-1 (cat. no. ab113850; Abcam) for 10 min
at room temperature. The cells were analyzed using an ACEA flow
cytometer (ACEA Bioscience, Inc.; Agilent Technologies, Inc.). Data
were analyzed using NovoExpress version 1.2.4 software (ACEA
Bioscience, Inc.; Agilent Technologies, Inc.).
Glucose uptake assay
A 2-NBDG uptake kit (cat. no. K682; BioVision, Inc.)
was used to examine the rate of glucose uptake according to the
manufacturer's protocols. Glucose uptake were analyzed using flow
cytometry at an excitation wavelength of 488 nm. The data were
analyzed using NovoExpress.
Glucose consumption assay
Glucose assay kits were purchased from Abcam (cat.
no. ab65333). At 72 h after transfection, the culture medium was
collected and centrifuged at 10,000 × g for 10 min at 4°C. The
supernatant was collected and glucose levels in the medium were
determined using glucose assay kits according to the manufacturer's
instructions.
ATP production assay
An ATP assay kit (cat. no. ab83355; Abcam) was used
to measure the rate of ATP production. Assays were performed
according to the manufacturer's protocols. Briefly, cells were
lysed, centrifuged at 13,000 × g at 4°C for 5 min, and filtered
using spin columns. Treated samples (5 µl) were mixed with ATP
reaction mix in 96-well plates. The mixture was measured at
excitation/emission wavelengths of 535/587 nm.
ChIP
ChIP assays were performed using a Magna ChIP G
Assay kit (EMD Millipore) according to the manufacturer's
instructions. Briefly, cells were crosslinked, pelleted and
resuspended in lysis buffer, then sonicated. The supernatant was
incubated overnight at 4°C with c-Myc (1:50; cat. no. 9402; Cell
Signaling Technology, Inc.) or IgG antibody (1:50; cat. no. 2729;
Cell Signaling Technology, Inc.) and Protein G-magnetic beads. The
beads were washed, and the chromatin complexes were collected,
purified and de-crosslinked. The DNA fragments were quantified via
qPCR analysis. Binding sites for c-Myc in the GLUT1 promoter were
predicted using TRANSFAC 7.0 (http://gene-regulation.com/pub/databases.html). The
primers for ChIP were as follows: GLUT1 position 1 forward,
5′-GTGGAGAGACTTTGAGGAGGG-3′ and reverse,
5′-CACTTGTCCTCTGTCTGCCTT-3′; and GLUT1 position 2 forward,
5′-GTCTCAAGTAAGGCACTGGTC-3′ and reverse,
5′-TTCCCACACCAATCTCATTGCT-3′.
Statistical analysis
SPSS 17.0 (SPSS, Inc.) was used for statistical
analysis. All experiments were repeated three times. The data were
presented as the mean ± standard deviation. The associations
between BCAT1 expression and clinical parameters were analyzed
using χ2 test or Fisher's exact test. Kaplan-Meier
analysis with log-rank test was used to estimate the overall
survival (OS) of patients with HNSCC. Mann-Whitney U tests were
used for the comparison of BCAT1 mRNA levels between cancerous and
normal tissues. Kruskal-Wallis test with Dunn's multiple comparison
post hoc test was used to evaluate the relationship between
clinical T stage and BCAT1 expression. For comparison of GLUT1 mRNA
data among multiple groups, ANOVA analysis with Tukey's multiple
comparisons post hoc test was performed. Other data were compared
using Student's t-test. P<0.05 was considered to indicate a
statistically significant difference.
Results
BCAT1 is upregulated in HNSCC
BCAT1 protein levels were examined in 106 cases of
HNSCC and 23 normal tissues using immunohistochemistry. BCAT1
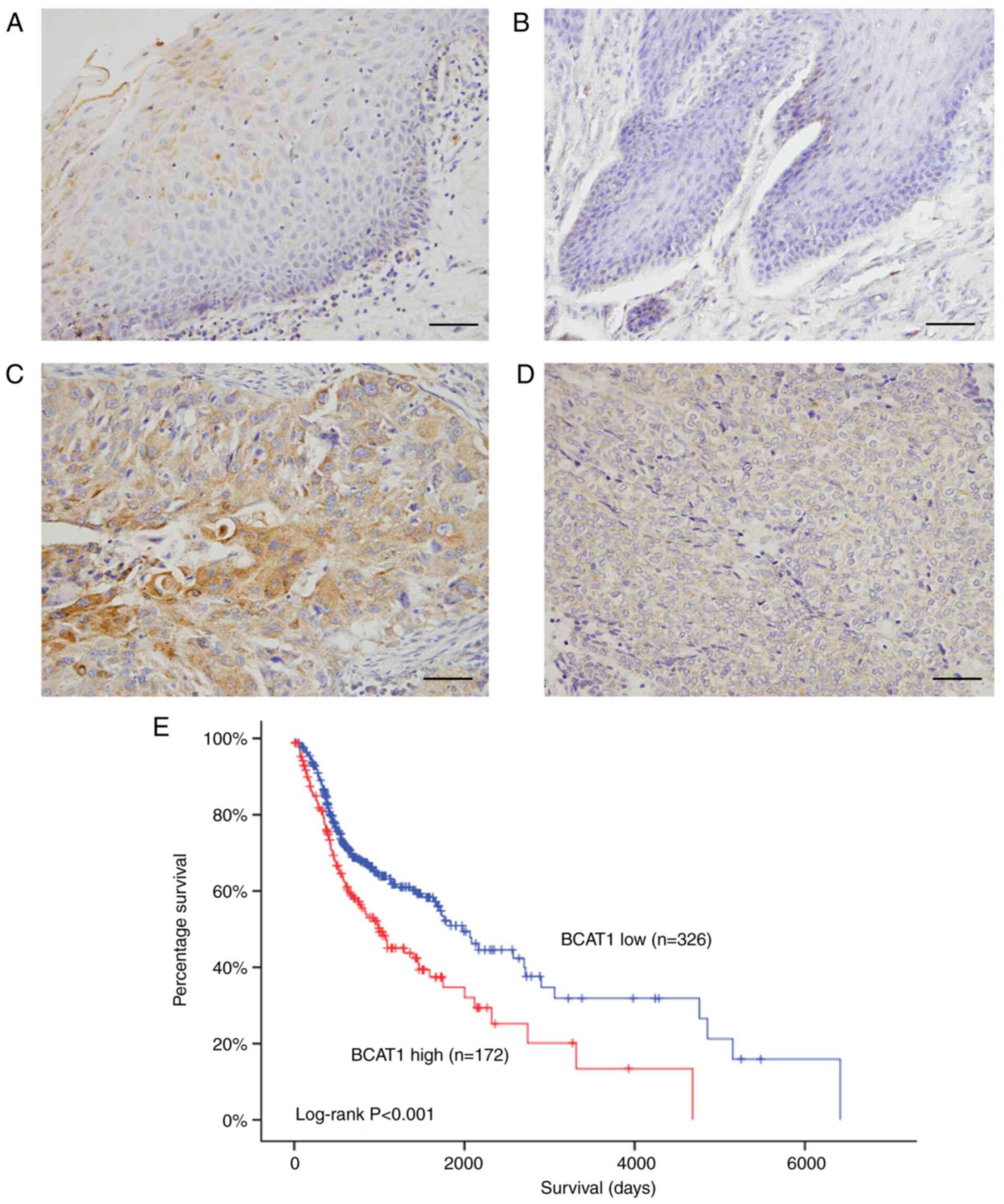
protein was detected in normal throat tissues (Fig. 1A) and oral mucosa (Fig. 1B). BCAT1 showed negative/low staining
in most normal cells. In contrast, BCAT1 protein expression was
elevated in tumor tissues (Fig. 1C and
D). In 106 cases examined, 52.8% (56/106) exhibited high BCAT1
expression. As presented in Table I,
high BCAT1 expression was positively associated with
tumor-node-metastasis (TNM) stage, lymph nodal metastasis and
higher T stage. In addition, a cohort of patients with HNSCC frin
TCGA showed that high BCAT1 expression was associated with shorter
overall survival (Fig. 1E). TCGA data
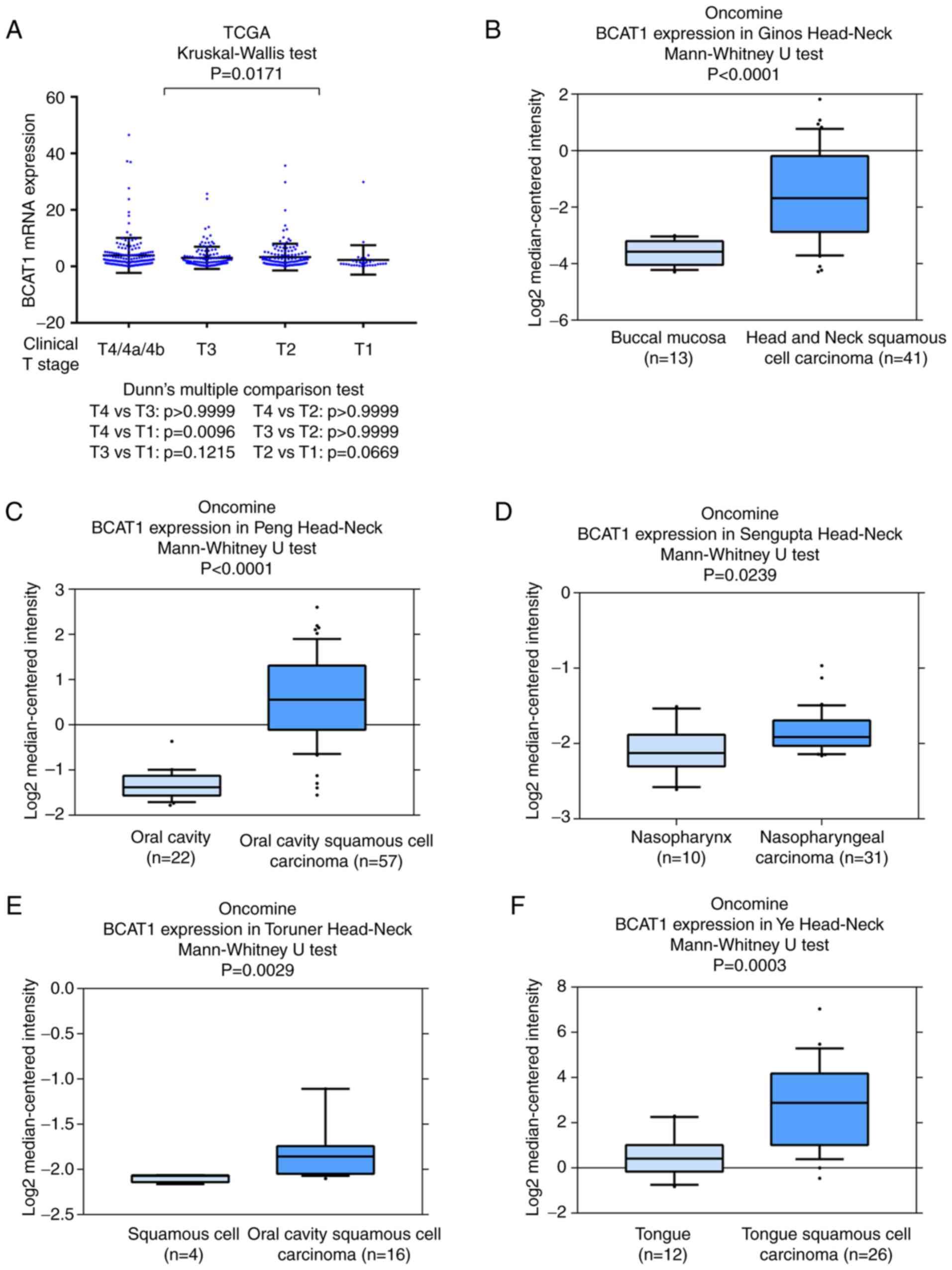
also indicated that BCAT1 mRNA was higher in HNSCC cases with a
higher clinical T stage (Fig. 2A).
Data from the Oncomine database were also analyzed. The Ginos,
Peng, Sengupta, Toruner and Ye head-neck datasets in Oncomine
indicated that BCAT1 mRNA was significantly elevated in cancer
tissues compared with normal tissues (Fig. 2B-F). These data indicated that BCAT1
expression was upregulated in HNSCC and associated with malignant
clinical features.
 | Table I.Association between
clinicopathological features and BCAT1 expression in head and neck
squamous cell carcinoma. |
Table I.
Association between
clinicopathological features and BCAT1 expression in head and neck
squamous cell carcinoma.
|
Characteristics | Number of
patients | Low BCAT1
expression | High BCAT1
expression | P-value |
|---|
| Age, years |
|
|
| 0.2677 |
|
<60 | 59 | 25 | 34 |
|
|
≥60 | 47 | 25 | 22 |
|
| Sex |
|
|
| 0.3101 |
|
Male | 83 | 37 | 46 |
|
|
Female | 23 | 13 | 10 |
|
| Node
metastasis |
|
|
| 0.0132 |
|
Absent | 77 | 42 | 35 |
|
|
Present | 29 | 8 | 21 |
|
| Tumor stage |
|
|
| 0.0207 |
| T1 +
T2 | 87 | 46 | 41 |
|
| T3 +
T4 | 19 | 4 | 15 |
|
| TNM stage |
|
|
| 0.021 |
| I +
II | 64 | 39 | 25 |
|
| III +
IV | 42 | 11 | 31 |
|
|
Differentiation |
|
|
| 0.1546 |
|
Well | 58 | 31 | 27 |
|
|
Moderate/poor | 48 | 19 | 29 |
|
BCAT1 promotes the proliferation and
invasion of FaDu cells
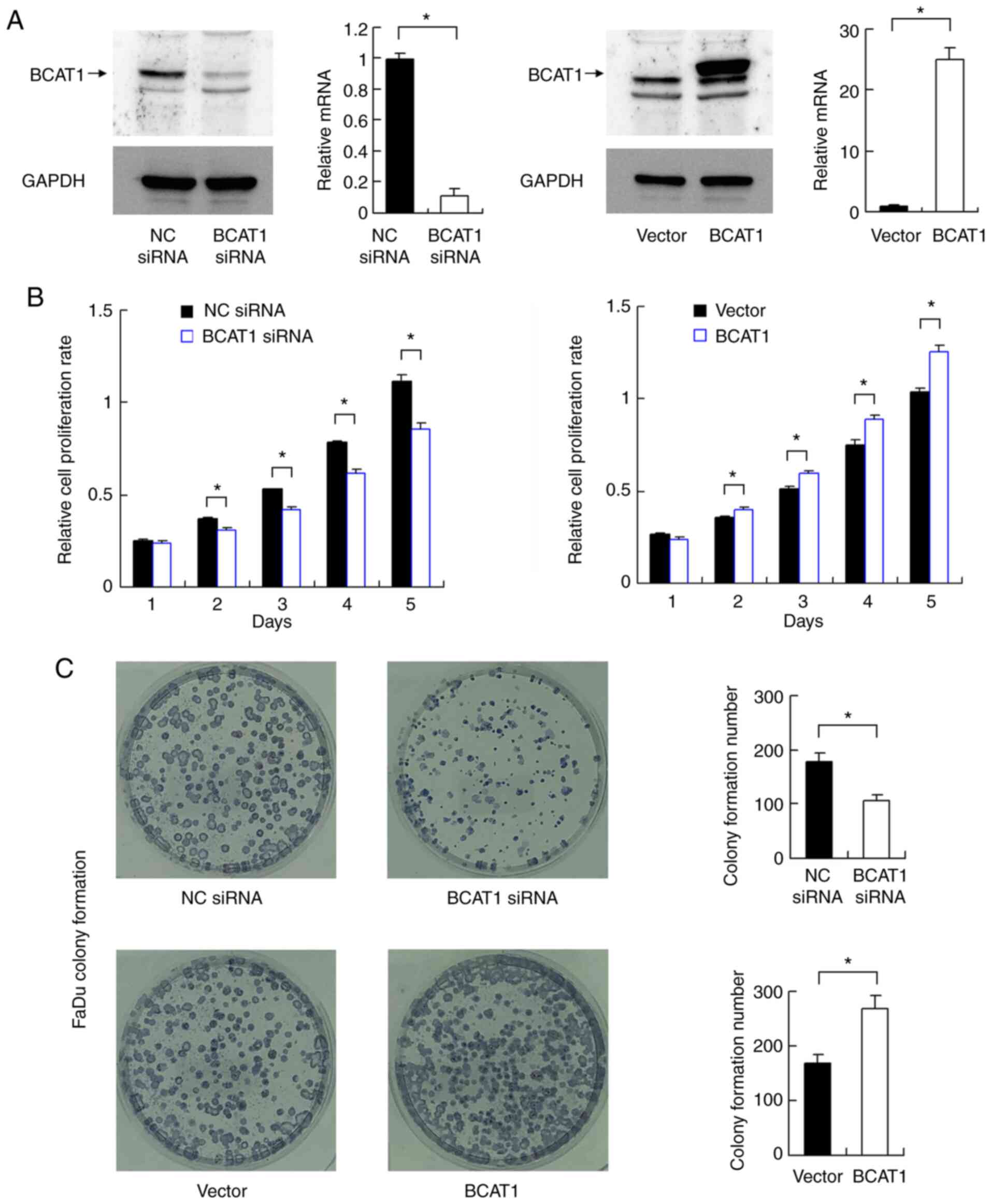
BCAT1 knockdown and overexpression were performed in
FaDu cells. The transfection efficiencies of BCAT1 siRNA and
overexpression vector were validated via western blotting and
RT-qPCR (Fig. 3A). CCK-8 assays
demonstrated that BCAT1 knockdown downregulated the proliferation
rate of FaDu cells, while BCAT1 overexpression increased the
cellular proliferation rate (Fig.
3B). Colony formation assays demonstrated that BCAT1 knockdown
decreased colony formation ability, while BCAT1 overexpression
upregulated colony formation ability (Fig. 3C).
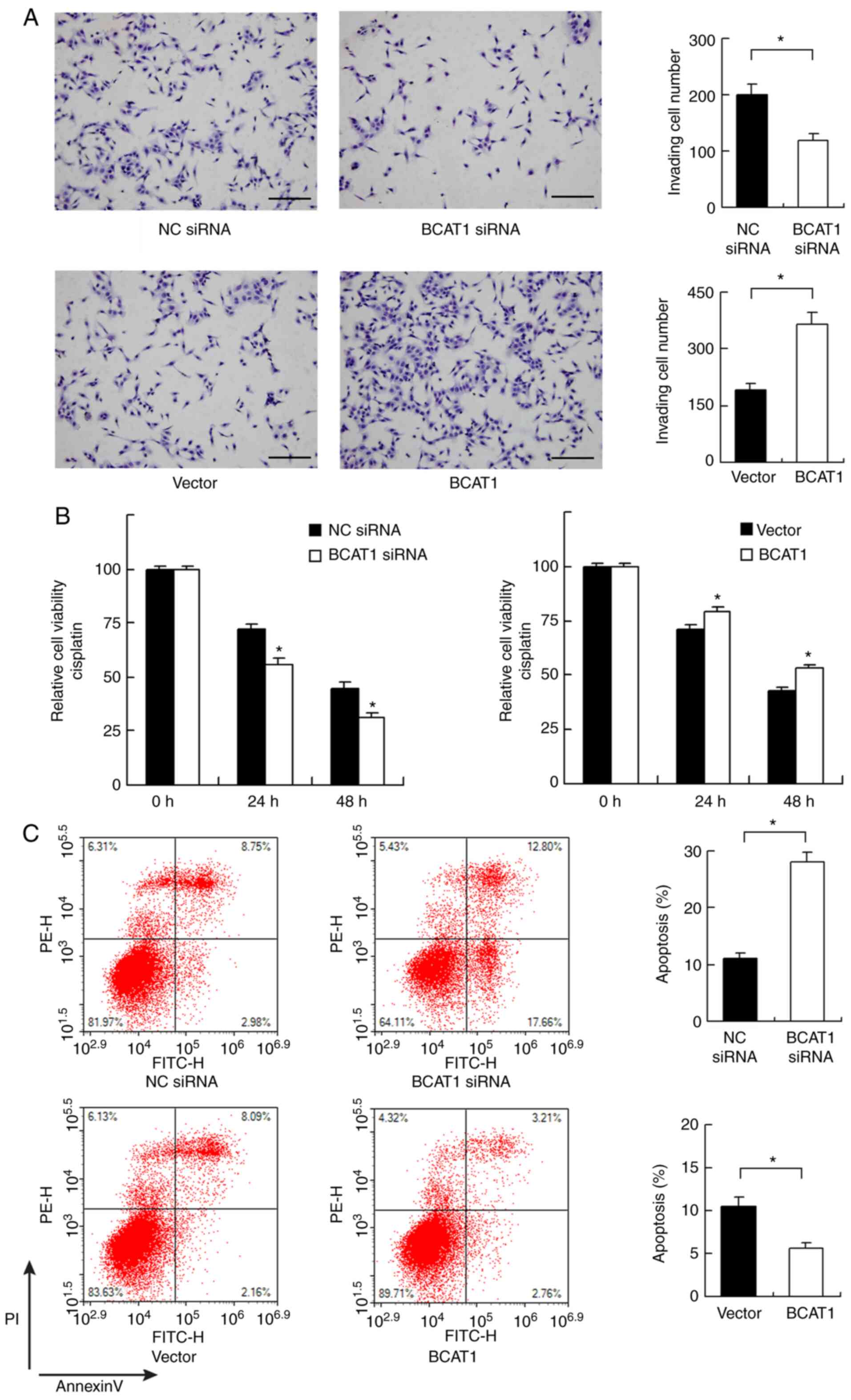
As there was a significant association between high
BCAT1 levels and nodal metastasis in clinical specimens, the
invasive ability of FaDu cells was determined after BCAT1
overexpression and knockdown. BCAT1 depletion inhibited invasion,
while BCAT1 overexpression increased the invasive ability of FaDu
cells (Fig. 4A).
BCAT1 regulates chemosensitivity and
mitochondrial function
The role of BCAT1 in chemosensitivity was also
investigated. Cisplatin (2 µM) was used to treat FaDu cells after
BCAT1 siRNA or plasmid transfection. CCK-8 assays showed that BCAT1
knockdown increased cisplatin-mediated inhibition of cells after 24
and 48 h of treatment, whereas BCAT1 overexpression induced
opposing effects (Fig. 4B). Annexin
V/PI staining showed that BCAT1 knockdown upregulated the levels of
cisplatin-induced apoptosis after 24 h of cisplatin treatment.
BCAT1 overexpression reduced the apoptosis rate, suggesting that
BCAT1 could induce resistance to chemotherapeutic drugs (Fig. 4C).
Chemotherapeutic drugs such as cisplatin can reduce
the mitochondrial membrane potential, which plays a pivotal role in
cell survival, as its loss induces apoptosis (16). FaDu cells were transfected with
pCMV6-BCAT1, BCAT1 siRNA or corresponding negative controls, and
were treated with subsequently treated with cisplatin. Then, JC-1
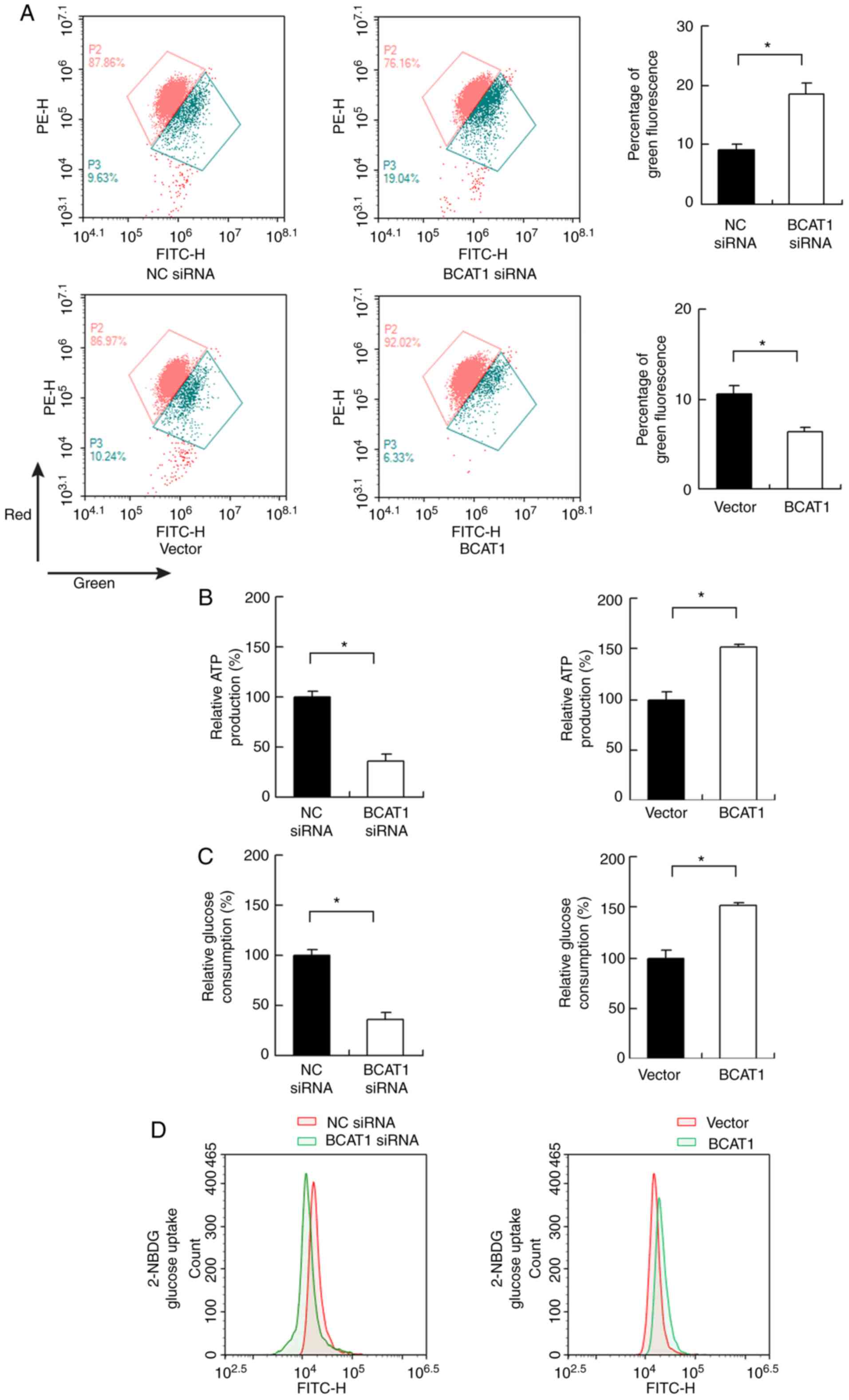
staining was performed to determine the change in Δψm. JC-1
exhibits red fluorescence in normal conditions but exhibits green
fluorescence when Δψm is downregulated. BCAT1 siRNA increased the
percentage of green staining, while BCAT1 overexpression decreased
green staining, suggesting that BCAT1 plays an important role in
maintaining normal Δψm (Fig. 5A).
BCAT1 regulates ATP production and
glucose uptake
Cancer cells adopt glucose metabolism to produce
energy (ATP), which is essential for cancer cell survival and
proliferation. Changes in ATP production were measured in FaDu
cells, and it found that BCAT1 knockdown decreased ATP levels,
whereas BCAT1 overexpression upregulated ATP levels (Fig. 5B). Subsequently, changes in glucose
consumption and uptake were evaluated. As shown in Fig. 5C, BCAT1 siRNA reduced glucose
consumption, whereas BCAT1 overexpression increased it. 2-NBDG
uptake showed that BCAT1 knockdown inhibited the levels of glucose
uptake, while BCAT1 overexpression upregulated glucose uptake
(Fig. 5D).
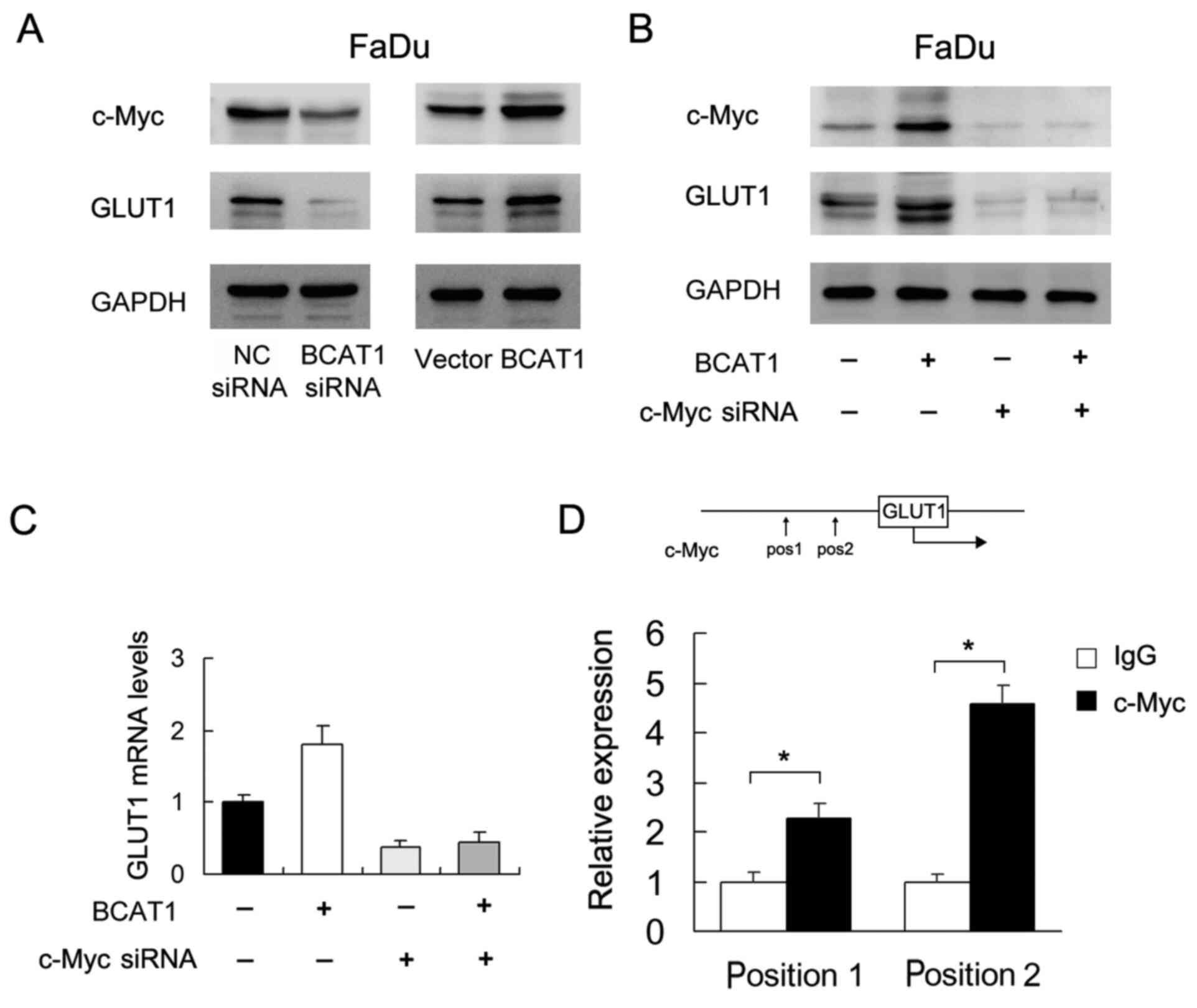
BCAT1 regulates c-Myc/GLUT1
signaling
To explore the mechanism underlying the role of
BCAT1 in HNSCC, several potentially associated proteins (including
GLUT1, GLUT2, GLUT3 and GLUT4) were screened. Of these, western
blotting showed that BCAT1 positively regulated GLUT1 protein
expression (Fig. 6A). It was also
found that BCAT1 siRNA downregulated c-Myc, which was upregulated
by BCAT1 overexpression. c-Myc has been reported as a
transcriptional regulator of glucose metabolism (17). To validate the association between
c-Myc and BCAT1-induced GLUT1 upregulation, c-Myc siRNA was used in
FaDu cells co-transfected with the BCAT1 plasmid. c-Myc siRNA
suppressed the protein and mRNA expression of GLUT1 (Fig. 6B and C). c-Myc siRNA also attenuated
the effect of BCAT1 plasmid on GLUT1 expression at both the protein
and mRNA levels (Fig. 6B and C).
Furthermore, it was investigated as to whether c-Myc binds to the
GLUT1 promoter. Binding sites for c-Myc in the GLUT1 promoter were
predicted using TRANSFAC. Binding of c-Myc to GLUT1 promoter was
examined using ChIP assays followed by RT-qPCR (Fig. 6D). The results indicated that the
effect of BCAT1 on GLUT1 was at least partly dependent on
c-Myc.
Discussion
In the present study, it was demonstrated that BCAT1
was upregulated in human HNSCC and promoted cell proliferation,
invasion, cisplatin resistance and glucose uptake, potentially via
c-Myc/GLUT1 signaling. Previous studies have focused on the role of
BCAT1 in BCAA metabolism (11,18), but
its potential roles in other pathways have not been fully explored.
The present data highlighted the role of BCAT1 in c-Myc/GLUT1
signaling and HNSCC progression.
Recent studies have shown that BCAT1 acts as a
cancer-related protein (9–12,19). A
recent study showed that BCAT1 was overexpressed in human non-small
cell lung cancer (NSCLC) tissues and cell lines (20). To date, there has been no such study
regarding BCAT1 in human HNSCC, to the best of the authors'
knowledge. The present findings showed that BCAT1 upregulation was
associated with TNM stage and nodal status, which was supported by
analyses of data from TCGA and Oncomine. TCGA data also showed that
BCAT1 levels were associated with poor prognosis, suggesting the
potential use of BCAT1 as a clinical indicator of malignant HNSCC.
It was noted that BCAT1 overexpression was not associated with the
differentiation of HNSCC. Generally, poorly differentiated HNSCC is
biologically more aggressive and tends to metastasize to regional
lymph nodes (21). However, HNSCC is
a heterogeneous cancer with a variety of histological subtypes.
Differentiation is not generally considered to be a direct
indicator of malignancy, and nodal/distal metastasis and clinical
stage are more useful for predicting patient prognosis (22,23).
CCK-8 and colony formation assays demonstrated that
BCAT1 overexpression promoted in vitro cell proliferation.
Matrigel invasion assays showed that BCAT1 facilitated HNSCC cell
invasion. In addition, it was determined that BCAT1 overexpression
decreased cisplatin sensitivity and reduced cisplatin-induced
apoptosis. Mitochondrial function serves an important role in
regulating chemosensitivity (24). A
decrease in Δψm can induce mitochondrial permeability, which
accelerates cytochrome c release and triggers apoptosis
(25–29). It was revealed that BCAT1
overexpression preserved Δψm after cisplatin treatment, suggesting
its role as a protector of mitochondrial function.
Mitochondrial balance in tumor cells relies on
energy supply (30,31). Glucose uptake is an important step
during ATP production. It was demonstrated that BCAT1
overexpression increased glucose uptake and consumption, and
upregulated ATP production, revealing BCAT1 as a positive regulator
of glucose metabolism. It should be noted that while BCAA
metabolism may play a part in cancer growth, other signaling
pathways may also contribute to malignant features induced by
BCAT1. These data therefore provide evidence for novel roles of
BCAT1 in regulating glucose metabolism in HNSCC cells.
In accordance with the increased glucose uptake, it
was further demonstrated that BCAT1 upregulated GLUT1 expression.
GLUT family proteins, which mediate the transport of glucose across
membranes, were reported to be elevated in human cancers (28,32). It
has been reported that GLUT1 regulated the proliferation and
survival of HNSCC cells (33). It was
also found that BCAT1 upregulated c-Myc expression. c-Myc has been
reported to control genes regulating glucose metabolism (34). Using siRNA-mediated knockdown of
c-Myc, it was shown that c-Myc mediated BCAT1-induced GLUT1
upregulation, which was further supported by ChIP data showing that
c-Myc directly bound to the GLUT1 promoter in FaDu cells. A recent
study in NSCLC also indicated the regulatory role of BCAT1 on
c-Myc, potentially via regulation of Wnt signaling (20). BCAT1 has also been reported as a
target gene of c-Myc (35). Thus,
there may be a positive feedback loop between BCAT1 and c-Myc in
HNSCC. Collectively, the present findings revealed a link between
BCAT1, c-Myc and GLUT1.
In conclusion, the present study revealed biological
roles of BCAT1 in human HNSCC. The results showed that BCAT1
overexpression promoted proliferation, invasion, cisplatin
resistance and glucose uptake in HNSCC cells. Its oncogenic effects
may be associated with its interactions with c-Myc/GLUT1 signaling,
suggesting the therapeutic possibility of targeting BCAT1 in
HNSCC.
Acknowledgements
Not applicable.
Funding
The present study was funded by The Natural Science
Foundation of Liaoning (grant no. 20180540097).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author upon reasonable
request.
Authors' contributions
HW and FW performed the experiments and drafted the
manuscript. HW and WO collected and analyzed the data. XJ and YW
conceived, designed and supervised the study, and drafted the
manuscript. All authors read and approved the final manuscript and
agree to be responsible for all aspects of the work in ensuring
that the accuracy or integrity of any part of the work are
appropriately investigated and resolved.
Ethics approval and consent to
participate
The present study was approved by the Institute
Research Ethics Committee of the First Affiliated Hospital of China
Medical University and written informed consent was obtained from
all patients involved.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Chen W, Zheng R, Baade PD, Zhang S, Zeng
H, Bray F, Jemal A, Yu XQ and He J: Cancer statistics in China,
2015. CA Cancer J Clin. 66:115–132. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2015. CA Cancer J Clin. 65:5–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Leemans CR, Braakhuis BJ and Brakenhoff
RH: The molecular biology of head and neck cancer. Nat Rev Cancer.
11:9–22. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yang CX, Sedhom W, Song J and Lu SL: The
role of MicroRNAs in recurrence and metastasis of head and neck
squamous cell carcinoma. Cancers (Basel). 11:3952019. View Article : Google Scholar
|
|
5
|
Eden A and Benvenisty N: Involvement of
branched-chain amino acid aminotransferase (Bcat1/Eca39) in
apoptosis. FEBS Lett. 457:255–261. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ben-Yosef T, Eden A and Benvenisty N:
Characterization of murine BCAT genes: Bcat1, a c-Myc target, and
its homolog, Bcat2. Mamm Genome. 9:595–597. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Garcia-Espinosa MA, Wallin R, Hutson SM
and Sweatt AJ: Widespread neuronal expression of branched-chain
aminotransferase in the CNS: Implications for leucine/glutamate
metabolism and for signaling by amino acids. J Neurochem.
100:1458–1468. 2007.PubMed/NCBI
|
|
8
|
Tonjes M, Barbus S, Park YJ, Wang W,
Schlotter M, Lindroth AM, Pleier SV, Bai AHC, Karra D, Piro RM, et
al: BCAT1 promotes cell proliferation through amino acid catabolism
in gliomas carrying wild-type IDH1. Nat Med. 19:901–908. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhang L and Han J: Branched-chain amino
acid transaminase 1 (BCAT1) promotes the growth of breast cancer
cells through improving mTOR-mediated mitochondrial biogenesis and
function. Biochem Biophys Res Commun. 486:224–231. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Raffel S, Falcone M, Kneisel N, Hansson J,
Wang W, Lutz C, Bullinger L, Poschet G, Nonnenmacher Y, Barnert A,
et al: BCAT1 restricts αKG levels in AML stem cells leading to
IDHmut-like DNA hypermethylation. Nature. 551:384–388. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hattori A, Tsunoda M, Konuma T, Kobayashi
M, Nagy T, Glushka J, Tayyari F, McSkimming D, Kannan N, Tojo A, et
al: Cancer progression by reprogrammed BCAA metabolism in myeloid
leukaemia. Nature. 545:500–504. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Qi LN, Xiang BD, Wu FX, Ye JZ, Zhong JH,
Wang YY, Chen YY, Chen ZS, Ma L, Chen J, et al: Circulating tumor
cells undergoing EMT provide a metric for diagnosis and prognosis
of patients with hepatocellular carcinoma. Cancer Res.
78:4731–4744. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang Z, Jensen MA and Zenklusen JC: A
practical guide to the cancer genome atlas (TCGA). Methods Mol
Biol. 1418:111–141. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Rhodes DR, Kalyana-Sundaram S, Mahavisno
V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ,
Kincead-Beal C, Kulkarni P, et al: Oncomine 3.0: Genes, pathways,
and networks in a collection of 18,000 cancer gene expression
profiles. Neoplasia. 9:166–180. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Erxleben A: Mitochondria-targeting
anticancer metal complexes. Curr Med Chem. 26:694–728. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Marbaniang C and Kma L: Dysregulation of
glucose metabolism by oncogenes and tumor suppressors in cancer
cells. Asian Pac J Cancer Prev. 19:2377–2390. 2018.PubMed/NCBI
|
|
18
|
Gu Z, Liu Y, Cai F, Patrick M, Zmajkovic
J, Cao H, Zhang Y, Tasdogan A, Chen M, Qi L, et al: Loss of EZH2
reprograms BCAA metabolism to drive leukemic transformation. Cancer
Discov. 9:1228–1247. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Xu Y, Yu W, Yang T, Zhang M, Liang C, Cai
X and Shao Q: Overexpression of BCAT1 is a prognostic marker in
gastric cancer. Hum Pathol. 75:41–46. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lin X, Tan S, Fu L and Dong Q: BCAT1
overexpression promotes proliferation, invasion, and wnt signaling
in non-small cell lung cancers. Onco Targets Therapy. 13:3583–3594.
2020. View Article : Google Scholar
|
|
21
|
Fortin A, Couture C, Doucet R, Albert M,
Allard J and Tetu B: Does histologic grade have a role in the
management of head and neck cancers? J Clin Oncol. 19:4107–4116.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Economopoulou P, de Bree R, Kotsantis I
and Psyrri A: Diagnostic tumor markers in head and neck squamous
cell carcinoma (HNSCC) in the clinical setting. Front Oncol.
9:8272019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Bossi P, Alfieri S, Strojan P, Takes RP,
López F, Mäkitie A, Saba NF, Rodrigo JP, Bradford C, Suarez C, et
al: Prognostic and predictive factors in recurrent and/or
metastatic head and neck squamous cell carcinoma: A review of the
literature. Crit Rev Oncol Hematol. 137:84–91. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Fu L, Dong Q, He J, Wang X, Xing J, Wang
E, Qiu X and Li Q: SIRT4 inhibits malignancy progression of NSCLCs,
through mitochondrial dynamics mediated by the ERK-Drp1 pathway.
Oncogene. 36:2724–2736. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Guerra F, Arbini AA and Moro L:
Mitochondria and cancer chemoresistance. Biochim Biophys Acta
Bioenerg. 1858:686–699. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Kim JS, Lee JM, Chwae YJ, Kim YH, Lee JH,
Kim K, Lee TH, Kim SJ and Park JH: Cisplatin-induced apoptosis in
Hep3B cells: Mitochondria-dependent and -independent pathways.
Biochem Pharmacol. 67:1459–1468. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhao W, You CC, Zhuang JP, Zu JN, Chi ZY,
Xu GP and Yan JL: Viability inhibition effect of gambogic acid
combined with cisplatin on osteosarcoma cells via
mitochondria-independent apoptotic pathway. Mol Cell Biochem.
382:243–252. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Li H, Fu L, Liu B, Lin X, Dong Q and Wang
E: Ajuba overexpression regulates mitochondrial potential and
glucose uptake through YAP/Bcl-xL/GLUT1 in human gastric cancer.
Gene. 693:16–24. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Battogtokh G, Cho YY, Lee JY, Lee HS and
Kang HC: Mitochondrial-Targeting anticancer agent conjugates and
nanocarrier systems for cancer treatment. Front Pharmacol.
9:9222018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Horbay R and Bilyy R: Mitochondrial
dynamics during cell cycling. Apoptosis. 21:1327–1335. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Stockburger C, Miano D, Pallas T,
Friedland K and Muller WE: Enhanced neuroplasticity by the
metabolic enhancer piracetam associated with improved mitochondrial
dynamics and altered permeability transition pore function. Neural
Plast. 2016:80759032016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zambrano A, Molt M, Uribe E and Salas M:
Glut 1 in cancer cells and the inhibitory action of resveratrol as
a potential therapeutic strategy. Int J Mol Sci. 20:33742019.
View Article : Google Scholar
|
|
33
|
Li S, Yang X, Wang P and Ran X: The
effects of GLUT1 on the survival of head and neck squamous cell
carcinoma. Cell Physiol Biochem. 32:624–634. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Osthus RC, Shim H, Kim S, Li Q, Reddy R,
Mukherjee M, Xu Y, Wonsey D, Lee LA and Dang CV: Deregulation of
glucose transporter 1 and glycolytic gene expression by c-Myc. J
Biol Chem. 275:21797–21800. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhou W, Feng X, Ren C, Jiang X, Liu W,
Huang W, Liu Z, Li Z, Zeng L, Wang L, et al: Over-expression of
BCAT1, a c-Myc target gene, induces cell proliferation, migration
and invasion in nasopharyngeal carcinoma. Mol Cancer. 12:532013.
View Article : Google Scholar : PubMed/NCBI
|